Solid-phase engineering of molecularly imprinted nanoparticles (NanoMIPs): how template and solid-phase drive polymer composition and binding performance
Teresa
Bosch-Tamayo
,
Tamara M.
Iakimova
 ,
Amy Q.
Shen
,
Amy Q.
Shen
 * and
Mark V.
Sullivan
* and
Mark V.
Sullivan
 *
*
Micro/Bio/Nanofluidics Unit, Okinawa Institute of Science and Technology Graduate University, 1919-1 Tancha, Onna-son, Okinawa 904-0495, Japan. E-mail: mark.sullivan@oist.jp
First published on 31st October 2025
Abstract
Molecularly imprinted polymer nanoparticles (nanoMIPs) represent a promising class of synthetic recognition elements with growing potential as robust alternatives to antibodies in diagnostic and sensing technologies. Despite widespread use, limited attention has been given to how solid-phase synthesis parameters, particularly the nature of the solid support and template identity, affect nanoMIP composition and function. Herein, we present a systematic investigation comparing popular glass bead and magnetic nanoparticle solid-phase protocols for nanoMIP synthesis targeting protein templates bovine haemoglobin (BHb) and bovine serum albumin (BSA). Using an identical functional monomer feed and surface plasmon resonance (SPR)-based affinity assays, we demonstrate that the choice of solid-phase significantly influences particle size, yield, and binding affinity, with nanoMIPs synthesized on glass beads exhibiting up to a tenfold enhancement in binding performance compared to those produced on magnetic nanoparticles. Furthermore, 1H NMR analysis reveals substantial deviations between initial monomer feed ratios and final polymer compositions, with polymer structure being highly dependent on both the solid phase and template characteristics. These findings highlight the importance of rational nanoMIP design, by challenging assumptions of uniform polymer composition and revealing how template and solid-phase interactions shape material properties. Our work establishes a framework for engineering high-performance synthetic receptors with tuneable properties and offers key insights for the optimisation of nanoMIP-based applications and sets new benchmarks for material consistency, reproducibility, and potential commercialisation.
Introduction
Molecular recognition – the selective interaction of two or more molecules via non-covalent forces, such as hydrogen bonding, metal coordination, hydrophobic effects, and van der Waals forces, is central to many biological processes.1,2 These interactions play significant roles in various biological processes, specifically where receptor–ligand binding occurs, including antigen–antibody,3 DNA–protein,4 sugar–lectin,5 and RNA–ribosomes interactions.6 Insights into these natural molecular recognition mechanisms have become fundamental to the development of disease treatments such as antibiotics, as well as developing a range of biosensors and diagnostics.7–9Antibodies are probably the most used natural recognition materials and are particularly used in medical diagnostics, especially in biosensors, lateral flow tests, and biochemical assays for disease diagnostics and monitoring.10–12 This is due to their long-standing reputation for a high degree of recognition and specificity towards a chosen target. This means that numerous standardized protocols have already been established for the production and application of antibodies in medical diagnostics.13 For all their success, antibodies can have issues particularly around their stability and long-term usability.13 Antibodies are Y-shaped proteins, which means they are susceptible to denaturation from the effects of UV light exposure, high temperature, and high/low pH, thus significantly affecting their performance.14,15 This has resulted in the search for more stable synthetic alternatives.
Molecularly imprinted polymers (MIPs) are a class of synthetic recognition materials that are often touted as a viable alternative to antibodies and have found success in the areas of biosensing and diagnostics.16 MIPs are generally produced using a self-assembly methodology, whereby a series of functional monomers are mixed with a template (target of choice) and allowed to preassemble, thereby forming a favourable complex. The monomers are then linked together with a suitable crosslinker, typically via free radical polymerisation.17 While the MIPs synthesized through conventional methods offered an acceptable level of performance, the development of molecularly imprinted nanoparticles (nanoMIPs), additionally being more suited for solid-phase extraction and chromatography,18 have drastically increased the success and versatility of these synthetic recognition materials.19,20 While there are multiple ways to produce MIP nanoparticles, e.g., via core–shell,21 precipitation,22 electrochemical23 and emulsion polymerisation,24 the use of a solid-phase synthesis has gained prominence due to its ability to yield uniform and high-affinity particles.
The solid-phase protocol for nanoMIP production involves the immobilisation of the template/target molecule onto a solid support before polymerisation and because of polymerization reactions around the template/target molecule, subsequent nanoMIP is produced.25 Immobilisation of the template onto a solid support allows for a directional template, thus offering more control over the production of the imprinted binding cavities. This provides more uniform binding sites and the potential for 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding kinetics with higher performance. Additionally, the solid phase serves as a means for MIP/template separation method, allowing for the template to be easily removed from a high affinity nanoMIP. Magnetic nanoparticles and glass bead microparticles are the most used solid support, as they are easily functionalised with amino/carboxy silanes. This enables the use of a range of coupling chemistry (EDC/NHS, succinimidyl iodoacetate, glutaraldehyde) thus a variety of templates (small molecules, peptides and proteins) with bearing different functionality (carboxyl, thiol, or amino groups) are easily immobilised. The overall process is illustrated in Fig. 1, which schematically depicts key stages of solid-phase nanoMIP synthesis.
1 binding kinetics with higher performance. Additionally, the solid phase serves as a means for MIP/template separation method, allowing for the template to be easily removed from a high affinity nanoMIP. Magnetic nanoparticles and glass bead microparticles are the most used solid support, as they are easily functionalised with amino/carboxy silanes. This enables the use of a range of coupling chemistry (EDC/NHS, succinimidyl iodoacetate, glutaraldehyde) thus a variety of templates (small molecules, peptides and proteins) with bearing different functionality (carboxyl, thiol, or amino groups) are easily immobilised. The overall process is illustrated in Fig. 1, which schematically depicts key stages of solid-phase nanoMIP synthesis.
The nanoMIPs synthesized via this solid-phase approach typically incorporate acrylamide and acrylic acid-based monomers, with ammonium persulfate (APS) and tetramethylethylenediamine (TEMED) as the free radical initiator and catalyst, respectively. This provides relatively mild reaction conditions, whereby water is typically the solvent of choice. Using these conditions, a hydrogel-based nanoMIP is produced with particle sizes that range between 50–250 nm and have been successfully used in sensing and assay-based applications.26–28 Whilst application and performance of these particles is widely studied, knowledge is limited with regards to the material composition. Furthermore, little is reported on the molecular weight (MW) of the material, with the work of Guerreiro et al., making the best approximation of apparent molarities, using the expression given in eqn (1):
 | (1) |
Although this approximation is regularly used for estimating nanoMIPs molarities, and subsequent binding affinities, it relies on multiple simplifying assumptions. This model assumes that one nanoMIP constitutes one particle, and that each particle is uniform in shape and size, thus having the same molecular weight (MW). However, nanoMIPs typically exhibit polydispersity, with varying size range distributions depending on the synthesis conditions. Furthermore, the model assumes that the density of the particle is consistent throughout, with the density (ρ) estimation based on a non-imprinted nanoparticle consisting of N-tert-butylacrylamide (TBAm), N-isopropylacylamide (NIPAm), acrylic acid (AAc), crosslinked with N,N′-methylenebisacrylamide (BIS), without the mention of monomer crosslinker ratios.30 This density (ρ) estimation neglects changes in polymer composition with the use and addition of different functional monomers. Moreover, it also fails to consider the effects of the imprinting and resulting binding cavity on particle density and assumes that the monomer: assuming the final polymer composition mirrors the initial monomer feed. These assumptions could seriously affect the molarity estimations and can have majority implications, especially when used for binding affinity calculations, such as those produced by surface plasmon resonance (SPR) when the target is immobilised to the sensor surface.31,32 Henderson et al. addressed these challenges by immobilizing the nanoMIP on the sensor chip, thereby avoiding the need for precise molarity estimates. This, however, also presents potential issues, predominantly arising from the binding of a small molecule onto the immobilised, much larger nanoMIP.33,34 This generates smaller differences in binding signal intensities, which become more difficult to interpret, potentially limiting the sensitivity of detection and the reliability of the kinetic data.
These issues highlight the broader challenge in the field: the lack of standardized methodologies for nanoMIP synthesis, and a lack of consistency when evaluating their affinity and subsequent binding performance.35 This negatively impacts the use of these materials, particularly within commercial settings that rely on consistency within a product.36
This study aims to systematically evaluate the influence of commonly used solid-phase carriers (glass bead vs. magnetic nanoparticle) and with template characteristics on the synthesis of molecularly imprinted nanoparticles (nanoMIPs). The proteins bovine haemoglobin (BHb) and bovine serum albumin (BSA) were used as exemplary templates due to their similarity in size and hydrophobic solvent accessible surfaces areas (SASA),37,38 whilst having significantly different surface residues and surface binding cavities. These differences potentially affect functional monomer binding within the self-assembly phase. A mixture of functional monomers, acrylic acid, acrylamide, N-(hydroxyl methyl) acrylamide (chosen due to their hydrogen bond donor and acceptor) and N-isopropyl acrylamide (chosen due to its thermosensitive properties) (Fig. 2), were used.39,40 By understanding the relationship between sold-phase, template and monomer choice will lay the groundwork for rational nanoMIP design with improved reproducibility and performance.
Results and discussion
The nanoMIPs were synthesised using solid-phase protocols adapted from Hix-Janssens et al. for the magnetic nanoparticle solid-phase,41 and Canfarotta et al. for the glass bead solid-phase.42 These methodologies are widely considered as favourable ways to produce uniform, high-affinity nanoMIPs. The use of the magnetic nanoparticles as the solid-phase allows for ease of separation of the nanoMIP from the template using thermo-responsive materials and magnet separation.43 Whilst the use of the glass bead microparticles allows for easy separation of the nanoMIP from the template using thermo-responsive materials and filtration/solid-phase extraction techniques.44 To evaluate how these solid-phase materials affect nanoMIP properties, we compared their influence on polymer yield, size, composition, and binding performance using two protein templates—bovine haemoglobin (BHb) and bovine serum albumin (BSA)—which differ in isoelectric point (pI) and surface residue characteristics. As summarized in Table 1, BHb and BSA have similar molecular weights (66.5 and 64.5 kDa, respectively) but differ in their pI values, with BHb being nearly neutral (pI 7.1) and BSA more acidic (pI 4.7). These distinctions offer a controlled basis for assessing how surface charge and topology affect monomer self-assembly and imprinting efficiency.| Protein | Molecular weight (KDa) | pI Value |
|---|---|---|
| BHb | 66.5 | 7.1 |
| BSA | 64.5 | 4.7 |
The nanoMIPs were produced using a range of functional monomers at the constant molar ratio feed shown in Table 2. The functional monomers were chosen due to their water solubility and range of functional groups (NH2, CO, COOH, OH), and their well-established selective binding properties as hydrogen-bond donor and acceptor for protein-based hydrogel nanoMIPs.39 Furthermore, the inclusion of NIPAm as a functional monomer, enables temperature-controlled responsive properties, thus allowing the easy release of the template/target molecule, by increasing the temperature.33 This ratio is considered a bona fide favourable ratio for the popular solid-phase nanoMIP methodologies45 and produces a total monomer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) crosslinker percentage ratio of 34
crosslinker percentage ratio of 34![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. This is widely used for a range of targets that includes low molecular weight molecules (antibiotics, drugs of interest, performance and image enhancing drugs),33,46 peptides,19 and large macromolecules (proteins, enzymes, antibodies, viruses).47,48
1. This is widely used for a range of targets that includes low molecular weight molecules (antibiotics, drugs of interest, performance and image enhancing drugs),33,46 peptides,19 and large macromolecules (proteins, enzymes, antibodies, viruses).47,48
| Monomer | Monomer ratio (%) | |
|---|---|---|
| Functional monomer | NIPAm | 57.0 |
| AAm | 19.9 | |
| NHMAm | 13.6 | |
| AAc | 6.6 | |
| Crosslinker | BIS | 2.8 |
To establish a comparative protocol between the solid-phases, 5 g of template-derivatized magnetic nanoparticles was used vs. 30 g of template-derivatized glass beads. As well as different template amounts for immobilisation (7.5 mg for template-derivatized magnetic nanoparticles vs. 10 mg for template-derivatized glass beads). This difference reflects the substantially higher surface area and template loading capacity of the magnetic nanoparticles., which provides a greater reactive interface per unit mass compared to glass beads.20 Consequently, a smaller quantity of magnetic nanoparticles was sufficient to achieve an equivalent level of template presentation and comparative polymerisation yield. The yields reported were normalised to the amount of template-derivatised solid phase used in each synthesis, allowing direct comparison between the two supports. This proportional scaling is consistent with previous nanoMIP synthesis methodologies.20,42
NanoMIP materials were successfully produced and developed using a magnetic and glass bead solid-phase approach for the targets BHb and BSA. Approximately 100 mL of nanoMIP solution was produced for each nanoMIP, with the yield of the material estimated by taking a 3 mL sample and incubating in a drying oven at 60 °C until dry and measuring the particle masses with a 6-point balance. The hydrodynamic diameter of the nanoMIPs was then estimated using dynamic light scattering (DLS), to measure a dilute solution of the nanoMIPs. Table 3 and Fig. 3 present the comparative yield (in mg) and size (in nm) of nanoMIPs synthesized using magnetic nanoparticles or glass beads, for both BHb and BSA templates.
| Protein template | Glass bead | Magnetic nanoparticle | ||
|---|---|---|---|---|
| Yield (mg) | Size (nm) | Yield (mg) | Size (nm) | |
| BHb | 19.53 ± 3.2 | 25.8 ± 3.8 | 9.85 ± 3.8 | 49.0 ± 7.0 |
| BSA | 13.04 ± 2.8 | 23.2 ± 2.9 | 9.50 ± 1.3 | 53.8 ± 13.8 |
Table 3 shows significant differences between the yield and size of the nanoMIPs produced by the different solid-phase protocols (magnetic nanoparticles vs. glass bead). The nanoMIPs produced using the glass bead method produces a much smaller size (25.8 nm and 23.2 nm for BHb and BSA, respectively) compared with the nanoMIPs produced using the magnetic nanoparticle as the solid-phase (49.0 nm and 53.8 nm for BHb and BSA, respectively). These nanoMIPs are overall much smaller than those presented in literature,29,31 potentially due differences in polymer composition. Additionally, this is the first demonstration of nanoMIP size differences as result of different solid phase used for production. The SEM images (Fig. 4) show that the magnetic solid-phase nanoparticles form clusters (when dried), whereas more uniform and spherical nanoparticles in appearance can be appreciated via the glass bead solid-phase, furthermore estimation of nanoMIP size these images (Fig. 4), confirm the sizes from DLS (Fig. S1 and Table S1). The yield of the nanoMIPs produced also show a variation that is dependent of the solid phase used, with the glass bead method producing 19.53 mg and 13.04 mg (for BHb and BSA, respectively) compared with the magnetic nanoparticle method producing 9.85 mg and 9.50 mg (for BHb and BSA, respectively). Although these yields are consistent with that the amount of nanoMIPs that are typically produced using a solid-phase protocol,46 this work demonstrates yet another clear difference in the nanoMIPs produced via the different protocols. With these observed differences in nanoMIP yield and particle size potentially attributed to variations in the physicochemical properties of the solid-phase carriers. Magnetic nanoparticles possess a highly curved, high surface–area interface that allows for greater monomer diffusion and polymer chain extension before detachment, resulting in larger particles. Whereas the relatively flat and less reactive surface of the glass beads promotes localised polymerisation with restrictive chain growth leading to the formation of smaller more uniform nanoMIPs.20,44
To assess the binding affinity and specificity of the synthesized nanoMIPs, we employed surface plasmon resonance (SPR), a label-free optical technique capable of quantifying real-time molecular interactions.49 Each nanoMIP formulation was immobilized onto a carboxymethyl dextran-coated gold chip via EDC/NHS coupling, exploiting the presence of primary amine groups within the polymer matrix. This immobilization strategy enables direct evaluation of target binding without requiring assumptions about the nanoMIP's molar concentration, which is often poorly defined due to particle polydispersity and uncertain internal density.
Unlike conventional approaches where the target is immobilized and nanoMIPs are injected as analytes,31 our setup of immobilizing the nanoMIPs on to the surface, circumvents uncertainties in molarity estimates and allows for more accurate determination of binding constants. Although the total quantity of nanoMIP bound to the chip is not precisely known, the use of excess material during immobilization ensures maximal surface coverage. Given the design of solid-phase nanoMIPs and their expected monovalent architecture (one dominant binding site per particle), we modelled the interaction using a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 Langmuir binding framework.
1 Langmuir binding framework.
Fig. 5 presents representative SPR sensorgrams showing the interaction of five different concentrations of each protein target with their corresponding immobilized nanoMIP. Panels A and B illustrate BHb binding to nanoMIPs synthesized on glass beads and magnetic nanoparticles, respectively; panels C and D show the equivalent for BSA. In all cases, the sensorgrams exhibit concentration-dependent association and dissociation phases, allowing for robust determination of equilibrium binding constants.
The KD values derived from Fig. 5 and presented in Table 4 show that, overall, the nanoMIPs display excellent affinity towards the chosen targets, with KD values within the nanomolar range. These values are consistent with those currently presented in literature for protein targeted nanoMIPs, albeit with a different functional monomer construction. The values presented in Table 4 also show that differences in the affinity of the materials produced occur dependent upon the solid-phase used. While these differences are minor, it appears that the nanoMIPs produced using the glass bead solid-phase show an approximately 10-fold improvement in affinity compared with the nanoMIPs produced using the magnetic nanoparticle solid-phase. This could potentially be due to the differences in size of the nanoMIPs that were produced, with the glass bead methodology producing a smaller nanoMIP compared with the magnetic nanoparticle methodology. Typically, the smaller nanoMIP size produces a higher surface-to-volume ratio. These smaller particles expose more binding sites on their surface, allowing faster mass transfer and recognition, leading to an increase in sensitivity and binding affinity.20 Furthermore, the use of a solid-phase to immobilise templates, allows for precise control over the polymerisation conditions and binding site formations. This leads to highly reproducible affinity characteristics, and with the well optimised synthesis methodology, KD values remain within a single order of magnitude between batches.20,42 Thus when immobilisation and washing steps are carefully standardised, batch-to-batch variation becomes negligible.50
| Imprinted nanoMIP | K D (M) | |
|---|---|---|
| Glass bead | Magnetic nanoparticle | |
| BHb | 2.57 × 10−9 | 2.96 × 10−8 |
| BSA | 5.30 × 10−8 | 8.43 × 10−7 |
The specificity of the nanoMIPs was investigated through the binding of a non-target protein onto the nanoMIP coated chips. This allows for cross-reactivity and non-specific binding exploration, with BHb binding to a BSA imprinted nanoMIP and BSA binding to a BHb imprinted nanoMIP. The KD values for these interactions are present in Table 5, with the comparative SPR sensorgrams shown in Fig. S2.
| Imprinted nanoMIP | K D (M) | |
|---|---|---|
| Glass bead | Magnetic nanoparticle | |
| BHb | 4.76 × 10−6 | 2.23 × 10−5 |
| BSA | 1.26 × 10−6 | 3.03 × 10−5 |
When investigated with non-target proteins, the nanoMIPs materials show that they are specific for the target proteins, with an approximately 100–1000-fold decrease in affinity towards a non-target protein. This is to be expected with nanoMIPs produced using a solid-phase, as other works also show similar decreases in affinities towards a non-target protein. These affinity differences are also consistent across different solid phases (glass bead vs. magnetic nanoparticle). In this case, the nanoMIPs produced using a magnetic solid phase show an approximate 10-fold decrease in affinity, compared with the nanoMIPs produced using the glass bead solid phase. This suggests that the nanoMIPs that are produced using the magnetic solid phase are overall less selective (by approximately 10-fold) compared with those produced using the glass bead solid phase. The proteins used in this study where specifically chosen based on their structural and functional similarity, allowing for comparative analysis. Thus, representing the most stringent and physiologically relevant challenge to the biding sites. The use of the solid-phase protocol has consistently shown to produce nanoMIPs with homogeneity and low non-specific binding.20,42 Since the nanoMIPs in this work demonstrated negligible cross-reactivity under these conditions, it should be expected that less related proteins would also not show any binding.
There is a current assumption that the polymer composition of the nanoMIP, produced via the solid-phase, is consistent (in terms of molar ratios) with the monomer mixture feed. However, this assumption neglects any impact the template may have on the polymer composition during nanoMIP formation. With differences in template surface functionality, it is then logical to suggest that different functional monomers will be attracted to these binding sites, particularly during a self-assembly process. The variation in target molecule should therefore significantly impact the final output on nanoMIP polymer composition, with variation in template molecules. NMR spectroscopy is a well-established method for the characterization of polymers and has been applied for the determination of monomer sequence and reactivity ratios and polymer microstructure.51 Utilising 1H NMR, an assessment on the number of repeating units in the overall nanoMIP was achieved with an estimation of the relative composition of the nanoMIP. The calculation of the relative number of moles of a polymer block (denoted as x) can be calculated using eqn (2):
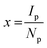 | (2) |
With, for one type of polymer block, Ip being the integration value of the polymer block's corresponding peak, and Np being the total number of H protons (or active nuclei) for that polymer block. By determining the relative number of moles per polymer block, the composition of the nanoMIP as ratio can be estimated.51
The 1H NMR spectra shown in Fig. S3–S6, show the polymer composition of the nanoMIPs, with a specific resonance for each monomer. Utilising eqn (2), the overall polymer composition of the nanoMIPs was estimated and shown in Table 6, with a full breakdown of calculations showin in eqn (S1)–(S9).
| NanoMIP | Polymer composition (%) | Monomer: crosslinker ratio | ||||
|---|---|---|---|---|---|---|
| NIPAm | AAm | NHMAm | AAc | BIS | ||
| BHb G | 30 | 11 | 41 | 4 | 14 | 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| BHb M | 20 | 10 | 4 | 17 | 6 | 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| BSA G | 20 | 11 | 39 | 0 | 30 | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| BSA M | 12 | 17 | 47 | 17 | 17 | 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
The polymer composition (%) shown in Table 6 reveal interesting differences between the composition of the monomer/crosslinker mixture inputted compared to the monomer/crosslinker composition of the polymers produced. The overall monomer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) crosslinker ratio that is inputted was calculated to be 34
crosslinker ratio that is inputted was calculated to be 34![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (monomer: crosslinker, respectively), which has been deemed favourable in either glass bead or magnetic nanoparticle-based solid phase protocols. Surprisingly, the overall monomer: crosslinker ratio that is produced within the nanoMIPs is drastically different compared to the ratio in the feed: the final monomer
1 (monomer: crosslinker, respectively), which has been deemed favourable in either glass bead or magnetic nanoparticle-based solid phase protocols. Surprisingly, the overall monomer: crosslinker ratio that is produced within the nanoMIPs is drastically different compared to the ratio in the feed: the final monomer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) crosslinker ratio is equal to a range of 2
crosslinker ratio is equal to a range of 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1–9
1–9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (monomer
1 (monomer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) crosslinker, respectively). The monomer
crosslinker, respectively). The monomer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) crosslinker ratios are much more consistent with the monomer
crosslinker ratios are much more consistent with the monomer![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) crosslinker ratios (9
crosslinker ratios (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) that were seen in traditional bulk polyacrylamide hydrogel-based MIPs,39,52 which themselves were adapted from polyacrylamide gels used for electrophoresis.53,54 This suggests that this lower monomer: crosslinker is favourable under these simple polymerisation conditions.
1) that were seen in traditional bulk polyacrylamide hydrogel-based MIPs,39,52 which themselves were adapted from polyacrylamide gels used for electrophoresis.53,54 This suggests that this lower monomer: crosslinker is favourable under these simple polymerisation conditions.
Table 6 also shows the polymer composition varies considerably between template used, as well as with varying solid phase. Differences between polymer composition with varying the target can be rationally explained due to differences in the surface of the protein targets. Sullivan et al. show that protein templates produce multiple potential binding cavities (for functional monomers) that are evenly distributed around the surface of the protein.39 These binding cavities are used by the functional monomers during monomer–template complexation with varying degrees of affinity depending on the site or monomer.39 Potential binding cavity analysis of the protein targets BHb and BSA performed by molecular modeling (CavitOmix) 655,56 reveal that BHb has 27 potential binding cavities evenly spaced around the surface of the protein (Fig. 6A), whereas BSA has 68 potential binding cavities around the surface of the protein (Fig. 6B). Along with having a significantly different amount of potential binding cavities, these cavities also differ in shape and size and amino acid functionality. This is shown with the differences in the solvent-accessible surface area (SASA) calculated using APBS (v3.4.1),57 and shown in Fig. 6C and D, for BHb and BSA, respectively, with BSA showing much more positive electrostatic potential (blue) regions compared with BHb. This means that different monomers will favourably bind to the different potential binding cavities,39 thus affecting the polymer composition of nanoMIP that is produced. More surprisingly, the changing of the solid phase, while keeping the same template, appears to be influencing the final polymer composition of the nanoMIP. Table 6 shows that using the glass bead as the solid-phase produces a polymer with a higher amount of NHMAm and a lower amount of AAc in the final composition compared to the magnetic solid-phase nanoparticles. These findings are also consistent for both the BHb and BSA template nanoMIPs, and potentially due to differences within the solid-phase size and composition.
 | ||
| Fig. 6 Potential binding cavity analysis of the target proteins BHb (A) and BSA (B) using CavitOmix and solvent-accessible surface area (SASA)55,56 of the target proteins BHb (C) and BSA (D) calculated using APBS (v3.4.1).57 The blue and red gradients correspond to positive electrostatic potential and negative electrostatic potential, respectively. The BSA shows much more positive electrostatic potential regions compared with BHb. | ||
The glass bead solid phase is predominately made of SiO2,58 while the magnetic nanoparticles are Fe3O4.59 Furthermore, the glass beads are also approximately 350 times larger than the magnetic nanoparticles. Glass beads present hydrophilic, hydroxyl-rich surfaces that differ in charge density and hydrogen-bonding capacity from the typically functionalised magnetic nanoparticles. These distinctions influence template immobilisation, local monomer orientation, and polymer chain propagation during polymerisation. Moreover, the markedly higher curvature and surface energy of magnetic nanoparticles alter monomer packing and radical diffusion dynamics at the interface, producing compositional and structural differences in the imprinted polymer layer.20,26,44 These differences will influence the functionalisation and immobilisation of the template to the solid-phase and thus will also impact the template–monomer interactions, along with solid-phase–monomer interactions, during the complexation process.
Collectively, these findings represent a rigorous investigation into how solid-phase choice and protein template impact nanoMIP synthesis at the compositional level. By linking physical imprinting conditions to polymer architecture and ultimately to functional performance, this study provides a framework for rational nanoMIP design. This work shows that the template and choice of solid phase can have a significant impact on the final polymer composition, which in turn can have a substantial impact on the performance of the nanoMIP. This suggests that the nanoMIPs should be rationally designed, with monomer choice dependent upon the solid phase used and the template needed, to achieve maximum performance. This highlights that there is no “one size fits all” with regards to pre-polymerisation monomer mixture or choice of solid phase, with careful considerations needed. Particularly, these highly specific and selectivity recognition materials offer a viable alternative in terms of performance to biological recognition materials (antibodies), but with the additional benefits of being low-cost, resistant to high and low pH and temperatures values, and having simple storage conditions (room temperature) without degradation that affects the performance.
Experimental
Materials and methodology
Acrylamide (AAm), acrylic acid (AAc), 3-aminopropyltriethyloxy-silane (APTES), ammonium persulfate (APS), bovine hemoglobin (BHb), bovine serum albumin (BSA), dimethyl sulfoxide-d6, ethanolamine, 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC), glutaraldehyde (GA), glycine, magnetic nanoparticles (100 nm diameter), N-(hydroxylmethyl)acrylamide (NHMAm), N-hydroxysuccinimide (NHS), N-isopropylacrylamide (NIPAm), N,N′-methylenebisacrylamide (BIS), phosphate buffer saline (PBS) pellets, potassium chloride (KCl), sodium chloride (NaCl), sodium hydroxide (NaOH), tetramethylethyldiamide (TEMED), and Tween20 were all purchased and used without purification from Sigma-Aldrich (Japan).Glass beads (35 μm) were purchased and used without purification from Axel (Japan).
Acetone, acetonitrile (anhydrous), ethanol (anhydrous), isopropanol, methanol, toluene (anhydrous), dipotassium phosphate, and disodium phosphate were all purchased and used without purification from Nacalai Tesque (Japan).
Preparation of protein immobilised magnetic nanoparticles
The magnetic nanoparticle protocol was adapted from Hix-Janssens et al.41 Iron oxide (magnetic) particles (30 g, 100 nm diameter) were activated by ultrasonication in 20 mL anhydrous ethanol for 3 hours. The iron oxides particles were then incubated with 2 mL APTES overnight at room temperature. Next, the particles were washed once with anhydrous ethanol then five times with MilliQ. Afterwards, 7% (v/v) glutaraldehyde was added and the mixture was left to react for two hours at room temperature. The target protein (10.0 mg at 0.5 mg mL−1) was introduced to the glutaraldehyde-modified beads in 1 mL phosphate buffered saline (PBS, 10 mM) pH 7.4, with the solution being left to incubate at room temperature overnight. The particles were washed thoroughly with MilliQ water and dried under vacuum. After this step, the iron oxide particles were used straight away for the synthesis of the imprinted nanoparticles without further storage.Preparation of protein immobilised glass beads
The glass bead solid-phase synthesis protocol was adapted from Canfarotta et al.42 30 g glass beads (35 μm diameter, Axel) were activated by boiling in 4 M NaOH (24 mL) for 15 min, then washed thoroughly with MilliQ water (eight times with 100 mL, for 30 g of beads) and with acetone (100 mL, two washes). The glass beads were then dried at 60 °C for 2 h and were then left on a roller with APTES (3%, v/v) in anhydrous toluene overnight at 60 °C. The glass beads were washed with 8 volumes (100 mL) of acetone, followed by 2 volumes (100 mL) of methanol, and dried in an oven at 60 °C for one hour. The beads were then incubated in a 7% (v/v) glutaraldehyde solution (0.5 mL of solution per gram of beads) for 2 h at room temperature. After once again washing the glass beads with 8 volumes of MilliQ water, the target protein (7.5 mg at 0.5 mg mL−1) was introduced to the glutaraldehyde-modified beads in 15 mL phosphate buffered saline (PBS, 10 mM) pH 7.4, with the solution being left to incubate at room temperature overnight, sealed under nitrogen. The derivatized beads were washed thoroughly with MilliQ water and dried under vacuum. After this step, the glass beads were used straight away for the synthesis of the imprinted nanoparticles without further storage.Polymerization reaction
A polymerization mixture consisting of 20 mg N-isopropylacrylamide (NIPAm), 1 mg N,N′-methylenebisacrylamide (BIS), 4.8 mg N-(hydroxymethyl)acrylamide (NHMAm), 7 mg acrylamide (AAm), and 2.2 μL acrylic acid (AAc) in 49 mL of double distilled water was produced. The chemical structures of the functional monomers—AAm, NHMAm, NIPAm, and AAc—are shown in Fig. 2. Double distilled water was added to the flask with the polymerization mixture to adjust the total volume to 50 mL. This solution was degassed under vacuum, while sonicating for 10 min. Following this, the polymerization mixture was bubbled for 20 min with a slow stream of N2. Next, 5 g of template-derivatized magnetic nanoparticles or 30 g of template-derivatized glass beads were transferred into a 100 mL sealable bottle and degassed by purging with N2 for 10 min. Next, the 50 mL of degassed solution of monomers was poured onto the glass beads, followed by the rapid addition of 12.5 μL of TEMED and 15 mg of APS dissolved in 250 μL of double distilled water to allow for polymerization to be initiated. Everything was gently agitated via swirl motion and left to polymerize for 2 h at room temperature, gently swirling from time to time to limit risk of abrasion. After the synthesis, the solid phases were filtered through a 11 μm filter paper using gravity filtration, then washed with 8 × 30 mL of water at room temperature to remove the impurities, unreacted monomers, and low affinity nanoMIPs. To elute high affinity nanoMIPs, the beads were heated in 40 mL of water at 60 °C, then filtered again using gravity filtration. The beads were then further washed with volumes of 20 mL of water at 60 °C until ≈100 mL of the eluted nanoparticles in water was collected. The solutions of all nanoMIPs were stored at 4 °C until measurements were taken.Yield, size and morphology estimations of molecularly imprinted nanoparticles (NanoMIPs)
Four 3 mL aliquots of the solution were oven-dried at 60 °C and the mass of the particles measured using a 6-point balance, allowing for a concentration (in μg mL−1) of the initial solution to be calculated.Particle size at 25 °C (effective hydrodynamic radius) and polydispersity index were measured using dynamic light scattering in MilliQ with Zeta-potential & Particle size Analyzer ELSZ-2000.
Scanning electron microscopy (JEOL JSM-7900F) was used to observe the morphology and dispersion of the synthesized nanoMIPs. The sample was prepared by mixing the MIPs with 1 mL MilliQ and vortexing the solution for 1 minute, then leaving one drop on one gold chip and leaving it to dry overnight. The next day, the chip is coated with gold and ready for SEM observation.
Binding and affinity performance by surface plasmon resonance (SPR) of molecularly imprinted nanoparticles (NanoMIPs)
Immobilisation of the nanoMIP onto the SPR sensors surface occurs through a carboxymethyl dextran hydrogel coated Au chip initially being pre-conditioned by a PBS pH 7.4 and 0.01% Tween 20 running buffer at 10 μL min−1 within the SPR. 1 mL of aqueous solution containing 40 mg EDC and 10 mg NHS was passed over the chip (6 min at 10 μL min−1). 300 μg of nanoMIPs in 1 mL of running buffer and 10 mM sodium acetate were injected over the left channel (working channel) of the chip for 1 min. The amine groups on the nanoMIP react with the functionalized surface, leading to particle immobilization. An 8 min injection over both channels (working and reference) of quenching solution (1 M ethanolamine, pH 8.5) was added, followed by a continuous flow of running buffer at 10 μL min−1. All injections were taken from a stable baseline.Kinetic analysis in rebinding of analyte (target and cross-reactivity) to the nanoMIP was performed in set pattern of 2 min association (PBST with analyte in range of 4.0–64.0 nM), 5 min dissociation (running buffer only), and a regeneration cycle (regeneration buffer 10 mM glycine–HCl, pH 2 for 1 min) followed by a final stabilization cycle (running buffer for 1 min). An initial injection of running buffer (blank) was used as the first run with increasing analyte concentration for subsequent runs. After the analyses were completed, signals from reference channel were subtracted from signals from the working. In all cases, rebinding was studied in triplicate. The SPR responses were fitted to a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 Langmuir fit bio-interaction (BI) model using the Biacore T200 evaluation software. Association rate constants (ka), dissociation rate constants (kd), and maximum binding (Bmax) were fitted globally, whereas the BI signal was fitted locally. Equilibrium dissociation constants (KD) were calculated by kd/ka. For each nanoMIP/analyte combination, a calibration curve was generated across the concentration range 4.0–64.0 nM taking n = 3 average. Where signal saturation was observed (noted in more complex matrices), the linear section of the curve was used for this calculation.
1 Langmuir fit bio-interaction (BI) model using the Biacore T200 evaluation software. Association rate constants (ka), dissociation rate constants (kd), and maximum binding (Bmax) were fitted globally, whereas the BI signal was fitted locally. Equilibrium dissociation constants (KD) were calculated by kd/ka. For each nanoMIP/analyte combination, a calibration curve was generated across the concentration range 4.0–64.0 nM taking n = 3 average. Where signal saturation was observed (noted in more complex matrices), the linear section of the curve was used for this calculation.
1H NMR analysis of molecularly imprinted nanoparticles (NanoMIPs) for polymeric composition
1H NMR analysis was performed on a Bruker Topspin 500 MHz spectrophotometer with 16 scans. For analysis, 5 mg of the nanoMIP material was dissolved with gentle stirring (overnight) and heat (80 °C) into in 1 mL DMSO-d6.60 Obtained spectra were then processed and analysis using TopSpin 4.5.0.Computational protein surface analysis of protein (BHb and BSA) templates
Potential binding cavities across the surface of BSA (PDB code: 4F5S) and BHb (PDB code: 2QSS) were calculated using CavitOmix (v1.0, 2022, Innophore GmbH). The corresponding hydrophobicity module of the program VASco55 was used to analyse the hydrophobicity of the cavities. The cavities were calculated using a modified LIGSITE algorithm.56 The settings include a minimum of five site points and a maximum of 100 site points per site. The sites generated were produced with a grid spacing of 0.7 between potential binding cavities.The surface of the protein was analysed by using APBS (v3.4.1), with the protein initially prepared using PDB2PQR (v3.6.1) at a pKa of 7.0 and an AMBER forcefield, ensuring that waters are removed, new atoms not rebuilt too close to current aaatoms, and optimizing the hydrogen bonding network.57 Adaptive Poisson–Boltzmann solver (APBS) was then subsequently used with mg-auto calculation time, to determine electrostatic surface visualization.57
Conclusions
This study provides a comprehensive evaluation of how both template identity and solid-phase material influence the polymer composition and binding performance of molecularly imprinted nanoparticles (nanoMIPs) synthesized via solid-phase protocols. Through the combined use of surface plasmon resonance (SPR) and nuclear magnetic resonance (NMR) spectroscopy, we show that seemingly minor changes in synthesis conditions can lead to measurable differences in polymer structure and function. SPR analysis revealed that nanoMIPs prepared using glass beads exhibited approximately ten-fold greater binding affinity and selectivity than those produced using magnetic nanoparticles, despite using identical pre-polymerization mixtures. These findings were further supported by NMR-derived compositional data, which demonstrated significant variations in monomer incorporation dependent on both the template protein and the solid-phase substrate. Specifically, nanoMIPs synthesized on glass beads contained higher proportions of NHMAm and lower amounts of AAc compared to those synthesized on magnetic nanoparticles.In parallel, protein surface analysis using computational cavity mapping and electrostatic surface potential modelling revealed that differences in the number, distribution, and physicochemical nature of binding sites on the template proteins likely contribute to template-specific polymer compositions. For instance, BSA, with its higher number of binding cavities and greater surface polarity, drove a distinct monomer incorporation profile compared to BHb.
Taken together, these findings challenge the assumption of compositional uniformity in nanoMIPs and highlight the importance of rational design in imprinted polymer synthesis. Our results suggest that template structure, solid-phase material, and monomer selection should be co-optimized to achieve high-performance nanoMIPs with enhanced affinity, selectivity, and reproducibility. This work lays the groundwork for more predictive and standardized approaches to nanoMIP fabrication, with broad implications for their application as synthetic recognition elements in diagnostics, sensing, and therapeutic platforms.
Author contributions
MVS and AS designed the study, interpreted the data, and wrote the manuscript; TT and TI performed nanoMIP synthesis and characterisation. MS performed NMR polymer composition and computational protein analysis. All authors contributed to manuscript writing and editing.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported here.Data availability
The data supporting this article have been included as part of the supplementary information (SI). See DOI: https://doi.org/10.1039/d5tb01580d.Acknowledgements
The authors gratefully acknowledge the support of Okinawa Institute of Science and Technology Graduate University (OIST) with subsidy funding from the Cabinet Office, Government of Japan. We are also grateful for the help and support provided by the Scientific Imaging Section and Instrumental Analysis Section of Core Facilities at OIST.Notes and references
- L. Y. Chu, T. Yamaguchi and S. Nakao, Adv. Mater., 2002, 14, 386–389 CrossRef CAS.
- M. J. Langton, C. J. Serpell and P. D. Beer, Angew. Chem. Int. Ed., 2016, 55, 1974–1987 CrossRef CAS PubMed.
- R. Reverberi and L. Reverberi, Blood Transfus., 2007, 5, 227–240 Search PubMed.
- R. Rohs, S. M. West, A. Sosinsky, L. Peng, R. S. Mann and B. Honig, Nat. London, 2009, 461, 1248–1253 CrossRef CAS PubMed.
- V. Wittmann and R. J. Pieters, Chem. Soc. Rev., 2013, 42, 4492–4503 RSC.
- A. Bastide and A. David, Biomol., 2018, 8, 100 Search PubMed.
- K. C. Nicolaou and S. Rigol, J. Antibiot., 2018, 71, 153–184 CrossRef CAS PubMed.
- S. Firdous, S. Anwar and R. Rafya, Laser Phys. Lett., 2018, 15, 65602 CrossRef.
- B. Van Dorst, J. Mehta, K. Bekaert, E. Rouah-Martin, W. De Coen, P. Dubruel, R. Blust and J. Robbens, Biosens. Bioelectron., 2010, 26, 1178–1194 CrossRef CAS PubMed.
- N. Bhalla, P. Jolly, N. Formisano and P. Estrela, Essays Biochem., 2016, 60, 1–8 CrossRef PubMed.
- P. Mehrotra, J. Oral. Biol. Craniofac. Res., 2016, 6, 153–159 CrossRef PubMed.
- M. A. Morales and J. M. Halpern, Bioconjugate Chem., 2018, 29, 3231–3239 CrossRef CAS PubMed.
- C. I. L. Justino, A. C. Freitas, R. Pereira, A. C. Duarte and T. A. P. Rocha Santos, TrAC, Trends Anal. Chem., 2015, 68, 2–17 CrossRef CAS.
- V. J. B. Ruigrok, M. Levisson, M. H. M. Eppink, H. Smidt and J. Van Der Oost, Biochem. J., 2011, 436, 1–13 CrossRef CAS PubMed.
- P. Chames, M. Van Regenmortal, E. Weiss and D. Baty, Br. J. Pharmacol., 2009, 157, 220–233 CrossRef CAS PubMed.
- G. Vasapollo, R. Del Sole, L. Mergola, M. R. Lazzoi, A. Scardino, S. Scorrano and G. Mele, Int. J. Mol. Sci., 2011, 12, 5908–5945 CrossRef CAS PubMed.
- C. Widstrand, E. Yilmaz, J. Billing and A. Rees, Am. Lab., 2006, 38, 12 CAS.
- P. A. G. Cormack and A. Z. Elorza, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2004, 804, 173–182 CrossRef CAS PubMed.
- Y. Hoshino, T. Kodama, Y. Okahata and K. J. Shea, J. Am. Chem. Soc., 2008, 130, 15242–15243 CrossRef CAS PubMed.
- A. Poma, A. P. Turner and S. A. Piletsky, Trends Biotechnol., 2010, 28, 629–637 CrossRef CAS PubMed.
- M. V. Sullivan, B. Dean, A. Mates, M. E. Farrow, C. Fletcher, M. German, R. Patel and N. W. Turner, Nano Express, 2023, 4, 025002 CrossRef CAS.
- R. M. Roland, S. A. Bhawani and M. N. M. Ibrahim, BMC Chem., 2023, 17, 165 CrossRef CAS PubMed.
- P. S. Sharma, A. Pietrzyk-Le, F. D’Souza and W. Kutner, Anal. Bioanal. Chem., 2012, 402, 3177–3204 CrossRef CAS PubMed.
- M. V. Sullivan, C. Fletcher, R. Armitage, C. Blackburn and N. W. Turner, Nanoscale Adv., 2023, 5, 5352–5360 RSC.
- T. Muhammad, D. Zhao, A. Guerreiro, I. Muhammad, M. Aimaitiniyazi, B. Ding, Y. Zheng, T. Yimamumaimaiti, L. Chen and S. A. Piletsky, TrAC, Trends Anal. Chem., 2025, 184, 118134 CrossRef CAS.
- F. Canfarotta, S. A. Piletsky and N. W. Turner, Methods Mol. Biol., 2020, 2073, 183–194 CrossRef CAS PubMed.
- F. Canfarotta, J. Czulak, K. Betlem, A. Sachdeva, K. Eersels, B. van Grinsven, T. J. Cleij and M. Peeters, Nanoscale, 2018, 10, 2081–2089 RSC.
- T. Hix-Janssens, S. Shinde, R. Abouhany, J. Davies, J. Neilands, G. Svensäter and B. Sellergren, ACS Omega, 2023, 8, 15259–15265 CrossRef CAS PubMed.
- A. Guerreiro, A. Poma, K. Karim, E. Moczko, J. Takarada, I. P. De Vargas-Sansalvador, N. Turner, E. Piletska, C. S. De Magalhães, N. Glazova, A. Serkova, A. Omelianova and S. Piletsky, Adv. Healthcare Mater., 2014, 3, 1426–1429 CrossRef CAS PubMed.
- J. D. Debord and L. A. Lyon, Langmuir, 2003, 19, 7662–7664 CrossRef CAS.
- A. H. M. Safaryan, A. M. Smith, T. S. Bedwell, E. V. Piletska, F. Canfarotta and S. A. Piletsky, Nanoscale Adv., 2019, 1, 379–3714 RSC.
- J. Bezdekova, F. Canfarotta, F. Grillo, H. Yesilkaya, M. Vaculovicova and S. Piletsky, Nanoscale Adv., 2023, 5, 2602–2609 RSC.
- A. Henderson, M. V. Sullivan, R. A. Hand and N. W. Turner, J. Mater. Chem. B, 2022, 10, 6792–6799 RSC.
- M. V. Sullivan, A. Henderson, R. A. Hand and N. W. Turner, Anal. Bioanal. Chem., 2022, 414, 3687–3696 CrossRef CAS PubMed.
- L. Carballido, T. Karbowiak, P. Cayot, M. Gerometta, N. Sok and E. Bou-Maroun, Chem., 2022, 8, 2330–2341 CAS.
- R. Raghuwanshi and M. Singh, Challenges to molecularly imprinted polymers in competition with biosensors at commercial scale, Elsevier, 2023, pp. 345–366 Search PubMed.
- S. Wu, W. Tan and H. Xu, Anal, 2010, 135, 2523–2527 RSC.
- Z. Hua, Z. Chen, Y. Li and M. Zhao, Langmuir, 2008, 24, 5773–5780 CrossRef CAS PubMed.
- M. V. Sullivan, S. R. Dennison, G. Archontis, J. M. Hayes and S. M. Reddy, J. Phys. Chem. B, 2019, 123, 5432–5443 CrossRef CAS.
- M. V. Sullivan, P. Lasserre, C. Blackburn, N. W. Turner and B. Sellergren, Responsive Mater., 2025, 3, e20240032 CrossRef CAS.
- T. Hix-Janssens, J. R. Davies, N. W. Turner, B. Sellergren and M. V. Sullivan, Anal. Bioanal. Chem., 2024, 416, 7305–7316 CrossRef CAS PubMed.
- F. Canfarotta, A. Poma, A. Guerreiro and S. A. Piletsky, Nat. Protoc., 2016, 11, 443–455 CrossRef CAS PubMed.
- H. Musarurwa and N. T. Tavengwa, Microchem. J., 2022, 181, 107750 CrossRef CAS.
- F. Canfarotta, A. Cecchini and S. Piletsky, In Polym. Chem. Ser., 2018, 1–27 CAS.
- T. S. Bedwell, N. Anjum, Y. Ma, J. Czulak, A. Poma, E. Piletska, M. J. Whitcombe and S. A. Piletsky, RSC Adv., 2019, 9, 27849–27855 RSC.
- C. Blackburn, M. V. Sullivan, M. I. Wild, A. J. O’ Connor and N. W. Turner, Anal. Chim. Acta, 2024, 1285, 342004 CrossRef CAS PubMed.
- A. Gómez-Caballero, A. Elejaga-Jimeno, G. García del Caño, N. Unceta, A. Guerreiro, M. Saumell-Esnaola, J. Sallés, M. A. Goicolea and R. J. Barrio, Microchim. Acta, 2021, 188, 368 CrossRef PubMed.
- P. X. Medina Rangel, S. Laclef, J. Xu, M. Panagiotopoulou, J. Kovensky, B. Tse Sum Bui and K. Haupt, Sci. Rep., 2019, 9, 3923 CrossRef PubMed.
- Y. Tang, X. Zeng and J. Liang, J. Chem. Educ., 2010, 87, 742–746 CrossRef CAS PubMed.
- S. Subrahmanyam, A. Guerreiro, A. Poma, E. Moczko, E. Piletska and S. Piletsky, Eur. Polym. J., 2013, 87, 100–105 CrossRef.
- J. U. Izunobi and C. L. Higginbotham, J. Chem. Educ., 2011, 88, 1098–1104 CrossRef CAS.
- D. Tong, C. S. Heényi, Z. S. Bikádi, J.-P. Gao and S. Hjertén, Chromatogr., 2001, 54, 7–14 CrossRef CAS.
- K. Takegami, Y. Kaneko, T. Watanabe, T. Maruyama, Y. Matsumoto and H. Nagawa, Ultrasound Med. Biol., 2004, 30, 1419–1422 CrossRef PubMed.
- M. O. Culjat, D. Goldenberg, P. Tewari and R. S. Singh, Ultrasound Med. Biol., 2010, 36, 861–873 CrossRef PubMed.
- G. Steinkellner, R. Rader, G. G. Thallinger, C. Kratky and K. Gruber, BMC Bioinf., 2009, 10, 32 CrossRef PubMed.
- M. Hendlich, F. Rippmann and G. Barnickel, J. Mol. Graphics Model., 1997, 15, 359–363 CrossRef CAS PubMed.
- E. Jurrus, D. Engel, K. Star, K. Monson, J. Brandi, L. E. Felberg, D. H. Brookes, L. Wilson, J. Chen, K. Liles, M. Chun, P. Li, D. W. Gohara, T. Dolinsky, R. Konecny, D. R. Koes, J. E. Nielsen, T. Head-Gordon, W. Geng, R. Krasny, G. Wei, M. J. Holst, J. A. McCammon and N. A. Baker, Protein Sci., 2018, 27, 112–128 CrossRef CAS PubMed.
- M. A. Presley and P. R. Christensen, J. Geophys. Res.: Planets, 1997, 102, 6551–6566 CrossRef CAS.
- I. Fernández-Barahona, M. Muñoz-Hernando and F. Herranz, Mol., 2019, 24(7), 1224 CrossRef PubMed.
- S. Deguchi and B. Lindman, Polym., 1999, 40(25), 7163–7165 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2025 |





