Accelerating the discovery of disordered multi-component solid-state electrolytes using machine learning interatomic potentials
Yanhao
Deng
 a,
Yan
Li
a,
Gopalakrishnan
Sai Gautam
a,
Yan
Li
a,
Gopalakrishnan
Sai Gautam
 b,
Bonan
Zhu
b,
Bonan
Zhu
 *cd and
Zeyu
Deng
*cd and
Zeyu
Deng
 *a
*a
aDepartment of Materials Science and Engineering, National University of Singapore, 9 Engineering Drive 1, 117575, Singapore. E-mail: msedz@nus.edu.sg
bDepartment of Materials Engineering, Indian Institute of Science, Bengaluru 560012, India
cSchool of Aerospace Engineering, Beijing Institute of Technology, Beijing, 100081, China. E-mail: bzhu@bit.edu.cn
dState Key Laboratory of Environment Characteristics and Effects for Near-space, Beijing, 100081, China
First published on 5th September 2025
Abstract
Machine learning interatomic potentials (MLIPs) are rapidly emerging as powerful tools for materials simulations, offering a promising pathway to explore complex systems beyond the reach of traditional methods. This study investigates the application of MLIPs focused on the MACE architecture, to multi-component, disordered solid-state electrolytes (SSEs), a critical class of materials for the next-generation solid-state batteries. We first benchmark the performance of MACE against established SSE families, Na1+xZr2SixP3−xO12 and Li3+xP1−xGexS4−4xO4x, confirming their general applicability while identifying key considerations for robust potential development in chemically diverse systems. This workflow, emphasizing the selection of representative configurations, provides critical insights for constructing reliable models in complex, multi-components environments. We further demonstrate the predictive power of this approach by constructing a high-performance MLIP for the novel halide system Li3InxY1−xBr6yCl6−6y (x, y ∈ [0, 1] and x + y ≥ 1), leading to the identification of the Li3In0.5Y0.5Br3Cl3 stoichiometry with the most favorable predicted ion transport properties. By analyzing the molecular dynamics (MD) trajectories generated in this work using our MLIP, we identified two distinct Li-ion migration pathways in this material. The trained model facilitates the computational investigation of intricate mixed cation/anion substitutions in halide SSEs, offering new insights into higher-entropy systems incorporating multi-components. Our results underscore the capability of MLIPs to accelerate the discovery cycle of complex functional materials and provide a robust computational framework for designing advanced SSEs.
Introduction
Combining quantum precision with classical efficiency, machine learning interatomic potentials (MLIPs) are emerging as transformative tools in computational materials science1,2 that enable simulations several orders of magnitude faster than ab initio methods.3,4 This dual advantage of enhanced speed and reliable accuracy enables the computational investigation of complex material systems and large-length, long-time scale molecular dynamics (MD) simulations, significantly broadening the horizons of computational materials research.5,6MLIPs have been developed based on approaches such as the Gaussian approximation potential (GAP),7 moment tensor potentials (MTP),8 and neural network models.9,10 Graph Neural Networks (GNNs)11 have become a widely used framework for modeling atomic systems and developing interatomic potentials. Notable examples include CGCNN,12 M3GNet,13 and CHGNet.14 By encoding structural symmetry information, invariant and particularly equivariant GNNs—such as NequIP15 and MACE16—have achieved enhanced efficiency without compromising accuracy.17,18 The MACE architecture,16 which extends the atomic cluster expansion (ACE) framework19 and employs high body-order equivariant features in each layer, enabling accurate modeling with just two layers of message passing, thus achieving low computational cost, on the order of nanoseconds per day on a GPU.
Meanwhile, training universal MLIPs (uMLIPs) using large, diverse datasets such as the Materials Project (MP),20 AFLOW,21 Open Quantum Materials Database (OQMD),22 Alexandria,23 Open Materials 2024 (OMAT24),24 and MatPES25 has generated significant interest, enabling direct deployment across materials encompassing >90% of application-relevant elements in the periodic table, including inorganic crystals, organic compounds, and theoretically predicted structures.5,13 As their reliability and generalizability improve, uMLIPs are rapidly becoming mainstream tools for materials modeling.26–28 Recent state-of-the-art uMLIPs, such as MACE-MP,29 eSEN,30 SevenNet,17 and BAMBOO,31 training on those diverse and combined datasets, enable transferability across a wide range of materials systems.
In this work, we systematically evaluate the performance of the MACE architecture by fine-tuning the MACE-MP (version: MACE-MP-0a).29 We apply this model to representative SSEs, which are crucial for advancing safer lithium- and sodium-based batteries by replacing conventional flammable liquid electrolytes with solid fast-ion conductors.32–34 While there have been increasing applications of MLIPs in modeling SSEs,4,35–41 achieving broad compositional coverage, low computational cost, high prediction accuracy, and fidelity remains challenging.42 To address this, fine-tuning a uMLIP offers a promising strategy, as it enables adaptation to new material systems with minimal additional data while leveraging knowledge from diverse chemistries. We implement this scheme by fine-tuning the MACE-MP model using a multi-head strategy, which requires only a small dataset to achieve stable results and helps prevent catastrophic forgetting that can occur during naive fine-tuning. Furthermore, an efficient training data selection algorithm is introduced to further enhance the effectiveness of this fine-tuning process.
As a benchmark system, we first evaluate the model on Na1+xZr2SixP3−xO12 (NaSICON), a class of SSEs first reported more than 40 years ago.43 Its extensive experimental and computational history makes it an ideal candidate to validate the MLIP performance and reconcile predictions with previous studies.43–49 Our investigation also includes materials from the oxysulfide SSE family. Specifically, we focus on the pseudo-binary system (1 − x)Li3PS4–xLi4GeO4 (LPSGO).50 This system aims to combine the high Li-ion conductivity of sulfides like Li3PS4 (LPS) with the electrochemical stability of oxides like Li4GeO4 (LGO), offering an excellent platform to assess the ability of MLIPs to capture complex hybrid SE chemistries.
Beyond these benchmarks, our investigation extends to lithium ternary halide systems (Li3MX6), with a focus on the mixed halide system represented by the ternary compound phase diagram Li3YBr6–Li3InBr6–Li3InCl6. Each compound serves as a vertex of a compositional triangle. This ternary system is described by the general chemical formula Li3InxY1−xBr6yCl6−6y, where x, y ∈ [0, 1] and x + y ≥ 1. Among these, Li3InBr6 exhibits high room-temperature conductivity (0.1–1 mS cm−1),51–53 while Li3YBr6 achieves similar performance (0.03–1.7 mS cm−1).54 Recently, Li3InCl6 has demonstrated excellent ionic conductivity (0.79–4.03 mS cm−1) and excellent chemical and electrochemical stability, benefiting from reduced grain boundary effects.55–59 Although simple mixed-halide SSEs (e.g., Li3InBrxCl6−x) are well-studied, investigating complex cases involving simultaneous double elemental substitutions (where x ≠ 0 and y ≠ 0 in Li3InxY1−xBr6yCl6−6y) remains challenging. Herein, leveraging MACE architecture, we demonstrate that a single, fine-tuned model can accurately capture the entire energy landscape across this quaternary compositional space, including experimentally unexplored regions, thereby facilitating the discovery of new high-performance compositions.
Results
It is widely recognized that the distribution of the training dataset significantly influences the performance of MLIP models.60 To construct a comprehensive training dataset for a disordered, multi-component system, we developed a robust workflow, as illustrated in the upper panel of Fig. 1. First, we enumerate possible structures for each distinct composition within the investigated chemical system. Their energies were estimated using the uMLIP (MACE-MP), enabling the construction of a thermodynamic convex hull across the entire compositional space. Structures were then selected probabilistically based on their energy above the convex hull (Ehull) per formula unit, with a probability proportional to the Boltzmann factor, p ∝ exp(−Ehull/kBT), where kB is the Boltzmann constant. In this study, the temperature T for the Boltzmann distribution was set to 800 K. This probabilistic approach ensures a realistic balance between highly stable, low-energy configurations and thermodynamically accessible higher-energy structures, reflecting their thermal population. Additionally, the lowest-energy structure for each composition was always included to guarantee complete baseline coverage in the dataset. | ||
| Fig. 1 Workflow for fine-tuning the MLIPs developed in this work. The upper panel (in a dashed box) illustrates the procedure for constructing the training dataset. | ||
Subsequently, for each selected configuration, we performed MD simulations to sample the potential energy surface under non-equilibrium conditions using the same MACE-MP model in a canonical NVT ensemble at 800 K for 15 ps with a 1 fs timestep. The first 5 ps were allocated for equilibration and the remaining 10 ps were used for the production phase. This step aims to comprehensively explore the configuration space of each system. For each MD simulation, 30 structures were randomly selected, resulting in a total of around 2500 structures. A random strain, within ±3%, is also applied for each configuration to enrich the stress values sampled. Finally, single-point DFT calculations were performed on these diverse configurations to construct an accurate training dataset. The strongly constrained and appropriately normed semilocal density functional (SCAN)61 is employed in these DFT calculations to achieve high fidelity, compared to the Perdew–Burke–Ernzerhof (PBE) functional62 used in the original MACE-MP model. This multi-fidelity training strategy has been shown to be feasible and effective for improving model accuracy.6,63 Further methodological details of DFT are provided in the Methodology section.
Benchmark on NaSICON and LPGSO
The first system investigated is NaSICON (Na1+xZr2SixP3−xO12, x ∈ [0, 3], Δx = 0.25), focusing on its high-temperature phase in R![[3 with combining macron]](https://www.rsc.org/images/entities/char_0033_0304.gif) c (no. 167) space group.44 The crystal structure of NaSICON is presented in Fig. 2(a). Na ions occupy two partially occupied crystallographic sites – Na(1) and Na(2) – represented as yellow and orange balls, respectively, which contribute to the high ionic conductivity of NaSICON materials.44 ZrO6 units are depicted as green polyhedra, while PO4/SiO4 polyanionic units are represented as purple polyhedra.
c (no. 167) space group.44 The crystal structure of NaSICON is presented in Fig. 2(a). Na ions occupy two partially occupied crystallographic sites – Na(1) and Na(2) – represented as yellow and orange balls, respectively, which contribute to the high ionic conductivity of NaSICON materials.44 ZrO6 units are depicted as green polyhedra, while PO4/SiO4 polyanionic units are represented as purple polyhedra.
 | ||
| Fig. 2 (a) Crystal structure of Na1+xZr2SixP3−xO12 where Na(1) and Na(2) sites depicted as yellow and orange spheres, respectively. ZrO6 and PO4/SiO4 units are represented as green and purple polyhedra. (b) Quantitative comparison of validation performance between models fine-tuned with different sizes of training dataset. (c) Formation energy (Ef) as a function of composition x in Na1+xZr2SixP3−xO12. The thermodynamic convex hull is shown as black lines, with energies calculated using a model fine-tuned with approximately 2500 DFT calculations. (d) Activation energy barrier and conductivity at 300 °C for Na3Zr2Si2PO12 predicted by models trained on datasets of different sizes. Experimental results43,44 and simulation using MACE-MP without fine-tuning are included. Error bars for activation energy reflect standard deviations from Arrhenius plots (see Methodology section for more details). When the training data size is approximately 2500, structures with different PO4/SiO4 orderings are used for MD simulations. The lowest point is calculated from the ground state structure. | ||
We systematically fine-tune the MACE-MP model with varying training data sizes. Fig. 2(b) illustrates the performance of the model, quantified by the root mean square error (RMSE) on energies, forces, and stresses on the validation set, as a function of training dataset size. As the training dataset size increases, the RMSEs of all three properties decrease consistently. In particular, the continued reduction in the RMSEs of force and stress for training dataset sizes above 1000 suggests that additional data augmentation could further enhance the model's accuracy. Given the sufficiently low errors for reliable property predictions (0.5 meV per atom, 17.5 meV Å−1, and 0.4 meV Å−3 for energy, force, and stress, respectively), we adopt the model fine-tuned with the full dataset of approximately 2500 structures for subsequent calculations. The original MACE-MP model is excluded from this comparison because it is trained with calculated data that uses PBE functional, while we generated the training data for fine-tuning employing SCAN, making direct energy comparisons invalid.
The formation energy and phase stability of Na1+xZr2SixP3−xO12 are predicted, as shown in Fig. 2(c). Three stable compositions at x = 0, 2, and 3 are identified, consistent with the previous work.44 We then evaluate the performance of fine-tuned models as a function of training dataset size and compare it with the MACE-MP model. Specifically, we examine the ionic conductivity of the ground-state structure of Na3Zr2Si2PO12, comparing our findings with experimental results43,44 (Fig. 2(d) and Table S1). The Arrhenius plots obtained by MACE-MP and each fine-tuned model are shown in Fig. S1. The results indicate that the activation energies predicted by the various fine-tuned models, even with limited training data, align well with experimental measurements to within a difference of 0.02 eV. Furthermore, the calculated ionic conductivities at 300 °C also show approximate order-of-magnitude consistency with experimental observations.
Moreover, Fig. 2(d) suggests that increasing the training data size of the fine-tuned model generally leads to more stable predictions and smaller errors in the estimated activation energy. While the original MACE-MP model predicts σ300°C with the highest accuracy among all models, its activation energy deviates significantly from the experimental values, about 0.15 eV lower. The conductivities predicted by the fine-tuned models are stable across different training data sizes. The difference between the predicted and experimental conductivity values is small and acceptable, at 0.05–0.12 S cm−1. While there are some differences—potentially due to variations in the exchange–correlation functionals used to calculate the training data—the results are still reasonable, especially considering that the activation energy predictions demonstrate even greater precision. We also analyzed the MD trajectory of Na ions in Na3Zr2Si2PO12 at 800 K using GEMDAT software.64 The shape of the probability density (Fig. S2) is consistent with the literature.65 Besides, it has been reported that different PO4/SiO4 orderings can lead to variations in ionic conductivity (Fig. S3).44 To provide further validation, a series of Na3Zr2Si2PO12 configurations were generated using a Monte Carlo (MC) sampling approach37 on a 4 × 4 × 4 supercell. Subsequent MD simulations were performed on these structures, using the potential model trained on a dataset of 2500 points. The resulting ionic conductivities, presented in Fig. 2(d) (in transparent orange star points), demonstrate a clear increase from 0.08 S cm−1 to about 0.1 S cm−1 for the MC phases relative to the crystalline ground state.
Overall, using the NaSICON system as a benchmark, we demonstrate that fine-tuning with an appropriately sized dataset enables predictions comparable to experimental results. Our findings suggest that datasets of approximately 2500 structures provide a reliable balance between computational feasibility and predictive precision for NaSICON material, while a careful convergence test is always needed.
For the LPGSO system, we adopted structures for Li3+xP1−xGexS4−4xO4x (x ∈ [0, 1]) from the work of Li et al., where Pnma (no. 62) phase is used as the host model for both parent materials β-Li3PS4 and Li4GeO4 (Fig. 3(a)).50 The same strategy has been adopted for creating the training dataset from x ∈ [0, 1], Δx = 0.125, comprising approximately 2500 structures in total. A convergence test was also performed to determine the optimal training data size for the fine-tuned model (Fig. S4).
 | ||
| Fig. 3 (a) Crystal structure of Li3+xP1−xGexS4−4xO4x in the Pnma space group. Li atoms are depicted by green spheres, P/Ge atoms as purple, and S/O atoms as dark blue. (b) Computed formation energy, Ef, as a function of composition x in Li3+xP1−xGexS4−4xO4x. Thermodynamically stable structures are marked by black points on the convex hull. (c–e) Comparison of activation energies (Ea) and ionic conductivities (σ) between our model and reference (AIMD and/or experiments) values for (c) Li3PS4,66,67 (d) Li3.5P0.5Ge0.5S2O2,50,68 and (e) Li4GeO4 (Cmcm).69 Experimental and AIMD reference data are plotted as lines extrapolated to high temperatures from lower-temperature measurements/simulations. | ||
The formation energies of Li3+xP1−xGexS4−4xO4x compositions as predicted by the trained MLIP, are presented in Fig. 3(b). These results exhibit a convex hull profile closely resembling the DFT-calculated results obtained using the SCAN functional.50 Most importantly, our model predicts a stable phase Li3.5P0.5Ge0.5S2O2, which possesses the same structure and ordering as the DFT calculations and exhibits similar formation energy (−178 meV per atom for the fine-tuned model compared to −182 meV per atom from DFT). Furthermore, for its Li-ion kinetics, the model yields an activation energy of ∼0.29 eV for Li3.5P0.5Ge0.5S2O2, as shown in Fig. 3(d). Since this material remains unexplored experimentally, we compare the predicted ionic conductivities with those obtained from previous ab initio molecular dynamics (AIMD) simulations.50,68 These AIMD results exhibit discrepancies with conductivities spanning orders of magnitude, whereas our model predicts values that lie between these extremes while showing good agreement in activation energies.
Given the computational efficiency of fine-tuned models, which offers a distinct advantage over traditional, more expensive DFT simulations, it is of significant interest to examine all available phases of Li3PS4 beyond the β polymorph specifically investigated in the LPGSO system, to test the generalizability. It is known that Li3PS4 exhibits three distinct phases at different temperatures: the room-temperature γ phase (Pmn21), which transitions to the β phase (Pnma) at 573 K, and subsequently to the α phase (Cmcm) at 746 K.66,67,70–73 Here, the ionic conductivity of the two high-temperature phases of Li3PS4 (α and β) is plotted in Fig. 3(c). The predicted ionic conductivities align closely with experimental measurements, as evidenced by the near-overlap of theoretical and experimental data. Specifically, the predicted activation energies are ∼0.26 eV for the α phase and ∼0.44 eV for the β phase. These values compare favorably to experimental values of 0.20 eV and 0.41 eV for the α and β phases, respectively. Fig. S5 illustrates the probability density isosurface of Li ion diffusion in β-Li3PS4. The data, which was extracted from MLIP-MD simulations performed with our fine-tuned model, demonstrates migration pathways in good agreement with previous analysis by Kim et al.74
While the Pnma phase is consistently used as the host structural model in the LPGSO system, it is important to note that this phase for pure oxide Li4GeO4 is hypothetical and metastable. According to our MLIP calculations, the Pnma phase is about 30 meV per atom higher in energy than the experimentally observed Cmcm phase.75 To best facilitate validating our model against experimentally relevant structures that are not included in the training dataset, we specifically employ the Cmcm structure of Li4GeO4 to compute its kinetic properties. As shown in Fig. 3(e), the predictions of our model for Li-ion migration in this phase, yielding an activation energy of 0.77 ± 0.27 eV, show excellent agreement with experimental results of 0.81 eV, particularly in the higher temperature range (>900 K).69 The calculated ionic conductivities show deviations from ideal linear Arrhenius behavior at lower temperatures. This may be attributed to an insufficient number of ion migration events occurring in that temperature range because of poor ionic conductivity properties of this material.
This successful application to materials with different space group symmetries (e.g., Pmn21 and Cmcm) underscores the generality and versatility of the fine-tuned model. The pre-trained uMLIP provides knowledge for transfer, while fine-tuning allows for more accurate predictions and reduced errors.
Training from scratch vs. fine-tuning
In addition to fine-tuning, we also explored training MLIPs from scratch on the same datasets for the two benchmark material systems, NaSICON and LPGSO. Compared to fine-tuning, training from scratch generally reaches a slightly lower accuracy in both energy and force predictions (Table S2). Moreover, it is observed that models trained from scratch tend to be less stable during MD simulations, frequently resulting in unphysical behavior, e.g., the divergence of the total energy. For the LPGSO system in particular, the absence of prior knowledge about Li–Li interactions leads to incorrect Li–Li pair repulsion force, which severely alters the resulting diffusion behavior and produces unphysical MD trajectories, as plotted in Fig. S6 and S7. More detailed analysis is in SI.Exploring LIYBC system
In addition to sulfide and oxide SEs, the Li3MX6 family of lithium ternary halides (M = Sc, Y, In, Er, Sm–Lu; X = F, Cl, Br)76,77 has emerged as a promising class of SSEs, owing to its distinct advantages such as high ionic conductivity, wide electrochemical stability window, and good mechanical properties.54,78 In this section, we fine-tuned the MACE-MP model to study the miscibility within the complex halide system comprising Li3InCl6, Li3InBr6, and Li3YBr6. This forms the partial ternary system Li3InxY1−xBr6yCl6−6y. Our investigation specifically examines compositions defined by discrete values of x ∈ {0, 0.25, 0.5, 1} and y ∈ {0, 0.25, 0.5, 0.75, 1}, subject to the condition that x + y ≥ 1, as illustrated in Fig. 4(b).The three vertex compositions, namely Li3InCl6, Li3InBr6, and Li3YBr6, are isostructural and have all been reported to crystallize in a disordered C2/m space group within a cubic close-packed (CCP) framework.53,79 The conventional cell of each material comprises two formula units (20 atoms), with two symmetrically inequivalent Li atoms fractionally residing at positions with Wyckoff labels and multiplicities of 4g and 4h, respectively, as observed in the experimentally reported structures (ICSD: 122395 (ref. 80) for Li3InBr6, 29957 (ref. 80) for Li3YBr6).
The lowest-energy configuration for each member was identified from 2000 symmetrically distinct enumerations based on Ewald energies, encompassing configurations within the conventional cell, 1 × 1 × 2, and 1 × 2 × 2 supercells. The static lattice energies of those enumerations were then determined using the fine-tuned model. The validation results of the fine-tuned model are shown in Fig. S8, suggesting that a dataset of approximately 2500 samples is sufficient to train the model effectively. This is evidenced by its lower RMSE for energy, force, and stress, with error values of 0.6 meV per atom, 19.2 meV Å−1, and 0.3 meV Å−3, respectively.
Fig. 4(a) depicts the ball-and-stick representation of the host C2/m model for the Li3InxY1−xBr6yCl6−6y system. The two Li sites, 4g and 4h, are both partially occupied. The optimized lattice constants for each composition are listed in Table S4. While the equilibrium Li positions in each lowest-energy ordering may vary with composition, the arrangement of the [MX6]3− building blocks remains throughout the investigated compositional range. In each stoichiometric composition, Li ions are found to preferentially occupy the interplanar 4g-type sites. Conversely, the 4h sites exhibit partial occupancy, ranging from 50% to 62.5% across different compositions. These unoccupied 4h positions thus constitute natural interstitial sites, potentially facilitating Li ion hopping. Interestingly, in the ground-state structure of Li3In0.5Y0.5Br3Cl3, which features an equimolar mixture of In and Y metal cations as well as Br and Cl halides, we observed that each polyhedron is coordinated exclusively by either Br or Cl ions, with no evidence of mixed halide coordination within a single polyhedron. Instead, Cl preferentially coordinates with Y, and Br preferentially coordinates with In, forming pure [YCl6]3− and [InBr6]3− polyhedra, respectively (Fig. 5). This preference can perhaps be attributed to factors such as ionic size matching and specific bond strengths between the specific M–X pairs.
The phase diagram in the pseudo-ternary Li3InCl6–Li3InBr6–Li3YBr6 composition space is depicted in Fig. 4(b). It was found that Li3InBr6, Li3YBr6, Li3InCl6, Li3In0.5Y0.5Br6, and Li3In0.5Y0.5Br3Cl3 are stable (represented by solid circles), on the convex hull relative to their constituent vertex compositions. Other compositions (in dashed circle) all exhibit only trivial Ehull values, not exceeding 3 meV per atom, as detailed in Table S5. Even without considering entropy at finite temperatures, the low Ehull values suggest these materials are metastable and synthesizable, so they are included in the further study of kinetic performance.
For the lowest-energy configurations of each composition, we performed a series of MD simulations at temperatures ranging from 800 to 1100 K, since we observed that some compositions tend to melt at temperatures higher than 1100 K. Ionic conductivity data collected at 800 K is chosen as the performance metrics for this material. The calculated activation energy for Li ion diffusion is represented by the color in each circle in Fig. 4(b), and the contour plot, interpolated from the data on the tie-lines, shows the predicted ionic conductivity at 800 K. Arrhenius plots for each structure are presented in Fig. S9. Although ionic conductivity varies among compositions, all values remain the same order of magnitude, ranging from σ = 0.3 to 0.62 S cm−1 at 800 K.
The reliability of our predicted properties was validated by comparing with previously reported AIMD results for the vertex compositions, Li3YBr6 (ref. 81) and Li3InCl6.82 As shown in Fig. S10, both the ionic conductivity and activation energy agree with AIMD results. Particularly, for Li3YBr6, our predicted Ea is 0.29 eV, which agrees well AIMD result of 0.28 eV. For the case of Li3InCl6, our predicted Ea is 0.24 eV, compared to AIMD results of 0.20 eV. Furthermore, the differences in ionic conductivity for both Li3YBr6 and Li3InCl6 between AIMD and our predictions are within the same order of magnitude, with notable overlap observed within certain temperature ranges.
Analysis of Li-ion diffusion in LIYBC system
Among the whole LIYBC system, the composition Li3In0.5Y0.5Br3Cl3 shows the best ionic conductivity of ∼0.62 S cm−1 at 800 K, along with an activation energy of 0.28 eV. Other two mixed compositions, Li3InBr3Cl3 and Li3In0.5Y0.5Br6 also reaches the highest Li-ion conductivities 0.51 S cm−1 and 0.54 S cm−1 in each tie-line. This demonstrates the great potential of those mixed compositions in Li3MX6 systems for application as solid electrolytes. In addition, the vertex composition Li3InCl6 with conductivity of 0.57 S cm−1 also exhibits good performance.Here, we investigate the diffusion mechanism of Li ions in Li3In0.5Y0.5Br3Cl3 using MD simulation results at 800 K, with trajectory analysis performed by the GEMDAT package. Fig. 5 presents the three-dimensional probability density distribution of Li ions, where an isosurface value of 10−4 Å−3 was employed for visualization. The green spheres correspond to crystallographic host Li sites in the ground-state structure, which are mapped from the fully occupied 4g and 50% occupied 4h-type Li positions observed in experimental structures. The GEMDAT analysis of Li ion trajectories reveals that Li ions frequently visit interstitial sites adjacent to the host 4h-type sites, as indicated by the gray spheres in Fig. 5. These important interstitial sites, including those corresponding to unoccupied 4h positions, demonstrate crucial intermediates in the Li ion diffusion process within this material and are likely generalizable to other SSE systems.83
Two prevalent Li ion migration pathways are identified by analyzing the connectivity within the Li probability density isosurfaces and their respective occupation sites. The first pathway represents an intraplanar route connecting 4h sites within the ab plane, as indicated by the red arrows in the left panel of Fig. 5. The hopping of a Li ion from one host 4h site (labeled h1) to another (h2) is facilitated by three intermediate interstitial sites, i1, i2, and i3, forming a characteristic zigzag trajectory. Notably, i2 corresponds to a natural interstitial site derived from an unoccupied 4h position within the host lattice, while i1 and i3 are other symmetrically equivalent interstitial sites. The second representative pathway involves cross-plane migration (connecting intraplanar to interplanar regions), as shown by the blue arrows in the right panel of Fig. 5, illustrating migration within the ac plane. This migration path consists of a hop from host 4h (labeled h3) to host 4g (g1), bridged by an interstitial site (i4). The hopping then extends from g1 to g2 in the neighboring planes, assisted by additional intermediate sites i5 and i6, where the latter (i6) is a natural interstitial 4h-type site within the structure. These two pathways discussed above are also consistent with the previous study on Li3MX6 family.82 We also validate our results by performing AIMD simulations and the results are shown in Fig. S12 and S13. The probability density plots of Li ions are qualitatively consistent between MLIP-MD and AIMD.
We further quantify the Li ion transport properties by computing diffusivities and corresponding activation energies for the identified migration pathways. The planar transport characteristics (ab- and ac-plane), along with the overall three-dimensional results for comparison, are presented in Fig. S11 and Table S5. The diffusivity within a single plane surely does not represent the diffusivity along the entire migration pathway, while we use it here only for a straightforward comparison. For each composition, the diffusivities along different directions are generally within half an order of magnitude. The differences in corresponding activation energies are also comparable to the uncertainties in these activation energies. These results indicate that there is no clear directional preference for Li-ion diffusion in the LIYBC system.
Discussion and conclusions
In this work, we developed a comprehensive workflow for constructing MLIPs for disordered multi-component SSEs efficiently, which significantly reduces the time required for materials simulations from several months to a few days. The effectiveness of this approach is demonstrated through benchmarking on two complex materials systems, Na1+xZr2SixP3−xO12 and Li3+xP1−xGexS4−4xO4x. Performing MD simulations with a pre-trained uMLIP, rather than relying on AIMD, offers greater efficiency in building the training dataset for these systems. As compared in Fig. S14, for a system with the same number of atoms, running MLIP-MD with GPU can reach much higher efficiency, with almost the same accuracy as AIMD. Moreover, the current system sizes may not fully utilize the computational capacity of running MLIPs on GPUs. Larger systems are therefore expected to be handled with comparable efficiency. For each system, a dataset of approximately 2500 configurations was constructed. This dataset was sufficient to fine-tune the pre-trained PBE-level uMLIP (MACE-MP) to specific chemical spaces. High-fidelity SCAN data and a multi-head fine-tuning strategy were used in this process. Importantly, it is still recommended to perform convergence tests on the training dataset size. After fine-tuning with this proposed strategy, the formation energies and convex hulls can be obtained with low computational cost while retaining high accuracy. Overall, our model shows excellent agreement with previous experimental observations and DFT predictions of thermodynamic phase behavior. It also aligns well with experimental and AIMD results for Li/Na-ion kinetics, such as ionic conductivities and activation energies.This agreement demonstrates that fine-tuning is an effective strategy for developing accurate MLIPs. Beyond substantially reducing computational costs, fine-tuning a uMLIP can also mitigate its sensitivity to the training data compared to training from scratch, thereby yielding improved stability and generalizability. Moreover, fine-tuning can help address systematic softening observed in uMLIPs.84 Furthermore, by incorporating data computed at higher levels of theory, such as meta-GGA, this approach further enhances the fidelity of the resulting potential.
Building on the insights gained from benchmarking studies, we applied our workflow to a previously unexplored halide SSEs Li3InxY1−xBr6yCl6−6y with x + y ≥ 1. The anion sublattice enables both two-dimensional intralayer and three-dimensional cross-layer Li-ion migration pathways, offering desirable Li/Na-ion kinetics. Among the studied compositions, Li3In0.5Y0.5Br3Cl3 exhibits the highest ionic conductivity, reaching 0.62 S cm−1 at 800 K. Other compositions within the compositional space also demonstrate favorable Li-ion kinetics, with conductivity lower by one order of magnitude compared to Li3In0.5Y0.5Br3Cl3. Moreover, the polyanion mixing introduces configurational entropy, which could further enhance ion transport by reducing activation energies.85 These results suggest that the Li3InxY1−xBr6yCl6−6y system is promising, and merits further experimental exploration.
The MD trajectories reveal that the Li-ion diffusion in LIYBC involves both intralayer and interlayer Li-ion migration pathways. However, preference in pathways is different for different compositions. These variations arise from the distinct ground-state arrangements of anions/polyanions in each composition, which can influence factors such as diffusion bottlenecks and percolation of ion-conducting paths. Achieving optimal conductivity requires a holistic design strategy that integrates multiple factors.86 Further investigation into the controlling factors for ionic conductivity within this system is of great importance for guiding future material design.
To summarize, we have demonstrated that our workflow for developing an MLIP combining structural enumeration with fine-tuning of pre-trained uMLIPs enables rapid and accurate exploration of complex compositional and configurational spaces. We expect that this approach will be broadly applicable to other SSEs, providing a powerful computational framework to accelerate the discovery and optimization of the next-generation solid state batteries. Furthermore, MLIPs offer significant avenues for future exploration, including the incorporation of error quantification, model distillation to enhance accuracy, and further acceleration of simulations.
Methodology
DFT calculations were performed using the Vienna ab initio Simulation Package (VASP), version 6.3.2.87,88 All structures were selected from MD trajectories using MACE-MP-0a. The exchange–correlation energy in DFT was approximated by the strongly constrained and appropriately normed (SCAN) semi-local meta-generalized gradient approximation (meta-GGA) functional.61 Wave functions were expanded in terms of plane-wave basis set, truncated at a kinetic energy cutoff of 520 eV. Projector Augmented Wave (PAW) potentials were employed to describe core–valence electron interactions.89 The PAW (PBE_54) potentials used were Na (19 Sep 2006, 2p63s1), Zr (04 Jan 2005, 4s24p64d25s2), Si (05 Jan 2001, 3s23p2), P (06 Sep 2000, 3s23p3), Li (10 Sep 2004, 1s22s1), Ge (03 Jul 2007, 3d104s24p2), S (06 Sep 2000, 3s23p4), Y (25 May 2007, 4s24p65s24d1), In (06 Sep 2000, 4d105s25p1), Cl (06 Sep 2000, 3s23p5), Br (06 Sep 2000, 4s24p5), and O (08 Apr 2002, 2s22p4). The first Brillouin zone was integrated using a Γ-centered Monkhorst–Pack homogeneous sampling with spacing of ∼0.2 Å−1 in each lattice direction. Total energies were converged within 10−5 eV per cell during the electronic minimization, without preserving any symmetry.The pre-trained model used in this work is the medium-size MACE-MP-0a, trained on the MPTraj dataset.14 To focus on the effectiveness of our fine-tuning strategy, we restrict our discussion to MACE-MP-0a, although comparisons with other pre-trained uMLIPs were also conducted.
We performed multi-head fine-tuning on this model, rather than naive fine-tuning, to avoid catastrophic forgetting.29 The model was trained for a maximum of 500 epochs, employing an early stopping strategy if the validation loss did not improve for 20 consecutive epochs. The learning rate was set to 0.01.
For comparison, we also trained models from scratch, using an architecture closely aligned with MACE-MP-0a (medium version). Specifically, we adopted a message passing framework with 128 hidden channels, 2 interaction layers, a maximum correlation order of 3, and a radial cutoff of 5.0 Å. The training setup for models trained from scratch utilized the same learning rate and early stopping settings as the fine-tuned models.
To evaluate ionic conductivity properties, we used 2 × 2 × 2 supercell models for all systems under investigation. MD simulations were performed in the canonical (NVT) ensemble, using a Berendsen thermostat90 to control the temperature. For temperatures above 800 K, each simulation was run for 500 ps, which was deemed sufficient to collect statistics on ion hopping. To ensure that the system reached equilibrium, the initial 100 ps of the trajectory was discarded, and the remaining 400 ps was used for the diffusion analysis. For temperatures below 800 K, longer simulations (1 ns) were conducted to ensure adequate sampling. At these lower temperatures, the first 200 ps were excluded from further analysis.
The temperature-dependent mean squared displacement (MSD) of mobile ions of interest was calculated from their trajectories using the pymatgen package91 based on the equation:
 | (1) |
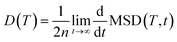 | (2) |
D(T) = D0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) e−Ea/(kBT) e−Ea/(kBT) | (3) |
 | (4) |
Author contributions
Z. D. designed and supervised the project. Y. D., and Z. D. performed the all the simulations with discussions with B. Z., G. S. G., and Y. L. Y. D. and Z. D. wrote the first draft. All the authors contributed to the data analysis, discussion, and final version of this manuscript.Conflicts of interest
The authors declare no competing financial interest.Data availability
Data for this article, including structure models used for MD simulations, interstitial sites for LIYBC analyzed from MD simulations, training datasets, and training scripts for each system, can be found at https://doi.org/10.5281/zenodo.15771782.Supplementary information is available: Detailed conductivity and activation energy data, Arrhenius plots, probability density of ions for each material system; comparison between fine-tuning and training from scratch; lattice parmeters and energy above hull, 2D diffusion data of LIYBC; computation cost. See DOI: https://doi.org/10.1039/d5ta05321h.
Acknowledgements
Z. D. acknowledges the support from his Lee Kuan Yew Postdoctoral Fellowship (22-5930-A0001), A*STAR under its MTC-YIRG (M24N8c0108), NRF-Singapore under its Competitive Research Programme (NRF-CRP30-2023-0001), and the Ministry of Education, Singapore, under the Academic Research Fund Tier 1 (FY2025). B. Z. acknowledges the support by the National Natural Science Foundation of China (Grant No. 12404068). Computational work involved in this research work is partially supported by NUS IT's Research Computing group using grant number NUSREC-HPC-00001.Notes and references
- L. C. Erhard, J. Rohrer, K. Albe and V. L. Deringer, Nat. Commun., 2024, 15, 1927 CrossRef CAS PubMed.
- Y. Zhou, W. Zhang, E. Ma and V. L. Deringer, Nat. Electron., 2023, 6, 746–754 CrossRef CAS.
- C. Chang, V. L. Deringer, K. S. Katti, V. Van Speybroeck and C. M. Wolverton, Nat. Rev. Mater., 2023, 8, 309–313 CrossRef PubMed.
- V. L. Deringer, M. A. Caro and G. Csányi, Adv. Mater., 2019, 31, 1902765 CrossRef CAS PubMed.
- J. Riebesell, R. E. A. Goodall, P. Benner, Y. Chiang, B. Deng, G. Ceder, M. Asta, A. A. Lee, A. Jain and K. A. Persson, Nat. Mach. Intell., 2025, 7, 836–847 CrossRef.
- J. Kim, J. Kim, J. Kim, J. Lee, Y. Park, Y. Kang and S. Han, J. Am. Chem. Soc., 2025, 147, 1042–1054 CrossRef PubMed.
- A. P. Bartók, M. C. Payne, R. Kondor and G. Csányi, Phys. Rev. Lett., 2010, 104, 136403 CrossRef PubMed.
- A. V. Shapeev, Multiscale Model. Simul., 2016, 14, 1153–1173 CrossRef.
- H. Wang, L. Zhang, J. Han and W. E, Comput. Phys. Commun., 2018, 228, 178–184 CrossRef CAS.
- J. Behler and M. Parrinello, Phys. Rev. Lett., 2007, 98, 146401 CrossRef PubMed.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals and G. E. Dahl, Proceedings of the 34th International Conference on Machine Learning – Volume 70, Sydney, NSW, Australia, 2017, pp. 1263–1272 Search PubMed.
- T. Xie and J. C. Grossman, Phys. Rev. Lett., 2018, 120, 145301 CrossRef CAS PubMed.
- C. Chen and S. P. Ong, Nat. Comput. Sci., 2022, 2, 718–728 CrossRef PubMed.
- B. Deng, P. Zhong, K. Jun, J. Riebesell, K. Han, C. J. Bartel and G. Ceder, Nat. Mach. Intell., 2023, 5, 1031–1041 CrossRef.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. P. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. Kozinsky, Nat. Commun., 2022, 13, 2453 CrossRef CAS PubMed.
- I. Batatia, D. P. Kovacs, G. Simm, C. Ortner and G. Csanyi, Advances in Neural Information Processing Systems, 2022, pp. 11423–11436 Search PubMed.
- Y. Park, J. Kim, S. Hwang and S. Han, J. Chem. Theory Comput., 2024, 20, 4857–4868 CrossRef CAS PubMed.
- F. Fuchs, D. Worrall, V. Fischer and M. Welling, Advances in Neural Information Processing Systems, 2020, pp. 1970–1981 Search PubMed.
- R. Drautz, Phys. Rev. B, 2019, 99, 014104 CrossRef CAS.
- A. Jain, S. P. Ong, G. Hautier, W. Chen, W. D. Richards, S. Dacek, S. Cholia, D. Gunter, D. Skinner, G. Ceder and K. A. Persson, APL Mater., 2013, 1, 011002 CrossRef.
- S. Curtarolo, W. Setyawan, G. L. W. Hart, M. Jahnatek, R. V. Chepulskii, R. H. Taylor, S. Wang, J. Xue, K. Yang, O. Levy, M. J. Mehl, H. T. Stokes, D. O. Demchenko and D. Morgan, Comput. Mater. Sci., 2012, 58, 218–226 CrossRef CAS.
- J. E. Saal, S. Kirklin, M. Aykol, B. Meredig and C. Wolverton, JOM, 2013, 65, 1501–1509 CrossRef CAS.
- J. Schmidt, H.-C. Wang, T. F. T. Cerqueira, S. Botti and M. A. L. Marques, Sci. Data, 2022, 9, 64 CrossRef CAS PubMed.
- L. Barroso-Luque, M. Shuaibi, X. Fu, B. M. Wood, M. Dzamba, M. Gao, A. Rizvi, C. L. Zitnick and Z. W. Ulissi, Open Materials 2024 (OMat24) Inorganic Materials Dataset and Models, arXiv, 2024, preprint, arXiv:2410.12771 [cond-mat.mtrl-sci], DOI:10.48550/arXiv.2410.12771, http://arxiv.org/abs/2410.12771.
- A. D. Kaplan, R. Liu, J. Qi, T. W. Ko, B. Deng, J. Riebesell, G. Ceder, K. A. Persson and S. P. Ong, A Foundational Potential Energy Surface Dataset for Materials, arXiv, 2025, preprint, arXiv:2503.04070 [cond-mat.mtrl-sci], DOI:10.48550/arXiv.2503.04070, http://arxiv.org/abs/2503.04070.
- Z. El-Machachi, D. Frantzov, A. Nijamudheen, T. Zarrouk, M. A. Caro and V. L. Deringer, Angew. Chem., Int. Ed., 2024, 63, e202410088 CrossRef CAS PubMed.
- C. Ben Mahmoud, J. L. A. Gardner and V. L. Deringer, Nat. Comput. Sci., 2024, 4, 384–387 CrossRef PubMed.
- H. Yu, M. Giantomassi, G. Matteo, J. Wang and G. Rignanese, Mater. Genome Eng. Adv., 2024, 2(3), 58 CrossRef.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon, W. J. Baldwin, F. Berger, N. Bernstein, A. Bhowmik, S. M. Blau, V. Cărare, J. P. Darby, S. De, F. Della Pia, V. L. Deringer, R. Elijošius, Z. El-Machachi, F. Falcioni, E. Fako, A. C. Ferrari, A. Genreith-Schriever, J. George, R. E. A. Goodall, C. P. Grey, P. Grigorev, S. Han, W. Handley, H. H. Heenen, K. Hermansson, C. Holm, J. Jaafar, S. Hofmann, K. S. Jakob, H. Jung, V. Kapil, A. D. Kaplan, N. Karimitari, J. R. Kermode, N. Kroupa, J. Kullgren, M. C. Kuner, D. Kuryla, G. Liepuoniute, J. T. Margraf, I.-B. Magdău, A. Michaelides, J. H. Moore, A. A. Naik, S. P. Niblett, S. W. Norwood, N. O'Neill, C. Ortner, K. A. Persson, K. Reuter, A. S. Rosen, L. L. Schaaf, C. Schran, B. X. Shi, E. Sivonxay, T. K. Stenczel, V. Svahn, C. Sutton, T. D. Swinburne, J. Tilly, C. van der Oord, E. Varga-Umbrich, T. Vegge, M. Vondrák, Y. Wang, W. C. Witt, F. Zills and G. Csányi, A foundation model for atomistic materials chemistry, arXiv, 2024, preprint, arXiv:2401.00096 [physics.chem-ph], DOI:10.48550/arXiv.2401.00096, http://arxiv.org/abs/2401.00096.
- X. Fu, B. M. Wood, L. Barroso-Luque, D. S. Levine, M. Gao, M. Dzamba and C. L. Zitnick, Learning Smooth and Expressive Interatomic Potentials for Physical Property Prediction, arXiv, 2025, preprint, arXiv:2502.12147 [physics.comp-ph], DOI:10.48550/arXiv.2502.12147, http://arxiv.org/abs/2502.12147.
- S. Gong, Y. Zhang, Z. Mu, Z. Pu, H. Wang, X. Han, Z. Yu, M. Chen, T. Zheng, Z. Wang, L. Chen, Z. Yang, X. Wu, S. Shi, W. Gao, W. Yan and L. Xiang, Nat. Mach. Intell., 2025, 7, 543–552 CrossRef.
- T. Famprikis, P. Canepa, J. A. Dawson, M. S. Islam and C. Masquelier, Nat. Mater., 2019, 18, 1278–1291 CrossRef CAS PubMed.
- J. C. Bachman, S. Muy, A. Grimaud, H.-H. Chang, N. Pour, S. F. Lux, O. Paschos, F. Maglia, S. Lupart, P. Lamp, L. Giordano and Y. Shao-Horn, Chem. Rev., 2016, 116, 140–162 CrossRef CAS PubMed.
- Q. Zhao, S. Stalin, C.-Z. Zhao and L. A. Archer, Nat. Rev. Mater., 2020, 5, 229–252 CrossRef CAS.
- Y. He, Q. Chen and W. Lai, Solid State Ionics, 2023, 399, 116298 CrossRef CAS.
- L. Xu, W. Shao, H. Jin and Q. Wang, J. Phys. Chem. C, 2023, 127, 24106–24117 CrossRef CAS.
- A. P. Maltsev, I. V. Chepkasov and A. R. Oganov, ACS Appl. Mater. Interfaces, 2023, 15, 42511–42519 CrossRef CAS PubMed.
- J. Choi, B. Jun and Y. Jung, Chem. Eng. J., 2025, 516, 163847 CrossRef CAS.
- Z. A. H. Goodwin, M. B. Wenny, J. H. Yang, A. Cepellotti, J. Ding, K. Bystrom, B. R. Duschatko, A. Johansson, L. Sun, S. Batzner, A. Musaelian, J. A. Mason, B. Kozinsky and N. Molinari, J. Phys. Chem. Lett., 2024, 15, 7539–7547 CrossRef CAS PubMed.
- J. Wang, A. A. Panchal, G. S. Gautam and P. Canepa, J. Mater. Chem. A, 2022, 10, 19732–19742 RSC.
- A. Seth, R. P. Kulkarni and G. S. Gautam, ACS Mater. Au, 2025, 5, 458–468 CrossRef CAS PubMed.
- A. C. C. Dutra, B. A. Goldmann, M. S. Islam and J. A. Dawson, Nat. Rev. Mater., 2025, 1–18 Search PubMed.
- J. B. Goodenough, H. Y.-P. Hong and J. A. Kafalas, Mater. Res. Bull., 1976, 11, 203–220 CrossRef CAS.
- Z. Deng, T. P. Mishra, E. Mahayoni, Q. Ma, A. J. K. Tieu, O. Guillon, J.-N. Chotard, V. Seznec, A. K. Cheetham, C. Masquelier, G. S. Gautam and P. Canepa, Nat. Commun., 2022, 13, 4470 CrossRef CAS PubMed.
- Z. Wang, S. Park, Z. Deng, D. Carlier, J.-N. Chotard, L. Croguennec, G. S. Gautam, A. K. Cheetham, C. Masquelier and P. Canepa, J. Mater. Chem. A, 2022, 10, 209–217 RSC.
- Z. Wang, T. P. Mishra, W. Xie, Z. Deng, G. S. Gautam, A. K. Cheetham and P. Canepa, ACS Mater. Lett., 2023, 5, 2499–2507 CrossRef CAS.
- B. Ouyang, J. Wang, T. He, C. J. Bartel, H. Huo, Y. Wang, V. Lacivita, H. Kim and G. Ceder, Nat. Commun., 2021, 12, 5752 CrossRef CAS PubMed.
- Q. Ma, C.-L. Tsai, X.-K. Wei, M. Heggen, F. Tietz and J. T. S. Irvine, J. Mater. Chem. A, 2019, 7766–7776 RSC.
- J. P. Boilot, G. Collin and P. Colomban, J. Solid State Chem., 1988, 73, 160–171 CrossRef CAS.
- Y. Li, Z. Deng and C. Chen, Chem. Mater., 2024, 36, 7877–7886 CAS.
- Y. Tomita, A. Fuji-i, H. Ohki, K. Yamada and T. Okuda, Chem. Lett., 1998, 27, 223–224 CrossRef.
- K. Yamada, K. Kumano and T. Okuda, Solid State Ionics, 2006, 177, 1691–1695 CrossRef CAS.
- Y. Tomita, H. Matsushita, K. Kobayashi, Y. Maeda and K. Yamada, Solid State Ionics, 2008, 179, 867–870 CrossRef CAS.
- X. Li, J. Liang, X. Yang, K. R. Adair, C. Wang, F. Zhao and X. Sun, Energy Environ. Sci., 2020, 13, 1429–1461 RSC.
- J. O. Bonsu, A. Bhadra and D. Kundu, Adv. Sci., 2024, 11, 2403208 CrossRef CAS PubMed.
- R. E. Skyner, J. B. O. Mitchell and C. R. Groom, CrystEngComm, 2017, 19, 641–652 RSC.
- P. Molaiyan, S. E. Mailhiot, K. Voges, A. M. Kantola, T. Hu, P. Michalowski, A. Kwade, V.-V. Telkki and U. Lassi, Mater. Des., 2023, 227, 111690 CrossRef CAS.
- X. Li, J. Liang, J. Luo, M. N. Banis, C. Wang, W. Li, S. Deng, C. Yu, F. Zhao, Y. Hu, T.-K. Sham, L. Zhang, S. Zhao, S. Lu, H. Huang, R. Li, K. R. Adair and X. Sun, Energy Environ. Sci., 2019, 12, 2665–2671 RSC.
- F. Stainer and H. M. R. Wilkening, Phys. Rev. B, 2024, 109, 174304 CrossRef CAS.
- F. Musil, A. Grisafi, A. P. Bartók, C. Ortner, G. Csányi and M. Ceriotti, Chem. Rev., 2021, 121, 9759–9815 CrossRef CAS PubMed.
- J. Sun, A. Ruzsinszky and J. Perdew, Phys. Rev. Lett., 2015, 115, 036402 CrossRef PubMed.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- X. Huang, B. Deng, P. Zhong, A. D. Kaplan, K. A. Persson and G. Ceder, Cross-functional transferability in universal machine learning interatomic potentials, arXiv, 2025, preprint, arXiv:2504.05565 [cond-mat.mtrl-sci], DOI:10.48550/arXiv.2504.05565, http://arxiv.org/abs/2504.05565.
- V. Azizi, S. Smeets, A. K. Lavrinenko, S. Ciarella, T. Famprikis, V. Landgraf and A. Vasileiadis, GEMDAT, 2025, https://zenodo.org/records/15235004 Search PubMed.
- L. Pradhan and P. Padma Kumar, J. Phys. Chem. C, 2025, 129, 13756–13767 CrossRef.
- J.-S. Kim, W. D. Jung, S. Choi, J.-W. Son, B.-K. Kim, J.-H. Lee and H. Kim, J. Phys. Chem. Lett., 2018, 9, 5592–5597 CrossRef CAS PubMed.
- N. J. de Klerk, E. van der Maas and M. Wagemaker, ACS Appl. Energy Mater., 2018, 1, 3230–3242 CrossRef CAS PubMed.
- B. Zhang, M. Weng, Z. Lin, Y. Feng, L. Yang, L.-W. Wang and F. Pan, Small, 2020, 16, 1906374 CrossRef CAS PubMed.
- S. B. Yahya, I. Garoui, M. Zaghrioui, A. Oueslati and B. Louati, RSC Adv., 2025, 15, 9295–9304 RSC.
- Y. Liu, X. He and Y. Mo, npj Comput. Mater., 2023, 9, 1–13 CrossRef CAS.
- K. Homma, M. Yonemura, T. Kobayashi, M. Nagao, M. Hirayama and R. Kanno, Solid State Ionics, 2011, 182, 53–58 CrossRef CAS.
- T. Kimura, T. Inaoka, R. Izawa, T. Nakano, C. Hotehama, A. Sakuda, M. Tatsumisago and A. Hayashi, J. Am. Chem. Soc., 2023, 145, 14466–14474 CrossRef CAS PubMed.
- N. D. Lepley, N. A. W. Holzwarth and Y. A. Du, Phys. Rev. B: Condens. Matter Mater. Phys., 2013, 88, 104103 CrossRef.
- J. H. Kim, B. Jun, Y. J. Jang, S. H. Choi, S. H. Choi, S. M. Cho, Y.-G. Kim, B.-H. Kim and S. U. Lee, Nano Energy, 2024, 124, 109436 CrossRef CAS.
- I. M. Hodge, M. D. Ingram and A. R. West, J. Am. Ceram. Soc., 1976, 59, 360–366 CrossRef CAS.
- T. Asano, A. Sakai, S. Ouchi, M. Sakaida, A. Miyazaki and S. Hasegawa, Adv. Mater., 2018, 30, 1803075 CrossRef PubMed.
- X. He, Y. Zhu and Y. Mo, Nat. Commun., 2017, 8, 15893 CrossRef CAS PubMed.
- R. Li, P. Lu, X. Liang, L. Liu, M. Avdeev, Z. Deng, S. Li, K. Xu, J. Feng, R. Si, F. Wu, Z. Zhang and Y.-S. Hu, ACS Energy Lett., 2024, 9, 1043–1052 CrossRef CAS.
- T. Jeon and S. C. Jung, J. Mater. Chem. A, 2023, 11, 4334–4344 RSC.
- B. Helm, R. Schlem, B. Wankmiller, A. Banik, A. Gautam, J. Ruhl, C. Li, M. R. Hansen and W. G. Zeier, Chem. Mater., 2021, 33, 4773–4782 CrossRef CAS.
- S. Wang, Q. Bai, A. M. Nolan, Y. Liu, S. Gong, Q. Sun and Y. Mo, Angew. Chem., Int. Ed., 2019, 58, 8039–8043 CrossRef CAS PubMed.
- D. Park, H. Park, Y. Lee, S.-O. Kim, H.-G. Jung, K. Y. Chung, J. H. Shim and S. Yu, ACS Appl. Mater. Interfaces, 2020, 12, 34806–34814 CrossRef CAS PubMed.
- Y. Li and N. A. W. Holzwarth, Phys. Rev. Mater., 2022, 6, 025401 CrossRef CAS.
- B. Deng, Y. Choi, P. Zhong, J. Riebesell, S. Anand, Z. Li, K. Jun, K. A. Persson and G. Ceder, npj Comput. Mater., 2025, 11, 9 CrossRef CAS.
- Y. Zeng, B. Ouyang, J. Liu, Y.-W. Byeon, Z. Cai, L. J. Miara, Y. Wang and G. Ceder, Science, 2022, 378, 1320–1324 CrossRef CAS PubMed.
- K. Jun, Y. Chen, G. Wei, X. Yang and G. Ceder, Nat. Rev. Mater., 2024, 1–19 CAS.
- G. Kresse and J. Furthmüller, Comput. Mater. Sci., 1996, 6, 15–50 CrossRef CAS.
- G. Kresse and J. Furthmüller, Phys. Rev. B: Condens. Matter Mater. Phys., 1996, 54, 11169–11186 CrossRef CAS PubMed.
- G. Kresse and D. Joubert, Phys. Rev. B: Condens. Matter Mater. Phys., 1999, 59, 1758–1775 CrossRef CAS.
- H. J. C. Berendsen, J. P. M. Postma, W. F. Van Gunsteren, A. DiNola and J. R. Haak, J. Chem. Phys., 1984, 81, 3684–3690 CrossRef CAS.
- S. P. Ong, W. D. Richards, A. Jain, G. Hautier, M. Kocher, S. Cholia, D. Gunter, V. L. Chevrier, K. A. Persson and G. Ceder, Comput. Mater. Sci., 2013, 68, 314–319 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2025 |



