Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Multi-stimuli responsive nanomaterials assembled from spiropyran-containing peptoids
Progyateg
Chakma
 ab,
Bradley S.
Harris
ab,
Bradley S.
Harris
 a,
Chenyang
Shi
a,
Chenyang
Shi
 a,
Botao
Hao
a,
Botao
Hao
 a,
Kyle
Whitaker
b,
Marcel D.
Baer
a,
Kyle
Whitaker
b,
Marcel D.
Baer
 a and
Chun-Long
Chen
a and
Chun-Long
Chen
 *ac
*ac
aPhysical Sciences Division, Pacific Northwest National Laboratory, Richland, WA 99352, USA. E-mail: Chunlong.Chen@pnnl.gov
bDepartment of Chemistry and Biochemistry, Kennesaw State University, Kennesaw, GA 30144, USA
cDepartment of Chemical Engineering, University of Washington, Seattle, WA 98195, USA
First published on 12th November 2025
Abstract
Stimuli-responsive tunable supramolecular assemblies are highly desirable for controlling functionalities via morphological changes and alternations in crystallinity. Here, we report that amphiphilic peptoids with a multifunctional spiropyran (SP) group assemble into responsive 2D nanosheets and 1D nanotubes through light and heat-mediated isomerization. Computational analysis by periodic density functional theory (DFT) calculations on dimers of SP and its isomer, merocyanine (MC), reveals significantly more favorable packing energetics for MC compared to SP, further supporting the high crystallinity of assemblies composed of peptoids with MC. Furthermore, we demonstrate how stimulus-responsive control over peptoid assembly crystallinity can be utilized in tuning catalytic activity in mimicking carbonic anhydrase (CA) enzymes.
Introduction
Molecular self-assembly into supramolecular structures is essential for creating functionality in biological systems.1–4 Inspired by nature, significant efforts in supramolecular chemistry and materials science have focused on constructing synthetic supramolecular assemblies with sophisticated and complex functions.1–3,5,6 In particular, there is considerable interest in stimuli-responsive assemblies of organic molecules, which can be modulated to achieve specific morphologies and functionalities.1,7–9 Often, the macroscopic properties of these supramolecular assemblies can be reversibly manipulated through external stimuli such as light, pH value, heat, mechanical force, and salts.1,3,10 In particular, light as a stimulus has significant advantages, as it can be applied to a specific location with responsive moieties activated at different wavelengths, with varying energy input, and in different directions.11–13 Among the photoswitches, spiropyran (SP) is highly unique as it reversibly isomerizes into the “open-ring” merocyanine (MC) form, exhibiting significant changes in physicochemical properties.14,15 For example, SP is a non-polar and non-planar molecule. In contrast, isomerization into MC induces a planar structure with extended pi–pi interaction and a change in dipole moments (from ∼4–6 D to ∼14–18 D) (Fig. 1). Interestingly, in addition to being a photoswitch, reversible SP to MC isomerization can be induced further by temperature, mechanical force, solvent polarity, pH, metal ions, and ionic concentration.14,16 The light-triggered pH switch enables the transition between SP and MC, affording Sp-containing materials with great potential in applications such as CO2 capture and sequestration.13,17,18 Due to their multi-stimuli-responsive and multifunctional properties, SP moieties have been frequently explored and utilized to construct stimuli-responsive functional materials.14,19–22 However, minimal effort has been made to connect SPs with sequence-defined synthetic polymers for the development of biomimetic materials with multi-stimuli responsiveness and protein-like high information content. | ||
| Fig. 1 Stimuli-responsive reversible transformation of spiropyran and merocyanine and their functionalization into peptoids, leading to tunable control of assembly into supramolecular structures. | ||
Peptoids, known as poly-N-substituted glycines, are a type of well-advanced, sequence-defined peptidomimetics.1,2,23,24 Unlike peptides, where self-assembly is primarily driven by hydrogen bonding, peptoids rely on controlled hydrophobic interactions and side-chain chemistry for their self-assembly. This leads to enhanced stability while maintaining biocompatibility.1,2,23,25 Recently, we and others have designed and synthesized a series of amphiphilic peptoids that self-assemble into highly crystalline, hierarchical nanomaterials with various morphologies and functionalities, as well as excellent chemical and thermal stability.1,2,25–27 In particular, peptide- and peptoid-based stimuli-responsive biomaterials have recently gained significant attention due to their excellent prospects in biomedical and biotechnology applications.1,2,6,28 However, SP has never been used as a functional group for the development of responsive peptoid assembly materials.
Here, for the first time, we report the covalent attachment of SP at the N-terminus of self-assembling peptoids for the development of light- and heat-responsive peptoid nanomaterials with nanosheet or nanotube morphologies. We demonstrate that assembly morphology and crystallinity of these peptoid nanomaterials can be modulated through light- and heat-mediated SP and MC isomerization (Fig. 1). To better understand the molecular origin of these stimuli-responsive behaviors, we also performed quantum chemical calculations on representative SP and MC dimer structures. These computations reveal distinct energetic preferences for MC-based stacking, which help rationalize the experimentally observed differences in crystallinity and assembly kinetics. Finally, we explored how stimulus-responsive control of the crystallinity of assembled peptoid nanomaterials can be utilized to tune catalytic activity in mimicking carbonic anhydrase (CA) enzymes.
Results and discussion
Design and synthesis of peptoid sequences conjugated with a spiropyran group
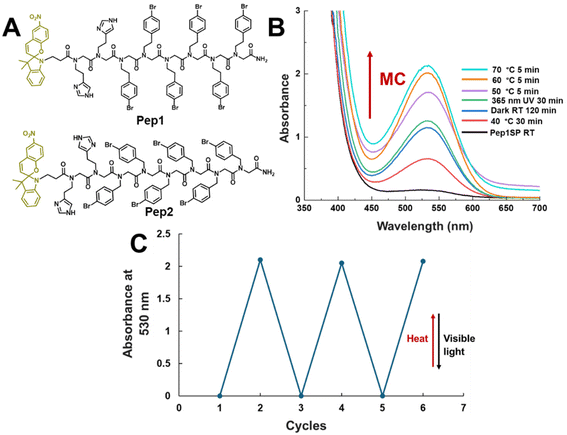
To study the effect of reversible photochromism of SP, we designed and synthesized amphiphilic peptoids with either six Nbrpe = N-([2-(4-bromophenyl)ethyl]glycine) or Nbrpm = N-[(4-bromophenyl)methyl] glycine were introduced as side chains for the hydrophobic domains that are responsible for assembly into highly crystalline nanosheets or nanotubes via aromatic interactions.2,27,29–33 Here, the π–π interaction of phenyl rings in six Nbrpe groups leads to the formation of a flat, sheet-like structure.29,34–36 In contrast, the molecular packing of peptoids containing six Nbrpm groups leads to the formation of a tubular structure.26,27,30,35,37–39 On the other hand, two [2-(4-imidazolyl)ethyl]glycine (Nhis) were introduced in the polar domain to help with solubility and processability. Here, in addition to facilitating solubility, Nhis side chain was used to mimic the active sites of carbonic anhydrase (CA) enzymes, thus creating responsive CA-mimics for enhanced CO2 hydration and sequestration.35 Finally, the SP moiety was functionalized at the N-terminus of self-assembly peptoids to study the effect of changes in polarity, solubility, steric, and supramolecular interactions through reversible SP to MC photochromism. We hypothesized that, while the self-assembly of amphiphilic peptoids into crystalline nanomaterials is dominated by enhanced hydrophobic interactions (e.g., π–π stacking) among hydrophobic domains,40–45 drastic changes in the physicochemical properties of the functional group at the N-terminus can significantly influence the properties of the peptoid assembly process. Peptoids with Nbrpe, Nbrpm, and Nhis side chains are shown in Fig. 2A. All peptoids were synthesized using the submonomer solid-phase synthesis method previously reported by our group.26,27,32,46 Synthesized peptoids were purified using reverse-phase high-performance liquid chromatography (HPLC), and their purity was confirmed by ultra-performance liquid chromatography equipped with mass spectrometry (UPLC-MS) (Fig. S1 and 2). The detailed synthesis and characterization methodology for these peptoids can be found in the Methods and SI sections.Reversible isomerism of spiropyran-functionalized peptoids
Before proceeding with the self-assembly of the peptoids, we initially studied the reversible photochromism of SP to MC using ultraviolet–visible (UV-Vis) spectroscopy with Pep1 to determine optimum conditions for this transformation. In the presence of ambient visible light and room temperature, Pep1 remains in SP form (Pep1SP), as no characteristic MC absorbance is observed in the 500–550 nm region. Typically, UV irradiation is utilized to initiate SP to MC transformation.14 Interestingly, minimal success was observed when irradiating the Pep1SP solution (0.1 mM in acetonitrile/water; v/v = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) with long-wavelength 365 nm UV light for 30 minutes or when leaving it in the absence of light, as shown in Fig. 2B. Recently, Wang et al. reported that the reverse photochromism of SP to MC can be effectively promoted by heating.22 Inspired by their report, when the Pep1SP solution was heated from 40 °C to 70 °C, a significant increase in the characteristic MC absorbance at 530 nm was observed (Fig. 2B), indicating that the opening of the SP moiety and transformation to Pep1MC can be effectively mediated by relatively mild heating, without any UV irradiation. Interestingly, reverse transformation to Pep1SP was very rapid upon exposure to warm white light (1 minute) or ambient visible light (5 minutes). The heat-light-responsive SP to MC transformation can be repeated for at least three cycles without any noticeable degradation (Fig. S5).
1) with long-wavelength 365 nm UV light for 30 minutes or when leaving it in the absence of light, as shown in Fig. 2B. Recently, Wang et al. reported that the reverse photochromism of SP to MC can be effectively promoted by heating.22 Inspired by their report, when the Pep1SP solution was heated from 40 °C to 70 °C, a significant increase in the characteristic MC absorbance at 530 nm was observed (Fig. 2B), indicating that the opening of the SP moiety and transformation to Pep1MC can be effectively mediated by relatively mild heating, without any UV irradiation. Interestingly, reverse transformation to Pep1SP was very rapid upon exposure to warm white light (1 minute) or ambient visible light (5 minutes). The heat-light-responsive SP to MC transformation can be repeated for at least three cycles without any noticeable degradation (Fig. S5).
Assembly of SP- and MC-peptoids
Based on the initial finding of heat-mediated, highly efficient transformation of SP to MC, we proceeded to study the self-assembly of Pep1 under different conditions. Typically, Pep1, containing six Nbrpe side groups, should lead to the formation of crystalline 2D nanosheets.34–36,46 We were interested in studying how the incorporation of a bulky and non-polar SP moiety, followed by its transformation into a planar and polar MC isomer, affects the self-assembly properties of amphiphilic peptoids. For the assembly of Pep1SP, lyophilized peptoid powder was dissolved in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) acetonitrile/water solution and left at 4 °C in the presence of ambient visible light to facilitate slow crystallization. On the other hand, for the assembly of Pep1MC, the Pep1MC solution in acetonitrile/water (v/v = 1
1 (v/v) acetonitrile/water solution and left at 4 °C in the presence of ambient visible light to facilitate slow crystallization. On the other hand, for the assembly of Pep1MC, the Pep1MC solution in acetonitrile/water (v/v = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
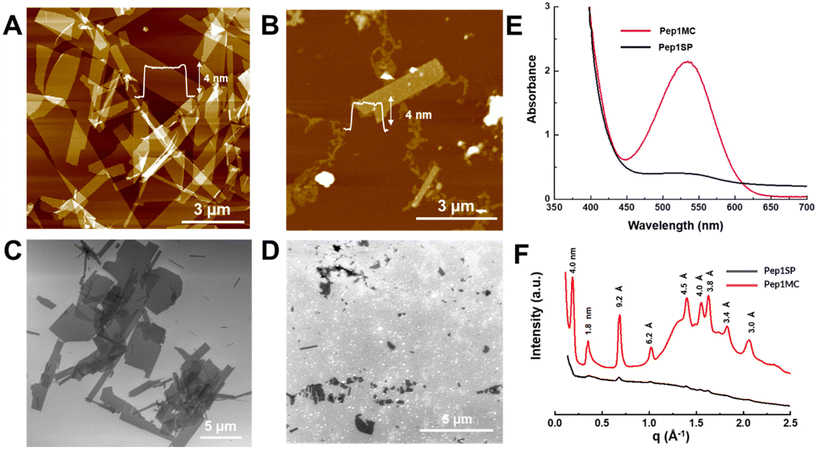
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) was heated at 60 °C for 20 minutes to facilitate the SP to MC transformation. A complete conversion of Pep1SP to Pep1MC was confirmed by UV-Vis spectroscopy. The resulting Pep1MC solution was then left at 4 °C in the dark to allow for slow evaporation, which triggered the self-assembly. Interestingly, a deep red, gel-like solution was obtained after about one week (Fig. S5). In contrast, the self-assembly of Pep1SP was much slower; a slightly yellowish, gel-like solution with slurry was obtained after at least two weeks of slow evaporation of the Pep1SP solution (Fig. S5). These results indicate that the MC form is much more favorable for the peptoid gelation (i.e., crystallization) than the SP form. In contrast, in the SP form, self-assembly of Pep1 into crystalline nanosheets is very challenging. To assess the morphology of assembled Pep1MC and Pep1SP materials, atomic force microscopy (AFM), scanning electron microscopy (SEM), and transmission electron microscopy (TEM) were employed. Both AFM and SEM characterizations of the assembled Pep1MC materials (Fig. 3A and C) reveal the formation of 2D nanosheets with lateral dimensions exceeding 5.0 µm, straight edges, and a thickness of approximately 4.0 nm. The formation of such large 2D nanosheets was further confirmed by liquid-cell AFM and TEM characterizations (Fig. S7A and S7B). On the contrary, AFM and SEM results show that assembled Pep1SP materials primarily consist of amorphous-like aggregates, with fewer nanosheets of approximately 4.0 nm thickness (Fig. 3B and D). UV-Vis spectroscopy analysis of the assembled Pep1SP and Pep1MC materials reveals that the majority of the self-assembling peptoid conjugate is in SP and MC forms, respectively (Fig. 3E), which is further confirmed by UPLC characterization (Fig. S10A and B). Interestingly, as seen in Fig. S8, the surface roughness of the nanosheets calculated from the AFM images of assembled Pep1MC and Pep1SP materials shows a significantly higher root mean square (RMS) roughness value (Rq) in Pep1SP sheets (Rq = 1.680 nm) than Pep1MC sheets (Rq = 0.108 nm). This indicates considerably greater height variation or a rougher surface in Pep1SP nanosheets compared to Pep1MC nanomaterials, resulting in high crystallinity and highly ordered functional groups. Finally, the crystallinity of the assembled Pep1MC and Pep1SP materials was evaluated by synchrotron-based X-ray diffraction (XRD) measurements. XRD data of Pep1MC materials confirm the high crystallinity of Pep1MC nanosheets, as evidenced by very sharp and distinct peaks (Fig. 3F), which are like those previously observed in the case of assembled nanosheets from other Nbrpe6-containing sequences.29,36,46 Conversely, the XRD data of Pep1SP materials show a broad and diffuse pattern, suggesting low crystallinity (Fig. 3F). This experimental observation is further supported by periodic DFT calculations (See discussion below), which show that MC dimers are energetically more favorable than SP dimers for ordered packing due to their planar structure and favorable dipole alignment. To study how SP and MC moieties affect the assembly of these responsive peptoid conjugates, peptoid conjugates were self-assembled with a lower composition of MC. To achieve this, the Pep1SP solution was maintained at 4 °C in a dark environment, either without initial heating or after 2 hours of 365 nm long-wavelength UV irradiation. As no heat-promoted SP to MC transformation was involved in either case, the MC composition should be lower than that of the ideal heat-mediated transformation. AFM and SEM characterizations of these assembled samples show mainly amorphous-like structures with few nanosheet formations, albeit smaller in size than those of Pep1MC assembled nanosheets, indicating non-optimal assembly (Fig. S10). We postulate that this could be due to a comparatively lower composition of MC and the presence of the SP isomer in the Pep1 solution, as indicated by UPLC traces (Fig. S10C and D).
1) was heated at 60 °C for 20 minutes to facilitate the SP to MC transformation. A complete conversion of Pep1SP to Pep1MC was confirmed by UV-Vis spectroscopy. The resulting Pep1MC solution was then left at 4 °C in the dark to allow for slow evaporation, which triggered the self-assembly. Interestingly, a deep red, gel-like solution was obtained after about one week (Fig. S5). In contrast, the self-assembly of Pep1SP was much slower; a slightly yellowish, gel-like solution with slurry was obtained after at least two weeks of slow evaporation of the Pep1SP solution (Fig. S5). These results indicate that the MC form is much more favorable for the peptoid gelation (i.e., crystallization) than the SP form. In contrast, in the SP form, self-assembly of Pep1 into crystalline nanosheets is very challenging. To assess the morphology of assembled Pep1MC and Pep1SP materials, atomic force microscopy (AFM), scanning electron microscopy (SEM), and transmission electron microscopy (TEM) were employed. Both AFM and SEM characterizations of the assembled Pep1MC materials (Fig. 3A and C) reveal the formation of 2D nanosheets with lateral dimensions exceeding 5.0 µm, straight edges, and a thickness of approximately 4.0 nm. The formation of such large 2D nanosheets was further confirmed by liquid-cell AFM and TEM characterizations (Fig. S7A and S7B). On the contrary, AFM and SEM results show that assembled Pep1SP materials primarily consist of amorphous-like aggregates, with fewer nanosheets of approximately 4.0 nm thickness (Fig. 3B and D). UV-Vis spectroscopy analysis of the assembled Pep1SP and Pep1MC materials reveals that the majority of the self-assembling peptoid conjugate is in SP and MC forms, respectively (Fig. 3E), which is further confirmed by UPLC characterization (Fig. S10A and B). Interestingly, as seen in Fig. S8, the surface roughness of the nanosheets calculated from the AFM images of assembled Pep1MC and Pep1SP materials shows a significantly higher root mean square (RMS) roughness value (Rq) in Pep1SP sheets (Rq = 1.680 nm) than Pep1MC sheets (Rq = 0.108 nm). This indicates considerably greater height variation or a rougher surface in Pep1SP nanosheets compared to Pep1MC nanomaterials, resulting in high crystallinity and highly ordered functional groups. Finally, the crystallinity of the assembled Pep1MC and Pep1SP materials was evaluated by synchrotron-based X-ray diffraction (XRD) measurements. XRD data of Pep1MC materials confirm the high crystallinity of Pep1MC nanosheets, as evidenced by very sharp and distinct peaks (Fig. 3F), which are like those previously observed in the case of assembled nanosheets from other Nbrpe6-containing sequences.29,36,46 Conversely, the XRD data of Pep1SP materials show a broad and diffuse pattern, suggesting low crystallinity (Fig. 3F). This experimental observation is further supported by periodic DFT calculations (See discussion below), which show that MC dimers are energetically more favorable than SP dimers for ordered packing due to their planar structure and favorable dipole alignment. To study how SP and MC moieties affect the assembly of these responsive peptoid conjugates, peptoid conjugates were self-assembled with a lower composition of MC. To achieve this, the Pep1SP solution was maintained at 4 °C in a dark environment, either without initial heating or after 2 hours of 365 nm long-wavelength UV irradiation. As no heat-promoted SP to MC transformation was involved in either case, the MC composition should be lower than that of the ideal heat-mediated transformation. AFM and SEM characterizations of these assembled samples show mainly amorphous-like structures with few nanosheet formations, albeit smaller in size than those of Pep1MC assembled nanosheets, indicating non-optimal assembly (Fig. S10). We postulate that this could be due to a comparatively lower composition of MC and the presence of the SP isomer in the Pep1 solution, as indicated by UPLC traces (Fig. S10C and D).
Encouraged by the effect of SP to MC transformation in nanosheet assembly, we further explored the influence of the reverse photochromism of SP on Pep2 containing six Nbrpm side chains (Fig. 2A). Typically, di-block-like peptoids containing six Nbrpm groups in the hydrophobic domain form 1D nanotubes. Pep2SP and Pep2MC were self-assembled in the same conditions as Pep1SP and Pep1MC. Similar to the assembly of sheet materials, a deep red, gel-like solution was obtained from Pep2MC after only one week of self-assembly. In contrast, a solution of aggregates and slurries was obtained from Pep2SP after at least three weeks of slow evaporation. Ex situ AFM characterization of Pep2MC gel-like materials reveals the formation of 1D nanotubes with a tube height of around 6.5 nm (Fig. 4A). Formation of uniform 1D nanotubes was further confirmed by liquid cell AFM (Fig. S6b) and SEM (Fig. 4C). Negatively stained transmission electron microscopy (TEM) images revealed that Pep2MC nanotubes have a wall thickness of 3.8 ± 0.6 nm and an average tube diameter of 30.9 ± 0.9 nm, similar to the previously reported tube thickness and average diameter of our peptoid nanotubes.27,30,38 On the other hand, AFM and SEM results indicate that Pep2SP forms a very few smaller, non-uniform nanotubes, with mostly amorphous-like aggregates (Fig. 4B and D). Finally, XRD analysis of the assembled Pep2MC materials unveils high crystallinity of the nanotubes, characterized by very sharp and distinct peaks (Fig. S11), similar to the characteristic peaks observed in other nanotubes assembled from peptoids containing six Nbrpm groups.27,30,38 On the contrary, the XRD data of Pep2SP assemblies reveal comparatively low crystallinity with broader peaks and a diffuse pattern (Fig. S11). To assess whether the SP to MC transformation induces similar effects on the assembly properties of amphiphilic peptoids lacking Nhis groups, Pep3 with six N-([2-(4-chlorophenyl)ethyl]glycine) (Nclpe) groups in the hydrophobic domain, and six N-(2-methoxyethyl)glycine (Nme) in the hydrophilic domain was synthesized (Fig. S3). Here, six Nme groups were used to assist with solubility. Like Pep1MC and Pep2MC, the assembly of the Pep3MC solution also resulted in the formation of crystalline nanosheets, as confirmed by AFM, SEM, and XRD results (Fig. S12). In contrast, amorphous materials were obtained from the Pep1SP (Fig. S12). These findings suggest that the transformation of bulky, sterically hindered, nonpolar SP moieties into polar and planar MC moieties, with MC–MC dipole–dipole and π-interactions, plays a key role in obtaining highly crystalline nanomaterials.
 | ||
| Fig. 4 Representative AFM images of assembled (A) Pep2MC and (B) Pep2SP. Representative SEM images of assembled (C) Pep2MC and (D) Pep2SP. | ||
Light-responsive disassembly of MC-conjugated peptoid nanomaterials
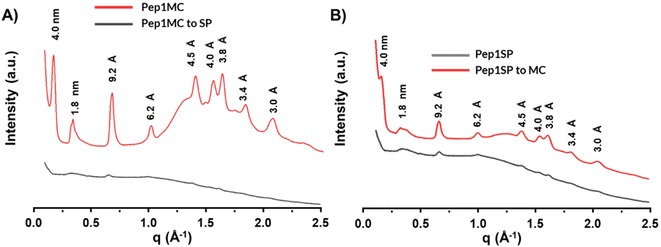
After exploring the heat- and light-responsive assembly properties of SP-conjugated peptoids, we were intrigued to see how crystalline nanomaterials assembled from MC-conjugated peptoids would behave upon exposure to visible light. When the assembled Pep1MC gel solution was exposed to ambient visible light for different durations, a change from deep red to yellowish color was observed. AFM characterization of gel-like materials after being exposed to varying periods of ambient visible light exposure reveals the disassembly of peptoid nanosheets into amorphous-like materials (Fig. 5A–E). UV-Vis spectra of the assembled Pep1MC materials after 96 h of visible light exposure reveal the disappearance of the characteristic MC peak, suggesting the complete conversion of Pep1MC into Pep1SP (Fig. S13). We hypothesize that the disassembly of Pep1MC nanosheets may be due to the disruption of MC–MC interactions, resulting in significant changes in physicochemical properties, such as steric hindrance, polarity, and solubility, within the assembled nanomaterials. Interestingly, the disassembly process can be accelerated upon exposure to warm white light using a photoreactor (Fig. 5F). Nonetheless, the MC to SP conversion in the assembled materials was significantly slower than in the Pep1MC monomer solution. We hypothesize that this is likely attributed to the significant energy barrier associated with disrupting MC–MC interactions within the crystalline peptoid assemblies, required for their conversion to the close-ring spiropyran form. Additionally, the SP to MC transformation can occur spontaneously through “negative photochromism” in the absence of light, particularly in polar solvents such as water.14,47 To evaluate that, when the aqueous solution of assembled Pep1SP materials was left in the absence of light for three weeks, a significantly high amount of nanosheets with nearly no amorphous-like materials was observed, as shown in the AFM characterization (Fig. S14). This result suggests that the conversion of Pep1SP to Pep1MC facilitates MC–MC interactions in a dark environment, thereby promoting the formation of nanosheets in aqueous solution. These findings were further confirmed by SEM results (Fig. S15). This light-responsive change in both assembly structures and crystallinity was further analyzed and confirmed by the XRD result (Fig. 6). | ||
| Fig. 5 Representative AFM images showing time-dependent change in crystallinity and assembly of Pep1MC, when exposed to ambient visible light (A–E), and warm white light (F). | ||
When the stability of Pep1MC nanosheets and Pep2MC nanotubes was studied in the absence of light at elevated temperatures or in organic solvents, it was found that both nanomaterials are stable and retain their nanostructures after heating or exposure to organic solvents (Fig. S16 and S17). These results suggest that MC-containing peptoid nanosheets and nanotubes, with MC functionality, retain their high stability in the absence of light, even at elevated temperatures or in the presence of excess organic solvents. These results are similar to those of our previously reported Peptoid assemblies, highlighting the unique advantages of these synthetic materials compared to biological systems.2,27,32 Exposure to visible light irradiation is necessary for the transformation of PepMC to PepSP, leading to the disassembly of the PepMC crystalline nanomaterials. To further confirm that the disruption of MC–MC interactions and a significant change in the physicochemical property of self-assembling PepMC are responsible for the disassembly of peptoid crystalline nanomaterials, Pep1MC was co-assembled with Pep4, an amphiphilic peptoid similar to Pep1SP but without MC conjugation. Co-assembly of Pep4 with Pep1MC in an 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 ratio, i.e., 80% Pep4 and 20% Pep1MC, resulted in the formation of crystalline nanosheets, as seen from AFM and SEM images (Fig. S18A and B) and SEM images (Fig. S18C and D). When these nanosheets were exposed to ambient visible light for 48 hours, no significant change in the assembly morphology was observed. We hypothesized that in the co-crystallization of Pep4
20 ratio, i.e., 80% Pep4 and 20% Pep1MC, resulted in the formation of crystalline nanosheets, as seen from AFM and SEM images (Fig. S18A and B) and SEM images (Fig. S18C and D). When these nanosheets were exposed to ambient visible light for 48 hours, no significant change in the assembly morphology was observed. We hypothesized that in the co-crystallization of Pep4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Pep1MC (80
Pep1MC (80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20), this process is primarily driven by hydrophobic interactions (e.g., π–π interactions) among hydrophobic Nbrpe side chains.40–45 Because this system contains only 20% Pep1MC, the low density of the MC group within the peptoid sheets makes MC–MC interactions unlikely.48 As a result, upon exposure to visible light, no substantial change in physicochemical properties can be observed, leading to the retention of assembled crystalline nanostructures.
20), this process is primarily driven by hydrophobic interactions (e.g., π–π interactions) among hydrophobic Nbrpe side chains.40–45 Because this system contains only 20% Pep1MC, the low density of the MC group within the peptoid sheets makes MC–MC interactions unlikely.48 As a result, upon exposure to visible light, no substantial change in physicochemical properties can be observed, leading to the retention of assembled crystalline nanostructures.
Computational analysis of SP and MC dimer packing energetics
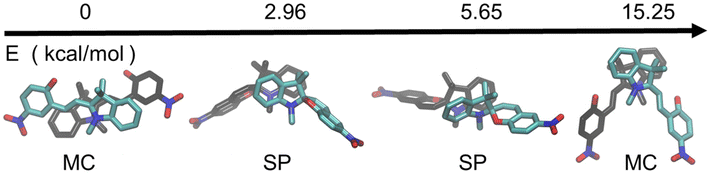
To investigate the molecular basis for the enhanced crystallinity observed in MC-functionalized peptoids, we conducted density functional theory (DFT) calculations on representative dimer structures of both the SP and MC forms. These dimers were selected to mimic backbone-backbone packing motifs relevant to the experimentally observed assembled nanostructures.For each isomer, the lowest-energy dimer conformations were used to construct periodic one-dimensional stacks with packing constraints that reflect the ∼0.45 nm spacing found in XRD data.29 Geometry optimizations were performed under periodic boundary conditions to assess the relative stabilities of these packing arrangements.
As shown in Fig. 7, MC dimers exhibit a significant energetic stabilization, approximately 3 kcal mol−1 lower in energy than their SP counterparts. This difference underscores the role of MC's planar, zwitterionic structure in promoting favorable π–π stacking and dipole–dipole interactions, which likely drive the formation of highly ordered nanosheets and nanotubes. In contrast, SP dimers are hindered by non-planarity and steric bulk, resulting in less favorable packing and reduced crystallinity.
To test whether this energetic preference persists in extended assemblies, molecular dynamics simulations were performed on fully solvated membrane models constructed from the two lowest-energy DFT-predicted SP and MC 1D packing motifs (see SI, Fig. S20). Using a customized peptoid force field, the MC membranes show a 10–15 kcal mol−1 per monomer stronger solute–solvent stabilization than their SP counterparts, consistent with the enhanced stability and crystallinity observed experimentally.
Together, the DFT and MD analyses provide a coherent mechanistic explanation for the experimental observations: SP-to-MC isomerization enhances the thermodynamic favorability of ordered assembly, directly linking molecular conformation to macroscopic morphology (see SI for details).
Stimuli-responsive peptoid crystalline nanomaterials as carbonic anhydrase (CA) mimics
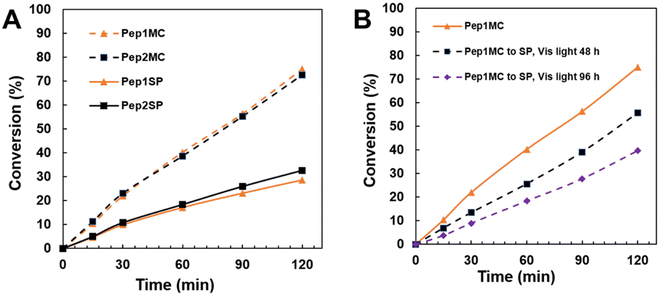
Our recent study showed that peptoid crystalline nanomaterials containing Nhis groups can be used as CA mimics for enhanced hydration and sequestration of CO2.35 Because the hydrophilic domains of Pep1 and Pep2 both contain Nhis groups, we hypothesize that the assembled, stimuli-responsive nanomaterials from Pep1MC can be used as CA mimics exhibiting responsive catalytic properties. The high catalytic efficiency and specificity found in natural enzymes are from their ability to fold into well-ordered active site conformations through self-assembly.34,49,50 We speculate that the assembly of highly crystalline Pep1MC nanosheets and Pep2MC nanotubes, respectively, which have a high density of ordered functional imidazole groups, will exhibit much higher efficiency as CA mimics compared to those assembled from Pep1SP or Pep2SP, which exhibit significantly lower crystallinity. To evaluate this, we employed a colorimetric assay that involved the hydrolysis of 4-nitrophenyl acetate (4-NPA) to 4-nitrophenol (4-NP) at room temperature. This reaction has been commonly used to evaluate the efficiency of CA mimics.35 By monitoring the UV-Vis absorbance of 4-NP at 400 nm, the hydrolysis rate of the peptoid assemblies was determined. As shown in Fig. 8A, both Pep1MC and Pep2MC crystalline assemblies demonstrated significantly higher activity in the hydrolysis of 4-NPA compared to their SP-containing counterparts. These notable differences in hydrolysis efficiency can be attributed to the high crystallinity of the MC-containing peptoid assemblies, which contain highly ordered imidazole groups, thereby enhancing ester hydrolysis activity. This is consistent with the improved packing stability of MC dimers predicted by DFT. These findings align with our recent reports, in which peptoid-based highly crystalline nanosheets and nanotubes demonstrated significantly higher activity as biomimetic catalysts compared to their amorphous counterparts.26,34,35,46 No significant difference was observed for the effect of assembly morphology on the catalytic activity, as both assembled Pep1MC and Pep2MC nanomaterials exhibited similar hydrolysis activities. To further illustrate the significance of crystallinity and the organized arrangement of active groups for catalytic activity, Pep1MC was exposed to ambient visible light for varying durations. This exposure resulted in a reduction in crystallinity and the disassembly of the tightly packed nanostructure as observed in Fig. 5. Subsequent hydrolysis tests with these altered samples revealed a decrease in their hydrolysis activity, which is likely due to the disruption of the crystalline assemblies (Fig. 8B). Pseudo rate constants derived from the hydrolysis rate also reveal superior activity from the MC-containing peptoid assemblies compared to the SP-containing assemblies (Table S1). Interestingly, both assembled Pep1MC and Pep2MC samples collected after the completion of the catalytic hydrolysis reactions retained their assembly morphology (Fig. S19), highlighting the excellent stability of these peptoid nanostructures. We expect that this stimuli-responsive crystalline peptoid assembly system could be used as a platform for the development of efficient CA mimics for enhanced hydration and sequestration of CO2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 35 and for the development of other enzyme mimics.
35 and for the development of other enzyme mimics.
Conclusions
In conclusion, we report the use of the conversion of spiropyran (SP) into merocyanine (MC) for the development of multi-stimuli responsive nanomaterials. We found that heat-mediated SP to MC transformation is highly efficient, and this property can be utilized for the tunable assembly properties of SP and MC functionalized peptoids. When peptoids were self-assembled in MC form, highly crystalline nanomaterials were obtained, whereas with SP form, amorphous-like materials were observed. MC peptoid nanomaterials showed excellent stability in the absence of light; only upon exposure to visible light, disassembly of the crystalline nanostructures was observed due to the transformation into SP moieties. Periodic DFT calculations on SP and MC dimers further support these findings by revealing significantly more favorable packing energetics for MC, consistent with its enhanced crystallinity and assembly behavior observed experimentally. Finally, we demonstrated that the catalytic activity in ester hydrolysis can be tuned by the stimuli-responsive, reversible control of the crystallinity of SP- and MC-functionalized assembled nanomaterials. We envision that the functionalization of SP moieties into peptoids for stimuli-responsive assembly and disassembly properties will inspire a wide range of applications, including biomedical technology and biomimetic crystallization.Methods
Materials
All materials were sourced from commercial suppliers and were used as received unless otherwise indicated.Peptoid synthesis and self-assembly
The synthesis of all peptoid sequences was performed on rink amide resins, utilizing either manual or automated solid-phase synthesis methods detailed in our earlier publications, unless otherwise specified.26,27 The specific synthesis protocols for each peptoid can be found in the SI. To cleave the synthesized peptoids from the resins, they were treated with 95% (v/v) trifluoroacetic acid (TFA) in water in a cartridge for one hour with agitation. The solution was then collected by filtration, and TFA was removed by evaporation under reduced pressure. After TFA evaporation, the crude samples were completely dissolved in an acetonitrile/water mixture and purified using reverse-phase high-performance liquid chromatography (HPLC) set up with an XBridgeTM Prep C18 OBDTM column, 10 μm, 19 mm × 100 with a linear gradient of 60–80% acetonitrile in water with 0.1% TFA. The collected fractions were analyzed with a UPLC (Waters, ACQUITY reverse-phase at 0.4 ml min−1 over 7 min at 40 °C with an ACQUITYBEH C18, 17 μm, and 2.1 mm × 50 mm column) connected with the MS system (Waters SQD2). The collected pure fractions were lyophilized to yield a slightly yellowish powder.For peptoid self-assembly, one µmol of the corresponding lyophilized peptoid was dissolved in 200 µL of a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 acetonitrile/water mixture, resulting in a slightly yellowish, clear solution of 5.0 mM concentration. The resulting SP-peptoid was allowed to sit at 4 °C for slow evaporation in the presence of ambient light, facilitating self-assembly. In the case of MC-peptoids, SP-peptoids were transformed into MC-peptoids by heating them at 60 °C for 20 minutes in the absence of light, resulting in a deep red solution. MC-peptoids were also self-assembled by initially exposing them to 365 nm UV light for one hour or by leaving them in a dark environment without heating. The resulting MC-peptoid solution was left at 4 °C in the absence of light to promote the slow evaporation of acetonitrile and facilitate self-assembly.
1 acetonitrile/water mixture, resulting in a slightly yellowish, clear solution of 5.0 mM concentration. The resulting SP-peptoid was allowed to sit at 4 °C for slow evaporation in the presence of ambient light, facilitating self-assembly. In the case of MC-peptoids, SP-peptoids were transformed into MC-peptoids by heating them at 60 °C for 20 minutes in the absence of light, resulting in a deep red solution. MC-peptoids were also self-assembled by initially exposing them to 365 nm UV light for one hour or by leaving them in a dark environment without heating. The resulting MC-peptoid solution was left at 4 °C in the absence of light to promote the slow evaporation of acetonitrile and facilitate self-assembly.
Characterizations
In situ AFM imaging was performed using silicon probes (SNL-C, spring constant: 0.24 N m−1, tip radius: 2 nm; Bruker) in tapping mode on a Cypher ES AFM (Asylum Research). Peptoid assembly solutions were diluted with deionized water, and 80 μL of the diluted solution was deposited onto a freshly cleaved mica surface at room temperature. Image processing and analysis were conducted using Gwyddion SPM data analysis software.
Computational methods
Relative total energies were used to compare the stability of MC and SP packing motifs under equivalent conditions. Energetic trends were verified by scanning the inter-dimer spacing along the x-axis to confirm a local minimum at approximately 0.45 nm.
Author contributions
C. L. C. directed the project. P. C. conducted all the experimental study and characterization except as mentioned otherwise. B. S. H. and M. D. B. conducted the atomistic modelling. C. L. C. conducted XRD data collection of the peptoid assemblies. P. C. and C-L. C. analyzed the XRD data. C. S. assisted with in situ AFM characterization. B. H. conducted the TEM characterizations. K. W. assisted with NMR collection and analysis. P. C., B. S. H., M. D. B., and C. L. C. wrote the manuscript. All authors discussed the results and commented on the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting the findings of this study are available within the article and its supplementary information (SI). All data are also available upon reasonable request from the corresponding author. Supplementary information is available. See DOI: https://doi.org/10.1039/d5nr03491d.Acknowledgements
The development of peptoid nanosheets and DFT calculations were supported by the Department of Energy (DOE), Office of Basic Energy Sciences (BES), Division of Materials Science and Engineering, Biomolecular Materials Program under award FWP-65357 at Pacific Northwest National Laboratory (PNNL). The development of responsive peptoid materials was supported by DOE-BES as part of the Energy Frontier Research Centers program: CSSAS – The Center for the Science of Synthesis Across Scales – under Award Number DE-SC0019288 [FWP 72448 at PNNL]. The carbonic anhydrase mimic study was supported by the DOE-BES, Division of Materials Science and Engineering, Biomolecular Materials Program under an award FWP 80124 at PNNL. The force field development and simulation were supported by the DOE-BES, Division of Materials Science and Engineering, Biomolecular Materials Program, under an award FWP 77876 at PNNL. Work at the Molecular Foundry and X-ray diffraction at the Advanced Light Source (ALS) at Lawrence Berkeley National Laboratory (LBNL) was supported by the Office of Science, Office of Basic Energy Sciences, of the U.S. DOE under Contract DE-AC02-05CH11231. This research used resources of the National Energy Research Scientific Computing Center (NERSC), a DOE Office of Science User Facility located at LBNL, operated under Contract No. DE-AC02-05CH11231 using NERSC award BES-ERCAP0024465. PNNL is a multi-program national laboratory operated for the Department of Energy by Battelle under Contract No. DE-AC05-76RL01830. P. C. acknowledges Kennesaw State Academic Affairs for support of the NMR facility.References
- L. Shao, J. Ma, J. L. Prelesnik, Y. Zhou, M. Nguyen, M. Zhao, S. A. Jenekhe, S. V. Kalinin, A. L. Ferguson, J. Pfaendtner, C. J. Mundy, J. J. De Yoreo, F. Baneyx and C.-L. Chen, Chem. Rev., 2022, 122, 17397–17478 CrossRef CAS PubMed.
- Z. Li, B. Cai, W. Yang and C.-L. Chen, Chem. Rev., 2021, 121, 14031–14087 CrossRef CAS PubMed.
- J. Zhu, N. Avakyan, A. Kakkis, A. M. Hoffnagle, K. Han, Y. Li, Z. Zhang, T. S. Choi, Y. Na, C.-J. Yu and F. A. Tezcan, Chem. Rev., 2021, 121, 13701–13796 CrossRef CAS PubMed.
- S. Fleming and R. V. Ulijn, Chem. Soc. Rev., 2014, 43, 8150–8177 RSC.
- A. Levin, T. A. Hakala, L. Schnaider, G. J. L. Bernardes, E. Gazit and T. P. J. Knowles, Nat. Rev. Chem., 2020, 4, 615–634 CrossRef CAS.
- L. Shao, D. Hu, S.-L. Zheng, T. K. H. Trinh, W. Zhou, H. Wang, Y. Zong, C. Li and C.-L. Chen, Angew. Chem., Int. Ed., 2024, 63, e202403263 CrossRef CAS PubMed.
- L.-J. Chen and H.-B. Yang, Acc. Chem. Res., 2018, 51, 2699–2710 CrossRef CAS PubMed.
- R. J. Mart, R. D. Osborne, M. M. Stevens and R. V. Ulijn, Soft Matter, 2006, 2, 822–835 RSC.
- S. Panja and D. J. Adams, Chem. Soc. Rev., 2021, 50, 5165–5200 RSC.
- M. P. Hendricks, K. Sato, L. C. Palmer and S. I. Stupp, Acc. Chem. Res., 2017, 50, 2440–2448 CrossRef CAS PubMed.
- J.-F. Gohy and Y. Zhao, Chem. Soc. Rev., 2013, 42, 7117–7129 RSC.
- K. Sun, Q.-Y. Lv, X.-L. Chen, L.-B. Qu and B. Yu, Green Chem., 2021, 23, 232–248 RSC.
- K. Tashiro, M. Otori and S. Satokawa, Chem. Commun., 2023, 59, 4304–4307 RSC.
- R. Klajn, Chem. Soc. Rev., 2014, 43, 148–184 RSC.
- L. Kortekaas and W. R. Browne, Chem. Soc. Rev., 2019, 48, 3406–3424 RSC.
- H. Görner and A. K. Chibisov, J. Chem. Soc., Faraday Trans., 1998, 94, 2557–2564 RSC.
- A. M. Alfaraidi, B. Kudisch, N. Ni, J. Thomas, T. Y. George, K. Rajabimoghadam, H. J. Jiang, D. G. Nocera, M. J. Aziz and R. Y. Liu, J. Am. Chem. Soc., 2023, 145, 26720–26727 CrossRef CAS PubMed.
- A. de Vries, K. Goloviznina, M. Reiter, M. Salanne and M. R. Lukatskaya, Chem. Mater., 2024, 36, 1308–1317 CrossRef CAS PubMed.
- J. Sheng, J. Perego, S. Bracco, W. Czepa, W. Danowski, S. Krause, P. Sozzani, A. Ciesielski, A. Comotti and B. L. Feringa, Adv. Mater., 2024, 36, 2305783 CrossRef CAS PubMed.
- G. Das, T. Prakasam, N. Alkhatib, R. G. AbdulHalim, F. Chandra, S. K. Sharma, B. Garai, S. Varghese, M. A. Addicoat, F. Ravaux, R. Pasricha, R. Jagannathan, N. i. Saleh, S. Kirmizialtin, M. A. Olson and A. Trabolsi, Nat. Commun., 2023, 14, 3765 CrossRef CAS PubMed.
- H. Inaba, M. Sakaguchi, S. Watari, S. Ogawa, A. M. R. Kabir, A. Kakugo, K. Sada and K. Matsuura, ChemBioChem, 2023, 24, e202200782 CrossRef CAS.
- W. Wang, J. Hu, M. Zheng, L. Zheng, H. Wang and Y. Zhang, Org. Biomol. Chem., 2015, 13, 11492–11498 RSC.
- J. Sun and R. N. Zuckermann, ACS Nano, 2013, 7, 4715–4732 CrossRef CAS PubMed.
- N. Gangloff, J. Ulbricht, T. Lorson, H. Schlaad and R. Luxenhofer, Chem. Rev., 2016, 116, 1753–1802 CrossRef CAS PubMed.
- B. Cai, Z. Li and C.-L. Chen, Acc. Chem. Res., 2021, 54, 81–91 CrossRef CAS PubMed.
- T. Jian, Y. Zhou, P. Wang, W. Yang, P. Mu, X. Zhang, X. Zhang and C.-L. Chen, Nat. Commun., 2022, 13, 3025 CrossRef CAS PubMed.
- H. Jin, Y.-H. Ding, M. Wang, Y. Song, Z. Liao, C. J. Newcomb, X. Wu, X.-Q. Tang, Z. Li, Y. Lin, F. Yan, T. Jian, P. Mu and C.-L. Chen, Nat. Commun., 2018, 9, 270 CrossRef PubMed.
- Y. Li, G. Yang, L. Gerstweiler, S. H. Thang and C.-X. Zhao, Adv. Funct. Mater., 2023, 33, 2210387 CrossRef CAS.
- J. A. Hammons, M. D. Baer, T. Jian, J. R. I. Lee, T. M. Weiss, J. J. De Yoreo, A. Noy, C.-L. Chen and A. Van Buuren, J. Phys. Chem. Lett., 2021, 12, 6126–6133 CrossRef CAS PubMed.
- Y. Luo, Y. Song, M. Wang, T. Jian, S. Ding, P. Mu, Z. Liao, Q. Shi, X. Cai, H. Jin, D. Du, W.-J. Dong, C.-L. Chen and Y. Lin, Small, 2019, 15, 1902485 CrossRef CAS PubMed.
- F. Jiao, X. Wu, T. Jian, S. Zhang, H. Jin, P. He, C.-L. Chen and J. J. De Yoreo, Angew. Chem., Int. Ed., 2019, 58, 12223–12230 CrossRef CAS PubMed.
- H. Jin, F. Jiao, M. D. Daily, Y. Chen, F. Yan, Y.-H. Ding, X. Zhang, E. J. Robertson, M. D. Baer and C.-L. Chen, Nat. Commun., 2016, 7, 12252 CrossRef CAS PubMed.
- F. Jiao, Y. Chen, H. Jin, P. He, C.-L. Chen and J. J. De Yoreo, Adv. Funct. Mater., 2016, 8960–8967, DOI:10.1002/adfm.201602365.
- T. K. H. Trinh, B. Jin, R. N. Zuckermann and C.-L. Chen, ACS Appl. Mater. Interfaces, 2025, 17, 44347–44359 CAS.
- P. Chakma, Y. Chen, B. S. Harris, Y. W. Elhady, R. Zheng, M. E. Bowden, V. Shutthanandan, A. B. Bard, T. K. H. Trinh, X. Zheng, C. J. Mundy, M. D. Baer and C.-L. Chen, Nat. Commun., 2025, 16, 7348 CrossRef CAS PubMed.
- J. Ma, B. Cai, S. Zhang, T. Jian, J. J. De Yoreo, C.-L. Chen and F. Baneyx, Nano Lett., 2021, 21, 1636–1642 CrossRef CAS PubMed.
- H. Du, H. Trinh, O. C. Brandon, R. Zheng, H. Wang, K. Corry, T. R. Wood, C.-L. Chen and E. Nance, bioRxiv, 2025, 2025.2003.2006.641448. DOI:10.1101/2025.03.06.641448.
- Y. Song, X. Cai, M. Wang, D. Du, Y. Lin and C.-L. Chen, Nano Res., 2024, 17, 788–796 CrossRef CAS.
- J. Ma, B. Jin, K. N. Guye, M. E. Chowdhury, N. Y. Naser, C.-L. Chen, J. J. De Yoreo and F. Baneyx, Adv. Mater., 2023, 35, 2207543 CrossRef CAS PubMed.
- C. L. Chen and A. M. Beatty, J. Am. Chem. Soc., 2008, 130, 17222–17223 CrossRef CAS.
- C. L. Chen, H. Y. Tan, J. H. Yao, Y. Q. Wan and C. Y. Su, Inorg. Chem., 2005, 44, 8510–8520 CrossRef CAS PubMed.
- C. L. Chen, A. M. Goforth, M. D. Smith, C. Y. Su and H. C. zur Loye, Angew. Chem., Int. Ed., 2005, 44, 6673–6677 CrossRef CAS PubMed.
- C. L. Chen, C. Y. Su, Y. P. Cai, H. X. Zhang, A. W. Xu, B. S. Kang and H. C. zur Loye, Inorg. Chem., 2003, 42, 3738–3750 CrossRef CAS PubMed.
- C. L. Chen, C. Y. Su, Y. P. Cai, H. X. Zhang, A. W. Xu and B. S. Kang, New J. Chem., 2003, 27, 790–792 RSC.
- C. L. Chen, J. Y. Zhang and C. Y. Su, Eur. J. Inorg. Chem., 2007, 2997–3010, DOI:10.1002/ejic.200700187.
- T. K. H. Trinh, T. Jian, B. Jin, D.-T. Nguyen, R. N. Zuckermann and C.-L. Chen, ACS Appl. Mater. Interfaces, 2023, 15, 51191–51203 CrossRef CAS PubMed.
- J. Zhou, Y. Li, Y. Tang, F. Zhao, X. Song and E. Li, J. Photochem. Photobiol., A, 1995, 90, 117–123 CrossRef CAS.
- Y. Song, M. Wang, S. Akkineni, W. Yang, J. J. Hettige, H. Jin, Z. Liao, P. Mu, F. Yan, M. Baer, J. J. De Yoreo, D. Du, Y. Lin and C.-L. Chen, ACS Mater. Lett., 2021, 3, 420–427 CrossRef CAS.
- M. Raynal, P. Ballester, A. Vidal-Ferran and P. W. N. M. van Leeuwen, Chem. Soc. Rev., 2014, 43, 1734–1787 RSC.
- A. R. Hirst, S. Roy, M. Arora, A. K. Das, N. Hodson, P. Murray, S. Marshall, N. Javid, J. Sefcik, J. Boekhoven, J. H. van Esch, S. Santabarbara, N. T. Hunt and R. V. Ulijn, Nat. Chem., 2010, 2, 1089–1094 CrossRef CAS PubMed.
- D. Weininger, J. Chem. Inf. Comput. Sci., 1988, 28, 31–36 CrossRef CAS.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 CrossRef CAS PubMed.
- J. Zhang and M. Dolg, Phys. Chem. Chem. Phys., 2015, 17, 24173–24181 RSC.
- J. Hutter, M. Iannuzzi, F. Schiffmann and J. VandeVondele, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2014, 4, 15–25 CAS.
- Y. Zhang and W. Yang, Phys. Rev. Lett., 1998, 80, 890–890 CrossRef CAS.
- S. Grimme, J. Comput. Chem., 2006, 27, 1787–1799 CrossRef CAS PubMed.
- J. VandeVondele and J. Hutter, J. Chem. Phys., 2007, 127, 114105 CrossRef PubMed.
- S. Goedecker, M. Teter and J. Hutter, Phys. Rev. B: Condens. Matter Mater. Phys., 1996, 54, 1703–1710 CrossRef CAS PubMed.
- B. S. Harris, K. K. Bejagam and M. D. Baer, J. Phys. Chem. B, 2023, 127, 6573–6584 CrossRef CAS PubMed.
- Y. W. Elhady, B. S. Harris, C. J. Mundy and M. D. Baer, J. Phys. Chem. B, 2025, 129, 5901–5912 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2025 |