Open Access Article
Open Access ArticleThe development of a PET radiotracer for imaging alpha synuclein aggregates in Parkinson's disease†
Gui-Long
Tian
 a,
Chia-Ju
Hsieh
a,
Dinahlee Saturnino
Guarino
a,
Thomas J. A.
Graham
a,
Zsofia
Lengyel-Zhand
a,
Chia-Ju
Hsieh
a,
Dinahlee Saturnino
Guarino
a,
Thomas J. A.
Graham
a,
Zsofia
Lengyel-Zhand
 a,
Alexander
Schmitz
a,
Wai Kit
Chia
a,
Anthony J.
Young
a,
Alexander
Schmitz
a,
Wai Kit
Chia
a,
Anthony J.
Young
 a,
John-Grey
Crosby
a,
Konstantinos
Plakas
a,
Tianyu
Huang
b,
Hao
Jiang
b,
Yanbo
Yu
a,
John-Grey
Crosby
a,
Konstantinos
Plakas
a,
Tianyu
Huang
b,
Hao
Jiang
b,
Yanbo
Yu
 b,
Catherine
Hou
b,
Catherine
Hou
 a,
Hsiaoju
Lee
a,
E. James
Petersson
c,
Sam
Giannakoulias
c,
Jennifer
O'Shea
d,
Paul
Kotzbauer
d,
Zhude
Tu
a,
Hsiaoju
Lee
a,
E. James
Petersson
c,
Sam
Giannakoulias
c,
Jennifer
O'Shea
d,
Paul
Kotzbauer
d,
Zhude
Tu
 b,
Chester A.
Mathis
b,
Chester A.
Mathis
 e and
Robert H.
Mach
e and
Robert H.
Mach
 *a
*a
aDepartment of Radiology, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, USA. E-mail: rmach@pennmedicine.upenn.edu
bDepartment of Radiology, Washington University, School of Medicine, Saint Louis, MO, USA
cDepartment of Chemistry, University of Pennsylvania, Philadelphia, PA, USA
dDepartment of Neurology, Washington University School of Medicine, Saint Louis, MO, USA
eDepartment of Radiology, University of Pittsburgh, Pittsburgh, PA, USA
First published on 3rd April 2025
Abstract
M503-1619 was identified as a promising ligand for positron emission tomography (PET) imaging of α-synuclein (α-Syn) pathology in Parkinson's disease (PD). An exemplar for binding site 9 (residues GLY-86, ILE-88, PHE-94 and LYS-96) of α-Syn fibrils was generated. An in silico ultrahigh throughput screening campaign was conducted using a 42 million compound library. Secondary in silico methods followed by visual inspection were used to select 6 compounds as candidates for in vitro binding studies. M503-1619 was found to have a high binding affinity (Ki = 6.5 nM versus the site 9 radioligand [3H]BF-2846) to α-Syn fibrils and low affinity for beta amyloid (Ki = 390 nM versus [3H]PiB) in competition binding assays. Saturation binding assays of [3H]M503-1619 in human tissues confirmed its high affinity to α-Syn (PD tissue, KD = 2.5 nM; Alzheimer's disease tissue, KD = 37 nM; progressive supranuclear palsy tissue, KD = 55 nM). Autoradiography studies demonstrated a higher binding of this radioligand in PD brain sections than in multiple system atrophy brain sections. PET studies with [11C]M503-1619 showed high brain uptake and rapid washout (whole brain peak to 60 min ratio = 3.2) in non-human primates. The results of this study suggest that [11C]M503-1619 is a lead compound for radiotracer development imaging α-Syn with PET.
1. Introduction
Parkinson's disease (PD) is the second most common and the fastest-growing neurodegenerative disease.1,2 Insoluble α-synuclein (α-Syn) aggregates are the histopathologic hallmark of the synucleinopathies that include both PD and multiple system atrophy (MSA).3,4 The development of a positron emission tomography (PET) radiotracer for imaging α-Syn is important for early diagnosis, monitoring of disease progression, and evaluation of the efficacy of therapeutic interventions to reduce α-Syn burden. The success in the development of PET radiotracers for imaging beta amyloid (Aβ)5,6 and tau7 in Alzheimer's disease (AD) has highlighted the need for a PET radiotracer for imaging α-Syn in the synucleinopathies. However, the pathway for developing a PET radiotracer for imaging α-Syn has proven to be far more challenging than the development of PET radiotracers for Aβ and tau.There are several obstacles that must be overcome in developing a PET radiotracer for α-Syn. First, the concentration of α-Syn aggregates within the brain is lower than that of Aβ or tau.8–10 Consequently, a high affinity for α-Syn aggregates is required for a PET radiotracer to succeed in translational imaging studies. Also, since α-Syn coexist and colocalize with Aβ and tau,8,9,11 a PET radiotracer for α-Syn must have a high binding selectivity for α-Syn over Aβ and tau.12 Finally, α-Syn is located intracellularly, hence, a successful α-Syn PET radiotracer should not only cross the blood–brain barrier, but it must also cross the neuronal cell membrane. These obstacles, together with the additional properties, a PET radiotracer must possess for central nervous system (CNS) imaging studies, such as appropriate physicochemical properties (e.g., molecular weight, topological polar surface area, Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D, etc.) and favorable peripheral metabolism leading to polar metabolites, present a steep challenge in developing an α-Syn PET radiotracer.
D, etc.) and favorable peripheral metabolism leading to polar metabolites, present a steep challenge in developing an α-Syn PET radiotracer.
In spite of the above limitations, there has been some recent success in the development of PET radiotracers for imaging α-Syn in vivo (Fig. 1). For example, [18F]ACI-12589 is a PET radiotracer that has shown promise in imaging α-Syn in MSA but not PD patients.13 In addition, Ye et al. have reported the radiotracer, [18F]F0502B,14 which is an analog of PBB315 (a mixed tau/α-Syn PET radiotracer) is capable of imaging α-Syn in autoradiography studies of PD brain. Although [18F]F0502B is not a useful PET radiotracer for in vivo imaging studies, it is a good lead compound for further structure–activity relationship studies aimed at identifying a ligand having a better off-target binding profile and higher brain uptake. More recently, Endo et al. demonstrated that [18F]C05-05 was able to image α-Syn in both PD and MSA patients.16 However, the low brain uptake and high affinity of this radiotracer for tau limits its utility in PET imaging studies of α-Syn.
Based on the solid state nuclear magnetic resonance structure of α-Syn fibrils, our group identified multiple binding sites for small molecules in the “Greek key region” of the protein.17 Using photo-crosslinking studies with mass spectrometry, we confirmed that there are two binding sites which can bind small molecules with nanomolar affinity, site 2 and site 9. “Site 2” is located towards the N-terminus of the Greek key motif and contains TYR-39, SER-42 and THR-44, whereas “site 9” is located towards the C-terminus of the Greek key motif and contains GLY-86, ILE-88, PHE-94 and LYS-96. We also conducted an ultrahigh throughput in silico screen of commercially available compound libraries by generating a pseudo-ligand or “Exemplar” to identify compounds binding to site 2 in recombinant α-Syn fibrils.18 We confirmed that the exemplar19 method for screening large compound libraries is an effective method for identifying novel compounds that bind to α-Syn.
Since site 2 contains TYR-39, a site that is phosphorylated in α-Syn in PD brain,20,21 we employed a similar strategy on a different compound library by using the exemplar generated from the prior work18 for site 9, which does not contain any amino acid residues that undergo post-translational modification in PD brain.22,23 This strategy resulted in the identification of a new ligand, M503-1619, which binds to α-Syn with high affinity and has low off-target binding to Aβ and tau. In this report, we describe the in vitro characterization of [3H]M503-1619 in postmortem samples of PD brain, and the radiosynthesis and in vivo evaluation of [11C]M503-1619 in PET imaging studies in non-human primates (NHPs). Our data suggest that [11C]M503-1619 is a promising lead compound for PET radiotracer for imaging α-Syn fibrils in PD.
2. Results and discussion
2.1. In silico ultrahigh throughput screening
A site 9 Exemplar generated from prior work18 was used for the ultrahigh throughput in silico screening on a library of 42 million compounds. The top 3 million in silico hits were narrowed down to 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 compounds by using the secondary in silico filters, including pan-assay interference compounds (PAINS)24 screening, central nervous system multiparameter optimization (CNS MPO)25 scoring, and additional chemical properties filtering (e.g., number of aromatic rings, molecular weight, and cLog
000 compounds by using the secondary in silico filters, including pan-assay interference compounds (PAINS)24 screening, central nervous system multiparameter optimization (CNS MPO)25 scoring, and additional chemical properties filtering (e.g., number of aromatic rings, molecular weight, and cLog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D). Visual inspection led to identification of 6 compounds (compound M503-1619, P035-0108, P706-0700, P706-0073, P706-0209, and P706-0701) for further in vitro binding studies. The structures of the candidate ligands are shown in Table 1.
D). Visual inspection led to identification of 6 compounds (compound M503-1619, P035-0108, P706-0700, P706-0073, P706-0209, and P706-0701) for further in vitro binding studies. The structures of the candidate ligands are shown in Table 1.
| Compounds | Structure | α-Syn fibrils (Ki; nM)a | AD brain tissue homogenates (Ki; nM)b | CLogP | Molecular weight | Topological polar surface area |
|---|---|---|---|---|---|---|
| a K i values were determined from competitive radioligand binding full curves using recombinant α-Syn fibrils with [3H]BF-2846. b K i values were determined from competitive radioligand binding full curves using AD brain tissue homogenates with [3H]PiB. | ||||||
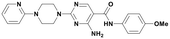
| M503-1619 |
|
6.5 ± 2.9 | 390 ± 120 | 2.89 | 404.45 | 109.50 |
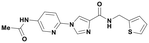
| P035-0108 |
|
388.4 ± 178.9 | >1000 | 2.00 | 341.39 | 117.15 |
| P706-0700 |
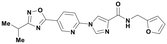
|
12.6 ± 1.0 | >1000 | 2.32 | 378.38 | 111.87 |
| P706-0073 |
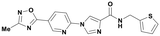
|
>1000 | >1000 | 2.09 | 366.40 | 126.97 |
| P706-0209 |
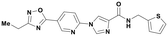
|
>1000 | >1000 | 2.62 | 380.42 | 126.97 |
| P706-0701 |
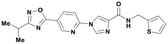
|
12.1 ± 2.9 | >1000 | 3.02 | 394.45 | 126.97 |
2.2. In vitro radioligand competition binding assays
For testing the binding affinity to α-Syn fibrils, competition binding assays were initially conducted in recombinant α-Syn fibrils using [3H]BF-2846 as the radioligand. This radioligand was found to bind preferentially to site 9 in α-Syn fibrils.17M503-1619, P706-0700 and P706-0701 had a high affinity to α-Syn fibrils (Ki = 6.5 nM, 12.6 nM, and 12.1 nM vs. [3H]BF-2846; Table 1). For determining their selectivity towards Aβ, the three ligands were further tested in AD brain tissue homogenates using [3H]PiB as the competing radioligand. All the three ligands were found to have a low affinity to Aβ (Ki = 390 nM for M503-1619 and Ki > 1000 for P706-0700 and P706-0701vs. [3H]PiB; Table 1).Competition binding assays for tau were also conducted using several tritiated tau radioligands, including [3H]MK-6240,26 [3H]PI-2620,27 [3H]Z-234028 and [3H]CBD-2115,29 in post-mortem samples of AD tissues (Table 2). [3H]Chrysamine-G ([3H]CG), which binds to a different site on Aβ than PiB,30 was also included in this assay as an alternative measure of binding to Aβ (Table 2). The results of these studies revealed very low binding of M503-1619 to tau and Aβ.
| Radioligand | K i (nM) |
|---|---|
| [3H]CG | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
| [3H]PI-2620 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
| [3H]Z-2340 | 3700 |
| [3H]MK-6240 | 240 |
| [3H]CBD-2115 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
2.3. Chemistry
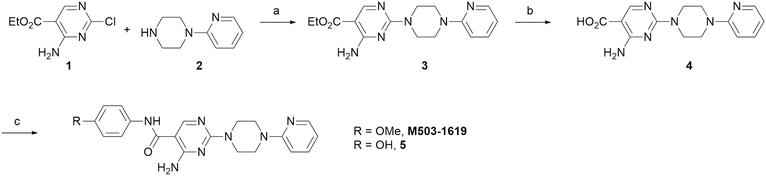
The route for the synthesis of M503-1619 and its precursor 5 for radiolabeling studies with tritium and carbon-11 is shown in Scheme 1. Coupling the commercially available compound 1 and 1-(pyridin-2-yl)piperazine 2 using triethylamine (Et3N) as base afforded intermediate 3 in moderate yield. Hydrolysis of the ester of 3 generated the carboxylic acid 4. This acid coupled with 4-methoxyaniline or 4-aminophenol delivered M503-1619 and its precursor 5 in moderate yield.2.4. In vitro saturation binding assays
To confirm the affinity and selectivity of M503-1619 for α-Syn versus Aβ and tau, the corresponding tritiated compound was synthesized for direct binding assays. Saturation binding assays of [3H]M503-1619 in different human brain tissue homogenates (Table 3) confirmed the high binding affinity to synthetic α-Syn fibrils (Kd = 4.4 nM) and PD brain tissue homogenates (Kd = 2.5 nM). These results are consistent with the data from the competition binding assays. [3H]M503-1619 had a relatively low affinity for AD (Kd = 37 nM), progressive supranuclear palsy (PSP; Kd = 55 nM), and corticobasal degeneration (CBD; Kd = 94) tissues. The results of this study support the selectivity of this radioligand for α-Syn found in PD brain.| Tissue | K d (nM) | B max (nM) |
|---|---|---|
| a The Kd values are expressed as the mean ± SD by three individual experiments, all assays were performed in triplicate. | ||
| α-Syn fibrils (n = 4) | 4.4 ± 1.0 | 1156 ± 112 |
| PD (n = 4) | 2.5 ± 0.3 | 59 ± 10 |
| AD (n = 4) | 37 ± 2 | 160 ± 28 |
| PSP (n = 4) | 55 ± 3 | 399 ± 24 |
| CBD (n = 3) | 94 ± 13 | 540 ± 130 |
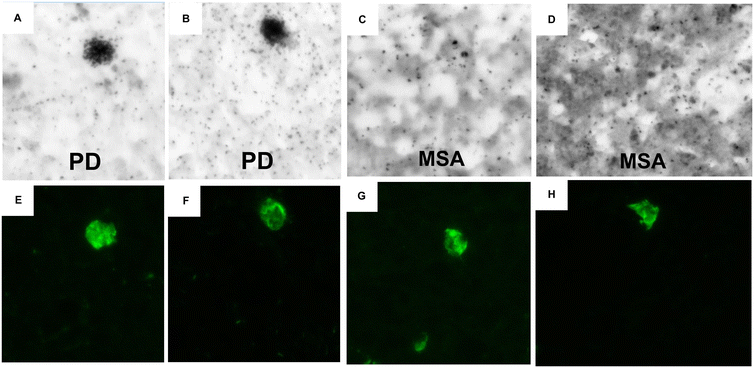
2.5. Autoradiography
A previous study from our lab demonstrated that the site 9 ligand, [3H]HY-2-15 (Fig. 1) showed preferential binding α-Syn in postmortem brain sections of MSA and not PD.31 An earlier study with a fluorescent probe displayed the opposite specificity, labeling α-Syn in Lewy bodies of PD brain and not the glial cytoplasmic inclusions (GCIs) in MSA brain.32 Therefore, nuclear emulsion autoradiography of [3H]M503-1619 was performed on PD and MSA tissue sections to assess if it has specificity for the different forms of α-Syn (Fig. 2). High-resolution autoradiography visualized target engagement of [3H]M503-1619 to Lewy bodies in PD brain (Fig. 2A and B) but not to GCIs present in MSA (Fig. 2C and D). α-Syn neuropathology visualized using anti-pS129 α-Syn immunohistochemistry (IHC) and high-resolution autoradiography with [3H]M503-1619 on the same section showed a clear overlap in the PD brain but not in the MSA (Fig. 2E–H). These data suggest that [3H]M503-1619 may be capable of differentiating α-Syn in PD versus MSA.To further confirm these results from nuclear emulsion autoradiography, autoradiography studies (Fig. 3) were conducted in tissue sections from PD (hippocampus) and MSA (basal ganglia) cases. These images clearly showed a higher binding of the radioligand [3H]M503-1619 in PD (Fig. 3A) than in MSA (Fig. 3E), even though both brain specimens have a high burden of α-Syn measured with the antibody labeling conditions described above (Fig. 3B–D, and F–H). These results are consistent with the nuclear emulsion autoradiography and suggest that [3H]M503-1619 binds preferentially to α-Syn in PD and not MSA.
2.6. Radiosynthesis and PET imaging studies in NHPs
NHP PET imaging studies were conducted with [11C]M503-1619 to determine the brain uptake and rate of washout of the radiotracer from brain. [11C]M503-1619 was synthesized via O-alkylation of the phenol group of 5 with 11CH3I (Scheme 2).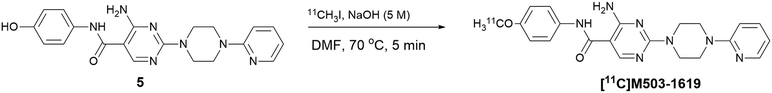 | ||
| Scheme 2 The procedure for radiosynthesis of [11C]M503-1619. Conditions: 11CH3I, 5 M NaOH, DMF, 70 °C, 5 min. | ||
The radiosynthesis required ∼30 min to complete and [11C]M503-1619 was prepared in radiochemical yield of 56–63% (decay corrected to end of bombardment (EOB)) and a molar activity of ∼1187 GBq μmol−1 (decay corrected to EOB).
PET imaging studies were conducted in two different species of NHPs, rhesus macaque and cynomolgus macaque. A total-body PET imaging study was conducted in a rhesus macaque on the PennPET Explorer33 scanner at the University of Pennsylvania (Fig. 4), and a brain imaging study in a cynomolgus macaque was conducted on a Siemens Focus 220 system at Washington University in St. Louis (Fig. S1†). The PET images with [11C]M503-1619 showed high initial brain uptake. The peak uptake of the whole brain occurred at 2–3 min with a standardized uptake value (SUV) of 2.5 in rhesus monkey (Fig. 4) and 2.3 in cynomolgus monkey (Fig. S1†). Meanwhile, there was a rapid washout of [11C]M503-1619 from brain. The whole brain peak to 60 min ratio was 3.2 in both rhesus and cynomolgus monkeys. These study results demonstrated that the high initial brain uptake of [11C]M503-1619 and rapid washout makes it a suitable candidate for further in vivo PET image studies. Venous blood samples were analyzed for metabolites, which largely consisted of polar metabolites (Fig. S2 and Table S1†). [11C]M503-1619 was rapidly metabolized, with a parent fraction of 57% by 7.5 minutes post-injection, presenting a potential limitation to long duration scans and image quantification using blood-input based pharmacokinetic modeling.
3. Conclusions
[11C]M503-1619 has a high affinity for α-Syn, moderate affinity for Aβ, and 10–20-fold selectivity vs. tau. Autoradiography studies suggest that it binds α-Syn in PD more robustly than in the MSA brain. The radiotracer has high brain uptake and rapid washout from the macaque brain. Our data suggest that M503-1619 has suitable properties as a lead compound for a PET radiotracer development. Additional structure–activity relationship studies are ongoing with M503-1619 to identify a suitable compound for translational imaging studies in PD patients.4. Experimental
4.1. In silico compound screen
The 42 million compound database from Mcule, Inc. was used for the initial screen against the site 9 exemplar that was generated from our prior work.18 The in silico conformers generation for compound library and the ultrahigh throughput screening were performed on the high-performance computing cluster at the Center for Biomedical Image Computing and Analytics at the University of Pennsylvania. Three-dimensional conformations of the compound database were generated using OMEGA (OpenEye Scientific Software, Inc.). Molecular alignment was performed using ROCS (OpenEye Scientific Software, Inc.). The top 3 million compounds as quantified by the sum Tanimoto coefficient of shape and pharmacophore similarities were retained for the secondary in silico screening. Compounds consisting of PAINS motifs, CNS MPO score lower than 4, number of aromatic ring lower than 2 or higher than 5, molecular weight higher than 500, and cLog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D lower than 1 or higher than 3.5 were excluded from the initial in silico hits for the final visual inspection. Visual inspection was performed by two experienced chemists, and compounds were selected based on the linearity of the structure, the number of aromatic rings, and the presence and position of nitrogen atoms (i.e., heteroaromatic rings) and substituents on the aromatic rings that could facilitate radiolabeling.
D lower than 1 or higher than 3.5 were excluded from the initial in silico hits for the final visual inspection. Visual inspection was performed by two experienced chemists, and compounds were selected based on the linearity of the structure, the number of aromatic rings, and the presence and position of nitrogen atoms (i.e., heteroaromatic rings) and substituents on the aromatic rings that could facilitate radiolabeling.
4.2. General
Commercial reagents and solvents were purchased from sources, including Sigma-Aldrich (MO, U.S.A.), Synthonix (NC, U.S.A.), Enamine (Kyiv, UA), Ambeed (IL, U.S.A), COMBI-BLOCKS (CA, U.S.A) or Thermo Fisher Scientific (MA, U.S.A.), and used as received. Thin layer chromatography (TLC) was performed using TLC silica gel 60 W F254S plates for monitoring reactions, and the spots were visualized under UV light (254 nm). Flash chromatography purifications were conducted on a Biotage Isolera One system (Uppsala, Sweden). 1H and 13C NMR spectra were recorded on a Bruker NEO-400 spectrometer (Bruker, MA, U.S.A.) with chemical shifts (δ) reported in parts per million (ppm) relative to the deuterated solvent as an internal reference. Mass spectra (m/z) were acquired on a 2695 Alliance LC-MS (Waters Corporation, MA, U.S.A.) with positive electrospray ionization (ESI), and high-resolution mass spectra (HRMS, m/z) were obtained using a waters LCT premier mass spectrometer (Waters Corporation, MA, U.S.A.). Recombinant α-Syn fibrils preparation including α-Syn protein expression and purification was performed according to the literature.32 The synthesis of [3H]M503-1619 is provided in the ESI† and was conducted by Novandi Chemistry AB (Södertälje, Sweden).4.3. Chemistry: the procedure for the synthesis of M503-1619 and its precursor 5
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 1
1 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to afford the desired product 3 as a white solid. 1H NMR (400 MHz, DMSO-d6)δ 8.52 (s, 1H), 8.12 (ddd, J = 4.9, 2.0, 0.8 Hz, 1H), 7.55 (ddd, J = 8.9, 7.1, 2.0 Hz, 1H), 7.47 (s, 1H), 7.41 (s, 1H), 6.86 (d, J = 8.6 Hz, 1H), 6.66 (dd, J = 6.7, 4.5 Hz, 1H), 4.21 (q, J = 7.1 Hz, 2H), 1.26 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO-d6)δ 166.1, 162.9, 161.5, 160.6, 158.9, 147.7, 137.8, 113.4, 107.4, 95.1, 59.9, 44.5, 43.0, 14.4. LCMS: 329 [M + H]+; HRMS (ESI) calculated for C16H21N6O2+ ([M + H+]) 329.1721, found: 329.1745. White solid, 100 mg, 51%.
1) to afford the desired product 3 as a white solid. 1H NMR (400 MHz, DMSO-d6)δ 8.52 (s, 1H), 8.12 (ddd, J = 4.9, 2.0, 0.8 Hz, 1H), 7.55 (ddd, J = 8.9, 7.1, 2.0 Hz, 1H), 7.47 (s, 1H), 7.41 (s, 1H), 6.86 (d, J = 8.6 Hz, 1H), 6.66 (dd, J = 6.7, 4.5 Hz, 1H), 4.21 (q, J = 7.1 Hz, 2H), 1.26 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO-d6)δ 166.1, 162.9, 161.5, 160.6, 158.9, 147.7, 137.8, 113.4, 107.4, 95.1, 59.9, 44.5, 43.0, 14.4. LCMS: 329 [M + H]+; HRMS (ESI) calculated for C16H21N6O2+ ([M + H+]) 329.1721, found: 329.1745. White solid, 100 mg, 51%.
A solution of the crude product 4, DIPEA (77 mg, 0.6 mmol), and HBTU (80 mg, 0.21 mmol) in 4 mL DMF was stirred for 15 min at room temperature. Either 4-methoxyaniline (26 mg, 0.21 mmol) or 4-aminophenol (23 mg, 0.21 mmol) was added to the solution. The mixture was stirred at room temperature overnight, washed with 1 N NaHCO3 solution extracted with DCM (3 × 25 mL), and dried over Na2SO4. The solvents were removed, and the residue was purified by flash column chromatography on silica gel, eluting with DCM/7 N NH3 in MeOH = 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 to afford M503-1619 or its precursor 5 as white solids.34
5 to afford M503-1619 or its precursor 5 as white solids.34
4.4. In vitro radioligand binding assays
4.5. In vitro real-time autoradiography
In vitro autoradiography was performed using 10 μm-thick fresh frozen sections derived from PD and MSA brains. Brain sections were first equilibrated for 30 min in 1× PBS (Dulbecco's phosphate buffered saline) and then incubated for 90 min at room temperature with 5 nM of [3H]M503-1619. The sections were rinsed two times in cold buffer 1× (PBS + 20% EtOH) for 5 minutes, followed by a quick dip in cold distillated water. Non-specific binding was determined using 1 μM unlabeled M503-1619. Slides were then allowed to air-dry before being exposed and scanned in a real-time autoradiography system (BeaQuant instrument, ai4R, Nantes, France) for 2 h. ROI delineation and quantification of signal were performed using image analysis software, Beamage ai4R. Specific binding was determined by subtracting the non-specific signal (NSB) from the total signal and expressed as counts min−1 mm−2. Immunohistochemistry was performed on sections adjacent to those used for autoradiography. Sections were fixed in ethanol for 15 min, washed 3 × 5 min in PBS Tween-20 buffer, permeabilized with 0.1% Triton X-100 for 10 min. Sections underwent hydrogen peroxide blocking for 15 min and PBS + 10% goat serum + 1% BSA + 0.1% Tween 20 blocking for 1 h at room temperature. Sections were immunostained using primary antibody alpha synuclein (phospho S129) antibody (p-Syn/81A), used at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 dilution overnight at 4 °C. After a series of thorough washes with PBS Tween 20 buffer, the slides were incubated with the secondary antibodies, Alexa Fluor 488at 1
500 dilution overnight at 4 °C. After a series of thorough washes with PBS Tween 20 buffer, the slides were incubated with the secondary antibodies, Alexa Fluor 488at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution for 1 h at RT. Subsequently, sections were mounted with Duolink in situ mounting media with DAPI and coverslips for microscopy. Images were captured with a Zeiss microscope at 10× magnification.
1000 dilution for 1 h at RT. Subsequently, sections were mounted with Duolink in situ mounting media with DAPI and coverslips for microscopy. Images were captured with a Zeiss microscope at 10× magnification.
4.6. Nuclear emulsion autoradiography
Frozen postmortem PD and MSA brain tissue samples were sectioned on a cryostat at 16 μm and mounted onto glass slides and stored at −80 °C. To quench lipofuscin autofluorescence, slides were incubated in 1× TrueBlack (Biotium) in 70% EtOH. Slides were incubated sequentially with anti-phospho–α-Syn (S129) antibody (p-Syn/81A, BioLegend) and Alexa Fluor 488 goat anti-mouse secondary antibody to immunostain LBs and LNs. Slides were then incubated in 5 nM or 15 nM of [3H]M503-1619 diluted in 30 mM Tris–HCl, pH 7.4 for 2 h at 37 °C. The slides were washed in 30 mM Tris–HCl, pH 7.4 for 1 min, 50% ethanol/30 mM Tris–HCl, pH 7.4 for 1 min, 30% ethanol/30 mM Tris–HCl, pH 7.4 for 1 min, 30 mM Tris–HCl, pH 7.4 for 1 min, and MQH2O for 1 s. The slides were allowed to completely air dry in a fume hood. To obtain high resolution autoradiographic information, the slides were dipped in autoradiography emulsion (Carestream Type NTB) and incubated in the dark for 4 days. The slides were developed in Ilford Phenisol Developer for 4 min, 200 mL of 1% acetic acid for 2 min, 200 mL of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 Ilford Hypam Fixer for 4 min, and 200 mL of MQH2O for 4 min (2×). The slides were allowed to completely air dry in the fume hood before a coverslip was applied with Fluoromount-G mounting medium. Photomicrographs of fluorescent antibody staining and tritium activated silver grains were obtained on a Nikon Eclipse TE2000-U Inverted Fluorescence Microscope (Nikon, Tokyo, Japan) microscope using fluorescence and brightfield/monochrome light, respectively.
4 Ilford Hypam Fixer for 4 min, and 200 mL of MQH2O for 4 min (2×). The slides were allowed to completely air dry in the fume hood before a coverslip was applied with Fluoromount-G mounting medium. Photomicrographs of fluorescent antibody staining and tritium activated silver grains were obtained on a Nikon Eclipse TE2000-U Inverted Fluorescence Microscope (Nikon, Tokyo, Japan) microscope using fluorescence and brightfield/monochrome light, respectively.
4.7. Radiosynthesis of [11C]M503-1619
The radiolabeling of [11C]M503-1619 was performed using an automated synthesizer, Synthra MeIPlus (Synthra GmbH, Germany). The precursor 5 (1 mg, 2.5 μmol) was dissolved in 400 μL anhydrous DMF and 3 μL of 5 M NaOH solution. Radioactive [11C]CH3I gas was bubbled into the solution within 2–4 minutes. Then the reaction vial was sealed and heated at 70 °C for 5 minutes with stirring. The reaction was quenched with 1.3 mL of HPLC mobile phase (acetonitrile/ammonium formate solution 51/49 (v/v), pH = ∼4.5); and then loaded onto a semi-preparative HPLC system equipped with a C18 column (Agilent C18 column, 250 × 9.6 mm, 5 μM) and purified with a mobile phase flow rate of 4.0 mL min−1. A UV detector set for a wavelength of 254 nm permitted purification. The target radioactive product was collected from 7 to 8 minutes using a glass vial prefilled with 50 mL Milli-Q water, then the solution was passed through a C-18 Sep-Pak cartridge (Part No. WAT020515, Waters Corporation, Milford, MA). After using 15 mL of Milli-Q water to rinse the C-18 cartridge, the radioactive product was eluted into a dose vial using 0.6 mL anhydrous ethanol, followed by diluting with 5.4 mL of saline to formulate the injection dose solution for administration to an animal.An aliquot of the dose sample (20 μL) was injected into the analytic HPLC system for authentication by co-injection with the cold standard compound M503-1619, the chemical and radiochemical purity, and molar activity of the final dose were determined. The analytic HPLC system for quality control consists of a reversed-phase HPLC analytical column (Agilent Zorbax SB-C18, 4.6 mm × 250), UV wavelength of 254 nm, mobile phase of acetonitrile/0.1 M ammonium formate buffer (v/v, 75/25, pH = 4.5) with a flow rate of 1.0 mL min−1. Under this condition, [11C]M503-1619 has a retention time of TR = 4.0 min, 56–63% radiochemical yield (decay corrected to EOB), >1187 GBq μmol−1 molar activity (decay corrected to EOB), >95% chemical purity and >99% radiochemical purity.
4.8. Radiometabolite study of [11C]M503-1619
Venous blood samples were collected throughout the PET scans to measure the radiometabolite profile. Blood samples were centrifuged for 5 minutes at 3000g to separate plasma. Plasma samples were treated with acetonitrile at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 by volume and centrifuged to recover deproteinated plasma supernatant. The supernatant was injected into an Agilent Infinity 1280 HPLC (Agilent Technologies, Santa Clara, CA) with a reverse-phase Agilent Zorbax XDB-C18 analytical HPLC column (4.6 mm × 150) and a mobile phase of 65% 0.1 M ammonium formate buffer and 35% acetonitrile at 1.2 mL min−1. Parent and metabolite activity fractions were measured with an in-line Posi-RAM detector (LabLogic Systems, Chantilly, VA) based on retention times of M503-1619 standards.
1 by volume and centrifuged to recover deproteinated plasma supernatant. The supernatant was injected into an Agilent Infinity 1280 HPLC (Agilent Technologies, Santa Clara, CA) with a reverse-phase Agilent Zorbax XDB-C18 analytical HPLC column (4.6 mm × 150) and a mobile phase of 65% 0.1 M ammonium formate buffer and 35% acetonitrile at 1.2 mL min−1. Parent and metabolite activity fractions were measured with an in-line Posi-RAM detector (LabLogic Systems, Chantilly, VA) based on retention times of M503-1619 standards.
4.9. PET studies of [11C]M503-1619 in NHPs
A filtered back projection method was used to reconstruct PET images (volume size: 12 × 128 × 95, voxel size: 1.898 × 1.898 × 0.796 mm3 in the x, y, and z direction) with dead time, scatter, and attenuation correction. To quantify the total tracer uptake in the brain, dynamic PET images were co-registered to a standardized monkey MRI template using PMOD software 4.02 (PMOD technologies, Zürich, Switzerland). A whole brain VOI was applied to the co-registered PET image to obtain the brain TAC. The uptake of radioactivity was normalized to body weight and the injected dose to obtain a SUV.
Data availability
The solid state NMR structure and exemplar for site 9 in alpha synuclein have been published previously.18 The molecular alignment was performed using ROCS from Open Eye Scientific Software, Inc. (Albuquerque, NM). The compound database used in the in silico screen was commercially available from Mcule, Inc. (https://mcule.com/database/).Author contributions
Conceptualization: R. H. M. Funding acquisition: R. H. M. Investigation, data curation, formal analysis, methodology and software: G.-L. T., C.-J. H., D. S. G., T. J. A. G., Z. L.-Z., C. T., W. K. C, A. J. Y., J.-G. C., K. P., T. H., H. J, Y. Y., C. H., S. G., H. L. and J. O. Project administration: C. A. M. and R. H. M. Resource, supervision, and validation: E. J. P, P. K., Z. T., C. A. M. and R. H. M. Visualization: C.-J. H. and R. H. M. Writing – original draft: G.-L. T., C-J. H. and R. H. M. Writing – review & editing: E. J. P., P. K., Z. T., C. A. M. and R. H. M.Conflicts of interest
There are no conflicts to declare.Acknowledgements
The authors would like to thank Mr. Mark Schneider for his excellent technical assistance. This work was funded by NIH grant U19-NS110456. The in silico studies were conducted on the high-performance Biomedical Image Computing and Informatics Cluster at the University of Pennsylvania, Center for Biomedical Image Computing and Analytics supported by the National Institutes of Health. We thank the University of Pennsylvania Physics and Instrumentation Group for the assistance with PennPET Explorer imaging studies.References
- E. R. Dorsey, T. Sherer, M. S. Okun and B. R. Bloem, J. Parkinson's Dis., 2018, 8, S3–S8 CrossRef PubMed.
- V. L. Feigin, A. A. Abajobir, K. H. Abate, F. Abd-Allah, A. M. Abdulle, S. F. Abera, G. Y. Abyu, M. B. Ahmed, A. N. Aichour, I. Aichour, M. T. E. Aichour, R. O. Akinyemi, S. Alabed, R. Al-Raddadi, N. Alvis-Guzman, A. T. Amare, H. Ansari, P. Anwari, J. Ärnlöv, H. Asayesh, S. W. Asgedom, T. M. Atey, L. Avila-Burgos, E. Frinel, G. A. Avokpaho, M. R. Azarpazhooh, A. Barac, M. Barboza, S. L. Barker-Collo, T. Bärnighausen, N. Bedi, E. Beghi, D. A. Bennett, I. M. Bensenor, A. Berhane, B. D. Betsu, S. Bhaumik, S. M. Birlik, S. Biryukov, D. J. Boneya, L. N. B. Bulto, H. Carabin, D. Casey, C. A. Castañeda-Orjuela, F. Catalá-López, H. Chen, A. A. Chitheer, R. Chowdhury, H. Christensen, L. Dandona, R. Dandona, G. A. de Veber, S. D. Dharmaratne, H. P. Do, K. Dokova, E. R. Dorsey, R. G. Ellenbogen, S. Eskandarieh, M. S. Farvid, S.-M. Fereshtehnejad, F. Fischer, K. J. Foreman, J. M. Geleijnse, R. F. Gillum, G. Giussani, E. M. Goldberg, P. N. Gona, A. C. Goulart, H. C. Gugnani, R. Gupta, V. Hachinski, R. Gupta, R. R. Hamadeh, M. Hambisa, G. J. Hankey, H. A. Hareri, R. Havmoeller, S. I. Hay, P. Heydarpour, P. J. Hotez, M. B. Jakovljevic, M. Javanbakht, P. Jeemon, J. B. Jonas, Y. Kalkonde, A. Kandel, A. Karch, A. Kasaeian, A. Kastor, P. N. Keiyoro, Y. S. Khader, I. A. Khalil, E. A. Khan, Y.-H. Khang, A. Tawfih, A. Khoja, J. Khubchandani, C. Kulkarni, D. Kim, Y. J. Kim, M. Kivimaki, Y. Kokubo, S. Kosen, M. Kravchenko, R. V. Krishnamurthi, B. Kuate Defo, G. A. Kumar, R. Kumar, H. H. Kyu, A. Larsson, P. M. Lavados, Y. Li, X. Liang, M. L. Liben, W. D. Lo, G. Logroscino, P. A. Lotufo, C. T. Loy, M. T. Mackay, H. M. A. El Razek, M. M. A. El Razek, A. Majeed, R. Malekzadeh, T. Manhertz, L. G. Mantovani, J. Massano, M. Mazidi, C. McAlinden, S. Mehata, M. M. Mehndiratta, Z. A. Memish, W. Mendoza, M. A. Mengistie, G. A. Mensah, A. Meretoja, H. B. Mezgebe, T. R. Miller, S. R. Mishra, N. M. Ibrahim, A. Mohammadi, K. E. Mohammed, S. Mohammed, A. H. Mokdad, M. Moradi-Lakeh, I. M. Velasquez, K. I. Musa, M. Naghavi, J. W. Ngunjiri, C. T. Nguyen, G. Nguyen, Q. L. Nguyen, T. H. Nguyen, E. Nichols, D. N. A. Ningrum, V. M. Nong, B. Norrving, J. J. N. Noubiap, F. A. Ogbo, M. O. Owolabi, J. D. Pandian, P. G. Parmar, D. M. Pereira, M. Petzold, M. R. Phillips, M. A. Piradov, R. G. Poulton, F. Pourmalek, M. Qorbani, A. Rafay, M. Rahman, M. H. Rahman, R. K. Rai, S. Rajsic, A. Ranta, S. Rawaf, A. M. N. Renzaho, M. S. Rezai, G. A. Roth, G. Roshandel, E. Rubagotti, P. Sachdev, S. Safiri, R. Sahathevan, M. A. Sahraian, A. M. Samy, P. Santalucia, I. S. Santos, B. Sartorius, M. Satpathy, M. Sawhney, M. I. Saylan, S. G. Sepanlou, M. A. Shaikh, R. Shakir, M. Shamsizadeh, K. N. Sheth, M. Shigematsu, H. Shoman, D. A. S. Silva, M. Smith, E. Sobngwi, L. A. Sposato, J. D. Stanaway, D. J. Stein, T. J. Steiner, L. J. Stovner, R. S. Abdulkader, C. E. I. Szoeke, R. Tabarés-Seisdedos, D. Tanne, A. M. Theadom, A. G. Thrift, D. L. Tirschwell, R. Topor-Madry, B. X. Tran, T. Truelsen, K. B. Tuem, K. N. Ukwaja, O. A. Uthman, Y. Y. Varakin, T. Vasankari, N. Venketasubramanian, V. V. Vlassov, F. Wadilo, T. Wakayo, M. T. Wallin, E. Weiderpass, R. Westerman, T. Wijeratne, C. S. Wiysonge, M. A. Woldu, C. D. A. Wolfe, D. Xavier, G. Xu, Y. Yano, H. H. Yimam, N. Yonemoto, C. Yu, Z. Zaidi, M. E. S. Zaki, J. R. Zunt, C. J. L. Murray and T. Vos, Lancet Neurol., 2017, 16, 877–897 CrossRef PubMed.
- M. Baba, S. Nakajo, P. H. Tu, T. Tomita, K. Nakaya, V. M. Lee, J. Q. Trojanowski and T. Iwatsubo, Am. J. Pathol, 1998, 152, 879–884 CAS.
- M. G. Spillantini, M. L. Schmidt, V. M. Lee, J. Q. Trojanowski, R. Jakes and M. Goedert, Nature, 1997, 388, 839–840 CrossRef CAS PubMed.
- C. M. Clark, J. A. Schneider, B. J. Bedell, T. G. Beach, W. B. Bilker, M. A. Mintun, M. J. Pontecorvo, F. Hefti, A. P. Carpenter, M. L. Flitter, M. J. Krautkramer, H. F. Kung, R. E. Coleman, P. M. Doraiswamy, A. S. Fleisher, M. N. Sabbagh, C. H. Sadowsky, E. M. Reiman, S. P. Zehntner, D. M. Skovronsky and T. A.-A. S. Group, JAMA, J. Am. Med. Assoc., 2011, 305, 275–283 CrossRef CAS PubMed.
- W. E. Klunk, H. Engler, A. Nordberg, Y. Wang, G. Blomqvist, D. P. Holt, M. Bergström, I. Savitcheva, G.-F. Huang, S. Estrada, B. Ausén, M. L. Debnath, J. Barletta, J. C. Price, J. Sandell, B. J. Lopresti, A. Wall, P. Koivisto, G. Antoni, C. A. Mathis and B. Långström, Ann. Neurol., 2004, 55, 306–319 CrossRef CAS PubMed.
- C.-F. Xia, J. Arteaga, G. Chen, U. Gangadharmath, L. F. Gomez, D. Kasi, C. Lam, Q. Liang, C. Liu, V. P. Mocharla, F. Mu, A. Sinha, H. Su, A. K. Szardenings, J. C. Walsh, E. Wang, C. Yu, W. Zhang, T. Zhao and H. C. Kolb, Alzheimer's Dementia, 2013, 9, 666–676 CrossRef PubMed.
- J. L. Eberling, K. D. Dave and M. A. Frasier, J. Parkinson's Dis., 2013, 3, 565–567 CrossRef CAS PubMed.
- C. A. Mathis, B. J. Lopresti, M. D. Ikonomovic and W. E. Klunk, Semin. Nucl. Med., 2017, 47, 553–575 CrossRef PubMed.
- D. P. Bagchi, L. Yu, J. S. Perlmutter, J. Xu, R. H. Mach, Z. Tu and P. T. Kotzbauer, PLoS One, 2013, 8, e55031 CrossRef CAS PubMed.
- E. T. L'Estrade, M. Erlandsson, F. G. Edgar, T. Ohlsson, G. M. Knudsen and M. M. Herth, Neuropharmacology, 2020, 172, 107830 CrossRef PubMed.
- M. Shah, J. Seibyl, A. Cartier, R. Bhatt and A. M. Catafau, J. Nucl. Med., 2014, 55, 1397–1400 CrossRef CAS PubMed.
- R. Smith, F. Capotosti, M. Schain, T. Ohlsson, E. Vokali, J. Molette, T. Touilloux, V. Hliva, I. K. Dimitrakopoulos, A. Puschmann, J. Jögi, P. Svenningsson, M. Andréasson, C. Sandiego, D. S. Russell, P. Miranda-Azpiazu, C. Halldin, E. Stomrud, S. Hall, K. Bratteby, E. Tampio L'Estrade, R. Luthi-Carter, A. Pfeifer, M. Kosco-Vilbois, J. Streffer and O. Hansson, Nat. Commun., 2023, 14, 6750 CrossRef CAS PubMed.
- J. Xiang, Y. Tao, Y. Xia, S. Luo, Q. Zhao, B. Li, X. Zhang, Y. Sun, W. Xia, M. Zhang, S. S. Kang, E.-H. Ahn, X. Liu, F. Xie, Y. Guan, J. J. Yang, L. Bu, S. Wu, X. Wang, X. Cao, C. Liu, Z. Zhang, D. Li and K. Ye, Cell, 2023, 186(3350–3367), e3319 Search PubMed.
- H. Wood, Nat. Rev. Neurol., 2013, 9, 599 CrossRef PubMed.
- H. Endo, M. Ono, Y. Takado, K. Matsuoka, M. Takahashi, K. Tagai, Y. Kataoka, K. Hirata, K. Takahata, C. Seki, N. Kokubo, M. Fujinaga, W. Mori, Y. Nagai, K. Mimura, K. Kumata, T. Kikuchi, A. Shimozawa, S. K. Mishra, Y. Yamaguchi, H. Shimizu, A. Kakita, H. Takuwa, H. Shinotoh, H. Shimada, Y. Kimura, M. Ichise, T. Suhara, T. Minamimoto, N. Sahara, K. Kawamura, M.-R. Zhang, M. Hasegawa and M. Higuchi, Neuron, 2024, 599 Search PubMed.
- C. J. Hsieh, J. J. Ferrie, K. Xu, I. Lee, T. J. A. Graham, Z. Tu, J. Yu, D. Dhavale, P. Kotzbauer, E. J. Petersson and R. H. Mach, ACS Chem. Neurosci., 2018, 9, 2521–2527 CrossRef CAS PubMed.
- J. J. Ferrie, Z. Lengyel-Zhand, B. Janssen, M. G. Lougee, S. Giannakoulias, C. J. Hsieh, V. V. Pagar, C. C. Weng, H. Xu, T. J. A. Graham, V. M. Lee, R. H. Mach and E. J. Petersson, Chem. Sci., 2020, 11, 12746–12754 RSC.
- D. K. Johnson and J. Karanicolas, PLoS Comput. Biol., 2013, 9, e1002951 CrossRef CAS PubMed.
- B. Sonustun, M. F. Altay, C. Strand, K. Ebanks, G. Hondhamuni, T. T. Warner, H. A. Lashuel and R. Bandopadhyay, Cells, 2022, 11, 906 CrossRef CAS PubMed.
- K. Zhao, Y. J. Lim, Z. Liu, H. Long, Y. Sun, J. J. Hu, C. Zhao, Y. Tao, X. Zhang, D. Li, Y. M. Li and C. Liu, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 20305–20315 CrossRef CAS PubMed.
- S. X. Pancoe, Y. J. Wang, M. Shimogawa, R. M. Perez, S. Giannakoulias and E. J. Petersson, J. Mol. Biol., 2022, 434, 167859 CrossRef CAS PubMed.
- S. Zhang, R. Zhu, B. Pan, H. Xu, M. F. Olufemi, R. J. Gathagan, Y. Li, L. Zhang, J. Zhang, W. Xiang, E. M. Kagan, X. Cao, C. Yuan, S.-J. Kim, C. K. Williams, S. Magaki, H. V. Vinters, H. A. Lashuel, B. A. Garcia, E. J. Petersson, J. Q. Trojanowski, V. M.-Y. Lee and C. Peng, Nat. Neurosci., 2023, 26, 213–225 CrossRef CAS PubMed.
- J. B. Baell and G. A. Holloway, J. Med. Chem., 2010, 53, 2719–2740 CrossRef CAS PubMed.
- T. T. Wager, X. Hou, P. R. Verhoest and A. Villalobos, ACS Chem. Neurosci., 2016, 7, 767–775 CrossRef CAS PubMed.
- A. M. Walji, E. D. Hostetler, H. Selnick, Z. Zeng, P. Miller, I. Bennacef, C. Salinas, B. Connolly, L. Gantert, M. Holahan, S. O'Malley, M. Purcell, K. Riffel, J. Li, J. Balsells, J. A. OBrien, S. Melquist, A. Soriano, X. Zhang, A. Ogawa, S. Xu, E. Joshi, J. D. Rocca, F. J. Hess, J. Schachter, D. Hesk, D. Schenk, A. Struyk, K. Babaoglu, T. G. Lohith, Y. Wang, K. Yang, J. Fu, J. L. Evelhoch and P. J. Coleman, J. Med. Chem., 2016, 59, 4778–4789 CrossRef CAS PubMed.
- H. Kroth, F. Oden, J. Molette, H. Schieferstein, F. Capotosti, A. Mueller, M. Berndt, H. Schmitt-Willich, V. Darmency, E. Gabellieri, C. Boudou, T. Juergens, Y. Varisco, E. Vokali, D. T. Hickman, G. Tamagnan, A. Pfeifer, L. Dinkelborg, A. Muhs and A. Stephens, Eur. J. Nucl. Med. Mol. Imaging, 2019, 46, 2178–2189 CrossRef CAS PubMed.
- T. J. Graham, A. Lindberg, J. Tong, J. S. Stehouwer, N. Vasdev, R. H. Mach and C. A. Mathis, J. Med. Chem., 2023, 66, 10628–10638 CrossRef CAS PubMed.
- A. Lindberg, A. C. Knight, D. Sohn, L. Rakos, J. Tong, A. Radelet, N. S. Mason, J. S. Stehouwer, B. J. Lopresti, W. E. Klunk, J. Sandell, A. Sandberg, P. Hammarström, S. Svensson, C. A. Mathis and N. Vasdev, ACS Chem. Neurosci., 2021, 12, 596–602 CrossRef CAS PubMed.
- C. A. Mathis, N. S. Mason, B. J. Lopresti and W. E. Klunk, Semin. Nucl. Med., 2012, 42, 423–432 CrossRef PubMed.
- H. Y. Kim, W. K. Chia, C.-J. Hsieh, D. S. Guarino, T. J. Graham, Z. Lengyel-Zhand, M. Schneider, C. Tomita, M. G. Lougee, H. J. Kim, V. V. Pagar, H. Lee, C. Hou, B. A. Garcia, E. J. Petersson, J. O'Shea, P. T. Kotzbauer, C. A. Mathis, V. M.-Y. Lee, K. C. Luk and R. H. Mach, J. Med. Chem., 2023, 66, 12185–12202 CrossRef CAS PubMed.
- Z. Lengyel-Zhand, J. J. Ferrie, B. Janssen, C. J. Hsieh, T. Graham, K. Y. Xu, C. M. Haney, V. M. Lee, J. Q. Trojanowski, E. J. Petersson and R. H. Mach, Chem. Commun., 2020, 56, 3567–3570 RSC.
- B. Dai, M. E. Daube-Witherspoon, S. McDonald, M. E. Werner, M. J. Parma, M. J. Geagan, V. Viswanath and J. S. Karp, Phys. Med. Biol., 2023, 68, 095007 CrossRef CAS PubMed.
- T. Ryckmans, A. A. Aubdool, J. V. Bodkin, P. Cox, S. D. Brain, T. Dupont, E. Fairman, Y. Hashizume, N. Ishii, T. Kato, L. Kitching, J. Newman, K. Omoto, D. Rawson and J. Strover, Bioorg. Med. Chem. Lett., 2011, 21, 4857–4859 CrossRef CAS PubMed.
- W. E. Klunk, Y. Wang, G. F. Huang, M. L. Debnath, D. P. Holt, L. Shao, R. L. Hamilton, M. D. Ikonomovic, S. T. DeKosky and C. A. Mathis, J. Neurosci., 2003, 23, 2086–2092 CrossRef CAS PubMed.
- W. E. Klunk, Y. Wang, G.-f. Huang, M. L. Debnath, D. P. Holt and C. A. Mathis, Life Sci., 2001, 69, 1471–1484 CrossRef CAS PubMed.
- A. R. Pantel, V. Viswanath, M. E. Daube-Witherspoon, J. G. Dubroff, G. Muehllehner, M. J. Parma, D. A. Pryma, E. K. Schubert, D. A. Mankoff and J. S. Karp, J. Nucl. Med., 2020, 61, 144–151 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d5md00057b |
| This journal is © The Royal Society of Chemistry 2025 |