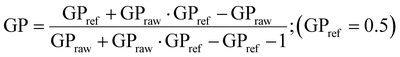Open Access Article
Open Access ArticleNucleolin perturbation alters membrane lipid homeostasis
Eitan Erez
Zahavi
a,
Ida
Rishal
a,
Juan A.
Oses-Prieto
 c,
Alexander
Brandis
d,
Sergey
Malitsky
d,
Maxim
Itkin
c,
Alexander
Brandis
d,
Sergey
Malitsky
d,
Maxim
Itkin
 d,
Šárka
Pokorná†
e,
Florencia
Cabrera-Cabrera
b,
Natjan-Naatan
Seeba
b,
Robert
Risti
d,
Šárka
Pokorná†
e,
Florencia
Cabrera-Cabrera
b,
Natjan-Naatan
Seeba
b,
Robert
Risti
 b,
Aivar
Lõokene
b,
Anthony H.
Futerman
b,
Aivar
Lõokene
b,
Anthony H.
Futerman
 e,
Alma L.
Burlingame
c,
Mike
Fainzilber
e,
Alma L.
Burlingame
c,
Mike
Fainzilber
 a and
Indrek
Koppel
a and
Indrek
Koppel
 *ab
*ab
aDepartments of Biomolecular Sciences and Molecular Neuroscience, Weizmann Institute of Science, 7610001 Rehovot, Israel. E-mail: indrek.koppel@taltech.ee
bDepartment of Chemistry and Biotechnology, Tallinn University of Technology, Akadeemia tee 15-234, Tallinn, Estonia
cDepartment of Pharmaceutical Chemistry, University of California, San Francisco, CA 94158, USA
dLife Science Core Facilities, Weizmann Institute of Science, 7610001 Rehovot, Israel
eDepartment of Biomolecular Sciences, Weizmann Institute of Science, 7610001 Rehovot, Israel
First published on 28th October 2025
Abstract
AS1411 is a G-rich DNA aptamer that targets the multifunctional RNA-binding protein nucleolin. AS1411 has both antiproliferative and cell size-regulating activities and has been evaluated for clinical utility, reaching phase II trials as an anticancer agent. The mechanisms underlying cell size effects of AS1411 are not well understood and broad characterization of its molecular effects is lacking. Here, we used a multi-omics approach to profile transcriptome, proteome and lipidome changes in AS1411-treated NIH-3T3 cells, which increase in size in response to the aptamer. We found that AS1411 caused downregulation of cholesterol biosynthesis pathway enzymes at both mRNA and protein levels, without an accompanying reduction in cellular cholesterol levels or cholesterol uptake. In addition, AS1411 induced changes in several lipid classes, including increases in phosphatidylethanolamine levels. Ratiometric imaging of Di-4-ANEPPS-labeled cells showed that AS1411 decreases the fluidity of intracellular membranes. Thus, aptamer engagement of nucleolin affects lipid biosynthesis and homeostasis, likely contributing to its roles in cell size control.
1. Introduction
Nucleolin is a ubiquitous RNA-binding protein (RBP) in eukaryotic cells, with roles in many cellular processes, including regulation of cell division, viability, and size.1–3 The earliest identified molecular functions of nucleolin were associated with its nucleolar localization, where it regulates transcription of rRNA and ribosome maturation.4 Nucleolin was also shown to have a role in polymerase II transcription, posttranscriptional regulation, and chromatin remodeling.1,2,5 In addition to being one of the most abundant nucleolar proteins, it is also shuttled to the cytoplasm and the plasma membrane.6–8 Overexpression and enhanced plasma membrane localization of nucleolin has been reported in several cancers,1,2,9 offering a target for antitumor therapies.9,10 AS1411 is a nucleolin-binding 26-mer G-rich DNA aptamer with antiproliferative effects in a broad range of tumor cells, and has shown limited efficacy in clinical trials for acute myeloid leukemia and renal cell carcinoma.10,11We have recently identified a key role for nucleolin in neuronal mRNA transport complexes, where it acts as an adapter between axonal kinesin motors and mRNA cargo.12–14 We have further suggested that cytoplasmic and axonal nucleolin functions in cell size and neuron growth regulation.12,13,15,16 The AS1411 aptamer interferes with the formation of a nucleolin–kinesin complex for axonal mRNA transport,13 by binding to the C-terminal glycine arginine-rich (GAR) domain of nucleolin.12 How this mechanism might be linked to size and growth-promoting effects of AS1411 remains unclear. Several nucleolin-dependent and nucleolin-independent mechanisms for AS1411 effects have been proposed, including cell cycle arrest, Bcl-2 inhibition, interference with EGFR signaling and hyperactivated macropinocytosis; however its precise mechanism of action remains elusive.10
Here, we conducted a multi-omics investigation to analyze the molecular pathways affected by AS1411, focusing our analyses on a cell type that responds with specific size changes upon exposure to AS1411. We combined RNA-seq transcriptomics, SILAC proteomics and lipidomics analyses to this end. Unexpectedly, these analyses revealed broad effects on cellular lipids and lipid biosynthesis pathways.
2. Experimental section
2.1 Cell culture and AS1411 treatment
NIH-3T3 cells (mouse embryonic fibroblasts, ATCC CRL-1658) and HeLa cells (HeLa 229, ATCC CCL-2.1) were grown at 37 °C and 5% CO2 in DMEM (Gibco), supplemented with 10% fetal bovine serum (Gibco), 100 U mL−1 penicillin and 100 μg mL−1 streptomycin (Biological Industries). Cells were seeded at densities of 2000 cells per cm2 (NIH-3T3) or 3000 cells per cm2 (HeLa). AS1411 (GGTGGTGGTGGTTGTGGTGGTGGTGG) and control DNA (CCTCCTCCTCCTTCTCCTCCTCCTCC) oligonucleotides (Integrated DNA Technologies) were dissolved in culture medium (vehicle) as 10× stock solutions. Unsupplemented DMEM was used as a vehicle in treatments (10% of total medium volume).2.2 Cell volume and proliferation measurements
For cell volume measurements, cells were seeded at a density of 3000 cm−2 on 6-well plates and treated with AS1411 or controls as indicated. Cells were detached from plates with 200 μL 0.25% trypsin – 0.05% EDTA (Biological Industries), resuspended in 2 mL growth medium, washed and resuspended in 1 mL ice-cold PBS with propidium iodide and passed through a cell strainer to obtain a single-cell suspension. Cell volume was measured by forward scatter (FSC-A) using the LSR II flow cytometer (BD Biosciences), excluding dead cells by gating out the population with high propidium iodide signal. Cell volumes are expressed as FCS-A readings relative to vehicle-treated controls. For cell proliferation analysis, the MTT assay kit (ab211091, Abcam) was used according to the manufacturer's instructions.2.3 SILAC proteomics
NIH-3T3 cells were cultivated in Stable Isotope Labeling by Amino acids in cell Culture (SILAC) DMEM medium (Thermo Fisher Scientific) supplemented with 10% dialyzed fetal calf serum (Biological Industries), 100 U mL−1 penicillin and 100 μg mL−1 streptomycin, containing isotopically labelled “heavy” 13C6-lysine (0.8 mM) and 13C6-arginine (0.4 mM) (Cambridge Isotope Laboratories). Control cells were grown in SILAC DMEM medium supplemented with the same concentrations of “light” (with natural 13C abundances) lysine and arginine. Cell cultures were maintained at 37 °C and 5% CO2 and collected after at least 5 doubling times. Cells were treated with 10 μM AS1411 or control aptamer for 24 or 48 h, rinsed with ice-cold PBS 3 times, collected by gentle scraping, transferred in PBS to 1.5 mL low protein binding tubes, pelleted at 200 g and supernatant carefully discarded, then cell pellets were flash frozen in liquid nitrogen and kept at −80 °C until digestion. Cell pellets were thawed and homogenized in 200 μl of 8 M urea on ice, using a probe sonicator. Samples were centrifuged at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 10 min, and supernatants recovered. The total protein content in the extracts was determined utilizing a Micro BCA Protein Assay Kit (Thermo Scientific). Equal amounts (200 μg) of heavy and light-labelled samples (AS1411 treated and untreated, respectively) were mixed, and the combined sample added ammonium bicarbonate to a concentration of 80 mM, then treated with 8.8 mM DTT at 56 °C for 10 min, followed by a 30-min incubation at room temperature in the dark with 15 mM iodoacetamide. For tryptic digestion, the samples were then diluted 4-fold with 100 mM ammonium bicarbonate to reduce urea concentration to 2 M and then added 5% (W/W) modified trypsin (Promega, Madison, WI). The pH was adjusted to 8.0 with 250 mM ammonium bicarbonate, and the samples were incubated for 12 h at 37 °C. After that, another aliquot of trypsin was added (2% W/W) and digested for an additional 6 hours. After this, samples were acidified with formic acid to a final concentration of 5%. The digests were then desalted using a MAX-RP Sep Pak® classic C18 cartridge (Waters) following the manufacturer's protocol. Sep Pak eluates were dried-evaporated in preparation for chromatographic fractionation. Digested samples were fractionated on an AKTA purifier system utilizing a Phenomenex Gemini 5u C18 110A 150 × 4.60 mm column, operating at a flow rate of 0.550 mL min−1. Buffer A consisted of 20 mM ammonium formate (pH 10), and buffer B consisted of 20 mM ammonium formate in 90% acetonitrile (pH 10). Gradient details were as follows: 1% to 9% B in 14 min, 9% B to 49% B in 4 min, 49% B to 70% B in 36 min, 70% B back down to 1% B in 3 min. 50 peptide-containing fractions were collected, evaporated and resuspended in 0.1% formic acid. Aliquots (containing around 3 μg of digested material) of 8 non-consecutive chromatographic fractions were run onto a 2 μm 75 μm × 50 cm PepMap RSLC C18 EasySpray column (Thermo Scientific). 3 h MeCN gradients (2–30% in 0.1% formic acid) were used to elute peptides, at a flow rate of 200 nL min−1, for analysis in a QExactive Plus (Thermo Scientific) in positive ion mode. MS spectra were acquired between 350 and 1500 m/z with a resolution of 70
000g for 10 min, and supernatants recovered. The total protein content in the extracts was determined utilizing a Micro BCA Protein Assay Kit (Thermo Scientific). Equal amounts (200 μg) of heavy and light-labelled samples (AS1411 treated and untreated, respectively) were mixed, and the combined sample added ammonium bicarbonate to a concentration of 80 mM, then treated with 8.8 mM DTT at 56 °C for 10 min, followed by a 30-min incubation at room temperature in the dark with 15 mM iodoacetamide. For tryptic digestion, the samples were then diluted 4-fold with 100 mM ammonium bicarbonate to reduce urea concentration to 2 M and then added 5% (W/W) modified trypsin (Promega, Madison, WI). The pH was adjusted to 8.0 with 250 mM ammonium bicarbonate, and the samples were incubated for 12 h at 37 °C. After that, another aliquot of trypsin was added (2% W/W) and digested for an additional 6 hours. After this, samples were acidified with formic acid to a final concentration of 5%. The digests were then desalted using a MAX-RP Sep Pak® classic C18 cartridge (Waters) following the manufacturer's protocol. Sep Pak eluates were dried-evaporated in preparation for chromatographic fractionation. Digested samples were fractionated on an AKTA purifier system utilizing a Phenomenex Gemini 5u C18 110A 150 × 4.60 mm column, operating at a flow rate of 0.550 mL min−1. Buffer A consisted of 20 mM ammonium formate (pH 10), and buffer B consisted of 20 mM ammonium formate in 90% acetonitrile (pH 10). Gradient details were as follows: 1% to 9% B in 14 min, 9% B to 49% B in 4 min, 49% B to 70% B in 36 min, 70% B back down to 1% B in 3 min. 50 peptide-containing fractions were collected, evaporated and resuspended in 0.1% formic acid. Aliquots (containing around 3 μg of digested material) of 8 non-consecutive chromatographic fractions were run onto a 2 μm 75 μm × 50 cm PepMap RSLC C18 EasySpray column (Thermo Scientific). 3 h MeCN gradients (2–30% in 0.1% formic acid) were used to elute peptides, at a flow rate of 200 nL min−1, for analysis in a QExactive Plus (Thermo Scientific) in positive ion mode. MS spectra were acquired between 350 and 1500 m/z with a resolution of 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. For each MS spectrum, up to 10 multiply charged ions over the selected threshold (1.7E4) were selected for MSMS with an isolation window of 4 m/z. Precursor ions were fragmented by HCD using a normalized collision energy of 25. MSMS spectra were acquired in centroid mode with a resolution of 17
000. For each MS spectrum, up to 10 multiply charged ions over the selected threshold (1.7E4) were selected for MSMS with an isolation window of 4 m/z. Precursor ions were fragmented by HCD using a normalized collision energy of 25. MSMS spectra were acquired in centroid mode with a resolution of 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 from m/z = 10. A dynamic exclusion window was applied which prevented the same m/z from being selected for 10 s after its acquisition. Peak lists were generated using PAVA in-house software.17 All generated peak lists were searched against the mouse subset of the UniProtKB 2013.6.17 database, using Protein Prospector18 with the following parameters: enzyme specificity was set as Trypsin, and up to 2 missed cleavages per peptide were allowed. Carbamidomethylation of cysteine residues was allowed as a fixed modification. N-Acetylation of the N-terminus of the protein, loss of protein N-terminal methionine, pyroglutamate formation from of peptide N-terminal glutamines, oxidation of methionine and 13C(6) labelling in lysine or arginine were allowed as variable modifications. Search space was limited with the “linked” option (6C12K linked to 6C12R, and 6C13K linked to 6C13R), thus excluding theoretical peptides simultaneously including an unlabelled and a labelled lysine or arginine. Mass tolerance was 20 ppm in MS and 30 ppm in MS/MS. Identifications with 10 ppm tolerance in MS yielded very similar results, and were not used for further analysis. The false positive rate was estimated by searching the data using a concatenated database which contains the original UniprotKB database, as well as a version of each original entry where the sequence has been randomized. A 1% FDR was permitted at the protein and peptide level. For quantitation, only unique peptides were considered; peptides common to several proteins were not used for quantitative analysis. Relative quantization of peptide abundance was performed via integration with Protein Prospector of the areas of precursor ions of the identified peptides, using the 3 lightest ions of the isotopic envelopes for both the light (natural isotopic abundances) and heavy (13C(6) lysine and/or arginine containing) forms. Integration was performed from −10 s to +30 s from the time at which the MS/MS spectrum identifying the peptide was acquired. Relative abundances were calculated as ratios vs. control (light, untreated). For total protein relative levels, peptide ratios were aggregated to the protein levels using median values of the log2 ratios. P values were calculated from 3 biological replicates of the experiment.
500 from m/z = 10. A dynamic exclusion window was applied which prevented the same m/z from being selected for 10 s after its acquisition. Peak lists were generated using PAVA in-house software.17 All generated peak lists were searched against the mouse subset of the UniProtKB 2013.6.17 database, using Protein Prospector18 with the following parameters: enzyme specificity was set as Trypsin, and up to 2 missed cleavages per peptide were allowed. Carbamidomethylation of cysteine residues was allowed as a fixed modification. N-Acetylation of the N-terminus of the protein, loss of protein N-terminal methionine, pyroglutamate formation from of peptide N-terminal glutamines, oxidation of methionine and 13C(6) labelling in lysine or arginine were allowed as variable modifications. Search space was limited with the “linked” option (6C12K linked to 6C12R, and 6C13K linked to 6C13R), thus excluding theoretical peptides simultaneously including an unlabelled and a labelled lysine or arginine. Mass tolerance was 20 ppm in MS and 30 ppm in MS/MS. Identifications with 10 ppm tolerance in MS yielded very similar results, and were not used for further analysis. The false positive rate was estimated by searching the data using a concatenated database which contains the original UniprotKB database, as well as a version of each original entry where the sequence has been randomized. A 1% FDR was permitted at the protein and peptide level. For quantitation, only unique peptides were considered; peptides common to several proteins were not used for quantitative analysis. Relative quantization of peptide abundance was performed via integration with Protein Prospector of the areas of precursor ions of the identified peptides, using the 3 lightest ions of the isotopic envelopes for both the light (natural isotopic abundances) and heavy (13C(6) lysine and/or arginine containing) forms. Integration was performed from −10 s to +30 s from the time at which the MS/MS spectrum identifying the peptide was acquired. Relative abundances were calculated as ratios vs. control (light, untreated). For total protein relative levels, peptide ratios were aggregated to the protein levels using median values of the log2 ratios. P values were calculated from 3 biological replicates of the experiment.
2.4 RNA-seq analysis
NIH-3T3 cells (3 independent repeats) grown in 6-well dishes were treated for 24 h or 48 h with 10 μM AS1411 or 10 μM control aptamer and total RNA was extracted using RNeasy Mini kit according to manufacturer's instructions. RNA integrity was verified using TapeStation analysis (Agilent) and high-integrity samples (RIN > 9) were processed using an in-house polyA-based RNA seq protocol (INCPM mRNA Seq) at the Crown Institute for Genomics (G-INCPM, Weizmann Institute of Science). SR60 reads were sequenced on 1 lane of an Illumina HiSeq2500v4. The output was ∼22 million reads per sample. Poly-A/T stretches and Illumina adapters were trimmed from the reads using cutadapt;19 resulting reads shorter than 30 bp were discarded. Reads for each sample were aligned independently to the Mus musculus reference genome GRCm38 using STAR,20 supplied with gene annotations downloaded from Ensembl (and with EndToEnd option). Expression levels for each gene were quantified using htseq-count.21 Differentially expressed genes were identified, and analysis was performed using DESeq222 with the betaPrior, cooksCutoff and independentFiltering parameters set to False. Raw P values were adjusted for multiple tests using the procedure of Benjamini and Hochberg. Differentially expressed genes were determined by a p-adj of <0.05, fold changes >1.5 and max raw counts >30.2.5 Colorimetric and GC–MS cholesterol assays
For colorimetric detection, 106 cells were extracted with 200 μL of chloroform![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) isopropanol
isopropanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NP-40 (7
NP-40 (7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 11
11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.1), organic phase containing cholesterol was air-dried at 50 °C and total cholesterol was quantified using a Cell Biolabs kit (STA-384) according to the manufacturer's instructions. For GC–MS detection, 2 × 106 NIH-3T3 cells were dried in a lyophilizer. The obtained material was extracted sequentially with methanol (400 μL), methanol–chloroform (1
0.1), organic phase containing cholesterol was air-dried at 50 °C and total cholesterol was quantified using a Cell Biolabs kit (STA-384) according to the manufacturer's instructions. For GC–MS detection, 2 × 106 NIH-3T3 cells were dried in a lyophilizer. The obtained material was extracted sequentially with methanol (400 μL), methanol–chloroform (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 400 μL), and chloroform (200 μL), alternating vortex (30 s) with centrifugal precipitation of insoluble material (21
1, 400 μL), and chloroform (200 μL), alternating vortex (30 s) with centrifugal precipitation of insoluble material (21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g, 5 min) and isolation of supernatants. The combined extracts were dried in speedvac, and finally in a lyophilizer. The obtained sample residues were derivatized with N-methyl-N-trimethylsilyl trifluoroacetamide (MSTFA, 70 μL per sample) by shaking at room temperature for 15 min. The mixtures were transferred to a 2 mL autosampler glass vial with a 100 μl glass insert. The GC–MS system comprised an Agilent 7890A gas chromatograph equipped with split/splitless injector, and LECO Pegasus HT time-of-flight mass spectrometer (TOFMS). GC was performed on a 30 m × 0.25 mm × 0.25 μm Rxi-5Sil MS column (Restek). Samples were analyzed in the splitless mode; injector and transfer line temperatures were set at 280 °C. Analytes (1 μL injections) were separated using the following chromatographic conditions: helium was used as carrier gas at a flow rate of 1 mL min−1. The thermal gradient started at 170 °C, was held at this temperature for 2 min, ramped to 280 °C at 37 °C min−1 and then ramped to 300 °C at 1.5 °C min−1 and held at 300 °C for 5 min. Eluents were fragmented in the electron impact mode with an ionization voltage of 70 eV. The MS mass range was 50–750 m/z with an acquisition rate of 20 spectra per second. The ion source chamber was set to 230 °C and the detector voltage was 1650 V. Cholesterol was analyzed using unique mass 458. Quantitation was done by peak areas, using standard curve 1–50 μg per sample, and normalized by total protein in the pellet.
000g, 5 min) and isolation of supernatants. The combined extracts were dried in speedvac, and finally in a lyophilizer. The obtained sample residues were derivatized with N-methyl-N-trimethylsilyl trifluoroacetamide (MSTFA, 70 μL per sample) by shaking at room temperature for 15 min. The mixtures were transferred to a 2 mL autosampler glass vial with a 100 μl glass insert. The GC–MS system comprised an Agilent 7890A gas chromatograph equipped with split/splitless injector, and LECO Pegasus HT time-of-flight mass spectrometer (TOFMS). GC was performed on a 30 m × 0.25 mm × 0.25 μm Rxi-5Sil MS column (Restek). Samples were analyzed in the splitless mode; injector and transfer line temperatures were set at 280 °C. Analytes (1 μL injections) were separated using the following chromatographic conditions: helium was used as carrier gas at a flow rate of 1 mL min−1. The thermal gradient started at 170 °C, was held at this temperature for 2 min, ramped to 280 °C at 37 °C min−1 and then ramped to 300 °C at 1.5 °C min−1 and held at 300 °C for 5 min. Eluents were fragmented in the electron impact mode with an ionization voltage of 70 eV. The MS mass range was 50–750 m/z with an acquisition rate of 20 spectra per second. The ion source chamber was set to 230 °C and the detector voltage was 1650 V. Cholesterol was analyzed using unique mass 458. Quantitation was done by peak areas, using standard curve 1–50 μg per sample, and normalized by total protein in the pellet.
2.6 Lipidomics analysis
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 (v/v) mixture, containing following internal standards: 0.1 μg mL−1 of phosphatidylcholine (17:0/17:0) (Avanti), 0.4 μg mL−1 of phosphatidylethanolamine (17:0/17:0, 0.15) nmol mL−1 of ceramide/sphingoid internal standard mixture I (Avanti, LM6005), 0.0267 μg mL−1 d5-TG internal standard mixture I (Avanti, LM6000) and 0.1 μg mL−1 palmitic acid-13C (Sigma, 605573). The tubes were vortexed and then sonicated for 30 min in an ice-cold sonication bath (taken for a brief vortex every 10 min). Then, double deionized water (DDW)
3 (v/v) mixture, containing following internal standards: 0.1 μg mL−1 of phosphatidylcholine (17:0/17:0) (Avanti), 0.4 μg mL−1 of phosphatidylethanolamine (17:0/17:0, 0.15) nmol mL−1 of ceramide/sphingoid internal standard mixture I (Avanti, LM6005), 0.0267 μg mL−1 d5-TG internal standard mixture I (Avanti, LM6000) and 0.1 μg mL−1 palmitic acid-13C (Sigma, 605573). The tubes were vortexed and then sonicated for 30 min in an ice-cold sonication bath (taken for a brief vortex every 10 min). Then, double deionized water (DDW)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) methanol (3
methanol (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) solution (0.5 mL) was added to the tubes, followed by centrifugation. The upper organic phase was transferred into a 2 mL Eppendorf tube. The polar phase was re-extracted as described above, with 0.5 mL of MTBE. Both organic phases were combined and dried in speedvac and then stored at −80 °C until analysis. For analysis, the dried lipid extracts were re-suspended in 150 μl mobile phase B (see below) and centrifuged again at 20
1, v/v) solution (0.5 mL) was added to the tubes, followed by centrifugation. The upper organic phase was transferred into a 2 mL Eppendorf tube. The polar phase was re-extracted as described above, with 0.5 mL of MTBE. Both organic phases were combined and dried in speedvac and then stored at −80 °C until analysis. For analysis, the dried lipid extracts were re-suspended in 150 μl mobile phase B (see below) and centrifuged again at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800g and 4 °C for 5 min.
800g and 4 °C for 5 min.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) acetonitrile
acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) isopropanol 46
isopropanol 46![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 38
38![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 16 (v/v/v) with 1% 1 M NH4Ac, 0.1% glacial acetic acid. Mobile phase B composition is DDW
16 (v/v/v) with 1% 1 M NH4Ac, 0.1% glacial acetic acid. Mobile phase B composition is DDW![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) acetonitrile
acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) isopropanol 1
isopropanol 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 69
69![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 (v/v/v) with 1% 1 M NH4Ac, 0.1% glacial acetic acid. The column was maintained at 40 °C; the flow rate of the mobile phase was 0.4 mL min−1, the run time was 25 min. The gradient was as follows: mobile phase A was run for 1 min at 100%, then it was reduced to 25% for 11 min, followed by a decrease to 0% for 4 min. Then, mobile phase B was run at 100% for 5.5 min, followed by setting mobile phase A to 100% for 0.5 min. Finally, the column was equilibrated at 100% A for 3 min. MS parameters were as follows: the source and desolvation temperatures were maintained at 120 °C and 450 °C, respectively. The capillary voltage was set to 3 kV and 2 kV for positive and negative ionization mode, respectively; cone voltage was set to 40 V. Nitrogen was used as desolvation gas and cone gas at a flow rate of 800 L h−1 and 30 L h−1, respectively. The mass spectrometer was operated in full scan HDMSE resolution mode over a mass range of 50–2000 Da. For the high-energy scan function, a collision energy ramp of 20–80 eV was applied, and for the low-energy scan function −4 eV was applied.
30 (v/v/v) with 1% 1 M NH4Ac, 0.1% glacial acetic acid. The column was maintained at 40 °C; the flow rate of the mobile phase was 0.4 mL min−1, the run time was 25 min. The gradient was as follows: mobile phase A was run for 1 min at 100%, then it was reduced to 25% for 11 min, followed by a decrease to 0% for 4 min. Then, mobile phase B was run at 100% for 5.5 min, followed by setting mobile phase A to 100% for 0.5 min. Finally, the column was equilibrated at 100% A for 3 min. MS parameters were as follows: the source and desolvation temperatures were maintained at 120 °C and 450 °C, respectively. The capillary voltage was set to 3 kV and 2 kV for positive and negative ionization mode, respectively; cone voltage was set to 40 V. Nitrogen was used as desolvation gas and cone gas at a flow rate of 800 L h−1 and 30 L h−1, respectively. The mass spectrometer was operated in full scan HDMSE resolution mode over a mass range of 50–2000 Da. For the high-energy scan function, a collision energy ramp of 20–80 eV was applied, and for the low-energy scan function −4 eV was applied.
2.7 qRT-PCR analysis
RNA was isolated with RNeasy Mini kit (Qiagen) and treated on-column with DNase (Qiagen) according to manufacturer's instructions. cDNA was synthesized from 100–200 ng of RNA with Superscript IV reverse transcriptase (Thermo Scientific) using 1:1 mix of oligo(dT)20 and random hexamer primers. Quantitative PCR was performed on a LightCycler 480 instrument using HOT FIREPol EvaGreen qPCR Supermix (Solis BioDyne). All qPCR reactions were performed in duplicates and target expression was normalized to 18S or UBC mRNA levels. Relative expression of target genes was calculated using the ΔΔCt method. Primers used in this study are listed in Table S7.2.8 Western blot
Cells grown in 24-well dishes were washed with PBS and lysed directly in 100 μl Laemmli buffer. Proteins (20 μl of lysate) were separated by 10% PAGE and transferred to PVDF membrane using the Trans-Blot Turbo Transfer System (Bio-Rad). 5% skimmed milk in PBST buffer (0.1% Tween-20) was used for blocking membranes and 2% milk-PBST was used for incubations with primary and secondary antibodies. Membranes were incubated overnight at 4 °C with anti-HMGCR (ab242315, Abcam, 1:1000) or anti-tubulin beta (DSHB clone E7, 1:2000) antibodies. Anti-mouse HRP-conjugated antibody (A16066, Thermo Scientific, 1:5000) was used as the secondary antibody. Chemiluminescence was detected by SuperSignal West Atto Chemiluminescent Substrate (Thermo Scientific, for HMGCR blots) SuperSignal West Dura Substrate (Thermo Scientific, for tubulin beta), and quantified using ImageQuant LAS 4000 imager and ImageQuant TL software (GE Healthcare).2.9 Ratiometric imaging of membrane fluidity
NIH-3T3 cells were seeded in growth medium at the density of 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per dish on 35-mm glass-bottom dishes (MatTek) coated with poly-L-lysine (100 μg mL−1, P4832, Sigma) and treated on the following day with AS1411 (10 μM), control aptamer (10 μM) or vehicle for 48 h. Before imaging, medium was replaced with phenol red-free medium with Di-4-ANEPPS (2.5 μM). The cells were subsequently imaged live using a Leica TCS SP8 scanning confocal microscope inside a 37 °C, 5% CO2 incubation chamber. Imaging was carried out in a time window of 15–60 minutes following Di-4-ANEPPS labeling. Single confocal images at the widest cross-section of each cell were taken with a 60× oil-immersion objective and 488 nm laser excitation. Quantification was carried out using custom written Fiji (ImageJ) software.24 Membrane fluidity was quantified by calculating Di-4-ANEPPS general polarization (GP) per pixel:
000 cells per dish on 35-mm glass-bottom dishes (MatTek) coated with poly-L-lysine (100 μg mL−1, P4832, Sigma) and treated on the following day with AS1411 (10 μM), control aptamer (10 μM) or vehicle for 48 h. Before imaging, medium was replaced with phenol red-free medium with Di-4-ANEPPS (2.5 μM). The cells were subsequently imaged live using a Leica TCS SP8 scanning confocal microscope inside a 37 °C, 5% CO2 incubation chamber. Imaging was carried out in a time window of 15–60 minutes following Di-4-ANEPPS labeling. Single confocal images at the widest cross-section of each cell were taken with a 60× oil-immersion objective and 488 nm laser excitation. Quantification was carried out using custom written Fiji (ImageJ) software.24 Membrane fluidity was quantified by calculating Di-4-ANEPPS general polarization (GP) per pixel:where I is the fluorescnce intensity at defined wavelenght and G is a GP of a reference sample determined as described previously (Pokorna et al., 2022).
Relative GP was calculated as follows:
The mean GP signal at the plasma membrane was measured in a manually drawn region of interest (ROI) around the circumference of the cell (11 pixels wide, 0.065 μm per pixel) per cell. Intracellular GP signal was measured in the entire cell area excluding the designated plasma membrane ROI.
2.10 Cholesterol uptake analysis
Cholesterol-NBD (Thermo Fisher, N1148) was dissolved at 1 mg ml−1 in Ethanol and added directly into the media (final concentration 5 μg ml−1) of cells grown on glass bottom multiwell plates that were pre-treated for 48 h with 10 μM AS1411, control aptamer or media only (vehicle). After 2 hours, cells were washed once with fresh media and promptly imaged using an ImageXpress micro confocal microscope (MDC) inside a chamber maintaining 37 °C and 5% CO2 environment. NBD fluorescence (491 nm laser excitation) and transmission light images were taken using 20× air objective and were analyzed using Fiji (ImageJ) as follows: cells were selected from relatively sparse regions in the well, so only cells that are clearly separated from their neighbors were marked for analysis. The diffuse signal of NBD was used to manually trace the area of the cell and NBD signal was measured per cell. Mean background signal (regions without cells) was subtracted and mean and total signal per cell were calculated.LDL:cholesterol oleate-BODIPY uptake experiments were done as described previously with minor modifications.25 Briefly, human LDL was purified by sequential flotation ultracentrifugation26 Chylomicrons and VLDL were removed from the top layer at d = 1.006 g ml−1, followed by separation of LDL at d = 1.063 g ml−1. Total protein in LDL fractions was quantified using a BCA assay (Merck, #71285). BODIPY-cholesterol oleate solution in methyl acetate (Cayman Chemical, #39685) was dried under nitrogen and dissolved in DMSO to 0.5 mg mL−1. 50 nmol of BODIPY-cholesterol oleate per 1 mg mL−1 of LDL was added to the purified LDL solution. The reaction was carried out at 40 °C for 2 h while shaking at 350 rpm using an Eppendorf Thermomixer comfort Incubator. The mixture was loaded onto a Superose 6 Increase 10/300 GL column equilibrated with 1× PBS pH 7.4 for size exclusion. LDL fractions were collected. Fluorescence of fractions was measured at excitation 480 nm, emission 500–600 nm (Shimadzu RF-5301 PC spectrophotometer). LDL fractions that displayed the highest fluorescence intensity were subsequently pooled and concentrated using a 10 kDa MWCO Vivaspin 500 centrifugal filter (Merck, #GE28-9322-25). Protein concentration in the purified LDL:cholesterol oleate-BODIPY complex was measured using a BCA assay (Thermo Scientific). 50 μg of protein equivalent of the LDL:cholesterol oleate-BODIPY complex was added to serum-free DMEM medium together with 10 μM AS1411 or 10 μM control aptamer and incubated for 24 h, fixed with 4% PFA for 15 min, counterstained with Phalloidin-Alexa594 and imaged using Zeiss LSM900 confocal microscope. Signal measurements per cell were performed as described for NBD-cholesterol above, with the difference that cell area was delineated using Phalloidin counterstain.
3. Results
3.1 AS1411 affects NIH-3T3 size and viability
We carried out our multi-omics investigation on NIH-3T3 cells as this line strongly responds to AS1411 by an increase in cell size, accompanied by a reduction in viability (Fig. 1(A) and (B)).13 Control aptamer, in which deoxyguanosines were replaced by deoxycytidines, did not affect either of these measures compared to vehicle control without aptamer DNA (Fig. 1(A) and (B)).3.2 AS1411-induced transcriptome changes
Next, we studied the transcriptomic changes in AS1411-treated NIH-3T3 cells by RNA-seq analysis. Comparing AS1411 vs. control aptamer-treated cells, we identified 273 differentially expressed mRNAs at 24 h and 2204 differentially expressed mRNAs at 48 h (minimum fold-change 1.5 and p < 0.05, Fig. 1(C) and Table S1). Gene ontology analysis of the regulated genes identified the cholesterol biosynthesis pathway as significantly regulated by AS1411 (Fig. 1(D) and (E) and Table S2). Transcript levels for several enzymes in this pathway were downregulated after both 24 h and 48 h AS1411 treatment (Fig. 1(E)). Note that the average effect size of this pathway regulation was larger at 48 h than 24 h (average downregulation of 2.34-fold vs. 1.58-fold across pathway genes Acat2, Cyp51, Dhcr7, Dhcr24, Fdft1, Fdps, Hmgcr, Hmgcs1, Idi1, Lss, Msmo1, Mvd, Mvk, Nsdhl, Pmvk, Sqle).In addition, we observed changes in expression of several mRNAs associated with the expected antiproliferative and cytotoxic effects of AS1411. At 24 h we observed 1.67-fold upregulation of Nupr1, a cellular stress-response integrating transcriptional regulator, 1.57-fold increase in Gadd45g, a cell cycle regulator reponsive to DNA damage, and a 1.56-fold downregulation of a cell cycle regulator Aurka. Upon 48 h of AS1411 treatment, a broader and more pronounced cellular stress response was unfolding at the mRNA level. Gadd45g was upregulated 5.27-fold, the stress-response transcription factor Atf3 increased 6.96-fold and the cell cycle inhibitor Mxd1 increased 2.41-fold. (top GO category “negative regulation of transcription from RNA polymerase II promoter”, Fig. 1(D)).
3.3 AS1411-induced proteome changes
Next, we conducted a SILAC analysis27 to study the effects of AS1411 on the NIH-3T3 cell proteome (Fig. 2(A)). We identified 101 and 480 differentially expressed proteins at 24 h and 48 h, respectively (AS1411 vs. control aptamer 1.5× fold change and p < 0.05 threshold; Fig. 2(B) and Table S3). Among these hits, we identified a number of proteins associated with ribosome biogenesis and ribosomal RNA maturation (Fig. 2(C) and Fig. S1A). At 24 h, we observed downregulation of protein MAK16 homolog involved in ribosome biogenesis (Mak16, 1.73-fold downregulation), DEAD-box helicase 56, a nucleolar helicase involved in ribosome biogenesis (Ddx56, down 1.55-fold) and GTPase Era, a mitochondrial rRNA chaperone (Eral1, down 1.53-fold). At 48 h, this molecular program was more pronounced and we detected a broad downregulation of several ribosome biogenesis and rRNA maturation proteins (Fig. S1A). In addition, a small but consistent decrease in ribosome proteins was detected at 48 h (Fig. S1B). We also detected decreases in RNA polymerase I and III subunits – RNA polymerase I subunit RPA49 (Polr1e, down 1.5-fold at 48 h), RNA polymerase III subunit RPC7 (Polr3g, down 1.66-fold) and RNA polymerases I and III subunit RPAC1 (Polr1c, down 1.5-fold). Collectively, these changes indicate a reduction in ribosome biosynthesis and translation upon AS1411-mediated perturbation of nucleolin function.In agreement with the antiproliferative effect of AS1411, we detected downregulation of DNA replication origin recognition complex subunits 2,4,5 and 6 at 48 h (Orc2, 3-6, 1.86-fold, 2.32-fold, 1.54-fold and 1.81-fold, respectively). In addition, we observed downregulation of histone proteins H1.1 (Hist1h1a, 3.42-fold), H1.5 (Hist1h1b, 2.0-fold), H3.1 (Hist1h3a, 1.65-fold) and H4 (Hist1h4a, 1.62-fold), indicating chromatin remodeling by AS1411.
Gene ontology pathway analysis showed that AS1411 induced downregulation of cholesterol biosynthesis pathway enzymes also at the protein level (Fig. 2(C) and (D) and Table S4). We also noted a twofold increase in Npc1 (Niemann-Pick C1), a protein responsible for the transfer of endocytosed cholesterol from late endosomes to the endoplasmic reticulum,28 as well as a 70% increase in Scp2, a lipid carrier protein that has been implicated in intracellular cholesterol distribution.29 In addition to regulation of sterol metabolism, we detected a concerted upregulation of several fatty acid beta-oxidation pathway enzymes (Fig. 2(C) and (E)).
Taken together, AS1411 consistently downregulated the expression of cholesterol biosynthesis enzymes at both mRNA and protein levels (Fig. 2(F)), with mRNA and protein levels of cholesterol biosynthesis enzymes strongly correlated at 48 h after AS411 treatment (Fig. S1C).
3.4 Validation of cholesterol biosynthesis gene expression changes
We selected four cholesterol pathway genes that showed the most consistent regulation in both RNA-seq and SILAC analyses – Sqle, Fdft1, Hmgcr and Hmgcs1 – and validated these using qRT-PCR analysis after treating NIH-3T3 cells for 48 h with AS1411 or controls (Fig. 3(A)). In addition, we also validated downregulation of Hmgcr – a rate-limiting enzyme in cholesterol biosynthesis – by western blot (Fig. 3(B) and (C)). We then repeated these validation experiments in HeLa cells, where nucleolin knockdown has previously been shown to result in downregulation of the cholesterol biosynthesis pathway.30 In both HeLa and NIH-3T3 cells nucleolin can be found on the cell surface, potentially allowing internalization of AS1411 by a nucleolin-dependent mechanism.7,12 We also observed downregulation of SQLE, FDFT1, HMGCR and HMGCS1 mRNAs in HeLa cells (Fig. 3(D)), as well as reduced levels of the Hmgcr protein (Fig. 3(E) and (F)), suggesting a shared mechanism in the regulation of cholesterol biosynthesis pathway gene expression by AS1411 and nucleolin knockdown.3.5 AS1411-induced changes in cellular lipid composition
The findings above strongly suggest that nucleolin engagement by AS1411 causes changes in cellular lipids. We tested this prediction by analyzing AS1411-induced lipidome changes, using targeted assays for cholesterol and by performing lipid profiling of the cells (Fig. 4(A)). Two different assays for cholesterol quantification (GC–MS and colorimetric) showed that AS1411 treatment increased cholesterol levels per cell and did not change cholesterol levels per cellular dry mass (Fig. 4(B) and (C)); while at 48 h AS1411 treatment increased also cellular protein content (Fig. S2). Among the 178 identified lipid species, 73 exhibited differential accumulation, and 5 out of 12 lipid classes (when individual lipid intensities were summed) showed differential levels (Fig. 4(D) and Table S5). No significant AS1411-induced changes were observed in storage lipids (triglycerides, fatty acids and sterol esters) and ceramides. In contrast, the abundances of several phospholipid classes were affected (Fig. 4(D)). We observed a robust, ∼2-fold increase in phosphatidylethanolamines and a similar increase in lysophosphatidylcholines and lysophosphatidylethanolamines, two minor phospholipid classes that regulate inflammation, tumor cells invasiveness and neuronal growth signaling.31–33 Finally, we detected a significant decrease in glycosphingolipids as well as a modest decrease in sphingomyelins (Fig. 4(D)).3.6 AS1411 decreases intracellular membrane fluidity
Considering the amount of cellular cholesterol per dry cell mass remained unchanged, downregulation of the cholesterol synthesis pathway by AS1411 may be related to cholesterol redistribution in cells. If such redistribution indeed occurs and leads to higher ER accumulation, the fluidity of the corresponding membranes should decrease.24,34 We therefore examined the effect of AS1411 on NIH-3T3 membrane fluidity by staining cells with the ratiometric fluorescence probe Di-4-ANEPPS.35 Generalized polarization (GP) measurements of Di-4-ANEPPS in NIH-3T3 cells showed that AS1411 treatment did not affect plasma membrane fluidity, but decreased fluidity in membranes of intracellular compartments (Fig. 5(A)–(C)), suggesting possible redistribution of cholesterol in intracellular compartments upon AS1411 treatment.3.7 AS1411 does not increase cholesterol uptake
Next, we investigated if enhanced cellular uptake of cholesterol from the medium could cause transcriptional downregulation of the mevalonate pathway. To this end, we quantified cholesterol uptake into NIH-3T3 cells using fluorescent, NBD-labeled cholesterol which was freely dissolved in the growth media and incubated for 2 hours following 48 h of aptamer treatment.36 We did not detect increased cholesterol uptake using this assay (Fig. 5(D) and (E)). To further test the effect of AS1411 specifically on LDLR-mediated cholesterol uptake, we measured the uptake of LDL:cholesterol oleate-BODIPY complex,25 which was lower after AS1411 treatment compared to control aptamer and vehicle (Fig. S3).4. Discussion
The molecular mechanisms that underlie cell size and proliferation regulation upon nucleolin engagement by AS1411 remain poorly understood.10,12 Our study now reveals broad changes in lipid metabolism-associated mRNAs and proteins, as well as cellular abundance of several lipid classes. Nucleolin was previously implicated in the regulation of cellular lipid homeostasis, but the role of these effects was unclear. Nucleolin co-fractionates with lipid rafts in HeLa cells,37 and associates with flotillin37 and integrin38 in a lipid raft-dependent manner. Indeed, we previously demonstrated that localization of nucleolin to the plasma membrane is facilitated by its glycine arginine-rich (GAR) domain.12 In macrophages, nucleolin was identified as a receptor for acetylated low density lipoprotein (LDL) particles.39 Another study reported that nucleolin interacts and enhances the stability of ABCA1 mRNA encoding for a cholesterol efflux transporter in macrophages.40 Overexpression of nucleolin was shown to attenuate oxidized LDL-induced lipid overload and facilitate cholesterol efflux in macrophages.40One of our main findings is a coordinated downregulation of cholesterol biosynthesis gene expression, both at mRNA and protein levels. This is in accordance with a previous study reporting down regulation of sterol biosynthesis pathway genes after nucleolin knockdown in HeLa cells.30 That prior study found also lower cellular cholesterol levels in HeLa cells targeted by nucleolin siRNA.30 In contrast, we did not observe a reduction in cellular cholesterol by either of two analysis methods, GC-MS and a colorimetric enzyme-based assay (while both cholesterol and protein content per cell were found increased by AS1411 due to the gross increase in cell size). Interestingly, another study reported increased cholesterol to cellular dry mass upon Ncl knockdown by shRNA in RAW264.7 macrophages.40 Possible explanations for the non-concordant results between these three studies are the use of different cell types and nucleolin perturbation methods. The underlying regulation mechanism likely involves different nucleolin-dependent processes affecting lipid synthesis, uptake and cellular redistribution. Further studies are needed to address these questions in depth.
What mode of action can explain our findings that downregulated cholesterol biosynthesis gene program was not coupled with decreased cholesterol levels? At first, we hypothesized that downregulation of cholesterol biosynthesis pathways by AS1411 may be a homeostatic response to increased cholesterol uptake. AS1411 has been shown to enhance micropinocytosis,41 which could lead to engulfment of additional cholesterol bound to serum LDL proteins in the medium.42 However, under our experimental conditions, cholesterol uptake experiments did not reveal increased uptake in AS1411 treated cells, suggesting alternative explanations such as cholesterol redistribution in the cell. Possibly, AS1411-driven increased local cholesterol concentrations are detected at the cellular “sensor unit”, the SCAP-SREBP complex in the ER, leading to a decrease in SREBP-driven transcription of cholesterol pathway genes.43 In agreement with this, we observed a decrease in intracellular membrane fluidity, which may be caused by increased cholesterol content24,34 or increased phosphatidylethanolamines.44,45 In addition, we detected AS1411-induced upregulation of the cholesterol transporter Npc1, which may contribute to shuttling of endocytosed cholesterol to the ER.42
We also found that AS1411 reduced the expression of ribosomal and ribosome assembly components at the protein level. One of nucleolin's prominent roles is in ribosome biogenesis, binding pre-rRNA46,47 and acting as a nucleolar RNA chaperone.48 In addition, nucleolin can also interact with ribosomal proteins via its GAR domain,16,49 which we have recently shown to bind AS1411.12 Our results suggest that in addition to its roles in ribosome assembly, nucleolin may also regulate the expression of ribosome components. The mechanisms by which nucleolin performs this role and how it is perturbed by AS1411 will be of interest for future investigation.
5. Conclusions and limitations of the study
Here, we conducted a set of omics analyses to characterize transcriptome, proteome and lipidome changes induced by the nucleolin-targeted aptamer AS1411 in a cell type where the aptamer mainly affects cell size. This analysis revealed mRNA and protein expression changes in a number of lipid biosynthesis enzymes, abundance changes in several lipid classes and also in lipid-related biophysical properties of intracellular membranes. These changes in lipid metabolism and membrane fluidity may mediate the effects of nucleolin in cell size regulation. Together with two previous reports where nucleolin levels were manipulated by either knockdown30 or overexpression,40 our study highlights nucleolin's essential role in regulating lipid metabolism.The current study focused on multi-omics characterization of the effects of AS1411 using NIH-3T3 cells, due to their primary size response to the aptamer. This may be a limitation for assessment of AS1411 effects in a broad range of cell types, but has the advantage of directly addressing nucleolin functions in normal cells.
Author contributions
Hypotheses and experimental design: IK, MF, EEZ; experiments and data analysis: EEZ, IK, FC-C, IR, JAO-P, AB, SM, MI, ŠP, N-NS, RR; data curation: IK, JAO-P, SM, MI supervision: IK, MF, ALB, AHF, AL; writing: IK, EEZ, MF.Conflicts of interest
The authors report no conflict of interest.Data availability
RNA-seq data have been deposited in NCBI's Gene Expression Omnibus (GEO) and are accessible through GEO series accession number GSE274745. Mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD055162. Lipidomics data has been deposited in the Metabolights repository (provisional accession number REQ20250806212293) and access to raw data files is available upon request.Supplementary information is available. See DOI: https://doi.org/10.1039/d5mo00088b.
Acknowledgements
We thank Albina Lin, Nataliya Okladnikova, Ana E. Ventura, Vladimir Kiss, Reinat Nevo and Tali Shalit for excellent technical assistance and Dalia Gordon for helpful comments and discussions. This work was supported by funding from the Moross Integrated Cancer Center (MF and IK), the Dr Miriam and Sheldon G. Adelson Medical Research Foundation (MF and ALB), the Israel Science Foundation 349/24 (MF and IR), the Chaya Professorial Chair in Molecular Neuroscience (M. F.), an Israeli Academy of Sciences and Humanities (IASH) postdoctoral fellowship (EEZ), a Koshland postdoctoral fellowship (IK), and Estonian Research Council (grants MOBTP192, PRG2206 to IK, MOBJD1210 to FC-C). The work of SM and MI was supported by the Vera and John Schwartz Family Center for Metabolic Biology. The Thermo Scientific Q-Exactive Plus used for this study was supported by the NIH – NIGMS NIH P41GM103481 and 1S10OD016229 awards to ALB.References
- C. M. Berger, X. Gaume and P. Bouvet, Biochimie, 2015, 113, 78–85 CrossRef CAS PubMed.
- R. Cong, S. Das and P. Bouvet, in The Nucleolus, ed. M. O. J. Olson, Springer, New York, NY, 2011, pp. 185–212 Search PubMed.
- I. Rishal and M. Fainzilber, BMC Biol., 2019, 17, 36 CrossRef PubMed.
- H. Ginisty, H. Sicard, B. Roger and P. Bouvet, J. Cell Sci., 1999, 112(Pt 6), 761–772 CrossRef CAS.
- F. Mongelard and P. Bouvet, Trends Cell Biol., 2007, 17, 80–86 CrossRef CAS.
- R. A. Borer, C. F. Lehner, H. M. Eppenberger and E. A. Nigg, Cell, 1989, 56, 379–390 CrossRef CAS.
- A. G. Hovanessian, F. Puvion-Dutilleul, S. Nisole, J. Svab, E. Perret, J. S. Deng and B. Krust, Exp. Cell Res., 2000, 261, 312–328 CrossRef CAS PubMed.
- C. F. Semenkovich, R. E. Ostlund, M. O. Olson and J. W. Yang, Biochemistry, 1990, 29, 9708–9713 CrossRef CAS.
- A. Van den Avont and N. Sharma-Walia, Front. Mol. Biosci., 2023, 10, 1217769 CrossRef CAS.
- P. J. Bates, E. M. Reyes-Reyes, M. T. Malik, E. M. Murphy, M. G. O’Toole and J. O. Trent, Biochim. Biophys. Acta, Gen. Subj., 2017, 1861, 1414–1428 CrossRef CAS.
- P. J. Bates, D. A. Laber, D. M. Miller, S. D. Thomas and J. O. Trent, Exp. Mol. Pathol., 2009, 86, 151–164 CrossRef CAS.
- E. Doron-Mandel, I. Koppel, O. Abraham, I. Rishal, T. P. Smith, C. N. Buchanan, P. K. Sahoo, J. Kadlec, J. A. Oses-Prieto, R. Kawaguchi, S. Alber, E. E. Zahavi, P. Di Matteo, A. Di Pizio, D.-A. Song, N. Okladnikov, D. Gordon, S. Ben-Dor, R. Haffner-Krausz, G. Coppola, A. L. Burlingame, P. Jungwirth, J. L. Twiss and M. Fainzilber, EMBO J., 2021, 40, e107158 CrossRef CAS PubMed.
- R. B.-T. Perry, I. Rishal, E. Doron-Mandel, A. L. Kalinski, K. F. Medzihradszky, M. Terenzio, S. Alber, S. Koley, A. Lin, M. Rozenbaum, D. Yudin, P. K. Sahoo, C. Gomes, V. Shinder, W. Geraisy, E. A. Huebner, C. J. Woolf, A. Yaron, A. L. Burlingame, J. L. Twiss and M. Fainzilber, Cell Rep., 2016, 16, 1664–1676 CrossRef CAS.
- M. Terenzio, S. Koley, N. Samra, I. Rishal, Q. Zhao, P. K. Sahoo, A. Urisman, L. Marvaldi, J. A. Oses-Prieto, C. Forester, C. Gomes, A. L. Kalinski, A. Di Pizio, E. Doron-Mandel, R. B.-T. Perry, I. Koppel, J. L. Twiss, A. L. Burlingame and M. Fainzilber, Science, 2018, 359, 1416–1421 CrossRef CAS PubMed.
- I. Rishal, N. Kam, R. B.-T. Perry, V. Shinder, E. M. C. Fisher, G. Schiavo and M. Fainzilber, Cell Rep., 2012, 1, 608–616 CrossRef CAS PubMed.
- E. E. Zahavi, I. Koppel, R. Kawaguchi, J. A. Oses-Prieto, A. Briner, A. Monavarfeshani, I. Dalla Costa, E. van Niekerk, J. Lee, S. Matoo, S. Hegarty, R. J. Donahue, P. K. Sahoo, S. Ben-Dor, E. Feldmesser, J. Ryvkin, D. Leshkowitz, R. B.-T. Perry, Y. Cheng, E. Farber, O. Abraham, N. Samra, N. Okladnikov, S. Alber, C. A. Albus, I. Rishal, I. Ulitsky, M. H. Tuszynski, J. L. Twiss, Z. He, A. L. Burlingame and M. Fainzilber, Cell, 2025, 188, 4350–4365 CrossRef CAS PubMed.
- S. Guan, J. C. Price, S. B. Prusiner, S. Ghaemmaghami and A. L. Burlingame, Mol. Cell. Proteomics, 2011, 10, M111.010728 CrossRef.
- K. R. Clauser, P. Baker and A. L. Burlingame, Anal. Chem., 1999, 71, 2871–2882 CrossRef CAS.
- M. Martin, EMBnet. J., 2011, 17, 10–12 CrossRef.
- A. Dobin, C. A. Davis, F. Schlesinger, J. Drenkow, C. Zaleski, S. Jha, P. Batut, M. Chaisson and T. R. Gingeras, Bioinformatics, 2013, 29, 15–21 CrossRef CAS.
- S. Anders, P. T. Pyl and W. Huber, Bioinformatics, 2015, 31, 166–169 CrossRef CAS.
- M. I. Love, W. Huber and S. Anders, Genome Biol., 2014, 15, 550 CrossRef PubMed.
- S. Malitsky, C. Ziv, S. Rosenwasser, S. Zheng, D. Schatz, Z. Porat, S. Ben-Dor, A. Aharoni and A. Vardi, New Phytol., 2016, 210, 88–96 CrossRef CAS.
- S. Pokorna, A. E. Ventura, T. C. B. Santos, M. Hof, M. Prieto, A. H. Futerman and L. C. Silva, J. Photochem. Photobiol., B, 2022, 228, 112404 CrossRef CAS PubMed.
- K. Kanerva, R.-L. Uronen, T. Blom, S. Li, R. Bittman, P. Lappalainen, J. Peränen, G. Raposo and E. Ikonen, Dev. Cell, 2013, 27, 249–262 CrossRef CAS.
- R. J. Havel, H. A. Eder and J. H. Bragdon, J. Clin. Invest., 1955, 34, 1345–1353 CrossRef CAS PubMed.
- S.-E. Ong, B. Blagoev, I. Kratchmarova, D. B. Kristensen, H. Steen, A. Pandey and M. Mann, Mol. Cell. Proteomics, 2002, 1, 376–386 CrossRef CAS.
- S. G. Pfisterer, J. Peränen and E. Ikonen, Curr. Opin. Lipidol., 2016, 27, 282–287 CrossRef CAS PubMed.
- A. M. Gallegos, B. P. Atshaves, S. M. Storey, A. L. McIntosh, A. D. Petrescu and F. Schroeder, Biochemistry, 2001, 40, 6493–6506 CrossRef CAS PubMed.
- S. Kumar, E. C. Gomez, M. Chalabi-Dchar, C. Rong, S. Das, I. Ugrinova, X. Gaume, K. Monier, F. Mongelard and P. Bouvet, Sci. Rep., 2017, 7, 9017 CrossRef.
- A. Grzelczyk and E. Gendaszewska-Darmach, Biochimie, 2013, 95, 667–679 CrossRef CAS PubMed.
- K. Hisano, H. Yoshida, S. Kawase, T. Mimura, H. Haniu, T. Tsukahara, T. Kurihara, Y. Matsuda, N. Saito and T. Uemura, J. Biochem., 2021, 170, 327–336 CrossRef CAS.
- S.-H. Law, M.-L. Chan, G. K. Marathe, F. Parveen, C.-H. Chen and L.-Y. Ke, Int. J. Mol. Sci., 2019, 20, 1149 CrossRef CAS PubMed.
- T. Parasassi, E. Gratton, W. M. Yu, P. Wilson and M. Levi, Biophys. J., 1997, 72, 2413–2429 CrossRef CAS PubMed.
- L. Jin, A. C. Millard, J. P. Wuskell, H. A. Clark and L. M. Loew, Biophys. J., 2005, 89, L04–06 CrossRef CAS.
- D. Wüstner, Chem. Phys. Lipids, 2007, 146, 1–25 CrossRef PubMed.
- X. Chen, S. Shank, P. B. Davis and A. G. Ziady, Mol. Ther., 2011, 19, 93–102 CrossRef CAS PubMed.
- J. Bi, R. Wang, Y. Zhang, X. Han, K. K. Ampah, W. Liu and X. Zeng, Mol. Cells, 2013, 36, 507–517 CrossRef CAS.
- Y. Miki, Y. Tachibana, Y. Ohminato and Y. Fujiwara, Biol. Pharm. Bull., 2015, 38, 1420–1424 CrossRef CAS.
- Y. Li, B. Jiang, P. Liang, Z. Tong, M. Liu, Q. Lv, Y. Liu, X. Liu, Y. Tang and X. Xiao, Biochem. Biophys. Res. Commun., 2017, 486, 364–371 CrossRef CAS PubMed.
- E. M. Reyes-Reyes, Y. Teng and P. J. Bates, Cancer Res., 2010, 70, 8617–8629 CrossRef CAS PubMed.
- A. Skorda, A. R. Lauridsen, C. Wu, J. Huang, M. Mrackova, N. I. Winther, V. Jank, Z. Sztupinszki, R. Strauss, M. Bilgin, K. Maeda, B. Liu, Y. Luo, M. Jäättelä and T. Kallunki, Oncogene, 2023, 42, 2495–2506 CrossRef CAS PubMed.
- J. L. Goldstein, R. A. DeBose-Boyd and M. S. Brown, Cell, 2006, 124, 35–46 CrossRef CAS PubMed.
- K. A. Dill and D. Stigter, Biochemistry, 1988, 27, 3446–3453 CrossRef CAS.
- R. Dawaliby, C. Trubbia, C. Delporte, C. Noyon, J.-M. Ruysschaert, P. Van Antwerpen and C. Govaerts, J. Biol. Chem., 2016, 291, 3658–3667 CrossRef CAS.
- M. S. Azman, E. L. Alard, M. Dodel, F. Capraro, R. Faraway, M. Dermit, W. Fan, A. Chakraborty, J. Ule and F. K. Mardakheh, EMBO J., 2023, 42, e110902 CrossRef CAS PubMed.
- A. H. Herrera and M. O. Olson, Biochemistry, 1986, 25, 6258–6264 CrossRef CAS PubMed.
- K. Dörner, C. Ruggeri, I. Zemp and U. Kutay, EMBO J., 2023, 42, e112699 CrossRef.
- P. Bouvet, J. J. Diaz, K. Kindbeiter, J. J. Madjar and F. Amalric, J. Biol. Chem., 1998, 273, 19025–19029 CrossRef CAS.
Footnote |
| † Present address: J. Heyrovský Institute of Physical Chemistry of the Czech Academy of Sciences, Dolejškova 3, 182 23 Prague, Czech Republic. |
| This journal is © The Royal Society of Chemistry 2025 |