 Open Access Article
Open Access ArticleBacterial imprinted polymer-based detection of Escherichia coli using polydopamine on gold nanodendrite/graphene oxide modified electrodes
Yilei
Xue
ab,
Jessica
Chen
a,
Meissam
Noroozifar
a,
Ruby May A.
Sullan
 ab and
Kagan
Kerman
ab and
Kagan
Kerman
 *ab
*ab
aDepartment of Physical and Environmental Sciences, University of Toronto Scarborough, 1265 Military Trail, Toronto, Ontario M1C 1A4, Canada. E-mail: kagan.kerman@utoronto.ca
bDepartment of Chemistry, University of Toronto, 80 St. George Street, Toronto, Ontario M5S 3H6, Canada
First published on 26th March 2026
Abstract
Pathogenic bacteria pose a major global threat to public health, environmental safety, and food quality, highlighting the need for rapid, sensitive, and cost-effective detection methods. In this study, an electrochemical biosensor based on a glassy carbon electrode modified with graphene oxide (GO), gold nanodendrites (AuNDs), and polydopamine (PDA) was developed with Escherichia coli imprinted on the surface. The sensor combines the high conductivity of AuNDs and GO with the molecular recognition capability of the imprinted PDA layer, providing excellent selectivity toward E. coli. By optimizing the bacterial removal method and the thickness of PDA film, the biosensor exhibited a wide linear detection range (1.0 × 101–1.0 × 104 CFU mL−1) and a remarkably low detection limit of 1.5 CFU mL−1. It demonstrated high recovery (96–103%) when tested with real creek water and wastewater samples. The proposed bacterial imprinted polymer-based biosensor offers a promising platform for portable detection of E. coli in environmental monitoring applications.
1. Introduction
Foodborne and waterborne diseases pose a substantial global health burden, with transmission commonly occurring via ingestion, aerosolization, or contact with contaminated water containing bacterial, viral, or protozoan pathogens.1 The World Health Organization's Foodborne Disease Burden Epidemiology Reference Group estimates that unsafe food sickens roughly 600 million people, about 1 in 10, annually, with the heaviest toll concentrated in low- and middle-income regions.2 Among these agents, such as bacteria, viruses, and protozoa, bacteria are the leading cause of waterborne illness. Salmonella spp., Listeria monocytogenes, Staphylococcus aureus, and Escherichia coli in water, dairy, seafood, and fruits are the causative agents of two-thirds of foodborne illnesses worldwide, especially in developing countries.2 Conventional detection methods for pathogenic bacteria associated with waterborne illness, such as culture-based techniques and immunoassays, are time-consuming and labor-intensive because they require growing a visible colony from a single bacterial cell to amplify the signal.3 Alternative DNA technologies, such as polymerase chain reaction, have been developed for the sensitive and reliable detection of pathogenic bacteria.4 However, these methods are limited by assay time, high cost, and the enrichment of pathogens, which are often present in environmental samples in low amounts.5To prevent the spread of pathogenic bacteria and monitor water quality, there is an urgent need for cost-effective, rapid pathogen detection. Biosensors have emerged as a promising alternative for bacterial detection due to their high sensitivity, rapid response, and straightforward sample preparation.6 Biosensors use a bioreceptor to interact with the target analyte and a transducer to convert a biological response into a measurable signal.7 Among the transducers, electrochemical measurements have the potential to be easily integrated into miniaturized, portable systems, enabling real-time quantification of label-free samples.8,9 Electrochemical biosensors with a wide variety of receptors, including antibodies,10 enzymes,11 phages,12 and nucleic acids,13 have been under development for pathogenic bacteria detection by numerous researchers.
Nevertheless, biomolecules are often limited by their complex preparation, high cost, low pH, and temperature instability.14 One innovative approach to enhancing the sensitivity and selectivity of electrochemical biosensors is the use of molecularly imprinted polymers (MIPs). MIPs are synthetic polymers engineered to match the conformational features of the target molecule, such as size, shape, and functional groups, prepared by polymerizing functional monomers and cross-linkers around a template, which is later removed, leaving selective recognition sites for the target.15 Compared with biomolecules, MIPs exhibit greater stability, lower cost, and resistance to harsh environmental conditions such as temperature, pH, and chemical degradation as biorecognition layers.16 Bacteria-imprinted polymers are a subcategory of MIPs, adapted for large biological targets, and also create template-defined recognition sites. Due to the scale and complexity of whole bacterial cells, it is challenging to fully encapsulate or remove them from a bulk polymer, and bacteria imprinting is commonly fabricated via surface imprinting.17 In this method, bacterial cells act as molds on the polymer surface, forming cavities that complement both the cell envelope's morphological shape and key chemical features. Integrating MIPs with electrochemical techniques has further advanced the development of sensitive, selective detection platforms for long-term use. The polymer layer can be directly deposited on a transducer surface via different methods such as electrochemical polymerization, drop casting, layer-by-layer deposition, self-assembly, and in situ chemical polymerization.18
Recent studies have demonstrated that modifying the transducer surface with nanomaterials can significantly improve the sensitivity of MIP-based electrochemical biosensors.19 Carbon materials, such as carbon nanotubes, graphene, and graphene oxide, feature densely packed hexagonal crystal lattices in which carbon atoms are predominantly connected via sp2 hybridization.20 However, the presence of defects or edge sites can introduce sp3-hybridized carbon atoms, slightly disrupting the uniform structure.20 This structure imparts excellent thermal and chemical stability, a high surface area, and superior electrical conductivity. Liu et al.21 reported that the nanosheet structure of graphene oxide significantly enhanced the sensitivity of the MIP sensor of testosterone. In addition, metallic nanoparticles with porous, branched structures exhibit high conductivity, facilitating electron transfer and enhancing the sensor's sensitivity.22
This preliminary work presents the successful design of a molecularly imprinted electrochemical biosensor for the selective recognition of E. coli. The device was fabricated by modifying a glassy carbon electrode (GCE) via layer-by-layer assembly of graphene oxide (GO) and gold nanodendrites (AuNDs), followed by the electrochemical polymerization of dopamine (DA). The integration of AuNDs into the polydopamine (PDA) layer significantly enhanced the sensor's electrical conductivity and sensitivity. DA was selected for its excellent biocompatibility, self-polymerization, and versatile functional groups, making it an effective monomer for molecular imprinting.23,24 PDA can be prepared by electropolymerization under mild aqueous conditions, without the need for additional linkers, thereby preserving bacterial template integrity.25 The functional groups in PDA interact with the bacterial cell wall via noncovalent interactions during polymerization, such as hydrogen bonding, π–π stacking, van der Waals forces, and electrostatic interactions,26 creating precise chemical matching of the bacteria's surface. PDA was identified as the optimal functional film for E. coli imprinting by density functional theory calculations, which confirmed that PDA exhibited the most favorable binding energy with lipopolysaccharides on the E. coli cell envelope compared to other conductive polymers, such as polypyrrole and polyaniline, underscoring its superior suitability for forming specific, complementary recognition sites.27 Overall, the developed platform demonstrated promise for future adaptation as a portable, point-of-care device for rapid detection of E. coli in environmental and food safety applications.
2. Experimental
2.1. Chemicals and reagents
Dopamine hydrochloride, potassium ferricyanide (K3[Fe(CN)6]), potassium ferrocyanide trihydrate (K4[Fe(CN)6]·3H2O), potassium nitrate (KNO3), potassium chloride (KCl), potassium hydroxide (KOH), nitric acid (HNO3), acetic acid (CH3COOH), graphene oxide (GO), chloroauric acid, NafionTM, sodium dodecyl sulfate (SDS), zinc dust, and Luria–Bertani (LB) broth (Miller) were all purchased from Sigma-Aldrich (Oakville, ON, Canada). Phosphoric acid (H3PO4) was obtained from Thermo Fisher Scientific (Mississauga, ON, Canada). Concentrated hydrochloric acid (HCl) was procured from Caledon Laboratories Ltd (Georgetown, ON, Canada).2.2. Instrumentation
All measurements were performed using a three-electrode system, where a modified glassy carbon electrode (GCE) or screen-printed electrode (SPE), platinum wire, and a silver/silver chloride (Ag/AgCl, 3 M KCl) reference electrode (Metrohm AG, Herisau, Switzerland) served as the working, auxiliary, and reference electrodes, respectively. Cyclic voltammetry (CV), differential pulse voltammetry (DPV), and electrochemical impedance spectroscopy (EIS) measurements were conducted using Autolab PGSTAT302N potentiostat/galvanostat (Metrohm AG, Herisau, Switzerland), and the measurements were processed by NOVATM 2.1.2 (Metrohm AG, Herisau, Switzerland) software. To visualize morphological changes, the GCE was imaged using a Hitachi S-530 scanning electron microscope (SEM) (Hitachi, Chiyoda, Tokyo, Japan) and an atomic force microscope (AFM) after each modification step. The SEM images were obtained through secondary electron analysis. All samples were coated with an osmium conductive layer before imaging.A JPK NanoWizard® 4 (JPK Instruments, Berlin, Germany) coupled with an inverted optical fluorescence microscope (Zeiss Axio Observer 7) was utilized to image the morphology of SPEs via the Quantitative Imaging (QI) modality in PBS solution. An MLCT-bio probe with a spring constant equal to 0.6 N m−1 and a 20 nm tip radius was selected for imaging. A Thermo Fisher Scientific K-Alpha XPS spectrometer (Thermo Fisher Scientific-E, Grinstead, UK) was used to obtain the X-ray photoelectron spectroscopy (XPS) measurements. The parameters of the survey spectra were a nominal 400 μm spot, 200 eV pass energy (PE), 1 eV step size, and 150 eV pass energy with a 0.2 eV step size in the spectral regions of interest. Fluorescent imaging was performed using a Zeiss Axio Observer 7 with a filter set 92 HE LED. Excitation was provided by a 591/27 nm LED, and emission was collected through a long-pass filter at 688/145 nm.
2.3. Synthesis of MIP
Before modification, GCEs were polished using 1, 0.3, and 0.05 µm Alumina powders for 15 min in sequence. 3 μL of a 0.5 mg mL−1 GO solution was drop-cast on the working area of GCEs and dried for 1 h. The GCE underwent 10 cycles of CV ranging from −0.2 to +0.8 V in 10 mM KCl with a scan rate of 0.1 V s−1. Before dipping the GCE in a 25 mM HAuCl3·3H2O solution for 30 min, 10 μL of a 0.05% (w/w) Zn solution in 0.1% (w/v) NafionTM was drop-cast on the electrode surface and dried for 1 h. Following, the GCE was washed with 2 M HCl and water. The electrical polymerization of the electrode was performed in a 0.2 M phosphate buffer solution (pH 7.4) containing 5 mM DA and 1.0 × 108 CFU mL−1 of E. coli using cyclic voltammetry from −0.5 to +0.5 V for 10 cycles at a scan rate of 0.02 V s−1 after 15 min of degassing with nitrogen. To remove the bacteria templates, the electrode was sonicated in 0.01 equiv. M HNO3/SDS solution for 20 min, then rinsed with water. A non-imprinted polymer (NIP)-modified electrode was prepared using the same method but without adding the bacteria.2.4. Real sample preparation
Creek water and wastewater were prepared to demonstrate the detection of target bacteria in real-world scenarios. The creek water was obtained from Highland Creek (Toronto, Canada) and was diluted five-fold with 0.2 M PBS (pH 7.4) prior to measurements. Wastewater was obtained from Highland Creek Treatment Plant (Toronto, Canada) and diluted ten-fold with 0.2 M PBS (pH 7.4). Each sample was spiked with the target bacteria, and analyte recovery percentages were quantified using the standard addition method.2.5. Liquid culture preparation
Frozen bacterial stocks of E. coli AR3110, E. coli DH5α, Pseudomonas defensor WCS374r0, and Bacillus velezensis VFb49 were prepared in LB broth with 25% (v/v) glycerol and stored at −80 °C. P. defensor and B. velezensis were provided by Dr. Keiko Yoshioka's laboratory from the University of Toronto as a gift. Bacterial cells were first streaked on LB agar from frozen stocks and incubated for 24 h. The E. coli strains were grown at 37 °C, and P. defensor and B. velezensis were incubated at 28 °C. Liquid cultures of each strain were initiated by inoculating a single colony into fresh LB solution and incubating for 24 h. Bacterial cells were harvested by centrifugation at 2000 rpm for 5 min, washed 3 times with PBS, and diluted to a specific OD600 in PBS.3. Results and discussion
3.1. Characterization of the surface morphology and composition
To enhance electrode conductivity, nanomaterials, including GO and AuNDs, were applied to modify the electrode surface, and the step-by-step modifications of the MIP were thoroughly characterized using electrochemical (CV, EIS), spectroscopic (XPS), and high-resolution imaging (SEM and AFM) techniques. The morphological changes of the electrode surface resulting from modifications with MIP and NIP were evaluated using SEM, and the results are shown in Fig. 1a–d and S1, with a variety of magnifications. As shown in Fig. 1a, the SEM displays the sheet-like porous structure of the graphene oxide layer of GO, and a dendrite-like structure of the AuNDs that formed on top of the graphene oxide layer was observed on the GO/AuNDs surface, confirming successful synthesis and immobilization of AuNDs (Fig. 2b). A comparison of Fig. 1c and d, showing the bacteria-imprinted PDA film before and after bacteria removal, respectively, revealed the formation of cavities on the polymer surface, indicating successful removal. A NIP-modified electrode was prepared as a control, as shown in Fig. S1i and j, where the PDA was deposited on the GO/AuNDs modified electrodes.The morphological changes on the electrode surface after each modification were analyzed using AFM to visualize the smaller features, as shown in Fig. 1e–l. Initially, GO was drop-cast onto the electrode surface, revealing its characteristic sheet-like structure (Fig. 1e). The successful immobilization of AuNDs was confirmed by the visualization of their branched architecture (Fig. 1f). In the presence of the bacteria, the polymerization entrapped the bacterial cells in solution, as shown in Fig. 1g–i. The bacterial cells were measured approximately 1.5 ± 0.3 µm in length and 523 ± 85 nm in height (Fig. 1h). After washing off the trapped bacteria, their imprinted patterns were distinctly observed on the electrode surface (Fig. 1h–k), indicating the successful preparation of the MIP. Cross-sectional analysis further estimated the height of the imprinted cavity to be 24 ± 2 nm (Fig. 1l). Electropolymerization accurately controlled the PDA deposition thickness to the nanometer scale, typically ranging from 10 nm to 45 nm, determined by the experimental design.28–30 In thin PDA films, bacterial cells were partially embedded during polymerization, creating near-surface binding sites that allow bacterial templates to be removed efficiently and completely, and creating accessible binding sites.26
XPS was used for the analysis of the elemental composition, empirical formula, chemical state, and electronic state of the elements. The chemical compositions of the GCE modified with GO, GO/AuNDs, GO/AuNDs/PDA/Bacteria, and GO/AuNDs/PDA/(w/o) Bacteria, where bacteria were washed off, were characterized by XPS, as shown in Fig. 2a, revealing the presence of Au 4f, C 1s, N 1s, and O 1s on the surface of the respective catalysts. The two characteristic peaks appearing at 83.7 and 87.4 eV corresponded to the metallic state of Au 3d5/2 and 3d3/2, as observed in Fig. 2b for the GCE modified with GO/AuNDs and GO/AuNDs/PDA/Bacteria. The intensity of these two characteristic peaks for the latter modification was noticeably lower than that of GO/AuNDs, which was attributed to the presence of PDA, regardless of whether bacteria were present or absent. The high-resolution spectra for Au 4f, C 1s, N 1s, and O 1s are shown in Fig. 2b–e, respectively. The C 1s peak observed at approximately 284 eV was attributed to four components located at 284.9 eV (sp2 C–C bonds), 285.6 eV (C–OH bonds), 286.9 eV (C–O–C bonds), and 288.8 eV (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds) for GO (Fig. 2c and Fig. S2l).20 After the deposition of the PDA layer, the peak at 285.6 eV corresponded to C–N bonds in PDA, while the fitting located at 287.7 eV was attributed to C–O bonds (Fig. S2e and h). The binding energies associated with the oxygen-containing functional groups were observed at 285.6 eV C–OH, 286.8 eV for C–O–C, 288.2 eV for C
O bonds) for GO (Fig. 2c and Fig. S2l).20 After the deposition of the PDA layer, the peak at 285.6 eV corresponded to C–N bonds in PDA, while the fitting located at 287.7 eV was attributed to C–O bonds (Fig. S2e and h). The binding energies associated with the oxygen-containing functional groups were observed at 285.6 eV C–OH, 286.8 eV for C–O–C, 288.2 eV for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and 289.4 eV for O
O, and 289.4 eV for O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C–OH (Fig. S2).31,32 The N 1s spectrum of GO/AuNDs/PDA/(w/o) Bacteria included two peaks at 399.7 eV and 401.9 eV for –NH2 and –NH–, respectively (Fig. 2d and Fig. S2d).22–25
C–OH (Fig. S2).31,32 The N 1s spectrum of GO/AuNDs/PDA/(w/o) Bacteria included two peaks at 399.7 eV and 401.9 eV for –NH2 and –NH–, respectively (Fig. 2d and Fig. S2d).22–25
3.2. Electrochemical characterization
The preparation of the GO/AuNDs/PDA-modified electrode on GCE, with and without bacteria, was assessed using CV, DPV, and EIS techniques, as shown in Fig. 3. The CV and DPV (in both oxidation and reduction directions) (Fig. 3a and b) revealed prominent redox peaks for [Fe(CN)6]3−/4− on bare GCE, GO/GCE, GO/AuNDs/GCE, and GO/AuNDs/PDA/GCE in the presence and absence of bacteria. In both CV and DPV, the modifications with GO and GO/AuNDs enhanced peak intensities, attributed to increased active surface area and improved conductivity. However, after polymerization with PDA and bacteria, the peaks diminished significantly due to the bacteria's poor conductivity. The removal of the nonconductive bacteria templates exposed imprinted cavities, restoring the redox peaks.In the Nyquist plots of the EIS measurements (Fig. 3c), the modification with GO and AuNDs significantly reduced the charge transfer resistance (Rct) from 6.9 ± 0.2 kΩ for the bare electrode to 2.7 ± 0.05 kΩ and 2.0 ± 0.09 kΩ, respectively. This reduction confirmed the successful deposition of conductive GO and AuNDs layers on the electrode surface. During NIP formation, the Rct value increased to 108.2 ± 3.0 kΩ after DA polymerization, attributed to its poor conductivity. The Rct further increased to 247.0 ± 7.1 kΩ after incorporating the nonconductive bacteria into the PDA film. Upon removing the bacteria and exposing the imprinted cavities on the polymer surface, the Rct decreased to 35.5 ± 1.7 kΩ, confirming the successful removal of bacteria and exposure of the functionalized sites.
Additionally, in this study, E. coli AR3110 was transformed with a plasmid harboring the gene for the fluorescent protein mCherry under a strong, constitutively expressed promoter.33 Fluorescent E. coli was used as the MIP template, enabling the visualization of the removal process through fluorescence imaging. Fig. S3a displays the visible light image of the 0.01 equiv. M HNO3/SDS washing solution following the removal of the bacterial template, where the removed bacteria were clearly observed. Fig. S3b shows the corresponding fluorescence image of the same washing solution, confirming the fluorescence of the removed bacteria. A sample of liquid culture containing mCherry E. coli suspended in PBS was also imaged (Fig. S3c and d) as a control reference without washing.
3.3. Optimization of MIP and calibration study
The efficient removal of templates is a pivotal step in the development of MIP-based biosensors because residual analyte blocks imprinted cavities, restricts access to the redox probe, destabilizes the electrochemical baseline, and degrades reattachment capabilities.34 To optimize this process, two washing solutions, CH3COOH/SDS and HNO3/SDS, were evaluated and compared. CH3COOH/SDS was a common, milder eluent in MIP workflows to remove bacterial cells from PDA films.26,35 As illustrated in Fig. 4a, washing the electrode with a 5% (w/w) CH3COOH/SDS solution for 18 h reduced the Rct value to 104.1 ± 6.1 kΩ. However, using a 0.01 equiv. M HNO3/SDS solution resulted in a further reduction of the Rct value to 10.6 ± 0.9 kΩ. The significant decrease in resistance observed with the HNO3/SDS solution indicated its superior effectiveness in removing bacteria compared to CH3COOH/SDS. PDA films were relatively stable in low-concentration acidic solutions, and covalently cross-linked PDA remains firmly anchored to the substrate after acid treatment.36 As shown in Fig. S4, the PDA films before and after washing with HNO3/SDS had similar morphologies and height distributions, confirming that the PDA films experienced minimal damage during the wash.To further enhance the removal process, various washing methods were studied using the HNO3/SDS solution. As shown in Fig. 4b, sonication provided a faster and more effective approach for bacterial removal. After the electrode was washed under magnetic stirring for 2 h, the Rct value decreased to 32.9 ± 1.2 kΩ. In contrast, sonication in the washing solution for 10, 20, and 30 min resulted in significantly lower Rct values of 18.1 ± 0.1 kΩ, 7.2 ± 0.3 kΩ, and 7.3 ± 0.2 kΩ, respectively. Since no notable difference was observed between the Rct values after 20 and 30 min of sonication, 20 min of sonication in 0.01 equiv. M HNO3/SDS was identified as the optimal method for effective bacterial removal.
Another critical factor in the MIP-based sensor development was precisely controlling the thickness of the polymer layer to ensure the formation of accessible and near-surface imprinted cavities, which can minimize mass-transfer limitations and shorten recognition time.37 A thick polymer with deeply buried sites slows diffusion, increases Rct, inhibits bacterial reattachment, and reduces reusability, thereby negatively impacting sensor performance.17 To determine the optimal polymer thickness, electropolymerization of DA was performed in the presence and absence of bacteria with 10, 15, and 20 cycles of CVs. As shown in Fig. S5a, the measured current response decreased with increasing polymer thickness, highlighting the critical balance required for efficient MIP fabrication.
The responses of electrodes with varying thicknesses were tested for 15 and 20 cycles of electropolymerization and compared with those for 10 cycles by incubating them in different concentrations of E. coli. The results, shown in Fig. S5b–e, indicated that for electrodes with 15 and 20 cycles, the Rct value remained constant beyond 1.0 × 102 CFU mL−1. In contrast, the MIP with 10 cycles of polymerization demonstrated a linear detection range up to 1.0 × 104 CFU mL−1, which was twice that of the MIPs with 15 and 20 cycles of polymerization (Fig. 4c and d). After incubation with 1.0 × 104 CFU mL−1, all cavities were filled, saturating the electrode surface with bacteria. Consequently, further incubation did not result in a significant change to the Rct value. The plot of ΔRct demonstrated a linear relationship within the range of 1.0 × 101 to 1.0 × 104 CFU mL−1. The Rct and χ2 values obtained from the EIS measurements are listed in Table S1. The χ2 ranged from 0.13 to 0.04, indicating that the proposed circuit fits the EIS measurements well. The limit of detection (LOD) was calculated using eqn (1) and determined to be 1.5 CFU mL−1.
 | (1) |
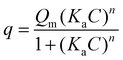 | (2) |
3.4. Selectivity of MIP-based sensor
To assess the sensor's selectivity and target recognition, the electrode was incubated with various bacterial strains at 1.0 × 103 CFU mL−1 for 30 min, including E. coli DH5α, P. defensor, B. velezensis, and E. coli AR3110, which served as the template bacteria. As illustrated in Fig. 4e and f, the sensor exhibited the most significant increase in Rct for the E. coli AR3110 strain, and the selective rebinding of the targets inhibited electron transfer between the electrode and the [Fe(CN)6]3−/4− redox couple, resulting in a marked increase in the Rct. In contrast, P. defensor and B. velezensis induced only minor Rct changes, attributable primarily to nonspecific adsorption. The slightly higher interference from E. coli DH5α relative to the non-E. coli species likely reflected similarities in cell size and Gram-negative envelope architecture. Compared with E. coli DH5α, curli-producing E. coli AR3110 exhibited stronger attachment and higher affinity to PDA-based MIPs. Curli are adhesive functional amyloid fibers produced by members of the Enterobacteriaceae that mediate adhesion to abiotic surfaces, promote cell–cell aggregation, and drive biofilm formation.39 To further enhance selectivity, dual-recognition systems based on MIPs have been developed. Recently, Agar et al.26 integrated aptamers with MIPs for the highly selective detection of S. aureus and E. coli, reporting that the combined aptamer and MIP architecture outperformed either component alone.3.5. Real sample study
To evaluate the sensor's applicability to environmental monitoring, its performance was tested on various water samples, including creek water and wastewater. The EIS and the standard addition method were employed to assess the sensor's ability to detect bacteria in these complex matrices. Prior to measurement, all samples were diluted fivefold with PBS (pH 7.4), which minimizes interference from ionic strength and pH on sensor performance. Diluting environmental samples with the standard PBS buffer prior to analysis is an effective step that normalizes pH and ionic strength, and reduces the concentration of potential organic matter to prevent electrode fouling.40 The standard addition method was used to mitigate matrix effects inherent to real samples. Initial bacterial concentrations were determined, followed by spiking samples with known amounts of bacteria. The percent recoveries were then calculated to verify the sensor's accuracy. As summarized in Table 1, the percent recoveries ranged from 96 to 103%, highlighting the sensor's effectiveness and reliability in detecting bacteria in complex real-world samples. To further minimize interference and non-specific absorption, we propose integrating membrane filtration as a sample-preparation step to separate bacteria from water samples. By adjusting pore size and surface properties, membrane filtration can efficiently capture and release bacteria for sample preparation in electrochemical detection.41 Using a membrane with a controlled pore size, typically in the microfiltration range (0.2–0.45 µm), bacterial cells can be separated from small, non-target molecules that contribute to non-specific adsorption, making this approach compatible with on-field applications.42| Real sample | Detected concentration (CFU mL−1 ± s.d.) | Spiked concentration (CFU mL−1) | Found concentration (CFU mL−1 ± s.d.) | Recovery (% ± s.d.) |
|---|---|---|---|---|
| Creek water | 13.5 ± 3.0 | 0 | — | — |
| 13.5 ± 3.0 | 50.0 | 61.5 ± 2.4 | 96.0 ± 4.7 | |
| Wastewater | 105.9 ± 1.0 | 0 | — | — |
| 105.9 ± 1.0 | 1000.0 | 1138.2 ± 16.9 | 103.0 ± 1.7 |
3.6. From GCE to SPE: a step toward portability
To adapt to field environmental applications, SPEs were popularly applied to develop portable electrochemical sensors for on-site monitoring of microbial contamination in water, offering several advantages, including low cost, disposability, and mass production.43 To further support practical applications, SPEs were modified with GO, AuNDs, and PDA using the same optimized parameters as for the GCE modification, and a comparative study was conducted between GCE- and SPE-based sensors.The sensitivity of the SPE-based sensors was evaluated by incubating them with varying concentrations of E. coli. As shown in Fig. S7a, the Rct value increased with rising bacterial concentrations. Fig. S7b illustrates that the change in Rct exhibited a linear response over the range of 1.0 × 101 to 1.0 × 103 CFU mL−1, beyond which no significant change was observed due to electrode surface saturation. The LOD calculated from the calibration curve was 1.3 CFU mL−1. Compared to the GCE-based sensors, the SPE-based ones demonstrated a narrower linear range (1.0 × 101–1.0 × 103 CFU mL−1vs. 1.0 × 101–1.0 × 104 CFU mL−1) but achieved a slightly lower LOD (1.3 CFU mL−1vs. 1.5 CFU mL−1).
Compared to uniform and smooth GCEs, the surface of SPEs contains carbon ink composed of graphite particles, polymer binders, and additives, and the polymer binders may influence redox activity and decrease electron transfer reactivity.44 Additionally, the fabrication process and carbon ink composition can significantly impact the electrochemical properties of SPE, including electron transfer kinetics, background current, and impedance.45 The narrower linear range of SPE compared to GCE can be attributed to its lower electron transfer efficiency and potential differences between the electrode materials. These factors limit the number of binding sites and compromise the sensor's linearity at higher bacterial concentrations. The chemically uniform, binder-free surface of GCE supports more uniform MIP growth, cleaner template removal, and lower capacitive noise. Although SPEs are affordable and compatible with portable devices, further enhancements, such as pretreatment, are necessary to extend the linear range and enhance sensor performance.
As summarized in Table 2, the MIP-based sensor developed in this study exhibited a comparable linear range and a lower LOD than those reported in the recent literature for E. coli detection using electrochemical techniques. Moreover, the MIP-based sensor demonstrated long-term stability and was suitable for integration with portable analytical instruments for practical applications. The GCEs were stored at room temperature for 45 days without a significant loss of performance, as shown in Fig. S8. This supports the feasibility of the sensor for on-site water quality monitoring, where an extended shelf life is essential. Although antibody-based electrochemical sensors with exceptional affinity and excellent selectivity are well-established,46 the stability of biomaterials depends on a delicate balance of hydrogen bonds, hydrophobic interactions, and ionic bonds, which is easily disrupted by changes in pH, temperature, and the organic matter in real samples.47 In contrast, the fully polymeric MIP represents a strategically advantageous, cost-effective, and durable alternative for the expensive and unstable biomolecules.
| Electrode | Electrochemical technique | Linear range (CFU mL−1) | LOD (CFU mL−1) | Ref. |
|---|---|---|---|---|
| SPE modified with magnetic and gold nanoparticles conjugated with antibodies | DPV | 1.0 × 101–1.0 × 106 | 10.0 | 48 |
| SPE modified with aptamer-NanoZyme based assay | CV | 1.0 × 101–1.0 × 109 | 10.0 | 49 |
| Gold SPE modified with antibodies | EIS | 1.0 × 101–1.0 × 108 | 30.0 | 50 |
| GCE modified with gold nanoparticles | DPV | 1.0 × 101–1.0 × 104 | 7.0 | 51 |
| GCE/GO/AuNDs/PDA | EIS | 1.0 × 101–1.0 × 104 | 1.5 | This work |
| SPE/GO/AuNDs/PDA | EIS | 1.0 × 101–1.0 × 103 | 1.3 |
4. Conclusions
In this study, a GO/AuNDs/PDA-modified electrode was developed as a selective and sensitive sensor for rapid E. coli detection. SEM analysis and AFM imaging confirmed the step-wise modifications applied to the electrode surface. The incorporation of GO/AuNDs significantly enhanced the sensor's sensitivity by promoting efficient electron transfer at the electrode interface. Compared to conventional detection methods, the fabricated electrochemical MIP-based sensor offered faster response times, higher sensitivity, and lower production costs. Under optimized conditions, the sensor exhibited a linear detection range from 1.0 × 101 to 1.0 × 104 CFU mL−1 with a LOD of 1.5 CFU mL−1 for E. coli AR3110. Moreover, the MIP-based sensor demonstrated excellent selectivity, effectively distinguishing the target E. coli AR3110 from non-target bacterial species, such as E. coli DH5α, P. defensor, and B. velezensis. Its practical applicability was further validated through the successful detection of E. coli in real water samples, confirming its potential for real-world bacterial monitoring.Author contributions
Yilei Xue: methodology, formal analysis, investigation, visualization, software, and writing – original draft; Jessica Chen: methodology, formal analysis, investigation, visualization, software, writing – original draft; Meissam Noroozifar: methodology, formal analysis, investigation, visualization, software, and writing – original draft; Ruby M. A. Sullan: conceptualization, supervision, funding acquisition, and writing – review & editing; Kagan Kerman: conceptualization, supervision, funding acquisition, and writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting the manuscript have been included as part of the supplementary information (SI). The SI file includes the SEM images and XPS data of various surface modifications of the electrode as well as the images of the liquid culture of mCherry E. coli taken by the fluorescence and visible light microscopes. AFM images showed the effects of electrode surface washes using the acid/surfactant mixture. The electropolymerization data were presented with Nyquist plots of EIS data and cyclic voltammograms. Langmuir–Freundlich isotherm fit line was employed to calculate the surface binding characteristics of the bacteria. The Nyquist plots were also shown to support the calibration curve data. See DOI: https://doi.org/10.1039/d5an01217a.Acknowledgements
The authors acknowledge financial support from the Canada Research Chair Tier-2 award for “Bioelectrochemistry of Proteins” (project no. 950-231116), Ontario Ministry of Research and Innovation (project no. 35272), Discovery Grant (project no. RGPIN-2020-07164) from Natural Sciences and Engineering Research Council of Canada (NSERC), and Canada Foundation for Innovation (project no. 35272).References
- H. Leclerc, L. Schwartzbrod and E. Dei-Cas, Crit. Rev. Microbiol., 2002, 28, 371–409 CrossRef CAS PubMed
.
- A. H. Havelaar, M. D. Kirk, P. R. Torgerson, H. J. Gibb, T. Hald, R. J. Lake, N. Praet, D. C. Bellinger, N. R. de Silva, N. Gargouri, N. Speybroeck, A. Cawthorne, C. Mathers, C. Stein, F. J. Angulo and B. Devleesschauwer, PLoS Med., 2015, 12, 1001923 CrossRef PubMed
.
- L. Varadi, J. L. Luo, D. E. Hibbs, J. D. Perry, R. J. Anderson, S. Orenga and P. W. Groundwater, Chem. Soc. Rev., 2017, 46, 4818–4832 RSC
.
- R. Y. C. Kong, M. M. H. Mak and R. S. S. Wu, Ocean Coast. Manag., 2009, 52, 355–358 CrossRef
.
- V. Velusamy, K. Arshak, O. Korostynska, K. Oliwa and C. Adley, Biotechnol. Adv., 2010, 28, 232–254 CrossRef CAS PubMed
.
- A. Ahmed, J. V. Rushworth, N. A. Hirst and P. A. Millner, Clin. Microbiol. Rev., 2014, 27, 631–646 CrossRef CAS PubMed
.
- N. Bhalla, P. Jolly, N. Formisano and P. Estrela, Essays Biochem., 2016, 60, 1–8 Search PubMed
.
- H. O. Kaya, A. E. Cetin, M. Azimzadeh and S. N. Topkaya, J. Electroanal. Chem., 2021, 882, 114989 CrossRef CAS PubMed
.
- J. Monzo, I. Insua, F. Fernandez-Trillo and P. Rodriguez, Analyst, 2015, 140, 7116–7128 RSC
.
- J. Bhardwaj, S. Devarakonda, S. Kumar and J. Jang, Sens. Actuators, B, 2017, 253, 115–123 CrossRef CAS
.
- J. Chen, Z. Jiang, J. D. Ackerman, M. Yazdani, S. Hou, S. R. Nugen and V. M. Rotello, Analyst, 2015, 140, 4991–4996 RSC
.
- M. Tolba, M. U. Ahmed, C. Tlili, F. Eichenseher, M. J. Loessner and M. Zourob, Analyst, 2012, 137, 5749–5756 RSC
.
- S. Shahrokhian and S. Ranjbar, Analyst, 2018, 143, 3191–3201 RSC
.
- J. Wu, R. Wang, Y. Lu, M. Jia, J. Yan and X. Bian, Anal. Chem., 2019, 91, 1027–1033 CrossRef CAS PubMed
.
- J. J. BelBruno, Chem. Rev., 2019, 119, 94–119 CrossRef CAS PubMed
.
- L. Wang, M. Pagett and W. Zhang, Sens. Actuators Rep., 2023, 5, 100153 CrossRef
.
- R. Wang, L. Wang, J. Yan, D. Luan, S. Tao, J. Wu and X. Bian, Talanta, 2021, 226, 122135 CrossRef CAS PubMed
.
- R. Gui, H. Jin, H. Guo and Z. Wang, Biosens. Bioelectron., 2018, 100, 56–70 CrossRef CAS PubMed
.
- A. A. Lahcen and A. Amine, Electroanalysis, 2018, 31, 188–201 CrossRef
.
- V. Georgakilas, J. A. Perman, J. Tucek and R. Zboril, Chem. Rev., 2015, 115, 4744–4822 CrossRef CAS PubMed
.
- W. Liu, Y. Ma, G. Sun, S. Wang, J. Deng and H. Wei, Biosens. Bioelectron., 2017, 92, 305–312 CrossRef CAS PubMed
.
- S. Prakash, T. Chakrabarty, A. K. Singh and V. K. Shahi, Biosens. Bioelectron., 2013, 41, 43–53 CrossRef CAS PubMed
.
- H. Lee, S. M. Dellatore, W. M. Miller and P. B. Messersmith, Science, 2007, 318, 426–430 CrossRef CAS PubMed
.
- Y. Liu, K. Ai and L. Lu, Chem. Rev., 2014, 114, 5057–5115 CrossRef CAS PubMed
.
- N. Bisht, N. Dwivedi, A. Khosla, D. P. Mondal, A. K. Srivastava and C. Dhand, J. Electrochem. Soc., 2022, 169, 107505 CrossRef CAS
.
- M. Agar, M. Laabei, H. S. Leese and P. Estrela, Biosens. Bioelectron., 2025, 272, 117136 CrossRef CAS PubMed
.
- X. Liu, M. Tian, Q. Zhu, Y. Wang, H. Huo, T. Chen and Y. Xu, Anal. Chem., 2025, 97, 8915–8922 CrossRef CAS PubMed
.
- B. Stöckle, D. Y. W. Ng, C. Meier, T. Paust, F. Bischoff, T. Diemant, R. J. Behm, K. E. Gottschalk, U. Ziener and T. Weil, Macromol. Symp., 2014, 346, 73–81 CrossRef
.
- S. Li, H. Wang, M. Young, F. Xu, G. Cheng and H. Cong, Langmuir, 2019, 35, 1119–1125 CrossRef CAS PubMed
.
- F. Bernsmann, J.-C. Voegel and V. Ball, Electrochim. Acta, 2011, 56, 3914–3919 CrossRef CAS
.
- F. Zeng, Z. Sun, X. Sang, D. Diamond, K. T. Lau, X. Liu and D. S. Su, ChemSusChem, 2011, 4, 1587–1591 CrossRef CAS PubMed
.
- B. Rui, M. Yang, L. Zhang, Y. Jia, Y. Shi, R. Histed, Y. Liao, J. Xie, F. Lei and L. Fan, J. Appl. Electrochem., 2020, 50, 407–420 CrossRef CAS
.
- E. L. Lagendijk, S. Validov, G. E. Lamers, S. de Weert and G. V. Bloemberg, FEMS Microbiol. Lett., 2010, 305, 81–90 CrossRef CAS PubMed
.
- M. Valentino, A. Imbriano, A. Tricase, F. Della Pelle, D. Compagnone, E. Macchia, L. Torsi, P. Bollella and N. Ditaranto, Anal. Methods, 2023, 15, 1250–1253 RSC
.
- S. Chen, X. Chen, L. Zhang, J. Gao and Q. Ma, ACS Appl. Mater. Interfaces, 2017, 9, 5430–5436 CrossRef CAS PubMed
.
- H. Wei, J. Ren, B. Han, L. Xu, L. Han and L. Jia, Colloids Surf., B, 2013, 110, 22–28 CrossRef CAS PubMed
.
- C. Mwanza, W.-Z. Zhang, K. Mulenga and S.-N. Ding, Green Chem., 2024, 26, 11490–11517 RSC
.
- J. C. Soares, A. C. Soares, P. A. Pereira, V. da C. Rodrigues, F. M. Shimizu, M. E. Melendez, C. Scapulatempo Neto, A. L. Carvalho, F. L. Leite, S. A. Machado and O. N. Oliveira Jr., Phys. Chem. Chem. Phys., 2016, 18, 8412–8418 RSC
.
- M. M. Barnhart and M. R. Chapman, Annu. Rev. Microbiol., 2006, 60, 131–147 CrossRef CAS PubMed
.
- D. Zabitler, E. Ülker, K. Turan, N. Ö. Erdoğan and G. Aydoğdu Tığ, Top. Catal., 2025, 69, 95–125 CrossRef
.
- R. O. Toganel, C. N. Ciurea, A. Cighir, A. D. Mare, R. L. Coseriu, C. Vintila, D. C. Cucoranu and A. Man, Microbiologyopen, 2025, 14, e70119 CrossRef CAS PubMed
.
- P. Chorti, A. P. Kazi, A.-M. J. Haque, M. Wiederoder and D. C. Christodouleas, Sens. Actuators, B, 2022, 351, 130965 CrossRef CAS
.
- M. Li, Y. T. Li, D. W. Li and Y. T. Long, Anal. Chim. Acta, 2012, 734, 31–44 CrossRef CAS PubMed
.
- J. Wang, B. Tian, V. B. Nascimento and L. Angnes, Electrochim. Acta, 1998, 43, 3459–3465 CrossRef CAS
.
- G. Cui, J. H. Yoo, J. S. Lee, J. Yoo, J. H. Uhm, G. S. Cha and H. Nam, Analyst, 2001, 126, 1399–1403 RSC
.
- N. Alizadeh and A. Salimi, Electroanalysis, 2018, 30, 2803–2840 CrossRef CAS
.
- C. Solier and H. Langen, Proteomics, 2014, 14, 774–783 CrossRef CAS PubMed
.
- Y. Wang and E. C. Alocilja, J. Biol. Eng., 2015, 9, 16 CrossRef PubMed
.
- R. Das, B. Chaterjee, A. Kapil and T. K. Sharma, Sens. Bio-Sens. Res., 2020, 27, 100313 CrossRef
.
- M. Cimafonte, A. Fulgione, R. Gaglione, M. Papaianni, R. Capparelli, A. Arciello, S. Bolletti Censi, G. Borriello, R. Velotta and B. Della Ventura, Sensors, 2020, 20, 274 CrossRef CAS PubMed
.
- Y. Li, H. Liu, H. Huang, J. Deng, L. Fang, J. Luo, S. Zhang, J. Huang, W. Liang and J. Zheng, Biosens. Bioelectron., 2020, 147, 111752 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |




