Data independent acquisition mass spectrometry (DIA-MS) in cancer studies: a concise review
Bharath S.
Kumar
 *
*
Department of Chemistry, Lehigh University, 6 E Packer Avenue, Bethlehem, PA 18015, USA. E-mail: bhs324@lehigh.edu
First published on 5th December 2025
Abstract
Advances in proteomics are reshaping our understanding of cancer biology by enabling the direct quantification of proteins and their modifications in complex biological systems. Among emerging mass spectrometry techniques, Data-Independent Acquisition (DIA) has established itself as a transformative approach for cancer proteomics. DIA offers unprecedented depth, reproducibility, and scalability by systematically fragmenting all precursor ions across defined mass ranges, allowing comprehensive proteome coverage and retrospective data analysis. This review highlights the fundamental principles of DIA-MS, recent technological developments—including spectral library-free workflows, and their impact on cancer research. We discuss the application of DIA in tumor classification, biomarker discovery, therapeutic target identification, and treatment response monitoring. Particular attention is given to its compatibility with clinical samples such as formalin-fixed paraffin-embedded (FFPE) tissues and its integration into large-scale efforts like CPTAC. Current challenges with the technique will be explored, including data analysis complexity and standardization, and future directions that could bring DIA-MS closer to clinical utility. DIA-MS is rapidly evolving into a cornerstone technology for precision oncology, with the potential to bridge research and clinical practice through robust, high-resolution proteomic profiling.
1 Introduction
Cancer remains one of the most complex and varied diseases because of a multitude of genetic, epigenetic, and proteomic alterations. While genomic profiling has provided significant insights into tumor classification and mutation landscapes, proteomics complements these efforts by directly measuring functional proteins and post-translational modifications (PTMs) that are critical in cancer pathogenesis and therapy response.1 However, traditional proteomic approaches, particularly data-dependent acquisition (DDA), often fall short in capturing the full spectrum of the cancer proteome, especially for low-abundance proteins and PTMs, due to limitations in sampling reproducibility and dynamic range.2Mass spectrometry (MS) has become an indispensable tool in proteomics, enabling the comprehensive analysis of complex protein mixtures. Traditional data-dependent acquisition (DDA) methods, while effective, often suffer from limitations such as stochastic sampling and limited reproducibility, particularly for low-abundance peptides.2 To address these challenges, Data-Independent Acquisition (DIA) has become a transformative approach in MS-based proteomics. DIA achieves comprehensive and reproducible proteomic analysis by fragmenting all precursor ions across defined m/z windows, producing a digital map of the proteome that can be mined retrospectively.3 In cancer research, this unbiased and systematic approach is especially valuable, given the need for high-throughput, sensitive, and reproducible methods to analyze heterogeneous tumor samples, longitudinal biopsies, and patient-derived material.
Current developments in DIA have significantly enhanced its applicability in cancer proteomics. Integration with technologies like parallel accumulation–serial fragmentation (PASEF) has improved peptide separation and increased identification rates, even in small or degraded clinical samples.4 Moreover, library-free DIA methods and deep learning-driven spectrum prediction tools (e.g., Prosit) now allow robust peptide identification without requiring experimental spectral libraries, making DIA more scalable and accessible for large cancer studies.5,6
DIA has been increasingly adopted in cancer biomarker discovery, patient stratification, and therapeutic response monitoring. For instance, the Clinical Proteomic Tumor Analysis Consortium (CPTAC) has used DIA-based workflows to profile thousands of tumor samples, identifying subtype-specific signatures in breast, lung, colorectal, and ovarian cancers.7,8 More recently, DIA has enabled the detection of actionable protein targets and resistance mechanisms in patient tumors that were not apparent from genomic analysis alone.9 In clinical applications, DIA's reproducibility and quantitative accuracy support its use in longitudinal and multi-center studies—critical for validating biomarkers and developing proteomic signatures for diagnosis, prognosis, and therapy selection. Importantly, DIA is well-suited for use with formalin-fixed paraffin-embedded (FFPE) tissues and other archival materials commonly available in oncology, overcoming one of the major practical barriers to clinical proteomics.10
The application of DIA has extended beyond traditional proteomics into clinical research, where its reproducibility and comprehensiveness of coverage are particularly beneficial. Studies have demonstrated the utility of DIA in biomarker discovery, disease classification, and therapeutic target identification across various diseases, including cancer and infectious diseases.9,11 The ability of DIA to generate permanent digital proteome maps allows for retrospective analysis, making it a valuable tool in longitudinal studies and large-scale clinical cohorts.
A pivotal yet sometimes underemphasized factor in the success of DIA-based workflows is the ionization source, which critically influences sensitivity, ionization efficiency, and reproducibility, especially when analyzing low-input or heterogeneous cancer samples. The most used ionization method in proteomics is electrospray ionization (ESI), which provides soft ionization compatible with liquid chromatography and high-throughput workflows. Advances in nano ESI and microflow ESI have improved ionization efficiency and stability, thereby enhancing peptide detectability and quantification, particularly important in clinical cancer proteomics where sample amounts are often limited.12
Ionization techniques such as atmospheric pressure photoionization (APPI) and matrix-assisted laser desorption/ionization (MALDI) have also been explored in combination with DIA for specialized cancer applications. For instance, MALDI-DIA imaging has shown promise in spatially resolved proteomics of tumors, providing both molecular content and localization, which is particularly relevant in solid tumor heterogeneity and microenvironment analysis.13 Meanwhile, high-pressure or high-field asymmetric waveform ion mobility spectrometry (FAIMS) integrated with ESI has been utilized to further improve signal-to-noise ratios and reduce interference, boosting the depth of proteome coverage in complex tumor tissues.14
In recent years, the combination of optimized ion sources and cutting-edge DIA methods has been instrumental in large-scale cancer studies. DIA-MS has been successfully applied to thousands of tumor samples by consortia such as CPTAC, revealing proteomic subtypes, actionable targets, and post-transcriptional regulation not evident from genomic data alone.8,9 The reproducibility of DIA and improved ionization methods has also enabled its deployment in longitudinal clinical trials, facilitating biomarker discovery and response monitoring with unprecedented precision.
Computational tools have also evolved to handle the complex data generated by DIA. Software platforms like DIA-NN, Spectronaut, and OpenSWATH have been instrumental in analyzing DIA data, offering both library-based and library-free approaches for peptide identification and quantification.6,15,16 Notably, library-free methods have gained prominence due to their ability to analyze data without the need for extensive spectral libraries, thus streamlining the workflow and reducing the reliance on prior experiments.3 Moreover, the integration of machine learning and deep learning algorithms into DIA workflows has enhanced the prediction of peptide fragmentation patterns and retention times, further improving the accuracy and efficiency of data analysis.5 These advancements have facilitated the move towards more standardized and high-throughput proteomic analyses, essential for the implementation of proteomics in clinical diagnostics and personalized medicine.
DIA-MS, empowered by innovations in ionization sources and acquisition strategies, has become a cornerstone in cancer proteomics. Its ability to produce high-quality, quantitative proteomic data from limited and clinically relevant samples makes it indispensable for precision oncology. As mass spectrometry instrumentation and ionization technologies continue to evolve, DIA-MS is poised to drive further breakthroughs in cancer diagnostics, therapeutics, and biomarker discovery.
2 Principles of DDA and DIA
MS has become a cornerstone technique in proteomics and metabolomics due to its ability to identify and quantify biomolecules with high sensitivity and precision. Among the various MS acquisition strategies, Data-Dependent Acquisition (DDA) and DIA represent two fundamentally different approaches to selecting and fragmenting precursor ions. Both methods have distinct principles, workflows, and analytical implications that influence the depth, reproducibility, and interpretability of proteomic datasets.DDA operates on a dynamic, intensity-driven model. In a typical DDA workflow, an initial MS1 scan captures all ions present in the sample. From this, the instrument automatically selects the most intense precursor ions—commonly the top 10 or 20, to be isolated and fragmented in the subsequent MS2 scan. This targeted fragmentation is governed by real-time ion intensity, with dynamic exclusion often applied to prevent repeated analysis of the same ion and to expand the sampling depth.2 The MS2 spectra obtained are typically clean and high-resolution, making them ideal for peptide identification through spectral library matching or database searching.
Conversely, DIA adopts a comprehensive and systematic approach. Instead of selectively targeting high-intensity ions, DIA divides the m/z range into sequential, overlapping windows (typically 10–30 Da wide) and fragments all precursor ions within each window during the analysis cycle. This generates highly multiplexed MS2 spectra, which contain fragment ions from multiple co-isolated precursors.3 A simplified visual overview of the DIA-MS workflow is shown in Fig. 1. While this results in more complex data, it allows for a near-complete sampling of the peptide content in a sample, enhancing reproducibility and quantitative depth.
3 Workflow differences
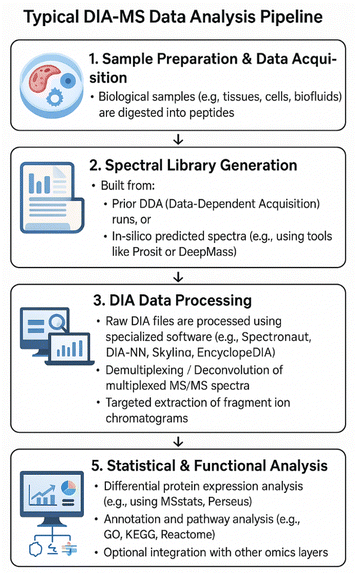
The DDA workflow begins with an MS1 scan to detect ion signals, followed by the selection of top N precursor ions for MS2 analysis. The cycle then repeats, often with exclusion rules to reduce redundancy. This method is efficient and produces high-quality spectra for abundant peptides but can miss low-abundance or transient species due to its stochastic sampling nature.17 As a result, DDA often suffers from limited reproducibility and inconsistent coverage between replicates.In contrast, the DIA workflow systematically steps through predefined m/z windows, fragmenting all ions within each window in a cycle. The resulting MS2 spectra are highly multiplexed and require computational deconvolution to assign fragments to their respective precursors. Advanced software tools such as Spectronaut, DIA-NN, and Skyline are employed to interpret these complex datasets, often leveraging spectral libraries or in silico predicted spectra.6,15 Although computationally intensive, this approach enables robust and reproducible quantification across multiple samples and experimental conditions. A comprehensive DIA-MS workflow is shown in Fig. 2.
4 DIA-MS workflow
4.1 Sample preparation
Sample preparation is foundational to DIA-MS performance, impacting peptide recovery, reproducibility, and proteome coverage.18 Innovations in this area have centered on automation, miniaturization, and compatibility with high-throughput platforms. Methods like SP3 (Single-Pot Solid-Phase-enhanced Sample Preparation) and S-Trap have become widely adopted due to their simplicity and high peptide recovery, with recent automation of SP3 workflows improving scalability and reducing user variability.19 For low-input applications such as single-cell proteomics, nanoPOTS and autoPOTS miniaturized platforms have significantly enhanced sensitivity while reducing losses. Additionally, although DIA is commonly used for label-free quantification, there's growing interest in combining it with TMT/iDiA-TMT isobaric labeling, enabling sample multiplexing while retaining the depth and consistency of DIA.20Weke and colleagues developed an optimized DIA-MS workflow for proteomic profiling of FFPE glioblastoma (GBM) tissues, addressing challenges such as protein cross-linking and poor extractability that typically limit FFPE analysis.21 After testing several lysis conditions, the authors established a protocol that enabled the identification of more than 1700 proteins and reliable quantification of over 1400 proteins from microdissected GBM specimens. Their dataset included accurate measurements of known GBM markers such as GFAP, FN1, VIM, and MBP, with median CV values below 12%, indicating strong precision across runs. Immune-related proteins including ILF2, MIF, and CD38 were also consistently detected. Overall, this work demonstrates that archival FFPE brain tumor tissues can support high-quality, reproducible DIA-MS proteomics, expanding access to valuable retrospective clinical samples.21
Kim et al. evaluated the feasibility of using DIA-MS in a clinical laboratory environment by analyzing twelve FFPE tumor biopsy specimens.22 Using a gas-phase fractionated library and single-shot DIA acquisitions, they quantified 3713 protein groups, including clinically actionable biomarkers. Unsupervised clustering based on the global proteome profiles clearly separated gastric cancer from colorectal cancer biopsies, and differential protein analysis identified promising diagnostic candidates. To assess translational potential, the group compared DIA-based protein quantification with SRM measurements for EGFR and TUBB3 and observed strong correlations, suggesting that DIA may serve as a viable alternative to targeted assays. They also correlated protein abundance with mRNA expression from matched samples, finding both concordant and discordant relationships that emphasize the unique value of proteomic data. Collectively, the study highlights the promise of DIA-MS for routine molecular diagnostics using FFPE biopsy material.22
Thomas and colleagues analyzed remnant peptides from more than 100 high-grade serous ovarian cancer (HGSOC) samples originally profiled by TCGA/CPTAC.24 They used SWATH-MS, a DIA variant, and evaluated its performance relative to CPTAC's traditional fractionated iTRAQ-DDA workflow. Although SWATH-MS produced shallower proteome coverage, it successfully recapitulated key subtype-associated proteomic signatures, including those defining the mesenchymal and homologous-recombination-deficient subtypes.24 Cross-platform comparisons showed strong correlations in protein abundance, and subtype cluster stability analyses revealed that sample size had a greater impact on classification robustness than the total number of quantified proteins. Importantly, the authors emphasized practical advantages of DIA-based SWATH-MS, including reduced sample consumption, simplified workflows, and higher throughput. Their findings highlight DIA approaches as efficient alternatives for large-scale clinical proteomic studies where reproducibility and resource constraints are major considerations.24
4.1.3.1 Methanol freshness. The freshness of methanol is critical during protein digestion because solvent degradation directly affects both peptide chemistry and enzyme performance. Over time, methanol undergoes slow oxidation when exposed to air or light, producing formaldehyde and formic acid. These reactive byproducts can modify peptide residues, especially lysine side chains and N-terminal amines leading to unwanted crosslinking or partial protein denaturation. Even trace amounts of these compounds can shift the digestion buffer's pH or destabilize trypsin, reducing its catalytic efficiency and resulting in incomplete digestion or an increased number of missed cleavages. Additionally, aged or impure methanol can alter protein solubility and disrupt the intended buffer composition, which may cause protein precipitation or reduce accessibility of cleavage sites. Using fresh methanol ensures a clean solvent environment, stable pH, preserved enzyme activity, and optimal peptide solubility for efficient and reproducible digestion.
4.1.3.2 Bubble removal. Removing bubbles prior to enzymatic digestion is essential because trapped air pockets interfere with the physical and chemical interactions required for efficient proteolysis. Air bubbles create barriers between proteins and trypsin, reducing their direct contact and slowing overall digestion kinetics. The air–liquid interface also promotes structural destabilization, increasing the likelihood of protein denaturation or trypsin autolysis detrimental to digestion efficiency. Furthermore, bubbles disrupt heat distribution and mixing uniformity within the digestion vessel, creating micro-gradients in temperature and reagent concentration that led to inconsistent cleavage patterns across the sample. Degassing the solution, whether by gentle vacuum, sonication, or centrifugation helps maintain a uniform liquid environment, ensuring consistent enzyme–substrate interactions, predictable reaction conditions, and improved reproducibility. Table 1 summarizes the effect of methanol freshness and bubble removal.
| Factor | Root cause | Effect on digestion | Preventive measure |
|---|---|---|---|
| Methanol freshness | Oxidation results in formation of formic acid/formaldehyde | Denature proteins & trypsin, causes incomplete cleavage | Use freshly opened, HPLC-grade methanol |
| Bubbles | Air–liquid interface, disrupted mixing | Inhibits enzyme contact, uneven digestion | Degas solvents and avoid vigorous pipetting |
4.2 Peptide separation (LC-MS)
Peptide separation prior to MS acquisition is critical for reducing sample complexity and improving identification and quantification. Recent advances in liquid chromatography (LC) have focused on increasing throughput without sacrificing resolution. High-performance columns such as µPAC (micro-pillar array columns) provide exceptional separation efficiency and peak capacity. In parallel, the Evosep One LC platform has emerged as a key innovation for high-throughput DIA workflows, supporting up to 300 samples per day while maintaining reproducibility and low carryover.21,25 Techniques like high-pH reversed-phase fractionation further enhance proteome depth, especially when used to generate deep spectral libraries or to improve identification in challenging sample matrices such as plasma.4.3 DIA acquisition
DIA acquisition strategies have evolved significantly, moving beyond fixed m/z window methods to more dynamic and efficient schemes. Traditional DIA windows (e.g., 25–30 Da) have given way to variable windowing and overlapping window techniques that optimize precursor density across the m/z range. A transformative innovation is diaPASEF (data-independent acquisition parallel accumulation–serial fragmentation), available on Bruker timsTOF Pro instruments, which combines ion mobility separation with DIA to boost speed, sensitivity, and proteome depth.22,26 Similarly, PulseDIA and scanning SWATH approaches dynamically vary acquisition parameters across runs to improve precursor coverage and sampling efficiency. More recently, real-time search-enabled DIA (RTS-DIA) platforms have emerged, allowing for adaptive fragmentation and acquisition refinement on the fly, increasing identification rates for low-abundance peptides.Multiplexed DIA (plexDIA): introduced in 2021, plexDIA increases throughput by parallelizing both samples and peptides, achieving multiplicative gains in data acquisition. MSX-DIA is an advanced acquisition method that extends traditional Data-Independent Acquisition (DIA) by multiplexing multiple precursor ions within a single MS/MS scan. This approach reduces the overall cycle time, increases throughput, and improves the signal-to-noise ratio by co-isolating and co-fragmenting ions with overlapping m/z values. The resulting spectra are then computationally demultiplexed to assign fragment ions to their respective precursors.23,27
(b) Enhanced quantification precision: studies have shown that MSX-DIA provides more accurate and precise quantification, particularly in complex samples such as histones.24,28
(c) Improved signal-to-noise ratio: the co-isolation and co-fragmentation of ions result in higher-intensity signals, which are beneficial for detecting low-abundance peptides.24,28
(b) Protein turnover studies: in pulse SILAC experiments, MSX-DIA has been utilized to profile protein turnover rates, showcasing its potential in dynamic proteomics.24,28
(c) Hybrid-DIA integration: the combination of MSX-DIA with targeted acquisition methods, such as Hybrid-DIA, allows for the simultaneous analysis of known and unknown peptides, enhancing the depth of proteomic profiling.26,29
(b) Optimization requirements: effective MSX-DIA requires careful optimization of acquisition parameters, including isolation window sizes and multiplexing schemes, to balance sensitivity and resolution.24,28
4.4 Spectral library generation
The spectral library is a cornerstone of DIA analysis, enabling accurate peptide identification by matching acquired fragment ion patterns against known references. Initially dependent on DDA (data-dependent acquisition) libraries, the field has rapidly moved toward in silico prediction using deep learning. Tools such as Prosit, DeepMass, and MS2PIP generate high-quality predicted MS/MS spectra for any peptide sequence, facilitating library-free and hybrid library workflows that are especially useful in non-model organisms or rare sample types.27,30 Additionally, experimental DDA libraries can be augmented with predicted spectra to improve coverage and identification rates. These innovations significantly reduce the need for time-consuming empirical library generation, making DIA more accessible and scalable for clinical and large-cohort studies.4.5 Data processing & analysis
Data analysis in DIA has progressed from traditional targeted tools to advanced, machine learning-powered platforms. DIA-NN has emerged as a leading open-source tool, enabling library-free analysis, deep proteome coverage, and interference correction through neural networks.28,31DIA-NN is widely regarded for its deep learning-enhanced spectral deconvolution and its ability to perform both library-based and library-free searches with exceptional sensitivity and speed. It excels at large-scale cohort processing, offers advanced interference correction, and typically produces some of the highest identification rates. Spectronaut, a commercial tool, provides a highly polished interface and robust statistical framework with industry-grade reproducibility. Its “directDIA” library-free mode and dynamic retention time correction make it a favored choice in clinical and translational studies where data quality and automation are paramount. OpenSWATH, part of the OpenMS ecosystem, is an open-source alternative known for its transparent algorithms and flexibility; although it requires more manual configuration, it remains valuable for method development and fully open workflows.
For targeted or hybrid analyses, Skyline is widely used due to its intuitive interface, strong community support, and versatility in handling both PRM/SRM and DIA data. While Skyline's library generation and peak scoring are not as automated as DIA-NN or Spectronaut, it offers precise manual curation tools that are ideal for validation studies. Other emerging tools such as EncyclopeDIA leverage chromatogram libraries to improve identification depth, while platforms like MSFragger-DIA bring ultrafast searching and compatibility with large cohorts. Together, these tools represent a spectrum of trade-offs: DIA-NN for maximal sensitivity and speed, Spectronaut for stability and high-confidence quantification, Skyline for targeted refinement and validation, and OpenSWATH for flexible, fully open-source workflows. The choice of software ultimately depends on study goals—discovery, large-scale quantification, clinical reproducibility, or targeted validation. Table 2 summarizes the different software tools used for data analysis.
| Tool | Type/license | Library mode | Key strengths | Typical limitations |
|---|---|---|---|---|
| DIA-NN | Open-source (GitHub) | Library-free (neural-network driven) and library-assisted | Extremely fast; excellent sensitivity and FDR control; built-in neural nets for spectrum prediction and interference correction; strong for dia-PASEF and high-throughput runs | Black-box ML models can be hard to interpret; needs careful parameter tuning for very noisy data |
| Spectronaut (biognosys) | Commercial (license) | Library-based and robust library-free (directDIA) | Polished GUI, extensive QC/reporting, excellent engineering for peptide quant and targeted scoring; widely used in clinical/CPTAC pipelines | Cost/licensing; some advanced workflows require commercial support |
| OpenSWATH/OpenMS | Open-source (OpenMS ecosystem) | Library-based (SWATH style targeted extraction) | Strong, well-documented targeted extraction framework; integrates with OpenMS tools and pyProphet for FDR; historically foundational for DIA | Historically more complex to install/configure; steeper learning curve than GUI tools |
| MaxDIA (MaxQuant 2+) | Open-source (MaxQuant) | Library-based & library-free (MaxDIA workflows) | Tight integration with MaxQuant ecosystem; strong protein inference and well-validated FDR control; good quant precision | Newer relative to legacy MaxQuant features for DDA—some users report different behavior vs. DIA-native tools; GUI oriented to MaxQuant users |
| FragPipe/MSFragger-DIA | Open-source (FragPipe suite) | Library-free and hybrid (MSFragger-DIA, build libraries for DIA-NN etc.) | Rapid database searching of DIA spectra, integrated pipeline (library building, quant extraction), good for exploratory and dia-PASEF data | Multi-component pipeline (more modules to configure); still maturing for some specialized DIA setups |
| Skyline | Open-source (windows GUI) | Targeted/hybrid (supports DIA extraction) | Excellent for method development, targeted verification, manual inspection and peak curation; integrates with panorama | Not optimized as an end-to-end automated discovery DIA pipeline for very large cohorts (better for verification/targeted steps) |
4.6 Quantification & statistical analysis
Quantification in DIA is highly reproducible due to its systematic and unbiased nature. New analytical frameworks are enhancing statistical power while maintaining rigor. MSstats and Triqler offer advanced statistical modeling, accounting for missing data, technical variation, and false positives. For single-cell and ultra-low-input applications, protocols like nanoPOTS-DIA combine miniaturized sample prep with DIA, enabling reproducible quantification of thousands of proteins from single cells.29,32 prolfqua, an R-based pipeline, supports flexible and user-friendly longitudinal analysis of DIA data. Together, these tools ensure that quantitative outputs are biologically meaningful and statistically robust, especially when analyzing complex clinical or time-course datasets.4.7 Visualization and biological interpretation
Post-quantification, visualizing and interpreting DIA-MS data is essential for generating actionable biological insights. Tools like Perseus, LFQ-Analyst, and Protti facilitate differential expression analysis, volcano plots, PCA, and clustering. Recent tools such as OmicScope integrate proteomics data with pathway and gene set enrichment analyses, providing a holistic view of biological systems. Additionally, multi-omics integration frameworks are increasingly used to contextualize proteomic changes with transcriptomic or metabolomic data. Platforms like Cytoscape and Ingenuity Pathway Analysis (IPA) allow for network and pathway modeling, while machine learning frameworks are being applied for biomarker discovery and patient stratification in clinical proteomics.• Variability in sample preparation (e.g., digestion, clean-up, enrichment) and chromatography (e.g., LC gradient, column length, mobile phases).
• Differences in instrument platforms, MS acquisition (window scheme, collision energies, scan cycle times) and ion mobility setups.
• Dependence on spectral libraries (or library-free approaches) for peptide identification and quantification: libraries differ in how they were built, what they cover, how retention time was calibrated, etc.
• Data-processing pipeline heterogeneity: software choice, scoring thresholds, FDR (false discovery rate) control, retention time alignment, quantification strategy, all affect final numbers. Because clinical translation demands high rigor (e.g., low intra- and inter-lab variability, linearity, known limits of detection/quantitation), standardization of DIA workflows and spectral libraries is not optional, it is essential.
Efforts to harmonize DIA-MS for clinical applications now operate on multiple, complementary fronts: community standards for data and libraries; large, shared spectral resources; consortium-led pipelines and reference materials; coordinated inter-laboratory benchmarking; and reproducible, containerized computational workflows. Together these reduce the two largest sources of variability in DIA studies: (a) differences in spectral libraries and their metadata/formatting, and (b) pre-analytical and chromatographic differences that affect retention time and ion signatures, and they provide routes to regulatory-grade, auditable analysis.33
Standardized spectral-library formats and fragment-annotation models have been a major recent advance. The HUPO Proteomics Standards Initiative (PSI) has developed machine-readable specifications (mzSpecLib for library containers and mzPAF for fragment-peak annotations) to capture peptide assays, provenance and fragment-ion interpretation in a consistent, software-agnostic way; these formats make it far easier to exchange, validate and re-use libraries across tools and repositories. Adoption of such standards (and their reference implementations) directly reduces errors that arise when converting between ad-hoc formats or when metadata are absent.34
Library standardization efforts resulted in,
(a). DIALib-QC evaluates spectral libraries in DIA proteomics: it examines 62 parameters of compliance, completeness and correctness of libraries. This helps users and labs to assess library quality and thus standardise library usage.35
(b). The algorithm STAVER is designed to reduce non-biological variation in large-scale DIA-MS data by leveraging standardised benchmark datasets and hybrid spectral libraries (generated from diverse platforms/labs) and thereby enhances reproducibility of protein quantification.36
(c). In plasma proteomics, an evaluation of four library workflows (fractionated DDA, fractionated DIA, predicted spectra, gas-phase fractionation (GPF) for diaPASEF) found that the GPF workflow outperformed traditional DDA library workflows in terms of number of identified and quantified proteins in a cohort of non-small cell lung cancer patients.37
Standardized spectral-library formats and fragment-annotation models have been a major recent advance. The HUPO Proteomics Standards Initiative (PSI) has developed machine-readable specifications (mzSpecLib for library containers and mzPAF for fragment-peak annotations) to capture peptide assays, provenance and fragment-ion interpretation in a consistent, software-agnostic way; these formats make it far easier to exchange, validate and re-use libraries across tools and repositories. Adoption of such standards (and their reference implementations) directly reduces errors that arise when converting between ad-hoc formats or when metadata are absent.38
At the same time, community pan-human and tissue-focused libraries provide shared assay backbones that laboratories can adopt rather than building small, idiosyncratic libraries that drive between-site differences. Examples include legacy Pan-Human assay collections and more recent, expanded DIA pan-human libraries such as the DPHL v2 resource, plus curated assay portals such as SWATHAtlas—these resources increase coverage and standardization for many human tissues and biofluids used in translational work. Using a validated pan-library (or thoroughly documented predicted library) reduces library-derived bias in discovery and targeted DIA analyses.
Consortia and standard-setting organizations play a central role in translating DIA to clinical use by producing reference datasets, recommended SOPs and performance metrics. The Clinical Proteomic Tumor Analysis Consortium (CPTAC) has produced harmonized proteogenomic datasets and standardized pipelines that serve as community benchmarks, while NIST continues to develop reference methods and reference materials (SRMs/RGTM) and metrics for quantitative proteomics that are needed for clinical validation. These efforts encourage laboratories to adopt agreed-upon system suitability tests, spike-in panels (e.g., iRT peptides), pooled QC samples and acceptance criteria that are essential for longitudinal and cross-site reproducibility.23
Independent benchmarking and multi-lab ring trials have quantified the remaining sources of variability and tested how well harmonization efforts work in practice. Recent benchmark studies comparing DIA software and workflows (and ABRF multi-laboratory investigations) show that, while DIA can achieve high reproducibility when common sample prep, LC-MS settings, QC and library resources are used, method-specific choices (sample handling, chromatography, library generation and search settings) still introduce measurable differences. These benchmarking exercises have therefore become standard practice for method validation and for demonstrating equivalence across clinical sites.33
On the computational side, reproducibility is being enforced by publishing fully specified, containerized workflows and by wide use of well-documented tools (OpenSWATH, DIA-NN, MSFragger-DIA/FragPipe, Skyline and others). Containerization or workflow managers (Docker/Singularity, Nextflow/Workflow Description Language) together with frozen parameter sets and versioned library files allow analyses to be rerun exactly and inspected during regulatory review. Parallel developments in machine-learning-based library prediction and direct (library-free) DIA search methods further give laboratories flexibility while simplifying the dependency on locally built DDA libraries—but these approaches must themselves be benchmarked against gold-standard references before clinical adoption.33
Despite progress, gaps remain. Universal adoption of mzSpecLib/mzPAF is still ongoing, many legacy libraries lack rich provenance metadata, standardized reference materials for low-abundance clinical biomarkers are incomplete, and inter-laboratory ring trials must expand to wider sample types and instruments (including new ion-mobility and high-throughput platforms). For a laboratory preparing a clinical DIA assay today, best practice therefore combines (1) use of a community-accepted library format (mzSpecLib/mzPAF), (2) reliance on validated pan-human or tissue-specific libraries when possible, (3) rigorous system suitability/QC (iRT, spike-ins, pooled reference), (4) participation in inter-lab benchmarking, and (5) deployment of containerized, versioned pipelines with documented parameter sets and acceptance thresholds.34
Scalability and accessibility depend heavily on local resources and study priorities. Modern DIA workflows paired with faster instruments (e.g., timsTOF/PASEF and newer Orbitrap families) and streamlined sample prep can reach high throughput and lower per-sample run times, and ongoing method simplifications and low-cost workflows are narrowing the cost gap to affinity assays—but the capital expenditure for high-end mass spectrometers and the need for experienced spectral-library/analysis expertise remain barriers for some clinical laboratories.42 Single-cell proteomics and highly multiplexed DIA variants are exciting and rapidly maturing, but currently add complexity and cost that limit immediate large-cohort deployment; they are most valuable when cell-level resolution is required rather than population-scale biomarker screening.43
In practice, many large clinical programs favor a hybrid strategy: use high-throughput affinity platforms or targeted MS for initial screening where speed, low cost per sample, and sensitivity for pre-defined panels matter, and deploy DIA-MS (or DIA-derived targeted assays) for discovery, orthogonal validation, and when peptide-level isoform/variant information is crucial. When choosing a platform for a large-scale clinical study weigh (a) required depth and peptide-level information, (b) acceptable per-sample costs (reagents, instrument time, personnel), (c) throughput timelines, and (d) the ability to produce reproducible, auditable pipelines—DIA often offers the best compromise for discovery-to-validation workflows, but the optimal choice is study dependent.39Table 3 presents a comparative view of DIA-MS vs. other emerging proteomic technologies.
| Platform | Capital cost | Typical per-sample cost (approx.) | Accessibility | Scalability/best use |
|---|---|---|---|---|
| DIA-MS (label-free, dia-PASEF or Orbitrap DIA) | High (Orbitrap/timsTOF) | Low-moderate (depends on LC time; reduced with short gradients) | Moderate-high expertise | High for discovery and medium-to-large cohorts with robust quality control44 |
| Affinity platforms (Olink, SomaScan) | Low for user labs (vendor-run) | Low-moderate per panel (Olink ≈$100 per panel reported; SomaScan higher for broader panels) | Low (vendor service; easy for clinical labs) | Very high for large cohorts; limited to panel content |
| TMT (multiplexed DDA) | High | Moderate-high (reagent kits cost thousands; per-sample cost falls with multiplexing) | High (MS expertise; complex prep) | Good via multiplexing but requires fractionation & careful batch design45 |
| SRM/PRM (targeted MS) | Moderate | Low per sample after assay development | Moderate (assay development skills required) | Excellent for validated small panels; poor for discovery scale40 |
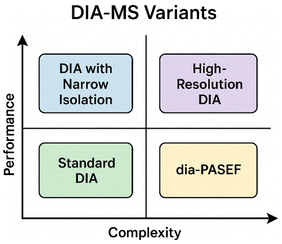
Narrow-window DIA strategies improve precursor selectivity and quantitation accuracy by reducing window widths (e.g., < 5 m/z), thereby decreasing spectral interference and boosting peptide identification confidence. For instance, a recent study optimizing narrow isolation windows on a ZenoTOF 7600 (“nSWATH”) showed improved identification by ∼10% and median CVs <6%.48 These methods can achieve deeper coverage of complex proteomes, albeit at the cost of longer cycle times or multiple injections, which reduce overall throughput.
High-resolution DIA further enhances analytical fidelity by leveraging high-performance Orbitrap or TOF analyzers to acquire fragment ions at high resolving power (e.g., >30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000), improving discrimination among co-eluting peptides and post-translational variants. A recent publication described a small-mass-range, narrow-window scheme achieving >9200 protein groups in HEK lysates with CV ≈ 3% using one-shot DIA.49 These approaches have shown excellent reproducibility and precision in complex clinical matrices but demand higher instrument cost and more intensive data processing pipelines.
000), improving discrimination among co-eluting peptides and post-translational variants. A recent publication described a small-mass-range, narrow-window scheme achieving >9200 protein groups in HEK lysates with CV ≈ 3% using one-shot DIA.49 These approaches have shown excellent reproducibility and precision in complex clinical matrices but demand higher instrument cost and more intensive data processing pipelines.
Finally, dia-PASEF (Parallel Accumulation–Serial Fragmentation combined with ion-mobility) integrates an additional separation dimension—ion mobility—on the Bruker timsTOF platform, substantially improving ion utilization, sensitivity and proteome depth while maintaining rapid duty cycles. A recent study demonstrated that diaPASEF enabled quantification of hundreds of host-cell proteins (HCPs) that were undetectable by standard DDA, improving throughput, coverage and data completeness.50 Although dia-PASEF requires specialized hardware and analysis tools (e.g., FragPipe, DIA-NN) it represents a major advance in throughput and sensitivity for large-scale proteomic studies.
Collectively, these DIA variants illustrate a continuum—from simple, high-throughput DIA workflows optimized for population-scale reproducibility to advanced, high-resolution ion-mobility-assisted methods aimed at maximizing proteomic depth and analytical precision. Understanding these trade-offs is increasingly important when choosing workflows for clinical studies that demand reproducibility, scalability and regulatory-grade robustness.
Targeted methods such as PRM and SRM provide the highest quantitative precision and specificity, making them the gold standard for absolute quantification and validation of clinically actionable biomarkers. SRM, performed on triple quadrupoles, offers excellent linearity, low limits of detection, and robust quantification but measures only predefined transitions. PRM, typically performed on high-resolution instruments, combines targeted isolation with full MS/MS spectra, improving selectivity and reducing interference. However, both PRM and SRM are limited in throughput because they can monitor only tens to hundreds of peptides per run. Compared with these targeted approaches, DIA-MS provides a powerful intermediate solution—far more comprehensive in proteome coverage while still maintaining excellent reproducibility, making it especially advantageous for discovery-to-validation pipelines, large clinical cohorts, and retrospective analysis of archived specimens.
DIA-MS offers the optimal balance between proteome depth, reproducibility, and scalability, making it well suited for large-scale discovery and translational studies. While DDA-MS remains valuable for exploratory proteomics and spectral library generation, PRM/SRM approaches excel in targeted quantification and clinical assay validation where absolute precision is required. Table 4 compares the techniques.
| Metric | DIA-MS | DDA-MS | PRM/SRM (targeted MS) |
|---|---|---|---|
| Depth of coverage | Very high (up to 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–12 000 proteins in deep profiling studies) 000–12 000 proteins in deep profiling studies) |
Moderate; typically 3000–6000 proteins depending on sample complexity | Low-moderate; focused on 10–100 s of predefined targets |
| Reproducibility | High; consistent sampling across all ions reduces stochastic variability | Moderate; stochastic precursor selection causes run-to-run variability | Very high; targeted acquisition ensures consistent quantification |
| Missing value rate | Low (<10% across large cohorts) | High (20–50%, especially in label-free experiments) | Very low (near-zero for targeted analytes) |
| Quantitative accuracy | High; label-free or library-based quantification with robust normalization | Moderate; affected by missing data and precursor interference | Very high; precise quantification using stable isotope standards |
| Throughput/scalability | High; suitable for large clinical cohorts | Moderate; requires fractionation or repeated runs for deep coverage | Low–moderate; limited multiplexing capacity |
| Best suited for | Discovery and translational proteomics; large-scale cohort studies | Exploratory proteomics; hypothesis generation | Targeted validation; clinical assay development |
Because the same m/z ranges are sampled in each duty cycle, DIA significantly reduces the missing value problem that often limits DDA-based quantification. Consequently, it offers enhanced precision and reproducibility, making it particularly well suited for clinical proteomics and large-cohort biomarker discovery studies.52
Another key advantage of DIA-MS is its capacity for deep proteome coverage and retrospective data mining. The comprehensive and systematic acquisition of fragment ion maps allows researchers to revisit existing datasets as spectral libraries or search algorithms improve, thereby maximizing data reuse and knowledge extraction.51,53 The combination of broad coverage and consistent sampling also enhances label-free quantification accuracy, supporting robust comparative analyses across biological replicates or study conditions. Moreover, advances in hardware and acquisition strategies—such as variable isolation window designs and staggered window schemes—have further optimized the balance between cycle time and sensitivity, enabling application of DIA to diverse workflows ranging from deep discovery proteomics to targeted verification and even single-cell studies.53,54
Recent computational advances have greatly extended the analytical power of DIA-MS. Deep-learning-based approaches, spectrum prediction, and library-free search algorithms have improved peptide identification and quantification from highly multiplexed spectra.54 Tools such as DIA-NN, Spectronaut, and FragPipe-DIA now enable precise analysis of highly complex proteomes while maintaining stringent false discovery rate (FDR) control. Collectively, these innovations have made DIA-MS a robust, scalable platform for quantitative proteomics, increasingly favored for translational and clinical applications.39
Despite these advantages, DIA-MS is not without limitations. The multiplexed nature of DIA spectra, in which fragment ions from multiple co-isolated precursors overlap, imposes a major analytical challenge. Accurate peptide identification and quantification depend on sophisticated deconvolution algorithms and high-quality spectral libraries, and suboptimal computational workflows can introduce false positives or reduce sensitivity.52,54 The design of isolation windows also involves trade-offs: narrower windows reduce spectral complexity but extend cycle times and may compromise chromatographic sampling, whereas wider windows improve throughput at the cost of sensitivity for low-abundance peptides.53
Although DIA improves reproducibility, the detection of very low-abundance proteins, rare proteoforms, or site-specific post-translational modifications remains challenging without prior enrichment, fractionation, or complementary targeted strategies.51 The approach also generates very large datasets that require considerable computational power, data-storage capacity, and rigorous quality-control frameworks to ensure reproducible results. Furthermore, the dependence on spectral library quality remains a critical consideration for library-based workflows, as incomplete or poorly matched libraries can bias identification and quantification outcomes. Hybrid or library-free strategies partially mitigate these issues, yet their performance still varies depending on sample type and instrument configuration.52,54,55
In summary, DIA-MS provides a powerful balance between proteome depth, reproducibility, and quantification accuracy, making it highly attractive for large-scale and clinical proteomic investigations. However, challenges related to spectral complexity, low-abundance detection, computational demands, and library dependency continue to drive methodological development. Continued innovation in acquisition design, machine-learning-based spectral interpretation, and library-free data processing will further enhance the sensitivity, throughput, and interpretability of DIA-MS workflows, solidifying its role as a cornerstone of modern quantitative proteomics. Table 5 compares DIA and DDA.
| Step | DDA | DIA |
|---|---|---|
| Precursor selection | Based on real-time ion intensity | Systematic across all ions in m/z windows |
| Fragmentation strategy | Targeted MS2 of selected ions | Fragmentation of all ions within set m/z windows |
| Spectral complexity | Cleaner MS2 spectra | Highly multiplexed MS2 spectra |
| Data analysis | Database search or spectral library matching | Requires spectral libraries or deep learning models |
| Reproducibility | Lower; stochastic sampling | Higher; consistent coverage |
4.7.6.1 Molecular characterization of cancer with DIA-MS. The increasing number of applications of molecular and omics-based approaches has allowed the development of novel classification approaches based on molecular changes and offered unparalleled comprehension of the biological basis of cancer, even though conventional categorization of tumors based primarily on histopathological evaluation has been crucial in the early detection and therapeutic handling of disease.56–58 Recent developments in MS technologies have opened new avenues for deep proteome profiling of clinical cohorts, which can be used to uncover significant biochemical pathways unique to a disease and improve the current cancer classification methods. Additionally, there is a great need for strong cancer markers for accurate and timely tumor diagnosis, treatment selection, and patient outcome prediction. In this context, DIA-MS has been used to perform proteome characterization of several cancer types, such as prostate, liver, kidney, and breast cancer, a few of those being highlighted in this section.52Table 6 presents recent cancer studies with DIA.
| Design | Sample | Proteome coverage | MS | Findings | Study |
|---|---|---|---|---|---|
| Prostate cancer (PC) | |||||
| Serum proteomic profiling of patients with PC and benign prostatic hyperplasia (BPH) | 30 BPH and 30 PC for MS analysis. 35 BPH and 51 PC samples analyzed using ELISA and immunoturbidimetry | 56 DEPs between early-stage PC and BPH and 47 DEPs between aggressive and non-aggressive PC patients | Hybrid qudrapole ion-trap | Serum SPP1 and CP could be effective biomarkers to differentiate aggressive PC from non-aggressive disease | Wang et al. (2022)68 |
| Proteomic characterization of primary and metastatic prostate cancer | Cohort consisted of 106 fresh frozen prostate tissues, including 48 primary and 48 metastatic PC and 10 benign prostate tissue controls (C) | 4415 Proteins were identified in the tumor and benign control tissues, including 158 up-regulated and 116 down-regulated proteins in AG tumors | Triple TOF | Dipeptidyl peptidase 4 (DPP4) plays a critical role in PC progression | Li et al. (2021)67 |
| Analze serium proteome and lipidome by LC-MS from samples of patients diagnosed with PC, benign prostatic hyperplasia (BPH) | Serum samples obtained from newly-diagnosed 127 PC patients, 37 patients with BPH, and 110 healthy controls (HC) | NA | TOF | The study identified proteins and lipids, including those in MAPK pathways and lipids containing oleic acid, that may serve as potential biomarkers to improve diagnostic specificity | Spick et al. (2023)124 |
| Quantification of multiple biomarkers to achieve an increased accuracy in predicting PC | Urine samples from 45 men with elevated prostate specific antigen (PSA) levels | 2735 Proteins across all samples, as well as potential biomarkers for the detection of all grades of PC | Orbitrap | The study presented a feasible method to reduce the number of unnecessary prostate biopsies and detect both healthy men and clinically significant PC | Alijaj et al. (2022)176 |
| Parallel reaction monitoring (PRM) assays for the detection of aggressive prostate cancer | Urine specimens collected from 69 nonaggressive and 73 aggressive (Gleason ≥ 8) PC patients | NA | Hybrid qudrapole orbitrap | PRM assays can serve as an adjunct to the serum prostate specific antigen (PSA) test to achieve better predictive power toward aggressive PC | Dong et al. (2021)177 |
| Proteomic profiling of extracellular vesicles (EVs) | 40 participants as the discovery cohort & 20 participants as the validation cohort | 18 EV proteins have been identified (specifially, ERPINA3, LRG1, and SCGB3A1were upregulated) | Hybrid qudrapole orbitrap | The study demonstrates a translational workflow to identify EV proteins as molecular markers to improve the clinical diagnosis of prostate cancer | Zhang et al. (2022)178 |
| Phosphoproteomics using LC-MS to comprehensively investigate changes in phosphorylation due to JPH203 administration | Human prostate cancer (PC) cell lines PC-3 (RRID: CVCL_0035) and DU145 (RRID: CVCL_0105) | NA | Quadrupole orbitrap | Cell cycle-related pathways were affected by JPH203 | Rii et al. (2023)162 |
| Protein content of exosomes secreted from PC cells through treatment with diferent AR-antagonists or SAL has been analyzed | Human PC LNCaP cells | 1070 Proteins were up regulated and 52 proteins downregulated by SAL whereas enzalutamide upregulated 151 proteins and downregulated 42 exosomal proteins | Quadrupole orbitrap | Cargo of exosomes is controlled by AR-agonist | Roozbahani et al. (2024)76 |
| Screening of novel biomarkers for acute kidney transplant rejection | Serum proteomic profiling of kidney transplant patients with T cell-mediated acute rejection (TCMR) and stable allograft function (STA) | NA | Quadrupole orbitrap | Higher concentrations of serum amyloid A protein 1 (SAA1) and insulin like growth factor binding protein 2 (IGFBP2), and lower fetuin-A (AHSG) concentration were found in TCMR patients when compared with STA patients | Wang et al. (2024)77 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Pancreatic cancer (PaC) | |||||
| Discover molecular signatures for the early detection of pancreatic ductal adenocarcinoma (PDAC) | 29 Patient-matched PDAC tissues and sera | 892 N-linked glycopeptides originating from 141 glycoproteins had PDAC-associated changes beyond normal variation | Tribrid MS | PDAC tissue-associated glycoproteins represent an inventory of serum-detectable PDAC-associated glycoproteins as candidate biomarkers | Lih et al. (2024)179 |
| DIA-MS based proteomic stragies | Library containing 10633 protein group and 184552 peptides | EMT was significant for CFPAC-1, AsPC-1, and PANC-1 cells | Q-exactive HF-MS | Library provides a valuable resource for in-depth proteomic analysis | Kong et al. (2022)144 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Ovarian cancer (OC) | |||||
| DIA-based MS analysis of serum proteometo identify potential diagnostic biomarkers for HGSOC | Depleted serum samples from 26 HGSOC cases and 24 healthy controls (HCs) | 1847 Proteins were identified across all samples, among which 116 proteinsshowed differential expressions between HGSOC patients and HC | Orbitrap MS with FAIMS | Biomarker panel obtained was able to discern HGSOC from HCs with high sensitivity and specificity | Huh et al. (2022)180 |
| DIA- and DDA-based proteomics to analyze differentially expressed proteins in tissue and cell samples of ovarian cancer | 50 Specimens of HGSOC (22 platinum-resistant and 28 platinum-sensitive tissue samples) from ovarian cancer patients | NA | Quadrupole orbitrap | CAAP1, reported to be involved in the regulation of apoptosis, may be negatively correlates with platinum resistance in ovarian cancer | Ni et al. (2023)181 |
| Identify potential prognostic biomarkers in urine specimens from patients with HGSOC | Urine samples from 56 HGSOC patients were collected before primary debulking surgery or initiation of neoadjuvant chemotherapy | NA | Quadrupole orbitrap | Four candidate biomarkers (ANXA1, G6PI, SPB3, and SPRR3) were validated | Ni et al. (2024)135 |
| High-throughput proteomics analysis of urine samples to discriminate between the two ovarian tumor types | 132 Urine samples from 73 malignant and 59 benign cases of ovarian carcinoma patients | 2199 Proteins were quantified and 69 identified with differential expression in benign and malignant groups | Quadrupole orbitrap | DIA-MS based urinary proteomics was combined with machine learning to establish a novel classifier for discriminating between malignant and benign ovarian tumor | Ni et al. (2021)81 |
| Potential of extracellular vesicles (EVs) circulating in blood, as a source for biomarker discovery | EVs isolated from blood plasma, ascites, and cell lines of patients. 30 HGSC patients and 30 control subjects | NA | Quadrupole orbitrap | The study helped identify biomarkers with high specificity and sensitivity | Cooper et al. (2024)140 |
| Proteomic profiling of ovarian clear cell carcinomas | Formalin-fixed paraffin-embedded (FFPE) CCOC tissue specimens from a cohort of 32 patients with CCOC &28 fresh frozen (FF) CCOC tissue specimens from an independent cohort of 24 patients with CCOC | Characterized 8697 protein from FFPE tissue specimens & 9409 proteins from fresh tissue speciments | Quadrupole orbitrap | The study presents a CCOC proteomic data resource, which could potentially predict the recurrence and survival of CCOC | Yue et al. (2024)92 |
| Characterize the proteome in menstrual blood from patients with endometriosis | Menstrual blood samples from patients with and without endometriosis were obtained on days 1, 2, or 3 of the menstrual cycle | A total of 95 differentially expressed proteins (DEPs) were identified in menstrual blood from women with and without endometriosis | Tribrid MS | The study contributes to the identification of putative endometriosis biomarkers from non-invasive samples and the roles of CXCL5 and IL1RN in the pathogenesis of endometriosis | Ji et al. (2023)182 |
| DIA-MS and PRM-MS analyses were conducted on urine samples | Urine samples from HGSOC patients were collected before surgery or neoadjuvant chemotherapy | NA | Quadrupole orbitrap | Our candidate biomarkers (ANXA1, G6PI, SPB3, and SPRR3) were identified | Ni et al. (2024)166 |
| Label-free quantification of protein markers of chemotherapy response | FIGO stage IIIC HGSOC tissues from 12 patients were obtained from patients | NA | QTOF | GSN, CALM1, and TXN may be useful biomarkers for predicting chemotherapy response and understanding the mechanisms of chemotherapy resistance | Arentz (2023)165 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Liver cancer (Lic) | |||||
| Establish a high-coverage human hepatocellular carcinoma (HCC) spectral library | 19 Tissue samples from patients with early-stage HCC were used | 6000 Proteins from HepG2 cells could be quantified | Quadrupole orbitrap | One-shot DIA identified proteins from HCC samples in lesser time | Weng et al. (2021)183 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Kidney cancer (KC) | |||||
| Phosphoproteomics of urinary extracellular vesicles | Urine samples were collected | 2584 Unique phosphosites were quantified | Quadrupole orbitrap | Results show that EV phosphoproteome analysis carried out in the study provide a powerful tool for future clinical applications | Hadisurya et al. (2023)112 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Rectal cancer (RC) | |||||
| Proteomics characterization of preoperative LARC biopsy samples | Rectal cancer FFPE samples obtained from 97 LARC patients treated with intense chemoradiotherapy, comprising radiation and chemotherapy | 915 Proteins were chartacterized | Quadrupole orbitrap | DIA-MS can unravel new tissue molecular features that might lead to different responses to nCRT | Stanoevic et al. (2025)154 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Lung cancer (LC) | |||||
| Proteomic study was performed to identify potential biomarkers | Exhaled breath condensate (EBC) samples were collected from 10 lung cancer patients and 10 healthy controls | 1151 Proteins were identified | Quadrupole orbitrap | S100A11, ANXA1, ENO1, and FABP5 might play a vital role in the EBC proteome | Ma et al. (2021)185 |
| Single-cell proteomics with DIA-MS | Adenocarcinoma cell line PC-9 | 1500 Protein groups across 20 single mammalian cells were identified | Tribrid MS | DIA workflow offers all-in-one cell characterization, analytical sensitivity and robustness | Gebreyesus et al. (2022)184 |
| Proteogenomic analysis of non-small cell lung cancer (NSCLC) | Tumor samples from a total of 192 patients | 6 Distinct proteome subtypes with differences in immune cell composition and subtype-specific expression of immune checkpoints | HiRIEF-LC-MS | The study linked immune evasion with LAG3 via STK11 mutationdependent HNF1A activation and FGL1 expression | Lehtiö et al. (2021)99 |
| Longitudinal plasma proteomic profiling of lung cancer | 159 longitudinal plasma samples were collected from 63 ALK-positive NSCLC patients treated with crizotinib | 737 Plasma proteins were profiled from 159 pre-treatment and on-treatment plasma samples of 63 ALK-positive NSCLC patients | Quadrupole orbitrap | The study contributes to the search for prognostic and drug-resistance biomarkers in plasma samples for ALK-TKI therapy | Wang et al. (2024)132 |
Among males, prostate cancer (PC) is among the most common types of cancer. Despite the remarkable advancements in detection and diagnostic procedures, PC continues to be the second most common cause of cancer-related mortality among men in Western nations.59 While high-grade PC patients typically have bone metastases, which can result in severe consequences and high mortality rates, most PC patients appear with indolent, slowly growing tumors.60,61 Active surveillance can safely monitor many PC cases, which typically manifest as slow-growing and/or limited disease without requiring invasive treatments. On the other hand, certain cancers exhibit aggressive activity that leads to further development, metastases, and death.37 Numerous recent research findings have also shown that, among males over 50, the incidence rate of advanced-stage disease is still increasing, while the incidence rate of localized disease is still declining.62–64
Numerous risk stratification systems, such as combining clinical and pathological parameters (e.g., Gleason score/ISUP grade, PSA levels, clinical and pathological staging), have been developed to distinguish between high-risk aggressive PC (AG) and low-risk non-aggressive indolent tumors (NAG). Nevertheless, these methods are still insufficient in forecasting the course and results of the disease.65,66 A proteome analysis of primary PC (containing both indolent NAG and AG subtypes) and metastatic PC was conducted using DIA-MS to gain a better understanding of the molecular characteristics of the PC progression.67 The study's main goals were to create a thorough proteome map of AG, NAG, and metastatic PC using a DIA technique and to find distinct proteins linked to AG that might be utilized to create a clinical test to distinguish AG from NAG PC.67
One of the study's distinctive features is its integrated investigation of several clinical phases of PC, such as metastatic PC with androgen deprivation therapy (M) and without (Nmet), PC with a Gleason score of 6 (NAG), and PC with a Gleason score over 7 that led to death (AG). Among the 4415 proteins identified in PC in this study, 158 were up-regulated and 116 were down-regulated in the AG subtype in comparison to NAG samples.67 A functional analysis showed that, in comparison to primary NAG PC and normal prostate tissues, the expression of specific proteinases, including dipeptidyl peptidase 4 (DPP4), carboxypeptidase E (CPE), and prostate-specific antigen (KLK3), was significantly changed in both primary aggressive PC and metastatic tumors.67 By specifically reexamining DIA-MS maps, the functional significance of DPP4 was further elucidated. NPY-cleaved peptides were reduced in AG tumors, while neuropeptide Y (NPY), a DPP4 substrate, was accumulated.67
An investigation was carried out by Wang et al. (2022) to find blood proteins that were both specific and sensitive enough to diagnose aggressive and early-stage PC.68 The bioinformatics analysis and DIA-MS were used to thoroughly examine the serum proteome profile of individuals with PC and benign prostatic hyperplasia (BPH).68 Utilizing the enzyme-linked immunosorbent assay (ELISA) and immunoturbidimetry test, the differentially expressed proteins (DEPs) significance was further confirmed.68 56 DEPs were found between early-stage PC and BPH, and 47 DEPs were found among aggressive and non-aggressive PC patients, indicating a statistically significant difference in abundance.68 Furthermore, as demonstrated by the verification results, the concentration of osteopontin (SPP1) and ceruloplasmin (CP) rose with a higher Gleason score, and the serum L-selectin concentration was considerably greater in Gleason 6 PC compared to BPH. Comparing PSA diagnosis alone with the addition of SPP1 and CP, the study showed that the separation of Gleason 7 (4 + 3) or above from Gleason 7 (3 + 4) or below was improved.68
The relationship between prostate tumor cells and their environment is widely recognized as one of the major factors impacting the disease's progression. This interaction affects several PC characteristics, such as proliferation, angiogenesis, metastasis, treatment resistance, and survival.69,70 It has been shown that exosomes facilitate communication within the tumor microenvironment.71 Numerous findings lend credence to the link between exosomes and cellular senescence,72 with exosome production being increased in conjunction with oxidative stress and cellular senescence brought on by radiation.73–75
Roozbahani et al. (2024) evaluated whether the androgenic receptor (AR) complex controls the protein composition of exosomes using AR-ligands. Determining the functional effects of exosomes on growth was another goal of the study.76 DIA-MS evaluation determined which proteins in exosomes are ligand-specific. The supraphysiological androgen level (SAL) up-regulated 1770 proteins and down-regulated 52, while enzalutamide up-regulated 151 proteins and down-regulated 42 exosomal proteins.76 According to the bioinformatic prediction, pro-proliferative pathways are regulated. The exosome hub factors AKT1, CALM1, PAK2, and CTNND1 are enhanced by AR ligands. Thus, the isolated exosomes from AR-ligand treated cells stimulate the proliferation of untreated PC cells, as confirmed by functional experiments. According to the study's findings, AR-agonists, as well as specific AR-antagonists, regulate the cargo of exosomes. Additionally, exosomes encourage proliferation that may have an impact on the TME. The intricate relationship between exosome-mediated interaction between PC cells and AR signaling is explained by this new finding.76
In a more recent study, Wang et al. (2025) applied DIA-MS to provide important insights into the molecular progression of intraductal papillary neoplasms (IPNs), the principal precursors to pancreatic ductal adenocarcinoma (PDAC).77 Using large-scale mass-spectrometry-based proteomic and glycoproteomic profiling of resected IPMNs, IOPNs, adjacent normal ducts, cyst fluid, and PDACs, the study identified more than 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins and over 22
000 proteins and over 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 glycopeptides, revealing distinct expression patterns that differentiate normal ducts, low-grade lesions, high-grade lesions, and invasive carcinoma. Compared with normal ducts, IPMNs showed significant upregulation of proteins involved in glycan biosynthesis, mucin-type O-glycosylation, immune signaling, and extracellular matrix (ECM) remodeling, indicating that microenvironmental alteration is an early event in neoplastic development. High-grade IPMNs exhibited further enrichment of ECM-related and tumor-progression pathways, and several proteins—such as PLOD3, LGALS9 (galectin-9), IRS2, and Trop-2—emerged as potential markers that distinguish high-grade from low-grade lesions in both tissue and cyst fluid. Integrative analyses demonstrated a molecular continuum from normal duct to IPMN to PDAC, with a subset of IPMNs clustering closely with invasive cancers, underscoring their high-risk biology. Non-negative matrix factorization (NMF) further identified three molecular subtypes across IPMNs and PDACs with distinct pathway signatures, suggesting that proteomic features may augment histopathologic classification. Overall, this multi-omic work highlights key biological processes—particularly glycosylation and ECM remodeling—that drive progression in pancreatic cystic neoplasia and provides promising biomarkers for improved risk stratification and early detection.77
000 glycopeptides, revealing distinct expression patterns that differentiate normal ducts, low-grade lesions, high-grade lesions, and invasive carcinoma. Compared with normal ducts, IPMNs showed significant upregulation of proteins involved in glycan biosynthesis, mucin-type O-glycosylation, immune signaling, and extracellular matrix (ECM) remodeling, indicating that microenvironmental alteration is an early event in neoplastic development. High-grade IPMNs exhibited further enrichment of ECM-related and tumor-progression pathways, and several proteins—such as PLOD3, LGALS9 (galectin-9), IRS2, and Trop-2—emerged as potential markers that distinguish high-grade from low-grade lesions in both tissue and cyst fluid. Integrative analyses demonstrated a molecular continuum from normal duct to IPMN to PDAC, with a subset of IPMNs clustering closely with invasive cancers, underscoring their high-risk biology. Non-negative matrix factorization (NMF) further identified three molecular subtypes across IPMNs and PDACs with distinct pathway signatures, suggesting that proteomic features may augment histopathologic classification. Overall, this multi-omic work highlights key biological processes—particularly glycosylation and ECM remodeling—that drive progression in pancreatic cystic neoplasia and provides promising biomarkers for improved risk stratification and early detection.77
The fifth most common cause of cancer-related death in women is ovarian cancer (OC), a widespread malignant disease.78 The pelvic cavity contains the ovaries, which are comparatively hidden. Early OC is also difficult to diagnose and differentiates from mild ovarian tumors due to a lack of clear clinical signs and efficient diagnostic techniques. OC is detected at an advanced stage in over 70% of cases.79 Even in countries like the USA and Canada, which have a wealth of medical resources, the survival rate after five years for OC is still about 47% after diagnosis, despite a little improvement in recent decades.80 To enhance the prognosis of individuals with OC, it is essential to differentiate tumors that are benign and malignant at an early stage or prior to surgery. The need for new, efficient techniques and biomarkers for quick, low-cost, and non-invasive surveillance of high-risk groups, as well as preoperative differentiation between benign and malignant ovarian cancers is therefore critical.
A high-throughput urine proteome evaluation in DIA mode was developed by Ni et al. in 2021.81 DIN-NN software, which separates genuine signals from noise using deep neural networks, as well as innovative quantification and interference-correction techniques, were used to process MS data.
Out of the 2199 proteins that were measured, 69 were found to exhibit differential expressions in the C1 cohort's benign and malignant groups. We trained and validated a classifier that included five proteins: WFDC2, PTMA, PVRL4, FIBA, and PVRL2. The test and validation datasets had AUC values of 0.970 and 0.952, respectively, according to an evaluation of the classifier's performance.81 AUCs of 0.966, 0.947, and 0.979 were obtained in all 132 patients using the classifier, blood CA125, and serum HE4, respectively. Classifier, serum CA125, and serum HE4 were used to accurately diagnosis seven, six, and four of the eight individuals with early-stage cancer. Auxiliary diagnosis could benefit from the new classifier, which might be commercially adapted into kits for efficient non-invasive monitoring of high-risk patients.81
One comparatively uncommon form of ovarian cancer (OC) that is highly resistant to conventional chemotherapy is clear cell ovarian carcinoma (CCOC). Geographical and ethnic characteristics significantly influence the occurrence of CCOC, which varies from 5% to 25% of all OC occurrences.82 Adjuvant chemotherapy with drugs like carboplatin and paclitaxel combined with debulking surgery is the current standard of care for EOCs.83 It has been shown that ADRID1A and PIK3CA work together to promote CCOC carcinogenesis,84 and that these mutations are common in CCOC tumors.85 CCOC with different clinical outcomes has been further stratified using gene expression patterns.86,87 The underlying molecular pathways are poorly understood, and predicting its prognosis following chemotherapy is still difficult. The throughput and depth of proteomics have significantly increased with the advent of sophisticated MS-based proteomic technologies, such as tandem mass tag (TMT) labeling-based data dependent acquisition (DDA) and data-independent acquisition mass spectrometry (DIA-MS), which show the dynamics of low-abundance protein expression in human tissue and cell lines.88–91
In their comprehensive DIA-MS proteomics analysis, Yue et al. (2024) examined 35 and 28 CCOC tissue specimens from two distinct CCOC patient populations (H1 and H2, respectively). DIA proteomics identified 9409 and 8697 proteins from both groups.92 Fifteen proteins were selected using bioinformatics analysis. The extracellular matrix (ECM), mitochondrial metabolism, and DNA damage response are the primary functions of these proteins. In the H1 population and an independent confirmation population (H3 cohort), the prognostic ability of the 15 proteins was confirmed using parallel reaction monitoring (PRM)-MS. IFITM1, or interferon-inducible transmembrane protein 1, was found to be a strong predictive indicator for CCOC in both immunohistochemistry (IHC) and PRM data. The work offers a proteomic data resource for CCOC as well as one intriguing protein, IFITM1, which may be able to predict the survival and recurrence of CCOC.92
With 2.1 million new cases identified and 1.8 million deaths overall, lung cancer has the greatest incidence and mortality rate in the world, discounting non-melanoma skin cancer prevalence.93 About 21% of those with lung cancer will survive for five years, according to SEER.94 Two-thirds of lung cancer cases are discovered when the disease is advanced, and most of the tumors have spread by then, making the prognosis poor. Most treatment options are palliative.95 Early identification is necessary to enable curative treatment prior to the tumor's local or metastatic spread.96 The importance of comprehending lung cancer at the systems level has been shown by exploratory DNA- and RNA-level omics analysis of clinical cancer cohorts.97,98
Lehtio et al. (2021) conducted a comprehensive examination of the proteome landscape of non-small cell lung cancer (NSCLC), encompassing all main histological subtypes of NSCLC and almost 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins using DIA-MS. Additionally, the researchers connected LAG3 to immune evasion through FGL1 expression and HNF1A activation that is dependent on STK11 mutations.99
000 proteins using DIA-MS. Additionally, the researchers connected LAG3 to immune evasion through FGL1 expression and HNF1A activation that is dependent on STK11 mutations.99
Renal cell carcinoma (RCC) accounts for over 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths yearly, affects around 300
000 deaths yearly, affects around 300![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 people globally, and is the eighth most common cause of cancer-related deaths in the United States.100,101 More than 90% of kidney malignancies are RCCs, which are derived from the renal cortex or renal epithelial cells.102,103 The frequency of RCC has been gradually rising over the past few decades, and a wide variety of RCC subtypes have been identified. A complementary test that may identify tumor category, aggressiveness, and the necessity for surgical intervention while avoiding the necessity for biopsy has been the elusive “holy grail” for the practicing urologist, despite recent improvements and increases in utilization of renal mass biopsy.104,105 Given that most of the tumors found are low-grade, an alternative approach might enable urologists to make decisions based on the cancer grades on an individual basis. This might necessitate active observation and alternate therapeutic approaches rather than surgery.
000 people globally, and is the eighth most common cause of cancer-related deaths in the United States.100,101 More than 90% of kidney malignancies are RCCs, which are derived from the renal cortex or renal epithelial cells.102,103 The frequency of RCC has been gradually rising over the past few decades, and a wide variety of RCC subtypes have been identified. A complementary test that may identify tumor category, aggressiveness, and the necessity for surgical intervention while avoiding the necessity for biopsy has been the elusive “holy grail” for the practicing urologist, despite recent improvements and increases in utilization of renal mass biopsy.104,105 Given that most of the tumors found are low-grade, an alternative approach might enable urologists to make decisions based on the cancer grades on an individual basis. This might necessitate active observation and alternate therapeutic approaches rather than surgery.
Extracellular vesicles (EVs), which are present in biofluids like urine, plasma, and saliva, have been suggested in recent research to be a prospective source for disease diagnostics.106–108 Exosomes are nanoscale vesicles with a spherical or cup-like topology that range in size from 30 to 120 nm, while microvesicles are more irregularly shaped and often have a wide range of sizes up to about 1500 nm.109,110 EVs released by cancer cells can influence the tumor microenvironment, stimulate cell proliferation and survival, and enhance metastatic activity.85 Moreover, EVs generated by cancer cells serve as important modulators of cancer cell communication and signaling, eliciting phenotypic responses from nearby or distant normal cells that support several aspects of tumor growth.111 A valuable and prospective resource for the non-invasive detection of cancer biomarkers is the stable presence of EVs in several bodily fluids, including urine and plasma.
To profile urine EV phosphoproteomics for renal cell carcinoma (RCC) grade differentiation, Hadisurya et al. (2023) investigated the creation of a reliable DIA-MS.112 Urinary EV phosphoproteomes were identified and measured using DIA-MS for EV phosphoproteomics in 57 persons who represented chronic kidney disease, high-grade clear cell RCC, low-grade clear cell RCC, and healthy control subjects. Several well-known cancer-related pathways, including ErbB signaling, renal cell carcinoma, and actin cytoskeleton regulation, were only increased in high-grade clear cell RCC, and 2584 distinct phosphosites were measured. The results indicate that EV phosphoproteome assessment in conjunction with phosphopeptide enrichment, EV isolation, and DIA based technique provides a useful tool for upcoming clinical applications.112
In a recent study Liu et al. (2025) highlighted the growing potential of data-independent acquisition mass spectrometry (DIA-MS) as a robust platform for large-scale clinical proteomics. Recent advances in DIA-MS have enabled comprehensive and reproducible quantification of thousands of proteins across extensive patient cohorts.113 Notably, a large-scale renal cell carcinoma (RCC) proteomic study quantified over 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins from 261 tumors, revealing protein expression signatures that distinguish tumor subtypes and normal tissues. These findings underscore DIA-MS's capacity to capture tumor heterogeneity and to identify clinically actionable biomarkers that may inform disease classification and therapeutic decision-making.113
000 proteins from 261 tumors, revealing protein expression signatures that distinguish tumor subtypes and normal tissues. These findings underscore DIA-MS's capacity to capture tumor heterogeneity and to identify clinically actionable biomarkers that may inform disease classification and therapeutic decision-making.113
Compared with conventional data-dependent acquisition (DDA) and affinity-based proteomic platforms, DIA-MS provides deeper proteome coverage, improved reproducibility, and reduced missing data across samples critical attributes for translational research and clinical application. Its scalability and capacity to generate retrospective, “digital” proteome maps make it particularly suitable for longitudinal and multi-center studies. Collectively, these advances position DIA-MS as a key technology for comprehensive molecular profiling, facilitating the integration of proteomic data with genomic and clinical information to advance precision oncology and biomarker discovery.113
4.7.6.2 Biomarker discovery. Using MS-based ‘omics’ approaches to analyze biofluids is an established technique to identify markers for the identification and/or prognosis of several illnesses, including PC.114–116 Numerous multi-omics techniques have been tested in research studies for the diagnosis of PC.117,118 Matrices such as blood, urine, tissue, and others have all been studied in this manner.119 Large high-dimensional datasets are often processed using machine-learning algorithms; however, since the models are trained on idealized cohorts that clearly distinguish between cases and healthy controls, there is sometimes a concern that these studies would introduce bias.120 Determining biomarkers in PC may be made more difficult by the blood-prostate barrier, which governs the flow of chemicals between the prostate tissue and peripheral blood.121–123 This barrier may prevent disease-specific biomarkers from entering peripheral blood. Improved diagnostic tests and risk assessment for PC cases are unmet needs that could enhance care and lessen overtreatment of indolent illness.
To increase biomarker specificity, Spick et al. (2023) recently assessed the serum proteome and lipidome of 262 research participants using LC-MS.124 These individuals included those with benign prostatic hyperplasia (BPH), PC, or other healthy volunteers. A two-class machine learning algorithm distinguished PC from controls with a sensitivity of 0.82 and a specificity of 0.95; however, the addition of BPH led to a statistically significant decrease in prostate cancer specificity to 0.76, with the model incorrectly classifying half of BPH patients as PC.124 A few biomarkers that distinguish between prostate cancer and BPH were found, such as lipids that contain oleic acid and proteins involved in MAP kinase pathways; these could provide a road toward more specificity. When machine learning algorithms are trained on idealized case-control discovery studies, there is an increased risk of overstating specificity by confounding general biomarkers for inflammation or poor health with specific biomarkers for a condition.124 These results also highlight a common problem in machine learning applied to “omics data”. Two-class cohorts cannot test the specificity of identified biomarkers with respect to closely related confounders, and they may yield inaccurate assessments of specificity and other metrics, even though they may still be valuable in certain use-cases due to cost or practicality considerations.124
As transplantation technology has advanced, kidney transplantation has emerged as the best option for treating end-stage renal disease.125,126 Acute rejection after kidney allograft remains an important factor of transplant loss, despite significant advances in short-term outcomes over the past few decades. 10% of kidney transplant recipients have acute rejection during the first year, according to studies127,128 which has a major impact on the long-term survival and function of allografts. Invasive and insufficiently sensitive, current monitoring techniques cannot identify acute rejection at a comparatively early stage. Therefore, the screening of new biomarkers for the diagnosis of acute rejection is of paramount importance. Recently, DIA-based nonlabelled quantitative technology has been proposed by researchers. Improved reproducibility is demonstrated by DIA for complex protein samples, particularly low-abundance protein samples, which have been used for biomarker screening in a variety of domains, including drug-like prediction, liver disease, and tumors.129–131
To identify biomarkers linked to acute rejection, Wang et al. (2024) recently compared control participants without rejection and T cell-mediated acute rejection (TCMR) using DIA-MS.132 Kidney transplant recipients can benefit from prompt and appropriate treatment recommendations by screening and investigation of rejection-related biomarkers, which can reverse or avoid rejection and increase the grafts' long-term survival rate. Additionally, the findings will lead to new mechanistic understandings of the pathophysiology of the immune system response and a better understanding of the molecular and signal pathways associated with the rejection process.132
Between patients with STA and TCMR, 131 DEPs were found, and between patients with mild and severe TCMR, 114 DEPs were found.132 When compared to STA patients, the results indicated that TCMR patients had significantly reduced levels of fetuin-A (AHSG) and significantly greater levels of serum amyloid A protein 1 (SAA1) and insulin like growth factor binding protein 2 (IGFBP2). Additionally, there was an elevated SAA1 concentration in the severe TCMR group than in the mild TCMR group. The combination of SAA1, AHSG, and IGFBP2 demonstrated exceptional performance in the acute rejection diagnosis, as further supported by the receiver operating characteristics (ROC) analysis.132
Of the malignant tumors of the female reproductive system, OC is the most prevalent.133 75% of OC patients receive their diagnosis at an advanced stage, which results in an elevated risk of recurrence and mortality. This is because there are no early signs specific to the disease and no reliable screening techniques. About 70% of cases of OC are high-grade serous ovarian cancer (HGSOC), one of the most prevalent histologic forms of OC with aggressive characteristics.134 Thus, an essential approach to enhance survival results may be the precise prognostication or 12-month relapse-free survival (RFS) rates of HGSOC patients. Prognosis prediction or 12-month RFS rates may therefore make it easier to identify patients who need closer monitoring.
The easy accessibility of urine makes it a perfect resource for disease surveillance. Urine also contains molecules that are more stable and less complicated than those found in various other liquid samples. A recent study by Ni et al. (2024) used the MSFragger-DIA methodology in conjunction with DIA-MS analysis to identify predictive protein biomarkers for HGSOC in urine samples.135 Parallel reaction monitoring (PRM)-MS analysis and regression models built using the LASSO approach to forecast the 12-month RFS rate in patients with HGSOC were used to further validate candidate biomarkers. Parallel reaction monitoring (PRM) analysis was used to validate four potential biomarkers (ANXA1, G6PI, SPB3, and SPRR3) in both the independent validation set and the discovery set. Further RFS and Cox regression analysis validated the usefulness of these potential biomarkers as separate prognostic variables influencing RFS in HGSOC patients.135 The 12 month RFS rate was predicted using regression models, which had area under the receiver operating characteristic curve (AUC) values between 0.847 and 0.905. Urine samples from patients with HGSOC were shown to include possible prognostic biomarkers, and prediction models for the 12-month RFS rate were developed.135
Epithelial ovarian cancer (EOC) is still the deadliest gynecological disease in affluent nations, despite growing knowledge of its biology and pathogenesis.136 The 5-year survival rate is still around 50%, and over 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 women worldwide receive a diagnosis each year.137 92% of patients who have high-grade serous carcinoma (HGSC) survive after early discovery, compared to 29% in instances that are detected later. This indicates that early detection is essential to improve outcomes. Unfortunately, non-specific symptoms (such as stomach pain) are experienced by 75% of women, who do not receive a diagnosis until the disease has advanced to stage 3 or later.
000 women worldwide receive a diagnosis each year.137 92% of patients who have high-grade serous carcinoma (HGSC) survive after early discovery, compared to 29% in instances that are detected later. This indicates that early detection is essential to improve outcomes. Unfortunately, non-specific symptoms (such as stomach pain) are experienced by 75% of women, who do not receive a diagnosis until the disease has advanced to stage 3 or later.
In raw tissue samples, the most effective method for separating biomarkers from high abundance proteins (HAPs) is still unreported. Enrichment of EVs may separate possible biomarkers from HAPs or other responsible proteins138 because EVs transport bioactive cargo either to or towards a metastatic cancer microenvironment.139
Cooper et al. (2024) used a two-pronged strategy to carefully probe the proteome of EVs from two different cohorts using both DDA and DIA proteomics.140 The goal of the study was to compare the proteomic fingerprints of EVs from women with HGSC to those with benign gynecological diseases to find new biomarkers for the early identification of HGSC. Using DDA proteomics, the first cohort included 19 donor-developed spectral libraries. DIA proteomics was utilized for a similar purpose in the following cohort, which comprised 30 HGSC patients and 30 control persons.140 In HGSC, the single-pass transmembrane protein MUC1 is markedly elevated. Combining MUC1 with other biomarkers revealed it to be a significant biomarker in both cohorts. An ELISA test performed on a selected number of benign, Stage I, and Stage II plasma samples confirmed MUC1's diagnostic value in the early diagnosis of HGSC.140
According to Siegel et al. (2023), pancreatic cancer (PC) has a 12% 5-year survival rate, making it one of the deadliest cancers.141 Additionally, PC ranks third among cancer-related causes of death for both men and women. 90% of PC cases are caused by pancreatic ductal adenocarcinoma (PDAC).142 According to Rawla et al. (2019), PDAC has been found to be more prevalent in men than women, and the incidence rate rises with age for both sexes.143 Numerous new biomarkers for the identification of PDAC have emerged because of the recent expansion of high-throughput biomarker screening technology. Using MS-based plasma proteomics could help with several issues, such as finding more accurate biomarkers that could aid in early disease detection and diagnosis. Comparatively speaking to blood, tissue collection is more intrusive and difficult to obtain.
A recent study by Dubazana (2024) sought to create a standard operating protocol for a workflow based on high-throughput MS that may be applied to studies of plasma-based biomarker development.144 For label-free quantification, SWATH-MS analysis was performed after the peptides were separated using low pH reverse-phase chromatography.144 The quantity of peptides found, automation ease, and reproducibility of the procedures were used to compare them. The chosen optimum approach was tested in a pilot trial with six PDAC patients, six people with chronic pancreatitis, and six healthy participants.144 25 proteins were found to be differently expressed between the PDAC patients and the healthy group in the preliminary analysis (three upregulated and 22 downregulated). These proteins are mostly linked to carcinogenesis. The work created a standard operating protocol for handling plasma samples, which can be used in subsequent research to find possible PDAC markers.144
According to predictions, PDAC will surpass lung cancer as the second most common cause of cancer-related deaths by 2030.145,146 Because of their clinical aggression and poor response to immunotherapy or chemotherapy, PDACs are very deadly. Also, PDAC is frequently discovered at a later stage, by which time resection may no longer be effective.147
These results could be improved by detecting PDAC early, whilst surgery is still possible. To enhance PDAC early detection, Lih et al. (2024) employed quantitative glycoproteomic and proteomic methods using data-independent acquisition (DIA) mass spectrometry (MS) to perform combined identification of PDAC proteins in sera from 53 PDAC patients, of whom 13 had matched normal neighboring pancreatic tissues (NATs) and 29 had matched tumor tissues.148 Glycopeptides (i.e., deglycosylated N-linked glycopeptides) and glycoproteins from each PDAC patient's serum sample were compared to 55 nondiseased controls. Of these, 892 glycopeptides were derived from 141 glycoproteins that had elevated expression in PDAC serum samples with both matched tumor tissue and case-matched NAT and had PDAC-associated changes that were more than two standard deviations from the mean of nondiseased controls in serum. Both tissue and serum-detectable glyco-signatures suggest the potential clinical use of these glyco-features in blood tests for PDAC identification.148
4.7.6.3 Therapy response. In 2020, there were about two million new instances of colorectal cancer (CRC), making it the third most frequent kind of cancer globally. Lung cancer is the leading cause of death, with colorectal cancer coming in second.149 Most often, the diagnosis is made at an advanced stage, when there are few effective treatment choices and the prognosis is dismal. Most often diagnosed as stage II (T3/4N0M0) or stage III (T1-4 N + M0), locally advanced rectal cancer (LARC) is the most common kind of RC.150 Neoadjuvant chemotherapy and radiation therapy (nCRT) along with aggressive surgery (complete mesorectal excision), are the usual treatments for LARC. But about 20–30% of patients respond completely clinically or pathologically to nCRT, and other patients will respond poorly or improve slowly during nCRT.151,152 The identification of predictive biomarkers for nCRT and the characterization of response pathways to therapy are unmet needs in LARC. In the event of a complete clinical response (cCR), patients who respond well to nCRT would be engaged in a watch-and-wait strategy or might be contenders for a less invasive surgical procedure. This would improve their standard of living and help lower treatment expenses overall.153 Using DIA-MS, a sensitive proteomic technique, Stanojevic et al. (2024) sought to thoroughly examine the LARC proteome to identify those with positive first reactions that would be appropriate for a watch-and-wait strategy.154
In this study, 97 LARC patients received intensive chemoradiotherapy, which consists of chemotherapy and radiation.154 For proteome profiling, 20 samples were chosen from the 97 LARC that were part of this investigation that had the most varied responses to nCRT. 915 proteins in all showed differential expression among the two groups; top candidates showed high prediction ability, while specific signaling pathways were linked to response mechanisms.154 The genes SMPDL3A, PCTP, LGMN, SYNJ2, NHLRC3, GLB1, and RAB43 encoded proteins that had a strong predictive capacity for an adverse treatment outcome, whereas RPA2, SARNP, PCBP2, SF3B2, HNRNPF, RBBP4, MAGOHB, DUT, ERG28, and BUB3 represented good predictive biomarkers of a favorable treatment outcome. Promising insights that potentially improve the treatment and welfare of LARC patients are offered by the proteins found and associated biological processes.154
Advanced malignancies are treated with chemotherapy, but androgen receptor (AR)-axis-targeted advanced malignancies are treated with chemotherapy, but androgen receptor (AR)-axis-targeted (ARAT) medications that target AR are typically used to treat metastatic PC. When docetaxel fails to cure metastatic castration-resistant prostate cancer (CRPC), cabazitaxel is the next recommended treatment.155 Drug resistance is a problem since it inevitably arises and, once it does, there are no longer any viable medications for PC, even if cabazitaxel has demonstrated some efficacy.156
The plasma membrane of many kinds of cancer cells expresses the amino acid transporter L-type amino acid transporter 1 (LAT1, SLC7A5).157 In urologic malignancies, such as PC, LAT1 activates the mammalian target of rapamycin (mTOR) pathway, which is essential for the growth of cancer cells.158–160 LAT1 has a significant function in CRPC and its expression is increased in PC.158 A LAT1-specific inhibitor called JPH203 has been shown to exhibit anti-cancer properties in several carcinomas, including urologic malignancies.159–161
Rii et al. (2024) investigated how LAT1 was expressed and how JPH203, a LAT1-specific inhibitor, affected prostate cancer cells that were resistant to cabazitaxel.162In vitro, cabazitaxel-resistant organisms' capacity to proliferate, migrate, and intrude was suppressed by the administration of JPH203. Phosphorylation changes brought on by JPH203 administration demonstrated that JPH203 impacted pathways linked to the cell cycle and that it markedly decreased the kinase activity of cyclin-dependent kinases 1 and 2. Additionally, JPH203 prevented cabazitaxel-resistant cells from proliferating in vivo.135 When combined, the findings of this study point to LAT1 as a potential therapeutic target for prostate cancer that is resistant to cabazitaxel. JPH203 may therefore be a useful treatment for those with CRPC that is resistant to cabazitaxel.162
Despite improvements in treatment and patient classification, ovarian cancer remains the deadliest gynecological malignancy. Treatment-resistant illness kills most patients after a recurrence, even if they respond initially to first-line treatment. First off, there isn't a practical screening method in place right now to consistently find ovarian cancer early on, while the malignancy is still only seen in the ovary. Many patients—more than 70% are diagnosed after the cancer has progressed to stage III or IV.163 Second, most patients will relapse even if 80% of them initially respond to first-line carboplatin + paclitaxel chemotherapy. Additionally, between 20 and 40 percent of patients have no response to treatment, indicating that their cancer has an inbuilt resistance of some kind.164
There isn't a trustworthy biomarker that could be applied to screening to find early diseases in the entire population. Arentz et al. (2023) used primary ovarian cancer samples using DIA-MS to find three prognostic protein biomarkers for chemotherapy response.165 They also demonstrated each biomarker's contemporaneous regulation in a chemoresistant cell line. Patients who responded to chemotherapy had higher levels of thioredoxin (TXN) and calmodulin (CALM1) in their high-grade serous ovarian cancer (HGSOC) tissues than those who did not; immunohistochemistry verified the varied expression of the three protein markers.165 According to the findings of the via the web GENT2 database, HGSOC had lower amounts of GSN, CALM1, and TXN mRNA than fallopian tube epithelium. In a Kaplan–Meier analysis of an extensive online repository of HGSOC patient data, higher levels of TXN mRNA expression and GSN mRNA expression were associated with higher overall and progression-free survival, respectively.165 Using DIA LC-MS analysis, the differential expression of the three protein markers was further validated by contrasting parental OVCAR-5 cells with carboplatin-resistant OVCAR-5 cells. According to our study findings, GSN, CALM1, and TXN could be helpful indicators for figuring out how chemotherapy works and forecasting how it will function.165
In a related investigation, Ni et al. (2024) analyzed urine samples from individuals suffering from HGSOC using DIA-MS and PRM-MS to find possible predictive indicators.166 Finally, the discovery and independent validation datasets were used to validate four potential biomarkers: ANXA1, G6PI, SPB3, and SPRR3.166 The above proteins were found to be useful as potential prognostic biomarkers influencing RFS in patients with HGSOC, according to the findings of RFS and Cox regression studies. Remarkably, lower RFS was linked to higher urine protein concentrations.166
Among patients with ALK-positive non-small cell lung cancer (NSCLC), anaplastic lymphoma kinase-tyrosine kinase inhibitors (ALK-TKIs) have shown remarkable therapeutic benefits. The use of ALK-TKIs as first-line treatments for advanced echinoderm microtubule-associated protein like 4 (EML4)-ALK positive non-small cell lung cancer has increased within the past ten years.167 As a result of the introduction of ALK-TKIs of the first through third generations, the median progression-free survival (PFS) of patients has risen from less than a year to almost three years.168–172
Using data-independent acquisition-mass spectrometry (DIA-MS), Wang et al. (2024) characterized 737 plasma proteins from 159 pre-treatment and on-treatment plasma samples of 63 patients with ALK-positive non-small cell lung cancer.173 The data was used to create a plasma-based predictive model for ALK-TKI therapy and to categorize ALK-positive NSCLC into various subtypes with unique prognoses. A follow-up study using longitudinal plasma proteome profiling was then conducted to examine the connection between patient treatment status and changes in plasma proteins.172 Lastly, an independent cohort was used to validate the predictive efficacy of plasma biomarkers. Depending on their longitudinal variations over the course of treatment, the results show that indicators including CRP, SAA1, and SAA2 significantly correlated with treatment efficacy in addition to having predictive value for patients' prognosis at baseline. Consequently, it may be possible to better monitor resistance to medication and clinical response by sequentially detecting plasma indicators during treatment.173
5 Future directions
5.1. Enhanced acquisition strategies
Future developments in acquisition strategies are expected to greatly refine DIA's ability to resolve and quantify complex proteomes. One key advancement involves narrowing the isolation windows used during MS/MS acquisition. As mass spectrometers become faster and more sensitive, it becomes feasible to use smaller windows without sacrificing cycle speed. This enhances precursor specificity and reduces the co-isolation of multiple peptides, which is one of the main challenges in DIA. Additionally, integrating ion mobility spectrometry (such as FAIMS or trapped ion mobility spectrometry, TIMS) introduces a gas-phase separation prior to mass analysis. This adds an orthogonal dimension of selectivity, effectively decreasing signal interference and improving sensitivity. Another exciting prospect is real-time adaptive DIA, where the acquisition scheme is modified on-the-fly based on the sample's spectral complexity or known peptide features. Such innovations will allow researchers to dynamically focus on the most informative m/z regions, thereby increasing both throughput and depth of coverage.5.2. Deep learning and AI-powered data analysis
One of the most transformative shifts in DIA-MS lies in the application of artificial intelligence, particularly deep learning, to spectral data analysis. Traditional DIA workflows rely heavily on empirical spectral libraries generated from data-dependent acquisition (DDA) runs, which can be labor-intensive and limit the analysis to peptides previously observed. Deep learning models, such as Prosit and DeepMass, are now capable of accurately predicting peptide fragmentation spectra and retention times in silico, enabling the creation of comprehensive predicted libraries. This dramatically broadens the scope of peptide identification and enhances reproducibility across labs. Furthermore, machine learning algorithms are being developed to directly analyze raw DIA data, identifying peptides without any spectral library—a step toward true de novo DIA proteomics. AI is also improving noise reduction, peak detection, and chromatographic alignment, allowing for more accurate quantification even in highly complex or noisy samples. As these tools mature, they will reduce computational bottlenecks and make DIA more accessible for routine use in both research and clinical settings.5.3. Single-cell and low-input proteomics
DIA's reproducibility and comprehensiveness make it an ideal technique for low-input and single-cell proteomics, a rapidly growing frontier in life sciences. Traditional mass spectrometry methods struggle with low-abundance samples due to stochastic sampling and variability, but DIA can systematically acquire data from all peptides within a defined mass range, making it more robust under low-input conditions. Future advancements in ultra-sensitive instrumentation, such as Orbitrap Eclipse and Bruker timsTOF SCP, combined with miniaturized nanoLC systems and refined sample preparation protocols, will push the detection limits of DIA down to the level of individual cells. Techniques like carrier proteome boosting and miniaturized workflows (e.g., nanoPOTS) are expected to further enhance sensitivity and quantitative reliability. These developments will empower researchers to explore cellular heterogeneity, track rare cell populations, and understand dynamic processes such as differentiation, disease progression, and immune responses at single-cell resolution.5.4 Integration with spatial and multimodal omics
The future of DIA also involves its integration with spatially resolved omics and imaging technologies, allowing researchers to map the molecular architecture of tissues with unprecedented resolution. While DIA-MS has traditionally been performed on homogenized bulk samples, new workflows are emerging that allow the combination of DIA with spatially defined sample preparation techniques like laser capture microdissection (LCM) or liquid microjunction extraction. These methods enable proteomic profiling of discrete anatomical regions or even individual cells within tissues. Furthermore, combining DIA with imaging mass spectrometry (e.g., MALDI-MS imaging), spatial transcriptomics, or label-free spectroscopic techniques such as infrared or Raman imaging can generate rich, multimodal datasets. These datasets are invaluable for systems biology approaches aimed at understanding the interplay between proteins, lipids, metabolites, and transcripts in spatial context. This convergence is particularly impactful in neuroscience, cancer biology, and developmental biology, where understanding spatial relationships is key to interpreting function and pathology.5.5 Clinical translation and biomarker discovery
As DIA workflows become more standardized, they are poised to make significant inroads into clinical proteomics. The high reproducibility, coverage, and quantitative accuracy of DIA make it well-suited for large-scale biomarker studies and longitudinal clinical investigations. One of the challenges in clinical proteomics has been the variability between runs and instruments; however, the systematic nature of DIA minimizes such variability and facilitates comparisons across large cohorts. In the coming years, efforts will likely focus on the development of standardized, regulatory-compliant DIA protocols, robust QC pipelines, and clinically validated spectral libraries. Additionally, DIA has the potential to replace or complement targeted methods such as SRM or PRM for clinical assays, especially as newer acquisition methods enable high precision and throughput. Applications may range from early cancer detection to personalized medicine and real-time monitoring of therapeutic responses, driving the adoption of DIA-MS in hospital laboratories and translational research centers.5.6 Expanded post-translational modification (PTM) analysis
Post-translational modifications (PTMs) are critical regulators of protein function and cellular signaling, and the future of DIA includes major advancements in PTM-specific workflows. Traditional DIA has been challenged in this area due to the lower abundance and greater complexity of modified peptides. However, recent improvements in enrichment strategies (e.g., IMAC for phosphopeptides or lectin affinity for glycopeptides), combined with optimized acquisition windows and predictive spectral libraries for modified peptides, are making DIA increasingly viable for PTM analysis. Future developments will likely include DIA-specific approaches for PTM site localization, quantification, and dynamics analysis. AI-driven tools will further assist in distinguishing modified from unmodified peptides and resolving isobaric modifications. This will open the door to large-scale, quantitative studies of PTMs in diverse biological contexts, such as kinase signaling, immune modulation, and epigenetic regulation, providing a more comprehensive understanding of cellular mechanisms.Artificial intelligence (AI) and deep learning are playing transformative roles in enhancing the analytical capabilities of Data-Independent Acquisition Mass Spectrometry (DIA-MS), especially in the realms of data interpretation, peptide identification, and quantitative accuracy. Unlike traditional rule-based algorithms, deep learning models can extract high-dimensional patterns from complex DIA datasets, making them particularly powerful for deconvoluting overlapping spectra and predicting peptide fragmentation. One of the most notable examples is Prosit, a deep neural network that predicts peptide tandem mass spectra and retention times with remarkable precision, significantly improving the quality and coverage of spectral libraries used in DIA workflows.5 Building on this, newer tools such as DIA-NN integrate deep neural networks with interference correction algorithms to deliver highly accurate and ultra-fast data processing even for low-input samples.6 Moreover, transformer-based models such as DiaTrans174 have emerged as cutting-edge tools for de novo peptide sequencing directly from DIA data, surpassing conventional database search methods in both speed and reliability. These models leverage attention mechanisms to focus on relevant sequence regions, enabling accurate identification of modified peptides and unexpected sequences. AI also facilitates real-time data analysis and adaptive acquisition, optimizing instrument parameters during runs based on learned patterns.175 The integration of deep learning into DIA-MS is not only accelerating throughput and reproducibility but is also democratizing access to high-quality proteomics by reducing dependency on expert intervention and proprietary tools. As the field continues to evolve, AI-driven platforms are expected to play a central role in enabling clinical and translational proteomics by making large-scale, reproducible protein quantification more accessible and interpretable.
5.7 Open-source platforms and community resources
A key enabler of DIA's growing adoption is the development of open-source tools and community-driven resources. Software platforms such as Skyline, DIA-NN, and OpenSWATH have matured significantly and now offer robust, user-friendly environments for analyzing DIA data. These platforms support diverse workflows, from small targeted studies to large-scale proteomic surveys, and their open-source nature promotes transparency, reproducibility, and customization. Community initiatives are also generating large, curated spectral libraries—such as the Human Proteome Project and various organism-specific repositories—that enhance the accessibility of high-quality references for DIA. Looking ahead, greater standardization of data formats, quality control metrics, and benchmark datasets will continue to drive cross-laboratory harmonization and regulatory acceptance. As more researchers and clinicians contribute to and benefit from these shared resources, the collective capability of the DIA-MS community will expand significantly.5.8 Cloud-based and real-time analytics
As DIA generates increasingly large and complex datasets, there is growing interest in cloud-based platforms for data processing, storage, and sharing. Cloud computing offers scalable computational power and collaborative environments, which are particularly valuable for multi-institutional studies or clinical consortia. Platforms such as Skyline Cloud and AWS-integrated pipelines allow researchers to analyze DIA data in real-time, reducing turnaround time and democratizing access to high-performance computing resources. The future may also bring real-time analytics embedded in the acquisition process itself, enabling adaptive experimental design or immediate feedback for clinical decision-making. This vision aligns with broader trends toward digital pathology and precision medicine, where rapid and accurate molecular profiling is crucial. Such innovations will also enhance data reproducibility and transparency, as entire analysis pipelines can be versioned, shared, and reproduced across research groups.6 Conclusion
Data-Independent Acquisition Mass Spectrometry (DIA-MS) has emerged as a transformative tool in proteomics, offering unparalleled reproducibility, depth, and scalability in protein identification and quantification. By circumventing the stochastic limitations of traditional data-dependent acquisition (DDA), DIA-MS enables comprehensive and consistent proteome coverage, making it particularly valuable in clinical, biomarker discovery, and systems biology studies.Recent advances in computational algorithms, spectral libraries, and machine learning-based data deconvolution have dramatically expanded the capabilities of DIA workflows, facilitating precise quantification even in complex and low-abundance samples. Furthermore, its compatibility with label-free and isobaric labeling strategies has enhanced throughput while maintaining quantitative accuracy.
As the field moves toward single-cell and spatially resolved proteomics, DIA-MS stands to play a central role, especially when integrated with emerging frontiers such as high-field asymmetric waveform ion mobility spectrometry (FAIMS), deep learning models, and high-resolution MS platforms. However, challenges remain, particularly in standardizing data analysis pipelines and ensuring interoperability across labs and platforms.
Looking ahead, the continued refinement of DIA-MS techniques—alongside more robust reference databases, real-time acquisition strategies, and user-friendly bioinformatics tools—will accelerate the adoption of DIA-MS as a gold standard in next-generation proteomic research.
Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included, and no new data were generated or analyzed as part of this review.References
- B. Zhang, J. Wang, X. Wang, J. Zhu, Q. Liu, Z. Shi, M. C. Chambers, L. J. Zimmerman, K. F. Shaddox, S. Kim and S. R. Davies, Proteogenomic characterization of human colon and rectal cancer, Nature, 2014, 513(7518), 382–387 Search PubMed.
- A. Michalski, J. Cox and M. Mann, More than 100,000 detectable peptide species elute in single shotgun proteomics runs but the majority is inaccessible to data-dependent LC– MS/MS, J. Proteome Res., 2011, 10(4), 1785–1793 CrossRef CAS PubMed.
- L. C. Gillet, P. Navarro, S. Tate, H. Röst, N. Selevsek, L. Reiter, R. Bonner and R. Aebersold, Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis, Mol. Cell. Proteomics, 2012, 11(6), 111–16717 CrossRef PubMed.
- F. Meier, A. D. Brunner, S. Koch, H. Koch, M. Lubeck, M. Krause, N. Goedecke, J. Decker, T. Kosinski, M. A. Park and N. Bache, Online parallel accumulation–serial fragmentation (PASEF) with a novel trapped ion mobility mass spectrometer, Mol. Cell. Proteomics, 2018, 17(12), 2534–2545 Search PubMed.
- S. Gessulat, T. Schmidt, D. P. Zolg, P. Samaras, K. Schnatbaum, J. Zerweck, T. Knaute, J. Rechenberger, B. Delanghe, A. Huhmer and U. Reimer, Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning, Nat. Methods, 2019, 16(6), 509–518 CrossRef CAS PubMed.
- V. Demichev, C. B. Messner, S. I. Vernardis, K. S. Lilley and M. Ralser, DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput, Nat. Methods, 2020, 17(1), 41–44 Search PubMed.
- N. J. Edwards, M. Oberti, R. R. Thangudu, S. Cai, P. B. McGarvey, S. Jacob, S. Madhavan and K. A. Ketchum, The CPTAC data portal: a resource for cancer proteomics research, J. Proteome Res., 2015, 14(6), 2707–2713 Search PubMed.
- P. Mertins, D. R. Mani, K. V. Ruggles, M. A. Gillette, K. R. Clauser, P. Wang, X. Wang, J. W. Qiao, S. Cao, F. Petralia and E. Kawaler, Proteogenomics connects somatic mutations to signalling in breast cancer, Nature, 2016, 534(7605), 55–62 CrossRef CAS PubMed.
- T. Guo and R. Aebersold, Recent advances of data-independent acquisition mass spectrometry-based proteomics, Proteomics, 2023, 23(7–8), 2200011 Search PubMed.
- R. Bruderer, O. M. Bernhardt, T. Gandhi, S. M. Miladinović, L. Y. Cheng, S. Messner, T. Ehrenberger, V. Zanotelli, Y. Butscheid, C. Escher and O. Vitek, Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues, Mol. Cell. Proteomics, 2015, 14(5), 1400–1410 CrossRef CAS PubMed.
- B. Wang, Q. Zhang, L. Wu, C. Deng, M. Luo, Y. Xie, G. Wu, W. Chen, Y. Sheng, P. Zhu and G. Qin, Data-independent acquisition-based mass spectrometry (DIA-MS) for quantitative analysis of patients with chronic hepatitis B, Proteome Sci., 2023, 21(1), 9 CrossRef CAS PubMed.
- B. S. Kumar, Advances and applications of mass spectrometry imaging in neuroscience: An overview, Mass Spectrom. Lett., 2023, 14(3), 57–78 CAS.
- J. M. Spraggins, D. G. Rizzo, J. L. Moore, M. J. Noto, E. P. Skaar and R. M. Caprioli, Next-generation technologies for spatial proteomics: integrating ultra-high speed MALDI-TOF and high mass resolution MALDI FTICR imaging mass spectrometry for protein analysis, Proteomics, 2016, 16(11–12), 1678–1689 CrossRef CAS PubMed.
- B. S. Kumar, Ambient Mass Spectrometry in Imaging and Profiling of Single Cells: An Overview, Mass Spectrom. Lett., 2023, 14(4), 121–140 CAS.
- R. Bruderer, O. M. Bernhardt, T. Gandhi, S. M. Miladinović, L. Y. Cheng, S. Messner, T. Ehrenberger, V. Zanotelli, Y. Butscheid, C. Escher and O. Vitek, Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues, Mol. Cell. Proteomics, 2015, 14(5), 1400–1410 Search PubMed.
- H. L. Röst, G. Rosenberger, P. Navarro, L. Gillet, S. M. Miladinović, O. T. Schubert, W. Wolski, B. C. Collins, J. Malmström, L. Malmström and R. Aebersold, OpenSWATH enables automated, targeted analysis of data-independent acquisition MS data, Nat. Biotechnol., 2014, 32(3), 219–223 CrossRef.
- M. Mauriat, J. C. Leplé, S. Claverol, J. Bartholomé, L. Negroni, N. Richet, C. Lalanne, M. Bonneu, C. Coutand and C. Plomion, Quantitative proteomic and phosphoproteomic approaches for deciphering the signaling pathway for tension wood formation in poplar, J. Proteome Res., 2015, 14(8), 3188–3203 CrossRef CAS PubMed.
- R. A. Scheltema, J. P. Hauschild, O. Lange, D. Hornburg, E. Denisov, E. Damoc, A. Kuehn, A. Makarov and M. Mann, The Q Exactive HF, a Benchtop mass spectrometer with a pre-filter, high-performance quadrupole and an ultra-high-field Orbitrap analyzer, Mol. Cell. Proteomics, 2014, 13(12), 3698–3708 Search PubMed.
- F. Meier, A. D. Brunner, M. Frank, A. Ha, I. Bludau, E. Voytik, S. Kaspar-Schoenefeld, M. Lubeck, O. Raether, N. Bache and R. Aebersold, diaPASEF: parallel accumulation–serial fragmentation combined with data-independent acquisition, Nat. Methods, 2020, 17(12), 1229–1236 CrossRef CAS PubMed.
- M. Dou, G. Clair, C. F. Tsai, K. Xu, W. B. Chrisler, R. L. Sontag, R. Zhao, R. J. Moore, T. Liu, L. Pasa-Tolic and R. D. Smith, High-throughput single cell proteomics enabled by multiplex isobaric labeling in a nanodroplet sample preparation platform, Anal. Chem., 2019, 91(20), 13119–13127 CrossRef CAS.
- K. Weke, S. Kote, J. Faktor, S. Al Shboul, N. Uwugiaren, P. M. Brennan, D. R. Goodlett, T. R. Hupp and I. Dapic, DIA-MS proteome analysis of formalin-fixed paraffin-embedded glioblastoma tissues, Anal. Chim. Acta, 2022, 1204, 339695 CrossRef CAS PubMed.
- Y. J. Kim, S. M. Sweet, J. D. Egertson, A. J. Sedgewick, S. Woo, W. L. Liao, G. E. Merrihew, B. C. Searle, C. Vaske, R. Heaton and M. J. MacCoss, Data-independent acquisition mass spectrometry to quantify protein levels in FFPE tumor biopsies for molecular diagnostics, J. Proteome Res., 2018, 18(1), 426–435 Search PubMed.
- CPTAC, https://gdc.cancer.gov/about-gdc/contributed-genomic-data-cancer-research/clinical-proteomic-tumor-analysis-consortium-cptac?utm_source.
- S. N. Thomas, B. Friedrich, M. Schnaubelt, D. W. Chan, H. Zhang and R. Aebersold, Orthogonal proteomic platforms and their implications for the stable classification of high-grade serous ovarian cancer subtypes, Iscience, 2020, 23(6), 1–17 CrossRef PubMed.
- N. Bache, P. E. Geyer, D. B. Bekker-Jensen, O. Hoerning, L. Falkenby, P. V. Treit, S. Doll, I. Paron, J. B. Müller, F. Meier and J. V. Olsen, A novel LC system embeds analytes in pre-formed gradients for rapid, ultra-robust proteomics, Mol. Cell. Proteomics, 2018, 17(11), 2284–2296 Search PubMed.
- F. Yu, G. C. Teo, A. T. Kong, K. Fröhlich, G. X. Li, V. Demichev and A. I. Nesvizhskii, Analysis of DIA proteomics data using MSFragger-DIA and FragPipe computational platform, Nat. Commun., 2023, 14(1), 4154 CrossRef CAS PubMed.
- R. Lou and W. Shui, Acquisition and analysis of DIA-based proteomic data: A comprehensive survey in 2023, Mol. Cell. Proteomics, 2024, 23(2), 100712 CrossRef CAS.
- S. Sidoli, R. Fujiwara and B. A. Garcia, Multiplexed data independent acquisition (MSX-DIA) applied by high resolution mass spectrometry improves quantification quality for the analysis of histone peptides, Proteomics, 2016, 16(15–16), 2095–2105 CrossRef CAS PubMed.
- B. Salovska, W. Li, O. M. Bernhardt, P. L. Germain, T. Gandhi, L. Reiter and Y. Liu, A comprehensive and robust multiplex-DIA workflow profiles protein turnover regulations associated with cisplatin resistance, bioRxiv, preprint, 2024, DOI:10.1101/2024.10.28.620709.
- A. Martínez-Val, K. Fort, C. Koenig, L. Van der Hoeven, G. Franciosa, T. Moehring, Y. Ishihama, Y. J. Chen, A. Makarov, Y. Xuan and J. V. Olsen, Hybrid-DIA: intelligent data acquisition integrates targeted and discovery proteomics to analyze phospho-signaling in single spheroids, Nat. Commun., 2023, 14(1), 3599 CrossRef.
- S. Gessulat, T. Schmidt, D. P. Zolg, P. Samaras, K. Schnatbaum, J. Zerweck, T. Knaute, J. Rechenberger, B. Delanghe, A. Huhmer and U. Reimer, Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning, Nat. Methods, 2019, 16(6), 509–518 CrossRef CAS PubMed.
- V. Demichev, C. B. Messner, S. I. Vernardis, K. S. Lilley and M. Ralser, DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput, Nat. Methods, 2020, 17(1), 41–44 CrossRef CAS PubMed.
- K. Fröhlich, E. Brombacher, M. Fahrner, D. Vogele, L. Kook, N. Pinter, P. Bronsert, S. Timme-Bronsert, A. Schmidt, K. Bärenfaller and C. Kreutz, Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity, Nat. Commun., 2022, 13(1), 2622 CrossRef.
- J. Klein, H. Lam, T. D. Mak, W. Bittremieux, Y. Perez-Riverol, R. Gabriels, J. Shofstahl, H. Hecht, P. A. Binz, S. Kawano and T. Van Den Bossche, The Proteomics Standards Initiative standardized formats for spectral libraries and fragment ion peak annotations: mzSpecLib and mzPAF, Anal. Chem., 2024, 96(46), 18491–18501 Search PubMed.
- M. K. Midha, D. S. Campbell, C. Kapil, U. Kusebauch, M. R. Hoopmann, S. L. Bader and R. L. Moritz, DIALib-QC an assessment tool for spectral libraries in data-independent acquisition proteomics, Nat. Commun., 2020, 11(1), 5251 CrossRef CAS.
- P. Ran, Y. Wang, K. Li, S. He, S. Tan, J. Lv, J. Zhu, S. Tang, J. Feng, Z. Qin and Y. Li, STAVER: a standardized benchmark dataset-based algorithm for effective variation reduction in large-scale DIA-MS data, Briefings Bioinf., 2024, 25(6), bbae553 CrossRef CAS PubMed.
- S. J. Rice and C. P. Belani, Optimizing data-independent acquisition (DIA) spectral library workflows for plasma proteomics studies, Proteomics, 2022, 22(17), 2200125 CrossRef CAS.
- J. Klein, H. Lam, T. D. Mak, W. Bittremieux, Y. Perez-Riverol, R. Gabriels, J. Shofstahl, H. Hecht, P. A. Binz, S. Kawano and T. Van Den Bossche, The Proteomics Standards Initiative standardized formats for spectral libraries and fragment ion peak annotations: mzSpecLib and mzPAF, Anal. Chem., 2024, 96(46), 18491–18501 CrossRef CAS PubMed.
- K. Fröhlich, M. Fahrner, E. Brombacher, A. Seredynska, M. Maldacker, C. Kreutz, A. Schmidt and O. Schilling, Data-independent acquisition: A Milestone and prospect in clinical mass spectrometry–based proteomics, Mol. Cell. Proteomics, 2024, 23(8), 100800 CrossRef PubMed.
- K. Bezstarosti, L. Van der Wal, W. A. Doff and J. A. Demmers, Parallel reaction monitoring targeted mass spectrometry as a fast and sensitive alternative to antibody-based protein detection, Front. Anal. Sci., 2024, 4, 1397810 CrossRef.
- G. H. Eldjarn, E. Ferkingstad, S. H. Lund, H. Helgason, O. T. Magnusson, K. Gunnarsdottir, T. A. Olafsdottir, B. V. Halldorsson, P. I. Olason, F. Zink and S. A. Gudjonsson, Large-scale plasma proteomics comparisons through genetics and disease associations, Nature, 2023, 622(7982), 348–358 CrossRef CAS PubMed.
- M. Thielert, E. C. Itang, C. Ammar, F. A. Rosenberger, I. Bludau, L. Schweizer, T. M. Nordmann, P. Skowronek, M. Wahle, W. F. Zeng and X. X. Zhou, Robust dimethyl-based multiplex-DIA doubles single-cell proteome depth via a reference channel, Mol. Syst. Biol., 2023, 19(9), e11503 CrossRef CAS PubMed.
- T. Truong and R. T. Kelly, What's new in single-cell proteomics, Curr. Opin. Biotechnol., 2024, 86, 103077 CrossRef CAS PubMed.
- V. Demichev, L. Szyrwiel, F. Yu, G. C. Teo, G. Rosenberger, A. Niewienda, D. Ludwig, J. Decker, S. Kaspar-Schoenefeld, K. S. Lilley and M. Mülleder, dia-PASEF data analysis using FragPipe and DIA-NN for deep proteomics of low sample amounts, Nat. Commun., 2022, 13(1), 3944 CrossRef CAS PubMed.
- Thermo Fisher, https://www.thermofisher.com/order/catalog/product/A44520?utm_source.
- B. C. Searle, L. K. Pino, J. D. Egertson, Y. S. Ting, R. T. Lawrence, B. X. MacLean, J. Villén and M. J. MacCoss, Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry, Nat. Commun., 2018, 9(1), 5128 CrossRef PubMed.
- https://www.evosep.com/project/dia-nn-software-identifies-more-than-5000-proteins/?utm_source .
- K. Gu, H. Kumabe, T. Yamamoto, N. Tashiro, T. Masuda, S. Ito and S. Ohtsuki, Improving proteomic identification using narrow isolation windows with Zeno SWATH data-independent acquisition, J. Proteome Res., 2024, 23(8), 3484–3495 CrossRef CAS.
- K. Frohlich, R. Furrer, C. Schori, C. Handschin and A. Schmidt, Robust, precise, and deep proteome profiling using a small mass range and narrow window data-independent-acquisition scheme, J. Proteome Res., 2024, 23(3), 1028–1038 CrossRef CAS PubMed.
- T. Tsukidate, A. Q. Stiving, S. Rivera, A. Sahoo, S. Madabhushi and X. Li, DiaPASEF Enables High-Throughput Proteomic Analysis of Host Cell Proteins for Biopharmaceutical Process Development, Anal. Chem., 2024, 96(32), 12999–13006 CrossRef CAS PubMed.
- R. B. Kitata, J. C. Yang and Y. J. Chen, Advances in data-independent acquisition mass spectrometry towards comprehensive digital proteome landscape, Mass Spectrom. Rev., 2023, 42(6), 2324–2348 CrossRef CAS PubMed.
- L. Krasny and P. H. Huang, Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology, Mol. Omics, 2021, 17(1), 29–42 RSC.
- C. Ludwig, L. Gillet, G. Rosenberger, S. Amon, B. C. Collins and R. Aebersold, Data-independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial, Mol. Syst. Biol., 2018, 14(8), e8126 CrossRef PubMed.
- F. Zhang, W. Ge, L. Huang, D. Li, L. Liu, Z. Dong, L. Xu, X. Ding, C. Zhang, Y. Sun and J. Gao, A comparative analysis of data analysis tools for data-independent acquisition mass spectrometry, Mol. Cell. Proteomics, 2023, 22(9), 1–14 CrossRef.
- P. Xiang, A. Liyu, Y. Kwon, D. Hu, S. M. Williams, D. Velickovic, L. M. Markillie, W. B. Chrisler, L. Pasa-Tolic and Y. Zhu, Spatial proteomics toward subcellular resolution by coupling deep ultraviolet laser ablation with nanodroplet sample preparation, ACS Meas. Sci. Au, 2023, 3(6), 459–468 CrossRef CAS PubMed.
- N. Eliyatkın, E. Yalçın, B. Zengel, S. Aktaş and E. Vardar, Molecular classification of breast carcinoma: from traditional, old-fashioned way to a new age, and a new way, J. Breast Health, 2015, 11(2), 59 CrossRef.
- B. S. Kumar, N and O-glycosylation Studies with Ion Mobility Mass Spectrometry (IM–MS): an Overview, Mass Spectrom. Lett., 2024, 15(3), 121–140 CAS.
- S. Zhu, W. Yu, X. Yang, C. Wu and F. Cheng, Traditional classification and novel subtyping systems for bladder cancer, Front. Oncol., 2020, 10, 102 CrossRef PubMed.
- J. Ferlay, M. Colombet, I. Soerjomataram, C. Mathers, D. M. Parkin, M. Piñeros, A. Znaor and F. Bray, Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods, Int. J. Cancer, 2019, 144(8), 1941–1953 CrossRef CAS PubMed.
- V. J. Gnanapragasam, T. Barrett, C. Massie, S. Pacey and A. Warren, Using prognosis to guide early detection and treatment selection in non-metastatic prostate cancer, BJU Int., 2019, 123(4), 562–563 CrossRef PubMed.
- B. S. Kumar, Desorption electrospray ionization mass spectrometry imaging (DESI-MSI) in disease diagnosis: an overview, Anal. Methods, 2023, 15(31), 3768–3784 RSC.
- A. Jemal, J. Ma, R. Siegel, S. Fedewa, O. Brawley and E. M. Ward, Prostate cancer incidence rates 2 years after the US Preventive Services Task Force recommendations against screening, JAMA Oncol., 2016, 2(12), 1657–1660 CrossRef PubMed.
- J. C. Hu, P. Nguyen, J. Mao, J. Halpern, J. Shoag, J. D. Wright and A. Sedrakyan, Increase in prostate cancer distant metastases at diagnosis in the United States, JAMA Oncol., 2017, 3(5), 705–707 CrossRef PubMed.
- A. Jemal, M. B. Culp, J. Ma, F. Islami and S. A. Fedewa, Prostate cancer incidence 5 years after US Preventive Services Task Force recommendations against screening, JNCI (J. Natl. Cancer Inst.), 2021, 113(1), 64–71 CrossRef.
- K. A. Houston, J. King, J. Li and A. Jemal, Trends in prostate cancer incidence rates and prevalence of prostate specific antigen screening by socioeconomic status and regions in the United States, 2004 to 2013, J. Urol., 2018, 199(3), 676–681 CrossRef PubMed.
- Q. K. Li, L. Chen, M. H. Ao, J. H. Chiu, Z. Zhang, H. Zhang and D. W. Chan, Serum fucosylated prostate-specific antigen (PSA) improves the differentiation of aggressive from non-aggressive prostate cancers, Theranostics, 2015, 5(3), 267 CrossRef CAS PubMed.
- Q. K. Li, J. Chen, Y. Hu, N. Höti, T. S. Lih, S. N. Thomas, L. Chen, S. Roy, A. Meeker, P. Shah and L. Chen, Proteomic characterization of primary and metastatic prostate cancer reveals reduced proteinase activity in aggressive tumors, Sci. Rep., 2021, 11(1), 18936 CrossRef CAS PubMed.
- C. Wang, G. Liu, Y. Liu, Z. Yang, W. Xin, M. Wang, Y. Li, L. Yang, H. Mu and C. Zhou, Novel serum proteomic biomarkers for early diagnosis and aggressive grade identification of prostate cancer, Front. Oncol., 2022, 12, 1004015 CrossRef CAS PubMed.
- T. Akoto and S. Saini, Role of exosomes in prostate cancer metastasis, Int. J. Mol. Sci., 2021, 22(7), 3528 CrossRef CAS PubMed.
- N. Ariotti, Y. Wu, S. Okano, Y. Gambin, J. Follett, J. Rae, C. Ferguson, R. D. Teasdale, K. Alexandrov, F. A. Meunier and M. M. Hill, An inverted CAV1 (caveolin 1) topology defines novel autophagy-dependent exosome secretion from prostate cancer cells, Autophagy, 2021, 17(9), 2200–2216 CrossRef CAS PubMed.
- Y. Sun, Tumor microenvironment and cancer therapy resistance, Cancer Lett., 2016, 380(1), 205–215 CrossRef CAS PubMed.
- M. Takasugi, R. Okada, A. Takahashi, D. Virya Chen, S. Watanabe and E. Hara, Small extracellular vesicles secreted from senescent cells promote cancer cell proliferation through EphA2, Nat. Commun., 2017, 8(1), 15729 CrossRef CAS PubMed.
- E. Chiaradia, B. Tancini, C. Emiliani, F. Delo, R. M. Pellegrino, A. Tognoloni, L. Urbanelli and S. Buratta, Extracellular vesicles under oxidative stress conditions: biological properties and physiological roles, Cells, 2021, 10(7), 1763 CrossRef CAS PubMed.
- S. A. Shaban, J. Rezaie and V. Nejati, Exosomes derived from senescent endothelial cells contain distinct pro-angiogenic miRNAs and proteins, Cardiovasc. Toxicol., 2022, 22(6), 592–601 CrossRef CAS PubMed.
- B. D. Lehmann, M. S. Paine, A. M. Brooks, J. A. McCubrey, R. H. Renegar, R. Wang and D. M. Terrian, Senescence-associated exosome release from human prostate cancer cells, Cancer Res., 2008, 68(19), 7864–7871 CrossRef CAS.
- R. G. Atri, M. Kokal-Ribaudo, M. Heidari Horestani, T. Pungsrinont and A. Baniahmad, The protein composition of exosomes released by prostate cancer cells is distinctly regulated by androgen receptor-antagonists and-agonist to stimulate growth of target cells, Cell Commun. Signal., 2024, 22(1), 219 CrossRef PubMed.
- Y. Wang, T. S. Lih, J. W. Lee, T. Ohtsuka, Y. Hozaka, M. Mino-Kenudson, N. V. Adsay, C. Luchini, A. Scarpa, A. V. Maker and G. E. Kim, Multi-omic profiling of intraductal papillary neoplasms of the pancreas reveals distinct patterns and potential markers of progression, Cancer Cell, 2025, 2156–2170 CrossRef CAS.
- R. L. Siegel, K. D. Miller, H. E. Fuchs and A. Jemal, Cancer statistics 2022, Ca-Cancer J. Clin., 2022, 72(1), 7–33 CrossRef PubMed.
- V. Dochez, H. Caillon, E. Vaucel, J. Dimet, N. Winer and G. Ducarme, Biomarkers and algorithms for diagnosis of ovarian cancer: CA125, HE4, RMI and ROMA, a review, J. Ovarian Res., 2019, 12(1), 28 CrossRef PubMed.
- B. S. Kumar, Recent developments and application of mass spectrometry imaging in N-glycosylation studies: an overview, Mass Spectrom., 2024, 13(1), A0142 CrossRef CAS PubMed.
- M. Ni, J. Zhou, Z. Zhu, J. Yuan, W. Gong, J. Zhu, Z. Zheng and H. Zhao, A novel classifier based on urinary proteomics for distinguishing between benign and malignant ovarian tumors, Front. Cell Dev. Biol., 2021, 9, 712196 CrossRef PubMed.
- A. Gadducci, F. Multinu, S. Cosio, S. Carinelli, M. Ghioni and G. D. Aletti, Clear cell carcinoma of the ovary: Epidemiology, pathological and biological features, treatment options and clinical outcomes, Gynecol. Oncol., 2021, 162(3), 741–750 CrossRef PubMed.
- S. Lheureux, C. Gourley, I. Vergote and A. M. Oza, Epithelial ovarian cancer, Lancet, 2019, 393(10177), 1240–1253 CrossRef.
- B. S. Kumar, Recent advances and applications of ambient mass spectrometry imaging in cancer research: An overview, Mass Spectrom., 2023, 12(1), A0129 CrossRef CAS PubMed.
- S. Zhu, C. Zhang, D. Cao, J. Bai, S. Yu, J. Chen, J. Wang, T. Ren, J. Yang, M. Yu and X. Xiao, Genomic and TCR profiling data reveal the distinct molecular traits in epithelial ovarian cancer histotypes, Oncogene, 2022, 41(22), 3093–3103 CrossRef CAS PubMed.
- T. Z. Tan, J. Ye, C. V. Yee, D. Lim, N. Y. Ngoi, D. S. Tan and R. Y. Huang, Analysis of gene expression signatures identifies prognostic and functionally distinct ovarian clear cell carcinoma subtypes, EBioMedicine, 2019, 50, 203–210 CrossRef CAS PubMed.
- V. Heong, T. Z. Tan, M. Miwa, J. Ye, D. Lim, C. S. Herrington, Y. Iida, M. Yano, M. Yasuda, N. Y. Ngoi and S. J. Wong, A multi-ethnic analysis of immune-related gene expression signatures in patients with ovarian clear cell carcinoma, J. Pathol., 2021, 255(3), 285–295 CrossRef CAS PubMed.
- Z. Wang, K. Yu, H. Tan, Z. Wu, J. H. Cho, X. Han, H. Sun, T. G. Beach and J. Peng, 27-Plex tandem mass tag mass spectrometry for profiling brain proteome in Alzheimer's disease, Anal. Chem., 2020, 92(10), 7162–7170 CrossRef CAS PubMed.
- D. J. Clark, S. M. Dhanasekaran, F. Petralia, J. Pan, X. Song, Y. Hu, F. da Veiga Leprevost, B. Reva, T. S. Lih, H. Y. Chang and W. Ma, Integrated proteogenomic characterization of clear cell renal cell carcinoma, Cell, 2019, 179(4), 964–983 CrossRef CAS PubMed.
- H. Zhang, T. Liu, Z. Zhang, S. H. Payne, B. Zhang, J. E. McDermott, J. Y. Zhou, V. A. Petyuk, L. Chen, D. Ray and S. Sun, Integrated proteogenomic characterization of human high-grade serous ovarian cancer, Cell, 2016, 166(3), 755–765 CrossRef CAS PubMed.
- Y. Kawashima, H. Nagai, R. Konno, M. Ishikawa, D. Nakajima, H. Sato, R. Nakamura, T. Furuyashiki and O. Ohara, Single-shot 10K proteome approach: over 10,000 protein identifications by data-independent acquisition-based single-shot proteomics with ion mobility spectrometry, J. Proteome Res., 2022, 21(6), 1418–1427 CrossRef CAS PubMed.
- L. Yue, T. Gong, W. Jiang, L. Qian, W. Gong, Y. Sun, X. Cai, H. Xu, F. Liu, H. Wang and S. Li, Proteomic profiling of ovarian clear cell carcinomas identifies prognostic biomarkers for chemotherapy, Proteomics, 2024, 24(6), 2300242 CrossRef CAS PubMed.
- F. Bray, J. Ferlay, I. Soerjomataram, R. L. Siegel, L. A. Torre and A. Jemal, Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries, Ca-Cancer J. Clin., 2018, 68(6), 394–424 CrossRef PubMed.
- T. Lu, X. Yang, Y. Huang, M. Zhao, M. Li, K. Ma, J. Yin, C. Zhan and Q. Wang, Trends in the incidence, treatment, and survival of patients with lung cancer in the last four decades, Cancer Manag. Res., 2019, 21, 943–953 CrossRef PubMed.
- N. Hasan, R. Kumar and M. S. Kavuru, Lung cancer screening beyond low-dose computed tomography: the role of novel biomarkers, Lung, 2014, 192(5), 639–648 CrossRef PubMed.
- A. K. Pathak, M. Bhutani, S. Kumar, A. Mohan and R. Guleria, Circulating cell-free DNA in plasma/serum of lung cancer patients as a potential screening and prognostic tool, Clin. Chem., 2006, 52(10), 1833–1842 CAS.
- Cancer Genome Atlas Research Network, Comprehensive genomic characterization of squamous cell lung cancers, Nature, 2012, 489, 519 CrossRef PubMed.
- Cancer Genome Atlas Research Network, Comprehensive genomic characterization of squamous cell lung cancers, Nature, 2014, vol. 489(7417), 519 CrossRef PubMed.
- J. Lehtiö, T. Arslan, I. Siavelis, Y. Pan, F. Socciarelli, O. Berkovska, H. M. Umer, G. Mermelekas, M. Pirmoradian, M. Jönsson and H. Brunnström, Proteogenomics of non-small cell lung cancer reveals molecular subtypes associated with specific therapeutic targets and immune-evasion mechanisms, Nat. Cancer, 2021, 2(11), 1224–1242 CrossRef PubMed.
- K. Attalla, S. Weng, M. H. Voss and A. A. Hakimi, Epidemiology, risk assessment, and biomarkers for patients with advanced renal cell carcinoma, Urol. Clin., 2020, 47(3), 293–303 CrossRef PubMed.
- R. L. Siegel, K. D. Miller and A. Jemal, Cancer statistics, 2019, Ca-Cancer J. Clin., 2019, 69(1), 7–34 CrossRef PubMed.
- S. A. Padala and A. Kallam, Clear Cell Renal Carcinoma (Archived). 2023 May 22. in StatPearls [Internet]. Treasure Island (FL), StatPearls Publishing, 2025 Search PubMed.
- J. J. Hsieh, M. P. Purdue, S. Signoretti, C. Swanton, L. Albiges, M. Schmidinger, D. Y. Heng, J. Larkin and V. Ficarra, Renal cell carcinoma, Nat. Rev. Dis. Primers, 2017, 3(1), 1–9 Search PubMed.
- B. Ljungberg, K. Bensalah, S. Canfield, S. Dabestani, F. Hofmann, M. Hora, M. A. Kuczyk, T. Lam, L. Marconi, A. S. Merseburger and P. Mulders, EAU guidelines on renal cell carcinoma: 2014 update, Eur. Urol., 2015, 67(5), 913–924 CrossRef PubMed.
- K. Sircar, P. Rao, E. Jonasch, F. A. Monzon and P. Tamboli, Contemporary approach to diagnosis and classification of renal cell carcinoma with mixed histologic features, Chin. J. Cancer, 2013, 32(6), 303 CrossRef CAS PubMed.
- J. Nilsson, J. Skog, A. Nordstrand, V. Baranov, L. Mincheva-Nilsson, X. O. Breakefield and A. Widmark, Prostate cancer-derived urine exosomes: a novel approach to biomarkers for prostate cancer, Br. J. Cancer, 2009, 100(10), 1603–1607 CrossRef CAS PubMed.
- S. A. Melo, L. B. Luecke, C. Kahlert, A. F. Fernandez, S. T. Gammon, J. Kaye, V. S. LeBleu, E. A. Mittendorf, J. Weitz, N. Rahbari and C. Reissfelder, Glypican-1 identifies cancer exosomes and detects early pancreatic cancer, Nature, 2015, 523(7559), 177–182 CrossRef CAS PubMed.
- Y. Sun, C. Huo, Z. Qiao, Z. Shang, A. Uzzaman, S. Liu, X. Jiang, L. Y. Fan, L. Ji, X. Guan and C. X. Cao, Comparative proteomic analysis of exosomes and microvesicles in human saliva for lung cancer, J. Proteome Res., 2018, 17(3), 1101–1107 CrossRef CAS PubMed.
- R. Xu, D. W. Greening, H. J. Zhu, N. Takahashi and R. J. Simpson, Extracellular vesicle isolation and characterization: toward clinical application, J. Clin. Investig., 2016, 126(4), 1152–1162 CrossRef PubMed.
- H. Kalra, G. P. Drummen and S. Mathivanan, Focus on extracellular vesicles: introducing the next small big thing, Int. J. Mol. Sci., 2016, 17(2), 170 CrossRef PubMed.
- W. H. Chang, R. A. Cerione and M. A. Antonyak, Extracellular Vesicles and Their Roles in Cancer Progression, in Cancer Cell Signaling, ed. M. Robles-Flores, Methods in Molecular Biology, Humana, New York, NY, 2021, vol. 2174, DOI:10.1007/978-1-0716-0759-6_10.
- M. Hadisurya, Z. C. Lee, Z. Luo, G. Zhang, Y. Ding, H. Zhang, A. B. Iliuk, R. Pili, R. S. Boris and W. A. Tao, Data-independent acquisition phosphoproteomics of urinary extracellular vesicles enables renal cell carcinoma grade differentiation, Mol. Cell. Proteomics, 2023, 22(5), 100536 CrossRef CAS PubMed.
- H. Liu, Z. Ma, T. M. Lih, L. Chen, Y. Hu, Y. Wang, Z. Sun, Y. Huang, Y. Xu and H. Zhang, New Method Enhanced Extraction of Protein Signatures of Renal Cell Carcinoma from Proteomics Data, J. Proteome Res., 2025, 5710–5726 CrossRef CAS PubMed.
- A. R. Lima, M. de Lourdes Bastos, M. Carvalho and P. G. de Pinho, Biomarker discovery in human prostate cancer: an update in metabolomics studies, Transl. Oncol., 2016, 9(4), 357–370 CrossRef PubMed.
- A. Muazzam, M. Spick, O. N. Cexus, B. Geary, F. Azhar, H. Pandha, A. Michael, R. Reed, S. Lennon, L. A. Gethings and R. S. Plumb, A novel blood proteomic signature for prostate cancer, Cancers, 2023, 15(4), 1051 CrossRef CAS PubMed.
- F. Crocetto, G. Russo, E. Di Zazzo, P. Pisapia, B. F. Mirto, A. Palmieri, F. Pepe, C. Bellevicine, A. Russo, E. La Civita and D. Terracciano, Liquid biopsy in prostate cancer management—Current challenges and future perspectives, Cancers, 2022, 14(13), 3272 CrossRef CAS PubMed.
- N. Gomez-Cebrian, J. L. Poveda, A. Pineda-Lucena and L. Puchades-Carrasco, Metabolic phenotyping in prostate cancer using multi-omics approaches, Cancers, 2022, 14(3), 596 CrossRef CAS PubMed.
- E. Nevedomskaya and B. Haendler, From omics to multi-omics approaches for in-depth analysis of the molecular mechanisms of prostate cancer, Int. J. Mol. Sci., 2022, 23(11), 6281 CrossRef CAS PubMed.
- M. Kdadra, S. Höckner, H. Leung, W. Kremer and E. Schiffer, Metabolomics biomarkers of prostate cancer: a systematic review, Diagnostics, 2019, 9(1), 21 CrossRef CAS PubMed.
- B. Mirza, W. Wang, J. Wang, H. Choi, N. C. Chung and P. Ping, Machine learning and integrative analysis of biomedical big data, Genes, 2019, 10(2), 87 CrossRef CAS PubMed.
- B. R. Fulmer and T. T. Turner, A blood-prostate barrier restricts cell and molecular movement across the rat ventral prostate epithelium, J. Urol., 2000, 163(5), 1591–1594 CrossRef CAS PubMed.
- Y. Shang, D. Cui and S. Yi, Opening Tight Junctions May Be Key to Opening the Blood–Prostate Barrier, Medical Science Monitor: International Medical Journal of Experimental and Clinical Research, 2014, vol. 20, p. 2504 Search PubMed.
- H. Xia, D. Yang, W. He, X. Zhu, Y. Yan, Z. Liu, T. Liu, J. Yang, S. Tan, J. Jiang and X. Hou, Ultrasound-mediated microbubbles cavitation enhanced chemotherapy of advanced prostate cancer by increasing the permeability of blood-prostate barrier, Transl. Oncol., 2021, 14(10), 101177 CrossRef CAS PubMed.
- M. Spick, A. Muazzam, H. Pandha, A. Michael, L. A. Gethings, C. J. Hughes, N. Munjoma, R. S. Plumb, I. D. Wilson, A. D. Whetton and P. A. Townsend, Multi-omic diagnostics of prostate cancer in the presence of benign prostatic hyperplasia, Heliyon, 2023, 9(12), e22604 CrossRef CAS PubMed.
- S. Voora and D. B. Adey, Management of kidney transplant recipients by general nephrologists: core curriculum 2019, Am. J. Kidney Dis., 2019, 73(6), 866–879 CrossRef PubMed.
- M. Cavaleri, M. Veroux, F. Palermo, F. Vasile, M. Mineri, J. Palumbo, L. Salemi, M. Astuto and P. Murabito, Perioperative goal-directed therapy during kidney transplantation: an impact evaluation on the major postoperative complications, J. Clin. Med., 2019, 8(1), 80 CrossRef CAS PubMed.
- A. Hart, J. M. Smith, M. A. Skeans, S. K. Gustafson, A. R. Wilk, A. Robinson, J. L. Wainright, C. R. Haynes, J. J. Snyder, B. L. Kasiske and A. K. Israni, OPTN/SRTR 2016 annual data report: kidney, Am. J. Transplant., 2018, 18, 18–113 CrossRef PubMed.
- Z. Zhang, Y. Tang, H. Zhuang, E. Lin, L. Xie, X. Feng, J. Zeng, Y. Liu, J. Liu and Y. Yu, Identifying 4 novel lncRNAs as potential biomarkers for acute rejection and graft loss of renal allograft, J. Immunol. Res., 2020, 2020(1), 2415374 CrossRef PubMed.
- X. Zheng, K. Xu, B. Zhou, T. Chen, Y. Huang, Q. Li, F. Wen, W. Ge, J. Wang, S. Yu and L. Sun, A circulating extracellular vesicles-based novel screening tool for colorectal cancer revealed by shotgun and data-independent acquisition mass spectrometry, J. Extracell. Vesicles, 2020, 9(1), 1750202 CrossRef CAS PubMed.
- L. Niu, M. Thiele, P. E. Geyer, D. N. Rasmussen, H. E. Webel, A. Santos, R. Gupta, F. Meier, M. Strauss, M. Kjaergaard and K. Lindvig, Noninvasive proteomic biomarkers for alcohol-related liver disease, Nat. Med., 2022, 28(6), 1277–1287 CrossRef CAS PubMed.
- M. Xu, J. Deng, K. Xu, T. Zhu, L. Han, Y. Yan, D. Yao, H. Deng, D. Wang, Y. Sun and C. Chang, In-depth serum proteomics reveals biomarkers of psoriasis severity and response to traditional Chinese medicine, Theranostics, 2019, 9(9), 2475 CrossRef CAS PubMed.
- C. Wang, G. Feng, J. Zhao, Y. Xu, Y. Li, L. Wang, M. Wang, M. Liu, Y. Wang, H. Mu and C. Zhou, Screening of novel biomarkers for acute kidney transplant rejection using DIA-MS based proteomics, Proteonomics Clin. Appl., 2024, 18(3), 2300047 CrossRef CAS PubMed.
- R. L. Siegel, K. D. Miller and A. Jemal, Cancer statistics, 2018, Ca-Cancer J. Clin., 2018, 68(1), 7–30 CrossRef PubMed.
- J. D. Seidman, I. Horkayne-Szakaly, M. Haiba, C. R. Boice, R. J. Kurman and B. M. Ronnett, The histologic type and stage distribution of ovarian carcinomas of surface epithelial origin, Int. J. Gynecol. Pathol., 2004, 23(1), 41–44 CrossRef PubMed.
- M. Ni, D. Wan, J. Wu, W. Gong, J. Wang and Z. Zheng, Candidate prognostic biomarkers and prediction models for high-grade serous ovarian cancer from urinary proteomics, J. Proteonomics, 2024, 304, 105234 CrossRef CAS PubMed.
- S. Lheureux, C. Gourley, I. Vergote and A. M. Oza, Epithelial ovarian cancer, Lancet, 2019, 393(10177), 1240–1253 CrossRef PubMed.
- B. M. Reid, J. B. Permuth and T. A. Sellers, Epidemiology of ovarian cancer: a review, Cancer Biol. Med., 2017, 14(1), 9–32 CAS.
- R. E. Lane, D. Korbie, M. M. Hill and M. Trau, Extracellular vesicles as circulating cancer biomarkers: opportunities and challenges, Clin. Transl. Med., 2018, 7(1), 14 CrossRef CAS PubMed.
- A. Becker, B. K. Thakur, J. M. Weiss, H. S. Kim, H. Peinado and D. Lyden, Extracellular vesicles in cancer: cell-to-cell mediators of metastasis, Cancer Cell, 2016, 30(6), 836–848 CrossRef CAS PubMed.
- T. T. Cooper, D. Z. Dieters-Castator, J. Liu, G. M. Siegers, D. Pink, L. Veliz, J. D. Lewis, F. Lagugné-Labarthet, Y. Fu, H. Steed and G. A. Lajoie, Targeted proteomics of plasma extracellular vesicles uncovers MUC1 as combinatorial biomarker for the early detection of high-grade serous ovarian cancer, J. Ovarian Res., 2024, 17(1), 149 CrossRef CAS PubMed.
- R. L. Siegel, K. D. Miller, N. S. Wagle and A. Jemal, Cancer statistics, 2023, Ca-Cancer J. Clin., 2023, 73(1), 17–48 CrossRef PubMed.
- A. Adamska, A. Domenichini and M. Falasca, Pancreatic ductal adenocarcinoma: current and evolving therapies, Int. J. Mol. Sci., 2017, 18(7), 1338 CrossRef PubMed.
- P. Rawla, T. Sunkara and V. Gaduputi, Epidemiology of pancreatic cancer: global trends, etiology and risk factors, World J. Oncol., 2019, 10(1), 10 CrossRef PubMed.
- R. Kong, X. Qian and W. Ying, Pancreatic cancer cells spectral library by DIA-MS and the phenotype analysis of gemcitabine sensitivity, Sci. Data, 2022, 9, 283 CrossRef CAS PubMed.
- A. S. Quante, C. Ming, M. Rottmann, J. Engel, S. Boeck, V. Heinemann, C. B. Westphalen and K. Strauch, Projections of cancer incidence and cancer-related deaths in Germany by 2020 and 2030, Cancer Med., 2016, 5(9), 2649–2656 CrossRef PubMed.
- L. Rahib, B. D. Smith, R. Aizenberg, A. B. Rosenzweig, J. M. Fleshman and L. M. Matrisian, Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States, Cancer Res., 2014, 74(11), 2913–2921 CrossRef CAS PubMed.
- J. D. Mizrahi, R. Surana, J. W. Valle and R. T. Shroff, Pancreatic cancer, Lancet, 2020, 395(10242), 2008–2020 CrossRef CAS PubMed.
- T. M. Lih, L. Cao, P. Minoo, G. S. Omenn, R. H. Hruban, D. W. Chan, O. F. Bathe and H. Zhang, Detection of pancreatic ductal adenocarcinoma-associated proteins in serum, Mol. Cell. Proteomics, 2024, 23(1), 100687 CrossRef CAS PubMed.
- H. Sung, J. Ferlay, R. L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal and F. Bray, Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries, Ca-Cancer J. Clin., 2021, 71(3), 209–249 CrossRef PubMed.
- Brierley J. D., Van Eycken E., Rous B. and Giuliani M., TNM Classification of Malignant Tumours, John Wiley & Sons, 2025 Search PubMed.
- F. Petrelli, F. Trevisan, M. Cabiddu, G. Sgroi, L. Bruschieri, E. Rausa, M. Ghidini and L. Turati, Total neoadjuvant therapy in rectal cancer: a systematic review and meta-analysis of treatment outcomes, Ann. Surg., 2020, 271(3), 440–448 CrossRef PubMed.
- M. Loos, P. Quentmeier, T. Schuster, U. Nitsche, R. Gertler, A. Keerl, T. Kocher, H. Friess and R. Rosenberg, Effect of preoperative radio (chemo) therapy on long-term functional outcome in rectal cancer patients: a systematic review and meta-analysis, Ann. Surg Oncol., 2013, 20(6), 1816–1828 CrossRef PubMed.
- F. López-Campos, M. Martín-Martín, R. Fornell-Pérez, J. C. García-Pérez, J. Die-Trill, R. Fuentes-Mateos, S. López-Durán, J. Domínguez-Rullán, R. Ferreiro, A. Riquelme-Oliveira and A. Hervás-Morón, Watch and wait approach in rectal cancer: Current controversies and future directions, World J. Gastroenterol., 2020, 26(29), 4218 CrossRef PubMed.
- A. Stanojevic, M. Samiotaki, V. Lygirou, M. Marinkovic, V. Nikolic, S. Stojanovic-Rundic, R. Jankovic, A. Vlahou, G. Panayotou, R. J. Fijneman and S. Castellví-Bel, Data-independent acquisition mass spectrometry analysis of FFPE rectal cancer samples offers in-depth proteomics characterization of the response to neoadjuvant chemoradiotherapy, Int. J. Mol. Sci., 2023, 24(20), 15412 CrossRef CAS PubMed.
- C. Pobel, E. Auclin, A. Procureur, A. Clément-Zhao, A. Simonaggio, N. Delanoy, Y. A. Vano, C. Thibault and S. Oudard, Cabazitaxel schedules in metastatic castration-resistant prostate cancer: a review, Future Oncol., 2021, 17(1), 91–102 CrossRef CAS PubMed.
- G. Galletti, B. I. Leach, L. Lam and S. T. Tagawa, Mechanisms of resistance to systemic therapy in metastatic castration-resistant prostate cancer, Cancer Treat Rev., 2017, 57, 16–27 CrossRef CAS PubMed.
- Y. Kanai, Amino acid transporter LAT1 (SLC7A5) as a molecular target for cancer diagnosis and therapeutics, Pharmacol. Therapeut., 2022, 230, 107964 CrossRef CAS PubMed.
- M. Xu, S. Sakamoto, J. Matsushima, T. Kimura, T. Ueda, A. Mizokami, Y. Kanai and T. Ichikawa, Up-regulation of LAT1 during antiandrogen therapy contributes to progression in prostate cancer cells, J. Urol., 2016, 195(5), 1588–1597 CrossRef PubMed.
- K. Higuchi, S. Sakamoto, K. Ando, M. Maimaiti, N. Takeshita, K. Okunushi, Y. Reien, Y. Imamura, T. Sazuka, K. Nakamura and J. Matsushima, Characterization of the expression of LAT1 as a prognostic indicator and a therapeutic target in renal cell carcinoma, Sci. Rep., 2019, 9(1), 16776 CrossRef PubMed.
- M. Maimaiti, S. Sakamoto, Y. Yamada, M. Sugiura, J. Rii, N. Takeuchi, Y. Imamura, T. Furihata, K. Ando, K. Higuchi and M. Xu, Expression of L-type amino acid transporter 1 as a molecular target for prognostic and therapeutic indicators in bladder carcinoma, Sci. Rep., 2020, 10(1), 1292 CrossRef CAS PubMed.
- S. Yothaisong, H. Dokduang, N. Anzai, K. Hayashi, N. Namwat, P. Yongvanit, S. Sangkhamanon, P. Jutabha, H. Endou and W. Loilome, Inhibition of l-type amino acid transporter 1 activity as a new therapeutic target for cholangiocarcinoma treatment, Tumor Biol., 2017, 39(3), 1010428317694545 Search PubMed.
- J. Rii, S. Sakamoto, A. Mizokami, M. Xu, A. Fujimoto, S. Saito, H. Koike, T. Tamura, T. Arai, Y. Yamada and Y. Goto, L-type amino acid transporter 1 inhibitor JPH203 prevents the growth of cabazitaxel-resistant prostate cancer by inhibiting cyclin-dependent kinase activity, Cancer Sci., 2024, 115(3), 937–953 CrossRef CAS PubMed.
- W. C. R. Fund, Ovarian Cancer Statistics, 2022, available online: https://www.wcrf.org/cancer-trends/ovarian-cancer-statistics/, accessed on 5 January 2023 Search PubMed.
- Australian Institute of Health and Welfare, Ovarian Cancer Statistics in Australia, 2022, available online: https://www.canceraustralia.gov.au/cancer-types/ovarian-cancer/statistics, accessed on 5 January 2023 Search PubMed.
- G. Arentz, P. Mittal, M. Klingler-Hoffmann, M. R. Condina, C. Ricciardelli, N. A. Lokman, G. Kaur, M. K. Oehler and P. Hoffmann, Label-free quantification mass spectrometry identifies protein markers of chemotherapy response in high-grade serous ovarian cancer, Cancers, 2023, 15(7), 2172 CrossRef CAS PubMed.
- M. Ni, D. Wan, J. Wu, W. Gong, J. Wang and Z. Zheng, Candidate prognostic biomarkers and prediction models for high-grade serous ovarian cancer from urinary proteomics, J. Proteonomics, 2024, 304, 105234 CrossRef CAS PubMed.
- A. C. Tan and D. S. Tan, Targeted therapies for lung cancer patients with oncogenic driver molecular alterations, J. Clin. Oncol., 2022, 40(6), 611–625 CrossRef CAS PubMed.
- B. J. Solomon, T. Mok, D. W. Kim, Y. L. Wu, K. Nakagawa, T. Mekhail, E. Felip, F. Cappuzzo, J. Paolini, T. Usari and S. Iyer, First-line crizotinib versus chemotherapy in ALK-positive lung cancer, N. Engl. J. Med., 2014, 371(23), 2167–2177 CrossRef PubMed.
- J. C. Soria, D. S. Tan, R. Chiari, Y. L. Wu, L. Paz-Ares, J. Wolf, S. L. Geater, S. Orlov, D. Cortinovis, C. J. Yu and M. Hochmair, First-line ceritinib versus platinum-based chemotherapy in advanced ALK-rearranged non-small-cell lung cancer (ASCEND-4): a randomised, open-label, phase 3 study, Lancet, 2017, 389(10072), 917–929 CrossRef CAS.
- D. R. Camidge, H. R. Kim, M. J. Ahn, J. C. Yang, J. Y. Han, M. J. Hochmair, K. H. Lee, A. Delmonte, M. R. García Campelo, D. W. Kim and F. Griesinger, Brigatinib versus crizotinib in advanced ALK inhibitor–naive ALK-positive non–small cell lung cancer: second interim analysis of the phase III ALTA-1L trial, J. Clin. Oncol., 2020, 38(31), 3592–3603 CrossRef CAS PubMed.
- T. Mok, D. R. Camidge, S. M. Gadgeel, R. Rosell, R. Dziadziuszko, D. W. Kim, M. Pérol, S. H. Ou, J. S. Ahn, A. T. Shaw and W. Bordogna, Updated overall survival and final progression-free survival data for patients with treatment-naive advanced ALK-positive non-small-cell lung cancer in the ALEX study, Ann. Oncol., 2020, 31(8), 1056–1064 CrossRef CAS PubMed.
- A. T. Shaw, T. M. Bauer, F. de Marinis, E. Felip, Y. Goto, G. Liu, J. Mazieres, D. W. Kim, T. Mok, A. Polli and H. Thurm, First-line lorlatinib or crizotinib in advanced ALK-positive lung cancer, N. Engl. J. Med., 2020, 383(21), 2018–2029 CrossRef CAS PubMed.
- S. Wang, X. Hao, L. Dai, N. Lou, G. Fan, R. Gao, M. Yang, P. Xing, Y. Liu, L. Wang and Z. Zhang, Longitudinal plasma proteomic profiling of EML4-ALK positive lung cancer receiving ALK-TKIs therapy, Lung Cancer, 2024, 189, 107503 CrossRef CAS PubMed.
- S. Ebrahimi, X. Guo, Transformer-based de novo peptide sequencing for data-independent acquisition mass spectrometry, in 2023 IEEE 23rd International Conference on Bioinformatics and Bioengineering (BIBE), IEEE, 2023, pp. 28–35 Search PubMed.
- L. Käll and O. Vitek, AI-powered mass spectrometry: intelligent software for intelligent experiments, J. Proteome Res., 2021, 20(5), 2345–2355, DOI:10.1021/acs.jproteome.0c00856.
- N. Alijaj, B. Pavlovic, P. Martel, A. Rakauskas, V. Cesson, K. Saba, T. Hermanns, P. Oechslin, M. Veit, M. Provenzano and J. H. Rüschoff, Identification of urine biomarkers to improve eligibility for prostate biopsy and detect high-grade prostate cancer, Cancers, 2022, 14(5), 1135 CrossRef CAS PubMed.
- M. Dong, T. S. Lih, N. Hoti, S. Y. Chen, S. Ponce, A. Partin and H. Zhang, Development of parallel reaction monitoring assays for the detection of aggressive prostate cancer using urinary glycoproteins, J. Proteome Res., 2021, 20(7), 3590–3599 CrossRef CAS PubMed.
- H. Zhang, G. Y. Zhang, W. C. Su, Y. T. Chen, Y. F. Liu, D. Wei, Y. X. Zhang, Q. Y. Tang, Y. X. Liu, S. Z. Wang and W. C. Li, High throughput isolation and data independent acquisition mass spectrometry (DIA-MS) of urinary extracellular vesicles to improve prostate cancer diagnosis, Molecules, 2022, 27(23), 8155 CrossRef CAS PubMed.
- T. M. Lih, L. Cao, P. Minoo, G. S. Omenn, R. H. Hruban, D. W. Chan, O. F. Bathe and H. Zhang, Detection of pancreatic ductal adenocarcinoma-associated proteins in serum, Mol. Cell. Proteomics, 2024, 23(1), 100687 CrossRef CAS PubMed.
- S. Huh, C. Kang, J. E. Park, D. Nam, S. I. Kim, A. Seol, K. Choi, D. Hwang, M. H. Yu, H. H. Chung and S. W. Lee, Novel diagnostic biomarkers for high-grade serous ovarian cancer uncovered by data-independent acquisition mass spectrometry, J. Proteome Res., 2022, 21(9), 2146–2159 CrossRef CAS PubMed.
- M. Ni, J. Zhou, W. Gong, R. Jiang, X. Li, W. Dai, Z. Yin, Z. Chen, Z. Zheng and J. Zhu, Proteomic analysis reveals CAAP1 negatively correlates with platinum resistance in ovarian cancer, J. Proteonomics, 2023, 277, 104864 CrossRef CAS PubMed.
- S. Ji, Y. Liu, L. Yan, Y. Zhang, Y. Li, Q. Zhu, W. Xia, S. Ge and J. Zhang, DIA-based analysis of the menstrual blood proteome identifies association between CXCL5 and IL1RN and endometriosis, J. Proteonomics, 2023, 289, 104995 CrossRef CAS PubMed.
- S. Weng, M. Wang, Y. Zhao, W. Ying and X. Qian, Optimised data-independent acquisition strategy recaptures the classification of early-stage hepatocellular carcinoma based on data-dependent acquisition, J. Proteonomics, 2021, 238, 104152 CrossRef CAS PubMed.
- S. T. Gebreyesus, A. A. Siyal, R. B. Kitata, E. S. Chen, B. Enkhbayar, T. Angata, K. I. Lin, Y. J. Chen and H. L. Tu, Streamlined single-cell proteomics by an integrated microfluidic chip and data-independent acquisition mass spectrometry, Nat. Commun., 2022, 13(1), 37 CrossRef CAS PubMed.
- L. Ma, J. E. Muscat, R. Sinha, D. Sun and G. Xiu, Proteomics of exhaled breath condensate in lung cancer and controls using data-independent acquisition (DIA): a pilot study, J. Breath Res., 2021, 15(2), 026002 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |