Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Antiviral activity of navy bean (Phaseolus vulgaris) extract against influenza A virus via haemagglutinin interaction and interferon pathway modulation
Ji-Yeong
Choi†
a,
Se-Young
Cho†
 bc,
Bipin
Vaidya†
bc,
Iksoon
Kang
d,
Chyer
Kim
e,
Dal Sik
Kim
f and
Duwoon
Kim
*abcg
bc,
Bipin
Vaidya†
bc,
Iksoon
Kang
d,
Chyer
Kim
e,
Dal Sik
Kim
f and
Duwoon
Kim
*abcg
aDepartment of Integrative Food, Bioscience and Biotechnology, Chonnam National University, Gwangju 61186, Republic of Korea. E-mail: dwkim@jnu.ac.kr
bDepartment of Food Science and Technology, Chonnam National University, Gwangju 61186, Republic of Korea
cFoodborne Virus Research Center, Chonnam National University, Gwangju 61186, Republic of Korea
dDepartment of Animal Science, California Polytechnic State University, San Luis Obispo, CA 93407, USA
eAgricultural Research Station, Virginia State University, Petersburg, VA 23806, USA
fDepartment of Laboratory Medicine, Green Cross Laboratories, Yongin, Gyeonggi 16926, Republic of Korea
gAgriBio Disaster Research Center, Chonnam National University, Gwangju 61186, Republic of Korea
First published on 7th October 2025
Abstract
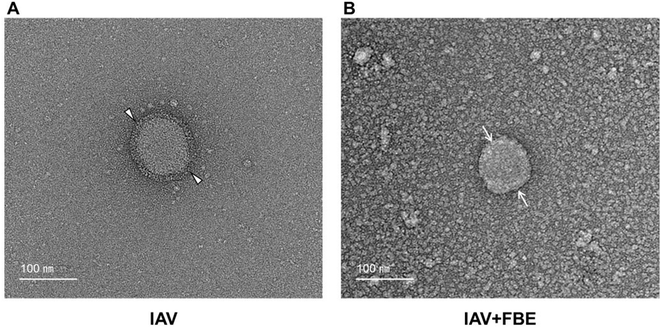
Phaseolus vulgaris agglutinins (PHAs), lectins derived from navy beans, are recognised for their limited antiviral effects and toxicity in Madin–Darby canine kidney (MDCK) cells. In this study, fermented navy bean extract (FBE) produced with Bacillus subtilis was analysed using ultra-performance liquid chromatography coupled with tandem mass spectrometry, identifying various lectins, including monomeric forms of PHA-L and PHA-E (∼32 kDa). Pre-treatment with FBE exhibited 2.6- and 12.6-fold greater inhibition of influenza A virus (IAV) H1N1 replication than commercially available PHA-E and PHA-L (12.5 μg mL−1), respectively. Additionally, a synthesised 10-amino acid peptide derived from PHA-L (antifungal lectin, AFL) exhibited antiviral properties, reducing the viral load by 1.95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log under pre-treatment conditions. This effect was attributed to AFL-induced type I interferon responses, which led to the upregulation of key antiviral genes (IFN-α, IFN-β, STAT1, and STAT2). Co-treatment with FBE and IAV H1N1 effectively inhibited viral–host interaction and entry. To further investigate the underlying mechanism, FBE was neutralised with haemagglutinin (HA)–His proteins, and immunoprecipitation–mass spectrometry analysis identified a 36 kDa protein (UniProt accession number: V7BGE3), previously annotated but newly implicated as an HA-binding protein. Transmission electron microscopy revealed that FBE binding to HA glycoproteins induced structural alterations on the outer surface of IAV H1N1 particles, potentially blocking viral entry. These findings highlight the multifunctional antiviral mechanisms of FBE and its potential as a novel therapeutic agent against IAV, warranting further exploration.
log under pre-treatment conditions. This effect was attributed to AFL-induced type I interferon responses, which led to the upregulation of key antiviral genes (IFN-α, IFN-β, STAT1, and STAT2). Co-treatment with FBE and IAV H1N1 effectively inhibited viral–host interaction and entry. To further investigate the underlying mechanism, FBE was neutralised with haemagglutinin (HA)–His proteins, and immunoprecipitation–mass spectrometry analysis identified a 36 kDa protein (UniProt accession number: V7BGE3), previously annotated but newly implicated as an HA-binding protein. Transmission electron microscopy revealed that FBE binding to HA glycoproteins induced structural alterations on the outer surface of IAV H1N1 particles, potentially blocking viral entry. These findings highlight the multifunctional antiviral mechanisms of FBE and its potential as a novel therapeutic agent against IAV, warranting further exploration.
Introduction
Influenza A viruses (IAVs) present a persistent global health challenge, causing seasonal epidemics and occasional pandemics. Despite efforts to mitigate IAV infections through vaccines and antiviral drugs, the viruses continue to impose a significant burden owing to their rapid mutation rate and development of drug resistance.1 IAV is an enveloped RNA virus characterised by surface glycoproteins, such as neuraminidase (NA) and haemagglutinin (HA), which facilitate viral entry by binding to receptors on host cells.2 HA, the major antigen of IAV, attaches to sialic acid receptors on host cells, initiating infection and prompting the step of neutralising antibody production.3,4 However, frequent mutations in HA lead to the emergence of novel IAV strains, undermining the efficacy of existing vaccines.5,6 Current antiviral strategies targeting the M2 ion channel and NA have been limited by the emergence of resistant IAV strains. Additionally, viral RNA polymerase inhibitors (e.g., favipiravir) exhibit variable therapeutic outcomes. These limitations underscore the urgent need for alternative antiviral therapies that address the challenges of IAV infections.7Lectins, carbohydrate-binding proteins of nonimmune origin, have shown potential as antiviral agents against IAV owing to their selective binding capabilities. This selectivity offers distinct advantages over current therapies utilising antibodies and enzymes.8,9 Plant-derived lectins, in particular, play an essential role in antiviral defence by binding to viral carbohydrates, effectively disrupting interactions between viral HA and host-cell sialic acid receptors.10 Lectins with affinities for mannose, glucose, and N-acetylglucosamine (GlcNAc) have demonstrated inhibitory effects against IAV.11,12 Several plant-derived lectins, including cyanovirin-N, hyacinth bean lectin, Hippeastrum hybrid agglutinin, and Galanthus nivalis agglutinin, act as IAV inhibitors by interacting with HA, thereby hindering virus entry and release.13,14 Engineered banana lectin has also demonstrated efficacy in protecting against IAV by blocking viral fusion with the endosome.15 However, the antiviral activity of lectins from certain plant sources, such as navy beans, remains underexplored.
Among bean-derived lectins, Phaseolus vulgaris agglutinin (PHA), a prominent component of navy beans, distinguishes itself owing to its well-defined structure and specific carbohydrate-binding domains that facilitate interactions with viral surface glycoproteins.16 The structural and binding properties of PHA suggest potential applications in disrupting virus–host cell interactions, a mechanism observed in other antiviral lectins.17 Although the carbohydrate-binding properties of PHA are well-documented, its antiviral potential, particularly against IAV H1N1, remains unexplored. Previously, our research team identified a novel lectin, Tcan, by fermenting sword beans (Canavalia gladiata) with Bacillus subtilis. This lectin demonstrated significant antiviral efficacy against SARS-CoV-2, highlighting the potential of fermentation-derived lectins in combating viral infections.18 Leveraging this success, we applied a similar fermentation method to navy beans using B. subtilis, hypothesising that the approach would exhibit anti-IAV H1N1 potential in vitro by mitigating cytotoxicity.
This study aimed to investigate the antiviral properties of fermented navy bean extract (FBE), identify the active antiviral components of FBE, and evaluate the efficacy of these components in inhibiting IAV H1N1. It focused on viral cell attachment and the modulation of immune-related gene expression in Madin–Darby canine kidney (MDCK) cells.
Materials and methods
Cell culture, virus propagation, and antifungal lectin (AFL) preparation
The MDCK cell line (KCLB #10034), sourced from the Korean Cell Line Bank, was cultured in Dulbecco's modified Eagle's medium (DMEM; Gibco, Thermo Fisher Scientific, Inc., Waltham, Massachusetts, USA) supplemented with 1% streptomycin–penicillin and 10% foetal bovine serum (FBS; Gibco, Thermo Fisher Scientific, Inc., Waltham, Massachusetts, USA). The cells were incubated at 37 °C in a 5%-CO2 incubator (VS-2180C; Vision Scientific, Daejeon, South Korea). For virus propagation, a low-pathogenic human IAV H1N1 strain A/PR/8/34 (ATCC VR-1469; American Type Culture Collection, Manassas, VA, USA) was cultured in MDCK cells using serum-free DMEM supplemented with 2 μg mL−1 of N-p-tosyl-L-phenylalanine chloromethyl ketone-treated trypsin (Sigma-Aldrich, Burlington, Massachusetts, USA) and 1% streptomycin–penicillin under 35 °C and 5% CO2 conditions (TPCK-treated DMEM0 + 1% S/P). To validate the sequence-specific antiviral activity of the synthetic peptide, a 10-amino acid peptide (SNDIYFNFQR), designated AFL, was synthesised based on the lectin domain of the Phaseolus vulgaris agglutinin (PHA) sequence. A scrambled version of this peptide, SNDIYFSFDR (denoted as AFLm), was also synthesised by Dandicure Co., Ltd (Cheongju, South Korea), where two residues were substituted to serve as a negative control for antiviral activity. PHA leucoagglutinin (PHA-L) and erythroagglutinin (PHA-E) were purchased from Sigma-Aldrich.FBE preparation
To prepare FBE, dried navy beans (P. vulgaris) were initially ground into a fine powder. The powdered beans were subsequently fermented under controlled conditions using B. subtilis at 37 °C for 4 days under constant agitation at 150 rpm. During fermentation, the dissolved oxygen level was maintained at 20–40% saturation, and the pH was kept at 7.0. After fermentation, the mixture was subjected to ultrafiltration to isolate the fraction containing components within the 10–100 kDa molecular-weight range. This fraction was then freeze-dried to produce a stable product, which was subsequently stored at −80 °C until further analysis.Cytotoxicity assay
MDCK cells were seeded at a density of 2 × 104 cells per well in a 96-well plate, with each well containing 100 μL of DMEM supplemented with 10% FBS (DMEM10). The cells were incubated for 24 h at 37 °C and 5% CO2. After incubation, the culture medium was carefully removed, and the cells were exposed to 100 μL of varying concentrations (25–400 μg mL−1) of FBE and other compounds prepared in DMEM10. The cells were incubated for an additional 48 h under similar conditions (37 °C, 5% CO2). For the negative control, cells were treated with DMEM10 without FBE or other compounds. Following treatment, 10 μL of Cell-Counting Kit-8 solution (CCK-8; Dojindo Molecular Technologies, Rockville, MD, USA) was added to each well and the plate incubated for 2 h. Absorbance was measured at 450 nm using a microplate spectrophotometer (Synergy™; BioTek, VT, USA). Cytotoxicity was determined by comparing the absorbance of the treated wells with that of the control wells. The concentration of FBE and other compounds resulting in a 50% reduction in cell viability (CC50) was determined.Anti-IAV assay in MDCK cells
The antiviral activity of FBE, the synthetic peptide AFL, and PHAs was evaluated in MDCK cells infected with influenza A virus (H1N1) using three treatment strategies: pre-treatment, co-treatment, and post-treatment. MDCK cells were seeded at a density of 2 × 104 cells per well in a 96-well plate, with 100 μL of medium added to each well, and incubated for 24 h at 37 °C in a 5%-CO2 incubator. Following incubation, the cultured cells were washed twice with phosphate-buffered saline (PBS) to remove any residual medium. In the pre-treatment assay, the cells were exposed to varying concentrations of FBE (0–100 μg mL−1), AFL (0–100 μg mL−1), PHA-E (0–12.5 μg mL−1), and PHA-L (0–12.5 μg mL−1) for 12 h at 37 °C in a 5% CO2 atmosphere, followed by two washes with PBS to remove unbound compounds. The cells were subsequently infected with H1N1 and incubated at 35 °C for 72 h in 5% CO2. In the co-treatment assay, equal volumes of the virus dilution and test compound were mixed and incubated at 32 °C for 2 h to facilitate direct interaction between viral particles and the treatment agents. Following incubation, the virus–compound mixture was directly applied to MDCK cells and incubated at 35 °C in 5% CO2 for 72 h. This approach aimed to determine whether the compounds could directly bind to and neutralise viral particles before their entry into host cells. In the post-treatment assay, MDCK cells were initially infected with H1N1 at 35 °C in 5% CO2 for 2 h. After removing unbound virus by washing with PBS, the cells were treated with the test compounds for an additional 12 h at 35 °C in 5% CO2. Following another PBS wash, the medium was replaced with TPCK-treated DMEM0 supplemented with 1% S/P and, the cells were incubated for 72 h. Infection with IAV H1N1 was performed at a multiplicity of infection of 10. After the treatment period, culture supernatants were harvested, and viral RNA was subsequently extracted and quantified using the quantitative reverse transcription-polymerase chain reaction (qRT-PCR), as detailed below.IAV H1N1 replication via qRT-PCR assays
To quantify viral copies in cells using qRT-PCR, cell supernatants were collected at the endpoint of infection. RNA was extracted from the collected supernatant using RNAiso Plus (Takara Bio, Inc., Shiga, Japan), according to the manufacturer's protocol. In brief, the extraction process involved chloroform addition for phase separation, RNA precipitation with isopropanol, centrifugation, 75% ethanol wash, and final resuspension in ribonuclease (RNase)-free water. The extracted RNA was quantified using a NanoVue™ Plus Spectrophotometer (GE Healthcare, Little Chalfont, Buckinghamshire, UK). For complementary DNA (cDNA) synthesis, 10 μL of quantified RNA was combined with 1 μL of a 6-mer random primer (Takara, Kyoto, Japan) and incubated at 65 °C for 10 min. The reaction mixture was subsequently supplemented with 4 μL of 5× PrimeScript® reverse transcription buffer, 2 μL of dithiothreitol, 2 μL of deoxynucleotide triphosphate, 0.5 μL of PrimeScript® reverse transcriptase, and 0.5 μL of RNase inhibitor. This mixture was incubated at 37 °C for 1 h, followed by heat inactivation at 95 °C for 5 min. cDNA was synthesised using a PCR Thermal Cycler Dice Gradient System (Takara). The synthesised cDNA was subsequently utilised to measure viral replication using IAV PA gene primers (forward: 5′-GAG CCT ATG TGG ATG GAT TC-3′ and reverse: 5′-CCC ATT CGG AAG TCT AAG TG-3′). qRT-PCR was performed with the following cycling parameters: 95 °C for 10 min initiation, 45 cycles of 95 °C for 5 s denaturation, 55 °C for 10 s annealing, and 72 °C for 20 s elongation. PA gene expression was determined by substituting the Ct values into the PA gene standard curve, which was constructed using known concentrations of the PA gene and expressed as copies per mL.Relative mRNA gene expression using the qPCR assay
MDCK cells were seeded into 96-well plates at a density of 1 × 105 cells per well in 100 μL of DMEM10. The cells were incubated at 37 °C in 5% CO2 for 24 h and subsequently washed twice with PBS. Thereafter, they were treated with varying concentrations of FBE or AFL at 37 °C for 12 h. Afterward, the cells were infected with IAV H1N1 and further incubated at 35 °C in 5% CO2 for 72 h. Following incubation, the supernatant was discarded, and the cells were lysed using 100 μL of RNAiso reagent. The lysed cells were subsequently incubated at 35 °C for 5 min to facilitate RNA extraction. RNA was extracted as described previously, and gene-specific primers (as detailed in Table S1) were used for the qPCR assay. Relative gene expression levels were calculated using the 2−ΔΔCt method, with gene expression in the lectin-treated samples normalised to that of the untreated control group.Sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE)
SDS-PAGE was performed using a 15% sodium dodecyl sulphate-polyacrylamide gel, following the manufacturer's protocol. Initially, the samples were mixed with a loading buffer containing 0.2% SDS and heated at 100 °C for 5 min to denature the proteins. The samples were subsequently loaded onto the SDS-PAGE gel and subjected to electrophoresis at a constant voltage of 100 V for 2 h. After electrophoresis, the gel was stained with Coomassie brilliant blue to visualise the protein bands. A protein molecular-weight marker (HiQ Pre-Stained Protein Marker; BioD, Korea), detecting molecular weights ranging from 11 to 75 kDa, was used to estimate the size of the lectin bands.Immunoprecipitation–mass spectrometry (IP–MS/MS)
To isolate His-tagged proteins, 10 μg of HA–His protein was diluted with 1× binding/washing buffer (50 mM sodium phosphate, pH 8.0, 300 mM NaCl, 0.01% Tween-20) and incubated with Dynabeads™ His-tag Isolation and Pulldown (Thermo Fisher Scientific, Inc. USA) for 5 min with gentle mixing. After magnetic separation and removal of the supernatant, the beads were washed four times with the same buffer. The FBE protein was subsequently diluted (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) with 2× pulldown buffer and incubated with the washed Dynabeads™ at 30 °C for 30 min. After incubation, the beads were washed with 1× binding/washing buffer to eliminate any non-specifically bound proteins. His-tagged proteins were eluted from the beads by adding His-elution buffer (300 mM imidazole, 50 mM sodium phosphate, pH 8.0, 300 mM NaCl, 0.01% Tween™-20) and collected for nano UPLC-MS/MS analysis.
1) with 2× pulldown buffer and incubated with the washed Dynabeads™ at 30 °C for 30 min. After incubation, the beads were washed with 1× binding/washing buffer to eliminate any non-specifically bound proteins. His-tagged proteins were eluted from the beads by adding His-elution buffer (300 mM imidazole, 50 mM sodium phosphate, pH 8.0, 300 mM NaCl, 0.01% Tween™-20) and collected for nano UPLC-MS/MS analysis.
Nano UPLC-MS/MS
Purified FBE and proteins eluted from the IP–MS/MS assay were processed into peptides using the tube-gel method.19 The resulting peptide mixture was analysed using a nano-ACQUITY Ultra-Performance Liquid Chromatograph™ (UPLC) equipped with a Synapt™ G2-Si High-Definition Mass Spectrometry System (Waters Corp., Milford, MA, USA), as described previously.20 Chromatographic separation was performed on an ACQUITY UPLC BEH C18 column (100 × 2.1 mm, 1.7 μm particle size; Waters Corp., Milford, MA, USA). The mobile phase comprised solvents A (0.1% formic acid in water) and B (0.1% formic acid in acetonitrile). The elution gradient was programmed as follows: 0–1 min, 5% B; 1–10 min, 5–40% B; 10–12 min, 40–85% B; 12–13 min, 85% B; 13–13.1 min, 85–5% B; and 13.1–15 min, 5% B. The flow rate was set at 0.3 mL min−1 and column temperature maintained at 40 °C. Mass spectrometry was performed in positive electrospray ionisation mode with a capillary voltage of 3.0 kV and source temperature of 120 °C. Raw data were processed using MassLynx software (version 4.1, Waters) and searched against the UniProt database (Phaseolus vulgaris; UP000000226) using Mascot (Matrix Science) for protein identification. Identified proteins were filtered using a false discovery rate of <4% at the protein level and categorised based on MS/MS spectra using the ProteinLynx Global Server algorithm. Subsequently, these proteins were compared with known lectin sequences using BLAST for further characterisation.Protein structure visualisation and analysis using PyMOL
The three-dimensional (3D) structures of AFL and AFLm, a modified variant of AFL, were obtained from the UniProt database as protein data bank files (UniProt IDs: Q8RVX9 for AFL and Q3LA43 for AFLm) and analysed using PyMOL (version 2.6; Schrödinger, LLC). Structural alignment and overlay were performed to identify conformational variations, with divergent regions highlighted using distinct colour coding. The charge distribution of amino acid residues was visualised using colour mapping, with positively charged, neutral, and negatively charged regions in red, cyan, and blue, respectively. Active-site residues were highlighted to facilitate a comparative analysis of functional differences between the two proteins. Additionally, structural measurements, such as interatomic distances and angles, were performed to evaluate potential conformational and functional implications. The resulting visualisations were exported for figure preparation and further analysis.Morphological analysis using transmission electron microscopy (TEM)
After 2 h incubation at 32 °C, morphological alterations in IAV particles, with or without FBE treatment, were observed via TEM. For sample preparation, 100 μL of each sample was applied to 200-mesh carbon-coated copper grids (Electron Microscopy Sciences, Hatfield, PA, USA). Afterward, the samples were negatively stained with 2% aqueous uranyl acetate for 30 s. Excess stain was blotted away, and the grids were rinsed three times with distilled water and air-dried. Imaging was performed on a Zeiss LEO 912AB TEM (Carl Zeiss, Oberkochen, Germany) at an accelerating voltage of 80 kV.Statistical analysis
All experiments were conducted in triplicate, and the results are presented as the mean ± standard deviation. GraphPad Prism for Windows (version 6; Graph Pad Software, Inc., San Diego, CA, USA) was used to generate all figures. The CC50 values of the MDCK cell lines were calculated using nonlinear regression analysis to fit a dose–response curve in GraphPad Prism. Statistical analysis was conducted using IBM SPSS Statistics software (version 27; IBM Corp., Armonk, NY, USA). Statistical significance was determined using one-way analysis of variance followed by Dunnett's test, with a p-value <0.01 considered statistically significant.Results and discussion
Cytotoxicities of FBE and AFL
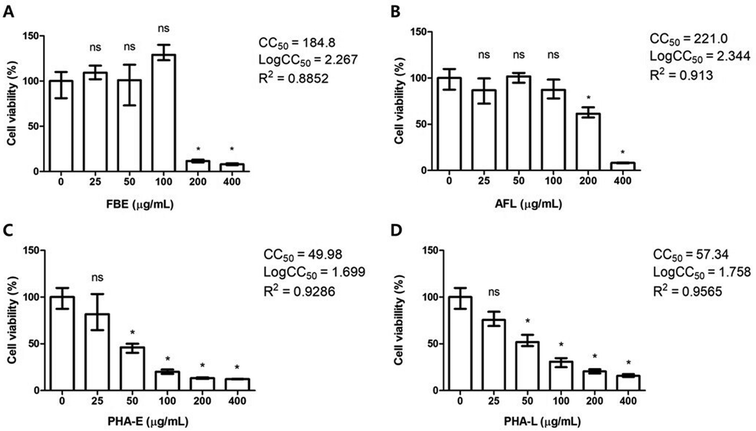
Fig. 1 displays the dose-dependent cytotoxicities of FBE, AFL, and the purchased standards (phytohaemagglutinin [PHA]-E and PHA-L) in MDCK cells. Both FBE and AFL exhibited no cytotoxicity at concentrations up to 100 μg mL−1, a promising indication of their potential therapeutic applications. However, at higher concentrations, their effects diverged: at 200 μg mL−1, FBE displayed significant cytotoxicity, reducing cell viability to 12%, whereas AFL was less cytotoxic, maintaining 61.5% cell viability at the same concentration. These findings were further supported by their CC50 values, which were determined to be 184.8 and 221.0 μg mL−1 for FBE and AFL, respectively, indicating the superior safety profile of FBE to that of AFL. In contrast, the purchased standards (PHA-E and PHA-L) demonstrated cytotoxicity effects starting at 50 μg mL−1, with CC50 values of 49.98 and 57.34 μg mL−1, respectively. These significantly lower CC50 values suggest greater cytotoxic potential, limiting their therapeutic applicability as antiviral agents. This aligns with previous studies that demonstrated their toxicity in both in vivo and in vitro settings.21,22Characterisation of lectin from fermented navy bean
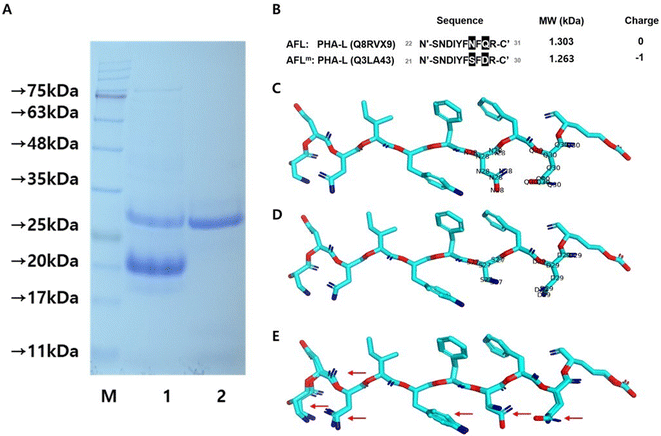
SDS-PAGE of FBE—derived from fermented navy bean powder and purified using a size cut-off ultrafiltration membrane—yielded a single prominent protein band at approximately 32 kDa (Fig. 2A). In contrast, the non-fermented navy bean water extract (NWE) exhibited three major protein bands: >100 kDa, ∼32 kDa, and <32 kDa, indicating the presence of unprocessed or non-selective proteins. The single dominant band in FBE highlights the enrichment of specific lectin proteins during fermentation and ultrafiltration. Further tandem mass spectrometry (MS/MS) analysis of FBE identified 26 lectin-related proteins, including phytohaemagglutinin variants such as leucoagglutinating phytohaemagglutinin (PHA-L) and erythroagglutinating phytohaemagglutinin (PHA-E), as well as additional lectin proteins homologous to other members of the P. vulgaris lectin family. These identified proteins, summarised in Table 1, provide insights into the diversity and specificity of lectin variants enriched during fermentation and purification.| Accession | Protein name | Gene | Score | Average mass | Matched products | Matched peptides | digestPeps | Seq cover (%) | AutoCurate |
|---|---|---|---|---|---|---|---|---|---|
| P84869 | Antifungal lectin PVAP (fragment) | — | 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 395.66 395.66 |
1303.397 | 12 | 1 | 1 | 100 | Green |
| V5YN37 | Erythroagglutinating | — | 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 874.61 874.61 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 796.564 796.564 |
205 | 13 | 18 | 55.64 | Green |
| V5QN77 | Erythroagglutinating phytohaemagglutinin | — | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 690.27 690.27 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 774.552 774.552 |
221 | 13 | 18 | 55.64 | Green |
| Q8RVX6 | Phytohaemagglutinin | pha-E | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 690.27 690.27 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 760.525 760.525 |
221 | 13 | 18 | 55.64 | Green |
| P05088 | Erythroagglutinating phytohaemagglutinin | DLEC1 | 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 874.61 874.61 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 746.504 746.504 |
215 | 13 | 18 | 55.64 | Green |
| Q8RVH2 | Phytohaemagglutinin | pha-L | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 680.03 680.03 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 331.986 331.986 |
131 | 10 | 18 | 53.48 | Green |
| Q43628 | Phytohaemagglutinin | — | 7563.183 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 480.034 480.034 |
114 | 9 | 16 | 50.36 | Green |
| Q8RVH3 | Phytohaemagglutinin | pha-E | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 069.37 069.37 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 824.612 824.612 |
176 | 12 | 17 | 48 | Green |
| I0J8I4 | PHA-E protein | Pha-E | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 069.37 069.37 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 770.48 770.48 |
176 | 12 | 18 | 48 | Green |
| P15231 | Leucoagglutinating phytohaemagglutinin | PDLEC2 | 4325.175 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 420.835 420.835 |
72 | 8 | 17 | 47.62 | Green |
| V7C654 | Legume lectin domain-containing protein | PHAVU_004G158200g | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 069.37 069.37 |
30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 262.095 262.095 |
176 | 12 | 18 | 47.31 | Green |
| Q8RVY1 | Phytohaemagglutinin | pha-E | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 743.05 743.05 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 559.216 559.216 |
128 | 11 | 15 | 45.82 | Green |
| Q8RVX5 | Lectin | lec4-B17 | 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 360.31 360.31 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 626.276 626.276 |
132 | 9 | 13 | 44.36 | Green |
| P05087 | Leucoagglutinating phytohaemagglutinin | DLEC2 | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 618.79 618.79 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 556.013 556.013 |
127 | 7 | 17 | 42.65 | Green |
| T2DPB5 | Leucoagglutinating phytohaemagglutinin | — | 4305.554 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 377.77 377.77 |
69 | 7 | 17 | 41.39 | Green |
| T2DM03 | Phytohaemagglutinin | — | 9967.701 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 858.721 858.721 |
126 | 10 | 18 | 36.73 | Green |
| Q8RVX9 | Phytohaemagglutinin | pha-L | 9087.686 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 516.1 516.1 |
93 | 8 | 19 | 36.63 | Green |
| V5N8T1 | Leucoagglutinating phytohaemagglutinin | — | 4101.986 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 583.343 583.343 |
56 | 6 | 18 | 35.53 | Green |
| V7C787 | Legume lectin domain-containing protein | PHAVU_004G158300g | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 596.11 596.11 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 556.013 556.013 |
116 | 6 | 17 | 32.72 | Green |
| V7C5Y6 | Legume lectin domain-containing protein | PHAVU_004G158000g | 9094.787 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 974.435 974.435 |
77 | 6 | 17 | 32.49 | Green |
| V7AIB2 | Chitinase | PHAVU_011G167000g | 3914.884 | 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 586.885 586.885 |
57 | 6 | 20 | 27.41 | Green |
| Q3LA43 | Phytohaemagglutinin | pha-L | 9124.707 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 446.792 446.792 |
75 | 5 | 15 | 27.21 | Green |
| Q8RVH1 | Lectin | lec3-A1 | 6904.168 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 651.001 651.001 |
79 | 5 | 14 | 25.45 | Green |
| Q8RVX4 | Arcelin | arc4-I | 919.758 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 489.963 489.963 |
27 | 3 | 20 | 10.53 | Green |
| Q43629 | Arcelin-4 | ARC4 | 919.758 | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 508.009 508.009 |
30 | 3 | 22 | 10.53 | Green |
| Q8RVY3 | Arcelin | Arc-II | 778.001 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 142.861 142.861 |
13 | 1 | 18 | 3.69 | Green |
The antifungal peptide AFL, comprising 10 amino acids (sequence: SNDIYFNFQR), corresponded to residues 22–31 in the N-terminal region of PHA-L (UniProt accession number, Q8RVX9) and shared 100% sequence identity with AFL PVAP (UniProt accession number, P84869). Identified via HPLC-MS/MS analysis of the FBE sample (Table 1), AFL was structurally modelled and visualised using PyMOL to elucidate its 3D conformation (Fig. 2B).23 To investigate the structure–function relationships, a modified variant of AFL, designated AFLm, was developed based on the PHA-L of P. vulgaris (UniProt accession number, Q3LA43). AFLm included specific substitutions (SNDIYFSFDR; Fig. 2B) that altered the net charge of the peptide from neutral (0) in AFL to −1 in AFLm. This final charge was attributed to the negatively charged aspartic acid (D) and hypothesised to influence peptide folding, stability, and interaction potential with biological targets.24,25 The differences at the 7th and 9th positions of the sequence likely altered the charge distribution and/or hydrogen bonding capabilities, potentially affecting the interactions or specificity of the peptides (Fig. 2B, C and E). For instance, at the 7th position, asparagine (N) and serine (S) exhibit similar hydrogen bonding capabilities; however, serine is slightly more hydrophilic. In AFL, asparagine can form more hydrogen bonds than serine in AFLm, potentially affecting interactions with water molecules and overall stability in aqueous solutions. At the 9th position, glutamine (Q) is neutral, while aspartic acid (D) is negatively charged. The substitution of glutamine with aspartic acid alters the net charge of the peptide, potentially affecting electrostatic interactions with binding partners. Additionally, the increased polar nature of serine and aspartic acid in AFLm may diminish its tendency to aggregate compared with that in AFL. AFLm is probably more sensitive to pH changes owing to the ability of aspartic acid to alter its protonation state.
The substitution of asparagine with serine and glutamine with aspartic acid may also alter hydrogen bond strength and electrostatic interactions with the HA protein.26 These modifications possibly affect the peptides’ ability to recognise and bind to specific regions of the HA protein, potentially altering their effectiveness as entry inhibitors.27 Additionally, these amino acid substitutions may influence the peptides’ secondary structure, compromising their ability to mimic or interact with HA's functional regions.27,28 PyMOL-based structural alignment and overlay of AFL and AFLm revealed significant differences in their 3D conformations, predominantly around the substituted residues, possibly impacting their interactional properties and stability (Fig. 2E). Sequence alignment confirmed these residue-level differences, emphasising the structural impact of the substitutions. Collectively, these findings provide a comprehensive structural and biochemical characterisation of the AFL variants. The observed structural differences between AFL and AFLm, especially the shift in net charge owing to the 9th position substitution, underscore the potential of modifying lectin sequences to explore functional diversity and improve bioactivity. Furthermore, the fermentation-based enrichment of specific lectins demonstrates the utility of processing methods in optimising bioactive protein profiles.
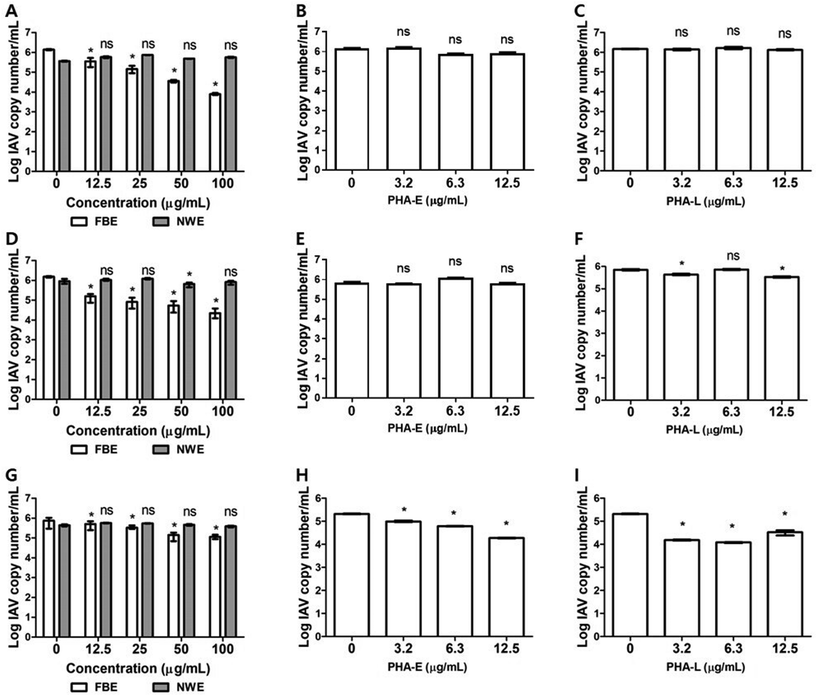
Anti-IAV H1N1 activity of pre-, co-, and post-treatment
The antiviral efficacies of FBE, AFL, AFLm, PHA-E, and PHA-L against IAV H1N1 were evaluated in MDCK cells under various treatment conditions by measuring reductions in viral (IAV H1N1) replication (Fig. 3 and 4). On comparing NWE with FBE, NWE did not exert any inhibitory effects on IAV H1N1 replication. In contrast, FBE significantly inhibited viral replication across all treatment conditions (Fig. 3A, D and G). These findings suggest that fermentation is requisite to producing bioactive compounds that inhibit viral replication.Under pre-treatment conditions, FBE inhibited viral replication in a dose-dependent manner, yielding a reduction of up to 2.24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log at 100 μg mL−1. PHA-E significantly reduced replication by <1
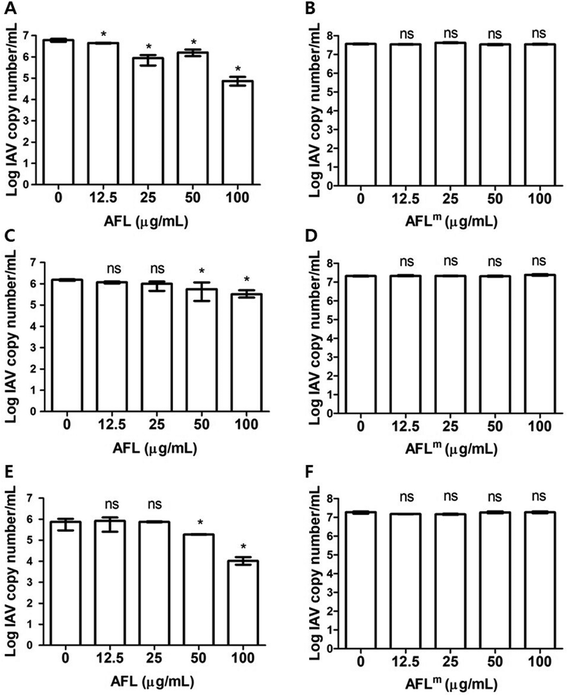
log at 100 μg mL−1. PHA-E significantly reduced replication by <1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log, whereas PHA-L exhibited no significant effect (Fig. 3A–C). Notably, treatment with AFL, a 10-amino acid peptide derived from PHA-L (Q8RVX9), inhibited viral replication by 1.95
log, whereas PHA-L exhibited no significant effect (Fig. 3A–C). Notably, treatment with AFL, a 10-amino acid peptide derived from PHA-L (Q8RVX9), inhibited viral replication by 1.95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log at 100 μg mL−1 in a dose-dependent manner, whereas its variant, AFLm, exerted no significant antiviral effects (Fig. 4A and B). Under co-treatment conditions, FBE exclusively inhibited viral replication in a dose-dependent manner, achieving a 1.89-log reduction at 100 μg mL−1 (Fig. 3D). In comparison, AFL inhibited viral replication by <1
log at 100 μg mL−1 in a dose-dependent manner, whereas its variant, AFLm, exerted no significant antiviral effects (Fig. 4A and B). Under co-treatment conditions, FBE exclusively inhibited viral replication in a dose-dependent manner, achieving a 1.89-log reduction at 100 μg mL−1 (Fig. 3D). In comparison, AFL inhibited viral replication by <1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log at the same concentration (Fig. 4C). PHA-E and PHA-L significantly inhibited IAV H1N1 replication, displaying a nonlinear dose–response pattern (Fig. 3E and F). AFLm wielded no significant antiviral effect (Fig. 4D). Under post-treatment conditions, FBE inhibited IAV H1N1 replication in a dose-dependent manner. However, its efficacy decreased compared with that under pre- or co-treatment conditions, achieving a 1-log reduction at 100 μg mL−1 (Fig. 3G). Post-treatment with PHA-E (up to 12.5 μg mL−1) reduced viral replication by 1.05
log at the same concentration (Fig. 4C). PHA-E and PHA-L significantly inhibited IAV H1N1 replication, displaying a nonlinear dose–response pattern (Fig. 3E and F). AFLm wielded no significant antiviral effect (Fig. 4D). Under post-treatment conditions, FBE inhibited IAV H1N1 replication in a dose-dependent manner. However, its efficacy decreased compared with that under pre- or co-treatment conditions, achieving a 1-log reduction at 100 μg mL−1 (Fig. 3G). Post-treatment with PHA-E (up to 12.5 μg mL−1) reduced viral replication by 1.05![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log in a dose-dependent manner. PHA-L exerted a nonlinear dose–response effect, reducing viral replication by <1
log in a dose-dependent manner. PHA-L exerted a nonlinear dose–response effect, reducing viral replication by <1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log (Fig. 3F and I). Remarkably, AFL displayed the most significant reduction in viral replication during post-treatment, yielding a 1.85-log decrease at 100 μg mL−1. Conversely, its variant, with specific amino acid modifications, did not exhibit any antiviral effects (Fig. 4E and F).
log (Fig. 3F and I). Remarkably, AFL displayed the most significant reduction in viral replication during post-treatment, yielding a 1.85-log decrease at 100 μg mL−1. Conversely, its variant, with specific amino acid modifications, did not exhibit any antiviral effects (Fig. 4E and F).
Based on the above results, we conclude that while PHA-L and PHA-E exert modest antiviral effects, their cytotoxicity limits their application as antiviral agents, unlike FBE. A previous study found PHA-L and PHA-E to induce apoptosis, cause direct cytotoxicity to various cell types, and trigger activation-induced cell death, rendering them unsuitable for general antiviral use.22 In contrast, pre-treatment with FBE and AFL yielded the most significant antiviral efficacy (2.24 and 1.95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log, respectively) at 100 μg mL−1, with FBE consistently outperforming AFL in mitigating IAV H1N1 replication under pre- and co-treatment conditions. Notably, pre- and co-treatment were more effective than post-treatment, possibly because of their ability to block viral entry.
log, respectively) at 100 μg mL−1, with FBE consistently outperforming AFL in mitigating IAV H1N1 replication under pre- and co-treatment conditions. Notably, pre- and co-treatment were more effective than post-treatment, possibly because of their ability to block viral entry.
The anti-IAV H1N1 effectiveness of FBE may differ from that of AFL owing to variations in binding affinity, viral particle aggregation, and/or immunomodulatory effects.29 FBE potentially enhances immune cell activation, facilitating early viral recognition and clearance, thus inhibiting viral replication. Furthermore, differences in carbohydrate-binding specificities potentially influence antiviral potency. For example, FBE may share glycan specificity with PHA-E by binding to GlcNAc,30 while AFL targets chitin, a polymer of GlcNAc.31 Both FBE and AFL can bind to GlcNAc but vary in their specificity depending on whether GlcNAc is free or polymerised. Numerous lectins, including FBE and AFL, demonstrate significant anti-IAV H1N1 activity, often related to their glycan-binding specificities. Mannose-binding lectins, in particular, display strong antiviral properties against IAV and other viruses. The highly glycosylated surface proteins of IAV, such as HA, contain mannose-rich glycans that can be targeted by these lectins.32,33 Studies have revealed that different lectins exhibit varying degrees of anti-IAV activity. For example, griffithsin (GRFT), an algae-derived lectin, has demonstrated broad-spectrum antiviral effects against IAV infection.34 GRFT and a GRFT-based bivalent entry inhibitor, GL25E, have proven effective in inhibiting IAV infection both in vitro and in vivo.34 Their mechanisms of action typically involve binding to glycans on viral surface proteins, such as HA for IAV, possibly interfering with the virus's ability to attach to and enter host cells.35 This interaction potentially disrupts the viral life cycle at diverse stages, including entry, replication, and release.36 While sialic acid serves as the primary receptor for the HA-mediated entry of IAV, lectins that bind to glycan structures on HA, such as mannose and GlcNAc, can also exhibit antiviral effects.37 The effectiveness of these lectins potentially varies with their specific binding properties and the glycan composition of the viral proteins. In contrast to AFL, AFLm, a variant sequence of AFL, exhibited no significant change in IAV H1N1 replication compared with the control across all three treatment methods. This lack of antiviral activity in AFLm suggests that the specific amino acid sequence of AFL is critical for maintaining its functional integrity and antiviral properties. The loss of antiviral activity in the altered sequence indicates that even minor changes in amino acid composition can significantly impact the ability of the protein to interfere with viral processes. These findings suggest that AFL may possess key properties required for its antiviral activity in FBE, specifically under pre-treatment conditions. Overall, the anti-IAV H1N1 potential of AFL highlights the importance of its specific amino acid sequence for antiviral activity. Previous studies also corroborate that mutations in the carbohydrate-recognition domain of lectins can significantly alter their binding affinity and specificity, potentially affecting their role in biological processes such as pathogen recognition and antiviral activity.38
Interaction of FBE with IAV H1N1 HA
TEM was utilised to qualitatively evaluate structural modifications in IAV H1N1 particles following treatment with FBE (Fig. 5 and S1). Untreated viral particles displayed a characteristic spherical morphology, with discernible surface projections indicative of H1N1 HA spikes. In contrast, FBE-treated particles appeared aggregated and exhibited less-defined surface structures. Although TEM does not serve as a quantitative or conclusive method for assessing antiviral efficacy, the observed morphological alterations suggest that FBE may interact with the viral surface, potentially affecting HA-mediated entry. This hypothesis is further corroborated by the co-treatment assays, which revealed a 1.89-log reduction in viral replication, as measured by qRT-PCR (Fig. 3D). Comparable morphological changes observed via TEM have been documented in antiviral research, such as the disruption of influenza virions by graphene oxide nanoparticles, which resulted in envelope glycoprotein spike loss and particle deformation.39 This morphological transformation indicates that FBE directly interacts with IAV H1N1 particles, likely inhibiting viral entry by binding to HA on the viral surface.14 These findings provide compelling evidence that FBE can neutralise IAV H1N1 particles, suggesting a promising mechanism for preventing viral infection. The interaction between FBE and IAV H1N1 was further analysed using IP-MS/MS. His-tagged HA antigens were incubated with FBE, and HA-interacting proteins were isolated using His-tag-specific magnetic beads. Immunoprecipitation followed by nano LC-MS/MS identified a 36 kDa protein (UniProt accession: V7BGE3) as the only FBE-derived candidate that selectively bound to the His-tagged haemagglutinin (HA) of IAV H1N1 (Tables S2 and S3). This protein was captured using HA-conjugated magnetic beads and remained after multiple washes, indicating a specific interaction with the viral surface glycoprotein. This mechanism may resemble the action of small-molecule inhibitors—described by Basu et al.—that target HA-mediated IAV fusion.40 V7BGE3 was consistently identified as a potential HA-binding protein through HA-targeted proteomic analysis, suggesting its involvement in the antiviral mechanism of FBE. Although direct functional evidence remains limited, ongoing and future studies employing recombinant protein-based assays are expected to clarify its mechanistic relevance.Antiviral immune response gene expression in IAV-infected MDCK cells treated with FBE and AFL
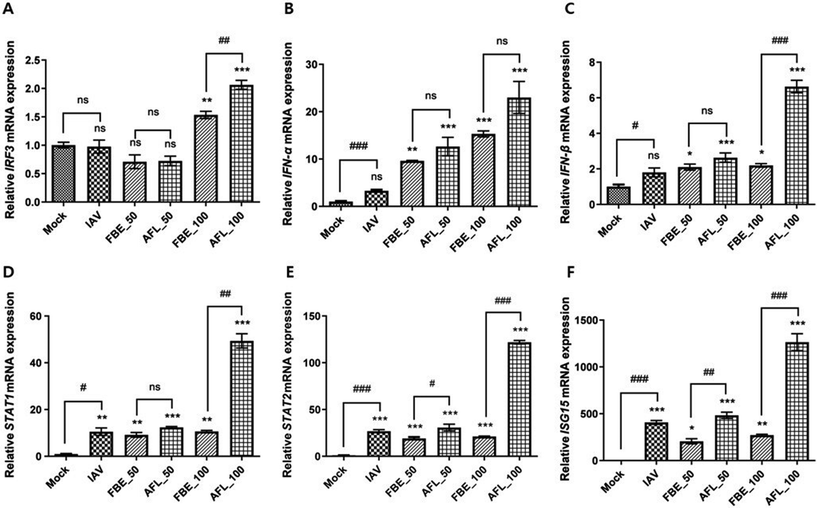
The effects of FBE and AFL on key antiviral immune response genes in IAV H1N1-infected MDCK cells revealed varied gene responses across treatments (Fig. 6). Interferon regulatory factor 3 (IRF3) expression significantly increased at high FBE and AFL concentrations (100 μg mL−1), yielding 1.53- and 2.06-fold increases, respectively, suggesting an enhanced antiviral response. Although the expression was not significantly different at low concentrations, this increase at higher concentrations aligns with the critical role of IRF3 in interferon production and interferon-stimulated gene (ISG) regulation.41Both treatments exhibited a dose-dependent activation of the interferon response in IAV-infected cells. FBE and AFL treatments significantly increased interferon alpha (IFN-α) expression in IAV-infected cells. AFL exerted a more pronounced effect on IFN-α expression than FBE at 50 and 100 μg mL−1, resulting in 12.64- and 23.00-fold increases compared with the control, while FBE produced 9.62- and 15.30-fold increases, respectively. These findings suggest that AFL may more effectively enhance the antiviral immune response, as IFN-α is essential to the innate immune defence against viral infections.42 At low concentration (50 μg mL−1), the treatment did not significantly affect interferon beta (IFN-β) expression; however, at 100 μg mL−1, AFL notably elevated IFN-β expression (3.02-fold) compared with FBE at the same concentration, further suggesting AFL's ability to enhance antiviral defences. Although IFN-β is crucial in the early stage of the immune response, its prolonged IFN-β signalling can lead to harmful inflammation,43 potentially exacerbating the severity of IAV infection in later stages.44 STAT1 and STAT2 were significantly upregulated in IAV-infected cells, yielding 10.53- and 26.82-fold increases, respectively, compared with mock treatments. AFL treatment further amplified their expression, particularly at higher doses, with STAT1 and STAT2 exhibiting 49.37- and 121.97-fold increases, respectively. This indicates a strong activation of the type I interferon pathway, consistent with the indispensable role of STAT1 and STAT2 in driving interferon-mediated antiviral responses.45,46 ISG15 expression increased dramatically (407.50-fold) in IAV-infected cells compared with that in the uninfected control, reflecting its vital role in the antiviral innate immune response against IAV infection.47,48 Interestingly, FBE treatment downregulated ISG15 expression in IAV-infected cells, whereas AFL significantly upregulated it, yielding a 3-fold upregulation at 100 μg mL−1. The contrasting effects of FBE and AFL on ISG15 expression indicate that FBE may control IAV replication through alternative mechanisms, possibly precluding the need for the ISG15 response. The significant alterations in ISG15 expression under different conditions suggest its potential as a biomarker for assessing both IAV infection and the antiviral treatment efficacy of antiviral treatments.47,49 These findings suggest that FBE and AFL exert antiviral activity through distinct mechanisms: FBE primarily via direct viral surface binding and AFL through the activation of type I interferon signalling. In this study, AFL significantly enhanced type I interferon responses in IAV-infected MDCK cells, including the notable upregulation of IFN-α, IFN-β, STAT1, STAT2, and ISG15. These results indicate a robust activation of interferon-stimulated gene (ISG) expression consistent with canonical antiviral signalling pathways. Notably, ISG15 was significantly elevated only in AFL-treated infected cells, whereas FBE, which exhibited strong antiviral effects, did not induce comparable ISG15 expression. This indicates that AFL and FBE may operate through distinct antiviral mechanisms: AFL primarily by potentiating host immune responses and FBE potentially through direct interaction with viral components. These findings suggest that AFL's immunomodulatory activity is likely infection-dependent rather than resulting from non-specific interferon stimulation. Moreover, no cytotoxicity was observed under the tested conditions, indicating a low likelihood of off-target inflammatory effects. Further in vivo studies are warranted to confirm the specificity and safety profile of AFL-mediated immune activation. These findings underscore the importance of considering multiple components of host–virus interactions when assessing antiviral agents.
Conclusions
This study establishes FBE as a promising antiviral agent against IAV H1N1. Its findings indicate that FBE exhibits multiple antiviral mechanisms. First, FBE effectively suppresses viral gene expression, especially that of the PA gene, which is crucial for viral replication. Second, the V7BGE3 protein identified in FBE probably interacts directly with IAV H1N1 particles by binding to HA, thereby modulating viral–host interactions and potentially inhibiting viral entry. Finally, AFL, an FBE-derived peptide, significantly boosts interferon responses, activating the antiviral defences of the host. These complex antiviral mechanisms accentuate the potential of FBE as a therapeutic agent against IAV H1N1. The combination of direct viral inhibition and host immune-response modulation renders FBE a valuable candidate for further research and development in antiviral therapeutics.Author contributions
J. Y. C.: investigation, methodology, validation, and writing – review & editing. S. Y. C.: conceptualisation, investigation, methodology, validation, writing – original draft, and writing – review & editing. B. V.: methodology, visualisation, and writing – review & editing. I. K.: formal analysis, funding acquisition, resources, and writing – review & editing. C. K.: formal analysis and writing – review & editing. D. S. K.: formal analysis, resources, and writing – review & editing. D. W. K.: conceptualisation, data curation, formal analysis, funding acquisition, investigation, and resources.Conflicts of interest
There are no conflicts to declare.Data availability
All data generated or analysed during this study are included within this submitted article and its supplementary information (SI). Additional data supporting the findings of this study are available from the corresponding author upon reasonable request.Supplementary information is available. See DOI: https://doi.org/10.1039/d5fo01554e.
Acknowledgements
This study was supported by the Basic Science Research Program (2021R1I1A3059219) and Brain Korea 21 Program for Leading Universities and Students (Fostering Outstanding Universities for Research, no. 4120240915070), both funded by the Ministry of Education (MOE) and National Research Foundation (NRF) of Korea, and by the Regional Innovation System & Education (RISE) through the Gwangju RISE Center, funded by the Ministry of Education (MOE) and the Gwangju Metropolitan Government, Republic of Korea (2025-RISE-05-011).References
- K. Das, J. M. Aramini, L. C. Ma, R. M. Krug and E. Arnold, Structures of influenza A proteins and insights into antiviral drug targets, Nat. Struct. Mol. Biol., 2010, 17, 530–538 CrossRef PubMed.
- D. P. Nayak, E. K. Hui and S. Barman, Assembly and budding of influenza virus, Virus Res., 2004, 106, 147–165 CrossRef PubMed.
- A. H. Ellebedy and R. J. Webby, Influenza vaccines, Vaccine, 2009, 27(Suppl 4), D65–D68 CrossRef PubMed.
- T. Han and W. A. Marasco, Structural basis of influenza virus neutralization, Ann. N. Y. Acad. Sci., 2011, 1217, 178–190 CrossRef PubMed.
- L. A. Grohskopf, L. H. Blanton, J. M. Ferdinands, J. R. Chung, K. R. Broder and H. K. Talbot, Prevention and Control of Seasonal Influenza with Vaccines: Recommendations of the Advisory Committee on Immunization Practices - United States, 2023–24 Influenza Season, MMWR Recomm. Rep., 2023, 72, 1–25 CrossRef.
- J. R. Chen, C. Ma and C. H. Wong, Vaccine design of hemagglutinin glycoprotein against influenza, Trends Biotechnol., 2011, 29, 426–434 CrossRef CAS PubMed.
- F. G. Hayden, R. P. Lenk, L. Stonis, C. Oldham-Creamer, L. L. Kang and C. Epstein, Favipiravir Treatment of Uncomplicated Influenza in Adults: Results of Two Phase 3, Randomized, Double-Blind, Placebo-Controlled Trials, J. Infect. Dis., 2022, 226, 1790–1799 CrossRef CAS PubMed.
- K. Muramoto, Lectins as Bioactive Proteins in Foods and Feeds, Food Sci. Technol. Res., 2017, 23, 487–494 CrossRef CAS.
- E. J. Van Damme, N. Lannoo and W. J. Peumans, in Advances in botanical research, Elsevier, 2008, vol. 48, pp. 107–209 Search PubMed.
- Y. Sato, K. Morimoto, T. Kubo, T. Sakaguchi, A. Nishizono, M. Hirayama and K. Hori, Entry Inhibition of Influenza Viruses with High Mannose Binding Lectin ESA-2 from the Red Alga Eucheuma serra through the Recognition of Viral Hemagglutinin, Mar. Drugs, 2015, 13, 3454–3465 CrossRef CAS PubMed.
- L. S. Ooi, W.-S. Ho, K. L. Ngai, L. Tian, P. K. Chan, S. S. Sun and V. E. Ooi, Narcissus tazetta lectin shows strong inhibitory effects against respiratory syncytial virus, influenza A (H1N1, H3N2, H5N1) and B viruses, J. Biosci., 2010, 35, 95–103 CrossRef CAS PubMed.
- Y. Sato, K. Morimoto, M. Hirayama and K. Hori, High mannose-specific lectin (KAA-2) from the red alga Kappaphycus alvarezii potently inhibits influenza virus infection in a strain-independent manner, Biochem. Biophys. Res. Commun., 2011, 405, 291–296 CrossRef CAS PubMed.
- E. Vanderlinden, N. Van Winkel, L. Naesens, E. J. Van Damme, L. Persoons and D. Schols, In vitro characterization of the carbohydrate-binding agents HHA, GNA, and UDA as inhibitors of influenza A and B virus replication, Antimicrob. Agents Chemother., 2021, 65(3), e01732-20 CrossRef PubMed.
- Y.-M. Liu, M. Shahed-Al-Mahmud, X. Chen, T.-H. Chen, K.-S. Liao, J. M. Lo, Y.-M. Wu, M.-C. Ho, C.-Y. Wu and C.-H. Wong, A carbohydrate-binding protein from the edible lablab beans effectively blocks the infections of influenza viruses and SARS-CoV-2, Cell Rep., 2020, 32, 108016 CrossRef CAS PubMed.
- E. M. Covés-Datson, S. R. King, M. Legendre, A. Gupta, S. M. Chan, E. Gitlin, V. V. Kulkarni, J. P. García, D. F. Smee, E. Lipka, S. E. Evans, E. B. Tarbet, A. Ono and D. M. Markovitz, A molecularly engineered antiviral banana lectin inhibits fusion and is efficacious against influenza virus infection in vivo, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 2122–2132 CrossRef PubMed.
- S.-M. Hsu and L. Raine, Discovery of a high affinity of Phaseolus vulgaris agglutinin (PHA) with gastrin-secreting cells, Am. J. Clin. Pathol., 1982, 77, 396–400 CrossRef CAS PubMed.
- C. A. Mitchell, K. Ramessar and B. R. O'Keefe, Antiviral lectins: Selective inhibitors of viral entry, Antiviral Res., 2017, 142, 37–54 CrossRef CAS PubMed.
- J. Kwon, E. Ko, S.-Y. Cho, Y.-H. Lee, S. Jun, K. Lee, E. Hwang, B. Vaidya, J.-H. Hwang and J.-H. Hwang, Bean extract-based gargle for efficient diagnosis of active COVID-19 infection using rapid antigen tests, Microbiol. Spectrum, 2022, 10, e01614–e01621 CAS.
- X. Lu and H. Zhu, Tube-gel digestion, Mol. Cell. Proteomics, 2005, 4, 1948–1958 CrossRef CAS PubMed.
- S.-Y. Cho, R. A. Protzman, Y. O. Kim, B. Vaidya, M.-J. Oh, J. Kwon and D. Kim, Elucidation of mechanism for host response to VHSV infection at varying temperatures in vitro and in vivo through proteomic analysis, Fish Shellfish Immunol., 2019, 88, 244–253 CrossRef CAS PubMed.
- S. Kumar, A. K. Verma, M. Das, S. Jain and P. D. Dwivedi, Clinical complications of kidney bean (Phaseolus vulgaris L.) consumption, Nutrition, 2013, 29, 821–827 CrossRef CAS PubMed.
- P. Wang, X. Leng, J. Duan, Y. Zhu, J. Wang, Z. Yan, S. Min, D. Wei and X. Wang, Functional component isolated from Phaseolus vulgaris lectin exerts in vitro and in vivo anti-tumor activity through potentiation of apoptosis and immunomodulation, Molecules, 2021, 26, 498 CrossRef CAS PubMed.
- W. L. DeLano, Pymol: An open-source molecular graphics tool, CCP4 Newsl, Protein Crystallogr., 2002, 40, 82–92 Search PubMed.
- H.-X. Zhou and X. Pang, Electrostatic interactions in protein structure, folding, binding, and condensation, Chem. Rev., 2018, 118, 1691–1741 CrossRef CAS PubMed.
- X. Zheng, L. Deng, E. S. Baker, Y. M. Ibrahim, V. A. Petyuk and R. D. Smith, Distinguishing d-and l-aspartic and isoaspartic acids in amyloid β peptides with ultrahigh resolution ion mobility spectrometry, Chem. Commun., 2017, 53, 7913–7916 RSC.
- M. Kruse, B. Altattan, E.-M. Laux, N. Grasse, L. Heinig, C. Möser, D. M. Smith and R. Hölzel, Characterization of binding interactions of SARS-CoV-2 spike protein and DNA-peptide nanostructures, Sci. Rep., 2022, 12, 12828 CrossRef CAS PubMed.
- Y. Behzadipour and S. Hemmati, Viral prefusion targeting using entry inhibitor peptides: the case of SARS-CoV-2 and Influenza A virus, Int. J. Pept. Res. Ther., 2022, 28, 42 CrossRef CAS PubMed.
- D. Lauster, M. Glanz, M. Bardua, K. Ludwig, M. Hellmund, U. Hoffmann, A. Hamann, C. Böttcher, R. Haag and C. P. Hackenberger, Multivalent peptide–nanoparticle conjugates for influenza–virus inhibition, Angew. Chem., Int. Ed., 2017, 56, 5931–5936 CrossRef CAS PubMed.
- W. Van Breedam, S. Pöhlmann, H. W. Favoreel, R. J. de Groot and H. J. Nauwynck, Bitter-sweet symphony: glycan–lectin interactions in virus biology, FEMS Microbiol. Rev., 2014, 38, 598–632 CrossRef CAS PubMed.
- M. Nagae, K. Soga, K. Morita-Matsumoto, S. Hanashima, A. Ikeda, K. Yamamoto and Y. Yamaguchi, Phytohemagglutinin from Phaseolus vulgaris (PHA-E) displays a novel glycan recognition mode using a common legume lectin fold, Glycobiology, 2014, 24, 368–378 CrossRef CAS PubMed.
- A. Mishra, A. Behura, S. Mawatwal, A. Kumar, L. Naik, S. S. Mohanty, D. Manna, P. Dokania, A. Mishra and S. K. Patra, Structure-function and application of plant lectins in disease biology and immunity, Food Chem. Toxicol., 2019, 134, 110827 CrossRef CAS PubMed.
- J. E. Stencel-Baerenwald, K. Reiss, D. M. Reiter, T. Stehle and T. S. Dermody, The sweet spot: defining virus–sialic acid interactions, Nat. Rev. Microbiol., 2014, 12, 739–749 CrossRef CAS PubMed.
- M. Matrosovich, G. Herrler and H. D. Klenk, Sialic acid receptors of viruses, in Sialoglyco chemistry and biology II: tools and techniques to identify and capture sialoglycans, 2015, pp. 1–28 Search PubMed.
- N. Cao, Y. Cai, X. Huang, H. Jiang, Z. Huang, L. Xing, L. Lu, S. Jiang and W. Xu, Inhibition of influenza A virus and SARS-CoV-2 infection or co-infection by griffithsin and griffithsin-based bivalent entry inhibitor, mBio, 2024, 15, e00741–e00724 Search PubMed.
- W. C. Ng, M. D. Tate, A. G. Brooks and P. C. Reading, Soluble host defense lectins in innate immunity to influenza virus, BioMed Res. Int., 2012, 2012, 732191 Search PubMed.
- D. S. Dimitrov, Virus entry: molecular mechanisms and biomedical applications, Nat. Rev. Microbiol., 2004, 2, 109–122 CrossRef PubMed.
- D. E. Mattox and C. Bailey-Kellogg, Comprehensive analysis of lectin-glycan interactions reveals determinants of lectin specificity, PLoS Comput. Biol., 2021, 17, e1009470 CrossRef CAS PubMed.
- M. Waespy, T. T. Gbem, L. Elenschneider, A.-P. Jeck, C. J. Day, L. Hartley-Tassell, N. Bovin, J. Tiralongo, T. Haselhorst and S. Kelm, Carbohydrate recognition specificity of trans-sialidase lectin domain from Trypanosoma congolense, PLoS Neglected Trop. Dis., 2015, 9, e0004120 CrossRef PubMed.
- H. Wang, J. Wang, J. Zhang, J. Song, D. Wang, J. Dong and H. Liu, Graphene oxide nanoparticles inhibit H9N2 influenza A virus infectivity by destroying viral coat proteins, Arch. Virol., 2024, 169, 192 CrossRef CAS PubMed.
- A. Basu, A. Antanasijevic, M. Wang, B. Li, D. M. Mills, J. A. Ames, P. J. Nash, J. D. Williams, N. P. Peet and D. T. Moir, New small molecule entry inhibitors targeting hemagglutinin-mediated influenza a virus fusion, J. Virol., 2014, 88, 1447–1460 CrossRef PubMed.
- T. H. Mogensen, IRF and STAT transcription factors-from basic biology to roles in infection, protective immunity, and primary immunodeficiencies, Front. Immunol., 2019, 9, 3047 CrossRef PubMed.
- F. McNab, K. Mayer-Barber, A. Sher, A. Wack and A. O′garra, Type I interferons in infectious disease, Nat. Rev. Immunol., 2015, 15, 87–103 CrossRef CAS PubMed.
- K. Högner, T. Wolff, S. Pleschka, S. Plog, A. D. Gruber, U. Kalinke, H.-D. Walmrath, J. Bodner, S. Gattenlöhner and P. Lewe-Schlosser, Macrophage-expressed IFN-β contributes to apoptotic alveolar epithelial cell injury in severe influenza virus pneumonia, PLoS Pathog., 2013, 9, e1003188 CrossRef PubMed.
- Y. Börgeling, M. Schmolke, D. Viemann, C. Nordhoff, J. Roth and S. Ludwig, Inhibition of p38 mitogen-activated protein kinase impairs influenza virus-induced primary and secondary host gene responses and protects mice from lethal H5N1 infection, J. Biol. Chem., 2014, 289, 13–27 CrossRef PubMed.
- N. Au-Yeung, R. Mandhana and C. M. Horvath, Transcriptional regulation by STAT1 and STAT2 in the interferon JAK-STAT pathway, JAKSTAT, 2013, 2, e23931 Search PubMed.
- J. E. Durbin, R. Hackenmiller, M. C. Simon and D. E. Levy, Targeted disruption of the mouse Stat1 gene results in compromised innate immunity to viral disease, Cell, 1996, 84, 443–450 CrossRef CAS PubMed.
- Q. Ye, T. Phan, W.-S. Hu, X. Liu, L. Fan, W.-S. Tan and L. Zhao, Transcriptomic characterization reveals attributes of high influenza virus productivity in MDCK cells, Viruses, 2021, 13, 2200 CrossRef CAS PubMed.
- Y.-C. Perng and D. J. Lenschow, ISG15 in antiviral immunity and beyond, Nat. Rev. Microbiol., 2018, 16, 423–439 CrossRef CAS PubMed.
- A. A. Hinay Jr, S. Kakee, S. Kageyama, A. Tsuneki-Tokunaga, W. Y. Perdana, Y. Akena, S. Nishiyama and K. Kanai, Pro-inflammatory cytokines and interferon-stimulated gene responses induced by seasonal influenza a virus with varying growth capabilities in human lung epithelial cell lines, Vaccines, 2022, 10, 1507 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2025 |