Lactic acid bacteria target NF-κB signaling to alleviate gastric inflammation
Shiying
Wu†
 *a,
Yuenuo
Luo†
a,
Fangtong
Wei
a,
Yanan
Li
b,
Jiayi
Fan
a,
Yongqiang
Chen
a,
Wenjie
Zhang
c,
Xuelong
Li
c,
Yang
Xu
a,
Ziqi
Chen
*a,
Yuenuo
Luo†
a,
Fangtong
Wei
a,
Yanan
Li
b,
Jiayi
Fan
a,
Yongqiang
Chen
a,
Wenjie
Zhang
c,
Xuelong
Li
c,
Yang
Xu
a,
Ziqi
Chen
 a,
Chenlan
Xia
a,
Mingyang
Hu
a,
Ping
Li
a,
Chenlan
Xia
a,
Mingyang
Hu
a,
Ping
Li
 a and
Qing
Gu
a and
Qing
Gu
 *a
*a
aKey Laboratory for Food Microbial Technology of Zhejiang Province, Zhejiang Gongshang University, Hangzhou, Zhejiang 310018, China. E-mail: wushiying@zjgsu.edu.cn; guqing2002@hotmail.com
bSchool of Food Science and Pharmaceutical Engineering, Nanjing Normal University, Nanjing, Jiangsu 210023, China
cHangzhou Helixinjian Industry Co., Ltd, No. 48 Zijinghua Road, Gudang Street, Xihu District, Hangzhou, Zhejiang 310050, China
First published on 25th March 2025
Abstract
Helicobacter pylori (H. pylori) infection and the resulting gastric inflammation are major contributors to gastric cancer development. Probiotics, particularly Lactobacillus, are promising for their anti-inflammatory potential, yet their exact mechanisms in inhibiting H. pylori-induced inflammation are unclear. In our previous study, Lactiplantibacillus plantarum ZJ316 (L. plantarum ZJ316) demonstrated strong anti-inflammatory effects against H. pylori infection in vivo, but its precise mechanisms were not fully understood. Here, we aimed to investigate how L. plantarum ZJ316 inhibits the inflammatory response to H. pylori infection. Our results demonstrated that L. plantarum ZJ316 effectively reduced the expression of pro-inflammatory cytokines in H. pylori-infected AGS cells. Mechanistically, L. plantarum ZJ316 inhibited the NF-κB signaling pathway by preventing the degradation of IκBα, suppressing p65 phosphorylation, and blocking the nuclear translocation of phosphorylated p65. Treatment with the NF-κB inhibitor BAY 11-7082 further decreased tumor necrosis factor-α (TNF-α), interleukin-8 (IL-8), and interleukin-1β (IL-1β) levels, confirming the inhibitory effect of L. plantarum ZJ316 on the NF-κB pathway. In H. pylori-infected mice, oral administration of L. plantarum ZJ316 significantly alleviated inflammatory cell infiltration, reduced TNF-α and pepsinogen II (PGII) levels, and increased interleukin-10 (IL-10) levels in serum. A comparative metagenomic analysis of the gastric microbiota revealed a decrease in Prevotella and Desulfovibrio, alongside an increase in Ligilactobacillus and Akkermansia, supporting the protective effects of L. plantarum ZJ316 and correlating with their decreased inflammatory response. In summary, administration of L. plantarum ZJ316 demonstrated robust anti-inflammatory effects against H. pylori infection by suppressing NF-κB signaling and promoting favorable changes in the gastric microbiota composition. Therefore, L. plantarum ZJ316 holds promise as a novel functional food for protecting the body against H. pylori infection.
1. Introduction
Helicobacter pylori (H. pylori) is the most significant risk factor for gastric cancer, which ranks as the third leading cause of cancer-related deaths globally.1 While only 2% of individuals harboring H. pylori eventually develop malignancies, the bacterium is also implicated in other gastric conditions such as ulcers and gastritis.2 The etiology of H. pylori involves several factors, including genetic predispositions, the host environment, and specific bacterial virulence factors.3 Moreover, elevated levels of inflammatory responses among long-standing H. pylori infestation are particularly critical, playing a substantial role in the development and advancement of gastric cancer.2H. pylori colonization and disease establishment involve several key steps that enable the bacterium to thrive in the acidic stomach environment and induce gastric damage.4 Initially, H. pylori adapts to the stomach's acidity by producing urease, which neutralizes gastric acid, allowing survival in that harsh environment.5 The bacterium then uses its flagella to navigate through the mucus layer and adhere to gastric epithelial cells6via adhesins such as blood group antigen-binding adhesin (BabA) and sialic acid-binding adhesin (SabA).7,8 Once attached, H. pylori secretes virulence factors like cytotoxin-associated gene A (CagA), which disrupts cellular signaling, weakens tight junctions between epithelial cells, and increases mucosal barrier permeability.9,10 This triggers a chronic inflammatory response, with the release of pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6,11,12 promoting cellular stress, oxidative damage, and DNA mutations in gastric epithelial cells.13,14 Additionally, H. pylori alters host signaling pathways, including NF-κB, MAPK, and JAK-STAT, thereby sustaining inflammation.15–17 Over time, this persistent inflammation leads to epithelial cell damage, metaplasia, and potential progression to gastritis and even gastric cancer.7,10
The persistent inflammation induced by H. pylori infection causes ongoing tissue damage, genetic mutations, and carcinogenesis, making it a major risk factor for gastric diseases.18 NF-κB and STAT3 pathways are prominently activated during carcinogenesis.19 NF-κB, a ubiquitous inflammatory pathway in animal cells, exhibits constitutive activation in gastric cancer,20,21 particularly mediated by CagA following H. pylori infestation. NF-κB activation induces the secretion of diverse inflammatory cytokines, which are implicated in processes including angiogenesis and the inhibition of apoptosis.22 Notably, NF-κB activation has been recognized as crucial in linking H. pylori-induced inflammation with cancer development. Yang et al. demonstrated that H. pylori triggers miR-223-3p upregulation via CagA-dependent mechanisms, with NF-κB directly stimulating gastric cancer cell proliferation by binding to the miR-223-3p promoter.23 Moreover, the lipopolysaccharide of this bacterium binds to the toll-like receptor on the surface of gastric epithelial cells,24 triggering subsequent inflammatory signaling pathways, including NF-κB.25,26 Inhibiting NF-κB activation could mitigate inflammation and potentially reduce cancer risk. Emerging evidence highlights the crucial importance of the gastric microbiota in regulating the development of H. pylori-related chronic diseases.27 Previous research has shown that in comparison with individuals without H. pylori, individuals who have been diagnosed with an H. pylori infection exhibit an enrichment of specific gastric bacterial genera, including Streptococcus, Neisseria, Prevotella, Rothia, Fusobacterium, Veillonella, and Haemophilus, which contribute to inflammatory processes.28–30
Early elimination of H. pylori infestation can potentially regress associated diseases.31–33 However, the effectiveness of eradication regimens has been considerably undermined globally by increasing antimicrobial resistance and uncertain impact on the gut microbiota, hindering widespread screening and eradication efforts.34–36 Moreover, current treatments for H. pylori eradication are beset by numerous drawbacks: high costs, complex protocols, frequent side effects, and low patient adherence due to complex drug regimens.37 Therefore, exploring alternative or antibiotic-free strategies is crucial to mitigate H. pylori-induced gastric diseases.
Over the past few years, the European Helicobacter and Microbiota Study Group has proposed updated H. pylori treatment guidelines that recommend specific probiotics as adjunct therapies, particularly for patients who experience significant antibiotic intolerance.38,39 Clinical research has supported the efficacy of probiotic administration, primarily using Lactobacillus strains, in alleviating disease severity and reducing antibiotic side effects.40–43Lactobacillus, widely acknowledged for its health benefits, positively impacts human health by modulating the gut microbiota, enhancing beneficial metabolite production, and regulating immune responses.44 In recent years, a large number of research studies have underscored its protective role against H. pylori-induced gastric diseases through inhibiting bacterial growth, enhancing the gastrointestinal microbiome community, and inducing apoptosis in gastric cancer cells.45,46 Notably, Urrutia-Baca et al. reported that reuterin, a bacteriocin derived from Limosilactobacillus reuteri, effectively suppresses the transcription of the VacA and FlaA genes of H. pylori.47 Several Lactobacillus species, including Lacticaseibacillus rhamnosus, Lactobacillus acidophilus, and Levilactobacillus brevis, demonstrate the ability to attach to gastric epithelial cells in vitro and mitigate H. pylori-triggered inflammation in animal models.48–50 Moreover, certain probiotic LAB strains alleviate H. pylori-induced inflammation by suppressing IL-8 secretion, potentially through the inactivation of Smad7 and NF-κB pathways.51,52 While probiotics, particularly Lactobacillus species, have shown promise in mitigating inflammation caused by H. pylori infection, the precise molecular mechanisms behind their anti-inflammatory effects remain unclear. Current research often emphasizes the anti-inflammatory effects of probiotics, but lacks an in-depth analysis of how these effects are linked to the changes in the gut microbiota. Additionally, while much of the existing research on H. pylori focuses on its direct impact on the gastric epithelium, there is a lack of comprehensive studies exploring how probiotics modulate the host's immune response to H. pylori.7
Our previous investigations have shown that L. plantarum ZJ316 successfully mitigates H. pylori-induced inflammatory responses in murine models.53,54 However, the specific mechanisms have remained elusive. In this study, we delve into the anti-inflammatory properties of L. plantarum ZJ316 both in vitro and in vivo during H. pylori infection. Compelling evidence showed that L. plantarum ZJ316 substantially suppresses H. pylori-induced NF-κB signaling activation, confirmed by inhibiting the NF-κB pathway with BAY 11-7082. Moreover, leveraging multi-omics analysis revealed that supplementation with L. plantarum ZJ316 enhances the abundance of Ligilactobacillus and Akkermansia, a widely recognized probiotic group known for its potential to alleviate inflammation. Overall, this study sets the stage for further exploration of novel probiotic functional foods, offering valuable insights for future research on effective strategies for preventing and managing H. pylori-associated inflammatory conditions.
2. Materials and methods
2.1. Bacterial strains and cell culture
Helicobacter pylori (CCTCC M20211218), a gift from Yongliang Zhu (Second Affiliated Hospital of Zhejiang University School of Medicine), was isolated from H. pylori positive gastritis patients. H. pylori was cultured on CBA supplemented with 7% v/v sheep blood and antibiotics (Hopebio, Qingdao, China) at 37 °C under microaerobic conditions.53Lactiplantibacillus plantarum ZJ316 (CCTCC M208077), originally selected from the laboratory, was cultivated in MRS medium at 37 °C for 36–48 hours.The AGS cells (human gastric adenocarcinoma epithelial cells) were acquired from the National Collection of Authenticated Cell Cultures (cat. no. TCHu232). AGS cells were propagated in Ham's F-12K medium fortified containing 10% fetal bovine serum (FBS, Procell, China) at 37 °C under 5% CO2.53 Then, the bacteria mentioned above were centrifuged, diluted in a fresh F-12K medium, and poured into AGS cells at a multiplicity of infection (MOI) of 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 for 2, 6 or 12 hours for the subsequent experimental assays. Cells were exposed to the NF-κB inhibitor BAY 11-7082 (MCE) at a concentration of 10 μM.
1 for 2, 6 or 12 hours for the subsequent experimental assays. Cells were exposed to the NF-κB inhibitor BAY 11-7082 (MCE) at a concentration of 10 μM.
2.2. RNA extraction, reverse transcription and RT-qPCR
Total RNA extraction from cells and tissue specimens was carried out using the TRIzol reagent (Sigma-Aldrich). Subsequently, the isolated RNA was converted into cDNA using the Severn Biotech synthesis kit. RT-qPCR analysis was conducted by using the TB Green promoter RTQ-960 PCR system using the SYBR Green kit (Severn Biotech), following the manufacturer's protocols. The expression patterns and Ct values were quantitatively evaluated via real-time PCR, and the gene expression changes were assessed using the 2−ΔΔCt method, normalized to the internal controls, with the control group set as 1 to facilitate relative comparisons between the groups. This standard approach was applied consistently across all the groups to account for inter-sample variations. All experiments were conducted in triplicate for robust statistical validation. The primer sequences used are detailed in Table 1.| Genes | Primers (5′–3′) |
|---|---|
| h GAPDH-F | GGCGACCTGCTGGAACATTACTG |
| h GAPDH-R | CATCGTTTAGGGCGTGGACTACC |
| h IL-6-F | AGACAGCCACTCACCTCTTCAG |
| h IL-6-R | TTCTGCCAGTGCCTCTTTGCTG |
| h IL-1β-F | CGGTTGCGTGGAAGACACTATGG |
| h IL-1β-R | GTGGAAGCGGGTAGGATGTTGTG |
| h TNF-α-F | TGAAAACCTCCAAGCAGCCCAAC |
| h TNF-α-R | GGCGTGTTTGATGTTGAGTCTGTTG |
| h IFN-γ-F | GAGTGTGGAGACCATCAAGGAAG |
| h IFN-γ-R | TGCTTTGCGTTGGACATTCAAGTC |
| h MAP3K14-F | CCACCTTTTCAGAACGCATTTTC |
| h MAP3K14-R | GTAGCATGGGCCACATTGTTG |
| h IκBα-F | TCCACTCCATCCTGAAGGCTAC |
| h IκBα-R | CAAGGACACCAAAAGCTCCACG |
| h IL-8-F | CGGATGATGTGATGGATGGTGTGG |
| h IL-8-R | GGGTATGCACGGTTACGAGTTTGG |
| m GAPDH-F | GGAGCGAGATCCCTCCAAAA |
| m GAPDH-R | GGCTGTTGTCATACTTCTCATGG |
| m IκBα-F | CACTTGGTGACTTTGGGTGCT |
| m IκBα-R | GCTGTATCCGGGTACTTGGG |
2.3. RNA-seq analysis of AGS cells
AGS cells underwent total RNA isolation with the TRIzol reagent (Sigma-Aldrich) as per the manufacturer's guidelines, followed by genomic DNA removal achieved using DNase I (Takara).55 Subsequently, the cDNA library was prepared following the protocol of the TruSeq™ RNA sample preparation kit,55 and library preparations were carried out by Nuohe Zhiyuan Biotechnology Co., Ltd. Differential expression analysis between the samples was conducted with the transcripts per million reads method. Genes were classified as differentially expressed if they exhibited a p-value <0.05 and a log2 fold-change ≥1, determined using DESeq2. Enrichment analyses for functional annotation, encompassing GO and KEGG pathway analyses, were conducted with the results reported at a false discovery rate (FDR) <0.05.2.4. Dual luciferase reporting system and western blot analysis
AGS cells, cultured in 24-well plates, were co-transfected with NF-κB luciferase plasmids and pRL-TK for 24 h. After a 4-hour exposure to H. pylori, the cells were harvested, and Renilla and firefly luciferase activities were measured using a dual luciferase reporter assay system. The Renilla luciferase activity was then normalized to the firefly luciferase activity. Cellular proteins were isolated using a RIPA lysis buffer, and their concentrations were assessed by using a BCA reagent kit (Solarbio). Proteins were then resolved by SDS-PAGE and transferred to PVDF membranes, blocked, and subsequently exposed overnight at 4 °C to primary antibodies targeting NF-κB p65 (8242S, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), phosphorylated NF-κB p65 (3033S, 1
1000), phosphorylated NF-κB p65 (3033S, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), IκBα (4814S, 1
1000), IκBα (4814S, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1200), and α-tubulin (2144S, 1
1200), and α-tubulin (2144S, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000) (Cell Signaling Technology). After TBST washes, the membranes were incubated with HRP-conjugated secondary antibodies and visualized by the enhanced chemiluminescence method (Omni-ECL).
1000) (Cell Signaling Technology). After TBST washes, the membranes were incubated with HRP-conjugated secondary antibodies and visualized by the enhanced chemiluminescence method (Omni-ECL).
2.5. Animal experiments
Six to eight-week-old SPF C57BL/6J mice were kept in a 12-hour light/dark cycle and provided ad libitum access to food and water. Room conditions were regulated at temperatures ranging from 18 to 26 °C and relative humidity between 40% and 70%. All animal procedures were conducted in compliance with the guidelines approved by the Animal Ethics Committee of Shanghai Public Health Clinical Center (approval number: 2022-A037-01). Following a one-week acclimation period, mice were assigned to four experimental groups in a randomized manner (n = 8 per group).2.6. ELISA, hematoxylin–eosin staining and histological immunofluorescence
Serum samples were obtained and subjected to IL-10, TNF-α, and PGII levels using ELISA kits (EK-Bioscience), following the manufacturer's guidelines.The gastric tissues were preserved in 4% paraformaldehyde, followed by embedding in paraffin, slicing into 4 mm slices,56 and staining with hematoxylin and eosin (H&E). The immunostaining of gastric tissue was conducted following the established protocols as previously described.57 After deparaffinization and rehydration, antigen retrieval was executed, and subsequently, blocking was carried out. Slides underwent overnight incubation at 4 °C with primary antibodies, followed by exposure to secondary antibodies and a subsequent 1-hour incubation in darkness. Finally, the samples were examined and photographed using a Zeiss LSM710 confocal microscope. Histological scoring was performed based on the following criteria: scoring was performed for the mucosal layer structure, adenoids, columnar epithelial cells, and inflammatory cell infiltration, divided into four levels, each with a score of 25, to reflect the severity of the injury. A score of 0–25 indicates no tissue damage, 26–50 indicates mild injury, 51–75 indicates moderate injury, and 76–100 indicates severe injury. The evaluation includes mucosal clarity, adenoid atrophy and necrosis, epithelial cell integrity, and inflammatory cell infiltration. The protein levels were quantified by IHC using ImageJ software (National Institutes of Health, Bethesda, USA) and the integrated optical density was analyzed using the color deconvolution method.58
2.7. Metagenomic sequencing
Gastric tissue samples from various groups were collected after experiments. DNA isolation was performed using the fast DNA spin kit following the manufacturer's protocol. Beijing Biomarker Biotechnology Co. Ltd (Beijing, China) conducted the library preparation and paired-end sequencing. Gastric microbiota analysis was carried out using BMK Cloud (https://www.biocloud.net).2.8. Statistical analysis
Values are presented as the mean ± standard deviation (M ± SD) from three independent experiments. One-way analysis of variance (ANOVA) was used to compare the differences among the groups, followed by a post hoc analysis with the least significant difference (LSD) test. A p-value of less than 0.05 was considered statistically significant. Pearson correlation coefficients were employed to assess the relationships between the variables. All statistical analyses were performed using GraphPad Prism software (version 9.0; GraphPad Software Inc., San Diego, CA, USA).3. Results
3.1. L. plantarum ZJ316 reduces the inflammatory response in AGS following H. pylori infection
Persistent H. pylori infestation triggers an inflammatory response, contributing to mucosal damage. We established an infection model to investigate how L. plantarum ZJ316 modulates the H. pylori-triggered inflammatory response in AGS cells. In the context of H. pylori infection, we noted a marked elevation in the expression levels of inflammatory factors, including TNF-α, IL-1β, IFN-γ, and IL-6 at 2, 6, and 12 h. Conversely, treatment with L. plantarum ZJ316 resulted in notable reductions in the pro-inflammatory cytokine levels compared to the untreated cells. Specifically, TNF-α levels decreased by 49.12%, 75.54%, and 52.48%, IL-1β by 70.04%, 38.35%, and 85.43%, IFN-γ by 78.99%, 21.57%, and 51.55%, and IL-6 by 77.95%, 60.94%, and 36.00%, respectively (Fig. 1A–D). These findings highlight the promising prospects of L. plantarum ZJ316 to mitigate cytokine production in AGS cells in the presence of H. pylori infection.3.2. Treatment with L. plantarum ZJ316 effectively reversed the alterations in AGS cell transcriptional profiles caused by H. pylori infection
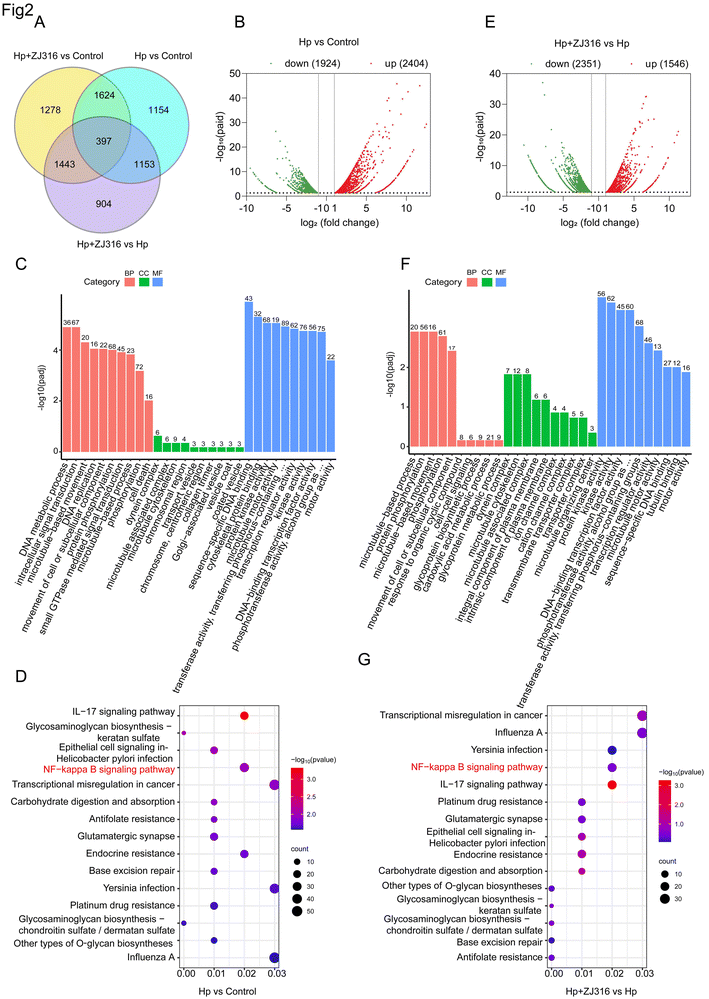
To examine the effects of L. plantarum ZJ316 on the primary inflammatory signaling pathway following H. pylori infection at the transcriptome level, we conducted a high-throughput RNA-seq analysis on AGS cells from three experimental groups, including control, Hp, and Hp + ZJ316. To ensure robustness in our cluster analysis, genes were selected based on the highest coefficient of variation across the entire dataset. Using this criterion, we identified a set of 397 identified differentially expressed genes (DEGs) that exhibited changes consistently across all experimental conditions, as visualized in the Venn plot (Fig. 2A). The volcano plot analysis highlighted the significant differences in the gene expression patterns among the experimental groups. Between the Hp group and the control group, a total of 4328 genes showed differential expressions, with 2404 genes being upregulated and 1924 genes downregulated (Fig. 2B). To elucidate the functions of the identified DEGs, we conducted GO analysis, categorizing DEGs into three primary functional groups: molecular function (MF), biological process (BP), and cellular component (CC) (Fig. 2C). In terms of biological processes, the DEGs were notably enriched in processes related to cell death, protein phosphorylation, phosphorylation, and intracellular signal transduction. In molecular functions, the enriched categories included protein kinase activity, transcription regulator activity and kinase activity (Fig. 2C). In our analysis of KEGG pathways, we observed a significant enrichment of DEGs in the top 15 pathways, notably including the IL-17 signaling pathway, NF-kappa B signaling pathway, and epithelial cell signaling in H. pylori infection. These enrichments highlight the alterations in gene regulation and nucleic acid interactions induced by H. pylori (Fig. 2D).Within the H. pylori-infected cells, comparison between the Hp group and the Hp + ZJ316 treatment group revealed 3897 differentially expressed genes, including 1546 that were upregulated and 2351 that were downregulated (Fig. 2E). These findings underscored the complex interplay of gene regulation influenced by both H. pylori infection and the addition of L. plantarum ZJ316, suggesting potential pathways through which L. plantarum ZJ316 could potentially alter the host response to H. pylori infection at the transcriptional level. The DEGs exhibited notable enrichment in GO function terms, related to protein phosphorylation, phosphorylation, protein kinase activity and kinase activity (Fig. 2F). Overall, these results indicated that the treatments with L. plantarum ZJ316 and H. pylori lead to significant changes in internal metabolism and transcriptional activities of specific genes in AGS cells. The KEGG pathway analysis revealed top 15 pathways enriched with statistically significant DEGs, which encompassed transcriptional misregulation in cancer, the NF-kappa B signaling pathway, and the IL-17 signaling pathway (Fig. 2G). The analysis revealed that H. pylori infection led to increased expressions of genes associated with inflammatory pathways like the NF-kappa B signaling pathway and IL-17 cell differentiation. Further analysis of these genes could provide deeper insights into the specific mechanisms through which L. plantarum ZJ316 exerts its effects on the inflammatory signaling pathway in this context.
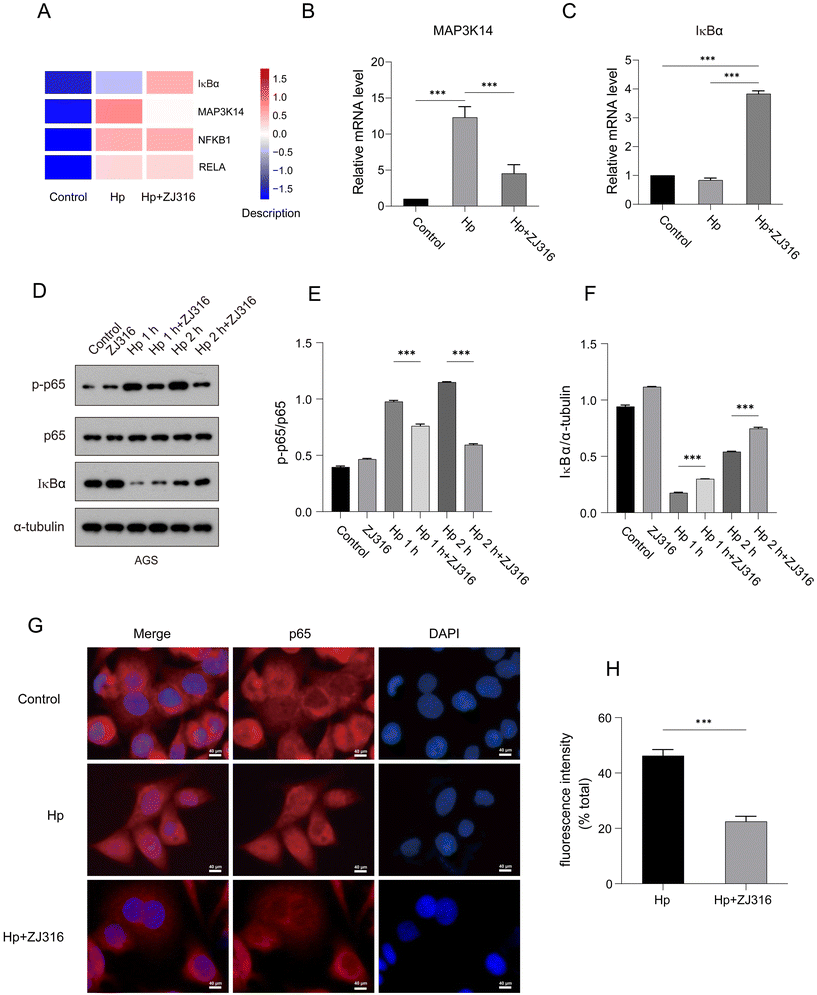
3.3. L. plantarum ZJ316 decreases the H. pylori-mediated activation of NF-κB in AGS cells
The NF-κB pathway is known to be constitutively active in various cancer cells.59–62L. plantarum ZJ316 has shown promise in inhibiting inflammatory factors. KEGG pathway analysis indicated that L. plantarum ZJ316 may suppress H. pylori-induced inflammation by modulating NF-κB activity. Then, we conducted gene set analysis to delve deeper into NF-κB-related gene changes and visualized the expression patterns of the top 4 significantly altered genes using a heat map. Our findings revealed that H. pylori infection dramatically up-regulated MAP3K14 (also known as NF-κB-inducing kinase or NIK), while down-regulating IκBα, a classic NF-κB inhibitory protein. Remarkably, supplementation with L. plantarum ZJ316 reversed these gene expression changes, as confirmed by RT-qPCR (Fig. 3A–C).Activation of the classical NF-κB pathway involves phosphorylation of p65 and degradation of IκBα. As shown in Fig. 3D–F, H. pylori infection induced rapid degradation of IκBα within 2 hours, accompanied by increased phosphorylation of p65. In contrast, treatment with L. plantarum ZJ316 effectively countered NF-κB activation by attenuating both IκBα degradation and p65 phosphorylation. Phosphorylated p65 nuclear translocation is crucial for initiating the expression of inflammatory mediators, including IL-8, IL-6, and TNF-α.63 To investigate L. plantarum ZJ316's role in suppressing NF-κB activation with H. pylori infection, we performed an immunofluorescence assay. Our findings showed that in comparison with the control group, H. pylori infection robustly elevated the percentage of cells with nuclear p65, whereas treatment with L. plantarum ZJ316 markedly reduced this nuclear translocation with H. pylori infestation (Fig. 3G and H). The results indicated that L. plantarum ZJ316 achieves its anti-inflammatory effects through modulating the NF-κB signaling pathway, specifically by suppressing classical IκBα degradation and p65 phosphorylation as well as its nuclear translocation.
3.4. BAY 11-7082 increases the anti-inflammatory effects of L. plantarum ZJ316
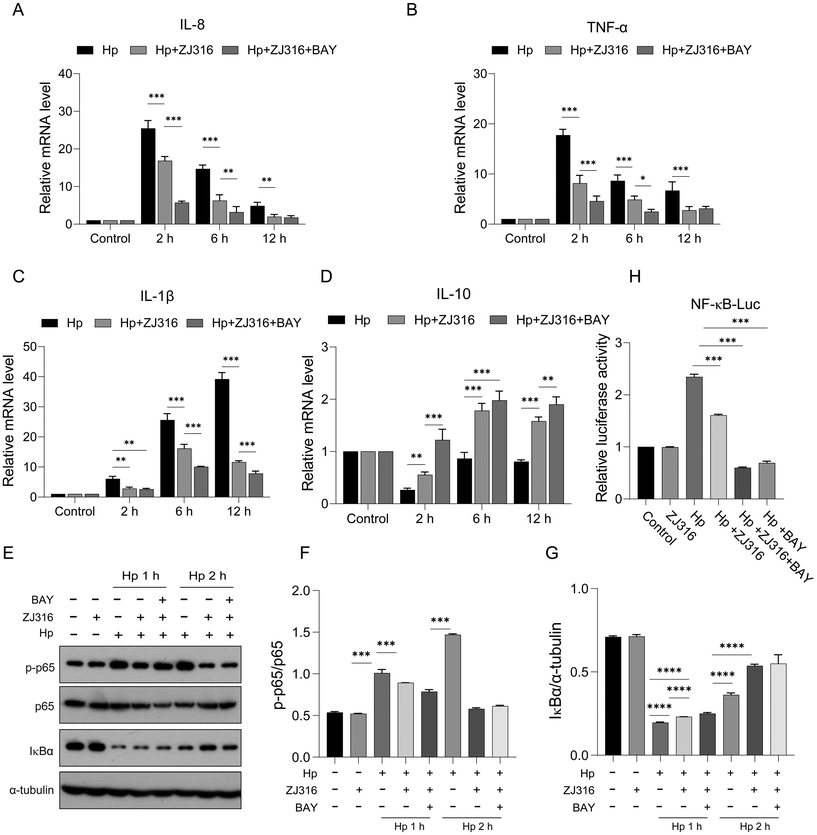
The degradation of IκBα and subsequent release of NF-κB hinge on phosphorylation at Ser32 and Ser36 residues.64,65 BAY 11-7082 selectively and irreversibly impedes IκBα phosphorylation.66 To explore the involvement of NF-κB signaling in the response of L. plantarum ZJ316 to H. pylori-induced inflammation, AGS cells were either left untreated or treated with 10 μM BAY 11-7082. The results indicated that H. pylori infection led to a significant increase in IL-8, TNF-α, and IL-1β levels at 2, 6, and 12 h compared to the control group, whereas treatment with L. plantarum ZJ316 led to a reduction in the levels of these inflammatory markers (Fig. 4A–C). Additionally, treatment with BAY 11-7082 augmented the protective effect of L. plantarum ZJ316 by suppressing IL-8, IL-1β, and TNF-α levels. Conversely, IL-10 levels, recognized for their anti-inflammatory cytokine role, displayed a contrasting trend (Fig. 4D). To delve into the impact of L. plantarum ZJ316 on the downstream targets of NF-κB, we examined IκBα degradation, p65 phosphorylation and NF-κB transcriptional activity. Western blot analysis additionally validated the heightened phosphorylation of p65 and IκBα degradation upon H. pylori stimulation, which was dampened by L. plantarum ZJ316 intervention. Importantly, additional administration of BAY-117082 also reduced these markers compared to the incubation with L. plantarum ZJ316 alone under H. pylori conditions (Fig. 4E–G). To confirm the impact of L. plantarum ZJ316 on the H. pylori-induced activation of NF-κB, we transfected a luciferase reporter controlled by an NF-κB promoter (NF-κB-Luc). The results showed that H. pylori infection triggered NF-κB-dependent transcriptional activation. Notably, NF-κB transcriptional activity was reduced by L. plantarum ZJ316 treatment alone and was significantly further attenuated when L. plantarum ZJ316 was combined with BAY 11-7082 (Fig. 4H). These findings strongly demonstrate that L. plantarum ZJ316 mitigates inflammation following H. pylori infection, partly through the modulation of the NF-κB pathway.3.5. Regulation of NF-κB signaling by L. plantarum ZJ316 in attenuating H. pylori-induced inflammation in vivo
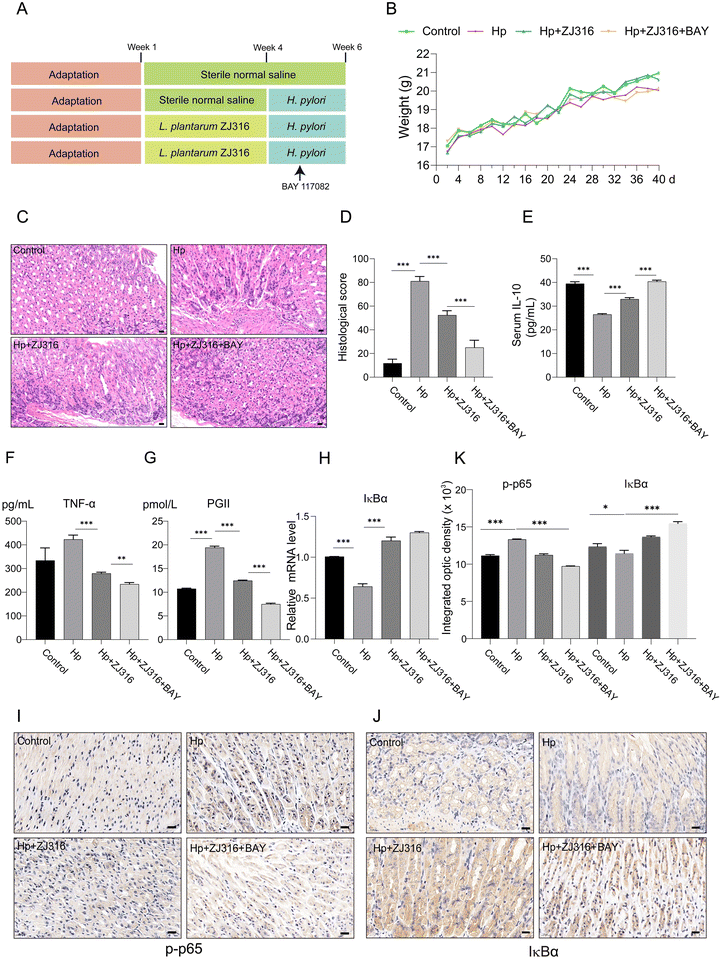
To explore the underlying mechanism by which L. plantarum ZJ316 dampens NF-κB activation in the presence of H. pylori in vivo, C57BL/6 mice were inoculated with H. pylori and received L. plantarum ZJ316 supplementation, with or without BAY 11-7082 (Fig. 5A). Notably, exposure to H. pylori, L. plantarum ZJ316, and BAY 11-7082 did not significantly affect the mouse body weight, as shown in Fig. 5B. Inflammatory molecules are pivotal in the pathogenesis of H. pylori infection, contributing significantly to tissue damage. Then, we evaluated the gastric tissue morphology using H&E staining. The control group exhibited a normal gastric mucosal structure, an intact glandular morphology, and a preserved columnar epithelium (Fig. 5C). Conversely, H. pylori infection induced notable histopathological changes characterized by increased inflammation and infiltration of inflammatory cells, resulting in tissue damage. Treatment with L. plantarum ZJ316 reduced infiltration of inflammatory cells in the infected gastric tissues. Furthermore, inhibition of NF-κB signaling using BAY 11-7082 further mitigated tissue damage compared to the treatment with L. plantarum ZJ316 alone (Fig. 5C). The assessment scores for tissue damage showed consistent results across the experiments (n = 5) (Fig. 5D). Furthermore, a detailed analysis of inflammatory cytokine levels in the mouse serum revealed a notable decrease in IL-10 and a pronounced elevation of TNF-α levels in the H. pylori-infected group, in comparison with the normal control group (Fig. 5E and F), while treatment with L. plantarum ZJ316 reversed these trends by increasing IL-10 levels and reducing TNF-α levels (Fig. 5E and F). Pepsinogens (PGs), enzyme precursors secreted by the gastric mucosa, serve as indicators of atrophic gastritis and are associated with a heightened gastric cancer risk.67,68 Previous studies have associated elevated PG II levels with chronic inflammation in H. pylori-related gastritis and gastric ulcers69,70 (Fig. 5G). Consistent with these findings, our study demonstrated elevated PG II levels following H. pylori infection, which were effectively reversed by treatment with L. plantarum ZJ316. Furthermore, administration of BAY 11-7082 augmented the protective effect of L. plantarum ZJ316 by further lowering PG II and TNF-α levels, while increasing IL-10 levels in serum (Fig. 5E–G).Finally, RT-qPCR and IHC were employed to elucidate the mechanisms underlying L. plantarum ZJ316's inhibition of NF-κB activity. Our results demonstrated that H. pylori infection significantly decreased the IκBα expression at both mRNA and protein levels (Fig. 5H and J), concurrently increasing the expression of phosphorylated p65 (Fig. 5I). Treatment with L. plantarum ZJ316 alone or in combination with BAY 11-7082 further reduced the IκBα levels, while enhancing the phosphorylated p65 expression (Fig. 5H–J), with statistical analysis confirming consistent experimental results (Fig. 5K). These comprehensive findings underscore the potential of L. plantarum ZJ316 in modulating NF-κB signaling in response to H. pylori infection, suggesting its role as a promising therapeutic agent for mitigating inflammation associated with gastrointestinal diseases.
3.6. Effect of L. plantarum ZJ316 on the gastric microbiota in mice with H. pylori infection
Recent studies have highlighted the crucial role of gastric microbes beyond H. pylori in gastric cancer progression. H. pylori infection has been linked to significant alterations in the microbiota composition, particularly in gastric cancer patients.71 In our investigation, we explored the microbial composition and changes in the gastric microbiome in H. pylori-infected mice treated with L. plantarum ZJ316, with or without BAY 11-7082 treatment. At the genus level, Lactobacillus and Prevotella were invariably predominant across all treatment groups. H. pylori treatment notably increased the abundance of Prevotella and Desulfovibrio, while decreasing the levels of Lactobacillus, Akkermansia, Oceanospirillum, and Ligilactobacillus compared to the control group. Treatment with L. plantarum ZJ316 completely mitigated these effects. Furthermore, when comparing H. pylori-infected mice treated with L. plantarum ZJ316 alone to those additionally treated with BAY 11-7082, BAY 11-7082 treatment significantly increased Lactobacillus abundance and reduced Prevotella, thus restoring the gastric microbiome towards a profile more akin to the control group (Fig. 6A). To discern the key taxa driving distinctions among the four groups, we employed a cladogram analysis across the four groups (Fig. 6B). We observed higher abundances of Alistipes, Prevotella, and Desulfovibrio in the Hp group. Conversely, in the Hp + ZJ316 group, Ligilactobacillus were notably more abundant. To elucidate the relationship between the top 15 genera of the gastric microbiota and inflammatory responses, as well as prostaglandins (PGs), we conducted Spearman's correlation analysis. We found that Clostridium strongly correlated with reduced IL-10 levels but increased PG II levels (p < 0.05). Lactobacillus was linked to lower TNF-α and PG II levels (p < 0.05). Additionally, Prevotella showed a significant association with TNF-α (p < 0.05) (Fig. 6C).4. Discussion
H. pylori infection is widespread globally, particularly in emerging nations, where infection rates can exceed 80%.72 Chronic inflammation resulting from H. pylori infestation is a primary driver that contributes to the onset of gastric cancer.12,73 Infected individuals are at a markedly increased risk of developing gastric cancer compared to uninfected individuals. The pathogenicity of H. pylori is mainly attributed to several virulence factors, including lipopolysaccharides, and vacuolating toxins such as VacA and CagA. These factors activate inflammatory signaling pathways such as NF-κB and MAPK.74,75 Extensive research underscores the central role of CagA in H. pylori-triggered inflammation and tumorigenesis, primarily through its activation of the NF-κB transcription factor.74,76–78 This chronic inflammation fosters a microenvironment that promotes the development of cancer-prone cells.79 Quadruple therapy is the established treatment regimen for eradicating H. pylori, yet it confronts significant challenges, including rising drug resistance and potential adverse effects such as gastrointestinal disorders and microbiota imbalance due to antibiotic misuse.80Lactobacillus strains hold promise as a potential solution to these challenges. Earlier research has pointed to a potential relationship between Lactobacillus strains and bolstering host immunity.81–83 Our earlier studies demonstrated that administration of L. plantarum ZJ316 improved gastric microbial balance and effectively reduced the inflammatory reaction triggered by H. pylori infection in mice.54 Nevertheless, the exact mechanisms driving these effects remain elusive. This study elucidates that L. plantarum ZJ316 markedly alleviates inflammation following H. pylori infestation by blocking NF-κB activation, both in vitro and in vivo. Furthermore, supplementation with L. plantarum ZJ316 improves gastric microbial diversity by reducing the abundance of inflammation-associated microbes such as Prevotella and Desulfovibrio, while increasing Ligilactobacillus and Akkermansia, which are linked to anti-inflammatory effects (graphical abstract). These findings suggest that L. plantarum ZJ316 holds promise as a therapeutic agent for alleviating H. pylori-induced inflammation.
Genetic variations in pro-inflammatory cytokine genes, comprising IL-1β and TNF-α, contribute significantly to the risk of gastric cancer.84–86 Specific combinations of polymorphisms in IL-1β, IL-1RN, TNF-α, and IL-10 can elevate the risk of gastric cancer by 27-fold,87 underscoring the pivotal role of these inflammatory cytokines in gastric cancer development.88–90 Research studies have shown that H. pylori infection upregulates the aforementioned inflammatory cytokines.91,92 Consistent with previous findings, our study demonstrated that H. pylori induces TNF-α, IL-1β, IFN-γ, and IL-6 production from co-infected AGS cells at 2 h, 6 h, and 12 h. Furthermore, administration of L. plantarum ZJ316 effectively reduced the production of these inflammatory cytokines induced by H. pylori across all observed time points. Similar findings have been reported for other probiotic Lactobacilli, including Lactobacillus johnsonii,93Ligilactobacillus salivarius,94 and Lactiplantibacillus plantarum,46 which demonstrate the suppression of TNF-α, IL-1β as well as IL-6 secretion or the modulation of gene expressions induced by H. pylori. This study underscores the beneficial impact of L. plantarum ZJ316 on AGS cells, highlighting its potential to mitigate the inflammatory response triggered by H. pylori.
RNA-Seq, an advanced method for profiling the transcriptome using deep-sequencing technologies, has found widespread application in genomic and molecular biology research.95 To delve deeper into how L. plantarum ZJ316 inhibits H. pylori-induced inflammation, we administered L. plantarum ZJ316 and H. pylori to AGS cells for 2 h and employed the RAN-Seq technology to explore the underlying signaling pathways and mechanisms. Our study identified 3897 DEGs between the Hp and Hp + ZJ316 treatment groups, with 1546 genes upregulated and 2351 genes downregulated, highlighting notable changes in the metabolic and transcriptional activities within AGS cells due to the treatments. Subsequent GO and KEGG analyses provided deeper insights into the specific mechanisms through which L. plantarum ZJ316 affects the inflammatory signaling pathways. Specifically, we discovered that L. plantarum ZJ316 treatment attenuated the up-regulation of genes triggered by H. pylori infection, particularly those enriched in the NF-κB signaling pathway and IL-17 signaling pathway according to the KEGG analysis. Notably, NF-κB, a transcription factor involved in inflammation, cell proliferation, and carcinogenesis, remained consistently activated during H. pylori infection96,97 and emerged as a potential target of L. plantarum ZJ316 in mitigating the inflammatory response.
NF-κB, initially bound to IκB proteins in the cytoplasm, undergoes activation upon phosphorylation and degradation of IκB proteins. The phosphorylation of p65 allows the p50–p65 complex to move into the nucleus, enabling NF-κB to bind to specific DNA sequences.98–101 This process facilitates interactions with transcriptional coactivators like p300/CBP, thereby promoting the activation of genes crucial for immune responses, apoptosis, inflammation, and cell survival.98–101 Therefore, inhibiting NF-κB signaling has become a promising approach for diverse medical conditions, given its pivotal role demonstrated in numerous disease studies.102 Yong et al. showed that Lactobacillus acidophilus improved H. pylori-induced gastric inflammation by blocking the Smad7 and NF-κB pathways. Additionally, conditioned media from Ligilactobacillus salivarius B101, Lacticaseibacillus rhamnosus B103, and Lactiplantibacillus plantarum XB7 were identified as protective agents against H. pylori-induced gastric inflammation in AGS cells, achieved through suppression of the NF-κB pathway.52 Our study revealed that probiotic therapy restored the transcriptional levels of certain NF-κB up-regulated genes following H. pylori infection, involving MAP3K14 and IκBα. The western blot analysis definitively showed that L. plantarum ZJ316 reversed NF-κB activation triggered by H. pylori, evidenced by decreased p65 phosphorylation and IκBα degradation in AGS cells. Upon phosphorylation, NF-κB translocates to the nucleus to activate various inflammatory genes. Immunofluorescence analysis confirmed that L. plantarum ZJ316 effectively blocks the H. pylori-induced nuclear translocation of p65, substantiating its inhibition of IκBα degradation and p65 phosphorylation. This mechanism reduces the NF-κB entry into the nucleus and suppresses inflammatory cytokine production. Similar effects were observed with human gastrogenic Weizmannia coagulans BCF-01, which inhibited NF-κB signaling in H. pylori-induced inflammation by blocking IκBα degradation and p65 phosphorylation in macrophages.103 In our study, suppressing the NF-κB activity with BAY 11-7082 demonstrated that treatment with BAY 11-7082 enhances the beneficial effects of L. plantarum ZJ316 on AGS cells against inflammatory damage induced by H. pylori infection. These findings strongly suggest that L. plantarum ZJ316 alleviates inflammation triggered by H. pylori, partly through modulation of the NF-κB pathway. However, the specific regulatory mechanism of L. plantarum ZJ316 on the NF-κB signaling pathway remains unclear. Generally, upon activation of Toll-like or TNF receptors in response to bacterial infection, the IKK complexes (IKKα, IKKβ and IKKγ) are activated,65,101 phosphorylating IκB proteins, which leads to their degradation and the subsequent release of NF-κB dimers.63,65,101 In our study, whether L. plantarum ZJ316 regulates the activation of the IKK complex, thereby regulating IκB phosphorylation and alleviating H. pylori-induced gastric inflammation, requires further investigation. H. pylori infection has been shown to activate STAT3, MAPKs, and AP-1, which interact with NF-κB signaling, forming a complex regulatory network in response to pathogens and inflammatory stimuli.76,104 The Lacticaseibacillus paracasei strain 06TCa19 and BIFICO capsules, which contain a blend of Enterococcus faecalis, Bifidobacterium animalis, and Lactobacillus acidophilus, have been reported to reduce the activation of NF-κB and p38 MAPK signaling pathways.105,106 Additionally, a combination of probiotics such as Lactiplantibacillus plantarum, Lacticaseibacillus rhamnosus, and Lactobacillus acidophilus has the potential to alleviate H. pylori-induced inflammation by upregulating SOCS-2 and SOCS-3 expressions or inhibiting JAK2 activation via the JAK-STAT signaling pathway.107 It is possible that L. plantarum ZJ316 is involved in regulating the MAPK and JAK/STAT pathways to suppress H. pylori-induced inflammation, but further investigation is needed.
Treating mice with H. pylori-induced gastritis using L. plantarum ZJ316 resulted in reduced infiltration of inflammatory cells and decreased secretion of TNF-α, alongside with an elevated IL-10 level compared to mice infected with H. pylori alone. These findings align with prior studies demonstrating the efficacy of probiotics, including Lacticaseibacillus rhamnosus,55,108Ligilactobacillus salivarius55 and Lacticaseibacillus rhamnosus JB3,109 in suppressing H. pylori-induced inflammation in mouse models. PG II, a precursor enzyme secreted by the gastric mucosa, along with serological markers including PG I, PG I/PG II ratio, and gastrin-17, are utilized in epidemiological investigations to assess the risk of gastric cancer.110–113 Gastric inflammation increases the release of both pepsinogens into the bloodstream, with PG II levels showing a more pronounced elevation than PG I.114 Studies consistently showed that infection with CagA-positive H. pylori significantly increases PG II levels and reduces the PG I/PG II ratios.115,116 Furthermore, our findings indicated that H. pylori colonization enhances serum PG II levels, while treatment with L. plantarum ZJ316 decreases PG II levels. Importantly, we observed an additive effect between the L. plantarum ZJ316 treatment group and the L. plantarum ZJ316 + inhibitor group.
Dysbiosis of the gastric microbiota has been linked to the development and advancement of several diseases. Research studies have indicated that H. pylori invasion can alter the composition of microbiota, especially in individuals with gastric cancer.117 Our research indicated that H. pylori colonization predominantly reduces the abundance of beneficial flora, including Lactobacillus, Akkermansia and Ligilactobacillus, while increasing Prevotella and Desulfovibrio, and treatment with L. plantarum ZJ316 reversed these changes by decreasing Prevotella and Desulfovibrio and increasing Ligilactobacillus and Akkermansia. Spearman's correlation analysis revealed that Prevotella showed a notable association with TNF-α. These findings align with previous research highlighting Prevotella as a Gram-negative anaerobic bacterium found in various human body sites including the mouth, respiratory tract, urogenital tract, and gastrointestinal tract, implicated in infections such as oral cavity issues and heart failure.118,119Desulfovibrio has been identified as a causative pathogen of bacteraemia and abdominal infections,120 with its outer membrane vesicles damaging the intestinal epithelial barrier and triggering intrinsic inflammation.121 Additionally, Clostridium was strongly correlated with decreased IL-10 levels but increased PG II levels. In contrast, Lactobacillus was associated with decreased levels of both TNF-α and PG II. Cáceres et al. found that the nutritional form of Clostridium chauvoei triggers an inflammatory response in macrophages.122 Due to the imbalance of the gut microbiota caused by H. pylori, Clostridium difficile can release toxins during intestinal colonization, leading to local tissue damage and severe inflammatory reactions.123 Moreover, Akkermansia has been shown to strengthen the intestinal barrier function and immune response and is inversely associated with inflammatory bowel disease.119,124 These findings imply that supplementation with L. plantarum ZJ316 reshaped the gastric microbiota by decreasing Prevotella and increasing beneficial bacteria such as Lactobacillus104,125 and Ligilactobacillus,126 supporting their protective effects and correlating with a decreased inflammatory response. In this study, we investigated the anti-inflammatory effects of L. plantarum ZJ316 in H. pylori-induced gastric inflammation, focusing primarily on the NF-κB signaling pathway. However, further investigation is needed to fully understand the specific molecular mechanisms by which L. plantarum ZJ316 inhibits NF-κB signaling, particularly its potential effects on the phosphorylation and activation of the IKK complex, a crucial step in the NF-κB pathway.63,65,101 In general, H. pylori infection is known to induce inflammation through various pathways, including STAT3, MAPKs, and AP-1,104,127 and it remains unclear whether L. plantarum ZJ316 also modulates these pathways. Additionally, while L. plantarum ZJ316 has been shown to improve the gastric microbiota disrupted by H. pylori infection, further research is needed to elucidate how these microbiota changes contribute to the observed anti-inflammatory effects. Finally, while the results from our mouse model are promising, clinical studies are essential to validate the effectiveness and safety of the application of L. plantarum ZJ316 in humans. Nonetheless, substantial work remains to be done before these findings can be translated into clinical applications.
5. Conclusions
This study demonstrates that L. plantarum ZJ316 effectively inhibits H. pylori-induced inflammatory responses by targeting the NF-κB signaling pathway. It achieves this by blocking IκBα degradation, preventing p65 phosphorylation, and inhibiting the nuclear translocation of p65. Additionally, L. plantarum ZJ316 reshapes the gastric microbiota by increasing the abundance of beneficial bacteria and decreasing the levels of harmful bacteria, particularly Prevotella, which is positively correlated with the secretion of inflammatory cytokines. These findings highlight the potential of L. plantarum ZJ316 as a probiotic therapy for mitigating H. pylori-induced inflammation and supporting gastrointestinal health. Further research is necessary to optimize dosage, delivery methods, and clinical applications for broader use.Author contributions
Shiying Wu: supervision, writing – review and editing, conceptualization and funding acquisition; Yuenuo Luo: writing – original draft, investigation and formal analysis; Fangtong Wei: investigation and formal analysis; Yanan Li: methodology; Jiayi Fan: data curation; Yang Xu: investigation; Yongqiang Chen: data curation; Wenjie Zhang: methodology; Xuelong Li: methodology; Ziqi Chen: validation; Chenlan Xia: validation; Mingyang Hu: formal analysis; Ping Li: supervision; and Qing Gu: resources, funding acquisition, and project administration.Data availability
The original data presented in the study are included in the article. For additional information, please contact the corresponding author.Conflicts of interest
The authors declare that there are no known competing financial interests or personal relationships that could have influenced the work reported in this paper.Acknowledgements
This work received funding from the National Natural Science Foundation of China (U20A2066), the Key Project of Zhejiang Gongshang University “Digital+” Discipline Construction (SZJ2022A006), the Zhejiang Gongshang University Provincial Colleges and Universities Basic Research Expenses (QRK23006), the Natural Science Foundation of Zhejiang Province (LY22C200012) and the National Natural Science Foundation of China (NSFC) Project (32001667).References
- L. Wang, H. Yao, D. C. Morgan, K. S. Lau, S. Y. Leung, J. W. K. Ho and W. K. Leung, Altered human gut virome in patients undergoing antibiotics therapy for Helicobacter pylori, Nat. Commun., 2023, 14, 2196 CrossRef CAS PubMed.
- Muzaheed, Helicobacter pylori, Oncogenicity: Mechanism, Prevention, and Risk Factors, Sci. World J., 2020, 2020, 3018326 CAS.
- T. L. Cover and R. M. Peek Jr., Diet, microbial virulence, and Helicobacter pylori-induced gastric cancer, Gut Microbes, 2013, 4, 482–493 CrossRef.
- H. C. Sharndama and I. E. Mba, Helicobacter pylori: an up-to-date overview on the virulence and pathogenesis mechanisms, Braz. J. Microbiol., 2022, 53, 33–50 CrossRef CAS PubMed.
- B. B. de Brito, F. A. F. da Silva, A. S. Soares, V. A. Pereira, M. L. C. Santos, M. M. Sampaio, P. H. M. Neves and F. F. de Melo, Pathogenesis and clinical management of Helicobacter pylori gastric infection, World J. Gastroenterol., 2019, 25, 5578–5589 CrossRef CAS PubMed.
- Y. Huang, Q. L. Wang, D. D. Cheng, W. T. Xu and N. H. Lu, Adhesion and Invasion of Gastric Mucosa Epithelial Cells by Helicobacter pylori, Front. Cell. Infect. Microbiol., 2016, 6, 159 Search PubMed.
- K. Moonens, Y. Hamway, M. Neddermann, M. Reschke, N. Tegtmeyer, T. Kruse, R. Kammerer, R. Mejias-Luque, B. B. Singer, S. Backert, M. Gerhard and H. Remaut, Helicobacter pylori adhesin HopQ disrupts trans dimerization in human CEACAMs, EMBO J., 2018, 37(13), e98665 CrossRef PubMed.
- Q. Sun, C. Yuan, S. Zhou, J. Lu, M. Zeng, X. Cai and H. Song, Helicobacter pylori infection: a dynamic process from diagnosis to treatment, Front. Cell. Infect. Microbiol., 2023, 13, 1257817 CrossRef CAS PubMed.
- S. C. Tran, K. N. Bryant and T. L. Cover, The Helicobacter pylori cag pathogenicity island as a determinant of gastric cancer risk, Gut Microbes, 2024, 16, 2314201 CrossRef PubMed.
- Y. Duan, Y. Xu, Y. Dou and D. Xu, Helicobacter pylori and gastric cancer: mechanisms and new perspectives, J. Hematol. Oncol., 2025, 18, 10 CrossRef PubMed.
- P. I. Seoane, B. Lee, C. Hoyle, S. Yu, G. Lopez-Castejon, M. Lowe and D. Brough, The NLRP3-inflammasome as a sensor of organelle dysfunction, J. Cell Biol., 2020, 219(12), e202006194 CrossRef CAS PubMed.
- H. Yang and B. Hu, Immunological Perspective: Helicobacter pylori Infection and Gastritis, Mediators Inflammation, 2022, 2022, 2944156 Search PubMed.
- Q. Yu, H. Shi, Z. Ding, Z. Wang, H. Yao and R. Lin, The E3 ubiquitin ligase TRIM31 attenuates NLRP3 inflammasome activation in Helicobacter pylori-associated gastritis by regulating ROS and autophagy, Cell Commun. Signaling, 2023, 21, 1 CrossRef CAS PubMed.
- D. Kidane, D. L. Murphy and J. B. Sweasy, Accumulation of abasic sites induces genomic instability in normal human gastric epithelial cells during Helicobacter pylori infection, Oncogenesis, 2014, 3, e128 CrossRef CAS PubMed.
- T. Baccili Cury Megid, A. R. Farooq, X. Wang and E. Elimova, Gastric Cancer: Molecular Mechanisms, Novel Targets, and Immunotherapies: From Bench to Clinical Therapeutics, Cancers, 2023, 15, 5075 CrossRef PubMed.
- A. Lamb and L. F. Chen, Role of the Helicobacter pylori-induced inflammatory response in the development of gastric cancer, J. Cell. Biochem., 2013, 114, 491–497 CrossRef CAS PubMed.
- W. Xu, Y. Huang, Z. Yang, Y. Hu, X. Shu, C. Xie, C. He, Y. Zhu and N. Lu, Helicobacter pylori promotes gastric epithelial cell survival through the PLK1/PI3K/Akt pathway, OncoTargets Ther., 2018, 11, 5703–5713 CrossRef CAS PubMed.
- X. Y. Zhang, P. Y. Zhang and M. A. Aboul-Soud, From inflammation to gastric cancer: Role of Helicobacter pylori, Oncol. Lett., 2017, 13, 543–548 CrossRef CAS PubMed.
- L. Xia, S. Tan, Y. Zhou, J. Lin, H. Wang, L. Oyang, Y. Tian, L. Liu, M. Su, H. Wang, D. Cao and Q. Liao, Role of the NFkappaB-signaling pathway in cancer, OncoTargets Ther., 2018, 11, 2063–2073 CrossRef PubMed.
- M. Alipour, Molecular Mechanism of Helicobacter pylori-Induced Gastric Cancer, J. Gastrointest. Cancer, 2021, 52, 23–30 CAS.
- X. Cui, M. Chang, Y. Wang, J. Liu, Z. Sun, Q. Sun, Y. Sun, J. Ren and W. Li, Helicobacter pylori reduces METTL14-mediated VAMP3 m(6)A modification and promotes the development of gastric cancer by regulating LC3C-mediated c-Met recycling, Cell Death Discovery, 2025, 11, 13 CAS.
- T. Liu, L. Zhang, D. Joo and S.-C. Sun, NF-κB signaling in inflammation, Signal Transduction Targeted Ther., 2017, 2, 17023 Search PubMed.
- F. Yang, Y. Xu, C. Liu, C. Ma, S. Zou, X. Xu, J. Jia and Z. Liu, NF-κB/miR-223–3p/ARID1A axis is involved in Helicobacter pylori CagA-induced gastric carcinogenesis and progression, Cell Death Dis., 2018, 9(1), 12 Search PubMed.
- B. Schmidinger, K. Petri, C. Lettl, H. Li, S. Namineni, H. Ishikawa-Ankerhold, L. F. Jimenez-Soto and R. Haas, Helicobacter pylori binds human Annexins via Lipopolysaccharide to interfere with Toll-like Receptor 4 signaling, PLoS Pathog., 2022, 18, e1010326 CAS.
- J.-H. Wong, K.-H. Ho, S. Nam, W.-L. Hsu, C.-H. Lin, C.-M. Chang, J.-Y. Wang and W.-C. Chang, Store-operated Ca2+ Entry Facilitates the Lipopolysaccharide-induced Cyclooxygenase-2 Expression in Gastric Cancer Cells, Sci. Rep., 2017, 7(1), 12813 Search PubMed.
- S. K. Pachathundikandi, J. Lind, N. Tegtmeyer, E. M. El-Omar and S. Backert, Interplay of the Gastric Pathogen Helicobacter pylori with Toll-Like Receptors, BioMed Res. Int., 2015, 2015, 192420 Search PubMed.
- J. Yan, Q. Xue, W. Chen, K. Wang, D. Peng, J. Jiang, P. Li and B. Du, Probiotic-fermented rice buckwheat alleviates high-fat diet-induced hyperlipidemia in mice by suppressing lipid accumulation and modulating gut microbiota, Food Res. Int., 2022, 155, 111125 CAS.
- L. Llorca, G. Perez-Perez, P. Urruzuno, M. J. Martinez, T. Iizumi, Z. Gao, J. Sohn, J. Chung, L. Cox, A. Simon-Soro, A. Mira and T. Alarcon, Characterization of the Gastric Microbiota in a Pediatric Population According to Helicobacter pylori Status, Pediatr. Infect. Dis. J., 2017, 36, 173–178 CrossRef PubMed.
- R. Miao, C. Wan and Z. Wang, The relationship of gastric microbiota and Helicobacter pylori infection in pediatrics population, Helicobacter, 2020, 25, e12676 CAS.
- R. Vasapolli, K. Schutte, C. Schulz, M. Vital, D. Schomburg, D. H. Pieper, R. Vilchez-Vargas and P. Malfertheiner, Analysis of Transcriptionally Active Bacteria Throughout the Gastrointestinal Tract of Healthy Individuals, Gastroenterology, 2019, 157, 1081–1092. e1083 CrossRef CAS PubMed.
- K. Fukase, M. Kato, S. Kikuchi, K. Inoue, N. Uemura, S. Okamoto, S. Terao, K. Amagai, S. Hayashi, M. Asaka and Japan Gast Study Group, Effect of eradication of Helicobacter pylori on incidence of metachronous gastric carcinoma after endoscopic resection of early gastric cancer: an open-label, randomised controlled trial, Lancet, 2008, 372, 392–397 Search PubMed.
- S. B. Yoon, J. M. Park, C. H. Lim, Y. K. Cho and M. G. Choi, Effect of Helicobacter pylori eradication on metachronous gastric cancer after endoscopic resection of gastric tumors: a meta-analysis, Helicobacter, 2014, 19, 243–248 CrossRef.
- Y. C. Lee, T. H. Chiang, C. K. Chou, Y. K. Tu, W. C. Liao, M. S. Wu and D. Y. Graham, Association Between Helicobacter pylori Eradication and Gastric Cancer Incidence: A Systematic Review and Meta-analysis, Gastroenterology, 2016, 150, 1113–1124. e1115 CrossRef PubMed.
- A. Savoldi, E. Carrara, D. Y. Graham, M. Conti and E. Tacconelli, Prevalence of Antibiotic Resistance in Helicobacter pylori: A Systematic Review and Meta-analysis in World Health Organization Regions, Gastroenterology, 2018, 155, 1372–1382. e1317 CrossRef.
- N. Farzi, A. Yadegar, A. Sadeghi, H. Asadzadeh Aghdaei, S. Marian Smith, J. Raymond, H. Suzuki and M. R. Zali, High Prevalence of Antibiotic Resistance in Iranian Helicobacter pylori Isolates: Importance of Functional and Mutational Analysis of Resistance Genes and Virulence Genotyping, J. Clin. Med., 2019, 8(11), 2004 CrossRef CAS PubMed.
- Z. Sadeghloo, P. Saffarian, M. Hakemi-Vala, A. Sadeghi and A. Yadegar, The modulatory effect of Lactobacillus gasseri ATCC 33323 on autophagy induced by extracellular vesicles of Helicobacter pylori in gastric epithelial cells in vitro, Microb. Pathog., 2024, 188, 106559 CrossRef CAS PubMed.
- S. Suzuki, M. Esaki, C. Kusano, H. Ikehara and T. Gotoda, Development of Helicobacter pylori treatment: How do we manage antimicrobial resistance?, World J. Gastroenterol., 2019, 25, 1907–1912 CrossRef CAS PubMed.
- A. Nabavi-Rad, A. Sadeghi, H. Asadzadeh Aghdaei, A. Yadegar, S. M. Smith and M. R. Zali, The double-edged sword of probiotic supplementation on gut microbiota structure in Helicobacter pylori management, Gut Microbes, 2022, 14, 2108655 CrossRef PubMed.
- P. Malfertheiner, F. Megraud, T. Rokkas, J. P. Gisbert, J. M. Liou, C. Schulz, A. Gasbarrini, R. H. Hunt, M. Leja, C. O’Morain, M. Rugge, S. Suerbaum, H. Tilg, K. Sugano, E. M. El-Omar and European Helicobacter and Microbiota Study group, Management of Helicobacter pylori infection: the Maastricht VI/Florence consensus report, Gut, 2022, 71, 1724–1762 CrossRef CAS PubMed.
- H. Y. Song, L. Zhou, D. Y. Liu, X. J. Yao and Y. Li, What Roles Do Probiotics Play in the Eradication of Helicobacter pylori? Current Knowledge and Ongoing Research, Gastroenterol. Res. Pract., 2018, 2018, 9379480 Search PubMed.
- N. I. Ismail, K. N. M. Nawawi, D. C. C. Hsin, K. W. Hao, N. Mahmood, G. L. C. Chearn, Z. Wong, A. M. Tamil, H. Joseph and R. A. Raja Ali, Probiotic containing Lactobacillus reuteri DSM 17648 as an adjunct treatment for Helicobacter pylori, infection: A randomized, double-blind, placebo-controlled trial, Helicobacter, 2023, 28, e13017 CrossRef CAS PubMed.
- H. R. Fang, G. Q. Zhang, J. Y. Cheng and Z. Y. Li, Efficacy of Lactobacillus-supplemented triple therapy for Helicobacter pylori infection in children: a meta-analysis of randomized controlled trials, Eur. J. Pediatr., 2019, 178, 7–16 CrossRef CAS.
- D. Dash, V. Mishra, M. K. Panda and S. K. Pathak, Effects of Lactobacillus spp. on Helicobacter pylori: A Promising Frontier in the Era of Antibiotic Resistance, Probiotics Antimicrob. Proteins, 2024 DOI:10.1007/s12602-024-10396-z.
- Z. Zhang, J. Lv, L. Pan and Y. Zhang, Roles and applications of probiotic Lactobacillus strains, Appl. Microbiol. Biotechnol., 2018, 102, 8135–8143 CrossRef CAS PubMed.
- H. Maleki-Kakelar, J. Dehghani, A. Barzegari, J. Barar, M. Shirmohamadi, J. Sadeghi and Y. Omidi, Lactobacillus plantarum induces apoptosis in gastric cancer cells via modulation of signaling pathways in Helicobacter pylori, Bioimpacts, 2020, 10, 65–72 CrossRef CAS PubMed.
- K. Zhao, Q. Xie, D. Xu, Y. Guo, X. Tao, H. Wei and C. Wan, Antagonistics of Lactobacillus plantarum ZDY2013 against Helicobacter pylori SS1 and its infection in vitro in human gastric epithelial AGS cells, J. Biosci. Bioeng., 2018, 126, 458–463 CrossRef CAS PubMed.
- V. H. Urrutia-Baca, E. Escamilla-Garcia, M. A. de la Garza-Ramos, P. Tamez-Guerra, R. Gomez-Flores and C. S. Urbina-Rios, In Vitro Antimicrobial Activity and Downregulation of Virulence Gene Expression on Helicobacter pylori by Reuterin, Probiotics Antimicrob. Proteins, 2018, 10, 168–175 CAS.
- N. de Klerk, L. Maudsdotter, H. Gebreegziabher, S. D. Saroj, B. Eriksson, O. S. Eriksson, S. Roos, S. Linden, H. Sjolinder and A. B. Jonsson, Lactobacilli Reduce Helicobacter pylori Attachment to Host Gastric Epithelial Cells by Inhibiting Adhesion Gene Expression, Infect. Immun., 2016, 84, 1526–1535 CAS.
- M. Gotteland, O. Brunser and S. Cruchet, Systematic review: are probiotics useful in controlling gastric colonization by Helicobacter pylori?, Aliment. Pharmacol. Ther., 2006, 23, 1077–1086 CAS.
- F. Fakharian, B. Asgari, A. Nabavi-Rad, A. Sadeghi, N. Soleimani, A. Yadegar and M. R. Zali, The interplay between Helicobacter pylori and the gut microbiota: An emerging driver influencing the immune system homeostasis and gastric carcinogenesis, Front. Cell. Infect. Microbiol., 2022, 12, 953718 CrossRef CAS PubMed.
- Y. J. Yang, C. C. Chuang, H. B. Yang, C. C. Lu and B. S. Sheu, Lactobacillus acidophilus ameliorates H. pylori-induced gastric inflammation by inactivating the Smad7 and NFkappaB pathways, BMC Microbiol., 2012, 12, 38 CAS.
- T. Thiraworawong, J. K. Spinler, D. Werawatganon, N. Klaikeaw, S. F. Venable, J. Versalovic and S. Tumwasorn, Anti-inflammatory properties of gastric-derived Lactobacillus plantarum XB7 in the context of Helicobacter pylori infection, Helicobacter, 2014, 19, 144–155 CAS.
- S. Wu, Y. Xu, Z. Chen, Y. Chen, F. Wei, C. Xia, Q. Zhou, P. Li and Q. Gu, Lactiplantibacillus plantarum ZJ316 Reduces Helicobacter pylori Adhesion and Inflammation by Inhibiting the Expression of Adhesin and Urease Genes, Mol. Nutr. Food Res., 2023, 67, e2300241 Search PubMed.
- Q. Zhou, B. Xue, R. Gu, P. Li and Q. Gu, Lactobacillus plantarum ZJ316 Attenuates Helicobacter pylori-Induced Gastritis in C57BL/6 Mice, J. Agric. Food Chem., 2021, 69, 6510–6523 CAS.
- C. He, C. Peng, X. Xu, N. Li, Y. Ouyang, Y. Zhu and N. Lu, Probiotics mitigate Helicobacter pylori-induced gastric inflammation and premalignant lesions in INS-GAS mice with the modulation of gastrointestinal microbiota, Helicobacter, 2022, 27, e12898 CAS.
- Z. Ren, Z. Xu, X. Chang, J. Liu and W. Xiao, STC1 competitively binding betaPIX enhances melanoma progression via YAP nuclear translocation and M2 macrophage recruitment through the YAP/CCL2/VEGFA/AKT feedback loop, Pharmacol. Res., 2024, 204, 107218 CAS.
- Z. Zhai, K. M. Niu, Y. Liu, C. Lin and X. Wu, The Gut Microbiota-Bile Acids-TGR5 Axis Mediates Eucommia ulmoides Leaf Extract Alleviation of Injury to Colonic Epithelium Integrity, Front. Microbiol., 2021, 12, 727681 Search PubMed.
- Z. Yu, M. Cao, J. Peng, D. Wu, S. Li, C. Wu, L. Qing, A. Zhang, W. Wang, M. Huang and J. Zhao, Lacticaseibacillus casei, T1 attenuates Helicobacter pylori-induced inflammation and gut microbiota disorders in mice, BMC Microbiol., 2023, 23, 39 CAS.
- S. Shishodia, H. M. Amin, R. Lai and B. B. Aggarwal, Curcumin (diferuloylmethane) inhibits constitutive NF-kappaB activation, induces G1/S arrest, suppresses proliferation, and induces apoptosis in mantle cell lymphoma, Biochem. Pharmacol., 2005, 70, 700–713 CAS.
- S. W. Kim, D. W. Oleksyn, R. M. Rossi, C. T. Jordan, I. Sanz, L. Chen and J. Zhao, Protein kinase C-associated kinase is required for NF-kappaB signaling and survival in diffuse large B-cell lymphoma cells, Blood, 2008, 111, 1644–1653 CAS.
- M. Karin and F. R. Greten, NF-kappaB: linking inflammation and immunity to cancer development and progression, Nat. Rev. Immunol., 2005, 5, 749–759 CrossRef CAS.
- A. Cras, B. Politis, N. Balitrand, D. Darsin-Bettinger, P. Y. Boelle, B. Cassinat, M. E. Toubert and C. Chomienne, Bexarotene via CBP/p300 induces suppression of NF-kappaB-dependent cell growth and invasion in thyroid cancer, Clin. Cancer Res., 2012, 18, 442–453 CrossRef CAS PubMed.
- B. Wang and J. Shen, NF-kappaB Inducing Kinase Regulates Intestinal Immunity and Homeostasis, Front. Immunol., 2022, 13, 895636 CrossRef CAS PubMed.
- S. Yazdi, M. Naumann and M. Stein, Double phosphorylation-induced structural changes in the signal-receiving domain of IkappaBalpha in complex with NF-kappaB, Proteins, 2017, 85, 17–29 CrossRef CAS PubMed.
- R. J. Antonia, R. S. Hagan and A. S. Baldwin, Expanding the View of IKK: New Substrates and New Biology, Trends Cell Biol., 2021, 31, 166–178 CrossRef CAS PubMed.
- J. W. Pierce, R. Schoenleber, G. Jesmok, J. Best, S. A. Moore, T. Collins and M. E. Gerritsen, Novel inhibitors of cytokine-induced IkappaBalpha phosphorylation and endothelial cell adhesion molecule expression show anti-inflammatory effects in vivo, J. Biol. Chem., 1997, 272, 21096–21103 CrossRef CAS PubMed.
- Y. Tong, H. Wang, Y. Zhao, X. He, H. Xu, H. Li, P. Shuai, L. Gong, H. Wu, H. Xu, Y. Luo, D. Wang, S. Liu and Z. Song, Diagnostic Value of Serum Pepsinogen Levels for Screening Gastric Cancer and Atrophic Gastritis in Asymptomatic Individuals: A Cross-Sectional Study, Front. Oncol., 2021, 11, 652574 CrossRef CAS PubMed.
- T. Kotachi, M. Ito, M. Yoshihara, T. Boda, M. Kiso, K. Masuda, T. Matsuo, S. Tanaka and K. Chayama, Serological Evaluation of Gastric Cancer Risk Based on Pepsinogen and Helicobacter pylori Antibody: Relationship to Endoscopic Findings, Digestion, 2017, 95, 314–318 CAS.
- N. Chapelle, J. Martin, M. Osmola, C. Hemont, M. Leroy, M. A. Vibet, D. Tougeron, D. Moussata, D. Lamarque, E. Bigot-Corbel, D. Masson, J. Blin, R. Josien, J. F. Mosnier and T. Matysiak-Budnik, Serum pepsinogens can help to discriminate between H. pylori-induced and auto-immune atrophic gastritis: Results from a prospective multicenter study, Dig. Liver Dis., 2023, 55, 1345–1351 CrossRef CAS PubMed.
- K. Muhsen, R. Sinnreich, D. Merom, H. Nassar, D. Cohen and J. D. Kark, Helicobacter pylori infection, serum pepsinogens as markers of atrophic gastritis, and leukocyte telomere length: a population-based study, Hum. Genomics, 2019, 13, 32 CrossRef PubMed.
- J. M. Noto and R. M. Peek Jr., The gastric microbiome, its interaction with Helicobacter pylori, and its potential role in the progression to stomach cancer, PLoS Pathog., 2017, 13, e1006573 CrossRef PubMed.
- L. Engstrand and D. Y. Graham, Microbiome and Gastric Cancer, Dig. Dis. Sci., 2020, 65, 865–873 CrossRef CAS PubMed.
- S. Salvatori, I. Marafini, F. Laudisi, G. Monteleone and C. Stolfi, Helicobacter pylori, and Gastric Cancer: Pathogenetic Mechanisms, Int. J. Mol. Sci., 2023, 24(3), 2895 CrossRef CAS PubMed.
- B. Chen, X. Liu, P. Yu, F. Xie, J. S. H. Kwan, W. N. Chan, C. Fang, J. Zhang, A. H. K. Cheung, C. Chow, G. W. M. Leung, K. T. Leung, S. Shi, B. Zhang, S. Wang, D. Xu, K. Fu, C. C. Wong, W. K. K. Wu, M. W. Y. Chan, P. M. K. Tang, C. M. Tsang, K. W. Lo, G. M. K. Tse, J. Yu, K. F. To and W. Kang, H. pylori-induced NF-kappaB-PIEZO1-YAP1-CTGF axis drives gastric cancer progression and cancer-associated fibroblast-mediated tumour microenvironment remodelling, Clin. Transl. Med., 2023, 13, e1481 CrossRef CAS PubMed.
- M. Yang, L. Wang, L. J. Gu and W. J. Yuan, Helicobacter pylori cytotoxin-associated gene A impairs the filtration barrier function of podocytes via p38 MAPK signaling pathway, Acta Biochim. Pol., 2017, 64, 471–475 CrossRef CAS PubMed.
- F. Yang, Y. Xu, C. Liu, C. Ma, S. Zou, X. Xu, J. Jia and Z. Liu, NF-kappaB/miR-223–3p/ARID1A axis is involved in Helicobacter pylori CagA-induced gastric carcinogenesis and progression, Cell Death Dis., 2018, 9, 12 CrossRef PubMed.
- B. E. Lee, S. J. Park, G. H. Kim, D. C. Joo and M. W. Lee, Anti-inflammatory effects of eupatilin on Helicobacter pylori CagA-induced gastric inflammation, PLoS One, 2024, 19, e0313251 CrossRef CAS PubMed.
- Y. Wen, H. Huang, T. Tang, H. Yang, X. Wang, X. Huang, Y. Gong, X. Zhang and F. She, AI-2 represses CagA expression and bacterial adhesion, attenuating the Helicobacter pylori-induced inflammatory response of gastric epithelial cells, Helicobacter, 2021, 26, e12778 CrossRef CAS PubMed.
- A. Mantovani, P. Allavena, A. Sica and F. Balkwill, Cancer-related inflammation, Nature, 2008, 454, 436–444 CrossRef CAS PubMed.
- Y. Hu, Y. Zhu and N. H. Lu, Novel and Effective Therapeutic Regimens for Helicobacter pylori, in an Era of Increasing Antibiotic Resistance, Front. Cell. Infect. Microbiol., 2017, 7, 168 CrossRef PubMed.
- C. Li, K. Peng, S. Xiao, Y. Long and Q. Yu, The role of Lactobacillus in inflammatory bowel disease: from actualities to prospects, Cell Death Discovery, 2023, 9, 361 CrossRef PubMed.
- A. Shapouri-Moghaddam, S. Mohammadian, H. Vazini, M. Taghadosi, S. A. Esmaeili, F. Mardani, B. Seifi, A. Mohammadi, J. T. Afshari and A. Sahebkar, Macrophage plasticity, polarization, and function in health and disease, J. Cell Physiol., 2018, 233, 6425–6440 CrossRef CAS PubMed.
- N. Erfanian, S. Nasseri, A. Miraki Feriz, H. Safarpour and M. H. Namaie, Characterization of Wnt signaling pathway under treatment of Lactobacillus acidophilus postbiotic in colorectal cancer using an integrated in silico and in vitro analysis, Sci. Rep., 2023, 13, 22988 CrossRef CAS PubMed.
- E. M. El-Omar, M. Carrington, W. H. Chow, K. E. McColl, J. H. Bream, H. A. Young, J. Herrera, J. Lissowska, C. C. Yuan, N. Rothman, G. Lanyon, M. Martin, J. F. Fraumeni Jr. and C. S. Rabkin, Interleukin-1 polymorphisms associated with increased risk of gastric cancer, Nature, 2000, 404, 398–402 CAS.
- E. M. El-Omar, C. S. Rabkin, M. D. Gammon, T. L. Vaughan, H. A. Risch, J. B. Schoenberg, J. L. Stanford, S. T. Mayne, J. Goedert, W. J. Blot, J. F. Fraumeni Jr. and W. H. Chow, Increased risk of noncardia gastric cancer associated with proinflammatory cytokine gene polymorphisms, Gastroenterology, 2003, 124, 1193–1201 CrossRef CAS PubMed.
- J. C. Machado, C. Figueiredo, P. Canedo, P. Pharoah, R. Carvalho, S. Nabais, C. Castro Alves, M. L. Campos, L. J. Van Doorn, C. Caldas, R. Seruca, F. Carneiro and M. Sobrinho-Simoes, A proinflammatory genetic profile increases the risk for chronic atrophic gastritis and gastric carcinoma, Gastroenterology, 2003, 125, 364–371 CAS.
- H. Oshima, T. Ishikawa, G. J. Yoshida, K. Naoi, Y. Maeda, K. Naka, X. Ju, Y. Yamada, T. Minamoto, N. Mukaida, H. Saya and M. Oshima, TNF-α/TNFR1 signaling promotes gastric tumorigenesis through induction of Noxo1 and Gna14 in tumor cells, Oncogene, 2013, 33, 3820–3829 CrossRef PubMed.
- M. C. Rosu, P. D. Mihnea, A. Ardelean, S. D. Moldovan, R. O. Popetiu and B. D. Totolici, Clinical significance of tumor necrosis factor-alpha and carcinoembryonic antigen in gastric cancer, J. Med. Life, 2022, 15, 4–6 CrossRef PubMed.
- Y. Ma, Y. Fu, X. Fan, Q. Ji, X. Duan, Y. Wang, Y. Zhang, Z. Wang and H. Hao, FAK/IL-8 axis promotes the proliferation and migration of gastric cancer cells, Gastric Cancer, 2023, 26, 528–541 CrossRef CAS PubMed.
- X. Y. Yuan, Y. Zhang, X. Zhao, A. Chen and P. Liu, IL-1beta, an important cytokine affecting Helicobacter pylori-mediated gastric carcinogenesis, Microb. Pathog., 2023, 174, 105933 CAS.
- N. R. Lim and W. C. Chung, Helicobacter pylori-associated Chronic Atrophic Gastritis and Progression of Gastric Carcinogenesis, Korean J. Gastroenterol., 2023, 82, 171–179 Search PubMed.
- K. A. Bockerstett and R. J. DiPaolo, Regulation of Gastric Carcinogenesis by Inflammatory Cytokines, Cell. Mol. Gastroenterol. Hepatol., 2017, 4, 47–53 CrossRef PubMed.
- S. Shen, F. Ren, H. Qin, I. Bukhari, J. Yang, D. Gao, A. C. Ouwehand, M. J. Lehtinen, P. Zheng and Y. Mi, Lactobacillus acidophilus NCFM and Lactiplantibacillus plantarum Lp-115 inhibit Helicobacter pylori colonization and gastric inflammation in a murine model, Front. Cell. Infect. Microbiol., 2023, 13, 1196084 Search PubMed.
- W. Panpetch, J. K. Spinler, J. Versalovic and S. Tumwasorn, Characterization of Lactobacillus salivarius strains, B37 and B60 capable of inhibiting IL-8 production in Helicobacter pylori-stimulated gastric epithelial cells, BMC Microbiol., 2016, 16, 242 CrossRef PubMed.
- Z. Wang, M. Gerstein and M. Snyder, RNA-Seq: a revolutionary tool for transcriptomics, Nat. Rev. Genet., 2009, 10, 57–63 CrossRef CAS PubMed.
- H. Yu, L. Lin, Z. Zhang, H. Zhang and H. Hu, Targeting NF-kappaB pathway for the therapy of diseases: mechanism and clinical study, Signal Transduction Targeted Ther., 2020, 5, 209 CrossRef CAS PubMed.
- M. Chen, X. Huang, M. Gao, Z. Yang, Z. Fang, J. Wei and B. Wu, Helicobacter pylori promotes inflammatory factor secretion and lung injury through VacA exotoxin-mediated activation of NF-kappaB signaling, Bioengineered, 2022, 13, 12760–12771 Search PubMed.
- I. M. Verma, J. K. Stevenson, E. M. Schwarz, D. Van Antwerp and S. Miyamoto, Rel/NF-kappa B/I kappa B family: intimate tales of association and dissociation, Genes Dev., 1995, 9, 2723–2735 CrossRef CAS PubMed.
- S. Ghosh, M. J. May and E. B. Kopp, NF-kappa B and Rel proteins: evolutionarily conserved mediators of immune responses, Annu. Rev. Immunol., 1998, 16, 225–260 Search PubMed.
- G. Natoli, S. Saccani, D. Bosisio and I. Marazzi, Interactions of NF-kappaB with chromatin: the art of being at the right place at the right time, Nat. Immunol., 2005, 6, 439–445 Search PubMed.
- N. Suzuki, N. Murata-Kamiya, K. Yanagiya, W. Suda, M. Hattori, H. Kanda, A. Bingo, Y. Fujii, S. Maeda, K. Koike and M. Hatakeyama, Mutual reinforcement of inflammation and carcinogenesis by the Helicobacter pylori CagA oncoprotein, Sci. Rep., 2015, 5, 10024 Search PubMed.
- P. Poma, NF-kappaB and Disease, Int. J. Mol. Sci., 2020, 21(23), 9181 CrossRef PubMed.
- Z. Chen, Z. Tang, W. Li, X. Deng, L. Yu, J. Yang, J. Liu, Y. Cheng, W. Huang, X. Guo, J. Shan, D. Zhou, W. Zeng, Y. Bai and H. Fan, Weizmannia coagulans BCF-01: a novel gastrogenic probiotic for Helicobacter pylori infection control, Gut Microbes, 2024, 16, 2313770 CrossRef PubMed.
- M. Kandpal, B. Baral, N. Varshney, A. K. Jain, D. Chatterji, A. K. Meena, R. K. Pandey and H. C. Jha, Gut-brain axis interplay via STAT3 pathway: Implications of Helicobacter pylori derived secretome on inflammation and Alzheimer's disease, Virulence, 2024, 15, 2303853 CrossRef PubMed.
- S. Takeda, K. Igoshi, C. Tsend-Ayush, T. Oyunsuren, R. Sakata, Y. Koga, Y. Arima and M. Takeshita, Lactobacillus paracasei strain 06TCa19 suppresses inflammatory chemokine induced by Helicobacter pylori in human gastric epithelial cells, Hum. Cell, 2017, 30, 258–266 CrossRef CAS PubMed.
- H. J. Yu, W. Liu, Z. Chang, H. Shen, L. J. He, S. S. Wang, L. Liu, Y. Y. Jiang, G. T. Xu, M. M. An and J. D. Zhang, Probiotic BIFICO cocktail ameliorates Helicobacter pylori induced gastritis, World J. Gastroenterol., 2015, 21, 6561–6571 Search PubMed.
- J. S. Lee, N. S. Paek, O. S. Kwon and K. B. Hahm, Anti-inflammatory actions of probiotics through activating suppressor of cytokine signaling (SOCS) expression and signaling in Helicobacter pylori infection: a novel mechanism, J. Gastroenterol. Hepatol., 2010, 25, 194–202 Search PubMed.
- Y. H. Chen, W. H. Tsai, H. Y. Wu, C. Y. Chen, W. L. Yeh, Y. H. Chen, H. Y. Hsu, W. W. Chen, Y. W. Chen, W. W. Chang, T. L. Lin, H. C. Lai, Y. H. Lin and C. H. Lai, Probiotic Lactobacillus spp. act Against Helicobacter pylori-induced Inflammation, J. Clin. Med., 2019, 8(1), 90 CAS.
- M. E. Chen, C. H. Su, J. S. Yang, C. C. Lu, Y. C. Hou, J. B. Wu and Y. M. Hsu, Baicalin, Baicalein, and Lactobacillus Rhamnosus JB3 Alleviated Helicobacter pylori Infections in Vitro and in Vivo, J. Food Sci., 2018, 83, 3118–3125 CAS.
- T. H. Chiang, S. Y. Chiu, S. L. Chen, A. M. Yen, J. C. Fann, C. Y. Liu, C. K. Chou, H. M. Chiu, C. T. Shun, M. S. Wu, J. T. Lin, Y. C. Lee, T. H. Chen and M. W. Lin, Serum Pepsinogen as a Predictor for Gastric Cancer Death: A 16-Year Community-based Cohort Study, J. Clin. Gastroenterol., 2019, 53, e186–e193 CAS.
- T. Kudo, S. Kakizaki, N. Sohara, Y. Onozato, S. Okamura, Y. Inui and M. Mori, Analysis of ABC (D) stratification for screening patients with gastric cancer, World J. Gastroenterol., 2011, 17, 4793–4798 Search PubMed.
- A. G. McNicholl, M. Forne, J. Barrio, C. De la Coba, B. Gonzalez, R. Rivera, M. Esteve, F. Fernandez-Banares, B. Madrigal, B. Gras-Miralles, A. Perez-Aisa, J. M. Viver-Pi-Sunyer, F. Bory, M. Rosinach, C. Loras, C. Esteban, S. Santolaria, F. Gomollon, J. Valle and J. P. Gisbert, Helicobacter pylori Study Group of Asociacion Espanola de, Accuracy of GastroPanel for the diagnosis of atrophic gastritis, Eur. J. Gastroenterol. Hepatol., 2014, 26, 941–948 Search PubMed.
- A. I. Koivusalo, M. P. Pakarinen and K. L. Kolho, Is GastroPanel serum assay useful in the diagnosis of Helicobacter pylori, infection and associated gastritis in children?, Diagn. Microbiol. Infect. Dis., 2007, 57, 35–38 CAS.
- H. Tu, L. Sun, X. Dong, Y. Gong, Q. Xu, J. Jing and Y. Yuan, Serum anti-Helicobacter pylori immunoglobulin G titer correlates with grade of histological gastritis, mucosal bacterial density, and levels of serum biomarkers, Scand. J. Gastroenterol., 2014, 49, 259–266 CAS.
- X. Y. Cao, Z. F. Jia, M. S. Jin, D. H. Cao, F. Kong, J. Suo and J. Jiang, Serum pepsinogen II is a better diagnostic marker in gastric cancer, World J. Gastroenterol., 2012, 18, 7357–7361 CrossRef PubMed.
- L. Yuan, J. B. Zhao, Y. L. Zhou, Y. B. Qi, Q. Y. Guo, H. H. Zhang, M. N. Khan, L. Lan, C. H. Jia, Y. R. Zhang and S. Z. Ding, Type I and type II Helicobacter pylori infection status and their impact on gastrin and pepsinogen level in a gastric cancer prevalent area, World J. Gastroenterol., 2020, 26, 3673–3685 CAS.
- W. Wang, W. Jiang, S. Zhu, X. Sun, P. Li, K. Liu, H. Liu, J. Gu and S. Zhang, Assessment of prevalence and risk factors of helicobacter pylori infection in an oilfield Community in Hebei, China, BMC Gastroenterol., 2019, 19, 186 Search PubMed.
- A. Tett, E. Pasolli, G. Masetti, D. Ercolini and N. Segata, Prevotella diversity, niches and interactions with the human host, Nat. Rev. Microbiol., 2021, 19, 585–599 CAS.
- I. G. Macchione, L. R. Lopetuso, G. Ianiro, M. Napoli, G. Gibiino, G. Rizzatti, V. Petito, A. Gasbarrini and F. Scaldaferri, Akkermansia muciniphila: key player in metabolic and gastrointestinal disorders, Eur. Rev. Med. Pharmacol. Sci., 2019, 23, 8075–8083 Search PubMed.
- Y. Okamoto, Y. Miyabe, M. Seki, Y. Ushio, K. Sato, E. Kasama, K. Akiyama, K. Karasawa, K. Uchida, K. Kikuchi, K. Nitta, T. Moriyama and J. Hoshino, First case of a renal cyst infection caused by Desulfovibrio: a case report and literature review, BMC Nephrol., 2022, 23, 194 Search PubMed.
- Y. Nie, X. Q. Xie, L. Zhou, Q. Guan, Y. Ren, Y. Mao, J. S. Shi, Z. H. Xu and Y. Geng, Desulfovibrio fairfieldensis-Derived Outer Membrane Vesicles Damage Epithelial Barrier and Induce Inflammation and Pyroptosis in Macrophages, Cells, 2022, 12(1), 89 CrossRef PubMed.
- C. S. Caceres, G. L. Gallo, F. A. Colocho, J. E. Silva, J. A. Garay and M. A. Mattar Dominguez, Vegetative forms of Clostridium chauvoei trigger apoptotic and inflammatory responses on macrophages, Anaerobe, 2022, 77, 102616 CrossRef CAS PubMed.
- M. R. Popoff, Clostridium difficile and Clostridium sordellii toxins, proinflammatory versus anti-inflammatory response, Toxicon, 2018, 149, 54–64 CrossRef CAS PubMed.
- L. Wang, L. Tang, Y. Feng, S. Zhao, M. Han, C. Zhang, G. Yuan, J. Zhu, S. Cao, Q. Wu, L. Li and Z. Zhang, A purified membrane protein from Akkermansia muciniphila or the pasteurised bacterium blunts colitis associated tumourigenesis by modulation of CD8(+) T cells in mice, Gut, 2020, 69, 1988–1997 Search PubMed.
- L. Wang, Z. Zhao, L. Zhao, Y. Zhao, G. Yang, C. Wang, L. Gao, C. Niu and S. Li, Lactobacillus plantarum DP189 Reduces alpha-SYN Aggravation in MPTP-Induced Parkinson's Disease Mice via Regulating Oxidative Damage, Inflammation, and Gut Microbiota Disorder, J. Agric. Food Chem., 2022, 70, 1163–1173 CrossRef CAS PubMed.
- C. Carbonne, S. Chadi, C. Kropp, L. Molimard, F. Chain, P. Langella and R. Martin, Ligilactobacillus salivarius CNCM I-4866, a potential probiotic candidate, shows anti-inflammatory properties in vitro and in vivo, Front. Microbiol., 2023, 14, 1270974 CrossRef PubMed.
- R. Tavares and S. K. Pathak, Induction of TNF, CXCL8 and IL-1beta in macrophages by Helicobacter pylori secreted protein HP1173 occurs via MAP-kinases, NF-kappaB and AP-1 signaling pathways, Microb. Pathog., 2018, 125, 295–305 CrossRef CAS PubMed.
Footnote |
| † These authors should be credited as having equal first authorship. |
| This journal is © The Royal Society of Chemistry 2025 |