Open Access Article
Open Access ArticleOpenLM: an open-source pixel super-resolution platform for lens-free microscopy with applications in bacterial growth monitoring and deep learning-based bacterial detection
Weiming
Xu
 ab,
Samiha
Ahmed
ab,
Majed
Althumayri
ab,
Samiha
Ahmed
ab,
Majed
Althumayri
 abd,
Azra Yaprak
Tarman
ab,
Mert Kerem
Ulku
ab,
Karston
Yong
abd,
Azra Yaprak
Tarman
ab,
Mert Kerem
Ulku
ab,
Karston
Yong
 a,
Muhammed
Veli
*c and
Hatice
Ceylan Koydemir
a,
Muhammed
Veli
*c and
Hatice
Ceylan Koydemir
 *ab
*ab
aDepartment of Biomedical Engineering, Texas A&M University, College Station, Texas 77843, USA. E-mail: hckoydemir@tamu.edu
bCenter for Remote Health Technologies and Systems, Texas A&M Engineering Experiment Station, College Station, Texas 77843, USA
cElectrical and Computer Engineering Department, University of California, Los Angeles, CA 90095, USA. E-mail: drmveli@gmail.com
dDepartment of Medical Equipment Technology, College of Applied Medical Sciences, Majmaah University, Al Majmaah, 11952, Saudi Arabia
First published on 15th October 2025
Abstract
Monitoring bacterial growth and detecting early-stage colony formation are essential tasks in biomedical research, clinical diagnostics, and food and water safety. However, conventional imaging systems for bacterial monitoring often require bulky optics, skilled operation, and high costs, making them unsuitable for scalable or field-deployable applications. Lens-free microscopy (LM) provides a promising alternative by enabling compact, low-cost imaging systems using only a light source and an image sensor, replacing the need for bulky objective lenses with computational algorithm. Still, a key limitation of LM is its resolution, which is fundamentally constrained by the sensor's pixel size. Pixel super-resolution techniques—especially when combined with multi-angle illumination using LED arrays—can significantly enhance resolution while maintaining a large field of view (FOV). We present OpenLM, an open-source lens-free microscopy platform integrated with a pixel super-resolution algorithm. The system is built from four affordable, off-the-shelf components: a Raspberry Pi camera, an optical filter, an LED array, and a Raspberry Pi board. Its 3D-printed housing enables easy replication and customization. User-friendly graphical interfaces for both Raspberry Pi OS and Windows provide camera control, real-time preview, image acquisition, and reconstruction—without requiring prior experience in lens-free imaging. To demonstrate its utility, we applied OpenLM to two bacterial imaging tasks: (1) long-term, time-lapse imaging of Escherichia coli (E. coli) colony growth, where colonies became visible within 30 minutes and complex spatial interactions emerged over time due to the wide FOV; and (2) early-stage colony detection using a YOLO (you only look once)-based deep learning model. With its affordability, high resolution, wide FOV, and ease of use, OpenLM is a practical and scalable tool for bacterial monitoring and other biomedical applications.
Introduction
Monitoring bacterial growth—especially in its early stages—is essential for timely diagnosis, treatment, and infection control.1 Rapid and accurate bacterial detection is crucial in healthcare particularly for the diagnosis of bacterial infections, where delays can exacerbate patient outcomes and increase risk of transmission.2–4Escherichia coli (E. coli) is part of the normal intestinal flora, but it is also a common bacterium responsible for various bacterial infections, including urinary tract infections (UTI), pneumonia, bacteremia, and peritonitis.5–7 In the United States, approximately 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 illnesses are caused by E. coli infections each year.8,9E. coli infections lead to a significant economic burden on healthcare systems and a high mortality rate.10E. coli bloodstream infections have a 30 day mortality rate of 9.6%.11 On the other hand, the healthcare costs of hospitalizations due to UTIs alone amount to approximately $2.8 billion annually in the United States.12 According to the CDC, bacterial culture is the standard method in clinical laboratories for diagnosing E. coli infections, however, this conventional approach requires two days of incubation before a diagnosis can be made.8 This traditional approach relies heavily on lens-based imaging systems. The trade-off between field of view (FOV) and resolution in lens-based imaging systems also extends the diagnosis time and limits continuous monitoring for early diagnosis.13 For example, a standard bright-field microscope equipped with a 20× objective—typically considered relatively low resolution in bright-field microscopy—provides a field of view (FOV) of about 1/20 of the sensor area, which is usually less than 1 mm2.14 In addition, these imaging systems are often expensive, require skilled operation, and limit their scalability in low-resource settings and point-of-care applications.15 Furthermore, incorporating lenses in a portable design adds weight and complexity, and limits flexibility of the system design.16
000 illnesses are caused by E. coli infections each year.8,9E. coli infections lead to a significant economic burden on healthcare systems and a high mortality rate.10E. coli bloodstream infections have a 30 day mortality rate of 9.6%.11 On the other hand, the healthcare costs of hospitalizations due to UTIs alone amount to approximately $2.8 billion annually in the United States.12 According to the CDC, bacterial culture is the standard method in clinical laboratories for diagnosing E. coli infections, however, this conventional approach requires two days of incubation before a diagnosis can be made.8 This traditional approach relies heavily on lens-based imaging systems. The trade-off between field of view (FOV) and resolution in lens-based imaging systems also extends the diagnosis time and limits continuous monitoring for early diagnosis.13 For example, a standard bright-field microscope equipped with a 20× objective—typically considered relatively low resolution in bright-field microscopy—provides a field of view (FOV) of about 1/20 of the sensor area, which is usually less than 1 mm2.14 In addition, these imaging systems are often expensive, require skilled operation, and limit their scalability in low-resource settings and point-of-care applications.15 Furthermore, incorporating lenses in a portable design adds weight and complexity, and limits flexibility of the system design.16
Lens-free microscopy (LM) offers a promising solution to overcome this challenge.1,17–20 As the name suggests, LM does not require lenses for imaging, resulting in a much simpler system architecture.21 It typically consists of two key components: a light source and an imaging sensor. This simplicity enables easy assembly, operation, and a low-cost design. The raw images captured by the LM's imaging sensor are holograms, representing the interference pattern between a reference wave and an object wave. Although the phase signal cannot be directly measured because the imaging sensor records only intensity, object information is still encoded in the interference pattern between the object wave and the reference wave. By assuming the reference wave to be a plane wave, as is common in lens-free microscopy, the object wave can be numerically reconstructed, allowing both amplitude and phase information to be retrieved.22 In lens-free microscopy (LM), the entire imaging sensor area dictates its field of view, with the resolution limit of a single hologram determined by the size of the sensor's pixels.23 CMOS sensors, commonly used in smartphones, are typically low-cost and feature small pixels, often around 1 μm or even smaller, making them useful for lens-free microscopy applications. For effective holography, the coherence of the light source is crucial. Therefore, lasers are ideal due to their high coherence. However, partially coherent light sources such as LEDs can also be used, provided that filters and pinholes are employed to enhance coherence. Given the advantages of low cost, ease of modification, and compact size, an LED array can serve as a suitable light source. The benefit of using an LED array is that it allows for the capture of multiple images of the same scene, which can be combined using pixel super-resolution algorithms to produce a high-resolution image, effectively overcoming the physical resolution limit inherent in LM.24,25
We present OpenLM, an open-source lens-free microscopy platform integrated with a pixel super-resolution algorithm. Designed for accessibility and affordability, the imaging system comprises four readily available components: a Raspberry Pi camera, an optical filter, an LED array, and a Raspberry Pi computer. The mechanical housing is fabricated via 3D printing, making the system both customizable and easy to replicate. To simplify operation, we developed two graphical user interface (GUI) applications—one for Raspberry Pi OS (Fig. S1) and another for Windows OS (Fig. S2). The Raspberry Pi application enables camera control, real-time preview, and communication with the Windows-based interface. The Windows application manages image acquisition, performs pixel super-resolution reconstruction, and sends capture commands to the Raspberry Pi. Together, these tools enable users to operate the system intuitively, with no prior experience in lens-free imaging and optical microscopy. Compared to previously published open-source lens-free imaging systems,26 our platform offers a higher level of integration and provides complete access to all source code. This allows users to operate and customize the entire image acquisition and processing pipeline. Additionally, our system delivers improved spatial resolution and features a more user-friendly interface. To demonstrate the utility of OpenLM, we present two applications. The first application is long-term time-lapse imaging of E. coli colony growth at room temperature. Colonies become visible within 30 minutes, and the system's large field of view (FOV) allows users to monitor inter-colony interactions at later stages. The second application involves early-stage bacterial detection using a you only look once (YOLO)-based deep learning model,27 showcasing the platform's compatibility with AI-powered analysis. OpenLM combines low cost, high resolution, large FOV, and user-friendly software, offering a versatile platform for diverse applications in biological research and education.
Methods
OpenLM system setup
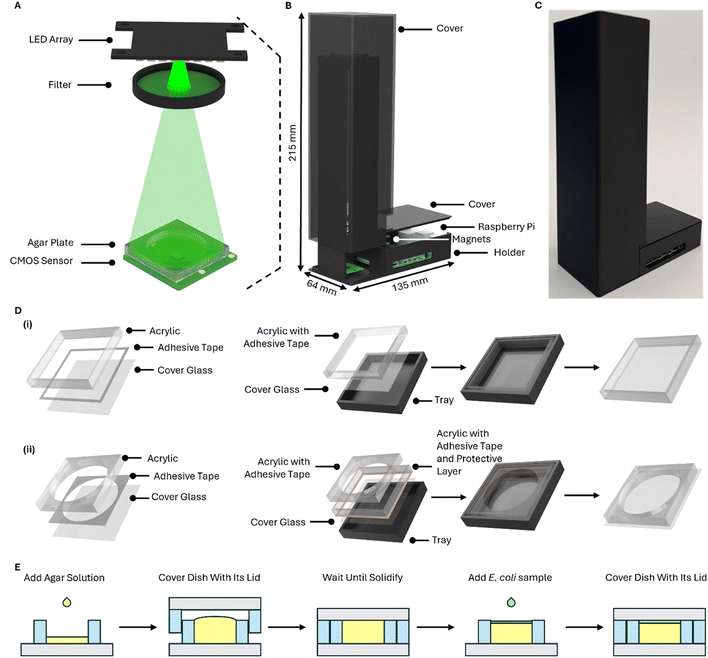
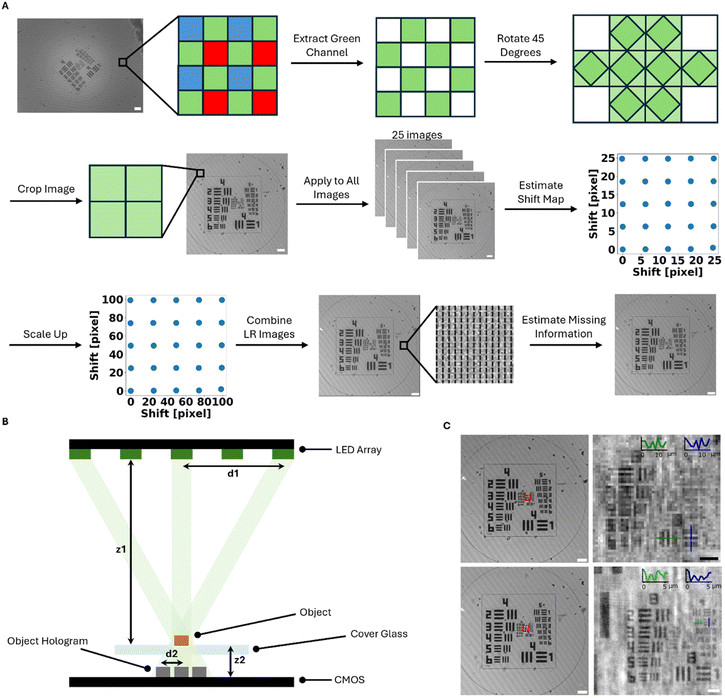
The OpenLM system consists of three key components (Fig. 1A): an LED array, a filter, and a CMOS sensor. The Raspberry Pi Camera Module 2 is selected as the imaging sensor, featuring a pixel size of 1.12 μm and a sensor resolution of 3280 × 2464, with a FOV of approximately 10.16 mm2. The lens on the top of the CMOS sensor is manually removed. The lens was enclosed in a plastic shell on top of the camera, and it was removed by carefully cutting and breaking the shell (Fig. S3). An 8 × 8 LED array (3444, Adafruit) serves as the light source to induce translational shifts of the object on the CMOS sensor. The LED array is directly connected to the Raspberry Pi via the MOSI and SCLK pins (Fig. S4). A band-pass filter (FLH532-4, Thorlabs) with a bandwidth of 4 nm and a center wavelength of 532 nm is employed to improve the temporal coherence of the light source. To ensure uniform illumination, 25 (5 × 5) LEDs are used to illuminate the sample, taking into account the size of the filter. The distance between the light source and the object is set to approximately 20 cm (Fig. 1B and C), ensuring that the CMOS sensor is fully illuminated by all LEDs and minimizing the object shift distance on the CMOS sensor (Fig. 1A and 2B). The relationship between the shifting distance and light source height is described by the following formula (eqn (1)).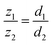 | (1) |
OpenLM system control
We developed two Python-based applications for system control: one for the Raspberry Pi to provide direct control, and another desktop application for remote control (Fig. S1). Both applications share core image-capturing functionalities, including capturing single images under user-selected LED illumination, capturing a set of 64 images—each illuminated by a different LED—and capturing time-lapse images based on user-defined settings. In time-lapse mode, 25 images are captured at each time point. During each capture event, the 25 central LEDs are sequentially turned on and off to acquire images under varying illumination conditions. Due to the limited computational power of the Raspberry Pi, its application only includes a basic focusing function for single images. All other image processing tasks are handled by the desktop application. Real-time processing is enabled by checking the “client” and “PSR” boxes in the Raspberry Pi interface (Fig. S1), which connects it to the desktop for processing requests. The desktop application also offers more advanced controls and adjustable parameters, such as adjusting the focal plane digitally, and provides greater flexibility in tuning the output images.Imaging substrate preparation
Substrate preparation for imaging resolution target while imaging the USAF resolution target (2017_Star_Dmnd, Ready Optics), the resolution target is placed face-down on a #0 cover glass, which is then positioned directly on top of the CMOS sensor.Substrate preparation for imaging bacterial growth on solid growth medium
The wall of a traditional Petri dish has a thickness of 1 mm, which prevents obtaining high-resolution images using OpenLM system when imaging bacterial colonies, Therefore, we developed a custom Petri dish (Fig. 1D) that reduces the distance between the object and the sensor to within 80–130 μm, which corresponds to the thickness range of #0 cover glass. Our custom Petri dish consists of three materials: #0 cover glasses, 3 mm-thick acrylic pieces, and double adhesive tape (468MP, 3 M), and is fabricated using a laser cutter (LS630, Boss Laser). The total dimensions are defined by the size of the cover glass, which is 22 mm × 22 mm. The height of entire Petri dish ranges from 3.16 mm to 3.21 mm due to variations in the thickness of the cover glass. The reservoir at the center has a diameter of 18 mm, with a total volume of about 763 μL. It is crucial to use a 3 mm acrylic sheet to maintain a well depth of 3 mm. If a 1 mm-thick acrylic sheet were used, the total volume of the Petri dish would be reduced to 254 μL, and the agar might solidify before it can be covered with the lid, resulting in a non-flat surface. The side thickness of the custom-made Petri dish cover is about 1 mm. To ensure a smoother fabrication process, the adhesive tape is applied to the acrylic sheet before cutting. We also fabricated a tray for component alignment using 3D printing (Fig. 1D and E) with PLA filament. The tray measures 22.6 mm × 22.6 mm × 3 mm, slightly larger than the cover glass to make it easier to remove the completed Petri dish bottom container and lid. The Petri dish bottom container and lid are assembled by sticking the three layers together (Fig. 1D). However, it is essential to note that while fabricating the bottom container of the Petri dish, the acrylic piece used for fabricating the lid also serves to secure the position of the acrylic piece used for the well (Fig. 1D).E. coli sample preparation
E. coli (25922™, ATCC) was cultured in tryptic soy broth (Bacto™ Tryptic Soy Broth, BD). For each experiment, the E. coli culture was diluted with phosphate-buffered saline (PBS) (P4417, Sigma). Tryptic soy agar (Difco™ Tryptic Soy Broth, BD) was used as the culture medium in custom-made Petri dishes. For this study, we prepared a concentration of 105 CFU mL−1 of E. coli to ensure colonies would be present in the field of view (FOV). First, the tryptic soy agar liquid solution was autoclaved, and it was pipetted into the reservoir of the Petri dish right after autoclaving (Fig. 1E). The pipette was set to 765 μL, but while injecting the solution, the plunger was only pushed to the first stop to prevent the generation of unwanted bubbles. Then, Petri dish was covered with the lid, and any excess agar solution was squeezed out. We placed a weight on top of the Petri dish to ensure that the lid was in complete contact with the bottom container. After the agar had fully solidified, the lid was carefully removed. Due to thermal expansion and contraction, the solidified agar surface was slightly lower than the surface of the bottom container. The resulting gap was large enough for E. coli to grow but small enough to allow for high-resolution imaging. Next, 5 μL of the E. coli-spiked PBS suspension was placed onto the agar and carefully spread to cover the agar surface. The bottom container was covered with a new lid just before flipping the petri dish. In the final step of sample preparation, the Petri dish was flipped and placed on the CMOS sensor for imaging.Imaging processing and analysis
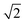 times larger than the original pixel size. To avoid pixel shifting during rotation, the image is initially cropped to a square shape with equal height and width. In our current automatic processing algorithm, the cropped image is 2464 × 2464 pixels, representing the largest square that can be extracted from the raw image. Following the rotation, the image is cropped again to eliminate the empty corners. The final image section is 1231 × 1231 pixels.
times larger than the original pixel size. To avoid pixel shifting during rotation, the image is initially cropped to a square shape with equal height and width. In our current automatic processing algorithm, the cropped image is 2464 × 2464 pixels, representing the largest square that can be extracted from the raw image. Following the rotation, the image is cropped again to eliminate the empty corners. The final image section is 1231 × 1231 pixels.
| H(x, y) = |R(x, y) + O(x, y)|2 | (2) |
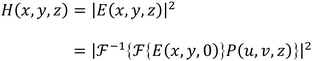 | (3) |
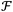 and
and 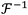 are the Fourier transform and its inverse, respectively. E(x, y, 0) is the field at the sensor plane, which is the captured hologram, and P(u, v, z) is the propagation function, defined in eqn (4) as:
are the Fourier transform and its inverse, respectively. E(x, y, 0) is the field at the sensor plane, which is the captured hologram, and P(u, v, z) is the propagation function, defined in eqn (4) as: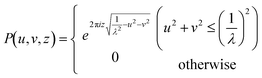 | (4) |
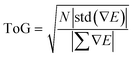 | (5) |
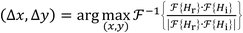 | (6) |
| ∇S4 = 0 | (7) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 728 unique colony instances were labelled across 19 independent experiments. Due to the overlapping nature of the patches, the final training dataset included 117
728 unique colony instances were labelled across 19 independent experiments. Due to the overlapping nature of the patches, the final training dataset included 117![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 463 annotated instances. The dataset was then randomly divided into training and validation subsets in an 85% to 15% ratio, respectively, to facilitate model development and evaluation.
463 annotated instances. The dataset was then randomly divided into training and validation subsets in an 85% to 15% ratio, respectively, to facilitate model development and evaluation.
The YOLOv11s model was selected as the base architecture for object detection. Training was initialized using pre-trained YOLOv11s weights to leverage transfer learning, accelerate convergence and reduce training time. All experiments were conducted in a consistent and controlled computational environment using Ultralytics version 8.3.115, Python 3.12.8, and PyTorch 2.7.0. Model training and inference were performed on a desktop equipped with an NVIDIA GeForce RTX 3070 Ti GPU (8 GB VRAM) and an Intel Core i7-12700KF CPU.
Results and discussion
High resolution image acquisition
To prevent moisture accumulation on the glass, three dummy images were captured before capturing the full set of 25 images for super resolution. As a result, a total of 28 images are captured at each time point during the time-lapse imaging process. Due to the writing speed and computational limitations of the Raspberry Pi, the OpenLM system takes approximately 50 seconds to complete the image capture task at each time point. The system's image capture speed may decrease over time due to the increased CPU temperature and reduced available computational resources. Although 25 valid images are captured at each time point, the scale factor in our super-resolution algorithm is set to 4. Since the generated hologram shift is still larger than a pixel, the subpixel information stored in the 25 images is insufficient to generate a super-resolution image with a scale factor of 5. After super-resolution, the effective sampling of the holographic fringes increases, enabling recovery of finer fringe content from smaller features and rendering them reconstructable (Fig. S5). The measured spatial resolution improves from 1.95 μm to 0.87 μm (Fig. 2C). Although the number of pixels increased by a scaling factor of 4, the resolution did not improve by a factor of 4. This is because the hologram shifts are larger than one pixel, resulting in empty pixels in the high-resolution (HR) image. These pixels are estimated based on surrounding pixel values, as described in the Methods section. Due to this missing information, the effective scaling factor is less than 4. Our desktop, equipped with an Intel i7-12700KF processor, takes about 22.4 seconds to generate a super-resolution image (4924 × 4924 pixels) from 25 low-resolution images (1231 × 1231 pixels).E. coli growth monitoring
To evaluate the performance of our imaging platform, we monitored the growth of E. coli colonies on agar plates maintained at room temperature (Fig. 3). Images were captured at 5 minute intervals over a 21.5-hour period. Time-lapse images of a single E. coli colony after reconstruction are presented in Fig. 3, illustrating the growth dynamics over time. As detailed in the Methods section, the lateral hologram shift induced by different LED illumination is a function of the axial object-to-sensor distance. Accordingly, the pixel super-resolution algorithm reconstructs the HR image by aligning and fusing LR holograms based on the lateral displacement characteristic of the E. coli colonies' focal plane. Holograms corresponding to objects located at differing axial depths undergo spatial misregistration during the fusion process, leading to their attenuation or suppression in the final reconstruction. This depth-selective integration inherently enhances image clarity at the target plane, such as E. coli colonies on agar plate, by diminishing out-of-focus contributions, such as moisture on the cover glass (Fig. 3).The first appearance of an E. coli colony was observed at approximately 30 minutes (Fig. 3). The presence of a diffraction fringe ring surrounding the colony indicates that it is a physical object rather than imaging noise. However, at this early stage, the colony is still too small to significantly distort the incident wavefront, resulting in limited contour definition. As time progresses and the colonies grow in size, their contours become increasingly well-defined, and the associated diffraction fringes become more pronounced. Notably, the ring surrounding each colony also corresponds to the edge of the twin-image artifact—a white region overlapping the colony center. This interference pattern arises from the missing phase information in lens-free imaging, which records only intensity and not phase, leading to reconstruction artifacts typical of in-line holography.36 The twin-image artifact plays a critical role in distinguishing real physical objects from random background noise. However, as a form of interference, it also degrades the fidelity of the reconstructed image. In the case of E. coli colonies, which grow with non-uniform height profiles, the twin-image typically appears with a brighter center and a darker edge, differing from the actual pattern of the object. The central bright region within the twin-image ring may be attributed to the thin colony edge thickness and dense internal microstructure. Over time, as the colony grows, the twin-image becomes more prominent. This is expected, as larger colonies produce more complex wavefronts, exacerbating phase ambiguity during intensity capture. After approximately 3.5 hours, it can dominate the reconstruction, obscuring the true colony contour and making it difficult to visually resolve the object's boundaries. Twin-image noise can be mitigated using multi-wavelength illumination for phase retrieval37 or mask-based phase retrieval techniques.38 However, our system currently employs a color CMOS sensor, which is inherently limited in its ability to capture consistent intensity information under multi-wavelength illumination. For example, in this study we utilized green light, which is detected only by the green pixels. If illumination is changed to another wavelength, such as red, the green pixels are unable to record the signal, resulting in images that are not directly comparable. Instead, only the red pixels contribute, and because they are fewer in number and distributed differently across the Bayer pattern, the effective sampling of the image differs from that of the green channel. Consequently, an image captured under red illumination differs both in resolution and sampling pattern from one captured under green illumination. Moreover, due to the very small size of early-stage colonies, mask-based methods are ineffective at this scale. Owing to the simplicity of our platform, users can readily replace the color CMOS sensor with a monochrome detector, thereby enabling multi-wavelength imaging and supporting accurate quantitative phase retrieval. The use of a monochrome sensor eliminates the Bayer filter, increases sensitivity, and allows all pixels to be utilized for information capture under multi-wavelength illumination.37 An alternative approach to achieve accurate phase recovery is to incorporate a z-axis translation stage beneath the imaging sensor, enabling image acquisition at different heights with a constant phase difference and phase recovery for each images.39 Although the twin-image artifact affects the reconstructed image quality, it does not significantly alter the apparent size or rough general contour of the E. coli colonies. Beyond 6 hours, the internal structure of the colony became too complex for accurate reconstruction due to multidirectional expansion. The object wavefront distortion surpassed the algorithm's reconstruction capability. However, at this stage, the colony was sufficiently large to generate a visible shadow image, enabling size tracking even in the absence of detailed structural reconstruction. And the expansive field of view enables continuous monitoring of spatial dynamics, including the fusion of adjacent E. coli colonies during growth (Video S1 and S2).
Early detection of E. coli colonies
YOLO is a state-of-the-art, continuously evolving object detection framework known for its high speed and accuracy. In this study, we selected the latest version, YOLOv11, to ensure reliable and accurate detection performance. YOLOv11 offers five model sizes, with parameter counts ranging from 2.6 million to 56.9 million. While larger models generally provide improved detection accuracy and training performance, they also impose higher computational and memory demands. YOLOv11s (9.4 million parameter) was selected as it is the largest model variant that could be fully accommodated within the 8 GB memory limit of our GPU. Training larger models, such as YOLOv11m (20.1 million parameter), would exceed this limit, forcing memory overflow operations to the CPU and significantly reducing training speed.After training, the model achieved a precision of 0.937, recall of 0.908, and a mean average precision (mAP) of 0.965 at an intersection over union (IoU) threshold of 0.5, referred to as mAP@0.5. A precision of 0.937 means that 93.7% of the colony predictions were correct, indicating excellent accuracy and a low false positive rate. The recall of 0.908 shows that 90.8% of actual colonies were successfully detected, demonstrating strong sensitivity with few missed detections. The mAP@0.5 of 0.965 reflects outstanding overall detection performance when predictions are considered correct at a 50% IoU threshold. Additionally, the more stringent mAP@0.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.95 was 0.771. This metric averages mAP scores across IoU thresholds ranging from 0.5 to 0.95 in 0.05 increments, and a value above 0.7 is typically considered very strong. This result indicates that the model performs robustly even under stricter localization criteria, maintaining high accuracy across a range of overlap requirements. To evaluate model performance, a blind test dataset was prepared using time-lapse images of E. coli growing on an agar plate over a 3 hour period. These images were entirely separate from those used for training and validation and were manually labeled. The number of E. coli colonies per image ranged from 27 to 48, depending on the variable growth rates of individual colonies.
0.95 was 0.771. This metric averages mAP scores across IoU thresholds ranging from 0.5 to 0.95 in 0.05 increments, and a value above 0.7 is typically considered very strong. This result indicates that the model performs robustly even under stricter localization criteria, maintaining high accuracy across a range of overlap requirements. To evaluate model performance, a blind test dataset was prepared using time-lapse images of E. coli growing on an agar plate over a 3 hour period. These images were entirely separate from those used for training and validation and were manually labeled. The number of E. coli colonies per image ranged from 27 to 48, depending on the variable growth rates of individual colonies.
The first colony detections by the model occurred at ∼35 minutes (Fig. 4A). The timing of initial visibility depends on several factors, including temperature, the density of colonies on agar plate, and the biological activity of the E. coli. In this setup, the E. coli suspension was applied immediately onto the agar, while the Petri dish was placed directly onto the CMOS sensor to minimize uncertainty in the growth start time. However, residual liquid between the agar and the chamber cover may have delayed the adhesion of E. coli to the agar surface. Not all colonies were detected at the 35 minute mark due to the challenges of identifying early-stage growth. Some small background particles exhibited patterns resembling early E. coli colonies, making it difficult for both the model and label experts to distinguish them without referring to later-stage images. This is particularly challenging given that the model processes only a single image at a time. Additionally, diffraction fringes and twin-image noise—common features surrounding all physical objects in holographic reconstructions—were not considered reliable indicators for labelling. These features are not specific to E. coli and evolve over time as the colony grows. Including such unstable and non-discriminative features in the training data would likely reduce model accuracy. In this study, colonies were only labelled when they exhibited clearly visible growth beyond the diffraction fringe, making them distinguishable to the naked eye. This same criterion was applied during evaluation to maintain consistency. At 115 minutes, approximately 87.5% of actual E. coli colonies had been successfully detected by the model (Fig. 4B). As the colonies continued to grow, their contours became more distinct and easier to differentiate from background particles, resulting in improved detection accuracy. Although the model's precision began to fluctuate over time, it consistently maintained a precision of approximately 0.9 throughout the blind test evaluation (Fig. 4C). The observed fluctuation is primarily due to the increasing variability in colony contours. At the early stages of growth, colonies contain only a small number of E. coli cells and tend to exhibit similar morphological patterns under our imaging platform. However, as colonies grow, the differences in their contours become more pronounced, making it more challenging for the model to accurately detect them—leading to fluctuations in precision. Despite this, the precision remains close to 0.9. We anticipate that increasing the dataset size and incorporating a broader range of colony contour variations will improve the model's overall precision.
Some false positives were observed, where the model identified E. coli colonies before they were visibly apparent in the images. Additional false positives occurred when objects with similar size and structure to the E. coli used in this study were mistakenly detected. This issue is primarily attributed to limitations in the training dataset. Although the model was trained on over 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 instances, certain scenarios were underrepresented or missing entirely—such as unexpected particulate matter at specific focal planes and random background noise. Additionally, the dataset contained relatively few background-only images. While including more background images could help the model better distinguish between colony and non-colony regions, it also presents a trade-off: increasing background diversity may improve specificity but can also raise the risk of false negatives. Conversely, having too few background images may lead the model to incorrectly assume that every image contains a detectable object, thereby increasing false positives. This limitation can be addressed through further expansion of the dataset to cover a wider range of imaging conditions, including more background and noise variations. Additionally, using a larger YOLO model variant—capable of learning more complex features—may improve detection accuracy. Furthermore, implementing a post-processing checkpoint that verifies whether predicted bounding boxes appear consistently across consecutive frames can help filter out transient false positives and improve the robustness of colony detection over time. In future studies, a wider range of growth scenarios could be included in the training dataset, such as varying bacterial concentrations and the presence of artifacts like microspheres or dust, to improve the model's reliability and robustness. Furthermore, data from multiple time points could be combined to track the growth dynamics of objects, which would help distinguish living bacterial colonies from non-living particles or other background noise. Incorporating such temporal information and diverse scenarios would enhance the model's generalization to real-world samples and increase its applicability in practical bacterial monitoring.
000 instances, certain scenarios were underrepresented or missing entirely—such as unexpected particulate matter at specific focal planes and random background noise. Additionally, the dataset contained relatively few background-only images. While including more background images could help the model better distinguish between colony and non-colony regions, it also presents a trade-off: increasing background diversity may improve specificity but can also raise the risk of false negatives. Conversely, having too few background images may lead the model to incorrectly assume that every image contains a detectable object, thereby increasing false positives. This limitation can be addressed through further expansion of the dataset to cover a wider range of imaging conditions, including more background and noise variations. Additionally, using a larger YOLO model variant—capable of learning more complex features—may improve detection accuracy. Furthermore, implementing a post-processing checkpoint that verifies whether predicted bounding boxes appear consistently across consecutive frames can help filter out transient false positives and improve the robustness of colony detection over time. In future studies, a wider range of growth scenarios could be included in the training dataset, such as varying bacterial concentrations and the presence of artifacts like microspheres or dust, to improve the model's reliability and robustness. Furthermore, data from multiple time points could be combined to track the growth dynamics of objects, which would help distinguish living bacterial colonies from non-living particles or other background noise. Incorporating such temporal information and diverse scenarios would enhance the model's generalization to real-world samples and increase its applicability in practical bacterial monitoring.
This study demonstrated the effectiveness of the OpenLM system in combination with an AI-based E. coli detection model. Future studies may explore the application of this system to other bacterial species, including mixed-species detection scenarios. Additionally, the detection process required only 3 seconds per image, suggesting that real-time detection could feasibly be integrated into the system for continuous monitoring of bacterial growth.
Comparison of image-based bacterial detection techniques for portable devices
Several image-based detection techniques have been developed and adapted for portable platforms, including traditional optical microscopy,40 subpixel perspective-sweeping microscopy,41 ptychography,42–44 and optofluidic scanning microscopy.45 Collectively, these approaches have demonstrated substantial potential as powerful tools for bacterial detection (Table S2). Nevertheless, they often rely on conventional optical elements or customized physical components, which inevitably introduce limitations in terms of portability, cost, and accessibility. These requirements can hinder the translation of otherwise promising technologies into field-ready, low-cost diagnostic platforms. A distinguishing feature of our system is that the imaging function—traditionally fulfilled by lenses,40 is instead realized entirely through computational reconstruction algorithms. By eliminating the need for physical focusing optics, our system achieves a significantly larger FOV than lens-based portable microscopes.40 This expanded FOV is not merely a convenience: it directly impacts throughput and detection efficiency, as it allows larger sample areas to be interrogated in a single capture. In the context of bacterial detection, where identifying rare events across broad sample regions is often crucial, this advantage substantially improves both speed and reliability.While certain lens-free methods, such as optofluidic microscopy45 and ptychography,42–44 also depart from traditional lens-based architectures, they generally replace lenses with other specialized optical components, including aperture arrays or diffusers. For example, an aperture array consists of a series of pinholes precisely arranged at specific orientations and distances. Although powerful, such components are difficult for end-users to fabricate, are not readily available off-the-shelf, and require expertise to integrate properly. In contrast, all optical and electronic elements in our system are inexpensive, commercially available components, and the only customized element is a 3D-printed sample holder. Importantly, this holder can be fabricated on entry-level consumer 3D printers or ordered from widely available printing services. This emphasis on standardization and accessibility lowers barriers to adoption, ensuring the system can be readily reproduced and deployed outside specialized laboratories. Another advantage of our platform is that object information is encoded in the interference pattern between the object and reference waves, rather than being limited to a pure intensity projection. Techniques such as subpixel perspective-sweeping microscopy41 enhance resolution primarily by collecting and computationally stitching large sets of intensity images. In contrast, our approach preserves both amplitude and phase information in the recorded hologram, enabling digital refocusing at different depths and facilitating true volumetric (3D) imaging. This digital focusing capability offers greater flexibility in sample handling and compatibility with diverse biological specimens, which often vary in thickness, morphology, and refractive properties. The trade-offs between resolution, frame count, and system complexity are exemplified by prior work in subpixel perspective-sweeping microscopy.41 For instance, one study demonstrated a resolution of 0.66 μm across a 24 mm2 FOV, but only after acquiring and processing 225 frames to generate a single high-quality reconstruction.41 While this result underscores the theoretical performance ceiling of lens-free approaches, it also highlights the associated burdens in terms of acquisition time and computational demand. Our system, although currently optimized for fewer frames, can be readily scaled to higher resolutions through straightforward upgrades. These include replacing the present LED array with a more densely packed emitter array or implementing a fiber array to achieve finer illumination steps, as well as incorporating a larger sensor to extend the attainable FOV. Importantly, these modifications remain both practical and cost-effective, preserving the accessibility of the platform while enabling advanced users to push performance further. This open-source framework therefore not only provides a reproducible baseline configuration but also offers the flexibility for users to adapt and upgrade the system according to their specific resolution and FOV requirements.
Compared with other portable image-based bacterial detection techniques, our system achieves a unique balance between performance, simplicity, and accessibility. By leveraging computational imaging in combination with inexpensive, readily available hardware, it delivers large-FOV, 3D-capable bacterial detection in a format that is reproducible, adaptable, and practical for deployment well beyond specialized laboratory environments.
Conclusions
In summary, the OpenLM system offers a unique combination of low cost, portability, and ease of use, making it an ideal tool for a wide range of applications, including bacterial growth monitoring and early detection. Early detection of E. coli colony formation as early as 30 min at room temperature was demonstrated, with the detection time potentially reduced even further when placed in an incubator, providing flexibility for various experimental setups. The system's large FOV and high resolution further enhance its utility, allowing for detailed monitoring of microbial growth and interactions. These features make OpenLM particularly well-suited for applications in food safety, environmental monitoring, and healthcare diagnostics, where rapid and accurate detection of microbial activity is crucial. Moreover, its user-friendly design and accessibility open up new possibilities for non-specialist users in diverse sectors, from clinical laboratories to field research, offering a versatile and efficient tool for real-time monitoring.Author contributions
W. X.: conceptualization, methodology, software, investigation, writing – original draft, writing – review & editing. S. A.: investigation. M. A.: investigation. A. Y. T.: investigation. M. K. U.: investigation. K. Y.: investigation. M. V.: writing – original draft, writing – review & editing, supervision. H. C. K.: conceptualization, methodology, writing – original draft, writing – review & editing, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
All 3D-printed part files, as well as system control and processing code, are publicly available at https://github.com/xuwimming/OpenLM. DOI: https://doi.org/10.5281/zenodo.15848567.Supplementary information file including component-cost, software screenshots, camera preparation and assembly workflows is available. See DOI: https://doi.org/10.1039/d5lc00719d.
Acknowledgements
The Koydemir Research Group at Texas A&M University acknowledges the support of the U.S. National Science Foundation (Award No. 1648451), the U.S. Department of Defense-Office of Naval Research (N00014-23-1-2225), and the U.S. National Institute of Health (NIH) NIGMS (R21GM150104). M. A. acknowledges Majmaah University and the Saudi Arabian Cultural Mission (SACM) for their support.References
- H. Wang, H. Ceylan Koydemir, Y. Qiu, B. Bai, Y. Zhang, Y. Jin, S. Tok, E. C. Yilmaz, E. Gumustekin, Y. Rivenson and A. Ozcan, Light: Sci. Appl., 2020, 9, 118 CrossRef PubMed.
- W. Xu, E. Venkat and H. Ceylan Koydemir, Curr. Opin. Biomed. Eng., 2023, 28, 100513 CrossRef CAS.
- S. Doron and S. L. Gorbach, in International Encyclopedia of Public Health, ed. H. K. Heggenhougen, Academic Press, Oxford, 2008, pp. 273–282, DOI:10.1016/B978-012373960-5.00596-7.
- W. Xu, M. Althumayri, A. Y. Tarman and H. Ceylan Koydemir, Biosens. Bioelectron., 2025, 283, 117539 CrossRef CAS PubMed.
- J. M. Mylotte, A. Tayara and S. Goodnough, Clin. Infect. Dis., 2002, 35, 1484–1490 CrossRef PubMed.
- J. D. McCue, J. Am. Geriatr. Soc., 1987, 35, 213–218 CrossRef CAS PubMed.
- S. Jain, H. S. Wesley, G. W. Richard, S. Fakhran, R. Balk, M. B. Anna, C. Reed, G. G. Carlos, J. A. Evan, D. M. Courtney, D. C. James, C. Qi, M. H. Eric, F. Carroll, C. Trabue, K. D. Helen, J. W. Derek, Y. Zhu, R. A. Sandra, K. Ampofo, W. W. Grant, M. Levine, S. Lindstrom, M. W. Jonas, M. K. Jacqueline, D. Erdman, E. Schneider, A. H. Lauri, A. M. Jonathan, T. P. Andrew, M. E. Kathryn and L. Finelli, N. Engl. J. Med., 2015, 373, 415–427 CrossRef CAS PubMed.
- C. A. Bopp, R. B. Carey, P. Gerner-Smidt, L. H. Gould, P. M. Griffin and N. A. Strockbine, MMWR Morb. Mortal. Wkly. Rep., 2009, 58, 1–14 Search PubMed.
- H. J. Shah, R. H. Jervis, K. Wymore, T. Rissman, B. LaClair, M. M. Boyle, K. Smith, S. Lathrop, S. McGuire, R. Trevejo, M. McMillian, S. Harris, J. Zablotsky Kufel, K. Houck, C. E. Lau, C. J. Devine, D. Boxrud and D. L. Weller, MMWR Morb. Mortal. Wkly. Rep., 2024, 73, 584–593 CrossRef PubMed.
- M. Camara, W. Green, C. E. MacPhee, P. D. Rakowska, R. Raval, M. C. Richardson, J. Slater-Jefferies, K. Steventon and J. S. Webb, npj Biofilms Microbiomes, 2022, 8, 42 CrossRef PubMed.
- M. C. MacKinnon, S. A. McEwen, D. L. Pearl, O. Lyytikainen, G. Jacobsson, P. Collignon, D. B. Gregson, L. Valiquette and K. B. Laupland, BMC Infect. Dis., 2021, 21, 606 CrossRef PubMed.
- K. Iskandar, R. Rizk, R. Matta, R. Husni-Samaha, H. Sacre, E. Bouraad, N. Dirani, P. Salameh, L. Molinier, C. Roques, G. Economics of the Antibiotic Resistance Research, A. Dimassi, S. Hallit, R. Abdo, P. A. Hanna, Y. Yared, M. Matta and I. Mostafa, Value Health Reg. Issues, 2021, 25, 90–98 CrossRef PubMed.
- J. Mertz, Introduction to Optical Microscopy, Cambridge University Press, Cambridge, 2nd edn, 2019 Search PubMed.
- S. L. Renne, Pathologica, 2023, 115, 302–307 Search PubMed.
- C. J. Potter, Z. Xiong and E. McLeod, Laser Photonics Rev., 2024, 18, 2400197 CrossRef.
- W. Xu, M. Althumayri, A. Mohammad and H. Ceylan Koydemir, Biosens. Bioelectron., 2023, 242, 115755 CrossRef CAS PubMed.
- T. Liu, Y. Li, H. C. Koydemir, Y. Zhang, E. Yang, M. Eryilmaz, H. Wang, J. Li, B. Bai, G. Ma and A. Ozcan, Nat. Biomed. Eng., 2023, 7, 1040–1052 CrossRef PubMed.
- Y. Li, T. Liu, H. C. Koydemir, H. Wang, K. O'Riordan, B. Bai, Y. Haga, J. Kobashi, H. Tanaka, T. Tamaru, K. Yamaguchi and A. Ozcan, ACS Photonics, 2022, 9, 2455–2466 CrossRef CAS.
- M. Roy, D. Seo, S. Oh, J. W. Yang and S. Seo, Biosens. Bioelectron., 2017, 88, 130–143 CrossRef CAS PubMed.
- Z. W. Qin, Y. Yang, Y. L. Ma, Y. B. Han, X. L. Liu, H. Y. Huang, C. S. Guo and Q. Y. Yue, Opt. Express, 2024, 32, 29329–29343 CrossRef CAS PubMed.
- A. Greenbaum, W. Luo, T. W. Su, Z. Gorocs, L. Xue, S. O. Isikman, A. F. Coskun, O. Mudanyali and A. Ozcan, Nat. Methods, 2012, 9, 889–895 CrossRef CAS PubMed.
- S. Mustafi and T. Latychevskaia, Photonics, 2023, 10(2), 153 CrossRef.
- J. Zhang, J. Sun, Q. Chen and C. Zuo, IEEE Trans. Comput. Imaging, 2020, 6, 697–710 Search PubMed.
- Z. Xiong, J. E. Melzer, J. Garan and E. McLeod, Opt. Express, 2018, 26, 25676–25692 CrossRef CAS PubMed.
- W. Bishara, U. Sikora, O. Mudanyali, T. W. Su, O. Yaglidere, S. Luckhart and A. Ozcan, Lab Chip, 2011, 11, 1276–1279 RSC.
- S. Amann, M. V. Witzleben and S. Breuer, Sci. Rep., 2019, 9, 11260 CrossRef PubMed.
- G. Jocher, J. Qiu and A. Chaurasia, YOLO by Ultralytics, 2023 Search PubMed.
- T. Latychevskaia, J. Geophys. Res. Atmos., 2019, 36, D31–D40 CAS.
- J. W. Goodman, Introduction to Fourier optics, Roberts and Company publishers, 2005 Search PubMed.
- Y. Zhang, H. Wang, Y. Wu, M. Tamamitsu and A. Ozcan, Opt. Lett., 2017, 42, 3824–3827 CrossRef PubMed.
- P. Sung Cheol, P. Min Kyu and K. Moon Gi, IEEE Signal Process. Mag., 2003, 20, 21–36 CrossRef.
- M. Guizar-Sicairos, S. T. Thurman and J. R. Fienup, Opt. Lett., 2008, 33, 156–158 CrossRef PubMed.
- C. K. Chui and H. N. Mhaskar, Appl. Comput. Harmon. Anal., 2010, 28, 104–113 CrossRef.
- S. B. Damelin and N. S. Hoang, Int. J. Math. Math. Sci., 2018, 2018, 1–8 CrossRef.
- F. C. Akyon, S. O. Altinuc and A. Temizel, 2022 IEEE International Conference on Image Processing (ICIP), 2022, pp. 966–970, DOI:10.1109/ICIP46576.2022.9897990.
- M. Guizar-Sicairos and J. R. Fienup, J. Opt. Soc. Am. A, 2012, 29, 2367–2375 CrossRef PubMed.
- Q. Wang, J. Ma and P. Su, Front. Photon., 2022, 3, 865666 CrossRef.
- O. Mudanyali, D. Tseng, C. Oh, S. O. Isikman, I. Sencan, W. Bishara, C. Oztoprak, S. Seo, B. Khademhosseini and A. Ozcan, Lab Chip, 2010, 10, 1417–1428 RSC.
- Y. Rivenson, Y. Wu, H. Wang, Y. Zhang, A. Feizi and A. Ozcan, Sci. Rep., 2016, 6, 37862 CrossRef CAS PubMed.
- J. S. Cybulski, J. Clements and M. Prakash, PLoS One, 2014, 9, e98781 CrossRef PubMed.
- G. Zheng, S. A. Lee, Y. Antebi, M. B. Elowitz and C. Yang, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 16889–16894 CrossRef CAS PubMed.
- S. Jiang, C. Guo, Z. Bian, R. Wang, J. Zhu, P. Song, P. Hu, D. Hu, Z. Zhang, K. Hoshino, B. Feng and G. Zheng, Biosens. Bioelectron., 2022, 196, 113699 CrossRef CAS PubMed.
- S. Jiang, C. Guo, P. Song, N. Zhou, Z. Bian, J. Zhu, R. Wang, P. Dong, Z. Zhang, J. Liao, J. Yao, B. Feng, M. Murphy and G. Zheng, ACS Photonics, 2021, 8, 3261–3271 CrossRef CAS.
- S. Jiang, P. Song, T. Wang, L. Yang, R. Wang, C. Guo, B. Feng, A. Maiden and G. Zheng, Nat. Protoc., 2023, 18, 2051–2083 CrossRef CAS PubMed.
- X. Heng, D. Erickson, L. R. Baugh, Z. Yaqoob, P. W. Sternberg, D. Psaltis and C. Yang, Lab Chip, 2006, 6, 1274–1276 RSC.
| This journal is © The Royal Society of Chemistry 2025 |