Predicting inflammatory response of biomimetic nanofibre scaffolds for tissue regeneration using machine learning and graph theory†
Lakshmi Yaneesha
Sujeeun
ab,
Itisha Chummun
Phul
 a,
Nowsheen
Goonoo
a,
Nowsheen
Goonoo
 a,
Nicholas A.
Kotov
b and
Archana
Bhaw-Luximon
a,
Nicholas A.
Kotov
b and
Archana
Bhaw-Luximon
 *a
*a
aBiomaterials Drug Delivery and Nanotechnology Unit, Centre for Biomedical and Biomaterials Research (CBBR), University of Mauritius, Réduit, Mauritius. E-mail: a.luximon@uom.ac.mu
bCenter for Complex Particle Systems (COMPASS), University of Michigan, Ann Arbor, Michigan 48109, USA
First published on 22nd January 2025
Abstract
Tissue regeneration after a wound occurs through three main overlapping and interrelated stages namely inflammatory, proliferative, and remodelling phases, respectively. The inflammatory phase is key for successful tissue reconstruction and triggers the proliferative phase. The macrophages in the non-healing wounds remain in the inflammatory loop, but their phenotypes can be changed via interactions with nanofibre-based scaffolds mimicking the organisation of the native structural support of healthy tissues. However, the organisation of extracellular matrix (ECM) is highly complex, combining order and disorder, which makes it difficult to replicate. The possibility of predicting the desirable biomimetic geometry and chemistry of these nanofibre scaffolds would streamline the scaffold design process. Fifteen families of nanofibre scaffolds, electrospun from combinations of polyesters (polylactide, polyhydroxybutyrate), polysaccharides (polysucrose, carrageenan, cellulose), and polyester ether (polydioxanone) were investigated and analysed using machine learning (ML). The Random Forest model had the best performance (92.8%) in predicting inflammatory responses of macrophages on the nanoscaffolds using tumour necrosis factor-alpha as the output. CellProfiler proved to be an effective tool to process scanning electron microscopy (SEM) images of the macrophages on the scaffolds, successfully extracting various features and measurements related to cell phenotypes M0, M1, and M2. Deep learning modelling indicated that convolutional neural network models have the potential to be applied to SEM images to classify macrophage cells according to their phenotypes. The complex organisation of the nanofibre scaffolds can be analysed using graph theory (GT), revealing the underlying connectivity patterns of the nanofibres. Analysis of GT descriptors showed that the electrospun membranes closely mimic the connectivity patterns of the ECM. We conclude that ML-facilitated, GT-quantified engineering of cellular scaffolds has the potential to predict cell interactions, streamlining the pipeline for tissue engineering.
1. Introduction
Polymer-based nanostructured scaffolds have emerged as an effective strategy for tissue regeneration, serving the dual role of structural supports and molecular platforms. These scaffolds are engineered to closely replicate the tissue microenvironment, thus facilitating and enhancing the wound healing process. Effective wound care scaffolds must support the three primary stages of tissue regeneration and wound healing: inflammation, cell proliferation, and tissue remodelling. These stages involve several interconnected phases that rely on the synergistic interactions between cells and the extracellular matrix (ECM).1 In acute wounds, the inflammatory phase typically lasts from hours to days, whereas in chronic wounds, this phase can extend from weeks to months. During the inflammatory phase, macrophages play a crucial role in clearing pathogens and foreign materials. This phase is accompanied by the migration of neutrophils and monocytes to the wound site, followed by cytokine production. Monocytes differentiate into macrophages, which act as primary responders during inflammation. These macrophages phagocytize the remaining neutrophils, secrete growth factors, and produce cytokines that promote tissue regeneration and cell migration, allowing the wound to transition to the proliferative phase.2There are three primary macrophage phenotypes: M0, M1, and M2. The M0 phenotype represents the tissue-resident inactivated state, while M1 is pro-inflammatory, phagocytic, and bactericidal. M2 is anti-inflammatory (or pro-reparative), immunosuppressive, and involved in scar resolution.3,4 For effective wound healing, it is crucial for the predominant macrophage phenotype to transition from pro-inflammatory (M1) to pro-reparative (M2). A deeper understanding and prediction of the specific roles of macrophage populations at different stages of tissue repair will support the development of targeted therapies for both acute and chronic wounds.3 The M1 phenotype produces cytokines such as tumour necrosis factor-alpha (TNF-α) and interleukin-6 (IL-6), whereas the M2 phenotype promotes the production of cytokines like vascular endothelial growth factor (VEGF) and transforming growth factor beta (TGF-β). During the inflammatory phase, failure of macrophages to transition to a tissue-healing phenotype can result in persistent inflammatory signals, creating a positive feedback loop that amplifies the M1 response and disrupts cytokine balance. Additionally, the M2 phenotype has been identified as a key modulator in cancer progression due to its tumour-promoting capabilities, including immunosuppression, angiogenesis, neovascularization, and stromal activation and remodelling.5
Successful scaffold tissue integration relies on a balance between activated (M1) macrophages, which clear the wound site, and anti-inflammatory (M2) macrophages, which promote tissue regeneration and wound healing.6 Therefore, scaffold materials and architecture, designed to mimic in vivo structural support with organisation characteristic of the healthy tissues, should favour the transition of macrophages to the M2 tissue-healing phenotype. However, the organisation of native structural support, such as the ECM, is very complex. It is based on highly interconnected network of nanofibres with large amount of disorder, which is difficult to describe and replicate. A promising pathway to their replication that has emerged over the last few years, is to learn and predict their organisation using machine learning (ML), which will enable the selection of appropriate scaffold materials and reduce the trial-and-error process in electrospinning, minimising the need for recurrent in vitro studies. ML methodology can also be combined with new approaches to the description of complex nanoscale matter, which can complement ML tools, making them more efficient and transparent.7,8
Classifying macrophages based on their functional phenotype is crucial for predicting their behaviour as either pro- or anti-inflammatory agents in the immune response. Macrophages are typically classified into M0, M1, or M2 subsets by quantifying various cell surface markers, transcription factors, and cytokine profiling. These methods are time-consuming and resource-intensive. Concurrently, different forms of ML have found expansive applications in biology, ranging from ribonucleic acid (RNA) profiling studies that identify over 50 phenotypes9 to the basic classification of two distinct cell types within a population.10 ML and, especially, deep learning (DL) algorithms are particularly well-suited for analysing data related to scaffolds and cellular interactions due to their efficiency in decoding the complexity of cell responses to hierarchical organisation of nanofibres in the matrix.11,12 Initial studies have highlighted the potential of ML in predicting cell–material interactions on scaffolds during the proliferative phase of wound healing,13 as well as in classifying the miscibility of polymer blends based on their physico-chemical attributes.14
In the current study, the focus is on applying ML methods to model the inflammatory responses of macrophage cells cultured on nanofibre scaffolds with various polymer functionalities, thereby optimising and predicting materials performance (Scheme 1). The study is divided into two main parts. The first part aims to predict the concentration of the pro-inflammatory cytokine TNF-α produced by macrophages (target outcome), using the physico-chemical properties of nanofibre scaffolds (pore diameter, fibre diameter, water contact angle, and Young's modulus) and the biological responses of macrophages (ruffling index and macrophage phenotype) as input data. An ML-based predictive model was developed using seven supervised learning regression algorithms. Eighty percent of the dataset was used for training, while 20% was used for testing. The efficacy of each algorithm was assessed using performance metrics to determine the most accurate model. The second part of this study attempted to classify macrophages according to their phenotype using scanning electron microscopy (SEM) images of macrophages on nanofibres. Two common techniques for image processing and classification were adopted. The first involved the use of CellProfiler to extract cell features from SEM images, which were then fed to classification algorithms in CellProfiler Analyst. The second technique used DL convolutional neural network (CNN) pre-trained models, which were fine-tuned for our specific cell-image classification task. The last section of this study highlights the emerging capabilities of graph theory (GT) for the description of complex biological materials, capturing intricate structural patterns found in native nanofibre-based tissues.15,16 A preliminary mapping of the scaffold architecture was conducted to explore correlations between nanofibre arrangement and mechanical properties.
2. Materials and methods
2.1. Polymeric blends
The present study involved 15 distinct families of 53 nanofibre scaffolds: polyhydroxybutyrate/kappa-carrageenan (PHB/KCG), poly(hydroxybutyrate-co-valerate) (PHBV)/KCG, polydioxanone/fucoidan (PDX/FUC), PDX/KCG, PDX/PHBV, PDX/polysucrose (PSuc), poly-L-lactide (PLLA)/PSuc, PDX/bagasse-cellulose, PLLA/bagasse-cellulose, PLLA/ulvan-cellulose, PLLA/bagasse-cellulose acetate (bagasse-CA), PLLA/ulvan-cellulose acetate (ulvan-CA), poly(D,L-lactic acid) (PDLLA)/ulvan-cellulose, PDLLA/ulvan-CA, and PDX/ulvan-CA. Each family of scaffolds included a minimum of four polymer blend compositions, with results presented in triplicate. The compositions of polymer content varied as follows: 100% polymer A and 0% polymer B; 0% polymer A and 100% polymer B; 90% polymer A and 10% polymer B; 80% polymer A and 20% polymer B; 70% polymer A and 30% polymer B; 60% polymer A and 40% polymer B; and an even blend of 50% polymer A and 50% polymer B.PHB (Sigma-Aldrich), KCG (Sigma-Aldrich), PHBV (12 mol% HV content, Sigma-Aldrich), PDX (ResomerX 206 S, inherent viscosity (IV) 2.0 g dl−1, Evonik), FUC (Fucoidan from Fucus vesiculosus ≥95%, Sigma-Aldrich), PSuc, PLLA (PURASORB PL 18, (IV) 1.8 g dl−1, Purac), and PDLLA (PURASORB PDL 20, (IV) 2.0 g dl−1, Netherlands) were used as purchased. Bagasse-cellulose was extracted from locally available sugarcane bagasse using a combination of mercerisation and bleaching techniques,17 yielding an average of 40% (±2). Bagasse-CA was synthesised from sugarcane bagasse-derived cellulose using an optimised acetylation method,18 with an average yield of 62% (±2).19 Ulvan-cellulose was extracted from locally available green seaweeds of the Ulva family, producing an average yield of 5.23% (±0.2) using a modified method.20 Ulvan-cellulose was then converted to ulvan-CA using an optimised method,18 achieving an average yield of 78.5% (±0.8).21
2.2. Nanofibre scaffold fabrication
Scaffolds were engineered using the electrospinning method (bottom-up NE300 laboratory scale electrospinner, Inovenso Company, Turkey). The electrospinning conditions were adjusted based on the polymers within they blend and on the blend composition to generate matrices of bead-free fibres with complex structures and high interconnectedness. PHB/KCG and PHBV/KCG fibres were produced as reported by Goonoo et al.22 Electrospinning parameters for PSuc-based and bagasse-cellulose-based fibres were reported by Chummun et al.23 and Ramphul et al.,19 respectively. The fabrication of PDX/KCG and PDX/FUC was detailed by Goonoo et al.24 The fabrication of scaffolds from ulvan-cellulose and ulvan-CA in combination with either PDX, PLLA, or PDLLA was described by Madub et al.21 Most blend solutions were prepared by mixing two solutions (solution A and solution B) with the exception of PDX/PHBV, PDX/PSuc, and PLLA/PSuc.23,252.3. Scaffold characterisation
2.4. Biological compatibility assessment
2.5. Computational studies
For each new cell type or assay, the software employs a pipeline composed of discrete modules, with each module uniquely processing the image according to a defined procedure. The pipeline consists of the following steps in sequential order: (i) image processing, (ii) object identification, and (iii) object measurements. Most of the modules are automated but CellProfiler also supports interactive modules. For example, a user can select or outline manually an area of interest in each image. To start an analysis, each macrophage SEM image was loaded into CellProfiler and processed by each module in order (Fig. 1).
2.6. Deep learning (DL) models for macrophage cell classification
Transfer Learning (TL) is a common approach for applying pre-trained DL models to small image datasets. Pre-trained models are networks that have already been trained on a large set of image data, typically on a large-scale image classification task. CNNs are a specific type of artificial neural networks (ANNs) inspired by the visual cortex of a human brain, where each individual neuron detects only signals from a small sub-region of the visual field, called a receptive field. Each ‘neuron’ in CNN performs a convolution of a kernel with an input image and produces a filtered output image often called feature map. The input image can consist of several channels, and each layer in the neural network holds as many channels of feature maps as we have neurons in this particular layer. The feature maps in the last layer can be interpreted as the final features learned by the network and are used for classification. The critical difference from traditional feature-based classification methods is that for CNN, no features, including the weights of the kernels, are predefined, but the algorithm learns them by itself.28We attempted to implement two pre-trained models: VGG-1629 and ResNet50,30 both available in the Keras Applications library. Both models are among the commonly used CNN models pre-trained on ImageNet for TL31 and have been reported to perform exceptionally well.32 VGG16 has 13 convolutional layers and three fully connected layers, for a total of 16 layers, whereas ResNet50 has a deeper network structure with a total of 50 layers. The aim was to apply transfer learning approaches to use these pre-trained models and train them on our custom macrophage images. Two macrophage phenotypes M1 and M0 were selected for modelling purposes as the majority of the dataset were images with either M1 or M0 phenotype, or a mixture of both M1 and M0. Since the purpose of this part was to only explore these models, a small dataset was manually prepared and customized: 200 macrophage images with M0 phenotype and 200 macrophage images with M1 phenotype. After the data preparation step, two pre-trained models VGG16 and ResNet50 were applied on the dataset. ImageNet dataset contains RGB (Red Green Blue) colour images (three channels) and SEM images are grayscale images (single channel). Thus, for this preliminary study, one approach was to convert the grayscale images to RGB images, i.e. to make the images “appear” to be RGB by repeating the image array three time on a new dimension in numpy. Data augmentation was used to generate additional images in the training set by rotating, mirroring and flipping the images using the ImageDataGenerator class in Keras. For VGG16, the training and validation sets were configured in batches of 20. As this study is a binary classification problem, the basic model was loaded, with changes made only to the final layer and all the other layers were set as “non-trainable” (i.e. frozen). The last fully-connected layer was created using basic settings, and the final model was built and fitted based on the training and validation sets created above, with 10 epochs. Similarly, for ResNet50, the base model was imported with the layers frozen and last layer modifiable. The model was built, compiled and fitted with 10 epochs.
2.7. Graph theory (GT) analysis of nanofibre networks
A preliminary analysis was conducted to explore the relationship between the complex organisation of three families of nanofibre scaffolds – PDX/KCG, PLLA/PSuc, and PHB/KCG – and the structural characteristics captured by GT descriptors. The goal was to establish an understanding of how variations in materials might influence the complex organisation of biomimetic scaffolds potentially affecting their biological performance. StructuralGT, a Python program for automated structural analysis was used to perform GT analysis on the SEM images. Details of the calculation of each GT parameter is described in the StructuralGT publication and on its Github page: https://github.com/drewvecchio/StructuralGT.16 Three SEM images were analysed for each polymer blend to provide exploratory insights. The SEM images were not enhanced or modified using additional image processing software. In the resulting graphs, dangling edges were removed to achieve a more accurate measure of connectivity.3. Results and discussion
3.1. Macrophage inflammatory responses
This study aimed to establish a relationship between the physico-chemical properties of nanofibre scaffolds and biological responses of macrophages, specifically the ruffling index and macrophage polarisation phenotype (Scheme 2). A wound that fails to transition effectively from the pro-inflammatory to the anti-inflammatory phase is at risk of becoming chronic. Therefore, understanding macrophage behaviour in response to scaffold material is essential for tissue engineering (TE), as it guides the selection of suitable scaffold materials that promote desirable healing outcomes.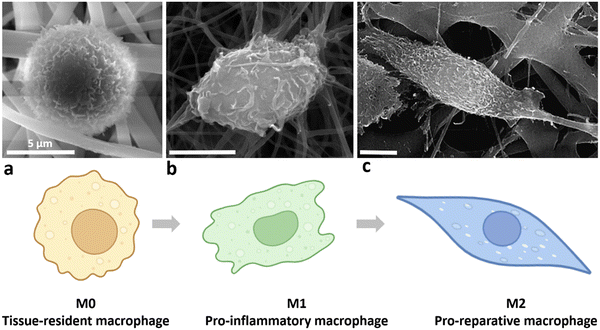 | ||
| Scheme 2 (a) M0, (b) M1, and (c) M2 phenotypes (SEM images from CBBR) and their representations (designed with https://BioRender.com). | ||
Fifteen families of nanofibre scaffolds were constructed using various blends of polymers (Table 1 and Table S1, ESI†). The physico-chemical parameters namely pore diameter, fibre diameter, water contact angle, and Young's modulus were determined for 53 scaffolds. Macrophages (RAW 264.7) were seeded on the scaffolds (Table 1). TNF-α levels were measured after three days to gauge the extent of inflammation induced by macrophages in response to the scaffolds, while SEM imaging provided a detailed assessment of macrophage phenotypes and ruffling index. In PDX/PHBV mats, a decreased in the concentration of TNF-α was observed with PHBV content above 20 wt%. The addition of KCG to PHBV led to an increase in TNF-α levels, whereas in the PHB/KCG blends, the addition of KCG slightly decreased TNF-α production. PDX/PHBV mats caused lower inflammatory reactions in RAW 264.7 cells than the PHB/KCG and PHBV/KCG mats. The addition of PHBV to PDX also promoted macrophage polarisation to the pro-healing phenotype when PHBV formed the shell of the fibres.25
| Scaffolds | Concentration of TNF-α/(pg ml−1) | Ruffling index | Main phenotypes observed |
|---|---|---|---|
| Polyhydroxybutyrate (PHB)/kappa-carrageenan (KCG) | |||
| 100/0 | 803.9 ± 56.4 | 2.8 ± 0.1 | M1 |
| 90/10 | 1123.7 ± 36.9 | 1.9 ± 0.3 | M1 |
| 80/20 | 947.0 ± 141.7 | 2.1 ± 0.2 | M1 |
| 70/30 | 631.2 ± 151.2 | 2.6 ± 0.2 | M1 |
| Poly(hydroxybutyrate-co-valerate (PHBV))/KCG | |||
| 100/0 | 784.6 ± 58.8 | 2.6 ± 0.1 | M1 |
| 90/10 | 1345.5 ± 63.6 | 0.2 ± 0.1 | M1 |
| 80/20 | 1293.0 ± 67.8 | 0.2 ± 0.1 | M1 |
| 70/30 | 865.6 ± 35.3 | 0.2 ± 0.1 | M0 |
| Polydioxanone (PDX)/fucoidan (FUC) | |||
| 100/0 | 261.1 ± 57.9 | 2.9 ± 0.1 | M1 |
| 90/10 | 854.5 ± 47.9 | 1.7 ± 0.1 | M1 |
| 80/20 | 644.8 ± 67.5 | 0.9 ± 0.1 | M1 |
| 70/30 | 1079.6 ± 2.1 | 1.2 ± 0.2 | M1 |
| PDX/KCG | |||
| 100/0 | 537.0 ± 125.2 | 2.5 ± 0.1 | M1 |
| 90/10 | 376.7 ± 76.6 | 3.0 ± 0.0 | M1 |
| 80/20 | 258.9 ± 49.9 | 1.2 ± 0.5 | M0 |
| 70/30 | 504.5 ± 62.1 | 0.3 ± 0.2 | M0 |
| PDX/PHBV | |||
| 100/0 | 196.5 ± 16.7 | 1.6 ± 0.3 | M1 |
| 90/10 | 722.3 ± 37.3 | 1.6 ± 0.7 | M1 |
| 80/20 | 1210.5 ± 61.8 | 0.2 ± 0.1 | M1 |
| 70/30 | 675.2 ± 63.8 | 2.4 ± 0.2 | M0 |
| PDX/polysucrose (PSuc) | |||
| 100/0 | 219.4 ± 49.7 | 2.9 ± 0.1 | M0 |
| 90/10 | 296.1 ± 90.0 | 3.0 ± 0.1 | M1 |
| 80/20 | 230.1 ± 88.1 | 2.8 ± 0.1 | M1 |
| 70/30 | 202.6 ± 41.6 | 2.9 ± 0.0 | M1 |
| 60/40 | 151.2 ± 11.8 | 3.0 ± 0.0 | M1 |
| 50/50 | 226.9 ± 57.9 | 2.9 ± 0.0 | M1 |
| Poly-L-lactide(PLLA)/PSuc | |||
| 100/0 | 163.7 ± 29.9 | 2.0 ± 0.0 | M2 |
| 90/10 | 149.0 ± 24.8 | 2.8 ± 0.0 | M2 |
| 80/20 | 168.7 ± 36.3 | 2.4 ± 0.1 | M1 |
| 70/30 | 173.1 ± 15.9 | 2.4 ± 0.3 | M1 |
| 60/40 | 245.6 ± 55.6 | 2.4 ± 0.7 | M1 |
| 50/50 | 240.3 ± 61.0 | 1.3 ± 0.1 | M1 |
| PLLA/cellulose acetate (CA) | |||
| 0/100 | 180.9 ± 26.9 | 0.4 ± 0.0 | M1 |
| 100/0 | 163.7 ± 29.9 | 2.0 ± 0.0 | M1 |
| 30/70 | 140.7 ± 12.8 | 1.0 ± 0.0 | M1 |
| 50/50 | 170.7 ± 3.3 | 1.8 ± 0.0 | M1 |
| PLLA/cellulose | |||
| 0/100 | 280.3 ± 7.0 | 2.8 ± 0.0 | M1 |
| 30/70 | 330.7 ± 27.2 | 2.9 ± 0.0 | M1 |
| 50/50 | 338.0 ± 102.0 | 2.9 ± 0.0 | M1 |
| PDX/CA | |||
| 100/0 | 239.4 ± 61.9 | 2.6 ± 0.0 | M1 |
| 30/70 | 300.0 ± 28.1 | 2.3 ± 0.0 | M1 |
| 50/50 | 298.9 ± 15.8 | 2.8 ± 0.0 | M1 |
| Poly(D,L-lactic acid) PDLLA/CA | |||
| 70/30 | 197.4 ± 9.9 | 1.0 ± 0.0 | M1 |
| 50/50 | 270.7 ± 30.2 | 1.0 ± 0.0 | M0 |
| PDLLA/cellulose | |||
| 0/100 | 280.3 ± 7.0 | 2.8 ± 0.0 | M1 |
| 30/70 | 350.9 ± 104.7 | 3.0 ± 0.0 | M1 |
| 50/50 | 346.8 ± 61.9 | 2.9 ± 0.0 | M1 |
In the presence of PSuc, a large number of round-shaped cells – M0 phenotype – dispersed among elongated spindle-like shaped macrophages (M2-like phenotype) visible on PLLA-PSuc mats. As for PDX 100, the cells formed flattened colonies compared to PDX-PSuc 70-30, onto which round-shaped cells were widely dispersed. Spindle-like macrophages on PLLA-PSuc mats are associated to M2 phenotype, indicative of induced in vitro biocompatibility of PLLA-PSuc blends. For PDX-PSuc electrospun mats, significant increase in TNF-α concentration was observed in the presence of PSuc.33 For bagasse-cellulose mats, as the amount of cellulose in the mats increased, the amount of adherent cells decreased, thus indicating its positive effect on reducing inflammatory response. The density of macrophages on the surface of PLLA 100% was also more prominent than on the blends. Bagasse-cellulose 100% mats displayed the lowest density of macrophage cells, thus indicating its positive effect towards inhibition of inflammation response due to its highly hydrophilic nature.34
On all the ulvan-cellulose and PDX-based scaffolds, large population of round-shaped macrophages exhibiting higher spreading and surface membrane activities were observed. While macrophages on 100% CA, ulvan-CA/PLLA, and PDLLA scaffolds displayed rather smooth surfaces, the presence of significant cell surface protrusions, i.e. surface ruffles, were noted on all ulvan-cellulose and PDX-based nanofibrous mats. Addition of PDX to CA and ulvan-cellulose to PLLA and PDLLA greatly enhanced the cells’ ability to undergo F actin-enriched membrane protrusions on their surfaces. Ulvan-cellulose and PDX-based scaffolds triggered higher release of TNF-α compared to the ulvan-CA/PLLA and PDLLA nanofibrous mats. Overall, these findings indicated that macrophage activity and thus, level of TNF-α are directly influenced by the surface chemistry of the biomaterial. Surface properties, hydrophilicity, and functional groups influence cell mechano-sensing and determine cell–material interactions.
3.2. ML Models to correlate physico-chemical properties with biological response
Data from the physico-chemical characterization of scaffolds and the biological parameters of macrophages were selected to develop a ML model to correlate in vitro inflammatory responses data with the properties of scaffolds. Data was collected, rearranged and pre-processed for exploratory analysis and feature selection. A high correlation filter calculated the correlation between scaled, independent numerical variables. The Pearson correlation matrix shown in Fig. 2(a) displayed correlation coefficients between −0.21 to 0.45, indicating no strong correlation among variables.Feature selection using the RF regressor identified the most predictive features based on their importance scores (Fig. 2(b)). Ruffling index ranked as the top feature (0.59), followed by pore diameter (0.13) and fibre diameter (0.09). Less influential features included Young's modulus (0.08), macrophage phenotype (0.06), and water contact angle (0.04). These findings were consistent with our previous results, where ML methods were applied to predict fibroblasts proliferation on nanofibre mats, with fibre diameter and pore size being the most influential properties.13 Since cellular responses are cell-specific and cannot be generalised, it was essential to re-investigate the scaffold properties influencing macrophage polarisation in this study. Ruffles are temporarily erected in response to stimuli and during cell migration and macrophages ruffling is enhanced in cells activated by pathogens.35 The key predictors identified in this study can help minimise trial and error in the development of nanofibre scaffolds. Parameters such as fibre diameter and pore size in electrospun scaffolds can be determined and utilised to predict the level of TNF-α expressed by macrophages. As a result, electrospinning parameters can be adjusted to fabricate scaffolds with optimised physico-chemical properties that elicit a lower immune response, thereby reducing the need of repeated in vitro experiments.
Seven supervised learning regression algorithms were trained with 80% of the data, and the remaining 20% was used for testing. After hyperparameter tuning, model performance of each model was evaluated using regression metrics (accuracy scores on the training and testing sets, MAPE, MAE, RMSE) (Fig. 2(c) and Table S2, ESI†). The RF regression model achieved an accuracy of 92.8% on the training set and 89.3% on the testing, coupled with a MAPE score of 22.9%, an MAE of 97.12, and an RMSE of 149.89, demonstrating strong predictive performance and robustness. This highlighted the RF regression model's accuracy and suitability for capturing the complexities of both physico-chemical and biological data, as well as our predictive targets. In our previous studies, the RF regressor and classifier consistently outperformed other algorithms.13,14 This suggested that RF models are particularly adept at managing non-linear relationships between variables, which is crucial in this study, where the correlation between TNF-α levels and various predictors was not robust. Actual data from the testing set was compared against predicted data from the testing set of the RF model through a regression plot. The analysis indicated a statistically significant fit (p-value < 4.27 × 10−5) that could potentially be improved with more representative samples in the testing set and additional data points closer to the regression line. The high proportion of the variance explained by the model (R2 = 0.89) suggested that the RF regression model closely fitted the actual dataset (Fig. 2(d)).
3.3. Identification of macrophage phenotypes from SEM images using CellProfiler
The pipeline for human HT29 cells available in CellProfiler was selected for preliminary macrophage SEM image analysis. This pipeline integrates modules to identify cells and measure cellular parameters such as morphology, count, intensity, and texture. The first step of image processing in CellProfiler included cropping the input image to select the area of interest (Fig. 3, steps 1 and 2), followed by applying illumination correction and filtering methods, as raw images degrade intensity measurements (Fig. 3, steps 3 and 4). This degradation may generate inaccurate cell identification/segmentation and adversely affect all types of measurements, from intensity to area and shape measurements. Object identification/segmentation is the most challenging step in image analysis, and its accuracy determines the reliability of the resulting cell measurements (Fig. 3, step 5). Similar to most biological images, SEM images of macrophages included cells in close proximity, often touching each other. In CellProfiler, clumped cells were detected as single objects, which were then separated by identifying dividing lines between them. Some resulting objects were subsequently merged together or discarded from the analysis. After primary object detection of the cells, CellProfiler computed feature measurements for each identified cell. These included standard features such as area or intensity, and also complex measurements like Zernike shape features, Haralick and Gabor texture features (Fig. 3, step 6). The data were exported in a tab-delimited spreadsheet format for further analysis (Fig. 3, step 7) (Table S3, ESI†). | ||
| Fig. 3 Image processing steps with CellProfiler to extract features related to macrophage phenotypes. | ||
Image-based profiling is a powerful quantitative method to measure cellular and sub-cellular features. Single-cell measurements are features that enrich biological dataset and increase robustness of statistical modelling. In this study, objects representing cells were detected by manually adjusting parameters of different modules for each image (Fig. 3). A full dataset of 41 SEM images (view field 10–20 μm) were analysed. Each cell was then assessed for a broad range of descriptors such as area, orientation, extent, shape, intensity, etc. A total of 225 measurements were acquired for each cell, and a dataset representing the full experiment/analysis was established for further phenotype classification analysis (Table S3, ESI†).
Detection of cellular features in images from automated software tools still requires optimisation due to the variability between cell phenotypes of different cell lines. The advantage of using CellProfiler remains its adaptability: a user can customise an image analysis pipeline from existing available pipelines or create a new one, tune segmentation parameters to perform well and detect cell phenotypes. The major hindrance would be analysing large datasets containing numerous phenotypes within a single image, which can complicate accurate segmentation and classification. For instance, it is extremely time-consuming to verify all the segmentation parameters while processing each image to ensure that it reliably segments images of all M0 and M1 phenotypes present in the dataset. CellProfiler enables batch processing of images once a pipeline has been established. However, due to particular cell morphology in our study, this option has not been considered. To be able to correct some automated object identification errors, manual object editing modules were used to select the objects and modify them accordingly. The main challenge in cell segmentation is to devise reliable features that will be able to identify cell boundaries with a high accuracy.
Our findings indicated that among the 15 different families of scaffolds, the M0 and M1 phenotypes were the most frequently observed using SEM image visual analysis (Table 2). Overall good agreement between CellProfiler image analysis and SEM image visual analysis was found. A more advanced analysis model would probably be required to differentiate and quantify between different phenotypes.
| Nanoscaffolds | Main phenotypes observed by SEM | Main phenotypes identified by CellProfiler image analysis | Main phenotypes identified by CNN |
|---|---|---|---|
| a Reported SEM images from literature were used to test the trained models.36,37 | |||
| M0 | — | M0 | M0 |
| M1 | — | M1 | M1 |
| M2 | — | M1 and M2 | NA |
| PHB/KCG | M1 | M0 and M1 | M0 and M1 |
| PHBV/KCG | M0 and M1 | M0 and M1 | M0 and M1 |
| PDX/FUC | M1 | M0 and M1 | M0 and M1 |
| PDX/KCG | M0 and M1 | M0 and M1 | M0 and M1 |
| PDX/PHBV | M0 and M1 | M0 and M1 | M0 and M1 |
| PDX/PSuc | M0 and M1 | M0 and M1 | M0 and M1 |
| PLLA/PSuc | M1 and M2 | M0 and M1 | M0 and M1 |
| PDX/bagasse-CA | M1 | M0 and M1 | M0 and M1 |
| PLLA/bagasse-cellulose | M1 | M0 and M1 | M1 |
| PLLA/ulvan-cellulose | M1 | M0 and M1 | M0 and M1 |
| PLLA/bagasse-CA | M1 | M0 and M1 | M0 and M1 |
| PLLA/ulvan-CA | M1 | M0 and M1 | M0 and M1 |
| PDLLA/ulvan-cellulose | M1 | M0 and M1 | M0 and M1 |
| PDLLA/ulvan-CA | M0 and M1 | M0 and M1 | M0 and M1 |
| PDX/ulvan-CA | M1 | M0 and M1 | M1 |
Thus processing SEM images and classification of macrophages can be performed with CellProfiler as an effective tool. However, to carry out statistically relevant image analysis by SEM, it is required to collect very large datasets. Processing such large image data is not possible in a time-efficient manner, thus the need to use DL models to address these limitations and facilitate the process.
3.4. Evaluation of CNN models for macrophage classification
Training a CNN model can be very complex and time-consuming, but the advantage of using pre-trained CNN models for image classification speeds up the training time. In general, the pre-training is performed with general images that come from outside the direct classification task domain. CNNs can learn appropriate features directly from the image data without the need for a predefined feature extraction process. Traditional methods would require cell features extracted from the images beforehand, such as in CellProfiler, whereas CNNs use the raw image data as input, providing better performance and more flexibility compared to traditional methods. Our aim was to assess the potential of applying CNN approaches to classify images of individual cells into two main phenotypes, M0 and M1. For the CNN models to learn from SEM images of cell phenotypes, the cells must be present in the image at an appropriate size so that their characteristic morphologies can be detected as objects. In our case, SEM images with 20 μm magnification were selected. An epoch is considered as a hyperparameter that defines the number of times that the learning algorithm will work through the entire training dataset. With 10 epochs, the VGG16 and ResNet50 models generated validation accuracies of 90.3% and 91.4% respectively, without any major changes to the models (Fig. 4 and Table 2). This indicated the ability of the CNN models to classify phenotypes of macrophage cells on the scaffolds independently of any other physico-chemical parameters. | ||
| Fig. 4 Graph of the accuracy at each epoch for both training and validation datasets for (a) VGG16 and (b) ResNet50. | ||
3.5. Preliminary correlation of complex nanofibrous architecture with scaffold materials and physico-chemical properties using graph theory
The SEM images of the scaffolds used in this study showed a complex arrangement of the nanofibres giving rise to a biomimetic organisation of the scaffolds, replicating the ECM (Fig. 5(a)–(c)). Can the geometrical patterns of such complex systems be correlated with scaffold materials and predicted based on material compositions? This fundamental hypothesis was investigated using GT, known for its application to complex systems analysis.16,38 In this case, GT was applied to explore potential relationships between fibre network structures and the material composition of the PDX/KCG, PLLA/PSuc, and PHB/KCG families of nanofibre scaffolds. These polymer blends were selected based on the origin and chemical characteristics of the polymers. PDX is a semi-crystalline synthetic polyester ether with a low glass transition temperature (Tg). KCG is a natural polysaccharide with negative sulphate groups and a helical structure. PSuc is a synthetic polysaccharide, PHB is a natural polyester with high crystallinity, and PLLA is a synthetic polyester with high crystallinity. All these materials, when blended, are used in various medical applications.GT analysis of the polymer blend networks – PDX/KCG, PLLA/PSuc, and PHB/KCG – revealed distinct trends in the scaffold architecture. Scaffolds with higher node and edge densities corresponded to more interconnected fibre networks. Node connectivity measures how well-connected each node is within the network, and clustering coefficient measures the local interconnectedness of the network. Pure polymers exhibited higher average degree (i.e., more connections per node) and node connectivity compared to the blends (Fig. 5(d)). Overall, as the proportion of the second component in the blend increased, a decrease in network connectivity was observed, with lower average degree and node connectivity. This suggested a more porous network and potentially increased immiscibility between the polymers in the blend, regardless of the specific polymer types.
There was no clear trend between crystallinity and structural density across the different material combinations. However, within the PLLA/PSuc family where a full spectrum of data for blend composition was available, a clear trend emerged: node density increased as blend crystallinity decreased, with a reversal of this behaviour when the amorphous PSuc became the predominant component (Fig. 5(d) and Fig. S1, ESI†). The variations in network topology should correlate with the mechanical properties of the scaffolds as denser networks are associated with smaller fibre and pore diameters, which influenced the mechanical performance. Indeed, the mechanical characterisation showed that the Young's modulus for the 70/30 PDX/KCG was 38.2 ± 5.5 MPa, while for PHB/KCG it was 90.6 ± 10.9 MPa (Table S1, ESI†). In terms of hydrophilicity, PSuc has higher water solubility than KCG at room temperature. It is assumed that the blend of PSuc (hydrophilic) with PLLA (hydrophobic) has more immiscible characteristics; thus making it more hydrophobic compared to the KCG blends with PHB (hydrophobic) or PDX (hydrophobic). The water contact angles for the 70/30 PDX/KCG, PHB/KCG, and PLLA/PSuc were 32.1 ± 0.0, 104 ± 0.9, and 126.3 ± 4.7, respectively (Table S1, ESI†).
 | ||
| Fig. 6 (a) SEM image of human small intestine grafts network converted into graph and (b) summary of GT parameters. (Licensed under CC-BY).39 | ||
The biomimetic structural characteristics of polymer nanofibre scaffolds were compared with those of the human small intestine39 following the methodology of comparative connectivity assessment (Fig. 7) described in our previous study.15 The overall trend indicated that the scaffolds exhibited network characteristics within the range of the natural ECM. For instance, PLLA/PSuc scaffolds demonstrated similar average node connectivity and clustering coefficients compared to the human ECM, reinforcing their biomimetic potential. PDX/KCG scaffolds exhibited higher edge and node densities, suggesting denser network structures, while PHB/KCG scaffolds showed lower edge densities but more balanced clustering, similar to the ECM. Our findings clearly showed that the complex structure of nanofibrous scaffolds have key parameters within the same range, further confirming their potential as biomimetic scaffolds for TE applications. These similarities highlight the promise of these nanofibrous networks in replicating essential features of the ECM, which could have important implications for cell behaviour and tissue regeneration.
The similarity in GT metrics for SEM and AFM data suggested that the scaffold's structural characteristics remain consistent across imaging techniques and within layers of nanofibre being independent of the image acquisition technique. This consistency also indicated that the scaffold maintained a uniform structural complexity in the different layers of nanofibres, which is particularly advantageous for TE applications, where ECM-like architecture is critical for promoting cell adhesion, migration, and differentiation. Performing GT analysis of both SEM and AFM images not only confirmed the scaffold's ECM-like architecture but also validated the use of GT as an analytical method for material characterization.
4. Conclusions
The focus of this study was to develop computational models to predict inflammatory responses, i.e. TNF-α levels in macrophages, on nanostructured electrospun scaffold, based on physico-chemical properties of nanofibres and complex geometry of the scaffolds. Among seven ML algorithms tested, the RF model outperformed the rest. Ruffling index, pore diameter, and fibre diameter emerged as the most important parameters influencing the concentration of TNF-α. These preliminary findings provided insights into cellular behaviour in the context of the tissue repair process, contributing to the improvement of material performance with evidence-based data. The second part of this study successfully demonstrated that CellProfiler is an effective tool in processing SEM images to extract diverse features and measurements related to cell phenotypes. Results from DL modelling indicated that CNN models are adept at classifying macrophage cells from SEM image based on their phenotypes. GT showed that it is possible to predict the correlation between materials and complex nanofibre arrangement thus providing a method to move further ahead of the scaffold development pipeline for tissue engineering. This study paves the way towards ML-facilitated GT-quantified scaffold development, with the potential to extend from material choice to nanofibre arrangements to in vitro–material interactions and finally in vivo–material interactions, thereby reducing the timeline and cost for translating scaffolds into clinical applications.Author contributions
LYS: formal analysis, investigation (computational studies and mathematical modelling), methodology, writing – original draft & editing; ICP: investigation (experimental data), writing – original draft; NG: investigation (experimental data), writing – original draft; NAK: conceptualization of complex nanofibre systems, supervision of graph theory modelling and training, resources, NSF funding acquisition and fellowship organisation; ABL: conceptualization, resources, writing – original draft, review & editing, supervision, project administration, funding acquisition.Data availability
The data supporting this article have been included as part of the ESI.†Conflicts of interest
There are no conflicts to declare.Acknowledgements
LYS is thankful to the Higher Education Commission (HEC) for a PhD fellowship. ABL would like to thank RT Knits Ltd for funding to the Biomaterials, Drug Delivery and Nanotechnology Unit, CBBR under project vote KB027. The authors are grateful for the support from the National Science Foundation (NSF), grant # 2243104, Centre for Complex Particle Systems (COMPASS). LYS and ABL thank the COMPASS-CERA Fellowship Program and Professor Martin Thuo, Deputy Director of COMPASS and Mrs Valerie Wehiong-Batta for great assistance in the fellowship organisation. N. A. K. also acknowledges additional support from NSF, grant # 2317423; Lock-And-Key Interactions Between Chiral Nanoparticles And Proteins.References
- V. A. Solarte David, V. R. Güiza-Argüello, M. L. Arango-Rodríguez, C. L. Sossa and S. M. Becerra-Bayona, Front. Bioeng. Biotechnol., 2022, 10, 821852 CrossRef PubMed.
- Y. Yu, Z. Yue, M. Xu, M. Zhang, X. Shen, Z. Ma, J. Li and X. Xie, PeerJ, 2022, 10, e14053 CrossRef PubMed.
- K. S. Smigiel and W. C. Parks, Curr. Rheumatol. Rep., 2018, 20, 17 CrossRef PubMed.
- A. Kishore and M. Petrek, Front. Immunol., 2021, 12, 678457 CrossRef CAS PubMed.
- J. Liu, X. Geng, J. Hou and G. Wu, Cancer Cell Int., 2021, 21, 389 CrossRef CAS PubMed.
- J. Huang, J. Wei, X. Xia, S. Xiao, S. Jin, Q. Zou, Y. Zuo, Y. Li and J. Li, Mater. Today Bio, 2024, 26, 101063 CrossRef CAS PubMed.
- L. Yang, H. Wang, D. Leng, S. Fang, Y. Yang and Y. Du, Chem. Eng. J., 2024, 156687 CrossRef CAS.
- K. A. Brown, S. Brittman, N. Maccaferri, D. Jariwala and U. Celano, Nano Lett., 2020, 20, 2–10 Search PubMed.
- T. R. Jones, A. E. Carpenter, M. R. Lamprecht, J. Moffat, S. J. Silver, J. K. Grenier, A. B. Castoreno, U. S. Eggert, D. E. Root, P. Golland and D. M. Sabatini, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 1826–1831 Search PubMed.
- H. M. Rostam, P. M. Reynolds, M. R. Alexander, N. Gadegaard and A. M. Ghaemmaghami, Sci. Rep., 2017, 7, 3521 CrossRef PubMed.
- E. Ahmed, P. Mulay, C. Ramirez, G. Tirado-Mansilla, E. Cheong and A. J. Gormley, Tissue Eng., Part A, 2024, 30, 662–680 Search PubMed.
- S. M. McDonald, E. K. Augustine, Q. Lanners, C. Rudin, L. Catherine Brinson and M. L. Becker, Nat. Commun., 2023, 14, 4838 CrossRef CAS PubMed.
- L. Y. Sujeeun, N. Goonoo, H. Ramphul, I. Chummun, F. Gimié, S. Baichoo and A. Bhaw-Luximon, R. Soc. Open Sci., 2020, 7, 201293 CrossRef CAS PubMed.
- L. Y. Sujeeun, N. Goonoo, K. M. Moutou, S. Baichoo and A. Bhaw-Luximon, Macromol. Res., 2023, 7, 201293 Search PubMed.
- H. Zhang, D. Vecchio, A. Emre, S. Rahmani, C. Cheng, J. Zhu, A. C. Misra, J. Lahann and N. A. Kotov, MRS Bull., 2021, 46, 576–587 Search PubMed.
- D. A. Vecchio, S. H. Mahler, M. D. Hammig and N. A. Kotov, ACS Nano, 2021, 15, 12847–12859 Search PubMed.
- M. F. Rosa, E. S. Medeiros, J. A. Malmonge, K. S. Gregorski, D. F. Wood, L. H. C. Mattoso, G. Glenn, W. J. Orts and S. H. Imam, Carbohydr. Polym., 2010, 81, 83–92 Search PubMed.
- D. A. Cerqueira, G. R. Filho and C. da S. Meireles, Carbohydr. Polym., 2007, 69, 579–582 Search PubMed.
- H. Ramphul, A. Bhaw-Luximon and D. Jhurry, Carbohydr. Polym., 2017, 178, 238–250 Search PubMed.
- A. Mihranyan, A. P. Llagostera, R. Karmhag, M. Strømme and R. Ek, Int. J. Pharm., 2004, 269, 433–442 Search PubMed.
- K. Madub, N. Goonoo, F. Gimié, I. Ait Arsa, H. Schönherr and A. Bhaw-Luximon, Carbohydr. Polym., 2021, 251, 117025 CrossRef CAS PubMed.
- N. Goonoo, A. Bhaw-Luximon, P. Passanha, S. Esteves, H. Schönherr and D. Jhurry, Mater. Sci. Eng., C, 2017, 76, 13–24 CrossRef CAS PubMed.
- I. Chummun, A. Bhaw-Luximon and D. Jhurry, J. Biomed. Mater. Res. A, 2018, 106, 3275–3291 CrossRef CAS PubMed.
- N. Goonoo, A. Bhaw-Luximon, U. Jonas, D. Jhurry and H. Schönherr, ACS Biomater. Sci. Eng., 2017, 3, 3447–3458 CrossRef CAS PubMed.
- N. Goonoo, F. Gimié, I. Ait-Arsa, C. Cordonin, J. Andries, D. Jhurry and A. Bhaw-Luximon, Biomater. Sci., 2021, 9, 5259–5274 Search PubMed.
- N. Goonoo, A. Fahmi, U. Jonas, F. Gimié, I. A. Arsa, S. Bénard, H. Schönherr and A. Bhaw-Luximon, ACS Appl. Mater. Interfaces, 2019, 11, 5834–5850 CrossRef CAS PubMed.
- A. E. Carpenter, T. R. Jones, M. R. Lamprecht, C. Clarke, I. H. Kang, O. Friman, D. A. Guertin, J. H. Chang, R. A. Lindquist, J. Moffat, P. Golland and D. M. Sabatini, Genome Biol., 2006, 7, R100 Search PubMed.
- O. Dürr and B. Sick, J. Biomol. Screening, 2016, 21, 998–1003 CrossRef PubMed.
- K. Simonyan and A. Zisserman, arXiv, 2014, preprint, arXiv:1409.1556 DOI:10.48550/arXiv.1409.1556.
- K. He, X. Zhang and S. Ren, Deep residual learning for image recognition, in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016, pp. 770–778, https://openaccess.thecvf.com/content_cvpr_2016/html/He_Deep_Residual_Learning_CVPR_2016_paper.html.
- M. A. Morid, A. Borjali and G. Del Fiol, Comput. Biol. Med., 2021, 128, 104115 CrossRef PubMed.
- O. Russakovsky, J. Deng, H. Su, J. Krause, S. Satheesh, S. Ma, Z. Huang, A. Karpathy, A. Khosla, M. Bernstein, A. C. Berg and L. Fei-Fei, Int. J. Comput. Vis., 2015, 115, 211–252 Search PubMed.
- I. Chummun, F. Gimié, N. Goonoo, I. A. Arsa, C. Cordonin, D. Jhurry and A. Bhaw-Luximon, Mater. Sci. Eng., C, 2022, 135, 112694 Search PubMed.
- H. Ramphul, F. Gimié, J. Andries, D. Jhurry and A. Bhaw-Luximon, Int. J. Biol. Macromol., 2020, 157, 296–310 Search PubMed.
- N. D. Condon, J. L. Stow and A. A. Wall, Bio-Protoc., 2020, 10, e3494 CAS.
- F. Heinrich, A. Lehmbecker, B. B. Raddatz, K. Kegler, A. Tipold, V. M. Stein, A. Kalkuhl, U. Deschl, W. Baumgärtner, R. Ulrich and I. Spitzbarth, PLoS One, 2017, 12, e0183572 CrossRef PubMed.
- A. R. B. Ribeiro, E. C. O. Silva, P. M. C. Araújo, S. T. Souza, E. J. da, S. Fonseca and E. Barreto, Spectrochim. Acta, Part A, 2022, 265, 120328 CrossRef CAS PubMed.
- D. A. Vecchio, M. D. Hammig, X. Xiao, A. Saha, P. Bogdan and N. A. Kotov, Adv. Mater., 2022, 34, e2201313 CrossRef PubMed.
- A. C. Oliveira, I. Garzón, A. M. Ionescu, V. Carriel, J. de la, C. Cardona, M. González-Andrades, M. del, M. Pérez, M. Alaminos and A. Campos, PLoS One, 2013, 8, e66538 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4tb02494j |
| This journal is © The Royal Society of Chemistry 2025 |






