Open Access Article
Open Access ArticleHuman-associated gut bacterial dysbiosis affects the anti-colitic activity of 3-DF and 3-HF
José Antonio
Haro-Reyes
 a,
Jayaprakash Kanijam
Raghupathi
a,
Jayaprakash Kanijam
Raghupathi
 ab,
Bruce
Hamaker
ab,
Bruce
Hamaker
 a and
Lavanya
Reddivari
a and
Lavanya
Reddivari
 *a
*a
aDepartment of Food Science, Purdue University, West Lafayette, IN 47907, USA. E-mail: lreddiva@purdue.edu
bDepartment of Chemistry, Acharya Nagarjuna University, Guntur, Andhra Pradesh, India-522510
First published on 4th November 2025
Abstract
Dysbiosis, defined as an imbalance in the gut microbiota, disrupts the gut barrier and impairs host immune and metabolic responses, playing a critical role in the pathogenesis of inflammatory bowel disease (IBD). Dietary flavonoids have been explored as safer alternatives to conventional drugs, showing efficacy in alleviating inflammation and restoring barrier integrity in colitis models. However, whether their anti-colitic activity is preserved under human microbial dysbiosis remains unclear. Here, we investigated the impact of ulcerative colitis (UC)-associated microbiota on the efficacy of 3-deoxyflavonoids (3-DFs) and 3-hydroxyflavonoids (3-HFs), using a humanized interleukin (IL)-10−/− mouse model. Four near-isogenic lines (NILs) of corn, Lines A (lacking 3-DF or 3-HF), B (expressing only 3-DF), C (expressing only 3-HF), and D (expressing both 3-DF + 3-HF), were incorporated into experimental diets, alongside a purified control. Mice were colonized with fecal microbiota from UC patients and fed the respective diets. The effects of flavonoid-rich diets on distal colon inflammatory markers, tight-junction expression, and disease activity varied according to the microbial composition of the three UC donors. Higher concentrations of gut metabolites, including short-chain fatty acids (SCFAs) and bile acids, were associated with reduced inflammation and enhanced barrier function. Overall, the composition of dysbiotic human microbiota influenced the anti-colitic activity of dietary flavonoids. When protective effects were observed, diets containing 3-HF were more effective than those with only 3-DF. These findings highlight the need to account for individual microbial profiles when designing flavonoid-based dietary interventions for IBD.
1. Introduction
Gut microbial dysbiosis is critical in the development of various gastrointestinal disorders, including inflammatory bowel disease.1 Dysbiosis refers to an imbalance in the gut microbiota, characterized by reduced microbial diversity, the loss of keystone taxa, the overgrowth of pathobionts or pathogenic species, and shifts in metabolic capacity.2,3 The origins of dysbiosis are multifactorial, involving genetic background, health status, diet, and exposure to xenobiotics, which can directly or indirectly influence microbiota composition. Dietary choices, for instance, can influence the risk and progression of the disease based on fiber content, fruit and vegetable consumption and energy density.4 Dysbiosis disrupts the gut barrier and impairs host immune response and metabolism, facilitating disease development.3Food phytochemicals have emerged as potential alternatives to prevent or ameliorate IBD symptoms, offering safer options compared to commercial drugs.5 Flavonoids, such as anthocyanins or 3-hydroxyflavonoids (3-HF), and phlobaphenes or 3-deoxyflavonoids (3-DF) in the food matrix, have demonstrated anti-colitic activity.6–8 The attributes of the food matrix, including composition and particle size, influence the bioaccessibility of flavonoids, affecting their delivery to the large intestine.9 In the colon, flavonoids can modulate the gut microbiota and its metabolic outputs and microbiota can, in turn, regulate flavonoid metabolism and bioactivity.10,11
Replicating the conditions of dysbiosis in the gut microbiota and evaluating modulation with flavonoids can be challenging. Humanizing germ-free mice with fecal microbiota from dysbiotic patients is a viable approach to replicate dysbiosis and to understand the role of flavonoids, despite limitations in colonization efficiency.12,13 Furthermore, gut microbial communities vary in their robustness or plasticity, maintaining or adapting their functional states despite compositional shifts after disturbances.14 This variability must be considered when using communities from different donors. Additionally, community resilience is lower in animals not exposed to environmental microbiota, such as those used in humanization experiments under gnotobiotic housing.15
Our approach delivers distinct flavonoid profiles within a food matrix to an IL-10−/− mouse model colonized with dysbiotic human microbiota. The knockout of IL-10 gene induces colitis through an immunological pathway, while dextran sulfate sodium (DSS) primarily disrupts the gut barrier.16 The IL-10−/− model therefore recapitulates colitis driven by IL-10 receptor mutations or gene deficiencies, both of which are strongly associated with early-onset IBD.17 Colonization with dysbiotic human microbiota in germ-free animals further establishes a characteristic microbial community linked to inflammatory responses.18 The objective of this study was to investigate the impact of dysbiotic microbiota derived from individuals with UC on the anti-colitic properties of two flavonoid groups, 3-deoxyflavonoids and 3-hydroxyflavonoids.
2. Materials and methods
2.1. Diets
For the dietary treatments, four near-isogenic lines (NILs) of corn were selected from the Penn State Russel Larson Agronomy Research Farm (Rocksprings, PA, USA), each exhibiting different profiles of phlobaphenes and anthocyanins. Specifically, Maize Line A lacked both flavonoids, Line B expressed only phlobaphenes, Line C expressed only anthocyanins, and Line D expressed both flavonoids. The NILs were developed by targeted backcrossing of maize inbred line 4Co63 (line A) with donor stocks carrying functional pigmentation alleles that regulate 3-DF or 3-HF within the phenylpropanoid pathway. The presence of phlobaphenes and abundance of anthocyanins in corn NILs were verified using stablished methods (SI Fig. S1 and Table S1).19 For these compounds to exert anti-colitic activity, dietary 3-DFs and 3-HFs must undergo metabolism by the gut microbiota, as demonstrated in murine models, where they restore gut barrier integrity and suppress inflammatory cytokine production, particularly 3-HFs.6,7 While the NF-κB pathway is the primary target studied in relation to anthocyanin-mediated effects, additional signaling cascades, including AMPK/mTOR, have also been implicated.8,20,21The diets were formulated based on the AIN-93G diet composition, incorporating 25% dry matter (DM) from one of the maize lines, as detailed in SI Table S2. To ensure nutritional balance, the diets were standardized for energy, fat, and protein content. A purified diet (P diet) without maize content was used as a control. All diets were manufactured by Envigo Teklad (Madison, WI, USA). Further details on the genetic background, metabolite content, and development of the NILs were published earlier.19
2.2. Experimental design
All animal procedures were conducted in accordance with the guidelines for animal care and use established by Purdue University and were approved by the Purdue Institutional Animal Care and Use Committee (protocol no. 1810001817). Six-week-old germ-free IL-10−/− mice (C57BL/6) from both sexes were obtained from the Purdue University Animal Facility (West Lafayette, IN, USA). These mice were colonized via oral gavage with human fecal microbiota obtained from ulcerative colitis (UC) patients. The fecal samples were collected at Seattle Children's Hospital from donors aged 14 to 18 years, all of whom had active disease at the time of collection, with calprotectin levels ranging from 1200 to 1800 μg g−1, indicating severe disease. Fecal sample collection and use were approved by the Seattle Children's Hospital Institutional Review Board (IRB STUDY00000619) and conducted in accordance with the Belmont Report, with written informed consent obtained from all patients. Although constrained by inherent limitations, fecal microbiota transplantation (FMT) from human patients remains an optimal approach for establishing causal relationships and elucidating mechanisms underlying microbiota-associated diseases.22 This strategy has expanded the range of research questions that can be addressed, including the inflammatory potential of microbiota from healthy versus UC-affected twins in the offspring and the association of UC-derived microbiota with altered expression of genes linked to anxiety and depression.23,24The study was conducted in accordance with the Declaration of Helsinki and approved by the Institutional Review Board of Purdue University IRB-2021-109 (12/9/2022). Mice were housed under controlled conditions with a 12 h light–dark cycle, provided food and water ad libitum, and their food intake was recorded (SI Fig. S2). Animals were randomly assigned to one of the experimental diets (P, A, B, C, and D), with three to five mice per donor, and maintained on these diets for a total of eight weeks (Fig. 1). In the final week of the study, mice received 1.25% dextran sodium sulfate (DSS) (CAS No. 9011-18-1, TdB Labs AB, Uppsala, Sweeden) in their drinking water for six days to induce mild colonic inflammation.
This specific DSS concentration was chosen to prevent the early onset of severe disease symptoms, as observed in preliminary trials. One day after the removal of DSS, the mice were euthanized by CO2 asphyxiation. Blood samples were collected via cardiac puncture, and organs including the colon, cecum, liver, kidneys, and spleen were harvested, weighed, and either processed immediately or stored for subsequent analysis. Additionally, fecal content from the colon and cecum was collected and stored at −80 °C.
2.3. Disease activity index (DAI)
The Disease Activity Index (DAI) was calculated based on a previously published method,25 with some modifications. Briefly, following DSS treatment, four parameters were assessed and graded according to the severity observed. The DAI was determined by summing the individual scores for weight loss (0–3, 0: no loss; 1: up to 5% loss; 2: from 5 to 10% loss; 3: >20% loss), stool consistency (0–3, 0: formed; 1:mild-soft; 2:very soft; 3: watery diarrhea), rectal bleeding/blood in the stool (0: normal; 1: week appearance of blood; 2: blood in stool; 3: gross bleeding), and overall condition of the mice (0: normal; 1: poor condition [respiratory distress, lethargy, ruffle coat and/or hunched posture]).2.4. Intestinal permeability
Intestinal permeability was evaluated using FITC-dextran (No. 46944, Sigma–Aldrich, St Louis, MO, USA). Mice were fasted for 3.5 hours prior to euthanasia. Two hours before euthanasia, they were administered with FITC-dextran solution at a concentration of 100 mg ml−1, corresponding to 0.6% of their body weight (w/v). Blood samples were collected via cardiac puncture. The fluorescence intensity of the samples was measured at 490 nm excitation and 520 nm emission, using a BioTek™ Cytation™1 Cell Imaging Reader (Thermo Fisher Scientific Inc., Winooski, VT, USA).2.5. Distal colon biomarkers
The mRNA expression levels of toll-like receptors, pro-inflammatory cytokines, nuclear transcription factors, tight-junction proteins, and Muc2 were analyzed. Total RNA was extracted from distal colon tissues using the PureLink Mini Kit (No. 12183018A, Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. A standardized concentration of 100 ng RNA/μl was used for cDNA synthesis, which was carried out using the High-Capacity cDNA Reverse Transcription Kit (No. 4374966, Applied Biosystems, Waltham, MA) following the manufacturer's protocol. Real-time PCR amplification and detection were conducted on a QuantStudio 3 PCR instrument. The PCR protocol included an initial denaturation step at 95 °C for 10 minutes, followed by 45 cycles of 95 °C for 10 seconds, annealing at 55 °C for 60 seconds. mRNA expression levels were normalized to β-actin. The primers used for DNA amplification are listed in SI Table S3.2.6. Gut microbiota 16S rRNA gene sequencing, diversity and compositional metrics
DNA was extracted from the cecal content of mice using the QIAmp PowerFecal Pro DNA Kit (No. 51804, QIAGEN, Germantown, MD, USA) according to the manufacturer's instructions. The DNA concentration was quantified using a Take3 microvolume plate (BioTek, Winooski, VT, USA) in a BioTek Cytation1 microplate reader (Winooski, VT, USA). The DNA extracts were then processed at the RUSH Genomics and Microbiome Core Facility (RUSH University, Chicago, IL, USA). The hypervariable V4 region (515F–806R) was amplified, and library preparation was performed using the Illumina MiniSeq Platform (Illumina, Inc., San Diego, CA, USA). Demultiplexed sequences were merged using the Pair-end read merger (PEAR) version 0.9.11 and analyzed with the Quantitative Insights into Microbial Ecology2 (QIIME2) Platform version 2022.18. The sequences were filtered and denoised using the DADA2 pipeline. Taxonomy assignment was performed at the family level using the Silva 138 database as a reference. Alpha and beta diversity indices were estimated in QIIME2. Only bacteria with a relative abundance greater than 1% were included in the taxa-bar plots.2.7. Extraction and quantification of targeted gut metabolites
SCFAs were extracted from murine cecal contents. Briefly, 50 mg of sample was treated with 0.5% phosphoric acid (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 12, v/w) and disrupted via bead beating. The homogenate underwent centrifugation (17
12, v/w) and disrupted via bead beating. The homogenate underwent centrifugation (17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g, 4 °C, 10 min), and the supernatant was extracted with ethyl acetate containing heptanoic acid as an internal standard (1
000g, 4 °C, 10 min), and the supernatant was extracted with ethyl acetate containing heptanoic acid as an internal standard (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v). Following vortexing and a secondary centrifugation, the ethyl acetate phase was recovered for analysis. SCFA standards, including acetate, propionate, and butyrate, were procured from Sigma-Aldrich (St Louis, MO, USA). Quantification was performed using gas chromatography (Agilent 7890A GC-FID) with a Nukon Supelco No. 40369-03A silica capillary column. The temperature gradient initiated at 50 °C (2 min), increased stepwise to 70 °C (10 °C min−1), then to 85 °C (3 °C min−1), 110 °C (5 °C min−1), and 290 °C (30 °C min−1), holding at 290 °C for 8 min. Helium was used as the carrier gas (1 mL min−1, constant flow).
1, v/v). Following vortexing and a secondary centrifugation, the ethyl acetate phase was recovered for analysis. SCFA standards, including acetate, propionate, and butyrate, were procured from Sigma-Aldrich (St Louis, MO, USA). Quantification was performed using gas chromatography (Agilent 7890A GC-FID) with a Nukon Supelco No. 40369-03A silica capillary column. The temperature gradient initiated at 50 °C (2 min), increased stepwise to 70 °C (10 °C min−1), then to 85 °C (3 °C min−1), 110 °C (5 °C min−1), and 290 °C (30 °C min−1), holding at 290 °C for 8 min. Helium was used as the carrier gas (1 mL min−1, constant flow).
Bile acids were extracted from mouse serum following a previously described method26 with slight modifications. Briefly, 50 μL of serum were combined with 150 μL of cold methanol containing 0.4 μg mL−1 of GCA as an internal standard. The mixture was incubated at −20 °C for 20 minutes, then centrifuged at 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 20 minutes at 4 °C. The supernatant was retained for analysis. Targeted bile acids were quantified by LC–MS (Agilent 1260 Infinity II HPLC coupled to an Ultivo triple quadrupole mass spectrometer). Primary unconjugated bile acids included α-muricholic acid (α-MCA) and β-muricholic acid (β-MCA; both from Avanti Research, Alabaster, AL, USA), as well as cholic acid (CA) and chenodeoxycholic acid (CDCA; both from Sigma–Aldrich, St Louis, MO, USA). Conjugated bile acids, including taurochenodeoxycholic acid (TCDCA), tauroursodeoxycholic acid (TUDCA), and glycocholic acid (GCA), were also from Sigma-Aldrich. The HPLC gradient and mass spectrometry conditions followed a previous method.27 Mobile phase solvents consisted of 20 mM ammonium acetate + 0.1% formic acid and methanol. A Supelco Ascentis Express No. 53829-U C18 150 × 4.6, 2.7 μm column was used for separation. Data were acquired in negative mode using targeted multiple reaction monitoring (MRM) for compound identification.
000g for 20 minutes at 4 °C. The supernatant was retained for analysis. Targeted bile acids were quantified by LC–MS (Agilent 1260 Infinity II HPLC coupled to an Ultivo triple quadrupole mass spectrometer). Primary unconjugated bile acids included α-muricholic acid (α-MCA) and β-muricholic acid (β-MCA; both from Avanti Research, Alabaster, AL, USA), as well as cholic acid (CA) and chenodeoxycholic acid (CDCA; both from Sigma–Aldrich, St Louis, MO, USA). Conjugated bile acids, including taurochenodeoxycholic acid (TCDCA), tauroursodeoxycholic acid (TUDCA), and glycocholic acid (GCA), were also from Sigma-Aldrich. The HPLC gradient and mass spectrometry conditions followed a previous method.27 Mobile phase solvents consisted of 20 mM ammonium acetate + 0.1% formic acid and methanol. A Supelco Ascentis Express No. 53829-U C18 150 × 4.6, 2.7 μm column was used for separation. Data were acquired in negative mode using targeted multiple reaction monitoring (MRM) for compound identification.
2.8. Statistical analysis
The analysis of the data was carried out using two distinct approaches for comparing treatments: one that pooled all results per treatment regardless of the donor, and another that compared treatments within individual donors. IBM SPSS Statistics 29.0 was used for all data analysis, except for bacterial sequencing, with GraphPad Prism 10 employed for graphing. To assess normality, the Shapiro–Wilk test was applied. When normality was confirmed, a one-way ANOVA was performed. Post-hoc comparisons of the means were conducted with Tukey's test, or, when Levene's test indicated heterogeneity of variances, with the Games-Howell test. Outliers were excluded from the analysis, and in cases of non-normality, non-parametric tests such as Kruskal–Wallis or Mann–Whitney were used. For microbial sequencing, data analysis was performed using R Studio version 2024.04.0 or Qiime2. Alpha diversity was analyzed using Kruskal–Wallis pairwise tests, while pairwise PERMANOVA was applied to assess beta diversity. Compositional differences were tested using ANCOM-BC. To examine associations between bacterial taxa and other biomarkers, Spearman's correlation was employed. Correlation strengths were categorized as mild (r < 0.4), moderate (0.4 < r < 0.6), or strong (r > 0.6). Statistical significance was reported as: #<0.10, *<0.05.3. Results
The effects of dietary interventions were analyzed by initially grouping all recipients irrespective of human fecal microbiota donors within the diet group, followed by disaggregation based on individual donor groups.3.1. Body weight, disease activity and anatomical changes
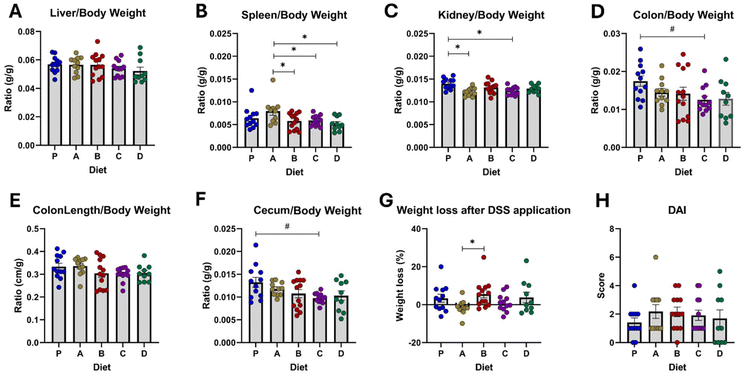
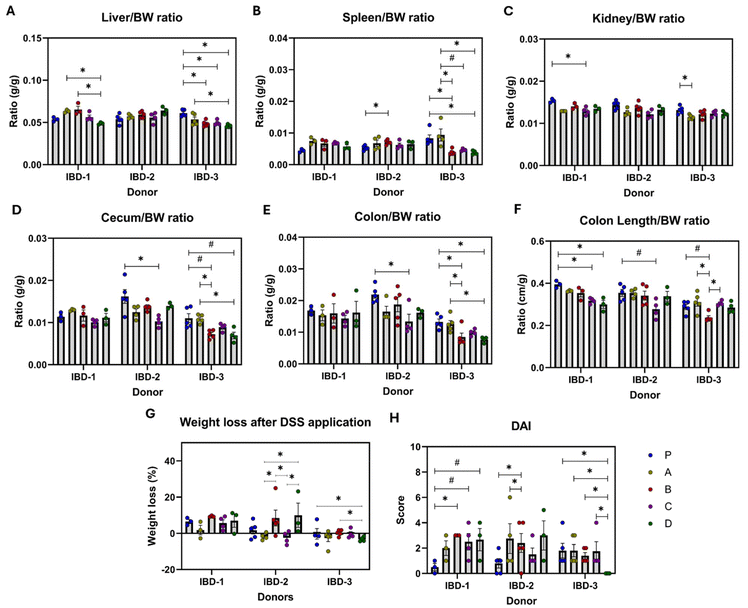
The corn matrix, with NILs containing or not anthocyanins (diets C and A respectively), reduced kidney weight compared to the control diet (Fig. 2C). Additionally, the 3-HF diet showed potential benefits, as evidenced by a reduction in colon and cecum weight. Conversely, the 3-DF diet showed the potential to exacerbate weight loss compared to the corn matrix (Fig. 2G). The differences observed through this overall analysis did not appear to be exclusively triggered by a single donor's microbiota (Fig. 3).From a single-donor perspective, flavonoid-rich diets exacerbated the disease activity index (DAI) in IBD-1 recipients. Specifically, diet C led to an increase in kidney weight, while both 3-HF-containing diets resulted in a reduction in colon length. In IBD-2 recipients, diet B worsened DAI and increased spleen weight, whereas diet C was effective in reducing colon and cecum weight but induced a decrease in colon length. Although none of the corn-based diets influenced weight loss significantly, the presence of 3-DFs showed a trend towards increasing weight loss when compared to corn-based diets (Fig. 3G). Notably, in IBD-3 recipients, diets containing flavonoids provided the most pronounced benefits. Diets B and D were associated with a reduction in liver, spleen, colon, and cecum weight, while only the combined administration of 3-DF + 3-HF alleviated weight loss and improved DAI (Fig. 3G and H). Diets A and C reduced liver and kidney weight, respectively (Fig. 3A and C).
3.2. Pattern recognition receptors (PRR) and inflammatory markers
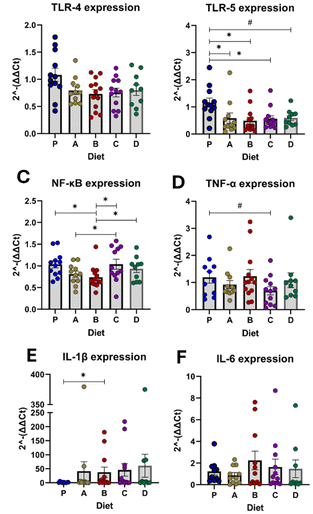
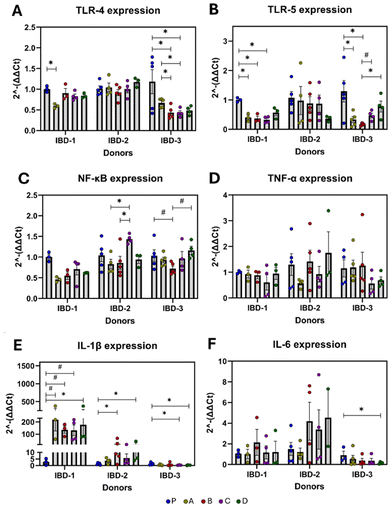
All corn matrix diets reduced toll-like receptor (TLR)-5 expression; however, only diet B significantly reduced nuclear factor (NF)-κB expression compared to 3-HF alone or in combination with 3-DF. The 3-DF diet also led to an increase in interleukin (IL)-1β levels (Fig. 4E), a finding consistent with the average responses across all three donor-recipient groups. Moreover, diet C exhibited a trend toward reducing tumor necrotic factor (TNF)-α expression.When the analysis was disaggregated by the recipient's donor group (Fig. 5), in IBD-1 recipients, a consistent pattern of proinflammatory activity was observed, characterized by elevated IL-1β expression in all flavonoid-rich diets, with diet D showing the most significant effect. Notably, all diets, except for diet D, reduced TLR-5 expression. In IBD-2 recipients, only diets containing 3-DF increased IL-1β, suggesting that the 3-HF diet had the least impact on inflammatory markers. Diet C also showed potential to elevate NF-κB expression, but only when compared to the corn-based diets. In contrast, in IBD-3 recipients, the flavonoid-rich diets demonstrated the most pronounced anti-inflammatory effects. Both diets B and C reduced TLR-4 expression compared to control and diet A. Only diet B reduced TLR-5 expression, which was not changed by the 3-HF-containing diets. Diets C and D exhibited the most anti-inflammatory activity, as evidenced by the reduced expression of IL-1β and IL-6 (Fig. 5E and F).
3.3. Tight-junction proteins and Gut barrier integrity
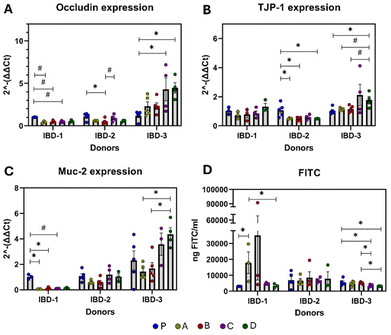
Tight-junction protein expression was unaffected by the corn matrix, regardless of flavonoid content (Fig. 6A and B). However, Muc2 expression was reduced by the corn matrix without flavonoids and by diet B (Fig. 6C) and the other diets did not significantly alter Muc2 expression. Additionally, gut permeability, assessed via fluorescein isothiocyanate (FITC), was decreased by diet D, indicating that the combination of 3-DF + 3-HF was more effective than when either component was included individually.In IBD-1 recipients, only diet D did not impact the barrier integrity markers. The remaining diets exhibited a trend toward reducing occludin and Muc2 expression, with diet A significantly increasing gut permeability. Conversely, in IBD-2 recipients, only tight-junction proteins were affected. Diet B reduced both occludin and tight junction protein (TJP)-1 expression, while diet D reduced only TJP-1, suggesting an effect of 3-DF in this recipient group. In contrast, flavonoid-rich diets demonstrated gut barrier protective effects in IBD-3 recipients. Diets containing 3-HF increased occludin expression and reduced gut permeability, suggesting gut barrier function restoration compared to the 3-DF diet (Fig. 7C and D). Furthermore, diet D also increased TJP-1 expression, exhibiting a pattern of enhancement greater than diets A and B.
3.4. Microbial diversity and composition
The corn matrix, and its interactions with flavonoids, did not significantly impact alpha diversity measurements across the dietary treatments (SI Fig. S3). Similarly, beta-diversity analysis (Fig. 8) showed no clustering by diet intervention, but recipient mice were clustered according to the microbiota composition of the human donor. Notably, in the Bray–Curtis plot, IBD-3 donor groups showed that all corn-based diets clustered separately from the P diet (Fig. 8B), indicating distinct composition and distribution. However, clusters approached when phylogenetic information was incorporated into the Weighted UniFrac analysis (Fig. 8D). Additionally, IBD-1 recipients differed from the IBD-2 and 3 recipients when the phylogenetic similarities were accounted for in the weighted UniFrac analysis (Fig. 8).Microbial composition showed limited modulation by the corn matrix and flavonoid-rich diets when all recipient donor groups were analyzed together (SI Table S4 and Fig. 9). However, Erysipelatoclostridiaceae was the taxonomic family with relative abundance >1%, which was significantly affected by diets A and B across donor groups. The reduction in this family was primarily driven by IBD-2 and IBD-3 recipient groups, highlighting the challenge of generalizing diet-induced changes across distinct microbiota communities.
When microbiota relationships were analyzed within human donor's recipient group, distinct dietary modulation was observed in each group (Fig. 10). In the IBD-1 recipient group, all corn-based diets decreased the abundance of Pasteurellaceae, Erysipelotrichaceae, and Oscillospiraceae. Moreover, the corn matrix without flavonoids increased Clostridiaceae abundance, while diet B reduced Lachnospiraceae, diet C reduced Peptostreptococcaceae, and diet D significantly increased Lachnospiraceae while reducing Clostridiaceae. In IBD-2 recipients, corn-based diets consistently increased the abundance of Sutterellaceae and Enterococcaceae, with diet D maintaining Enterococcaceae levels. Notably, diet D exhibited the most distinct modulation of the microbiota, particularly by promoting Clostridiaceae. Diet A increased Ruminococcaceae and reduced Erysipelatoclostridiaceae, with a microbiota profile similar to that of diet B. Diets B and C had similar effects on bacterial composition, but compared to the control, diet B promoted Enterobacteriaceae and reduced Lachnospiraceae, while diet C increased Erysipelotrichaceae. In IBD-3 recipients, all corn-based diets reduced Enterococcaceae levels. However, diet A had a higher level of Enterococcaceae compared to the flavonoid-rich diets. Simultaneously, diet A increased Erysipelotrichaceae but reduced Erysipelatoclostridiaceae. Diet B showed a similar profile to diet A, except for Enterococcaceae levels. In contrast, diet C reduced Enterococcaceae and maintained Sutterellaceae, whereas Sutterellaceae was depleted in the other treatments. Finally, diet D exhibited a pattern similar to diet C but led to a depletion of Sutterellaceae.
3.5. Microbial metabolic products: SCFA and bile acids
Corn-based diets and flavonoids did not improve the concentration of SCFAs (Fig. 11). However, diet D reduced butyrate levels, consistent with the patterns observed in each recipient group (SI Fig. S4), based on the human donor source.Overall, primary bile acids were mostly unaffected by corn-based diets (Fig. 12), except for diet B reducing CDCA and diet D increasing the concentration of CA, primarily in IBD-3 recipients (Fig. 13C and D). Interestingly, IBD-3 recipients accounted for the most notable differences between diets compared to other recipient groups. Specifically, in IBD-3 recipients, all corn-based diets elevated GC significantly (SI Table S5), while the corn diet without flavonoids increased CA and β-MCA. Concurrently, diets B and C reduced CDCA concentration in serum, while diet D increased CA and GCA, and decreased TUDCA. Furthermore, only α-MCA was reduced by diet D in IBD-2 recipients, and no significant differences were found in IBD-1 recipients for any of the bile acids measured in this study.
3.6. Associations between gut bacteria, gut metabolites, and inflammatory and barrier protection markers
Bacterial families were correlated with markers associated with the inflammation (PRRs and pro-inflammatory cytokines) and gut barrier function (mucus production and tight junction protein genes). Interestingly, biomarkers within each group responded similarly in their correlations to bacteria, except for NF-κB, which clustered with the gut barrier protection genes (SI Fig. S5).Bacteria with relative abundance higher than 1% were grouped based on their association with gut barrier protection and inflammation. One group was composed of Eggerthellaceae, Marinifilaceae, Tannerellaceae, and Rikenellaceae, which showed a strong positive correlation with occludin and Muc2, and a moderate correlation with TJP-1. Conversely, these bacteria were negatively correlated with IL-1β (high correlation), IL-6, and TLR-4 (moderate correlation). A second group consisted of Bacteroidaceae, Ruminococcaceae, and Lachnospiraceae. Although hierarchically separated from the first group, they exhibited similar correlation patterns for occludin, Muc2, and IL-1β. A third group, composed of Enterobacteriaceae, Pasteurellaceae, Enterococcaceae, Veillonellaceae, Peptostreptococcaceae, and Clostridiaceae, was highly positively correlated with IL-1β but negatively correlated with occludin and Muc2. Furthermore, Enterobacteriaceae and Enterococcaceae were highly negatively associated with TJP-1, and moderately positively associated with IL-6 and TLR-4. In addition to these groups, NF-κB had a unique correlation pattern. It moderately correlated with Bacteroidaceae (positively) and with Pasteurellaceae and Veillonellaceae (negatively). Finally, only weak correlations were found between TLR-5 and TNF-α with bacterial composition.
Among the SCFAs, propionate and butyrate showed opposite association dynamics (SI Fig. S6). Strong positive correlations between propionate and Bacteroidaceae were found, along with moderate correlations with Eggerthellaceae, Marinifilaceae, Tannerellaceae, Rikenellaceae, Ruminococcaceae, and Erysipelotrichaceae. Conversely, negative and moderate correlations were observed between propionate and Enterobacteriaceae, Pasteurellaceae, Peptostreptococcaceae, and Clostridiaceae. Regarding butyrate, Butyricicoccaceae, Eggerthellaceae, Marinifilaceae, Tannerellaceae, and Rikenellaceae exhibited negative and moderate correlations, in contrast to Pasteurellaceae, Peptostreptococcaceae, and Clostridiaceae, which showed positive correlations.
GCA exhibited a distinct response in its correlations with bacterial composition (SI Fig. S7). GCA was strongly positively correlated with Eggerthellaceae, Marinifilaceae, Tannerellaceae, Rikenellaceae, Ruminococcaceae, and moderately correlated with Lachnospiraceae, Butyricicoccaceae, and Streptococcaceae. Conversely, moderate negative correlations were observed with Enterobacteriaceae, Enterococcaceae, Pasteurellaceae, Peptostreptococcaceae, and Clostridiaceae. Other bile acids presented weaker interactions, except for β-MCA and CA. β-MCA exhibited moderate positive correlations with Veillonellaceae, Pasteurellaceae, Peptostreptococcaceae, and Clostridiaceae, but negative correlations with Bacteroidaceae and Butyricicoccaceae. Similarly, CA showed similar correlations with the same bacteria as β-MCA but was also negatively associated with Coriobacteriaceae and Sutterellaceae. In addition, CDCA exhibited similar correlations to β-MCA, except for Bacteroidaceae and Veillonellaceae.
The correlations between gut metabolites and inflammation and barrier function markers (Fig. 14) to better understand the consequences of dysbiotic microbiota and host health were analyzed. We categorized the parameters evaluated: markers of the NF-κB pathway, including PRRs, pro-inflammatory cytokines, and nuclear factors; markers of gut barrier integrity (occludin, TJP-1, Muc2); gut metabolites (SCFAs and bile acids); and DAI.
We observed the most relevant relationships within categories, focusing on strong correlations (r > 0.6). Among the gut barrier markers, occludin showed a very strong correlation with Muc2 and a strong correlation with TJP-1. Among the gut metabolites, strong positive correlations were found between CA and β-MCA, TUDCA and TCDCA, as well as negative correlations between propionate and β-MCA.
Secondly, when comparing correlations between categories, we found very strong negative correlations between IL-1β and Muc2 or occludin, demonstrating the inverse regulation between inflammation and barrier restoration. Additionally, propionate and GCA appeared to be associated with gut barrier protection, as they presented strong positive correlations with Muc2 and occludin, but negative correlations with IL-1β.
4. Discussion
Dysbiosis entails both compositional and functional shifts in the gut microbiota.28 Expansion of pathogens, such as Clostridium difficile, Bacteroides fragilis or Escherichia coli (AIEC) has been linked to IBD dysbiosis, yet patient microbiota profiles remain heterogeneous.29,30 Dysbiosis reshapes colonic immune responses through pathogen-associated molecular patterns and perturbs microbial metabolite production.31–34 Dietary flavonoids can counteract these disturbances after undergoing microbial hydrolysis, reduction, and deglycosylation. The resulting phenolic metabolites serve as energy substrates and modulate microbial composition and function.7,10,35,36 Although anti-colitic effects have been documented with healthy human microbiota,27 their interaction with dysbiotic, UC-associated communities remains largely unexplored and is the focus of this study.Overall, corn diets were ineffective in alleviating inflammation or gut barrier dysfunction (Fig. 4 and 6), except for diet D, which reduced barrier permeability (Fig. 6D). This improvement was primarily observed in IBD-3 recipients (Fig. 7D). Additionally, corn diets led to the downregulation of TLR-5 (Fig. 4B), which was associated with the disease, as observed in mice with DSS-induced colitis and in IBD patients.37,38 However, TLR-5 downregulation did not appear to be a determining factor in the inflammatory status or gut barrier function.
Conversely, a donor-based analysis revealed that flavonoid-rich diets alleviated colitis in one donor group (IBD-3 recipients) (Fig. 5A, E, F and 6A, B and D), with limited activity in the other two. In this group, when flavonoid-rich diets were effective against colitis, Enterococcaceae was significantly reduced (Fig. 10). We hypothesize that the reduction of Enterococcaceae may be associated with disease alleviation, as this commensal taxon is notably increased in IBD patients.39,40 Mucin-degrading species within the Enterococcaceae family are linked to a cross-feeding dynamic that culminates in gut barrier disruption.41 IBD-3 recipients also harbored Marinifilaceae and Tannerellaceae, which were largely absent in other groups (<1% relative abundance). Marinifilaceae is more abundant in healthy individuals than in UC patients.42Tannerellaceae is negatively correlated with lipocalin-2 and positively with body weight in a DSS model,43 consistent with weight gain patterns in IBD-3 recipients compared to the others (Fig. 3). Some Tannerellaceae members are H2 producers, and microbial hydrogen economy has been implicated in maintaining barrier integrity under DSS challenge.44 Notably, we observed that anthocyanin-rich diets in IBD-3 recipients restored gut barrier function, and when combined with phlobaphenes, they also reduced pro-inflammatory cytokines and disease activity. The benefits of the 3-DF + 3-HF combination surpassed the single action of 3-DFs in diet B (Fig. 5E, F, 6D, 7B and D). The anti-colitic effects observed are consistent with previous reports using the same NILs of corn in mouse models.7,19 Furthermore, the superior performance of the 3-DF + 3-HF diet suggests potential synergisms conditioned by the microbiota composition and metabolism. On the other hand, in recipients of IBD-1 or IBD-2 donors, corn diets did not show improvement in inflammation or barrier integrity-associated biomarkers, and some parameters were even aggravated. However, the microbiota compositional changes differed in each group. Interestingly, although IBD-2 and IBD-3 recipient communities were more similar according to Weighted UniFrac β-diversity (Fig. 8D), IBD-1 and IBD-2 recipients were more similar in their response to the diets (Fig. 5 and 7). This aligns with the significant inter-individual variability in human microbiota,1 particularly in IBD patients, whose communities are more distinct than those of healthy individuals.45 Prior studies have demonstrated that inter-individual differences in microbial diversity or baseline taxonomic abundance can lead to divergent responses to dietary interventions.46 The similar responses of IBD-1 and IBD-2 recipients may be explained by the functional redundancy of their microbiota communities,47 whereby distinct microbial compositions can yield comparable metabolic outcomes. In IBD-1, the more affectation of the gut barrier and higher Clostridiaceae abundance were observed when flavonoids were absent from the diets (Fig. 7C, D and 10). Conversely, when the diet was rich in both 3-DF and 3-HF, Clostridiaceae was reduced, and the disturbance of the gut barrier did not progress further, although inflammation remained significant (Fig. 5E). Some members of the Clostridiaceae family are more abundant in UC patients,48 and their reduction accompanies effective treatment of colitis symptoms, as shown with Clostridium_sensu_stricto in IL-10−/− mouse models.49 Additionally, the combined effect of 3-DF + 3-HF did not increase gut permeability as the corn matrix did (Fig. 7D). In IBD-2 recipients, anthocyanins improved a few anatomical responses (Fig. 2D and E) and did not exacerbate inflammatory and gut barrier damage (Fig. 5 and 7). Anthocyanins also increased the Erysipelotrichaceae relative abundance (Fig. 10) and elevated the propionate concentration (SI Fig. S4), which were moderately and positively correlated (SI Fig. S6). The role of Erysipelotrichaceae in IBD is controversial, although it is closely related to inflammatory disorders in colon cancer.50 This bacterial family was also elevated when 5% black raspberry extract was provided to mice in a DSS colitis model, but only under dietary conditions of a total western diet and not with a healthy diet.51 Altogether, in scenarios where microbial compositions limit the anti-colitic activity of flavonoid-rich matrices, 3-DF or 3-HF can alternatively prevent the worsening of the condition depending on their interaction with the particular microbiota profile.
Examining the targeted gut metabolites and their relationship with biomarkers, propionate and GCA were highly associated with the recovery of gut barrier integrity and the reduction of pro-inflammatory cytokines (Fig. 14). Both metabolite concentrations were higher in IBD-3 recipients (SI Fig. S4 and Table S5), where anti-colitic activity was observed. This is consistent with propionate's properties, such as reducing neutrophil release, decreasing pro-inflammatory molecules (NF-κB, TNF-α, IL-1β, and IL-6), and preserving tight junctions and gut permeability.52,53 Moreover, GCA has demonstrated anti-inflammatory activity in LPS-stimulated zebrafish macrophages, likely due to the stimulation of farnesoid receptor expression. Although associations between these metabolites and improvements in colitis parameters are consistent. Mechanistic studies are needed to determine real causality.
In summary, the anti-colitic effects of flavonoid-rich diets were dependent on the microbial composition of the dysbiotic donor. Anti-colitic effects were evident in recipients with higher concentrations of propionate and GCA, possibly as a product of the microbial composition and the food matrix. Reduction of metabolic products like short-chain fatty acids is common in IBD patients.54 Furthermore, human studies investigating the supplementation of polyphenol-rich extracts have demonstrated significant inter-individual variability in gut metabolic responses, suggesting gut microbiota as a potential source of variability.55 Therefore, it is feasible that even in dysbiosis, microbiota composition can alter the metabolic outcomes and health effects of bioactive compounds.
One limitation of our study is the relatively small sample size per donor, which may limit the generalizability of our findings. A larger cohort is essential to accurately capture bacterial dynamics and uncover more robust trends. Despite this limitation, our study highlights the substantial impact of dysbiotic microbiota composition on the activity of flavonoids. Future investigations should include disease severity and the physiological stages of patients as key factors, as these elements contribute to the complexity of the disease. Additionally, incorporating network analysis with a larger dataset, as well as metagenomic analyses, will contribute to the identification of microbiota key networks responsible for flavonoid-associated health impacts.
Conclusions
Our study demonstrates that the composition of dysbiotic human-associated microbiota significantly influences the anti-colitic activity of 3-deoxyflavonoids (3-DFs) and 3-hydroxyflavonoids (3-HFs). When protective effects were observed, 3-HF–containing diets reduced inflammation and enhanced barrier integrity. These outcomes were accompanied by higher concentrations of gut-derived metabolites.Author contributions
Conceptualization, L. R.; validation, J. H.-R. and L. R.; formal analysis, J. H.-R. and L. R.; investigation, J. H.-R., L. R., J. K. R. and B. H.; data curation, J. H.-R.; writing – original draft preparation, J. H.-R. and L. R.; writing – review and editing, J. H.-R., J. K. R., B. H. and L. R.; visualization, J. H.-R. and L. R.; supervision, L. R.; funding acquisition, L. R. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Sequence data generated in this study are publicly accessible and have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject ID PRJNA1185774 and can be found at https://www.ncbi.nlm.nih.gov/.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5fo03053f.
Acknowledgements
The authors gratefully acknowledge PRONABEC, Peruvian National Scholarship and Education Credit Program, for providing support to J. H.-R. during his Ph.D. studies.References
- S. E. Winter and A. J. Bäumler, Gut dysbiosis: Ecological causes and causative effects on human disease, Proc. Natl. Acad. Sci. U. S. A., 2023, 120(50), e2316579120 CrossRef . Available from: https://www.pnas.org/doi/full/10.1073/pnas.2316579120 [cited 2024 Nov 11].
- H. Brüssow, Problems with the concept of gut microbiota dysbiosis, Microb. Biotechnol., 2020, 13(2), 423–434 CrossRef . Available from: https://onlinelibrary.wiley.com/doi/abs/10.1111/1751-7915.13479 [cited 2024 Nov 20].
- T. Hrncir, Gut microbiota dysbiosis: triggers, consequences, diagnostic and therapeutic options, Microorganisms, 2022, 10(3), 578 CrossRef . Available from: https://www.mdpi.com/2076-2607/10/3/578 [cited 2024 Nov 20].
- W. Zeng, D. He, Y. Xing, J. Liu, N. Su and C. Zhang, et al., Internal connections between dietary intake and gut microbiota homeostasis in disease progression of ulcerative colitis: a review, Food Sci. Hum. Wellness, 2021, 10(2), 119–130 CrossRef . Available from: https://www.sciencedirect.com/science/article/pii/S2213453021000252 [cited 2024 Nov 8].
- J. C. Xue, S. Yuan, H. Meng, X. T. Hou, J. Li and H. M. Zhang, et al., The role and mechanism of flavonoid herbal natural products in ulcerative colitis, Biomed. Pharmacother., 2023, 158, 114086 CrossRef . Available from: https://www.sciencedirect.com/science/article/pii/S0753332222014755 [cited 2023 May 5].
- B. Wu, R. Bhatnagar, V. V. Indukuri, S. Chopra, K. March and N. Cordero, et al., Intestinal mucosal barrier function restoration in mice by maize diet containing enriched flavan-4-ols, Nutrients, 2020, 12(4), 896 CrossRef . Available from: https://www.mdpi.com/2072-6643/12/4/896 [cited 2023 May 5].
- B. Wu, A. D. Cox, H. Chang, M. Kennett, C. Rosa and S. Chopra, et al., Maize near-isogenic lines with enhanced flavonoids alleviated dextran sodium sulfate-induced murine colitis via modulation of the gut microbiota, Food Funct., 2023, 14(21), 9606–9616 RSC . Available from: https://pubs.rsc.org/en/content/articlelanding/2023/fo/d3fo02953k [cited 2023 Nov 16].
- S. Li, T. Wang, B. Wu, W. Fu, B. Xu and R. R. Pamuru, et al., Anthocyanin-containing purple potatoes ameliorate DSS-induced colitis in mice, J. Nutr. Biochem., 2021, 93, 108616 CrossRef . Available from: https://www.sciencedirect.com/science/article/pii/S095528632100036X [cited 2024 May 29].
- S. Kamiloglu, M. Tomas, T. Ozdal and E. Capanoglu, Effect of food matrix on the content and bioavailability of flavonoids, Trends Food Sci. Technol., 2021, 117, 15–33 CrossRef . Available from: https://www.sciencedirect.com/science/article/pii/S0924224420306531 [cited 2024 Sept 28].
- J. Zhang, H. Wang, C. Ai, R. Lu, L. Chen and J. Xiao, et al., Food matrix-flavonoid interactions and their effect on bioavailability, Crit. Rev. Food Sci. Nutr., 2023, 1–22, DOI:10.1080/10408398.2023.2232880 , [cited 2024 Sept 27].
- D. Hervert-Hernández and I. Goñi, Dietary polyphenols and human gut microbiota: a review, Food Rev. Int., 2011, 27(2), 154–169, DOI:10.1080/87559129.2010.535233 , [cited 2024 Sept 28].
- R. Lundberg, Humanizing the gut microbiota of mice: Opportunities and challenges, Lab. Anim., 2019, 53(3), 244–251, DOI:10.1177/0023677218787554 , [cited 2024 Sept 2].
- H. Nagao-Kitamoto, A. B. Shreiner, M. G. Gillilland, S. Kitamoto, C. Ishii and A. Hirayama, et al., Functional characterization of inflammatory bowel disease–associated gut dysbiosis in gnotobiotic mice, Cell. Mol. Gastroenterol. Hepatol., 2016, 2(4), 468–481 CrossRef . Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5042563/ [cited 2022 Sept 11].
- A. Eng and E. Borenstein, Taxa-function robustness in microbial communities, Microbiome, 2018, 6(1), 45, DOI:10.1186/s40168-018-0425-4 , [cited 2024 Nov 2].
- K. M. Ng, A. Aranda-Díaz, C. Tropini, M. R. Frankel, W. V. Treuren and C. T. O'Loughlin, et al., Recovery of the gut microbiota after antibiotics depends on host diet, community context, and environmental reservoirs, Cell Host Microbe, 2019, 26(5), 650–665 CrossRef . Available from: https://www.cell.com/cell-host-microbe/abstract/S1931-3128(19)30535-9 [cited 2024 Nov 3].
- R. González, I. Ballester, R. López-Posadas, M. D. Suárez, A. Zarzuelo and O. Martínez-Augustin, et al., Effects of Flavonoids and other Polyphenols on Inflammation, Crit. Rev. Food Sci. Nutr., 2011, 51(4), 331–362, DOI:10.1080/10408390903584094 , [cited 2024 Sept 28].
- E. Glocker, D. Kotlarz, C. Klein, N. Shah and B. Grimbacher, IL–10 and IL–10 receptor defects in humans, Ann. N. Y. Acad. Sci., 2011, 1246(1), 102–107 CrossRef PubMed . Available from: https://nyaspubs.onlinelibrary.wiley.com/doi/10.1111/j.1749-6632.2011.06339.x [cited 2025 Sept 20].
- I. A. Sheikh, J. Bianchi-Smak, D. Laubitz, G. Schiro, M. T. Midura-Kiela and D. G. Besselsen, et al., Transplant of microbiota from Crohn's disease patients to germ-free mice results in colitis, Gut Microbes, 2024, 16(1), 2333483, DOI:10.1080/19490976.2024.2333483 , [cited 2025 Sept 20].
- B. Wu, H. Chang, R. Marini, S. Chopra and L. Reddivari, Characterization of maize near-isogenic lines with enhanced flavonoid expression to be used as tools in diet-health complexity, Front. Plant Sci., 2021, 11, 1–13 Search PubMed . Available from: https://www.frontiersin.org/articles/10.3389/fpls.2020.619598 [cited 2023 May 5].
- Y. Liu, I. Fernandes, N. Mateus, H. Oliveira and F. Han, The Role of Anthocyanins in Alleviating Intestinal Diseases: A Mini Review, J. Agric. Food Chem., 2024, 72(11), 5491–5502, DOI:10.1021/acs.jafc.3c07741 , [cited 2025 Sept 28].
- K. J. Kim, Y. Kim, S. G. Jin and J. Y. Kim, Acai berry extract as a regulator of intestinal inflammation pathways in a Caco-2 and RAW 264.7 co-culture model, J. Food Biochem., 2021, 45(8), e13848 CrossRef CAS PubMed . Available from: https://onlinelibrary.wiley.com/doi/abs/10.1111/jfbc.13848 [cited 2024 May 29].
- J. Ruszkowski, Z. Kachlik, M. Walaszek, D. Storman, K. Podkowa and P. Garbarczuk, et al., Fecal microbiota transplantation from patients into animals to establish human microbiota-associated animal models: a scoping review, J. Transl. Med., 2025, 23(1), 662, DOI:10.1186/s12967-025-06645-6 , [cited 2025 Sept 28].
- M. K. Joo, J. W. Lee, J. H. Woo, H. J. Kim, D. H. Kim and J. H. Choi, Regulation of colonic neuropeptide Y expression by the gut microbiome in patients with ulcerative colitis and its association with anxiety- and depression-like behavior in mice, Gut Microbes, 2024, 16(1), 2319844 CrossRef . Available from: https://www.tandfonline.com/doi/full/10.1080/19490976.2024.2319844 [cited 2025 Sept 28].
- L. A. Knudsen, L. S. Zachariassen, M. L. Strube, J. F. Havelund, B. Pilecki and A. B. Nexoe, et al., Assessment of the Inflammatory Effects of Gut Microbiota from Human Twins Discordant for Ulcerative Colitis on Germ-free Mice, Comp. Med., 2024, 74(2), 55–69 CAS . Available from: https://aalas.kglmeridian.com/view/journals/72010023/74/2/article-p55.xml [cited 2025 Sept 28].
- J. Hou, M. Hu, L. Zhang, Y. Gao, L. Ma and Q. Xu, Dietary taxifolin protects against dextran sulfate sodium-induced colitis via NF-κB signaling, enhancing intestinal barrier and modulating gut microbiota, Front. Immunol., 2021, 11, 631809 CrossRef PubMed . Available from: https://www.frontiersin.org/articles/10.3389/fimmu.2020.631809 [cited 2022 July 28].
- R. M. Golonka, B. S. Yeoh, P. Saha, Y. Tian, J. Y. L. Chiang and A. D. Patterson, et al., Sex dimorphic effects of bile acid metabolism in liver cancer in mice, Cell. Mol. Gastroenterol. Hepatol., 2024, 17(5), 719–735 CrossRef CAS . Available from: https://www.sciencedirect.com/science/article/pii/S2352345X24000110.
- J. Haro-Reyes, J. K. Raghupathi and L. Reddivari, Composition of human-associated gut microbiota determines 3-DF and 3-HF anti-colitic activity in IL-10 -/- mice, Nutrients, 2024, 16(23), 4232 CrossRef CAS . Available from: https://www.mdpi.com/2072-6643/16/23/4232 [cited 2024 Dec 9].
- P. T. Santana, S. L. B. Rosas, B. E. Ribeiro, Y. Marinho and H. S. P. de Souza, Dysbiosis in inflammatory bowel disease: pathogenic role and potential therapeutic targets, Int. J. Mol. Sci., 2022, 23(7), 3464 CrossRef CAS PubMed . Available from: https://www.mdpi.com/1422-0067/23/7/3464 [cited 2022 Oct 24].
- G. J. Britton, E. J. Contijoch, I. Mogno, O. H. Vennaro, S. R. Llewellyn and R. Ng, et al., Microbiotas from humans with inflammatory bowel disease alter the balance of gut Th17 and RORγt+ regulatory t cells and exacerbate colitis in mice, Immunity, 2019, 50(1), 212–224 CrossRef CAS . Available from: https://www.sciencedirect.com/science/article/pii/S1074761318305636 [cited 2025 Jan 6].
- Y. Zhang, X. Si, L. Yang, H. Wang, Y. Sun and N. Liu, Association between intestinal microbiota and inflammatory bowel disease, Anim. Models Exp. Med., 2022, 5(4), 311–322 CrossRef . Available from: https://onlinelibrary.wiley.com/doi/abs/10.1002/ame2.12255 [cited 2022 Oct 24].
- X. C. Morgan, T. L. Tickle, H. Sokol, D. Gevers, K. L. Devaney and D. V. Ward, et al., Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment, Genome Biol., 2012, 13(9), R79, DOI:10.1186/gb-2012-13-9-r79 , [cited 2024 Dec 27].
- N. Iyer and S. C. Corr, Gut microbial metabolite-mediated regulation of the intestinal barrier in the pathogenesis of inflammatory bowel disease, Nutrients, 2021, 13(12), 4259 CrossRef CAS . Available from: https://www.mdpi.com/2072-6643/13/12/4259 [cited 2024 Dec 29].
- Z. Zhang, H. Zhang, T. Chen, L. Shi, D. Wang and D. Tang, Regulatory role of short-chain fatty acids in inflammatory bowel disease, Cell Commun. Signaling, 2022, 20(1), 64, DOI:10.1186/s12964-022-00869-5 , [cited 2024 Aug 28].
- T. Liu, L. Zhang, D. Joo and S. C. Sun, NF-κB signaling in inflammation, Signal Transduction Targeted Ther., 2017, 2(1), 1–9 Search PubMed . Available from: https://www.nature.com/articles/sigtrans201723 [cited 2023 May 31].
- L. Wang, M. Gao, G. Kang and H. Huang, The Potential Role of Phytonutrients Flavonoids Influencing Gut Microbiota in the Prophylaxis and Treatment of Inflammatory Bowel Disease, Front. Nutr., 2021, 8, 798038 CrossRef . Available from: https://www.frontiersin.org/articles/10.3389/fnut.2021.798038 [cited 2023 May 16].
- A. Faria, I. Fernandes, S. Norberto, N. Mateus and C. Calhau, Interplay between anthocyanins and gut microbiota, J. Agric. Food Chem., 2014, 62(29), 6898–6902, DOI:10.1021/jf501808a , [cited 2025 Jan 11].
- C. F. Ortega-Cava, S. Ishihara, M. A. K. Rumi, M. M. Aziz, H. Kazumori and T. Yuki, et al., Epithelial toll-like receptor 5 is constitutively localized in the mouse cecum and exhibits distinctive down-regulation during experimental colitis, Clin. Vaccine Immunol., 2006, 13(1), 132–138 CrossRef CAS . Available from: https://journals.asm.org/doi/full/10.1128/cvi.13.1.132-138.2006 [cited 2024 Aug 31].
- S. Feng, C. Zhang, S. Chen, R. He, G. Chao and S. Zhang, TLR5 Signaling in the Regulation of Intestinal Mucosal Immunity, J. Inflammation Res., 2023, 16, 2491–2501 CrossRef CAS . Available from: https://www.tandfonline.com/doi/abs/10.2147/JIR.S407521 [cited 2024 Nov 24].
- O. Zakerska-Banaszak, H. Tomczak, M. Gabryel, A. Baturo, L. Wolko and M. Michalak, et al., Dysbiosis of gut microbiota in Polish patients with ulcerative colitis: a pilot study, Sci. Rep., 2021, 11(1), 2166 CrossRef CAS . Available from: https://www.nature.com/articles/s41598-021-81628-3 [cited 2024 Nov 26].
- A. Lo Presti, F. Zorzi, F. Del Chierico, A. Altomare, S. Cocca and A. Avola, et al., Fecal and mucosal microbiota profiling in irritable bowel syndrome and inflammatory bowel disease, Front. Microbiol., 2019, 10(1655), 1655 CrossRef PubMed . Available from: https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2019.01655/full [cited 2024 May 9].
- S. Raimondi, E. Musmeci, F. Candeliere, A. Amaretti and M. Rossi, Identification of mucin degraders of the human gut microbiota, Sci. Rep., 2021, 11(1), 11094 CrossRef CAS . Available from: https://www.nature.com/articles/s41598-021-90553-4 [cited 2024 Sept 5].
- A. Teofani, I. Marafini, F. Laudisi, D. Pietrucci, S. Salvatori and V. Unida, et al., Intestinal Taxa Abundance and Diversity in Inflammatory Bowel Disease Patients: An Analysis including Covariates and Confounders, Nutrients, 2022, 14(2), 260 CrossRef CAS . Available from: https://www.mdpi.com/2072-6643/14/2/260 [cited 2025 Sept 29].
- Y. Fujiki, T. Tanaka, K. Yakabe, N. Seki, M. Akiyama and K. Uchida, et al., Hydrogen gas and the gut microbiota are potential biomarkers for the development of experimental colitis in mice, Gut Microbiome, 2024, 5, e3 CrossRef PubMed . Available from: https://www.cambridge.org/core/journals/gut-microbiome/article/hydrogen-gas-and-the-gut-microbiota-are-potential-biomarkers-for-the-development-of-experimental-colitis-in-mice/7D09C5192A79D779F367C332797CFFDA [cited 2024 Dec 27].
- L. Ge, J. Qi, B. Shao, Z. Ruan, Y. Ren and S. Sui, et al., Microbial hydrogen economy alleviates colitis by reprogramming colonocyte metabolism and reinforcing intestinal barrier, Gut Microbes, 2022, 14(1), 2013764 CrossRef PubMed . Available from: https://www.tandfonline.com/doi/full/10.1080/19490976.2021.2013764 [cited 2025 Sept 29].
- D. Taladrid, I. Zorraquín-Peña, N. Molinero, M. Silva, N. Manceñido and R. Pajares, et al., Polyphenols and Ulcerative Colitis: An Exploratory Study of the Effects of Red Wine Consumption on Gut and Oral Microbiome in Active-Phase Patients, Mol. Nutr. Food Res., 2022, 66(21), 2101073 CrossRef CAS . Available from: https://onlinelibrary.wiley.com/doi/abs/10.1002/mnfr.202101073 [cited 2024 Nov 27].
- G. R. Healey, R. Murphy, L. Brough, C. A. Butts and J. Coad, Interindividual variability in gut microbiota and host response to dietary interventions, Nutr. Rev., 2017, 75(12), 1059–1080, DOI:10.1093/nutrit/nux062 , [cited 2025 Feb 28].
- C. W. Ha, Y. Y. Lam and A. J. Holmes, Mechanistic links between gut microbial community dynamics, microbial functions and metabolic health, World J. Gastroenterol., 2014, 20(44), 16498–16517 CrossRef CAS . Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4248193/ [cited 2024 Dec 2].
- C. Rodríguez, E. Romero, L. Garrido-Sanchez, G. Alcaín-Martínez, R. J. Andrade and B. Taminiau, et al., Microbiota insights in Clostridium difficile infection and inflammatory bowel disease, Gut Microbes, 2020, 12(1), 1725220, DOI:10.1080/19490976.2020.1725220 , [cited 2024 Aug 25].
- Y. Geng, Y. Yue, Q. Guan, Y. Ren, L. Guo and Y. Fan, et al., Cereal Vinegar Sediment Alleviates Spontaneous Ulcerative Colitis in Il-10 Deficient Mice, Mol. Nutr. Food Res., 2021, 65(24), 2001227 CrossRef CAS PubMed . Available from: https://onlinelibrary.wiley.com/doi/abs/10.1002/mnfr.202001227 [cited 2024 Dec 1].
- N. O. Kaakoush, Insights into the Role of Erysipelotrichaceae in the Human Host, Front. Cell. Infect. Microbiol., 2015, 5, 84 Search PubMed . Available from: https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2015.00084/full [cited 2024 Dec 1].
- D. M. Rodriguez, K. J. Hintze, G. Rompato, A. J. V. Wettere, R. E. Ward and S. Phatak, et al., Dietary supplementation with black raspberries altered the gut microbiome composition in a mouse model of colitis-associated colorectal cancer, although with differing effects for a healthy versus a western basal diet, Nutrients, 2022, 14(24), 5270 CrossRef PubMed . Available from: https://www.mdpi.com/2072-6643/14/24/5270 [cited 2024 Aug 29].
- L. C. Tong, Y. Wang, Z. B. Wang, W. Y. Liu, S. Sun and L. Li, et al., Propionate ameliorates dextran sodium sulfate-induced colitis by improving intestinal barrier function and reducing inflammation and oxidative stress, Front. Pharmacol., 2016, 7, 253 Search PubMed . Available from: https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2016.00253/full [cited 2024 Dec 2].
- Y. Shi, D. Xin, H. Zhang, S. Wang, M. Yang and C. Liu, Propionate alleviates ulcerative colitis by modulating the PI3K/AKT signaling pathway and suppressing NLRP3 inflammasome activation, Food Funct., 2025, 16(17), 6925–6938 RSC . Available from: https://xlink.rsc.org/?DOI:=D5FO00772K [cited 2025 Sept 14].
- G. Lo Sasso, L. Khachatryan, A. Kondylis, J. N. D. Battey, N. Sierro and N. A. Danilova, et al., Inflammatory Bowel Disease-Associated Changes in the Gut: Focus on Kazan Patients, Inflammatory Bowel Dis., 2021, 27(3), 418–433 CrossRef PubMed.
- R. P. Feliciano, C. E. Mills, G. Istas, C. Heiss and A. Rodriguez-Mateos, Absorption, Metabolism and Excretion of Cranberry (Poly)phenols in Humans: A Dose Response Study and Assessment of Inter-Individual Variability, Nutrients, 2017, 9(3), 268 CrossRef PubMed . Available from: https://www.mdpi.com/2072-6643/9/3/268 [cited 2024 Nov 27].
| This journal is © The Royal Society of Chemistry 2025 |