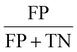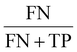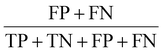Exploring the feasibility of near-infrared spectroscopy and machine learning for detecting cardiovascular diseases and diabetes mellitus in fingernails
Megan
Wilson
 *a,
Dhiya
Al-Jumeily
b,
Jason
Birkett
*a,
Dhiya
Al-Jumeily
b,
Jason
Birkett
 a,
Iftikhar
Khan
a,
Ismail
Abbas
c,
Matthew
Harper
d and
Sulaf
Assi
a,
Iftikhar
Khan
a,
Ismail
Abbas
c,
Matthew
Harper
d and
Sulaf
Assi
 a
a
aSchool of Pharmacy and Biomedical Sciences, Liverpool John Moores University, 3 Byrom Street, Liverpool, L3 3AF, UK. E-mail: m.wilson3@2019.ljmu.ac.uk
bSchool of Computer Science and Mathematics, Liverpool John Moores University, Liverpool, UK
cFaculty of Science, Lebanese University, Beirut, Lebanon
dDepartment of Archaeology, Classics and Egyptology, University of Liverpool, Liverpool, UK
First published on 16th February 2026
Abstract
Cardiovascular diseases (CVDs) and diabetes mellitus (DM) are significant conditions that impact lives around the globe. Frequently employed methods for detecting CVDs and/or DM such as blood work and cardiac catheterisation are often invasive, intrusive and can cause the patient additional physical and psychological harm. Vibrational spectroscopic methods including near-infrared (NIR) spectroscopy have emerged as novel methods for detecting medical conditions and diseases including amyotrophic lateral sclerosis, cancer, DM and periodontitis. NIR spectroscopy's ability to perform rapid and cost-effective analysis saves diagnostic waiting times, providing relief for strained healthcare systems. Moreover, their non-invasive, non-intrusive and non-destructive nature allow application to alternative biological matrices such as hair, fingernails and saliva. Therefore, this work explored the feasibility of NIR spectroscopy paired with machine learning (ML) for detecting CVDs and/or DM in fingernails. NIR spectroscopy successful characterised disease-related spectral features including key NIR regions related to the presence of advanced glycated end-products (AGEs), glycated proteins and DM. To further assess the detective capabilities of NIR spectroscopy, classification models were trained. Cubic and quadratic support vector machine (SVM) models demonstrated accuracy in terms of the classification of healthy, CVD and diabetic fingernails. Accuracy was further improved through binary classification models, which allowed the independent classification of CVD and DM spectra against healthy spectra. In summary, NIR spectroscopy combined with ML provided accurate detection for CVDs and DM in fingernails.
Introduction
Cardiovascular diseases (CVDs) refer to the chronic conditions of the heart or blood vessels such as coronary heart disease (CHD), cerebrovascular disease and rheumatic heart disease.1 Across the globe, CVDs cause a significant number of deaths (19.8 million) annually.1 Several factors such as alcohol/tobacco use, physical inactivity and unhealthy diets have been associated with the prevalence of CVDs; research showed the close link between CVDs and diabetes mellitus (DM).2 DM is a chronic metabolic disease that is characterised by elevated blood glucose levels that often causes severe damage to the blood vessels, eyes, heart, kidney and nerves.3 According to the International Diabetes Federation, 590 million people worldwide have DM.4 This number is said to increase to 853 million in 2050.4 In diabetic cases, CVDs are the most prevalent cause of morbidity and mortality, with several overlapping risk factors such as obesity and dyslipidaemia.2 As a result, diabetic patients are likely to suffer a cardiac event.The medical field has previously utilised invasive and intrusive diagnostic techniques for both CVDs and DM. For example, blood work is now considered the gold standard for the diagnosis of DM.5 This diagnostic test often utilises haemoglobin A1c (HbA1c) as an indicator of blood glucose levels over two to three months.6 Despite, its frequent employment for initial diagnosis and diagnostic monitoring/management, blood work with HbA1c is invasive and intrusive. Similarly, the diagnosis of CVDs has often been carried out through several invasive techniques including cardiac catheterisation and coronary angiography.7 Not only are the highlighted techniques invasive and intrusive, which can cause patients further physical and psychological harm, but require specialised equipment, training and personnel. Moreover, non-intrusive techniques such as computed topography (CT) scans and electrocardiograms (ECGs) require high-levels of user knowledge for application and interpretation, which is not often feasible in low-resource settings. This urges for alternative disease detection techniques, which allow for non-invasive, non-intrusive and rapid analysis, without the expense of detection accuracy.
Within the scientific community, there has been a movement towards the use of vibrational spectroscopic techniques for detecting diseases. Not only does vibrational spectroscopy offer rapid and non-destructive analysis but can be applied non-invasively and non-intrusively to several alternative matrices including fingernails, hair and saliva.8–11 Nogueira et al., demonstrated the successful combination of Fourier transform infrared (FTIR) spectroscopy and saliva for the detection of DM and periodontitis.11 This work utilised infrared (IR) spectra and a weighted K-nearest neighbour (KNN) model for the prediction of healthy controls, diabetic cases and diabetic periodontitis cases. This model was trained using 23 patients and obtained an area under the Receiver Operating Characteristic (ROC) curve of 0.92 and 0.95 when considering the diabetic or diabetic periodontitis groups as positive groups.12
Similarly, Carlomagno et al., showed the use of Raman spectroscopy and saliva for the diagnosis of amyotrophic lateral sclerosis (ALS).12 This work highlighted the correlation between Raman data and paraclinical scores, making evident multifactorial biochemical modifications of ALS’ pathology. Moreover, this research applied principal component analysis (PCA) for the classification of ALS, Alzheimer's disease (AD), Parkinson's disease (PD) cases and healthy controls. PCA demonstrated a partial overlap between healthy controls and AD cases. ALS scores were well-separated from control scores and indicated the feasibility of Raman spectroscopy for the detection of ALS.12
Despite the increased use of IR and Raman spectroscopy in detecting disease, there has been little use for near-infrared (NIR) spectroscopy for disease detection. Thus, this study aims to complement the literature by detecting the presence of CVDs and/or DM in fingernails using NIR spectroscopy. Therefore, taking a precision medicine approach for cost-effective and rapid disease detection, which will reduce patients’ time in the waiting room.
Methodology
Sample collection
Prior to the collection of fingernail clippings, ethical approval was granted by two institutions, Liverpool John Moores University (LJMU) (23/PBS/009A) and the Lebanese University (LU) (2022-0104). Participants were recruited through the LJMU's community site, where they received a participant information sheet, questionnaire and consent form. The administration of a questionnaire allowed the collection of the participant's characteristics including demographic (age, biological sex and ethnicity), medicinal (previous diagnosis and medicine use) and lifestyle (diet, exercise and smoking habits) factors. Those that took part in the research, provided a set of 6–10 fingernail clippings and a completed questionnaire. Fingernail clippings were stored and analysed in clear 2 mL glass vials (VWR) that allowed NIR radiation to pass through.Fingernails were sourced from female (n = 50) and male (n = 35) participants, who were aged 18–85 years old. Across the 85 participants, five ethnic groups were accounted for, those being: Arab (n = 33, 39%), Asian (n = 10, 12%), Indian (n = 2, 2%), Lebanese Arab (n = 15, 18%) and White (n = 25, 29%). Participants were classified into five groups including healthy (n = 48, 58%), unhealthy (n = 13, 16%), CVD (n = 11, 13%), diabetic (n = 3, 4%) and CVD-diabetic (n = 7, 9%). It is important to note that all participants received their medical diagnosis/diagnoses prior to participating within this study by a trained healthcare professional. The purpose of this study was to not diagnose but detect the presence of CVDs and/or DM in fingernails. Individuals categorised into the CVD group had been diagnosed with atrial fibrillation (AF) (n = 1, 9%), heart disease (n = 1, 9%), and hypertension (n = 9, 82%). Diabetic participants consisted of both type 1 diabetes mellitus (T1DM) (n = 1, 33.3%) and type 2 diabetes mellitus (T2DM) (n = 2, 66.6%). Individuals with a diagnosis of both a CVD and DM were placed into CVD-diabetic group. Participants classified as unhealthy had previously received a medical diagnosis from a healthcare professional that was unrelated to CVDs or DM. The anonymised dataset can be accessed upon request from the eSystem Engineering Society DataBank (https://dese.ai/medicaldatabank/).
Instrumentation
NIR spectra were collected using the PerkinElmer Spectrum Two N FT-NIR Spectrometer (PerkinElmer, Waltham, MA). The spectrometer was equipped with a temperature stabilised InGaAs detector. To hold the glass vial of fingernail clippings, a vial holder and NIR reflectance module (NIRM) sampling accessories were utilised. Spectra were collected over the spectral range of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–4000 cm−1, with a spectral resolution of 8 cm−1. Between spectral collection, the sampling accessory was removed and the instrument covered with a black blackout cloth (polyester-based), which acted as a dark correction and was obtained from the instrument manufacturer. Moreover, the black cloth helped prevent light leakage and minimised interference from external sources for accurate measurements. The black cloth was then removed and a background taken (Fig. 1).
000–4000 cm−1, with a spectral resolution of 8 cm−1. Between spectral collection, the sampling accessory was removed and the instrument covered with a black blackout cloth (polyester-based), which acted as a dark correction and was obtained from the instrument manufacturer. Moreover, the black cloth helped prevent light leakage and minimised interference from external sources for accurate measurements. The black cloth was then removed and a background taken (Fig. 1).
Fingernail measurements
Fingernails were sampled and measured in the form of an overhang intact clipping. Fingernail clippings possessed an overhang width (growth) of 0.3017–5.82 mm (median = 1.67 mm, interquartile range (IQR) = 1.34–2.034 mm), depth (thickness) of 0.138–1.14 mm (median = 0.37 mm, IQR = 0.313–0.45 mm) and length of 5.41–14.4 mm (median = 9.88 mm, IQR = 8.87–11.1 mm). The fingernail clippings filled approximately 0.5 mL of the 2 mL glass vial. Using diffuse reflectance, NIR light penetrates the fingernails gathering information from both the surface and interior of the fingernail clippings. For each ensemble of fingernails, a total of 22 spectra were taken. The vial holder accessory ensured that the fingernails were placed directly over the NIR source. To ensure that the distribution of endogenous compounds was accounted for across the fingernails, the glass vial was vortexed using the SciQuip Vortex VariMix (Rotherham, UK) between the collection of spectra. Spectral acquisition took 40 seconds per spectrum. All spectra were exported for online spectral interpretation and spectral quality assessments, as well as the application of classification models. The number (n) of bands indicated the presence of key endogenous compounds, alongside the NIR activity of fingernails. Other parameters for spectral quality included maximum band position/intensity and signal-to-noise (S/N) ratio.Data treatment
Normalisation ensures data consistency by aligning and clearing noise enhancing spectral analysis and was carried out throughout the PerkinElmer Spectrum Two N FT-NIR's software, Spectrum IR. Applies baseline correction and scaling to ensure consistent and accurate spectral analysis. Further pre-processing was carried out through the SpectraGryph. Multiplicative scatter correlation (MSC) was applied to NIR spectra for the correction of offset and baseline. This treatment also ensured that light-scattering effects were corrected by aligning the spectra to a reference.13 As NIR spectra are often characterised by broad and overlapping bands, derivatisation was imperative Therefore, NIR spectra were also treated using the first derivative (D1), which emphasises subtle spectral features where the data slope changes rapidly.Data analysis
NIR spectra were imported into MATLAB 2019a for spectral visualisation. ML classification models were then explored via the MATLAB 2019a Classification Learner Toolbox. Initially, 32 models were trained with 1403 spectra and the full range of features (n = 3000). It is worth mentioning that the 3000 datapoints were obtained at a 2 cm−1 increment over the range of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–4000 cm−1. While feature selection was initially explored, the accuracy of classifications models vastly decreased. A 10-fold cross-validation method was applied to validate the models. Moreover, 20% of the imported data was employed as a test set, which blindly assessed the models’ predictive abilities. The highest performing models included: bagged tree ensemble, boosted tree ensemble, cubic support vector (SVM), SVM kernel and quadratic SVM.
000–4000 cm−1. While feature selection was initially explored, the accuracy of classifications models vastly decreased. A 10-fold cross-validation method was applied to validate the models. Moreover, 20% of the imported data was employed as a test set, which blindly assessed the models’ predictive abilities. The highest performing models included: bagged tree ensemble, boosted tree ensemble, cubic support vector (SVM), SVM kernel and quadratic SVM.
Hyperparameters were then applied to the aforementioned classification models. In the case of bagged tree ensemble, a maximum number of splits was set to 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 956 along with 30 learner nodes.14 The boosted tree model utilised an ensemble method of AdaBoost and a decision tree learner type. A maximum of number of splits was set at 20, with 30 learners and a 0.1 learning rate.14 Hyperparameters for the cubic SVM model included a cubic kernel function, a box level of one and multiclass method of one-versus-one. The quadratic SVM model employed similar hyperparameters but utilised a quadratic kernel function.
956 along with 30 learner nodes.14 The boosted tree model utilised an ensemble method of AdaBoost and a decision tree learner type. A maximum of number of splits was set at 20, with 30 learners and a 0.1 learning rate.14 Hyperparameters for the cubic SVM model included a cubic kernel function, a box level of one and multiclass method of one-versus-one. The quadratic SVM model employed similar hyperparameters but utilised a quadratic kernel function.
Models were tested via a training: test split of 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20, with 80% of the data applied for training and 20% for testing.15 The testing data, assessed the ability of the utilised models to handle unknown data and therefore, demonstrated their predictive abilities for future diagnostic purposes. A K-left method was utilised to test the classification models and reduced their subjectivity. For this experiment, a 10-fold cross-validation was employed, with data being randomly divided ten part and tested using 10% of the overall data.16
20, with 80% of the data applied for training and 20% for testing.15 The testing data, assessed the ability of the utilised models to handle unknown data and therefore, demonstrated their predictive abilities for future diagnostic purposes. A K-left method was utilised to test the classification models and reduced their subjectivity. For this experiment, a 10-fold cross-validation was employed, with data being randomly divided ten part and tested using 10% of the overall data.16
To determine the predictive capabilities of models, confusion matrices were visualised and evaluated using several evaluation metrices, including accuracy, area under the curve (AUC), false negative rate (FNR), false positive rate (FPR), misclassificatio5n rate, recall, specificity, precision, prevalence and F1-score (Table 1).
The AUC was determined via the visualisation of receiver operating characteristics (ROCs). To plot the ROC, the FPR was plotted against the true positive rate (TPR). ROC curves that were in the upper diagonal were represented by an AUC value of >0.5, while ROC curves present within the lower diagonal was indicated by an AUC value of <0.5.17
Results and discussion
Spectral analysis of fingernails
To determine the feasibility of NIR spectroscopy as a technique for the detection of CVDs and/or DM, raw NIR spectra were subject to spectral interpretation (Table 2). The main absorption bands identified within the fingernail spectra were associated with key endogenous compounds including amino acids, lipids and proteins.| Wavenumber (cm−1) | Band assignment | Region | Overtone |
|---|---|---|---|
| 4186 | CH2, CHCl3 | III | First |
| 4248 | CH2, CHCl3 | III | First |
| 4330 | CH2, CH3, CHCl3 | III | First |
| 4864 | RNHR’, RCONHR’, ROH | III | First |
| 5148 | RNH2, RNHR’ RCONHR’ | III | First |
| 5784 | SH | II | Second |
| 5890 | CH3, SH | II | Second |
| 6106 | RCONHR’ | II | Second |
| 6362 | RCONHR’ | II | Second |
| 6632 | RNH2, RCONH2, ROH | II | Second |
| 7068 | CH2 | II | Second |
| 8410 | CH, CH3 | II | Second |
Across the five groups of fingernails, the number of absorption bands varied between 7–17 bands per spectra. Healthy spectra produced 8–16 bands (median = 11, inter quartile range (IQR) = 9–11), unhealthy spectra demonstrated 8–17 (median = 10, IQR = 8.75–13), CVD spectra displayed 6–16 (median = 11.5, IQR = 9.25–15.3), diabetic spectra showed 7–12 (median = 11, IQR = 10–12) and CVD-diabetic spectra 9–14 (median = 12, IQR = 11–13). Overall, the diabetic spectra produced the lowest number of bands (n = 7) and indicated the gradual alteration of intrinsic material properties and tissue damage caused by DM.17 As a result, the NIR activity and presence of endogenous compounds was reduced in diabetic fingernails in comparison to the healthy or remaining diseased fingernails.
Spectral visualisation revealed that the shape and trend of NIR spectra between the five groups were similar, however varied in terms of absorbance intensity (Fig. 2). Variation of absorbances were seen over two main regions, those being 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–5000 and 5000–4000 cm−1. Across the first region, CVD-diabetic fingernails showed the highest intensities of absorbance followed by heathy > unhealthy > CVD > DM. At the second, region CVD-diabetic fingernails yielded the highest overall absorbance values but was shadowed by healthy > CVD > unhealthy > DM. Overall, diabetic fingernails displayed the lowest absorbance values of all NIR spectra and this was attributed to the relationship between DM and circulation.20 Through chronic glucose exposure, small and large blood cells become damaged and the deposition of key endogenous compounds within the fingernails was impaired.21 As a result, NIR light interacted with fewer molecules and lower absorbance levels are produced.
000–5000 and 5000–4000 cm−1. Across the first region, CVD-diabetic fingernails showed the highest intensities of absorbance followed by heathy > unhealthy > CVD > DM. At the second, region CVD-diabetic fingernails yielded the highest overall absorbance values but was shadowed by healthy > CVD > unhealthy > DM. Overall, diabetic fingernails displayed the lowest absorbance values of all NIR spectra and this was attributed to the relationship between DM and circulation.20 Through chronic glucose exposure, small and large blood cells become damaged and the deposition of key endogenous compounds within the fingernails was impaired.21 As a result, NIR light interacted with fewer molecules and lower absorbance levels are produced.
 | ||
| Fig. 2 Average NIR spectra of healthy (green), unhealthy (orange), CVD (red), diabetic (blue) and CVD-diabetic (black) fingernails measured using the PerkinElmer Spectrum Two N FT-NIR spectrometer. | ||
Two water bands were identified across the five groups of fingernails and were located at 5148 and 7068 cm−1.19 At band 5148 cm−1, CVD-diabetic fingernails possessed the highest absorbance, with a peak intensity of 1.26 absorbance units. Healthy, unhealthy, CVD and diabetic fingernails demonstrated slightly lower absorbance with peak intensities of 1.14, 1.13, 1.092 and 1.088 absorbance units, respectively. In contrary, healthy fingernails demonstrated the highest absorbance at water band 7068 cm−1 and displayed a peak intensity of 0.837 absorbance units. This was then followed by unhealthy, CVD-diabetic, CVD and diabetic fingernails, with peak intensities of 0.827, 0.817, 0.812 and 0.785 absorbance units, respectively. The variation of water content of fingernails can be attributed to the fingernail plate's hydration level, which plays a significant role in the fingernail's permeability.21 As a result, endogenous constituents are more likely to deposit into hydrated fingernails than dehydrated fingernails. Hence, providing explainability for the lower absorbance of CVD and diabetic fingernail sets, which are both often characterised by dry, dehydrated brittle fingernails.
A close inspection of diabetic fingernails demonstrated the ability of NIR spectroscopy to detect protein glycation in fingernails.22 Diabetic patients often suffer from hyperglycaemia, which can cause non-enzymatic glycation of free amino protein groups.23 Across the full spectral range, three areas of interest were detected and related to the presence of glycation proteins and AGEs (Fig. 3a). The first region was located between 4400–4250 cm−1 and was attributed to combination bands of C–H stretching, C–H bending and O–H bending of glucose.22 An examination of this region demonstrated differences between healthy versus diabetic fingernails (Fig. 3b). Specifically, the diabetic spectrum showed a broader band within this region in comparison to the non-diabetic healthy spectrum. The broadness of this band indicated an increase in glycation levels and suggested the presence of disease. The second region detected was between 7100–6000 cm−1 and was associated with a combination of OH antisymmetric and symmetric stretching.22Fig. 3b demonstrated that the prominent band at 6636 cm−1 was attributed to fingernail glycation. Both the intensity and broadness of this band increased in the diabetic spectrum, while the healthy spectrum demonstrated a smaller intensity and broadness. The lower intensity of band 6636 cm−1 within the healthy spectrum lends itself to the successful metabolism of glucose. Similarly, between the region 5100–4600 cm−1, the diabetic spectrum demonstrated higher intensity than the healthy spectrum. This area lends itself to CONH2 stretching bands and a combination of NH stretching and bending.24 Furthermore, the sharp band located at 4864 cm−1 provided an indication of glycation and AGEs and in turn the presence of DM. Therefore, the aforementioned areas can be utilised for the initial detection of DM in fingernails, as well as for monitoring disease management.
Application of machine learning classification
Preliminary tests were performed using the MATLAB 2019a's Classification Toolbox to establish the suitability of classification models with NIR spectra. All models within the Classification Toolbox were trained and tested using the full NIR data (n = 1403). Hence, 32 models were tested. The models that showed the highest test accuracies included bagged tree ensemble (72.7%), boosted tree ensemble (74.4%), cubic SVM (69.8%), SVM kernel (62.0%) and quadratic SVM (69.2%). However, across the 32 tested models, accuracy was low and this was attributed to a class imbalance, whereby the majority class was favoured.24–27 Therefore, data levelling was imperative. The highest performing models were then trained with a balanced dataset, whereby each class possessed an equal number of spectra (n = 66). After testing, the bagged tree ensemble and SVM kernel models improved in terms of accuracy, while the accuracy of the boosted tree ensemble, cubic SVM and quadratic SVM reduced (Table 3). Therefore, suggesting that the bagged tree ensemble and SVM kernel models were the most successful in handling unknown data. Nevertheless, the overall accuracies of four out of five classification models improved after data levelling. The overall effectiveness of the classification models were assessed in terms of accuracy, AUC, FPR, F1-score, misclassification rate, precision, prevalence, specificity and recall (Table S1).| Model | Validation accuracy (%) | Test accuracy (%) |
|---|---|---|
| SVM: support vector machine. | ||
| Bagged lree ensemble | 79.2 | 81.8 |
| Boosted tree ensemble | 30.3 | 18.2 |
| Cubic SVM | 89.8 | 89.4 |
| SVM kernel | 78.4 | 80.3 |
| Quadratic SVM | 92.8 | 87.9 |
Cubic and quadratic SVM models demonstrated the overall highest test accuracy, with 89.4% and 87.9%, respectively. The high accuracy achieved by the aforementioned models was attributed to the successful pairing of NIR spectroscopy. For example, this successful combination has been utilised for the classification of food powders, food product quality and tobacco quality.28–30 Moreover, the pairing of NIR spectroscopy and SVM models have proven successful in the classification of cancer, endocrine diseases, neurological diseases and renal disease.30
For the cubic SVM model, 27 misclassifications were seen, 12 of which were healthy spectra misclassified as unhealthy (n = 10) and CVD (n = 2). However, it was interesting to note that no unhealthy or diabetic spectra were misclassified as healthy. The cubic SVM model showed a FNR of 14.5%, FPR of 93.3%, specificity of 85.5%, recall of 91.1%, precision of 95.3% and F1-score of 93.3%. Hence, suggesting that NIR spectrometer and the cubic SVM classification model were able to differentiate between healthy and diseased fingernails. A similar trend was seen in the quadratic SVM model, which highlighted 245 correct classifications but 19 misclassifications. Within this model, eight heathy participants were misclassified as unhealthy (n = 7) and CVD (n = 1). The quadratic SVM achieved a FNR of 6.34%, FPR of 10.2%, recall of 93.7%, specificity of 89.8% and precision of 96.9%. Therefore, an F1-score of 95.3% was observed.
As an additional proof of concept, binary classification models were explored, with each model being trained and validated via the input of two classes (Fig. 4). For example, healthy versus CVD spectra or healthy versus diabetic spectra. To ensure that a class imbalance was not reintroduced, both classes were represented by 66 spectra each. Binary classification models vastly improved the classification of healthy and diseased fingernails, with several of the validated and trained models demonstrating accuracies of >90%. For the classification of CVD fingernails decision tree (DT) models such as find tree, medium tree and coarse tree models showed accuracies of 95.3% during validation. After testing, the aforementioned accuracies increased to 92.3%. Each tree model produced an AUC value of 0.953. Misclassification of the tree models was very limited, with each model classifying three CVD spectra as healthy. Moreover, no healthy spectra were misclassified as CVD, therefore a FPR of 0% was achieved (Table 4). This demonstrated that NIR spectroscopy paired with binary classification models such as DTs were successful in the classification of healthy and CVD fingernails.
 | ||
| Fig. 4 Fine tree confusion matrices of healthy (1) and CVD (2) NIR spectra measured using the Perki n Elmer Spectrum Two N FT-NIR spectrometer. | ||
| Parameters | Fine tree | Medium tree | Coarse tree |
|---|---|---|---|
| AUC: area under the curve; FNR: false negative rate; FPR: false positive rate; SVM: support vector machine. | |||
| FNR (%) | 15.4 | 15.4 | 15.4 |
| FPR (%) | 0 | 0 | 0 |
| Misclassification rate (%) | 7.69 | 7.69 | 7.69 |
| Prevalence (%) | 50 | 50 | 50 |
| Specificity (%) | 84.6 | 84.6 | 84.6 |
| Precision (%) | 86.7 | 86.7 | 86.7 |
| Recall (%) | 100 | 100 | 100 |
| F1-score (%) | 92 | 92 | 92 |
| AUC | 0.953 | 0.953 | 0.953 |
Binary classification models also showed success for the classification of healthy versus diabetic spectra. For example, bilayered and trilayered neural network models produced high levels of accuracies, with validation and test accuracies of 97.2%/100% and 96.2%/100%, respectively. The high performance of multilayered neural network models was also supported by an AUC value of 0.992, which indicated a high level of discrimination between the healthy and diabetic spectra (Table 5).
| Parameters | Bilayered neural network | Trilayered neural network |
|---|---|---|
| AUC: area under the curve; FNR: false negative rate; FPR: false positive rate. | ||
| FPR (%) | 0 | 1.87 |
| FNR (%) | 3.77 | 3.77 |
| Misclassification rate (%) | 1.89 | 2.78 |
| Prevalence (%) | 50 | 50 |
| Specificity (%) | 100 | 98.1 |
| Precision (%) | 100 | 98.1 |
| Recall (%) | 96.2 | 96.2 |
| F1-score (%) | 98.1 | 97.1 |
| AUC | 0.985 | 0.987 |
Conclusions
In this study, NIR spectroscopy has been investigated as a means of detecting CVDs and/or DM within 85 sets of fingernails. Spectral visualisation demonstrated that diabetic fingernails showed the lowest intensity of absorbance across the five groups and this was attributed to high levels of unmetabolized glucose. As a result, glucose was displaced within the fingernails. In extreme cases (hyperglycaemia), an accumulation of AGEs and glycated proteins can cause damage to small and large arteries, limiting the deposition of endogenous compounds and causing tissue damage to the fingernail itself. Furthermore, spectral interpretation revealed three key areas of interest in relation to AGEs and glycated proteins within the fingernails that indicated the presence of DM. The broadness and intensity of bands present within regions 7100–6000, 5100–4600 and 4400 and 4250 cm−1 indicated the level of AGEs and glycated proteins deposited within fingernails. Thus, can be utilised in the future for detecting DM and monitoring of insulin adherence. During multiclass classification, SVM models paired well with NIR spectroscopy and produced the highest validation and test accuracy across the five selected models. In contrast, binary classification was best performed by DT models for the classification of CVDs and multilayered neural models scored high accuracy for the binary classification of DM. Overall, this work demonstrated the feasible application of NIR spectroscopy and ML for detecting CVDs and/or DM in fingernails.Few limitations were encountered in this study. The first was attributed to the sample size in terms of healthy and diseased participants, as well as the influence of confounding factors such as age, biological sex, ethnicity and diet. The issue of data imbalance was addressed through the equal number of spectra per class employed for classification models. The aforementioned issues were attributed to the convenience and pragmatic sampling approach utilised in this study. A challenge with this pragmatic sampling approach was related to unavailability of an independent external validation set and this impacts the availability of the results and the possibility of overfitting. Moreover, multiple resampling-based validation methods were also applied to substantially mitigate the risk of overfitting.
Author contributions
Megan Wilson: conceptualisation, data curation, formal analysis, investigation, methodology, project administration, software, validation, visualisation, writing. Dhiya Al-Jumeily: conceptualisation, methodology, supervision, validation. Jason Birkett: conceptualisation, supervision. Iftikhar Khan: conceptualisation, supervision. Ismail Abbas: data curation, investigation, resources, supervision. Matthew Harper: data curation, formal analysis, methodology, software, validation, visualisation. Sulaf Assi: conceptualisation, data curation, formal analysis, investigation, methodology, resources, software, validation.Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article, including the near-infrared spectra of healthy, cardiovascular disease and diabetes mellitus individuals are available at the databank of eSystem Engineering Society at https://dese.ai/medicaldatabank-viewdata/.Supplementary information (SI) is available. The enclosed dataset comprises near-infrared (NIR) spectra of human fingernails taken from healthy, unhealthy, cardiovascular, diabetic, cardiovascular-diabetic fingernails measured using the Perkin Elmer Spectrum Two N Fourier Transform (FT)-NIR spectrometer equipped with a NIR module (NIRM) (Perkin Elmer Spectrum Two N FT-NIR spectrometer). Cardiovascular diseases (CVDs) included atrial fibrillation, heart disease (unspecified) and hypertension. See DOI: https://doi.org/10.1039/d5an01061f.
Acknowledgements
The authors would like to acknowledge PerkinElmer for providing the PerkinElmer Spectrum Two N FT-NIR spectrometer.References
- World Health Organisation, https://www.who.int/health-topics/cardiovascular-diseases#tab=tab_1, accessed September 2025.
- B. M. Leon and T. M. Maddox, World J. Diabetes, 2015, 6, 1245–1258 CrossRef.
- World Health Organisation, https://www.who.int/health-topics/diabetes#tab=tab_1, accessed September 2025.
- International Diabetes Federation, https://idf.org/about-diabetes/diabetes-facts-figures/, accessed October 2024.
- T. Higgins, Endocrine, 2013, 43, 266–273 CrossRef CAS PubMed.
- S. I. Sherwani, H. A. Khan, A. Ekhzaimy, A. Maswood and M. K. Sakharkar, Biomarker Insights, 2016, 11, 95–104 CrossRef CAS PubMed.
- NHS, https://www.nhs.uk/conditions/coronary-angiography/, accessed September 2025.
- N. Kourkoumelis, G. Gaitanis, A. Velegraki and I. D. Bassukas, Med. Mycol., 2018, 56, 551–558 CrossRef CAS.
- J. Zhu, H. Xia, X. Xu, R. Zheng, C. Liu, J. Hong and Q. Huang, Spectrochim. Acta, Part A, 2024, 314, 1241185 CrossRef.
- C. Delru, S. D. Bruyne and M. M. Speekaert, J. Pers. Med., 2023, 13, 907 CrossRef.
- M. S. Nogueira, A. L. Barreto, M. Furukawa, E. S. Rovai, A. Bastos, G. Bertoncello and L. F. C. S. Carvalho, Photodiagn. Photodyn. Ther., 2022, 40, 103036 CrossRef CAS.
- C. Carlomagno, P. I. Banfi, A. Gualerzi, S. Picciolini, E. Volpato, M. Meloni, A. Lax, E. Colombo, N. Ticozzi, F. Verde, V. Silani and M. Bedoni, Sci. Rep., 2020, 10, 10175 CrossRef CAS PubMed.
- S. N. Thennadil, H. Martens and A. Kohler, Appl. Spectrosc., 2006, 60, 315–321 CrossRef CAS PubMed.
- M. Harper, PhD thesis, Liverpool John Moores University, 2023.
- B. Colakoglu, D. Alis and M. Yergin, J. Oncol. Pract., 2019, 2019, 1–7 Search PubMed.
- A. Jiménez-Valverde, Biodiversity Conserv., 2021, 30, 1–10 Search PubMed.
- P. Sihota, R. N. Yadav, V. Dhiman, S. K. Bhadada, V. Mehandia and N. Kumar, Sci. Rep., 2019, 9, 3193 CrossRef.
- D. Jee, unpublished work.
- A. Poznyak, A. V. Grechko, P. Poggio, V. A. Myasoedova, V. Alfieri and A. N. Orekhov, Int. J. Mol. Sci., 2020, 6, 1835 CrossRef.
- S. Baswan, G. B. Kastings, S. K. Li, R. Wickett, B. Adams, S. Eurich and R. Schamper, Mycoses, 2017, 60, 284–295 CrossRef.
- T. Monteyne, R. Coopman, A. S. Kishabongo, J. Himpe, B. Lapauw, S. Shadid, A. H. V. Aken, D. Berenson, M. M. Speechaert, T. D. Beer and J. R. Delanghe, Clin. Chem. Lab. Med., 2018, 56, 1551–1558 CrossRef CAS PubMed.
- B. Giri, S. Dey, T. Das, M. Sarkar, J. Banerjee and S. K. Dash, Biomed. Pharmacother., 2018, 107, 306–328 CrossRef CAS PubMed.
- N. V. Chawla, in Data Mining and Knowledge Discovery Handbook, ed. O. Maimon and L. Rokach, 2010, 2nd edn, ch. 8, pp. 854–867 Search PubMed.
- M. Hossin and M. N. Sulaiman, Int. J. Data Min. Knowl. Manage. Process, 2015, 5, 1–11 Search PubMed.
- A. Luque, A. Carrasco, A. Martín and A. D. L. Heras, Pattern Recognit., 2019, 91, 216–231 CrossRef.
- A. Ali, S. M. Shamsuddin and A. Ralescu, Int. J. Adv. Soft Comput. Appl., 2015, 7, 176–204 Search PubMed.
- S. Ozturk, A. Bowler, A. Rady and N. J. Watson, J. Food Eng., 2023, 341, 111339 CrossRef CAS.
- L. Xie and Y. Ying, J. Zhejiang Univ., Sci., B, 2009, 23, 6157 Search PubMed.
- D. Wang, L. Xie, S. X. Yang and F. Tian, Sensors, 2018, 18, 3222 CrossRef PubMed.
- R. Victorino, A. S. Barros, S. Guedes, D. C. Caixeta and R. Sabino-Silva, Photodiagn. Photodyn. Ther., 2023, 42, 103633 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |