Chemical reaction-enabled lipidomics: from sensitive structural analysis to biomedical applications
Qirui
Yu
and
Xiaoxiao
Ma
 *
*
State Key Laboratory of Precision Measurement Technology and Instruments, Department of Precision Instrument, Tsinghua University, Beijing, 100084, China. E-mail: maxx@tsinghua.edu.cn
First published on 11th February 2026
Abstract
Lipidomics has emerged as a vital discipline for understanding cellular metabolism and disease pathology. However, the immense structural diversity, wide dynamic range, and varying ionization efficiencies of lipids present significant analytical challenges. The MS analysis workflow often falls short in detecting low-abundance species and resolving complex structural isomers. To address these limitations, chemical derivatization has been widely adopted to manipulate the chemical properties of lipids prior to analysis. This review summarizes the significant progress in chemical derivatization-enabled lipidomics over the past decades, highlighting its pivotal role in bridging the gap between analytical capability and biological complexity. We critically discuss three core dimensions of improvement: (1) enhancement of detection sensitivity through derivatization strategies that increase the ionization efficiency of lipids; (2) refinement of structural elucidation, specifically using selective reactions to pinpoint carbon–carbon double bond locations and differentiate isomers; and (3) advancement of spectrometric specificity and quantification via mass-shift profiling, which enables precise quantification or high-throughput multiplex analysis. Finally, we discuss how these chemical tools are facilitating the discovery of novel lipid biomarkers and providing deeper insights into lipid metabolism in biomedical research.
Introduction
Lipids are widely distributed in living organisms and closely related to a variety of complex biological functions.1 Glycerophospholipids, sterols, and sphingolipids are fundamental components of cell membranes. The formation of the lipid bilayer ensures the integrity and selective permeability of the cell membrane, while cholesterol regulates membrane fluidity and the stability of the phospholipid bilayer.2,3 Fats serve as a major form of energy storage in mammals. Upon energy surplus, triglycerides are synthesized and stored in adipocytes.4 Furthermore, many lipids play crucial roles in signal transduction. For instance, phosphatidylinositol derivatives regulate the activity of membrane proteins in cellular signaling,5,6 and fatty acid-derived prostaglandins and leukotrienes are key inflammatory mediators.7 Moreover, a growing body of evidence indicates that lipid metabolism is associated with numerous diseases, such as cardiovascular diseases,8 Alzheimer's disease,9 COVID-19,10 and cancer.11–13 Given the increasing roles of lipids in biology and pathology, a comprehensive and detailed analysis of these molecules is of great importance.The structural diversity and complexity of lipids present significant challenges in their comprehensive analysis. As of December 9, 2025, the LIPID MAPS database has cataloged 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 288 lipid species (including 22
288 lipid species (including 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 034 computationally generated and 28
034 computationally generated and 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 254 curated entries). The rapid development of mass spectrometry techniques for in-depth lipidome characterization has made the large-scale identification and quantification of lipids feasible.14,15 To date, lipidomics has emerged as a distinct discipline for the study of lipid composition and functions. The term “lipidomics” was first proposed in 2003 by Han and Gross,16–19 the aim of which is to reveal the biological composition, functions and mechanisms of lipids, ultimately elucidating how lipid metabolism affects physiological or pathological processes. The primary objectives of lipidomics research include the identification of lipid structures, quantitative lipid analysis, elucidation of lipid metabolic pathways, and discovery of lipid biomarkers.
254 curated entries). The rapid development of mass spectrometry techniques for in-depth lipidome characterization has made the large-scale identification and quantification of lipids feasible.14,15 To date, lipidomics has emerged as a distinct discipline for the study of lipid composition and functions. The term “lipidomics” was first proposed in 2003 by Han and Gross,16–19 the aim of which is to reveal the biological composition, functions and mechanisms of lipids, ultimately elucidating how lipid metabolism affects physiological or pathological processes. The primary objectives of lipidomics research include the identification of lipid structures, quantitative lipid analysis, elucidation of lipid metabolic pathways, and discovery of lipid biomarkers.
Improvement in the analytical capability of mass spectrometers can enhance lipidomics analysis in multiple dimensions. These include high resolution, fast speed of analysis,20 comprehensive structural characterization,21,22 and a high dynamic range of quantitative analysis.23 Chemical derivatization refers to subjecting lipids to chemical reactions prior to analysis, leading to improved detection sensitivity, in-depth structural elucidation, and accurate quantitation. With increasingly powerful MS instrumentation, the potential of chemical derivatization in lipidomics research remains to be explored. This review summarizes the current progress in chemical derivatization for lipidomics analysis within the past decade.
Strategies for enhanced detection sensitivity
Lipids exhibit remarkable chemical diversity, structural complexity, and varied polarity, which together result in significant variations in their ionization efficiencies for MS analysis. Such disparities in ionization lead to inconsistent signal responses among different lipid classes, and lipids of low abundance are often poorly ionized or even missed during detection. To address this challenge, researchers have developed a variety of chemical derivatization strategies to enhance lipid ionization. By introducing charged or easily ionizable chemical groups, chemical derivatization can effectively improve the ionization efficiency and detection sensitivity of lipids of various classes.Methylation
Methylation was introduced to derivatize fatty acids (FAs), which possess relatively high molecular weights and extensive hydrogen bonding, since FAs are non-volatile and unsuitable for direct GC-MS analysis. To enable analysis, FAs must be converted to their methyl esters before injection.24 In LC-MS, such derivatization is generally not required, but lipids containing carboxyl or phosphate groups often have low ionization efficiency in the positive mode. Methylation provides an effective strategy to neutralize active hydrogens and thereby leads to enhanced sensitivity of detection. Phosphatidylinositol phosphates (PIPs) are signaling lipids in biological systems but are typically low in abundance and difficult to ionize compared with other phospholipids. In 2011, Hawkins et al. demonstrated that methylation of phosphatidylinositols using TMS-diazomethane (TMSD), followed by HPLC-MS analysis, significantly improved the detection sensitivity of PIPs (Fig. 1a).25 Using this workflow, the authors successfully quantified PI and PIP3 in human neutrophils. This approach can be broadly applied to various PIPs and can also facilitate structural elucidation through MS/MS analysis.26,27 Since then, this derivatization strategy has been widely adopted in the studies of PIP-related metabolic pathways and biological processes.28,29 | ||
| Fig. 1 (a) Methylation of phosphatidylinositol triphosphate (PIP3) via reaction with TMSD. Reproduced from ref. 25 with permission from Springer Nature, copyright 2011. (b) Methylation of PE and PS via reaction with diazomethane. Reproduced from ref. 30 with permission from the American Chemical Society, copyright 2014. (c) Methylation of CL via TMSD. Reproduced from ref. 34 with permission from the American Chemical Society, copyright 2017. (d) Dimethylation of DoIP via TMSD. Reproduced from ref. 35 with permission from the American Chemical Society, copyright 2023. | ||
In 2014, Wasslen and colleagues developed a derivatization technique known as trimethylation enhancement using diazomethane (TrEnDi), which enables methylation of peptides and amino-containing phospholipids such as phosphatidylethanolamine (PE) and phosphatidylserine (PS) (Fig. 1b). This modification introduces a permanent positive charge to lipids, thereby enhancing their ionization efficiency and detection sensitivity.30 They further incorporated 13C-labeled stable isotopes into the derivatization reagent, allowing for the differentiation of PC and PE species within complex biological samples. Using this approach, various phospholipids and sphingolipids could be analyzed.31 Subsequent developments have refined the 13C-TrEnDi strategy, including the creation of dedicated experimental platforms to improve safety and reproducibility.32 In addition, a 13C-TrEnDi database was established based on lipid extracts from yeast, facilitating the integration of this derivatization technique into diverse LC-MS-based lipidomics workflows.33
Cardiolipins (CL) represent a class of phospholipids with extremely low abundance, and their simultaneous detection with other phospholipids in biological samples remains challenging. To address this issue, the Moon group developed an isotope-labeled methylation derivatization method to enhance the mass spectrometric analysis of cardiolipins (Fig. 1c).34 This approach not only improved the detection sensitivity of CLs but also increased the accuracy of quantitative measurements. Similarly, dolichyl monophosphates (DolP) are difficult to analyze due to their low abundance and the lack of effective enrichment strategies. After methylation, however, DolPs can be analyzed by RPLC-MS with greatly improved sensitivity and quantitative precision (Fig. 1d).35
In 2011, the Bamba group developed a methylation-based mass spectrometric approach coupled with supercritical fluid chromatography (SFC), which was particularly suitable for the targeted analysis of polar lipids such as phosphatidic acid (PA), phosphatidylglycerol (PG), and PI.36 With further optimization, this method was expanded to enable high-sensitivity analysis of complex mixtures containing multiple polar lipid classes, including PS. Using this workflow, the authors achieved sensitive and reliable quantification of low-abundance polar lipids in mouse liver samples.37 These findings demonstrate that methylation-based derivatization provides multiple advantages for the analysis of low-abundance and highly polar lipids. Moreover, even when the overall abundance of analytes is reduced due to incomplete derivatization yields, methylation can still improve ionization efficiency and enhance signal intensity, thereby compensating for potential losses during reaction.38
Pyridinium or quaternary ammonium derivatization
In mass spectrometric analysis, molecules carrying a permanent positive charge are ionized much more efficiently than neutral species, especially in polar solvents where charged groups remain fully dissociated. Many lipid classes, however, lack a stable positive charge under mildly acidic or neutral conditions, which leads to poor ionization efficiency and low detection sensitivity. Charge-tagging derivatization addresses this limitation by introducing a fixed cationic group—such as a pyridinium or quaternary ammonium moiety—onto the lipid molecule, thereby converting weakly ionizable compounds into readily detectable ions. FAs, with their simple structures and well-defined functional groups, serve as ideal substrates for such derivatization strategies and have thus become a major testing ground for the development of charge-tagging reagents.In 2007, Yang and colleagues introduced 3-acyloxymethyl-1-methylpyridinium iodide (AMMP) onto FAs, achieving an approximately 2500-fold improvement in ionization efficiency compared with negative mode analysis.39 Quaternary ammonium and pyridinium salts exhibit similar effects, and appropriate selection of derivatization reagents and conditions can yield comparable results.40 To enhance the reactivity of different reagents with FAs, Li et al. first converted the carboxyl group of FAs into highly reactive acyl chlorides prior to derivatization.41 The Han group further optimized this strategy by screening various quaternary ammonium and pyridinium reagents and identified N-[4-(aminomethyl)phenyl]pyridinium (AMPP) as a highly effective tag (Fig. 2a). AMPP-derivatized FAs and hydroxy fatty acids were successfully quantified in human plasma lipid extracts, establishing a shotgun lipidomics workflow for biological applications.42,43 AMPP derivatives produce a characteristic 183 Da fragment ion under MS/MS CID, enabling targeted quantification through multiple reaction monitoring (MRM). The AMPP reagent has since been commercialized and widely adopted in LC-MS-based lipidomics for determining total FA composition.44
 | ||
| Fig. 2 (a) Enhanced positive-mode ionization of fatty acids via AMPP derivatization (commercial kit). Reproduced from ref. 42. (b) Fatty aldehyde derivatization with N-heterocyclic carbenes (NHCs) for LC-MS analysis. Reproduced from ref. 54. (c) Derivatization of steroids with Girard reagents. Reproduced from ref. 55 with permission from Elsevier, copyright 2013. Licensed under CC-BY 4.0. (d) Employing acetylpyridine instead of acetone in the Paternò–Büchi reaction to enhance product ionization via the pyridine moiety. Reproduced from ref. 57 with permission from the American Chemical Society, copyright 2018. (e) Multiple structural elucidation by charge switching derivatization. Reproduced from ref. 60 with permission from the American Chemical Society, copyright 2025. Licensed under CC-BY 4.0. | ||
Beyond LC-MS applications, charge-tagging derivatization also helps reduce matrix effects in MALDI-MS analysis. In 2009, Koulman and colleagues developed a pyridinium-based derivatization method for carboxylic acid metabolites, demonstrating significant improvements in sensitivity, dynamic range, and quantitative accuracy.45
Pyridine or amine group tagging
Introducing a tag that carries a permanent positive charge can greatly enhance ionization efficiency, but this modification often leads to fragmentation patterns dominated by the charge tag, resulting in the loss of valuable structural information in MS/MS spectra. To balance ionization enhancement with structural characterization, incorporating pyridine or amine groups provides a more flexible strategy. Liu and colleagues used DEEA to derivatize FAs, achieving a three-order-of-magnitude increase in signal intensity.46 The derivatives exhibited a characteristic 74 Da neutral loss under MS/MS CID, enabling FA quantification through neutral-loss scanning. Commercial aminoxy-TMT reagents can also be applied to FA labeling, allowing multiplexed analysis of total FA composition in positive mode.47 In another example, Jiang et al. introduced a pair of structurally related tags to derivatize FAs.48 Their design not only improved sensitivity but also produced distinct fragmentation ions, with the tag pair used separately for samples and internal standards to ensure quantitative accuracy.Because FAs offer limited structural information under routine MS/MS conditions, pyridine- or amine-based tagging has been more widely applied to lipid classes with greater structural complexity. Fatty aldehydes, for instance, rely heavily on charge-tagging strategies for reliable detection. Their high reactivity makes Schiff-base formation highly favorable, and more than ten derivatization reagents have been developed since the early 2000s,49–51 with comprehensive summaries available in several reviews.52,53 However, Schiff-base chemistry inevitably generates a C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond, which may introduce unnecessary chromatographic behavior and complicate separation. To address this limitation, the Feng group recently developed an N-heterocyclic carbene–based derivatization method that improved the linearity, reproducibility, and accuracy of fatty aldehyde analysis, while lowering both detection and quantification limits (Fig. 2b). Using this workflow, they quantified fatty aldehydes in serum samples from sepsis patients.54 Similarly, Girard reagents, which contain a hydrazide group, have long been used to derivatize steroids, enhancing their ionization (Fig. 2c).55 Sphingolipids, which possess a native amine group, can be labeled using naphthalene-2,3-dicarboxaldehyde to form an N-heteroaromatic product that behaves similarly to pyridinium-based tags.56
N bond, which may introduce unnecessary chromatographic behavior and complicate separation. To address this limitation, the Feng group recently developed an N-heterocyclic carbene–based derivatization method that improved the linearity, reproducibility, and accuracy of fatty aldehyde analysis, while lowering both detection and quantification limits (Fig. 2b). Using this workflow, they quantified fatty aldehydes in serum samples from sepsis patients.54 Similarly, Girard reagents, which contain a hydrazide group, have long been used to derivatize steroids, enhancing their ionization (Fig. 2c).55 Sphingolipids, which possess a native amine group, can be labeled using naphthalene-2,3-dicarboxaldehyde to form an N-heteroaromatic product that behaves similarly to pyridinium-based tags.56
Multifunctional tagging
Derivatization methods designed for structural elucidation or quantification often provide limited benefits when the target lipids are present at very low abundance. Without improved ionization efficiency, these lipids may still fall below the detection threshold. Performing two separate derivatization steps can in principle address this issue, but such workflows frequently suffer from low yields, incompatible reaction conditions, or poor reproducibility. Multifunctional tags offer a practical solution by combining multiple analytical functions—such as ionization enhancement and quantitative encoding—within a single reagent, allowing low-abundance lipids to be analyzed more reliably without complex multistep procedures. Esch and colleagues reported in 2018 a class of pyridine-containing tagging reagents that produced strong ionization enhancement for neutral unsaturated lipids (Fig. 2d), increasing signal intensity by more than 600 fold.57 Xie et al. evaluated a series of pyridine-based reagents and identified 2-AP as an efficient tag with high derivatization yields, which they applied to the analysis of cholesteryl esters (CE).58 Reagents of this type have since been incorporated into several established lipidomics workflows.44,59Because positive mode generally provides higher ionization efficiency than negative mode, most derivatization strategies for low-abundance lipids aim to introduce a permanent or easily formed positive charge. However, negative mode remains advantageous for structural analysis, particularly for obtaining fatty-acyl chain information from complex lipids. To address this need, the Yan group recently reported a derivatization strategy for unsaturated GPLs that introduces an aromatic moiety carrying a stable negative charge under basic conditions (Fig. 2e).60 This approach enhances ionization in negative mode while enabling the simultaneous assignment of acyl-chain composition and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C double-bond positions. All these lipid derivatization reagents discussed are listed in Table 1.
C double-bond positions. All these lipid derivatization reagents discussed are listed in Table 1.
| Reagent | Formula | Advantages | Permanent charge | Analyte | Ref. |
|---|---|---|---|---|---|
| AMMP |

|
Remarkable ionization enhancement (∼2500-fold) vs. negative mode; compatible with stable isotope coding | Positive | Fatty acid | 39 |
| AMPP |
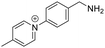
|
High reaction stability; commercial reagent; well-established analytical workflows | Positive | Fatty acid | 42–44 |
| HMEP |

|
Homogenizes solution-phase properties for MALDI compatibility; suppresses matrix background ions; compatible with differential isotope coding | Positive | Fatty acid | 45 |
| DEEA |

|
Rapid and high selectivity reaction; provides a signature NLS pattern | — | Fatty acid | 46 |
| AminoxyTMT |

|
Highly efficient and multiplexed labeling for precise quantification; commercial reagent | — | Fatty acid | 47 |
| Dns-PP |

|
Dual labeling of analytes and internal standards for accurate, high-throughput quantification | — | Fatty acid | 48 |
| MTPTC |
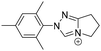
|
High selectivity; well-resolved single peaks in LC; addressed the issue of stereoisomeric products | — | Fatty aldehydes | 54 |
| Girard reagents |

|
Highly adaptable for further functionalization; improved solubility | — | Steroids | 55 |
| NDA |

|
High derivatization specificity; streamlined experimental protocol | — | SL | 56 |
| 2-AP (PB reagent) |

|
Enables detailed structural elucidation | — | Fatty acid TG, DG, PL, CE | 57–59 |
| DNPZ |

|
Bifunctional tagging enables detailed structural elucidation and fatty acyl chain identification | Negative | GPL | 60 |
Strategies for enhanced structural elucidation
In mass spectrometry–based lipid analysis, MS1 profiling provides accurate mass information, while MS/MS fragmentation offers structural details. However, MS/MS using collision-induced dissociation (CID) relies on efficient ion fragmentation. Many lipid classes share highly similar backbones or contain bonds that fragment poorly, making it difficult to obtain the structural features needed for confident identification. In parallel, multidimensional separation techniques, such as LC retention time (RT) and ion mobility drift time (DT), can supplement structural analysis, but lipids with only subtle structural differences often remain unresolved. In both situations, chemical derivatization can introduce reactive or diagnostic features that improve fragmentation behavior or enhance separation selectivity, thereby enabling deeper structural characterization of challenging lipid species.C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/h3_char_e001.gif) C location
C location
Unsaturated lipids containing carbon–carbon double bonds (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds) are widely distributed in biological systems, and the positional isomers of these C
C bonds) are widely distributed in biological systems, and the positional isomers of these C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds can influence multiple properties of cellular membranes.61,62 As a result, determining double-bond positions is an important aspect of lipid structural analysis. Conventional MS/MS using CID relies on the formation of diagnostic fragment ions, but C
C bonds can influence multiple properties of cellular membranes.61,62 As a result, determining double-bond positions is an important aspect of lipid structural analysis. Conventional MS/MS using CID relies on the formation of diagnostic fragment ions, but C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds have low polarity and relatively high bond dissociation energies. They therefore fail to fragment under typical CID conditions, making it difficult to distinguish positional isomers using standard lipidomics workflows.
C bonds have low polarity and relatively high bond dissociation energies. They therefore fail to fragment under typical CID conditions, making it difficult to distinguish positional isomers using standard lipidomics workflows.
Chemical derivatization provides an effective means of rendering the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond cleavable. In 2008, the Blanksby group introduced ozonolysis as a strategy to analyze unsaturated lipids.63 Ozone reacts selectively at C
C bond cleavable. In 2008, the Blanksby group introduced ozonolysis as a strategy to analyze unsaturated lipids.63 Ozone reacts selectively at C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds, generating characteristic product ions that directly reflect the position of the C
C bonds, generating characteristic product ions that directly reflect the position of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C site. Over the past decade, this approach has been extensively refined, and modern implementations can identify more than 1000 unsaturated lipid species directly from biological samples.64
C site. Over the past decade, this approach has been extensively refined, and modern implementations can identify more than 1000 unsaturated lipid species directly from biological samples.64
The Paternò–Büchi (PB) reaction, a [2 + 2] cycloaddition between a carbonyl compound and a C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond, offers another highly specific strategy for double-bond derivatization (Fig. 3a). In 2014, Ma and colleagues first applied this reaction to lipid analysis by using acetone as the derivatization reagent for unsaturated lipids.65 The resulting oxetane products yield diagnostic fragment ions upon MS/MS CID, allowing the unambiguous assignment of C
C bond, offers another highly specific strategy for double-bond derivatization (Fig. 3a). In 2014, Ma and colleagues first applied this reaction to lipid analysis by using acetone as the derivatization reagent for unsaturated lipids.65 The resulting oxetane products yield diagnostic fragment ions upon MS/MS CID, allowing the unambiguous assignment of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C positions. Beyond acetone, a variety of aromatic ketones have been introduced as PB reagents, each providing advantages for different sample types and analytical contexts.44,57,58,66 When combined with multistage tandem MS, PB chemistry can also support the elucidation of s/n-positional isomers in phospholipids.67 The PB reaction has shown broad applicability for probing double-bond isomers, and its utility has expanded into mass spectrometry imaging68–70 and single-cell lipid analysis,71 where it has enabled new biological insights. For instance, recent studies have achieved the characterization of C
C positions. Beyond acetone, a variety of aromatic ketones have been introduced as PB reagents, each providing advantages for different sample types and analytical contexts.44,57,58,66 When combined with multistage tandem MS, PB chemistry can also support the elucidation of s/n-positional isomers in phospholipids.67 The PB reaction has shown broad applicability for probing double-bond isomers, and its utility has expanded into mass spectrometry imaging68–70 and single-cell lipid analysis,71 where it has enabled new biological insights. For instance, recent studies have achieved the characterization of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C double bond positions in sebum.21 Such detailed profiling is critical for interpreting complex lipid changes, underscoring its potential in investigating neurodegenerative disorders such as Alzheimer's and Parkinson's diseases.72,73
C double bond positions in sebum.21 Such detailed profiling is critical for interpreting complex lipid changes, underscoring its potential in investigating neurodegenerative disorders such as Alzheimer's and Parkinson's diseases.72,73
 | ||
Fig. 3 (a) Analysis of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds in lipids using Paternò–Büchi reactions. Reproduced from ref. 65 with permission from Wiley, copyright 2014. (b) Several derivatization strategies can be applied to the C C bonds in lipids using Paternò–Büchi reactions. Reproduced from ref. 65 with permission from Wiley, copyright 2014. (b) Several derivatization strategies can be applied to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond, which facilitates the generation of fragments in CID. Reproduced from ref. 78 with permission from Wiley, copyright 2022. Licensed under CC BY-NC-ND 4.0. (c) Asymmetric N-tosylaziridination enhances the separation of cis/trans isomers by ion mobility. Reproduced from ref. 79 with permission from the American Chemical Society, copyright 2024. (d) High-yield thiyl radical-catalyzed C C bond, which facilitates the generation of fragments in CID. Reproduced from ref. 78 with permission from Wiley, copyright 2022. Licensed under CC BY-NC-ND 4.0. (c) Asymmetric N-tosylaziridination enhances the separation of cis/trans isomers by ion mobility. Reproduced from ref. 79 with permission from the American Chemical Society, copyright 2024. (d) High-yield thiyl radical-catalyzed C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C isomerization for analysis of cis/trans isomers. Reproduced from ref. 82 with permission from the American Chemical Society, copyright 2025. C isomerization for analysis of cis/trans isomers. Reproduced from ref. 82 with permission from the American Chemical Society, copyright 2025. | ||
Several additional derivatization strategies have been developed to target C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds. In 2018, Zhao and colleagues introduced a low-temperature plasma (LTP) probe to induce epoxidation of unsaturated fatty acids through reactive oxygen species generated from ambient air.74 This approach enabled quantitative analysis of double-bond positional isomers in human plasma. Subsequent work expanded the method to achieve rapid derivatization and to cover a broader range of phospholipid classes.75,76 In 2019, the Li group applied meta-chloroperoxybenzoic acid (m-CPBA) to perform fast and selective epoxidation of various unsaturated lipids, followed by MS/MS analysis. This strategy provided a simple and efficient means to localize C
C bonds. In 2018, Zhao and colleagues introduced a low-temperature plasma (LTP) probe to induce epoxidation of unsaturated fatty acids through reactive oxygen species generated from ambient air.74 This approach enabled quantitative analysis of double-bond positional isomers in human plasma. Subsequent work expanded the method to achieve rapid derivatization and to cover a broader range of phospholipid classes.75,76 In 2019, the Li group applied meta-chloroperoxybenzoic acid (m-CPBA) to perform fast and selective epoxidation of various unsaturated lipids, followed by MS/MS analysis. This strategy provided a simple and efficient means to localize C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds across multiple lipid subclasses.77
C bonds across multiple lipid subclasses.77
Each derivatization strategy presents distinct advantages tailored to specific analytical needs. Ozonolysis, despite its pioneering status and widespread early adoption, often necessitates specialized instrument modifications, creating a technical barrier for laboratories lacking such hardware. In contrast, the PB reaction has emerged as a widely applied method recently, although its reaction yield still needs improvement; it is favored for its high fragmentation yields and versatility, allowing for seamless integration with various analytical workflows. Epoxidation, however, offers an advantage in stereochemistry. Unlike methods that cleave the chain, it preserves the original double bond configuration, making it particularly superior for preserving and resolving the stereochemical information of geometric isomers. These advancements in C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond derivatization provide a broad toolkit for lipid structural analysis. For a more comprehensive overview of methods for analyzing C
C bond derivatization provide a broad toolkit for lipid structural analysis. For a more comprehensive overview of methods for analyzing C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds in lipids, readers are referred to a detailed review (Fig. 3b).78
C bonds in lipids, readers are referred to a detailed review (Fig. 3b).78
Separation selectivity
With the increasing use of multidimensional separation techniques in tandem with mass spectrometry, both chromatographic retention time and ion mobility drift time have become valuable for distinguishing lipid isomers. For example, MS/MS CID alone generally cannot differentiate cis and trans C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C isomers, whereas isomers with larger structural differences can sometimes be resolved by LC or ion mobility. Tang and colleagues improved the separation of Z/E isomers in ion mobility by derivatizing lipids with N-tosylaziridination reagents (Fig. 3c).79 This strategy was applied to map changes in cis–trans unsaturated lipids in bacterial membranes and to track their heterogeneous distribution along the injured spinal cord in spinal cord injury rats. The authors attributed the distinct CCS values of the stereoisomers to the formation of intramolecular hydrogen bonds within the derivatized structures. This hypothesis was theoretically corroborated by calculations based on optimized 3D conformers. The Chen group developed a series of carbonyl-based diradicals capable of inducing Z-to-E alkene isomerization of C
C isomers, whereas isomers with larger structural differences can sometimes be resolved by LC or ion mobility. Tang and colleagues improved the separation of Z/E isomers in ion mobility by derivatizing lipids with N-tosylaziridination reagents (Fig. 3c).79 This strategy was applied to map changes in cis–trans unsaturated lipids in bacterial membranes and to track their heterogeneous distribution along the injured spinal cord in spinal cord injury rats. The authors attributed the distinct CCS values of the stereoisomers to the formation of intramolecular hydrogen bonds within the derivatized structures. This hypothesis was theoretically corroborated by calculations based on optimized 3D conformers. The Chen group developed a series of carbonyl-based diradicals capable of inducing Z-to-E alkene isomerization of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds,80,81 providing MS-based lipidomics with useful indicators for distinguishing geometric isomers. Recently, Shi and co-workers reported a highly efficient method for converting geometric isomers and incorporated it into a sensitive RPLC-MS/MS workflow (Fig. 3d), revealing lipidomic alterations in Pseudomonas putida exposed to toluene, which revealed the critical role of Cti and its substrate preference.82
C bonds,80,81 providing MS-based lipidomics with useful indicators for distinguishing geometric isomers. Recently, Shi and co-workers reported a highly efficient method for converting geometric isomers and incorporated it into a sensitive RPLC-MS/MS workflow (Fig. 3d), revealing lipidomic alterations in Pseudomonas putida exposed to toluene, which revealed the critical role of Cti and its substrate preference.82
Strategies for mass-shift-based profiling
Although mass spectrometry is a powerful tool for lipidomics, its fundamental reliance on the mass-to-charge ratio (m/z) presents an intrinsic limitation: m/z is a singular scalar value that often lacks the specificity required for complex biological analysis. This limitation manifests in two critical aspects. First, the crowded lipidome is replete with isobaric species and contaminants that share identical masses, making them indistinguishable by MS1 detection alone. Second, in comparative studies, identical analytes from different biological samples yield the same signal, which precludes the possibility of differentiating sample origins during simultaneous analysis. To overcome these barriers, chemical derivatization offers a precise mechanism to manipulate the physicochemical identity of analytes by introducing a controlled and predictable “mass shift”. By conjugating lipids with specific chemical tags, researchers can artificially offset the mass of target molecules. This strategy not only segregates target analytes from the spectral background or isobaric interferences to achieve high-confidence identification (e.g., via isotope-coded derivatization) but also enables the encoding of sample identities through isotopically distinct labels for multiplexed analysis. Consequently, mass-shift-based profiling transforms a standard mass measurement into a highly specific dimension for both targeted recognition and high-throughput quantification.Improving MS1 specificity
Accurate quantitative analysis in lipidomics often relies on internal standards (IS); yet, suitable standards are frequently unavailable or prohibitively expensive. This need arises from a fundamental limitation of MS data: the m/z value alone cannot guarantee that an observed signal originates solely from the target analyte. Before high-resolution mass spectrometers became widely accessible, this issue was even more pronounced, as low-resolution MS measurements provided insufficient specificity to ensure reliable quantification. Although certain tandem MS strategies can mitigate this limitation, these approaches fall outside the scope of the present review. A widely used alternative is to employ chemical derivatization. By introducing mass tags, derivatized analytes gain enhanced specificity at the MS1 level, allowing samples to be distinguished or quantified without the need for compound-matched internal standards. This concept, often referred to as “coding”, forms the basis of many isotope-based derivatization strategies.In 2015, Narayana and colleagues derivatized FAs using four structurally similar charge tags of different masses, achieving simultaneous enhancement of ionization efficiency and improved MS1 specificity.83 However, expanding the number of channels by increasing carbon-chain length introduces structural differences that may alter chromatographic behavior, thereby limiting scalability. Isotope-coded derivatization (ICD) (Fig. 4a) offers an alternative solution: it introduces mass differences without significantly affecting the physicochemical properties of the analytes.84 An early example was reported by Tsukamoto et al. in 2005, who synthesized 7-(N,N-dimethylaminosulfonyl)-4-(aminoethyl)piperazino-2,1,3-benzoxadiazole (d6) for FA derivatization. This reagent enabled a linear dynamic range spanning four orders of magnitude, which was further expanded in subsequent studies.85,86 Many derivatization reactions discussed previously for enhancing FA ionization can also be adapted to ICD workflows, thereby improving both specificity and quantitative performance.39,40,45 The Guo group developed a DMMP reagent incorporating six deuterium atoms, providing improved MS specificity for FA analysis.87 Isotope-coded reagents have also been designed for fatty aldehydes,88,89 and later work from the Feng group demonstrated that profiles of serum carbonyl compounds may be associated with subclinical carotid atherosclerosis (SCA).90
 | ||
| Fig. 4 (a) Principle of ICD, which enables accurate duplex comparison in a single MS run. Reproduced from ref. 40 with permission from the American Chemical Society, copyright 2007. (b) Parallel quantification of PE and PS via 2-plex iTRAQ (commercial kit). Reproduced from ref. 91 with permission from Wiley, copyright 2005. Licensed under CC-BY 4.0. (c) A derivatization strategy enabling the application of AminoxyTMT (commercial kit) to phospholipids and sulfated glycolipids. Reproduced from ref. 104 with permission from Springer Nature, copyright 2022. | ||
For phospholipids that contain a primary amine, several commercial tagging reagents originally developed for proteomics can be directly applied for derivatization.91 As a lower-cost alternative, the Chen group employed acetone and deuterated acetone as labeling reagents, enabling accurate isotope-based quantification without relying on commercial kits.92,93 Methylation also offers a convenient route for introducing isotopic differences, as replacing protium with deuterium in the derivatization reagent requires minimal synthetic effort. As described earlier, cardiolipins, PIPs, and other phospholipid classes all have corresponding ICD workflows.34,94
Most ICD strategies rely on a single isotope pair and therefore support only 2-plex parallel analysis. To expand multiplexing capability, Chen and colleagues developed an 8-plex stable-isotope labeling method for sphingoids.95 The specificity introduced through ICD is not limited to MS profiling. Because the mass differences originate solely from the isotope-coded tags, any fragmentation of these tags allows neutral-loss scanning or precursor ion scanning to be used for parallel analysis as well. Building on this principle, the Feng group created a series of isotope-coded reagents for multiplexed tandem MS analysis of diverse metabolites, enabling non-targeted data acquisition without relying on MS signals.96–101
Multiplex parallel analysis
Biological applications of lipidomics often require comparing two or more sample groups, which places high demands on the parallelism and consistency of quantitative measurements. To minimize system-level variation introduced by the instrument, analytes from different samples can be labeled with distinct tags and analyzed together. Fluorescent dyes have long been used for this purpose in traditional biochemical assays, whereas mass spectrometry relies on mass-based tags for parallel analysis. In proteomics, a wide range of tandem mass tags, such as TMT102 and iTRAQ,103 have been developed to support multiplexed quantification. However, these reagents were designed to react with amino or carboxyl groups, which most lipids do not possess. As a result, early applications were limited to amine-containing lipids; for example, Berry et al. used iTRAQ to quantify PE and PS species in cultured cells (Fig. 4b).91 To overcome this limitation, the Li group developed a diazo-based tagging strategy that links phospholipids or sulfated glycolipids to commercial tandem mass tags through a click reaction (Fig. 4c), enabling multiplexed analysis beyond amine-containing lipids. Using 6-plex parallel analysis of mouse liver tissues, they characterized phospholipid alterations, pinpointing 146 species enriched in insulin-resistant obese mice and 28 in controls. While caution is warranted in translating animal data to human metabolic diseases, this study elucidates key lipid–disease associations.104 In addition, lipids with characteristic reactive groups, such as FAs, now have dedicated derivatization tags that support reliable parallel analysis.105Conclusions
Mass spectrometry-based lipidomics is a powerful and widely adopted technology for profiling cellular- and tissue-level lipid alterations. However, conventional lipidomics is often constrained by the intrinsic physicochemical properties of analytes, where the lack of easily ionizable or fragmented sites limits the ability to identify a vast number of detected features. The absence of enhanced ionization efficiency or specific structural diagnostic ions restricts the comprehensive coverage of the lipidome, particularly for low abundance species and complex isomers. Recent advances in chemical derivatization are transforming the acquisition mode of lipidomics, making it possible to extract deep structural and quantitative information from samples that were previously challenging to analyze. Strategies focused on detection sensitivity, such as methylation and charge-tagging, are particularly effective in visualizing lipid classes hidden beneath the high ionization efficiency classes. However, while these methods dramatically boost signal intensity, they primarily address abundance issues rather than structural ambiguity.In the realm of structural elucidation, reaction-based derivatization strategies have emerged to bridge the gap in isomer differentiation. Techniques such as the PB reaction and ozonolysis provide precise localization of double bonds, while other derivatization strategies provide separation of geometric isomers. This approach allows for the relative quantification of subtle structural variations that are biologically significant. Furthermore, to address the specificity and throughput limitations in quantitative analysis, mass-shift-based profiling has been widely used. These methods introduce a constant mass shift into analyte signals, enabling the identification of analytes with high confidence at the MS1 scale or multiplexed analysis. Nevertheless, the introduction of chemical reactions adds a layer of complexity to sample preparation, and the synthesis yield of these derivatizations must be carefully controlled to avoid quantitative bias. Consequently, the integration of these diverse chemical tools could establish a multidimensional framework—combining enhanced ionization, structural specificity, and quantitative precision—essential for advancing modern lipidomics.
The evolution of chemical derivatization in lipidomics can draw substantial inspiration from the mature methodologies of proteomics and metabolomics. Promising adaptations already include neutron encoding (NeuCode)106 and the use of open-modification search strategies to improve the annotation of intricate derivatized analytes.107 However, the integration of derivatization with data-independent acquisition (DIA) offers particularly transformative potential.108 This concept has been advanced in proteomics by the recent implementation of TMTpro labeling with a 4 Da mass difference, which circumvents isotopic crosstalk and enables wide-window DIA without the need for deconvolution.109 Although such high-fidelity, multi-channel workflows have not yet been fully translated to lipidomics, they offer a clear blueprint. Future efforts should focus on developing multifunctional lipid derivatization tags that incorporate similar design principles (e.g., optimized mass spacing), allowing the field to replicate these high-throughput, multiplexed capabilities.
Parallel to cross-disciplinary adaptation, the future of lipidomics lies in the integration of functionalities into unified chemical tools. Currently, most reagents optimize a single analytical metric: either sensitivity, structural resolution, or quantification. There is a need to develop next-generation, high-sensitivity, multifunctional derivatization tags. Such strategies would not only streamline the increasingly complex sample preparation workflows but also ensure that comprehensive molecular information is captured in a single analysis. Ultimately, these chemical innovations will bridge the gap between analytical discovery and clinical utility, transforming chemical derivatization-enabled lipidomics into a robust, standardized platform for deciphering the complex language of lipids in health and disease.
While the innovative methods highlighted here demonstrate immense potential, direct evidence from human clinical cohorts remains limited in some instances. This limitation stems largely from translational barriers, as cutting-edge analytical protocols are often slow to be adopted in hospital-based laboratories where patient samples are processed. We hope this review serves as a catalyst to accelerate the translation of these technologies, providing clinical researchers with powerful new tools to identify robust biomarkers and deepen our understanding of disease mechanisms.
Author contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This work was supported by the National Key R&D Program of China (2022YFC3401900).References
- D. Wang, H. M. Xiao, X. Lv, H. Chen and F. Wei, Crit. Rev. Anal. Chem., 2025, 55, 21–52 CrossRef CAS PubMed.
- O. Quehenberger, A. M. Armando and E. A. Dennis, Biochim. Biophys. Acta, Mol. Cell Biol. Lipids, 2011, 1811, 648–656 CrossRef CAS PubMed.
- E. Currie, A. Schulze, R. Zechner, T. C. Walther and R. V. Farese Jr., Cell Metab., 2013, 18, 153–161 CrossRef CAS PubMed.
- R. A. Coleman and D. G. Mashek, Chem. Rev., 2011, 111, 6359–6386 CrossRef CAS PubMed.
- K. E. Ile, G. Schaaf and V. A. Bankaitis, Nat. Chem. Biol., 2006, 2, 576–583 CrossRef CAS PubMed.
- M. G. Lete, A. Tripathi, V. Chandran, V. A. Bankaitis and M. I. McDermott, Adv. Biol. Regul., 2020, 78, 100740 CrossRef CAS PubMed.
- C. N. Serhan and B. D. Levy, J. Clin. Invest., 2018, 128, 2657–2669 CrossRef PubMed.
- M. M. S. Pallebage-Gamarallage, R. Takechi, V. Lam, S. Galloway, S. Dhaliwal and J. C. L. Mamo, Atheroscler. Suppl., 2010, 11, 49–54 CrossRef CAS PubMed.
- M. Mapstone, A. K. Cheema, M. S. Fiandaca, X. G. Zhong, T. R. Mhyre, L. H. MacArthur, W. J. Hall, S. G. Fisher, D. R. Peterson, J. M. Haley, M. D. Nazar, S. A. Rich, D. J. Berlau, C. B. Peltz, M. T. Tan, C. H. Kawas and H. J. Federoff, Nat. Med., 2014, 20, 415–418 CrossRef CAS PubMed.
- S. M. Lam, C. Zhang, Z. H. Wang, Z. Ni, S. H. Zhang, S. Y. Yang, X. H. Huang, L. S. Mo, J. Li, B. E. Lee, M. Mei, L. Huang, M. Shi, Z. Xu, F. P. Meng, W. J. Cao, M. J. Zhou, L. Shi, G. H. Chua, B. W. Li, J. B. Cao, J. Wang, S. L. Bao, Y. C. Wang, J. W. Song, F. J. Zhang, F. S. Wang and G. H. Shui, Nat. Metab., 2021, 3, 909–922 CrossRef CAS PubMed.
- J. J. Huang, J. Wang, H. He, Z. C. Huang, S. F. Wu, C. Chen, W. B. Liu, L. Xie, Y. G. Tao, L. Cong and Y. Q. Jiang, Int. J. Biol. Sci., 2021, 17, 4493–4513 CrossRef CAS PubMed.
- G. Deep and I. R. Schlaepfer, Int. J. Mol. Sci., 2016, 17(7), 1061 CrossRef PubMed.
- Y. P. Qiu, B. S. Zhou, M. M. Su, S. Baxter, X. J. Zheng, X. Q. Zhao, Y. Yen and W. Jia, Int. J. Mol. Sci., 2013, 14, 8047–8061 CrossRef PubMed.
- D. Schwudke, J. T. Hannich, V. Surendranath, V. Grimard, T. Moehring, L. Burton, T. Kurzchalia and A. Shevchenko, Anal. Chem., 2007, 79, 4083–4093 CrossRef CAS PubMed.
- M. R. Wenk, Cell, 2010, 143, 888–895 CrossRef CAS PubMed.
- X. L. Han and R. W. Gross, J. Lipid Res., 2003, 44, 1071–1079 CrossRef CAS PubMed.
- R. W. Gross and X. Han, Chem. Biol., 2011, 18, 284–291 CrossRef CAS PubMed.
- X. L. Han and R. W. Gross, Expert Rev. Proteomics, 2005, 2, 253–264 CrossRef CAS PubMed.
- X. L. Han and R. W. Gross, Mass Spectrom. Rev., 2005, 24, 367–412 CrossRef CAS PubMed.
- B. L. Su, L. F. Bettcher, W. Y. Hsieh, D. Hornburg, M. J. Pearson, N. Blomberg, M. Giera, M. P. Snyder, D. Raftery, S. J. Bensinger and K. J. Williams, J. Am. Soc. Mass Spectrom., 2021, 32, 2655–2663 CrossRef CAS PubMed.
- H. X. Shi and Y. Xia, Anal. Chem., 2024, 96, 5589–5597 CrossRef CAS PubMed.
- S. M. Camunas-Alberca, M. Moran-Garrido, J. Sáiz, A. Gil-de-la-Fuente, C. Barbas and A. Gradillas, Front. Mol. Biosci., 2023, 10, 1112521 CrossRef CAS PubMed.
- R. Almeida, J. K. Pauling, E. Sokol, H. K. Hannibal-Bach and C. S. Ejsing, J. Am. Soc. Mass Spectrom., 2015, 26, 133–148 CrossRef CAS PubMed.
- Z. Moldovan, E. Jover and J. M. Bayona, Anal. Chim. Acta, 2002, 465, 359–378 CrossRef CAS.
- J. Clark, K. E. Anderson, V. Juvin, T. S. Smith, F. Karpe, M. J. O. Wakelam, L. R. Stephens and P. T. Hawkins, Nat. Methods, 2011, 8, 267–U120 CrossRef CAS PubMed.
- A. Kielkowska, I. Niewczas, K. E. Anderson, T. N. Durrant, J. Clark, L. R. Stephens and P. T. Hawkins, Adv. Biol. Regul., 2014, 54, 131–141 CrossRef CAS PubMed.
- S. H. Kim, H. E. Song, S. J. Kim, D. C. Woo, S. Chang, W. G. Choi, M. J. Kim, S. H. Back and H. J. Yoo, J. Lipid Res., 2017, 58, 469–478 CrossRef CAS PubMed.
- G. Q. Gong, B. Bilanges, B. Allsop, G. R. Masson, V. Roberton, T. Askwith, S. Oxenford, R. R. Madsen, S. E. Conduit, D. Bellini, M. Fitzek, M. Collier, O. Najam, Z. H. He, B. Wahab, S. H. McLaughlin, A. W. E. Chan, I. Feierberg, A. Madin, D. Morelli, A. Bhamra, V. Vinciauskaite, K. E. Anderson, S. Surinova, N. Pinotsis, E. Lopez-Guadamillas, M. Wilcox, A. Hooper, C. Patel, M. A. Whitehead, T. D. Bunney, L. R. Stephens, P. T. Hawkins, M. Katan, D. M. Yellon, S. M. Davidson, D. M. Smith, J. B. Phillips, R. Angell, R. L. Williams and B. Vanhaesebroeck, Nature, 2023, 618(7963), 159–168 CrossRef CAS PubMed.
- S. Morioka, H. Nakanishi, T. Yamamoto, J. Hasegawa, E. Tokuda, T. Hikita, T. Sakihara, Y. Kugii, C. Oneyama, M. Yamazaki, A. Suzuki, J. Sasaki and T. Sasaki, Nat. Commun., 2022, 13(1), 83 CrossRef CAS PubMed.
- K. V. Wasslen, C. R. Canez, H. Lee, J. M. Manthorpe and J. C. Smith, Anal. Chem., 2014, 86, 9523–9532 CrossRef CAS PubMed.
- C. R. Canez, S. W. J. Shields, M. Bugno, K. V. Wasslen, H. P. Weinert, W. G. Willmore, J. M. Manthorpe and J. C. Smith, Anal. Chem., 2016, 88, 6996–7004 CrossRef CAS PubMed.
- S. W. J. Shields, C. A. Rosales, J. A. Roberts, P. J. Pallister, K. Wasslen, J. M. Manthorpe and J. C. Smith, Anal. Chem., 2021, 93, 1084–1091 CrossRef CAS PubMed.
- J. A. Roberts, C. A. Rosales, K. V. Wasslen, A. S. Radnoff, E. Godbout, J. S. Diallo, J. M. Manthorpe and J. C. Smith, J. Am. Soc. Mass Spectrom., 2023, 34, 2722–2730 CrossRef CAS PubMed.
- J. C. Lee, S. K. Byeon and M. H. Moon, Anal. Chem., 2017, 89, 4969–4977 CrossRef CAS PubMed.
- D. Kale, F. Kikul, P. Phapale, L. Beedgen, C. Thiel and B. Brügger, Anal. Chem., 2023 DOI:10.1021/acs.analchem.2c03623.
- J. W. Lee, T. Yamamoto, T. Uchikata, A. Matsubara, E. Fukusaki and T. Bamba, J. Sep. Sci., 2011, 34, 3553–3560 CrossRef CAS PubMed.
- J. W. Lee, S. Nishiumi, M. Yoshida, E. Fukusaki and T. Bamba, J. Chromatogr., A, 2013, 1279, 98–107 CrossRef CAS PubMed.
- T. Xia, H. L. Ren, W. P. Zhang and Y. Xia, Anal. Chim. Acta, 2020, 1128, 107–115 CrossRef CAS PubMed.
- W. C. Yang, J. Adamec and F. E. Regnier, Anal. Chem., 2007, 79, 5150–5157 CrossRef CAS PubMed.
- S. M. Lamos, M. R. Shortreed, B. L. Frey, P. J. Belshaw and L. M. Smith, Anal. Chem., 2007, 79, 5143–5149 CrossRef CAS PubMed.
- X. N. Li and A. A. Franke, Anal. Chem., 2011, 83, 3192–3198 CrossRef CAS PubMed.
- M. Wang, R. H. Han and X. L. Han, Anal. Chem., 2013, 85, 9312–9320 CrossRef CAS PubMed.
- C. F. Hu, M. Wang, Q. Duan and X. L. Han, Anal. Chim. Acta, 2020, 1105, 105–111 CrossRef CAS PubMed.
- J. Zhao, M. X. Fang and Y. Xia, J. Lipid Res., 2021, 62, 100110 CrossRef CAS PubMed.
- A. Koulman, D. Petras, V. K. Narayana, L. Wang and D. A. Volmer, Anal. Chem., 2009, 81, 7544–7551 CrossRef CAS PubMed.
- M. Liu, F. Wei, X. Lv, X. Y. Dong and H. Chen, Food Chem., 2018, 242, 338–344 CrossRef CAS PubMed.
- F. X. Sun, A. A. Choi and R. H. Wu, J. Proteome Res., 2018, 17, 1606–1614 CrossRef CAS PubMed.
- R. Jiang, Y. Jiao, P. Zhang, Y. Liu, X. Wang, Y. Huang, Z. Zhang and F. Xu, Anal. Chem., 2017, 89, 12223–12230 CrossRef CAS PubMed.
- M. Eggink, M. Wijtmans, R. Ekkebus, H. Lingeman, I. J. P. de Esch, J. Kool, W. M. A. Niessen and H. Irth, Anal. Chem., 2008, 80, 9042–9051 CrossRef CAS PubMed.
- C. Tie, T. Hu, Z. X. Jia and J. L. Zhang, Anal. Chem., 2016, 88, 7762–7768 CrossRef CAS PubMed.
- W. S. Qi, Y. J. Wang, Y. Q. Cao, Y. J. Cao, Q. Guan, T. Q. Sun, L. Zhang and Y. L. Guo, Anal. Chem., 2020, 92, 8644–8648 CrossRef CAS PubMed.
- T. Y. Zhang, S. Li, Q. F. Zhu, Q. Wang, D. Hussain and Y. Q. Feng, TrAC, Trends Anal. Chem., 2019, 119, 115608 CrossRef CAS.
- S. Zhao and L. Li, TrAC, Trends Anal. Chem., 2020, 131, 115988 CrossRef CAS.
- Y. Z. Wang, H. M. Xiao, P. R. Bai, X. Liu, X. Liu, Q. F. Zhu and Y. Q. Feng, Anal. Chem., 2025, 97, 9600–9605 CrossRef CAS PubMed.
- O. Lavrynenko, R. Nedielkov, H. M. Möller and A. Shevchenko, J. Lipid Res., 2013, 54, 2265–2272 CrossRef CAS PubMed.
- X. X. He, C. L. Huang and E. H. Schuchman, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2009, 877, 983–990 CrossRef CAS PubMed.
- P. Esch and S. Heiles, J. Am. Soc. Mass Spectrom., 2018, 29, 1971–1980 CrossRef CAS PubMed.
- X. B. Xie, J. Zhao, M. Lin, J. L. Zhang and Y. Xia, Anal. Chem., 2020, 92, 8487–8496 CrossRef CAS PubMed.
- T. Xia, F. Zhou, D. H. Zhang, X. Jin, H. X. Shi, H. Yin, Y. Q. Gong and Y. Xia, Nat. Commun., 2023, 14(1), 4263 CrossRef CAS PubMed.
- C.-L. Tsai, X. Chen, R. G. Reddy and X. Yan, Anal. Chem., 2025, 97, 8992–8999 CrossRef CAS PubMed.
- P. V. Subbaiah, D. Sircar, B. Aizezi and E. Mintzer, Biochim. Biophys. Acta, Biomembr., 2010, 1798, 506–514 CrossRef CAS PubMed.
- K. Vriens, S. Christen, S. Parik, D. Broekaert, K. Yoshinaga, A. Talebi, J. Dehairs, C. Escalona-Noguero, R. Schmieder, T. Cornfield, C. Charlton, L. Romero-Pérez, M. Rossi, G. Rinaldi, M. F. Orth, R. Boon, A. Kerstens, S. Y. Kwan, B. Faubert, A. Méndez-Lucas, C. C. Kopitz, T. Chen, J. Fernandez-Garcia, J. A. G. Duarte, A. A. Schmitz, P. Steigemann, M. Najimi, A. Hägebarth, J. A. Van Ginderachter, E. Sokal, N. Gotoh, K. K. Wong, C. Verfaillie, R. Derua, S. Munck, M. Yuneva, L. Beretta, R. J. DeBerardinis, J. V. Swinnen, L. Hodson, D. Cassiman, C. Verslype, S. Christian, S. Grünewald, T. G. P. Grünewald and S. M. Fendt, Nature, 2019, 566, 403 CrossRef PubMed –+.
- M. C. Thomas, T. W. Mitchell, D. G. Harman, J. M. Deeley, J. R. Nealon and S. J. Blanksby, Anal. Chem., 2008, 80, 303–311 CrossRef CAS PubMed.
- J. A. Michael, R. S. E. Young, R. Balez, L. J. Jekimovs, D. L. Marshall, B. L. J. Poad, T. W. Mitchell, S. J. Blanksby, C. S. Ejsing and S. R. Ellis, Angew. Chem., Int. Ed., 2024, 63(9), e202316793 CrossRef CAS PubMed.
- X. X. Ma and Y. Xia, Angew. Chem., Int. Ed., 2014, 53, 2592–2596 CrossRef CAS PubMed.
- H. Shi, Z. Tan, X. Guo, H. Ren, S. Wang and Y. Xia, Anal. Chem., 2023, 95, 5117–5125 CrossRef CAS PubMed.
- W. B. Cao, S. M. Cheng, J. Yang, J. X. Feng, W. P. Zhang, Z. S. Li, Q. H. Chen, Y. Xia, Z. Ouyang and X. X. Ma, Nat. Commun., 2020, 11(1), 375 CrossRef CAS PubMed.
- F. Tang, C. G. Guo, X. X. Ma, J. Zhang, Y. Su, R. Tian, R. Y. Shi, Y. Xia, X. H. Wang and Z. Ouyang, Anal. Chem., 2018, 90, 5612–5619 CrossRef CAS PubMed.
- F. Wäldchen, F. Mohr, A. H. Wagner and S. Heiles, Anal. Chem., 2020, 92, 14130–14138 CrossRef PubMed.
- A. Bednarík, S. Bölsker, J. Soltwisch and K. Dreisewerd, Angew. Chem., Int. Ed., 2018, 57, 12092–12096 Search PubMed.
- Y. L. Zhu, W. H. Wang and Z. B. Yang, Anal. Chem., 2020, 92, 11380–11387 CrossRef CAS PubMed.
- D. Sarkar, E. Sinclair, S. H. Lim, C. Walton-Doyle, K. Jafri, J. Milne, J. P. C. Vissers, K. Richardson, D. K. Trivedi, M. Silverdale and P. Barran, JACS Au, 2022, 2(9), 2013–2022, DOI:10.1021/jacsau.2c00300.
- S. Briganti, M. Truglio, A. Angiolillo, S. Lombardo, D. Leccese, E. Camera, M. Picardo and A. Di Costanzo, Metabolites, 2021, 11(12), 819 CrossRef CAS PubMed.
- Y. Y. Zhao, H. S. Zhao, X. Zhao, J. Jia, Q. Ma, S. C. Zhang, X. R. Zhang, H. Chiba, S. P. Hui and X. X. Ma, Anal. Chem., 2017, 89, 10270–10278 CrossRef CAS PubMed.
- X. Zhao, Y. Y. Zhao, L. Zhang, X. X. Ma, S. C. Zhang and X. R. Zhang, Anal. Chem., 2018, 90, 2070–2078 CrossRef CAS PubMed.
- W. B. Cao, X. X. Ma, Z. S. Li, X. Y. Zhou and Z. Ouyang, Anal. Chem., 2018, 90, 10286–10292 CrossRef CAS PubMed.
- Y. Feng, B. M. Chen, Q. Y. Yu and L. J. Li, Anal. Chem., 2019, 91, 1791–1795 CrossRef CAS PubMed.
- W. Zhang, R. Jian, J. Zhao, Y. Liu and Y. Xia, J. Lipid Res., 2022, 63(7), 100219 CrossRef CAS PubMed.
- S. Tang, H. Wang, H. H. Zhang, M. Y. Zhang, J. C. Xu, C. Yang, X. S. Chen and X. H. Guo, J. Am. Chem. Soc., 2024, 146, 29503–29512 CrossRef CAS PubMed.
- R. R. Fu, G. F. Feng, L. W. Wang, M. L. Hou, Z. J. Tang, X. Li, C. S. Xu, X. T. Qi, G. Y. Xu and S. M. Chen, Anal. Chem., 2025, 97, 555–564 CrossRef CAS PubMed.
- G. F. Feng, M. Gao, L. W. Wang, J. Y. Chen, M. L. Hou, Q. Q. Wan, Y. Lin, G. Y. Xu, X. T. Qi and S. M. Chen, Nat. Commun., 2022, 13(1), 2652 CrossRef CAS PubMed.
- H. X. Shi, H. J. Bai, H. T. Deng and Y. Xia, Anal. Chem., 2025, 97, 10378–10387 CrossRef CAS PubMed.
- V. K. Narayana, V. M. Tomatis, T. Wang, D. Kvaskoff and F. A. Meunier, Chem. Biol., 2015, 22, 1552–1561 CrossRef CAS PubMed.
- P. Bruheim, H. F. N. Kvitvang and S. G. Villas-Boas, J. Chromatogr., A, 2013, 1296, 196–203 CrossRef CAS PubMed.
- Y. Tsukamoto, T. Santa, H. Yoshida, H. Miyano, T. Fukushima, K. Hirayama, K. Imai and T. Funatsu, Biomed. Chromatogr., 2006, 20, 1049–1055 CrossRef CAS PubMed.
- Y. Tsukamoto, T. Santa, H. Yoshida, H. Miyano, T. Fukushima, K. Hirayama, K. Imai and T. Funatsu, Biomed. Chromatogr., 2006, 20, 358–364 CrossRef CAS PubMed.
- J. P. Leng, H. Y. Wang, L. Zhang, J. Zhang, H. Wang and Y. L. Guo, Anal. Chim. Acta, 2013, 758, 114–121 CrossRef CAS PubMed.
- N. Guo, C. Y. Peng, Q. F. Zhu, B. F. Yuan and Y. Q. Feng, Anal. Chim. Acta, 2017, 967, 42–51 CrossRef CAS PubMed.
- S. Zhao, M. Dawe, K. Guo and L. Li, Anal. Chem., 2017, 89, 6758–6765 CrossRef CAS PubMed.
- D. M. Wu, J. J. Zhang, N. Guo, S. J. Zheng, Q. F. Zhu, Y. Q. Feng, S. S. Peng, Y. H. Wu and M. H. Tuo, NeuroReport, 2018, 29, 1550–1557 CrossRef CAS PubMed.
- K. A. Z. Berry and R. C. Murphy, J. Lipid Res., 2005, 46, 1038–1046 CrossRef PubMed.
- H. F. Ma, F. Wei, B. F. Wu, C. Yang, Y. Xie, Z. Y. Wu, X. Lv and H. Chen, J. Lipid Res., 2019, 60, 121–134 CrossRef CAS PubMed.
- X. Wang, F. Wei, J. Q. Xu, X. Lv, X. Y. Dong, X. L. Han, S. Y. Quek, F. H. Huang and H. Chen, Anal. Chim. Acta, 2016, 902, 142–153 CrossRef CAS PubMed.
- T. X. Cai, Q. B. Shu, J. J. Hou, P. B. Liu, L. L. Niu, X. J. Guo, C. C. Liu and F. Q. Yang, Anal. Chem., 2015, 87, 513–521 CrossRef CAS PubMed.
- S. E. Chen, S. Y. Zhu, J. W. Hu, J. Sun, Z. J. Zheng, X. E. Zhao and H. W. Liu, Anal. Chim. Acta, 2020, 1124, 40–51 CrossRef CAS PubMed.
- Q. F. Zhu, Z. Zhang, P. Liu, S. J. Zheng, K. Peng, Q. Y. Deng, F. Zheng, B. F. Yuan and Y. Q. Feng, J. Chromatogr., A, 2016, 1460, 100–109 CrossRef CAS PubMed.
- P. Liu, C. B. Qi, Q. F. Zhu, B. F. Yuan and Y. Q. Feng, Sci. Rep., 2016, 6(1), 21443 CrossRef PubMed.
- L. Yu, P. Liu, Y. L. Wang, Q. W. Yu, B. F. Yuan and Y. Q. Feng, Analyst, 2015, 140, 5276–5286 RSC.
- J. M. Chu, C. B. Qi, Y. Q. Huang, H. P. Jiang, Y. H. Hao, B. F. Yuan and Y. Q. Feng, Anal. Chem., 2015, 87, 7364–7372 CrossRef CAS PubMed.
- P. Liu, Y. Q. Huang, W. J. Cai, B. F. Yuan and Y. Q. Feng, Anal. Chem., 2014, 86, 9765–9773 CrossRef CAS PubMed.
- Y. Q. Huang, Q. Y. Wang, J. Q. Liu, Y. H. Hao, B. F. Yuan and Y. Q. Feng, Analyst, 2014, 139, 3446–3454 RSC.
- A. Thompson, J. Schäfer, K. Kuhn, S. Kienle, J. Schwarz, G. Schmidt, T. Neumann and C. Hamon, Anal. Chem., 2003, 75, 1895–1904 CrossRef CAS PubMed.
- P. L. Ross, Y. L. N. Huang, J. N. Marchese, B. Williamson, K. Parker, S. Hattan, N. Khainovski, S. Pillai, S. Dey, S. Daniels, S. Purkayastha, P. Juhasz, S. Martin, M. Bartlet-Jones, F. He, A. Jacobson and D. J. Pappin, Mol. Cell. Proteomics, 2004, 3, 1154–1169 CrossRef CAS PubMed.
- T. J. Gu, P. K. Liu, Y. W. Wang, M. T. Flowers, S. L. Xu, Y. Liu, D. B. Davis and L. J. Li, Nat. Chem., 2024, 16(5), 762–770 CrossRef CAS PubMed.
- Y. Feng, Y. N. Lv, T. J. Gu, B. M. Chen and L. J. Li, Anal. Chem., 2022, 94(38), 13036–13042, DOI:10.1021/acs.analchem.2c01917.
- G. K. Potts, E. A. Voigt, D. J. Bailey, C. M. Rose, M. S. Westphall, A. S. Hebert, J. Yin and J. J. Coon, Anal. Chem., 2016, 88, 3295–3303 CrossRef CAS PubMed.
- G. A. Vitale, S. N. Xia, K. Dührkop, M. R. Z. Shahneh, H. Brötz-Oesterhelt, Y. Mast, C. Brungs, S. Böcker, R. Schmid, M. X. Wang, C. C. Hughes and D. Petras, Nat. Commun., 2025, 16(1), 6911 CrossRef CAS PubMed.
- J. Derks, A. Leduc, G. Wallmann, R. G. Huffman, M. Willetts, S. Khan, H. Specht, M. Ralser, V. Demichev and N. Slavov, Nat. Biotechnol., 2023, 41(1), 50–59 CrossRef CAS PubMed.
- Z. Wang, P.-K. Liu, H. Lu and L. Li, Anal. Chem., 2025, 97, 26419–26428 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |
