Open Access Article
Open Access ArticleImmune cell infiltration of patient derived glioblastoma cell spheroids in acoustic levitation in bulk acoustic wave devices
Xavier
Mousset
acd,
Shuake
Kuermanbayi
acd,
Chloé
Dupuis
 acd,
Nathan
Jeger-Madiot
acd,
Nathan
Jeger-Madiot
 acd,
Virgile
Delaunay
cd,
Ahmed
Ibdaih
b,
Elias A.
El-Habr
acd,
Virgile
Delaunay
cd,
Ahmed
Ibdaih
b,
Elias A.
El-Habr
 cde,
Hervé
Chneiweiss
cd,
Marie-Pierre
Junier
cde,
Hervé
Chneiweiss
cd,
Marie-Pierre
Junier
 cd,
Jean-Luc
Aider†
a and
Jean-Michel
Peyrin†
cd,
Jean-Luc
Aider†
a and
Jean-Michel
Peyrin†
 *cd
*cd
aCNRS UMR7636, Laboratoire de Physique et Mécanique des Milieux Hétérogènes (PMMH), Team AcoustoFluidics, Paris, France
bSorbonne Université, AP-HP, Institut du Cerveau - Paris Brain Institute - ICM, Inserm, CNRS, Hôpitaux Universitaires La Pitié Salpêtrière - Charles Foix, DMU Neurosciences, Service de Neuro-Oncologie-Institut de Neurologie, F-75013, Paris, France
cSorbonne Université, CNRS, Inserm, Neuro-SU, F-75005 Paris, France. E-mail: jean-michel.peyrin@sorbonne-universite.fr
dSorbonne Université, CNRS, Inserm, Institut de Biologie Paris-Seine, IBPS, F-75005 Paris, France
eDepartment of Biological Chemistry, Medical School, National and Kapodistrian University of Athens, Athens, Greece
First published on 5th November 2025
Abstract
We describe an acoustofluidic device that allows scaffold-free structuration and culture of multi-cellular tumoroids composed of patient-derived glioblastoma cells only or in combination with non-cancerous cells. A PDMS chip of controlled height was created to allow acoustic levitation of cells using a 2 MHz transducer held on top of the chip. Cells are introduced into the chip through a dedicated inlet upstream of the resonant cavity. The specific design of the cavity together with the acoustic field allows the formation of tumoroids of cells in a precise and controlled manner within the levitation chamber. The acoustic and fluidic environment of the device was determined through experiments confronted with numerical simulations. The control of the flow within the chip was optimized to allow long-term culture of tumoroids and injection of cell culture media without disturbing the tumoroids in levitation. The tumoroids can be also structured, with sequential injections of the different cell types. Using microglia, we show that the acoustofluidic device allows the formation and culture in acoustic levitation of tumoroids mixing cancer cells with other cells populating the tumor as well as immune cell infiltration within the tumoroids. These results demonstrate the suitability of acoustofluidic levitation as an original 3D culture method adapted to the exploration of cancer growth at multiple levels.
Introduction
Glioblastoma, the most common and aggressive malignant brain tumour, remains particularly resistant to current treatments, which typically combine surgical resection with chemotherapy and radiotherapy. Increasing evidence highlights the molecular and cellular heterogeneity of glioblastoma as key factors in treatment resistance. This heterogeneity results from the coexistence of distinct genomic clones, the exceptional plasticity of malignant cells, and the presence of normal cells co-opted by the tumour.Components of the tumour microenvironment (TME) participate in promoting tumour growth and aggressiveness.1 The TME comprises a diverse set of neural cells (astrocytes, oligodendrocyte progenitors, neurons) and non-neuronal cells (immune cells, cellular components of blood vessels), as well as components of the extracellular matrix. Glioblastoma cells interact with the TME through various mechanisms, which influence both tumour progression and therapeutic resistance. For example, they induce tumour-associated macrophages (TAMs), the most abundant immune cellular component of glioblastoma, to adopt a pro-tumour state.2 Similarly, glioblastoma cells interact with surrounding neuronal networks, creating a detrimental feedback loop that promotes tumour growth while disrupting neuronal activity.3 A thorough understanding of these complex interactions between the tumour and the microenvironment is essential to progress towards more effective therapies. This requires the development of appropriate experimental models, simplified enough to allow a fine functional and molecular dynamic dissection of specific cellular interactions in a three-dimensional setting.
3D cell culture models of glioblastoma have generated substantial scientific interest due to their potential to better mimic the in vivo tumour environment. In contrast to conventional two-dimensional (2D) cultures, 3D models allow glioblastoma cells to grow in a more physiologically relevant context, preserving the complex architecture and cell–cell interactions found in actual tumours. These models facilitate the study of tumour heterogeneity, cell invasion, and resistance mechanisms in a controlled yet realistic setting. By incorporating elements like extracellular matrix components, 3D cultures provide insights into the influence of the tumour microenvironment on glioblastoma growth and therapy responses. They are particularly complementary of conventional 2D high-throughput drug screening, as they can better predict the efficacy and toxicity of potential treatments compared to 2D cultures. Overall, 3D cell culture models are a powerful tool for advancing our understanding of glioblastoma biology and for accelerating the development of more effective therapies.
The large majority of 3D models rely on matrix-based methods in order to constrain cells in a 3D setting with the help of a variety of hydrogels containing or not extra-cellular matrix components. Although relatively easy to establish, these models are limited by their reliance on scaffolds of uncertain impact on the cells. Organ-on-a-chip and cell bio-printing methods offer alternative 3D models including cell elements of the tumour microenvironment. They have notably allowed exposure of tumour cell organoids to vascular-like structures.4,5 However, they are complex to implement and, for the bio-printing procedure, impose a strain on the cells. Most recent models have been developed based on ancient strategies grafting human glioblastoma cells in human nervous tissues obtained upon differentiation of human pluripotent stem cells6 and now from human iPSC-derived brain organoids.7 Finally, tissue from surgical resections of the patients' tumours has been found to be amplifiable under constant rotation in culture.8 These methods reproduce or conserve to a fair extent the tumour microenvironment but require months to be implemented or to access fresh surgical material.
Acoustic levitation offers a novel and intriguing method of cell manipulation and cell culture, with several advantages over traditional techniques.9,10 By using sound waves to aggregate cells in a nutrient-rich medium,11 acoustic levitation eliminates physical contact with a substrate, reducing stress and potential contamination. This method enables the rapid formation of aggregates of cells in a three-dimensional environment with only cell–cell interactions, more closely mimicking the natural conditions within the body, making it a promising technique for tissue engineering applications,12–15 as it enables the formation and culture of complex cellular structures and spheroids either in culture medium or hydrogels.16,17 Overall, acoustic levitation represents a cutting-edge and innovative approach to cell culture, with significant potential to advance research in cell biology and regenerative medicine.18,19 This technique is also widely used for cell sorting (acoustophoresis) or high-throughput screening.20,21
Because cells are freed from the mechanical stress and cell–wall interactions inherent in 2D cultures, acoustic levitation can improve or accelerate cell function development. This has been illustrated by the successful culture in acoustic levitation of fragile cells such as mesenchymal stromal stem cells22,23 or hepatic cells.24,25 In these studies, the formation of cell layers in acoustic levitation was very quick, followed by a step of self-organization of the cell layers into spheroids whose dynamics depends highly on the cell types. The viability of the cells forming the spheroids was always evaluated and was found to be as good as or better than the cells cultured in wells or on 2D substrates, even after 10 days of culture in acoustic levitation. Moreover improved cell proliferation and faster acquisition of cell functionalities, like albumin production by hepatocytes, were observed. Additionally, the acoustofluidic chips used in these previous studies allows precise control of cell positioning,22,23,26 facilitating studies of cellular behaviour and interactions in a dynamic environment. Nevertheless, all these studies were dedicated to culture of a single type of cell. The present study aims at introducing a secondary cell type and evaluating the potential of this technique for infiltration of tumoroids by a secondary cell type, which is crucial to develop more realistic tumoroid models.
In this study, we first show that acoustic levitation can be used to develop tumoroids from patient-derived glioblastoma cells (GB-PDCs) that recapitulate the complex organization of human glioblastoma. The viability and self-organization into tumoroids are verified, even for long cell culture times. We then show that it is indeed possible to manage cell infiltration of a tumoroid in acoustic levitation by a proper design of the acoustofluidic chip.
Methods
Acoustic levitation
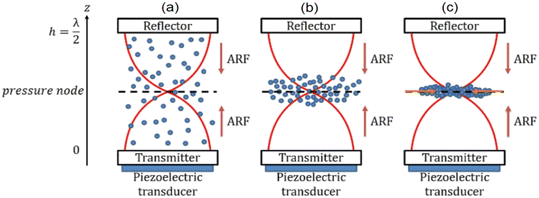
Acoustic levitation is used in this study to handle softly a large number of cells in well-defined positions inside the volume of a cavity where they first form large layers. After a few hours the cells spontaneously self-organize into spheroids that can be maintained in acoustic levitation as long as needed. These manipulations rely on the acoustic radiation force (ARF) applied to suspended objects in a resonant acoustic cavity (bulk acoustic wave – BAW device). Indeed, if the resonance condition is satisfied (h = nλac/2, with n an integer and λac the acoustic wavelength), a plane acoustic standing wave is generated inside the cavity (Fig. 1). In this case, every object in suspension in the fluid medium will undergo the axial component Frad of the ARF. If the objects are considered as compressible spheres with the diameter dp smaller than the acoustic wavelength (λac ≫ dp), then the ARF can be written as: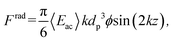 | (1) |
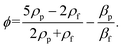 | (2) |
Microfluidic chip fabrication
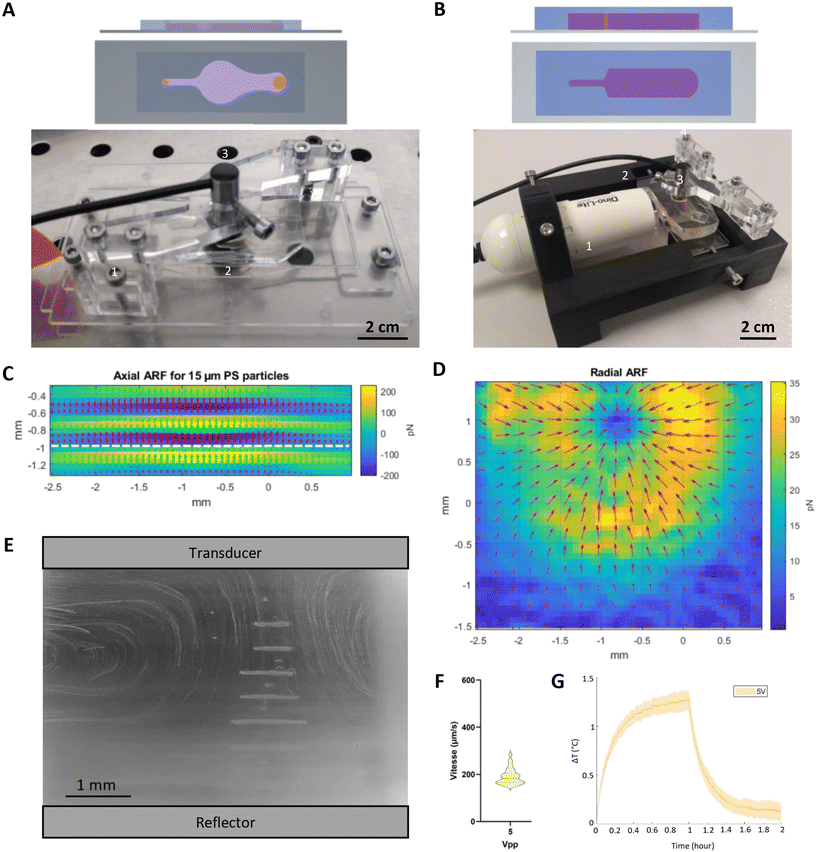
The acoustofluidic chips were designed to control the flow around the tumoroids trapped in acoustic levitation in the resonant cavity, between the inlet and the outlet. It is important to emphasize that the fluid injection is used only to inject the cells or to add a second cell type over an already formed spheroids in acoustic levitation. Two types of acoustofluidic cavities were used.A chip was dedicated to microscopy imaging, meaning an observation of the lowest aggregate from under the chip using an inverted microscope. This chip is 40 mm long, 10 mm large and 1.5 mm high. For an acoustic frequency fac = 2 MHz, the acoustic wavelength is λac = 750 μm. This means that the cavity can contain up to 3 aggregates. The total internal volume is 300 μL (Fig. 2A and S2A).
Another chip was dedicated to cell culture in an incubator and side observations. The objective is to create and culture multiple aggregates for further biological analysis. They are also built to allow an observation from the side. This is useful to monitor the dynamics of self-organization of the aggregates over time. It is 40 mm long, 8 mm large and 5 mm high (containing up to 14 aggregates) with a total internal volume of 1 mL (Fig. 2B).
The acoustofluidic chips were fabricated by pouring PDMS (poly-dimethylsiloxane), a gas-porous biocompatible material, using a protocol previously developed to grow neurons in microfluidic chips.27 Briefly, a master encoding the microfluidic chip was designed by laser cutting a 1.5 or 5 mm thick Plexiglas sheet and further glued on top of a polycarbonate Petri dish (Greiner). The PDMS chip was made by pouring a degassed solution of 90% Sylgard™ 184 silicon elastomer base (Neyco) mixed with 10% curing agent (w/w) in the molding Petri dish, with a thickness of at least 0.5 mm above the Plexiglas mold. The mold filled with PDMS was placed for 2 hours in an oven (Memmert) at 70 °C. The PDMS imprint was cleaned with double sided tape (3 M), and plasma bonded (plasma cleaner Diener electronic) on a 1 mm thick glass slide (VWR). The glass slide acts as a reflector for the ultrasound leading to stationary wave generation. Prior to plasma bonding, the glass slide was cleaned with precision wipes (Kimtech) soaked in isopropanol (VWR); PDMS slabs and glass slides were then exposed for 5 min to air plasma (RF plasma cleaner, Diener, Femto). After bonding, the chip was filled with water to preserve the hydrophilic properties of the PDMS acquired during plasma treatment. The sealed chips were exposed for 30 minutes to UV for sterilization, and stored at 4 °C up to 15 days prior to use. In order to prevent media evaporation, once filled with liquids, the inlet and outlet reservoirs were covered by an UV treated 1 mm-thick PDMS membrane. This coverage, coupled with immediate hydration of the chips after plasma treatment, was also instrumental in preventing bubble formation during cell culture.
Acoustofluidic device
The acoustofluidic chips consist of a PDMS/glass microchip. They can be considered as a “layered resonator” device as described in ref. 28. The device allows the location of the acoustic pressure nodes to be forced by choosing the correct thicknesses for the layers. It was also used later by Dron & Aider to move the axial location of the levitation plane of a single node device just by changing the acoustic frequency.26 It was more recently extended to a multiple node device similar to the ones used in the present study.23 The dimensions of the two microchips are available as STEP files in the SI.Another important feature is the choice of the acoustic source. To deal with the layered chip we use an ultrasonic transducer with a 2 MHz wide-band (TR0205, Signal Processing™) and with a 5 mm diameter piezoelectric element embedded in an 8 mm diameter casing. The use of a wide-band source authorizes the tuning of the acoustic frequency to maximize the acoustic energy inside the cavity.23 The ultrasonic transducer is held by a home-made Plexiglass holder for microscopy imaging of a single aggregate (Fig. 2A). A specific 3D-printed holder was designed to hold a digital USB microscope on the side of the ‘multi-node’ chip for the lateral imaging of the levitating tumoroids (Fig. 1B). The ultrasonic transducer was powered by a USB-powered digital arbitrary waveform generator (Handyscope HS5, Tiepie Engineering™). A thin layer of olive oil was used to ensure a good acoustic coupling between the transducer and the PDMS chip.
Cell cultures
Patient-derived cells (PDCs), named 6240 and R633, were obtained from neurosurgical resection samples of distinct primary glioblastoma.38 The cells were expanded and cultured in conventional cell culture flasks in defined culture medium (human Neurocult NSA-H basal medium, StemCell) containing heparin (1 μg mL−1, Sigma), EGF (20 ng mL−1, AbCys SA), and bFGF (10 ng mL−1, AbCys SA), as described.38 The human microglia clone 3 (HMC3) cell line (Russo 2018) was cultured in DMEM medium with 10% fetal bovine serum (Gibco). All cells were cultured in an incubator at 37 °C under 5% CO2. If necessary, HMC3 cells were labeled with the nontoxic cell tracker red dye (Invitrogen). Cells were incubated for 5 min at 37 °C in PBS (500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells ml−1) containing the dye at 1 μg ml−1 before being washed with PBS and resuspended in fresh culture medium.
000 cells ml−1) containing the dye at 1 μg ml−1 before being washed with PBS and resuspended in fresh culture medium.
Acoustofluidic tumoroid formation, culture & retrieval
Acoustofluidic chips were mounted with the transducer under a biosafety hood, and held in the Plexiglas holder. Olive oil was inserted between the transducer and the PDMS chip to allow for efficient acoustic coupling. Water inside the chip was replaced by the cell culture medium. Single cell suspensions were prepared using Accutase (StemPro). A 50 μL cell suspension was injected in the acoustofluidic device to reach a final concentration of 105 cells mL−1. It was then carefully mixed to homogenize the spatial distribution of cells inside the levitation chamber.The signal applied to the acoustic transducer was set at a frequency fac = 2 ± 0.05 MHz with a voltage amplitude of 7 V. These parameters maximize the efficiency of cell trapping in the levitation planes and their aggregation in the levitation planes. A good acoustic trapping also minimizes the risk of losing cells trapped in the pressure nodes during transport of the acoustofluidic chips from the hood to the incubator.
After injection of the cell culture medium with the cells and stabilized assembly of the cells, the chip's inlet and outlet holes were sealed with PDMS caps to prevent evaporation of the medium. The porosity of PDMS to gases allows exchanges between the medium and the air throughout the chip. Once the chips installed inside the incubator, the voltage was decreased to 5 V for long-term culture. At the end of the experiment, the transducer is turned off and the chip top opened. The tumoroids having developed in levitation are retrieved by aspiration with a 1 mL pipet tip. The tips are cut with a cutter to enlarge their opening. Retrieval of the cell clusters having developed outside of the acoustic field was then performed by forcing their detachment from the glass-bottom of the chip with up and down pipetting of media.
Characterisation of the acoustic pressure field
Experimental measurements of the acoustic pressure amplitude inside the cavity were achieved using a fiber optic hydrophone (Precision Acoustic™, UK) mounted on an XYZ translation stage. The sensor dimension is 10 μm and its sensitivity is approximately 200 mV MPa−1. The chip was cut at the side to allow the introduction of the hydrophone inside the cavity, perpendicularly to the acoustic propagation axis (angular flat response of the sensor). The pressure field was scanned in a volume of 5 × 3 × 1 mm3 inside the cavity and with a spatial resolution of λac/10 = 70 μm. The displacement of the probe was controlled using dedicated software to probe automatically the volume of the cavity. The signal was recorded for 150 μs with an acquisition frequency of 20 MHz for each measurement points.Live cell imaging and image analysis
The time-evolution of the size and shape of the cell aggregates created in the acoustic levitation planes was monitored using a USB digital microscope (AM2111, Dino-Lite™) with a resolution of 640 × 480 pixels, placed on the side of the chip. This small digital microscope was connected to a computer and controlled using the DinoCapture 2.0 software supplied by Dino-Lite. Snapshots of the cells were recorded every five-minutes throughout the length of cultures. A final image was recorded at the end of the experiment, just after turning off the transducer, once tumoroids had fallen at the bottom of the cavity. This image of the background without tumoroids was used in subsequent image processing. The images were processed using Matlab software (R2022a). Each image was converted to grayscale and binarized after background subtraction. Small objects were then removed and holes were filled to improve the detection of the parameters of interest. All detections were visually verified using a video file displaying the grayscale images along with the detected contour.Three parameters were computed from the snapshots: the circularity (Cir), eccentricity (Ecc) and tumoroid diameter (Deq) defined respectively as:
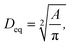 | (3a) |
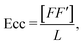 | (3b) |
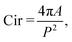 | (3c) |
Tumoroid retrieval, cell counting and viability testing
After culture, levitated and non-levitated cells were retrieved from the chips in two different Eppendorf tubes by pipetting tumoroids out from the chips with a modified P1000 pipette tip under a laboratory binocular. Levitated and non-levitated tumoroids were discriminated by their location in the chips and the fact that levitated tumoroids do not attach to the resonator glass slide while the non-levitated ones do. Tumoroids were then centrifuged at 3000 rpm for 1 min at room temperature. The tumoroids were exposed to Accutase (StemPro) for 5 min at 37 °C and final cell dissociation was achieved with gentle up and down pipetting. Ten μL of cell suspension was mixed with an equal volume of trypan blue and incubated for 3 min at room temperature prior to counting live and dead cells (Invitrogen Countess Automated Cell Counter, Thermofisher). The total number of counted cells ranged from 100 to 1000. Live/dead staining was performed with a LIVE/DEAD Cell Imaging Kit (ThermoFisher) according to the manufacturer instructions.Fixation, cryosections, tissue clearing and immunostaining
Tumoroids were fixed in 4% paraformaldehyde in PBS for 15 min at 4 °C, cryoprotected in 15% and 30% sucrose in PBS at 4 °C for 15 min and 30 min, respectively, embedded in tissue freezing medium (Cryofix Gel, BIOGNOST), frozen in isopentane at −40 °C and stored at −80 °C until used. The tumoroids were cut into 25 μm-thick cryostat slices. For immunostaining, the slices were washed with PBS for 15 min, and subsequently incubated in PBS containing 0.2% Triton (X100, Sigma) and 1% BSA (Sigma) for 5 min, and in PBS containing 10% BSA for 1 h at room temperature (RT). Primary antibodies were applied overnight at 4 °C in PBS containing 5% BSA (anti-Ki67 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000, Agilent, M7240, A488-coupled anti-EGFR, 352907, BioLegend, 1
1000, Agilent, M7240, A488-coupled anti-EGFR, 352907, BioLegend, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500), followed by rinsing and incubation for 2 hours at room temperature in PBS containing 5% BSA, 1
500), followed by rinsing and incubation for 2 hours at room temperature in PBS containing 5% BSA, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20
20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 DAPI (1050-A, Euromedex), and when relevant anti-mouse antibodies coupled to Alexa 633 (1/500, Invitrogen). After washing with PBS, slices were mounted using Mowiol (475904-M, Sigma) and stored at 4 °C.
000 DAPI (1050-A, Euromedex), and when relevant anti-mouse antibodies coupled to Alexa 633 (1/500, Invitrogen). After washing with PBS, slices were mounted using Mowiol (475904-M, Sigma) and stored at 4 °C.
For tissue clearing, tumoroids were fixed in 4% paraformaldehyde in PBS for 15 min, followed by washing with PBS for 15 min. Tumoroids were then transferred into 24 well plates and incubated overnight at 37 °C in permeabilization buffer containing 0.5% Triton in PBS. Tumoroids were then washed three times with PBS1X for 10 min each, and incubated with DAPI 1X 2 days at 4 °C, followed by washing three times with PBS 1X for 10 min each. Tumoroids were deposited into Gene Frame chambers (AB-0577, ThermoFisher) pre-adhered to coverslips, and pre-filled with RapiClear 1.47 (RC147000, SunJin Lab). The Gene Frame chambers were covered with glass coverslips prior to imaging.
Image acquisition and analysis
Immunofluorescence imaging was achieved using a Zeiss Axio-observer microscope and a Coolsnap HQ2 CCD camera, or a confocal microscope with Zen software (Zeiss LSM 980 Inverted, confocal laser scanning microscopy, Airyscan 2, fluorescence-lifetime imaging module, laser diodes ranging from 405 nm to 639 nm, Colibri 7 LED light source, Zeiss ZEN Blue 3). Acquired images were processed with FIJI (Schindelin, 2012), the StarDist plugin30 and MATLAB. Cells were segmented relying on the DAPI staining using the StarDist plugin. The segmentation allowed extraction of the nucleus position and shape and quantification to be performed. Quantification of Ki67 was determined by averaging the Ki67 fluorescence signal over the segmented nucleus or region of interest (ROI). The positive or negative state of cell proliferation was determined using a threshold value defined visually for each image. Cell density was equal to the number of detected cells divided by the area of a circle with radius R0. The position of each nucleus positive for the staining was detected by thresholding the minima of the second derivative of the image. The position of the centroid of the tumoroid was calculated by averaging the different positions of the nuclei in the section, and the distance of each nucleus with regard to the centre was calculated (R/R0). The distribution of these positions was then plotted using a sliding box method with a box width of 30 μm.Statistical methods
Statistical analyses were performed using the unpaired t-test with Welch's correction with Prism 6.0 software (GraphPad) and significance threshold set at p < 0.05. Results are shown as means ± SD (standard deviation).Results and discussion
Design and characterization of acoustofluidic cell culture devices
Acoustofluidic cell culture relies on the establishment of an ultrasound standing wave in a resonant cavity (Fig. 1) that must take into account fluidic control, biological constraints and geometries optimized to create an acoustic standing wave. In order to achieve long-term cultures of glioblastoma cells, we optimized two complementary configurations, one adapted to fluorescence microscopy (Fig. 2A) and the other adapted to cell culture of many aggregates in an incubator (Fig. 2B). Operating the transducer at 5 V was found optimal to obtain efficient and reproducible trapping of GB-PDC in both single and multi-node chips. In the multi-node chip, the maximum amplitude of the measured acoustic pressure field was found to be about 300 kPa, for a supplied voltage of 5 V (see SI, Movie S1). The acoustic velocity was deduced from the Euler equation in the harmonic regime: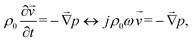 | (4) |
![[small nu, Greek, vector]](https://www.rsc.org/images/entities/i_char_e0ea.gif) the acoustic pressure and velocity, ρ0 the fluid density, and ω the angular frequency.
the acoustic pressure and velocity, ρ0 the fluid density, and ω the angular frequency.
The acoustic energy density could also be evaluated using the relation 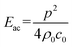 , leading to Eac ≈ 10 J m−3, a value comparable to other results found in previous studies on similar acoustic resonators.26,39
, leading to Eac ≈ 10 J m−3, a value comparable to other results found in previous studies on similar acoustic resonators.26,39
To find a rough estimate of the axial component of the ARF Frad applied to the cells, the ARF acting on a polystyrene (PS) particle with a 15 μm diameter has been calculated using Gorkov's theory.39 Indeed, the ARF can be defined as the gradient of an acoustic potential:
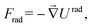 | (5) |
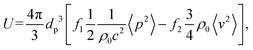 | (6) |
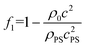 and
and 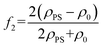 , dp the particle diameter, ρ0 and ρPS respectively the densities of fluid and particle PS, and c and cPS the sound speed respectively in the fluid and polystyrene. Using these definitions, it is possible to derive the expression of the axial ARF given in eqn (1).39
, dp the particle diameter, ρ0 and ρPS respectively the densities of fluid and particle PS, and c and cPS the sound speed respectively in the fluid and polystyrene. Using these definitions, it is possible to derive the expression of the axial ARF given in eqn (1).39
Using these relations and knowing the properties of the fluid and polystyrene beads, the ARF field has been computed. The result obtained for a 15 μm polystyrene bead in a 1.5 mm high cavity is shown in Fig. 2C in a vertical plane. The axial ARF is maximum on the pressure antinodes with a value Frad ≅ 200 pN. The particles or cells are then pushed by the axial ARF toward the pressure nodes. Once in the acoustic levitation plane (i.e. the pressure node) the axial component of the ARF vanishes, while the radial component FT of the ARF is maximum. The 2D field of FT in one of the horizontal levitation plane (white dashed line in Fig. 2C) is shown in Fig. 2D. One can see that the radial component of the ARF is oriented inward toward the cylindrical symmetry centre of the cavity, where the particles or cells will form an aggregate (see Movie S1). The maximum value of the transversal component of the ARF is FT ≅ 30 pN, much lower than the axial component, but strong enough to lead to the rapid formation of a large aggregate in acoustic levitation. Under these conditions, it was possible to form large layers of PDC in acoustic levitation that turned into tumoroids within 12 h, with low risk of falling.
It should be noted that these calculations are valid only during the first steps of creation of large layers of cells. They cannot be applied to the large tumoroids that spontaneously form after 12 hours in acoustic levitation since the ≃300 μm final size of the tumoroid does not respect the Rayleigh approximation (dp ≪ λac). As they grow, tumoroids modify their interactions with the acoustic wave – and therefore the spatial organization of the acoustic field – but also their densities, both phenomena impacting the ARF. To ensure trapping of acoustically levitated tumoroids throughout the culture, ARF was applied at a minimum value of 5 V to provide sufficiently powerful acoustic traps compensating for the weight of the tumoroids.
Acoustic wave propagation in liquid can trigger acoustic streaming,31–34 which may be an issue to stabilize the aggregates over a long time. To quantify the streaming in the chips, the GB-PDC displacement in the vicinity of the levitation planes was monitored using the side observation of the multimode chip by the digital microscope. A time-series of snapshots were recorded, allowing the tracking of the cells and an estimation of their velocities. Fig. 2E shows a side view of the cell trajectories after their injection into the chip and the initial formation of planar cell layers. The large re-circulations induced by the streaming can be clearly seen around the cell layers in acoustic levitation at the center of the chip. The velocities were then measured away from the central region, where aggregation takes place as can be seen in Fig. 2E. The piezoelectric is only 5 mm wide. One can see that most of the streaming happens on the side of the chip, mostly away from the region where the ARF is strong. Using particle tracking analyses, it was then possible to measure the average streaming velocity vst = 200 μm s−1. The velocity distribution is shown in Fig. 2F. Of note, the streaming does not perturb the levitation area where GB-PDC formed very stable aggregates.
Finally, the temperature was measured using a thermocouple within the multi-node chip filled with cell media. The time evolution of temperature when the acoustics is turned on is shown in Fig. 2G. One can see a temperature increase of 1.25 °C after one hour for these acoustic parameters (see Fig. S1). When the acoustics is turned off, the temperature decreases to reach its original value after one hour. To ensure a constant temperature of 37 °C for cell culture, the temperature of the incubator was set at 36 °C.
Acoustic levitation represents an innovative approach to advance cell culture studies, particularly for challenging contexts like glioblastoma which need more realistic cell culture environments. In this study, we therefore optimized and characterized a microfluidic acoustic bioreactor capable of generating stable acoustic standing waves in a closed system, enabling precise control of the fluidic environment and facilitating long-term cell culture of many spheroids. The optimized microchip design allowed manual medium exchange without causing disruptive drag forces, streamlining the maintenance of cell cultures.
Self-organization of glioblastoma PDC under acoustic levitation
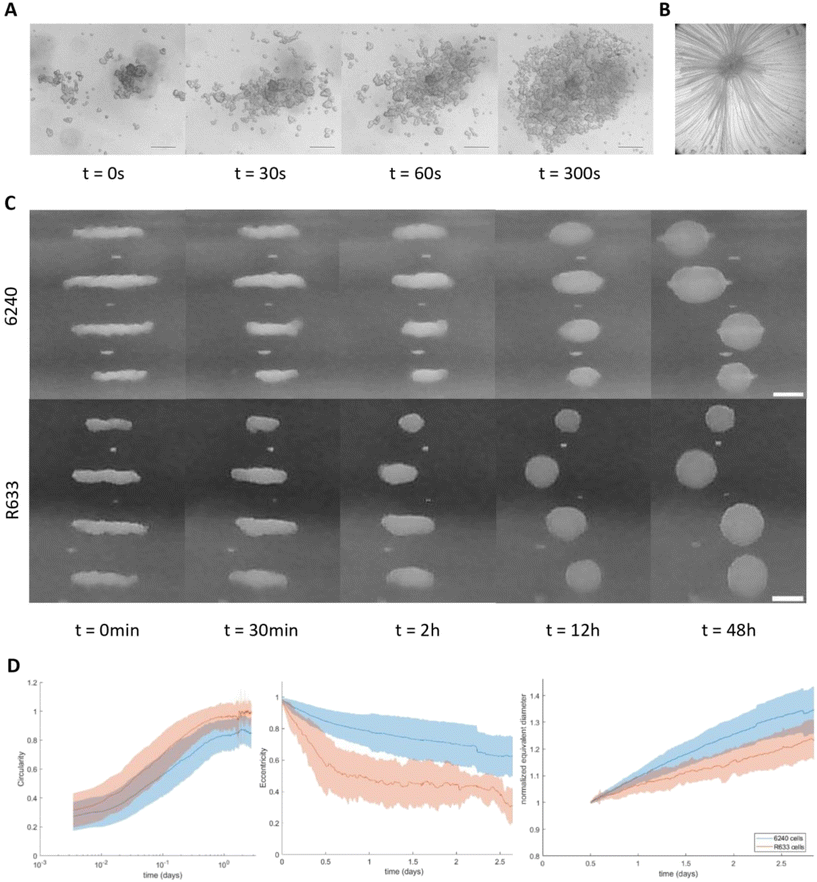
The self-organization of the layers of GB-PDCs into tumoroids when cultivated in acoustic levitation was monitored at short and long times after cell injection in either the mono or the multimode chips. Cells were injected through the inlet port of the microchips while the acoustic transducer was activated (Fig. 2A and B). Aggregation at pressure nodes takes place sequentially after injecting the cell suspension into the chips. Injected cells are trapped within seconds as loosely packed planar layers (Fig. 3A) under the influence of the axial component of the ARF (Fig. 2C). This is followed by progressive packing of the cells over minutes at the center of the levitation plane (Fig. 3A and B, Video S1), or pressure node, under the action of the radial component of the ARF (Fig. 2D) resulting in the formation of stable cell layers. We did not observe cells attracted to the bottom or top walls of the chips. The cells that were not trapped in acoustic levitation sedimented on the bottom wall outside of the region submitted to the acoustic field.We compared the behavior of two PDC lines (R633 and 6240) with different genomic backgrounds, different abilities to adhere to untreated culture vial surfaces, and distinct migratory properties35 (Fig. S2A and B). PDC progressively self-organized over time into dense spheroidal aggregates with tumoroid-like shapes (Fig. 3C). The kinetics of tumoroid formation was evaluated through the evolution of the morphometric parameters over time, as defined in the Methods section (see eqn (3a–c)). The aggregates achieved roughly circular shapes (Cir = 1) after approximately 24 hours of culture (Fig. 3D, left panel). Then the tumoroid size increased, suggesting active cell amplification inside the tumoroids (Fig. 1D, right panel). Both PDCs exhibit an overall similar behavior in levitation, although cells with lesser migratory and adhesion properties tend to form more ellipsoidal tumoroids (Fig. 3D, middle panel) while PDCs with the highest adhesion properties self-organized more quickly into more spheroidal aggregates. Altogether these results show that PDCs self-organize in a contactless environment into tumoroid-like spheroids under acoustic levitation, regardless of their intrinsic specificities.
Long term culture of PDC under acoustic levitation
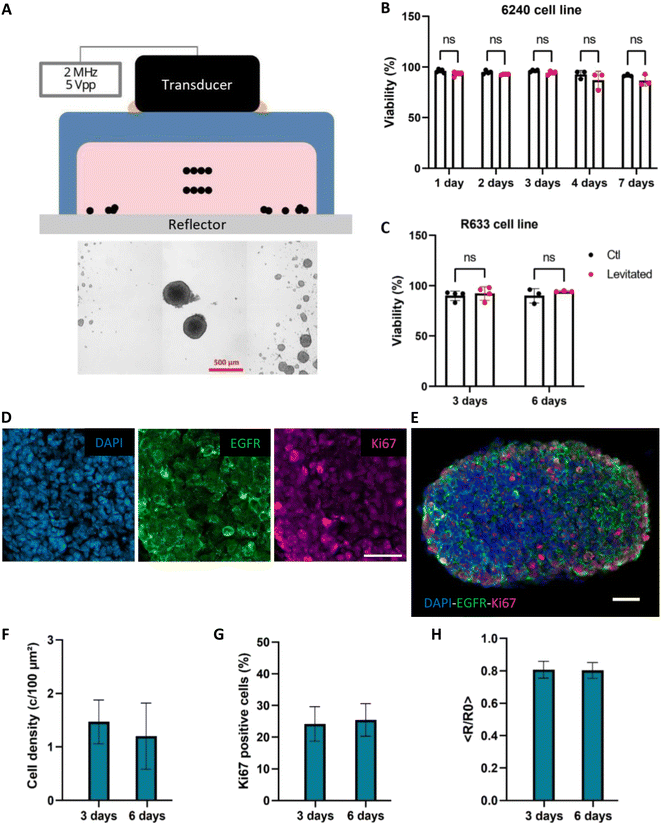
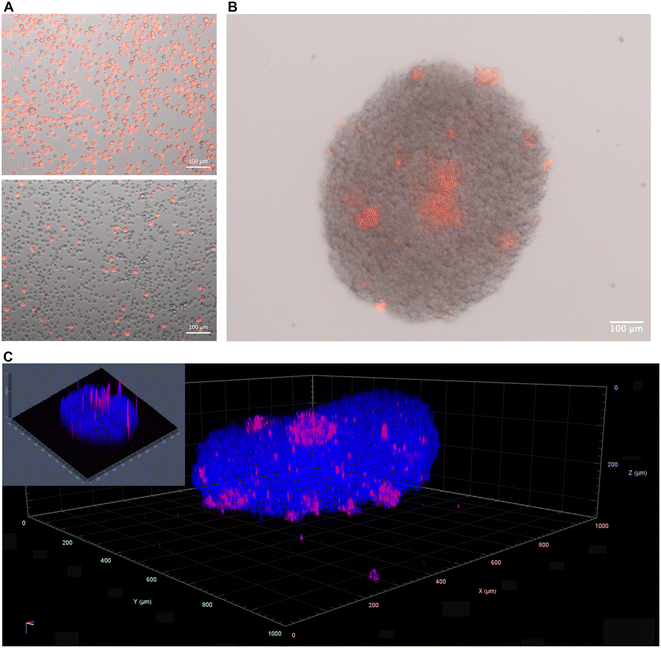
Cell survival was evaluated after 6–7 days of culture by comparing cells captured in levitation to cells having developed under the form of small tumoroids outside of the acoustic field (Fig. 4A). As shown in Fig. 4B and C, cell survival assessed with trypan blue exclusion assay showed that levitation did not alter PDC survival as compared to control non-levitated cells, over 95% live cells being recovered in either condition. The cells were homogeneously distributed throughout the tumoroids with no evidence of a necrotic core, as shown with DNA labelling with DAPI and immunolabelling of EGFR, a transmembrane tyrosine kinase receptor overexpressed by glioblastoma cells (Fig. 4D, left and middle panels, Fig. 4E). EGFR-immunolabeling showed that cells were arranged in close contact with their neighbors in the tumoroids (Fig. 4E). Cell densities were similar at both time-points evaluated (Fig. 4F). Finally, we used Ki67 staining (Fig. 4D, right panel, Fig. 4E) to determine whether PDC maintained active proliferation under acoustic levitation, as suggested by the increased size of the tumoroids over culture time. Quantification of Ki67-immunopositive cells showed that approximately 25% of PDC were undergoing proliferation at both time points evaluated (Fig. 4G). Determination of the repartition of Ki67-immunoreactive cells within the tumoroids showed enrichment of proliferating cells in the outer layers of the tumoroids (Fig. 4H). Altogether these results demonstrate that acoustic levitation allows fast and efficient self-organization of glioblastoma PDC into actively proliferating tumoroids.Here, we demonstrated that malignant cells derived from the patients' tumours cultured in acoustic levitation maintain active proliferation after self-organization into tumoroids that mirror the 3D morphology of tumour tissues. The tumoroids maintain their coherent structure once removed from the acoustic field instead of dispersing into individual cells as expected for aggregates whose maintenance would depend on continuous exposure to ARF. Notably, the system enabled the growth and three-dimensional self-organization of cells such as PDC-R633, which spontaneously grow under the form of 2D monolayers even in low-adhesion culture flasks. This underscores the potential of acoustic levitation to enforce rapid formation of tumoroids by promoting rapidly strong initial cell–cell contacts upon cell seeding. Interestingly, the spatial organization of the 2 distinct PDC lines presented subtle differences exemplified by self-organization of PDC-R633 into more ovoid tumoroids. While there is no direct explanation, this may be linked to their distinct adhesion and/or migration properties as suggested by the observation that R633 cell lines strongly adhere and form migratory streams in conventional 2D cell cultures systems while 6240 cells spontaneously form cellular spheres in these conventional culture systems. We thus succeeded in performing long-term culture of 3D tumoroids from 2 distinct patient-derived glioblastoma cell lines.
Structuration of PDC-tumoroids including immune cells
Next, we determined the suitability of acoustic levitation for implementing complex tumoroids that would integrate macrophage/microglia, i.e. non-malignant elements known to populate glioblastoma tumors, and to interact with malignant cells. The HMC3 microglia cell line was used, after verifying that its culture in PDC culture medium did not alter its survival and proliferation. The tumoroids were first generated using a cell suspension mixing PDC with human HMC3 microglia at a 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
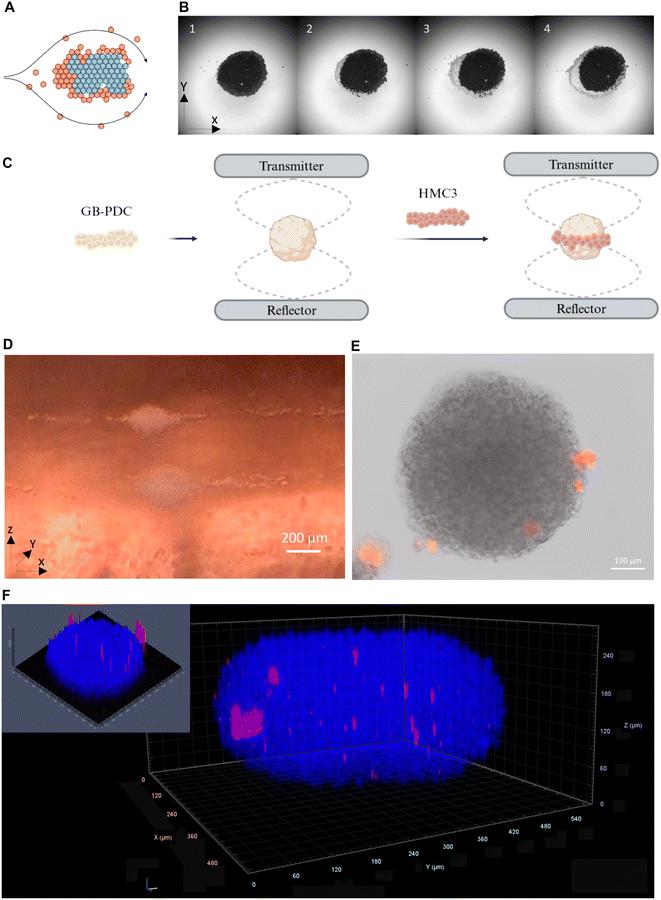
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio after staining microglia with a Cell Tracker Red membrane probe (Fig. 5A). The cultures were performed in multi-node acoustofluidic chips for 72 hours after injection of the mixed cell suspension. The patterning of the mixed cell suspension at acoustic pressure nodes was similar to that of the homogeneous PDC suspension. Fluorescence microscopy imaging of the resulting tumoroids showed patches of red labelled microglia at the periphery of and within the tumoroid (Fig. 5B). Tissue clearing and confocal imaging confirmed the presence of microglial cells inside the tumoroid masses (Fig. 5C), thus evidencing efficient generation of microglia/PDC mixed tumoroids.
1 ratio after staining microglia with a Cell Tracker Red membrane probe (Fig. 5A). The cultures were performed in multi-node acoustofluidic chips for 72 hours after injection of the mixed cell suspension. The patterning of the mixed cell suspension at acoustic pressure nodes was similar to that of the homogeneous PDC suspension. Fluorescence microscopy imaging of the resulting tumoroids showed patches of red labelled microglia at the periphery of and within the tumoroid (Fig. 5B). Tissue clearing and confocal imaging confirmed the presence of microglial cells inside the tumoroid masses (Fig. 5C), thus evidencing efficient generation of microglia/PDC mixed tumoroids.
We next assessed whether the acoustic levitation technique could be amenable to study tumor infiltration by microglial cells. We initially questioned the feasibility of structuring aggregates in concentric layers at the pressure nodes through sequential injection of, first blue-, and then white-colored polystyrene micro-beads (15 μm-diameter, Fig. 6A). As expected, injection of the first bead population resulted in the formation of a stable and densely packed layer of microbeads at the pressure node as shown in Fig 6B1 (axial view from under the aggregate). Subsequent injection of the second bead population resulted in their progressive accumulation around the first layer of beads (Fig 6B2 and B3), through the action of the radial component of the ARF, ending in the formation of 2 concentric layers of beads (Fig 6B4). This protocol was implemented to inject red labelled microglial cells onto preformed PDC tumoroids maintained in acoustic levitation (Fig. 6D). Lateral view imaging showed immediate trapping of microglial cells at pressure nodes through the axial ARF component, prior to their progressive gathering around PDC tumoroids under the action of the transverse component of the ARF (Fig. 6D and Movie S2). The tumoroids were recovered after 2 more days of culture in levitation (Fig. 6E). Fluorescence microscopy imaging and confocal imaging after tissue clearing were used to verify the presence of microglial cells within the tumoroids. Microglia cells were found to have infiltrated the tumoroids, as shown in Fig. 6F. Similar results were obtained using the mouse microglia cell line BV2 (Fig. S3).
In the second part of our work, we leveraged our acoustic bioreactor platform to model the intricate interactions between glioblastoma cells and other cell-intrinsic components of the tumour tissue. Microglia constitute an important fraction of the immune cell component of glioblastoma. While the effect of microglia on glioblastoma cells is mostly modelled through 2D co-culture assays, 3D glioblastoma/microglia cell co-cultures are technically challenging as both cell types have distinct adhesion/migration properties and require challenging techniques such as bioprinting.36,37 By combining fluidic control with the transverse acoustic radiation force, our bioreactor facilitated the formation of controlled 3D co-cultures, enabling the study of microglia infiltration and interactions within a more physiologically relevant context. This advancement addresses a critical challenge in tumour immunology, where traditional 2D co-culture systems fail to replicate the distinct adhesion properties and spatial dynamics of immune cells and glioblastoma cells. Disposing of a simple model accessible to experimental studies of interactions between immune and glioblastoma cells is likely to offer valuable insights into tumor immunology and hence support the development of immunotherapeutic approaches to combat this aggressive cancer. Easy retrieval of the tumoroids from the acoustic bioreactor enables indeed any type of analysis at either the tumoroid or the single cell level, including functional analysis on live cells. Structuration of tumoroids in acoustic levitation is in addition amenable to simulate varying microenvironmental pressures, for example by integrating additional immune cell types in various proportions, thereby enhancing the translational relevance of this model. Overall, our findings highlight the promise of acoustic levitation as a versatile tool for recreating complex tumour microenvironments. These insights open new avenues for exploring glioblastoma biology and notably the dynamics of interactions between malignant and normal cells that constitute this aggressive tumour.
Conclusions
In this study we have engineered a novel acoustic-levitation bioreactor that harnesses the radial component of acoustic radiation force to suspend and grow human glioblastoma spheroids in a scaffold free three-dimensional microenvironment. By precisely controlling the acoustic field, we achieved reproducible self-organization and stable levitation of tumor spheroids while maintaining their viability and proliferative state over several days. While acoustic levitation allowed rapid and reliable formation of heterologous 3D co-culture a notable outcome was the demonstration of the feasibility to achieve tumor spheroid infiltration by injecting additional cell types that migrated radially inward the tumoroid. This 3D co-culture system thus offers an unprecedented platform for studying glioma–immune interactions under physiologically relevant mechanical and biochemical conditions. Importantly, the acoustic approach eliminates the need for scaffold materials such as external matrix components, thereby preserving native cell–cell contacts and minimizing confounding variables. The ability to observe microglial infiltration in a levitated tumor opens new avenues for dissecting the molecular cues that govern recruitment of immune cells and their response to tumor cells. Moreover, the system's scalability and compatibility with live imaging make it an attractive tool for evaluating therapeutic strategies aimed at modulating the tumor microenvironment. By bridging the gap between simplistic 2D cultures and complex animal models, our acoustic levitation bioreactor provides a versatile and reproducible platform that is also ethically favourable. Allowing bioconstruction of concentric, layered, spheroids should be valuable for accelerating translational research in neuro-oncology, immunotherapy and neurosciences.Author contributions
Conceptualization: JMP, JLA, XM, EEH. Methodology: XM, JLA, JMP, EEH, MPJ, HC. Experimental investigations: XM, SK, CD, NJM. Data analyses: XM, SK, CD, NJM, EEH, MPJ, JLA, JMP. Writing – original draft: JMP, XM, MPJ. Review & editing: all. Project administration: JLA, HC. Funding acquisition: JLA, HC, MPJ.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information is available. See DOI: https://doi.org/10.1039/D5LC00249D.Data will be made available upon request.
Acknowledgements
This work was supported by grants from MERT (XM and CD, PhD Fellowship), Chinese Scientific Council (SK PhD Fellowship), Sorbonne Université Emergence Research Program (JLA, JMP), ANR OCTOBOL (JLA, JMP) and La Fondation pour la Recherche Médicale - Equipes FRM (HC). We thank the BRIGHT research team from the ICM institute for providing us with the HMC3 cell line.Notes and references
- A. Bikfalvi, C. A. da Costa, T. Avril, J.-V. Barnier, L. Bauchet, L. Brisson, P. F. Cartron, H. Castel, E. Chevet, H. Chneiweiss, A. Clavreul, B. Constantin, V. Coronas, T. Daubon, M. Dontenwill, F. Ducray, N. Enz-Werle, D. Figarella-Branger, I. Fournier, J.-S. Frenel, M. Gabut, T. Galli, J. Gavard, G. Huberfeld, J.-P. Hugnot, A. Idbaih, M.-P. Junier, T. Mathivet, P. Menei, D. Meyronet, C. Mirjolet, F. Morin, J. Mosser, E. C.-J. Moyal, V. Rousseau, M. Salzet, M. Sanson, G. Seano, E. Tabouret, A. Tchoghandjian, L. Turchi, F. M. Vallette, S. Vats, M. Verreault and T. Virolle, Trends Cancer, 2023, 9, 9–27 CrossRef CAS PubMed.
- A. Buonfiglioli and D. Hambardzumyan, Acta Neuropathol. Commun., 2021, 9, 54 CrossRef PubMed.
- V. Venkataramani, Y. Yang, M. C. Schubert, E. Reyhan, S. K. Tetzlaff, N. Wißmann, M. Botz, S. J. Soyka, C. A. Beretta, R. L. Pramatarov, L. Fankhauser, L. Garofano, A. Freudenberg, J. Wagner, D. I. Tanev, M. Ratliff, R. Xie, T. Kessler, D. C. Hoffmann, L. Hai, Y. Dörflinger, S. Hoppe, Y. A. Yabo, A. Golebiewska, S. P. Niclou, F. Sahm, A. Lasorella, M. Slowik, L. Döring, A. Iavarone, W. Wick, T. Kuner and F. Winkler, Cell, 2022, 185, 2899–2917.e31 CrossRef CAS PubMed.
- F. Wang, X. Liu, S. Li, C. Zhao, Y. Sun, K. Tian, J. Wang, W. Li, L. Xu, J. Jing, J. Wang, S. M. Evans, Z. Li, Y. Liu and Y. Zhou, Protein Cell, 2023, 14, 105–122 CAS.
- Y.-T. Tung, Y.-C. Chen, K. Derr, K. Wilson, M. J. Song and M. Ferrer, Adv. Healthcare Mater., 2024, 13, 2302831 CrossRef CAS PubMed.
- K. Nayernia, M. Li, L. Jaroszynski, R. Khusainov, G. Wulf, I. Schwandt, M. Korabiowska, H. W. Michelmann, A. Meinhardt and W. Engel, Hum. Mol. Genet., 2004, 13, 1451–1460 CrossRef CAS PubMed.
- V. Mangena, R. Chanoch-Myers, R. Sartore, B. Paulsen, S. Gritsch, H. Weisman, T. Hara, X. O. Breakefield, K. Breyne, A. Regev, K. Chung, P. Arlotta, I. Tirosh and M. L. Suva, Cancer Discovery, 2025, 15(2), 299–315 CrossRef CAS PubMed.
- F. Jacob, R. D. Salinas, D. Y. Zhang, P. T. T. Nguyen, J. G. Schnoll, S. Z. H. Wong, R. Thokala, S. Sheikh, D. Saxena, S. Prokop, D.-A. Liu, X. Qian, D. Petrov, T. Lucas, H. I. Chen, J. F. Dorsey, K. M. Christian, Z. A. Binder, M. Nasrallah, S. Brem, D. M. O'Rourke, G.-L. Ming and H. Song, Cell, 2020, 180, 188–204.e22 CrossRef CAS PubMed.
- M. A. B. Andrade, N. Pérez and J. C. Adamowski, Braz. J. Phys., 2018, 48, 190–213 CrossRef.
- M. X. Lim, B. VanSaders and H. M. Jaeger, Rep. Prog. Phys., 2024, 87(6), 1361 CrossRef PubMed.
- D. Bazou, A. Castro and M. Hoyos, Ultrasonics, 2012, 52, 842–850 CrossRef CAS PubMed.
- S. Li, P. Glynne-Jones, O. G. Andriotis, K. Y. Ching, U. S. Jonnalagadda, R. O. C. Oreffo, M. Hill and R. S. Tare, Lab Chip, 2014, 14, 4475–4485 RSC.
- C. Bouyer, P. Chen, S. Güven, T. T. Demirtaş, T. J. F. Nieland, F. Padilla and U. Demirci, Adv. Mater., 2016, 28, 161–167 CrossRef CAS PubMed.
- U. S. Jonnalagadda, M. Hill, W. Messaoudi, R. B. Cook, R. O. C. Oreffo, P. Glynne-Jones and R. S. Tare, Lab Chip, 2018, 18, 473–485 RSC.
- K. Olofsson, B. Hammarström and M. Wiklund, Micromachines, 2018, 9, 594 CrossRef PubMed.
- A. Tait, P. Glynne-Jones, A. R. Hill, D. E. Smart, C. Blume, B. Hammarstrom, A. L. Fisher, M. C. Grossel, E. J. Swindle, M. Hill and D. E. Davies, Sci. Rep., 2019, 9, 9789 CrossRef PubMed.
- K. Olofsson, V. Carannante, M. Takai, B. Önfelt and M. Wiklund, Micromachines, 2021, 12, 329 CrossRef PubMed.
- R. Rasouli, K. M. Villegas and M. Tabrizian, Lab Chip, 2023, 23, 1300–1338 RSC.
- T. Boudreaux, L. Freyhof, B. D. Riehl, E. Kim, R. M. Pedrigi and J. Y. Lim, Bioengineering, 2025, 12, 458 CrossRef PubMed.
- H. R. Sugier, L. Bellebon, J.-L. Aider, J. Larghero, J. Peltzer and C. Martinaud, Cytotherapy, 2023, 25, 891–899 CrossRef CAS PubMed.
- S. Surappa, P. Multani, U. Parlatan, P. D. Sinawang, J. Kaifi, D. Akin and U. Demirci, Lab Chip, 2023, 23, 2942–2958 RSC.
- N. Jeger-Madiot, L. Arakelian, N. Setterblad, P. Bruneval, M. Hoyos, J. Larghero and J.-L. Aider, Sci. Rep., 2021, 11, 8355 CrossRef CAS PubMed.
- N. Jeger-Madiot, X. Mousset, C. Dupuis, L. Rabiet, M. Hoyos, J.-M. Peyrin and J.-L. Aider, J. Acoust. Soc. Am., 2022, 151, 4165 CrossRef CAS PubMed.
- L. Rabiet, L. Arakelian, N. Jeger-Madiot, D. R. García, J. Larghero and J.-L. Aider, Biotechnol. Bioeng., 2024, 121(4), 1422–1434 CrossRef CAS PubMed.
- L. Rabiet, N. Jeger-Madiot, D. R. García, L. Tosca, G. Tachdjian, S. Kellouche, R. Agniel, J. Larghero, J.-L. Aider and L. Arakelian, Sci. Rep., 2024, 14, 21528 CrossRef CAS PubMed.
- O. Dron and J.-L. Aider, Ultrasonics, 2013, 53, 1280–1287 CrossRef CAS PubMed.
- J.-M. Peyrin, B. Deleglise, L. Saias, M. Vignes, P. Gougis, S. Magnifico, S. Betuing, M. Pietri, J. Caboche, P. Vanhoutte, J.-L. Viovy and B. Brugg, Lab Chip, 2011, 11, 3663–3673 RSC.
- M. Hill, J. Acoust. Soc. Am., 2003, 114, 2654–2661 CrossRef PubMed.
- W. Thielicke and R. Sonntag, J. Open Res. Softw., 2021, 9(1) DOI:10.5334/jors.334.
- U. Schmidt, M. Weigert, C. Broaddus and G. Myers, in Medical Image Computing and Computer Assisted Intervention – MICCAI 2018, ed. A. F. Frangi, J. A. Schnabel, C. Davatzikos, C. Alberola-López and G. Fichtinger, Springer International Publishing, Cham, 2018, pp. 265–273 Search PubMed.
- P. B. Muller and H. Bruus, Phys. Rev. E:Stat., Nonlinear, Soft Matter Phys., 2014, 90, 043016 CrossRef PubMed.
- P. B. Muller and H. Bruus, Phys. Rev. E:Stat., Nonlinear, Soft Matter Phys., 2015, 92, 063018 CrossRef PubMed.
- M. Hoyos and A. Castro, Ultrasonics, 2013, 53, 70–76 CrossRef PubMed.
- L. A. Castro and M. Hoyos, Microgravity Sci. Technol., 2016, 28, 11–18 CrossRef.
- M. S. Saurty-Seerunghen, T. Daubon, L. Bellenger, V. Delaunay, G. Castro, J. Guyon, A. Rezk, S. Fabrega, A. Idbaih, F. Almairac, F. Burel-Vandenbos, L. Turchi, E. Duplus, T. Virolle, J.-M. Peyrin, C. Antoniewski, H. Chneiweiss, E. A. El-Habr and M.-P. Junier, Cell Death Discovery, 2022, 13, 913 CrossRef CAS PubMed.
- M. A. Hermida, J. D. Kumar, D. Schwarz, K. G. Laverty, A. Di Bartolo, M. Ardron, M. Bogomolnijs, A. Clavreul, P. M. Brennan, U. K. Wiegand, F. P. Melchels, W. Shu and N. R. Leslie, Adv. Biol. Regul., 2020, 75, 100658 CrossRef CAS PubMed.
- M. Tang, Q. Xie, R. C. Gimple, Z. Zhong, T. Tam, J. Tian, R. L. Kidwell, Q. Wu, B. C. Prager, Z. Qiu, A. Yu, Z. Zhu, P. Mesci, H. Jing, J. Schimelman, P. Wang, D. Lee, M. H. Lorenzini, D. Dixit, L. Zhao, S. Bhargava, T. E. Miller, X. Wan, J. Tang, B. Sun, B. F. Cravatt, A. R. Muotri, S. Chen and J. N. Rich, Cell Res., 2020, 30, 833–853 CrossRef CAS PubMed.
- E. A. El-Habr, L. G. Dubois, F. Burel-Vandenbos, A. Bogeas, J. Lipecka, L. Turchi, F.-X. Lejeune, P. L. C. Coehlo, T. Yamaki, B. M. Wittmann, M. Fareh, E. Mahfoudhi, M. Janin, A. Narayanan, G. Morvan-Dubois, C. Schmitt, M. Verreault, L. Oliver, A. Sharif, J. Pallud, B. Devaux, S. Puget, P. Korkolopoulou, P. Varlet, C. Ottolenghi, I. Plo, V. Moura-Neto, T. Virolle, H. Chneiweiss and M.-P. Junier, Acta Neuropathol., 2017, 133(4), 645–660 CrossRef CAS PubMed.
- H. Bruus, Lab Chip, 2012, 12(6), 1014–1021 RSC.
Footnote |
| † Equal contributions. |
| This journal is © The Royal Society of Chemistry 2025 |