Isorhamnetin ameliorates hyperuricemia by regulating uric acid metabolism and alleviates renal inflammation through the PI3K/AKT/NF-κB signaling pathway†
Xiaoran
Kong
 ab,
Li
Zhao
ab,
He
Huang
ab,
Qiaozhen
Kang
a,
Jike
Lu
ab,
Li
Zhao
ab,
He
Huang
ab,
Qiaozhen
Kang
a,
Jike
Lu
 *ab and
Jiaqing
Zhu
*ab
*ab and
Jiaqing
Zhu
*ab
aSchool of Life Sciences, Zhengzhou University, Zhengzhou, 450001, China. E-mail: zjq0519qing@163.com; ljk002004@163.com; xrkong2023@163.com; zhli099@163.com; huang4826443@163.com; qzkang@zzu.edu.cn
bFood Laboratory of Zhongyuan, Zhengzhou University, Zhengzhou, 450001, Henan, China
First published on 7th March 2025
Abstract
Hyperuricemia is a chronic metabolic disease with high incidence, and it has become a severe health risk in modern times. Isorhamnetin is a natural flavonoid found in a variety of plants, especially fruits such as buckthorn. The in vivo hyperuricemia ameliorating effect of isorhamnetin and the specific molecular mechanism were profoundly investigated using a hyperuricemia mouse model in this study. Results indicated that isorhamnetin showed a significant uric acid-lowering effect in mice. Isorhamnetin was able to reduce uric acid production by inhibiting XOD activity. Furthermore, it reduced the expression of GLUT9 to inhibit uric acid reabsorption and enhanced the expression of ABCG2, OAT1, and OAT3 to promote uric acid excretion. Metabolomics analysis revealed that gavage administration of isorhamnetin restored purine metabolism and riboflavin metabolism disorders and thus significantly alleviated hyperuricemia in mice. Furthermore, the alleviating effect of isorhamnetin on hyperuricemia-induced renal inflammation and its specific mechanism were explored through network pharmacology and molecular validation experiments. Network pharmacology predicted that seven targets were enriched in the PI3K/AKT pathway (CDK6, SYK, KDR, RELA, PIK3CG, IGF1R, and MCL1) and four targets were enriched in the NF-κB pathway (SYK, PARP1, PTGS2, and RELA). Western blot analysis validated that isorhamnetin inhibited the phosphorylation of PI3K and AKT and down-regulated the expression of NF-κB p65. It indicated that isorhamnetin could inhibit the PI3K/AKT/NF-κB signaling pathway to reduce the levels of renal inflammatory factors (TNF-α, IL-β and IL-6) and ultimately ameliorate hyperuricemia-induced renal inflammation in mice. This study provides a comprehensive and strong theoretical basis for the application of isorhamnetin in the field of functional foods or dietary supplements to improve hyperuricemia.
1. Background
Hyperuricemia (HUA) is a common chronic metabolic disease characterized by a disorder of uric acid metabolism, resulting in abnormally elevated serum uric acid (UA) levels. HUA is diagnosed when serum uric acid concentrations are ≥420 μmol L−1 in males and ≥360 μmol L−1 in females.1,2 The incidence of HUA has been increasing globally in recent years. Moreover, it is not only a precursor to gout but also closely associated with the development of insulin resistance,3 cardiovascular disease,4 hypertension,5 chronic kidney disease,6 and so on,7 which impose a huge burden on global public health.HUA generally occurs via either excessive UA production or inadequate UA excretion. UA is mainly produced through the purine metabolic pathway in the liver, which is a complex process that relies on the synergistic action of several enzymes. Xanthine oxidase (XOD) is the key rate-limiting enzyme in UA synthesis, and it is responsible for oxidizing hypoxanthine to xanthine and further converting xanthine into UA in the final stage of metabolism.8 Once synthesized in the liver, UA enters the blood circulation, and then, most of the UA is excreted through the kidneys. Renal filtration and tubular reabsorption are the key links in the excretion of UA. These processes involve the participation of various UA transport proteins, including urate transport 1 (URAT1), glucose transporter 9 (GLUT9), organic anion transporter 1 (OAT1), and organic anion transporter 3 (OAT3).9 Among them, URAT110 and GLUT911 are able to reabsorb UA from the renal tubules back into the bloodstream, while OAT1 and OAT3 can transport UA into the lumen of renal tubules and excrete it out of the body.12,13 In addition, ATP-binding cassette super-family G member 2 (ABCG2) has also been found to play an important role in UA excretion in recent years.14,15 Abnormal expressions of the above transport proteins may lead to impaired UA excretion and trigger hyperuricemia.
Currently, the clinical drugs for HUA mainly include allopurinol (All), febuxostat and benzbromarone. Among them, allopurinol and febuxostat are inhibitors of XOD, while benzbromarone prevents the reabsorption of UA through inhibiting the activity of URAT1. Nevertheless, the side effects of these drugs severely limit their use. Allopurinol has a certain degree of nephrotoxicity and also causes gastrointestinal discomfort and rashes in some patients,16,17 while benzbromarone has been forbidden in America and some European countries owing to its potential hepatotoxicity.18,19 Therefore, it is still urgent to find safer alternatives for the treatment of hyperuricemia.
Compounds originating from natural products have been extensively studied for the treatment of HUA or developed into functional foods with HUA ameliorating effects on account of their safety to living organisms. Most of them are plant polyphenols and flavonoids such as naringenin,20 apigenin,21 chrysin,22 caffeic acid phenethyl ester,23 chlorogenic acid,24 ellagic acid,25 and epigallocatechin gallate.26 Naringenin was able to improve HUA through reducing the expression of GLUT9 and increasing the expression of ABCG2.20 Furthermore, 2,4-dihydroxybenzoic acid methyl ester could exert UA-lowering effects in HUA mice by inhibiting the XOD activity and down-regulating the expression of URAT1.27
In recent years, a growing body of epidemiologic and experimental evidence has suggested that UA acts as a danger signal capable of triggering immune and inflammatory responses. HUA can increase the risk of kidney injury through causing renal inflammation.28 Concurrently, the HUA renal inflammation ameliorating effects and the mechanisms of natural products have also been extensively researched in the last few years. It had been demonstrated that corn silk flavonoids alleviated inflammation and apoptosis by inhibiting the PI3K/AKT/NF-κB pathway.29 After apigenin administration, serum and renal inflammatory factor levels were reduced in HUA mice and apigenin could ameliorate HUA and renal injury through the JAK2/STAT3 pathway.21
Isorhamnetin is a flavonoid compound found in various natural plants and is abundant in the leaves and fruits of sea buckthorn and ginkgo. Isorhamnetin has a wide range of pharmacological activities such as cardiovascular protective, anti-inflammatory, antitumor, antioxidant, antibacterial and antiviral effects.30,31 It has been reported that isorhamnetin has a UA-lowering effect in vitro and in vivo,32,33 whereas the specific molecular mechanisms by which isorhamnetin ameliorates HUA and its mitigating effect on HUA-induced renal inflammation have not been clarified yet. Therefore, this study comprehensively investigated the UA-lowering effect and its potential mechanism of isorhamnetin and further explored its relieving function and mechanism on renal inflammation caused by HUA based on the constructed HUA mouse model. It will provide a comprehensive and penetrating theoretical basis for the development of functional food ingredients to ameliorate HUA.
2. Materials and methods
2.1 Chemicals and reagents
Isorhamnetin (purity ≥ 98%, CAS: 480-19-3) was purchased from Weikeqi Biological Technology Co., Ltd (Chengdu, China). Potassium oxonate (PO), hypoxanthine (HX) and carboxymethyl cellulose were purchased from Macklin Co., Ltd (Shanghai, China). Allopurinol was obtained from Aladdin Co., Ltd (Shanghai, China). UA (uric acid), creatinine (CRE), blood urine nitrogen (BUN) and xanthine oxidase (XOD) assay kits were purchased from Jiancheng Technology Co., Ltd (Nanjing, China). TNF-α (tumor necrosis factor-α), IL-1β (Interleukin-1β) and IL-6 (Interleukin-6) ELISA kits were purchased from Shanghai Enzyme-linked Biotechnology Co., Ltd (Shanghai, China). Primary antibodies against GLUT9 (ab223470, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3000) and OAT1 (ab135924, 1
3000) and OAT1 (ab135924, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000) were purchased from Abcam (Cambridge, UK). Antibodies against OAT3 (1
1000) were purchased from Abcam (Cambridge, UK). Antibodies against OAT3 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000) was purchased from Santa Cruz Technology (Dallas, Texas, USA). The following primary antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA): phosphoinositide 3-kinase (PI3K, 4292, 1
1000) was purchased from Santa Cruz Technology (Dallas, Texas, USA). The following primary antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA): phosphoinositide 3-kinase (PI3K, 4292, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), phospho-PI3K (p-PI3K, 4228, 1
1000), phospho-PI3K (p-PI3K, 4228, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), protein kinase B (AKT, 9272, 1
1000), protein kinase B (AKT, 9272, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), and phosphor-AKT (p-AKT, 9275, 1
1000), and phosphor-AKT (p-AKT, 9275, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000). The following primary antibodies were purchased from Proteintech (Chicago, USA): ABCG2 (27286-1-AP, 1
1000). The following primary antibodies were purchased from Proteintech (Chicago, USA): ABCG2 (27286-1-AP, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), nuclear factor kappa-B p65 (NF-κB p65, 10745-1-AP, 1
1000), nuclear factor kappa-B p65 (NF-κB p65, 10745-1-AP, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000), and glyceraldehyde 3-phosphate dehydrogenase (GAPDH, 10494-1-AP, 1
1000), and glyceraldehyde 3-phosphate dehydrogenase (GAPDH, 10494-1-AP, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20
20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000). Horseradish peroxidase-labeled anti-rabbit IgG (SA00001-2, 1
000). Horseradish peroxidase-labeled anti-rabbit IgG (SA00001-2, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000) and Horseradish peroxidase-labeled anti-mouse IgG (SA00001-1, 1
000) and Horseradish peroxidase-labeled anti-mouse IgG (SA00001-1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000) were obtained from Jackson Proteintech.
000) were obtained from Jackson Proteintech.
2.2 Animal experiment
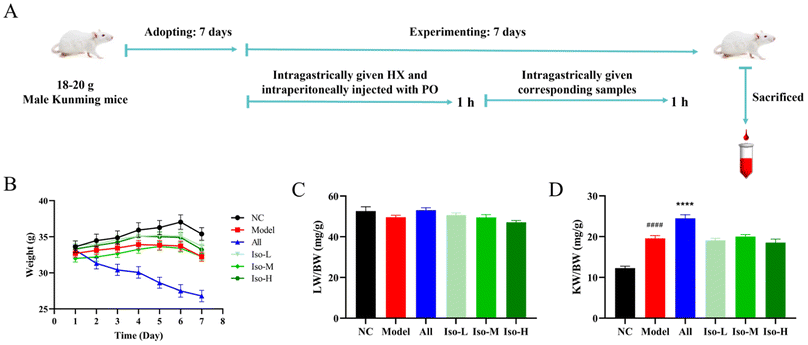
Male Kunming mice (weight 18–22 g) were purchased from the Experimental Animal Center of Henan Province (Zhengzhou, China). All mice were housed in a specific-pathogen-free animal facility with a constant temperature of 22 ± 2 °C, humidity of 50 ± 15%, and a 12 h light/12 h darkness cycle at Zhengzhou University. All animal procedures were performed in accordance with the Guidelines for Care and Use of Laboratory Animals of Zhengzhou University (China) and approved by the Animal Ethics Committee of Zhengzhou University (2023-328).After acclimatization for one week, KM mice were randomly divided into 6 groups (n = 8): normal control (NC), model (HUA), positive control (All, 10 mg kg−1), isorhamnetin low-dose (Iso-L, 25 mg kg−1), isorhamnetin medium-dose (Iso-M, 100 mg kg−1) and isorhamnetin high-dose (Iso-H, 175 mg kg−1). Based on the dose conversion from body surface area, the dose of 100 mg kg−1 of mice is equivalent to 8.1 mg kg−1 of the human dose. The content of isorhamnetin in dried Ningxia sea buckthorn fruit is 11.8 mg g−1, while the moisture content of the fruit can reach 80%. For an adult (70 kg), 8.1 mg kg−1 of the human dose means that he/she needs to consume 240.5 g fresh Ningxia sea buckthorn fruit. Mice except for the NC group were administered daily with PO (300 mg kg−1, intraperitoneal injection) and HX (500 mg kg−1, gavage) to construct an HUA mouse model. The NC group was administered equal volumes of 0.5% carboxymethylcellulose sodium (CMC-Na) solution separately. At 1 h after the induction of PO and HX, each group was gavaged with the corresponding doses of allopurinol and isorhamnetin, and the NC and model groups were gavaged with equal volumes of 0.5% CMC-Na solution. PO, HX, All and isorhamnetin suspension were configured using 0.5% CMC-Na. The above procedures continued for 7 days. An animal experiment was conducted as previously described with appropriate modifications,23,27 and the experiment protocol is illustrated in Fig. 1A.
2.3 Sample collection
After the last treatment, blood was collected from the mouse to obtain the serum. The liver and kidney were immediately removed and then stored either at −80 °C or in a 4% paraformaldehyde solution for further research.2.4 Biochemical analysis
The levels of serum UA and BUN, CRE and liver XOD were measured using commercial kits according to the corresponding protocols. Following the manufacturer's instructions, the levels of TNF-α, IL-1β and IL-6 were detected using Enzyme-linked immunosorbent assay (ELISA) kits.2.5 Hematoxylin and eosin (H&E) staining
Liver and renal tissues of mice kept in a 4% paraformaldehyde solution were taken out and fixed in neutral 10% formalin and then dehydrated with ethanol. The kidney tissues were embedded in paraffin, divided into 5 μm sections and then stained with H&E staining. The treated sections were observed using a microscope at ×200 magnification for pathological analysis.2.6 Network pharmacology
2.7 Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA of mouse kidney tissue was obtained using FastPure Cell/Tissue Total RNA Isolation Kit V2 (RC112, Vazyme, Nanjing, China) and reversely transcribed to cDNA using HiScript III All-in-one RT SuperMix Perfect for qPCR (R333, Vazyme, Nanjing, China). The mRNA expression was analyzed according to the instructions of ChamQ Universal SYBR qPCR Master Mix (Q711, Vazyme, Nanjing, China). The qRT-PCR was performed with initial heat denaturation at 95 °C for 30 s, and the PCR cycles were repeated 40 times under the following conditions: denaturation at 95 °C for 15 s and annealing at 60 °C for 30 s. Genetic relative quantification was achieved by the 2−ΔΔCT method using GAPDH as an internal reference. The primer was synthesized by Sangon technology (Shanghai, China), and the primer sequences are listed in Table 1.| Gene | GeneBank | Primer sequences (5′–3′) |
|---|---|---|
| GAPDH | NM_001289726.2 | Forward: AGGTCGGTGTGAACGGATTTG |
| Reverse: TGTAGACCATGTAGTTGAGGTCA | ||
| GLUT9 | NM_001102415.1 | Forward: CCTCCTTCCTGTGGACTCTG |
| Reverse: TCTTTGTCCTCCTCTGCTGG | ||
| ABCG2 | XM_006506150.5 | Forward: CACTGACCCTTCCATCCTCTTC |
| Reverse: ACCCTGTTTAGACATCCTTTTCA | ||
| OAT1 | NM_008766.3 | Forward: CACCTGCTAATGCCAACCTC |
| Reverse: CCATTGTGCGGGAAAGGAAA | ||
| OAT3 | NM_001411470.1 | Forward: CTGCCTTCTTCATCTTCTCCTTG |
| Reverse: CTTCCTCCTTCTTGCCGTTG | ||
| TNF-α | NM_013693.3 | Forward: CACCACGCTCTTCTGTCTACTGAAC |
| Reverse: AGATGATCTGAGTGTGAGGGTCTGG | ||
| IL-1β | XM_006498795.5 | Forward: CACTACAGGCTCCGAGATGAACAAC |
| Reverse: TGTCGTTGCTTGGTTCTCCTTGTAC | ||
| IL-6 | NM_001314054.1 | Forward: CTTCTTGGGACTGATGCTGGTGAC |
| Reverse: TCTGTTGGGAGTGGTATCCTCTGTG | ||
| PI3K | NM_001024955.2 | Forward: ACACCACGGTTTGGACTATGG |
| Reverse: GGCTACAGTAGTGGGCTTGG | ||
| AKT | XM_030242000.2 | Forward: ACGTGGTGAATACATCAAGACC |
| Reverse: GCTACAGAGAAATTGTTCAGGGG | ||
| NF-κB | NM_001402548.1 | Forward: AGGCTTCTGGGCCTTATGTG |
| Reverse: TGCTTCTCTCGCCAGGAATAC |
2.8 Western blot analysis
A portion of mouse kidney tissue was taken and lysed using a RIPA lysis buffer. Then they were centrifuged at 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm, 4 °C for 10 min to obtain the total proteins. The protein content was measured using a BCA protein assay kit in order to normalize the concentrations of proteins. After adding the corresponding volume of loading buffer, the protein was denatured via steaming at 95 °C for 10 min (membrane proteins were denatured at 37 °C for 30 min). Total protein (30 μg per lane) was then loaded and separated by 12% SDS-PAGE, which were then transferred onto a PVDF membrane. After blocking with 5% skimmed milk for 1.5 h, the membranes were hybridized with a primary antibody at 4 °C overnight. Next, the primary antibody was eluted and the secondary antibody was incubated for 1 h at room temperature. The bands were visualized with an ECL reagent using an iBright FL1500 imaging system (Thermo, MA, USA). Finally, the grayscale of the protein bands was quantified using the ImageJ software. GAPDH was used as an internal reference for all proteins.
000 rpm, 4 °C for 10 min to obtain the total proteins. The protein content was measured using a BCA protein assay kit in order to normalize the concentrations of proteins. After adding the corresponding volume of loading buffer, the protein was denatured via steaming at 95 °C for 10 min (membrane proteins were denatured at 37 °C for 30 min). Total protein (30 μg per lane) was then loaded and separated by 12% SDS-PAGE, which were then transferred onto a PVDF membrane. After blocking with 5% skimmed milk for 1.5 h, the membranes were hybridized with a primary antibody at 4 °C overnight. Next, the primary antibody was eluted and the secondary antibody was incubated for 1 h at room temperature. The bands were visualized with an ECL reagent using an iBright FL1500 imaging system (Thermo, MA, USA). Finally, the grayscale of the protein bands was quantified using the ImageJ software. GAPDH was used as an internal reference for all proteins.
2.9 Serum untargeted metabolomics analysis
Serum samples from the NC, Model and Iso H groups were taken and preprocessed for untargeted metabolomics analysis. Briefly, chromatography was carried out using an LC-30A UHPLC system (Shimadzu, Japan), and all samples were detected using a Waters ACQUITY Premier HSS T3 Column (2.1 mm × 100 mm, 1.8 μm; Waters, MA, United States). Mass spectrometry was performed using a TripleTOF 6600+ instrument (SCIEX, CA, United States) in both positive and negative ionization modes.Based on the metabolomics data obtained from metabolite detection, principal component analysis (PCA) and orthogonal partial least squares discriminant analysis (OPLS-DA) were performed to assess the separation between groups and the overall differences in metabolic profiles. Then, the quality of the OPLS-DA model was validated and the metabolites were visualized using S-plots. Combined multivariate and univariate statistics were used to screen for differential metabolites, with screening criteria of Variable Importance in Projection (VIP) > 1 and P < 0.05. Finally, the differential metabolites were analyzed for related metabolic pathways by searching the KEGG database (https://www.kegg.jp/).
2.10 Data statistical analysis
Statistical analysis was performed using one-way analysis of variance (ANOVA) in GraphPad Prism 9.0. Data are presented as mean ± standard error of the mean (SEM). Differences were considered statistically significant at the level of P < 0.05.3. Results and discussion
3.1 Effects of isorhamnetin on HUA and renal function in mice
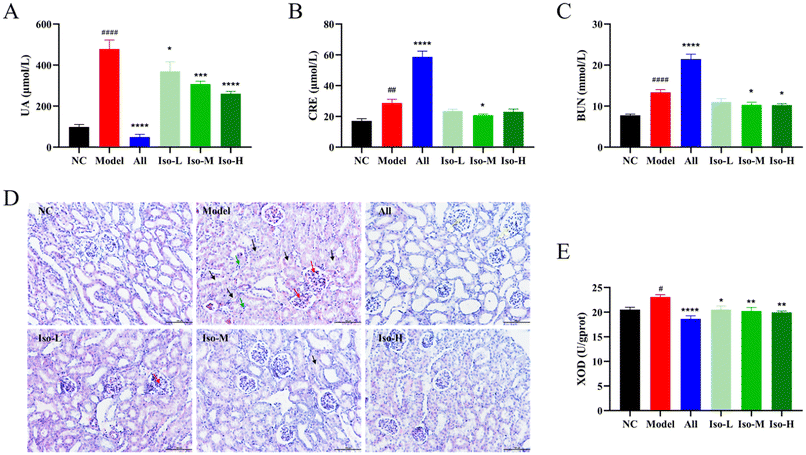
Body weight was one of the key parameters reflecting the physiological and pathological states of mice. By comparing the body weights of mice in each group, it could be initially determined whether isorhamnetin had negative effects on the growth of mice. The body weight growth curves for each group during the experiment are shown in Fig. 1B. The body weights of the mice in all groups except All group showed an increasing trend, while the weight loss observed on day 7 might have been due to water and food fasting on the previous day. The body weight of the All group continued to decrease throughout the gavage period. Mice in this group were also accompanied by signs of depression, dull fur, and loss of appetite, which might be due to the adverse effects of allopurinol. The organ coefficient was an important indicator for assessing organ damage, and it remained relatively stable under normal conditions.33 The liver and kidney coefficients were investigated in each group of mice and used to evaluate the potential hepatorenal toxicity of isorhamnetin. As shown in Fig. 1C, there was no significant difference in the liver coefficients of mice in each group (P > 0.05), indicating that neither the isorhamnetin nor the modeling drugs had negative effects on the liver of mice. The kidney coefficients of mice are shown in Fig. 1D. Compared with the NC group, the kidney coefficients of the model group were significantly elevated by 59.65% (P < 0.0001). Meanwhile, abnormalities such as swelling and whitening of the kidneys were also observed. It suggested that HUA might induce renal hyperplasia and hypertrophy, thus leading to renal injury in mice. The kidney coefficient was further elevated by 25.12% (P < 0.0001) in the All group as compared to the model group. These results indicated that allopurinol caused more severe renal damage on the basis of HUA and that it had serious toxic side effects on kidneys. After isorhamnetin gavage, although the kidney coefficients of HUA mice were not significantly reduced, it did not continue to increase, suggesting that isorhamnetin showed much less side-effects over allopurinol. Overall, allopurinol showed severe overall toxicity and nephrotoxicity in HUA mice as a commonly used therapeutic agent for HUA. In contrast, isorhamnetin was comparatively safe for HUA mice and no significant liver or kidney toxicity was observed in vivo.In most mammals, the produced UA could be further converted into the more soluble allantoin by uricase, thus maintaining UA at a low level in vivo.35 However, the functions of uricase were absent in humans and some primates due to the mutations of uricase genes, which could prevent UA from being broken down and it could only be excreted through the kidney and other organs. When the level of serum UA in the body was abnormally high, it often indicated the occurrence of HUA.1 PO was an inhibitor of uricase and could inhibit the degradation of UA in mice; HX was a precursor substance of UA, its increased level could promote the production of UA. The serum UA level was increased by gavage of HX (500 mg kg−1) combined with intraperitoneal injection of PO (300 mg kg−1) to construct an HUA mouse model. Fig. 2A demonstrates the serum UA levels of mice in each group. Compared with the NC group (98.45 μmol L−1), the serum UA level of model group (478.02 μmol L−1) was significantly higher by 4.86-fold (P < 0.0001), suggesting the successful establishment of the HUA mouse model. Compared with the model group, allopurinol reduced serum UA to 49.54 μmol L−1 (P < 0.0001), which was consistent with its expected effect and further validated the reliability of the HUA model. Gavage of low, medium and high doses of isorhamnetin reduced serum UA to 370.28, 307.74 and 260.68 μmol L−1 (P < 0.05, P < 0.001, P < 0.0001) in HUA mice, respectively. It indicated that isorhamnetin showed significant UA-lowering effects with dose-dependent characteristics. This is in agreement with the results of previous studies.27,33 Creatinine (CRE) is a type of waste product produced by muscle metabolism. The serum CRE level is usually measured to assess the renal function. When the kidneys are in a healthy state, they can effectively filter CRE from the body and maintain its concentration in the blood within a certain range. As shown in Fig. 2B, the serum CRE in the model group (28.71 μmol L−1, P < 0.01) was significantly increased compared with the NC group (17.15 μmol L−1), which indicated that the kidneys of the HUA mice were severely damaged. Compared with that in the model group, the All group mouse serum CRE continued to increase to 58.65 μmol L−1, which could be attributed to the toxic effects of allopurinol on the kidneys. The serum CRE level in the ISO-M group was significantly reduced to 20.84 μmol L−1 (P < 0.05). These results suggested that isorhamnetin had a potential ameliorative effect on renal functional impairment caused by HUA. Blood urea nitrogen (BUN), like serum CRE, was also a key indicator for assessing renal health. Its elevated levels were equally indicative of possible renal dysfunction.23 As can be seen in Fig. 2C, the BUN levels in the NC and model groups were 7.79 and 13.37 mmol L−1, with a 71.63% rise in the HUA mice compared to the NC group (P < 0.0001). BUN of the All group was 21.45 mmol L−1, which continued to rise by 60.43% compared to the model group (P < 0.0001), whereas, the mice BUN levels all showed a decreasing trend after isorhamnetin gavage. It was significantly reduced in the Iso-M (10.31 mmol L−1, P < 0.05) and Iso-H (10.26 mmol L−1, P < 0.05) groups. These results were consistent with serum CRE, which indicated that isorhamnetin might have reparative effects on HUA-induced renal injury.
The preliminary results showed that the kidney coefficient, serum CRE and BUN levels were abnormally elevated in HUA mice. HUA disrupted renal tubular filtration and impaired renal function in mice. Therefore, the histopathological changes in the kidney tissues were analyzed by H&E staining to further investigate the effects of HUA on the kidney and the role of isorhamnetin.36 The results of H&E staining are shown in Fig. 2D. Renal tissues of mice in the NC group had a normal morphology, and the tubular epithelial cells were arranged in a neat and orderly manner, with no obvious inflammatory reaction. The model group mice showed solidification of the glomeruli (red arrows), inflammatory cell infiltration (green arrows) and tubular edema (black arrows). The mice of the All group exhibited glomerular hypertrophy (yellow arrows). After gavage with isorhamnetin, kidney tissue edema and glomerular atrophy of HUA mice were attenuated, and inflammatory cells were reduced. These indicated that isorhamnetin could significantly restore the morphological damage and lesions of renal tissues induced by HUA, alleviate renal inflammation, and exert a protective effect on the kidneys.
Based on the above results, it was clear that isorhamnetin exhibited remarkable UA-lowering effects. Compared with the conventional therapeutic drug allopurinol, isorhamnetin showed much less side-effects in terms of renal safety and nephroprotection, highlighting its potential as a function-improving compound for the alleviation of HUA.
3.2 Isorhamnetin inhibits UA production by inhibiting the XOD activity
As shown in Fig. 2E, the XOD activity of mice in the NC group was 20.52 U per gprot, whereas it was significantly higher in the model group (23.09 U per gprot, P < 0.05). It might be caused by excessive intake of hypoxanthine, a precursor substance of uric acid. The activity of XOD in the All group was significantly lower (18.64 U per gprot, P < 0.0001) than that of the model group, which was consistent with the expectation that allopurinol was an inhibitor of XOD. Compared to the model group mice, the XOD activities in the Iso-L, Iso-M and Iso-H group mice were significantly reduced to 20.53 (P < 0.05), 20.22 (P < 0.01) and 19.96 (P < 0.01) U per gprot, respectively. The synthesis of UA in the body mainly occurred through the purine nucleotide metabolic pathway in the liver. In this process, adenosine phosphate was first phosphorylated to form adenosine, which was then deaminated to form inosine under the action of adenosine deaminase. Inosine was further decomposed into hypoxanthine, which was then oxidized to xanthine and UA by XOD.37 Among them, XOD played the key role in the above process and was the rate-limiting enzyme of UA-synthesis. The experimental results suggested that isorhamnetin could reduce the serum UA level by inhibiting the XOD activity, which was consistent with what was reported in the previous literature.383.3 Isorhamnetin promotes UA excretion by regulating GLUT9, ABCG2, OAT1 and OAT3 expression
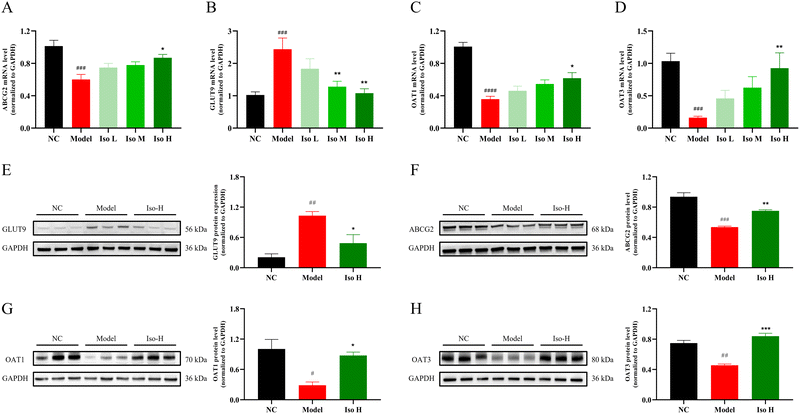
To investigate whether isorhamnetin alleviated HUA by affecting uric acid excretion, expressions of renal uric acid transporters GLUT9, ABCG2, OAT1 and OAT3 at the transcriptional level were determined using qRT-PCR. GLUT9 was encoded by the SLC2A9 gene, which was expressed on the cell membrane of renal tubules and in the proximal renal tubules. GLUT9 reabsorbed UA filtered by the glomerulus back into the blood.11 Therefore, high GLUT9 expression tended to elevate blood UA levels and increase the risk of suffering from HUA. As shown in Fig. 3A, the expression of GLUT9 mRNA was significantly higher in the model group than in the NC group (P < 0.001), while its expression was reduced after gavage of isorhamnetin, and the difference was significant in the Iso-M group (P < 0.01) and the Iso-H group (P < 0.01). Notably, the expression of GLUT9 in the Iso-H group was restored to a level close to that of the NC group. These results suggested that HUA modeling led to elevated expression of GLUT9 at the gene level in the kidney, and isorhamnetin gavage was able to reverse this phenomenon and reduce the expression of GLUT9 mRNA. ABCG2, distributed in both the kidney and intestinal tissues, plays a crucial role in the process of UA excretion. It is able to utilize ATP hydrolysis to generate energy and actively transport UA against the concentration gradient to excrete them out of the body.39 OAT1 and OAT3, located in the basolateral membrane of the proximal tubule of the kidney, are capable of performing UA uptake from the blood into the proximal tubular cells, thus secreting them into the urine for excretion.12 The three above-mentioned proteins are responsible for the excretion of UA, and play vital roles in maintaining the UA balance in vivo. The mRNA expression of ABCG2, OAT1 and OAT3 in the HUA model group was lower than that in the NC group (P < 0.001, P < 0.0001 and P < 0.001), and high-dose isorhamnetin gavage significantly down-regulated the mRNA expression of the three transporters (P < 0.05, P < 0.05, and P < 0.01) (Fig. 3B–D). These results suggested that isorhamnetin could promote the expression of ABCG2, OAT1 and OAT3 in the kidney of HUA mice at the gene level to facilitate UA excretion. In summary, isorhamnetin was able to regulate the expression of renal UA transporter proteins at the mRNA level. It could inhibit UA reabsorption by decreasing the expression of GLUT9 and promote UA excretion by elevating ABCG2, OAT1 and OAT3 mRNA expression to achieve UA-lowering effects in vivo.The qRT-PCR results showed that the isorhamnetin high-dose group exhibited significant regulatory effects on the four above-mentioned UA transporter proteins. Accordingly, the NC, Model and Iso-H groups were selected for further western blot experiments to validate the regulation effect of isorhamnetin on the expression of GLUT9, ABCG2, OAT1 and OAT3 at the protein level. The western blot results are presented in Fig. 3E–H. In comparison to the NC group, there was an observable increase in GLUT9 protein expression in the kidneys of model group mice (P < 0.01). Meanwhile, the expressions of ABCG2 (P < 0.001), OAT1 (P < 0.05), and OAT3 (P < 0.01) proteins were remarkably decreased. In contrast, after gavage with isorhamnetin, there was a significant reduction in the expression of GLUT9 protein (P < 0.05), accompanied by recovery expression trends of ABCG2 (P < 0.01), OAT1 (P < 0.05), and OAT3 (P < 0.001) proteins. The protein expressions shown by western blot experiments were consistent with the qRT-PCR results. In conclusion, isorhamnetin gavage could effectively decrease the renal reabsorption of UA by inhibiting the expression of GLUT9 protein and promote the excretion of UA by improving the expression of ABCG2, OAT1 and OAT3 proteins in HUA mice, thereby reducing the serum UA level and alleviating the symptoms of hyperuricemia.29,40,41
3.4 Isorhamnetin restores metabolic disorders in HUA mice
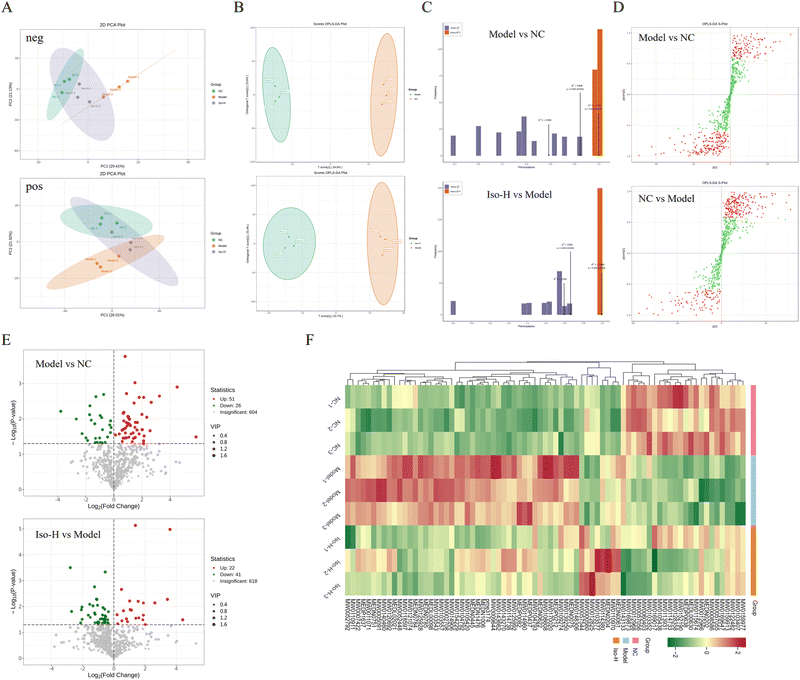
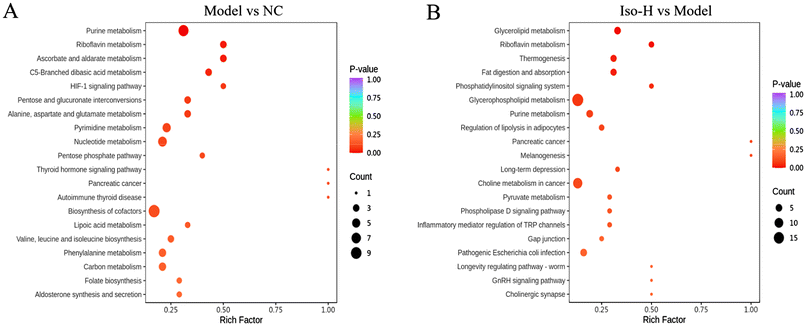
Several previous reports claimed that HUA would cause metabolic disorders in the body.42–45 Thus, the present study utilized metabolomics to investigate the regulatory effects of isorhamnetin on metabolic disorders in HUA mice. Untargeted metabolomics was performed on serum from mice in the NC, Model and Iso-H groups. A total of 773 metabolites were detected, including 308 in the negative and 465 in the positive ion mode. The differences in distribution between the sample groups were analyzed by PCA, and the results showed that significant separation of the NC, Model and Iso-H groups occurred on PC1 in the negative ion mode and on PC2 in the positive ion mode (Fig. 4A). In both modes, the Iso-H group had a tendency to move closer to the NC group, indicating that isorhamnetin reversed HUA-induced metabolic disorders in mice. To further explore the differences between the groups and screen for differential metabolites, OPLS-DA was performed. As shown in the OPLS-DA score plot (Fig. 4B), the model group was clearly separated from the NC and Iso-H groups. It suggested that HUA markedly altered the metabolites in mice, and that isorhamnetin might regulate the disordered metabolism. The R2Y and Q2 values derived from the permutation test are shown in Fig. 4C, which confirmed that the OPLS-DA model was credible and could be used as a predictable model to screen differential metabolites.46 The S-plot (Fig. 4D) revealed the metabolites that contributed to the separation between groups in the OPLS-DA score plot and illustrated the up- and down-regulation of metabolites. Candidate biomarkers were further screened using the VIP of the OPLS-DA model and the P-value of univariate analysis, with VIP > 1 and P < 0.05 as criteria. The screening results are shown in the volcano plots (Fig. 4E); 77 and 63 metabolites were significantly different between the model and NC groups, and Iso-H and Model groups, respectively. Most of the 77 significantly changed metabolites after HUA modeling were organic acids and their derivatives, nucleotides and their metabolites, etc. Their change trends were shown in the heatmap (Fig. 4F). Detailed information about them is provided in Table S1.† After isorhamnetin gavage, the levels of most differential metabolites were regressed. Twenty-four of them were significantly (P < 0.05) restored, including 3 metabolites in the purine metabolic pathways, inosine, xanthosine, and 5′-phosphoribosyl-N-formylglycinamide.The screened significantly different metabolites were imported into the KEGG database for pathway enrichment analysis. As shown in Fig. 5A, the occurrence of metabolic disorders is mainly related to the following pathways after HUA model construction: purine metabolism, riboflavin metabolism, ascorbate and aldarate metabolism, pyrimidine metabolism, nucleotide metabolism, etc. Pathways that were significantly altered after isorhamnetin gavage include glycerolipid metabolism, riboflavin metabolism, fat digestion and absorption, and purine metabolism (Fig. 5B). Purine metabolism and riboflavin metabolism were enriched in both comparison groups. Purine metabolism was a pathway known to be closely related to the development of HUA, and UA was the end product of purine catabolism in the organism. The disturbance of purine metabolism often causes abnormal elevation of the UA level. Purine metabolism plays an important role in the pathogenesis of HUA.47 Riboflavin, one type of B vitamin, is an indispensable coenzyme of many oxidative enzyme systems in vivo. It is involved in many complex processes in sugar, fat and protein metabolism. The deficiency of riboflavin often leads to metabolic disorders, which seriously affect the progress of material metabolism and organ function.48 The above-mentioned results indicated that isorhamnetin might restore metabolic homeostasis through purine metabolism and riboflavin metabolism in HUA mice. The metabolomics results are consistent with the results presented in part 3.2. To summarize, isorhamnetin could reduce uric acid production by inhibiting the XOD activity, regulate the expression of renal UA transporter proteins GLUT9, ABCG2, OAT1, and OAT3 to promote UA excretion, and restore purine metabolism and riboflavin metabolism disorders, and thus, significantly alleviate hyperuricemia in mice.
3.5 Effects of isorhamnetin on renal inflammatory factors in HUA mice
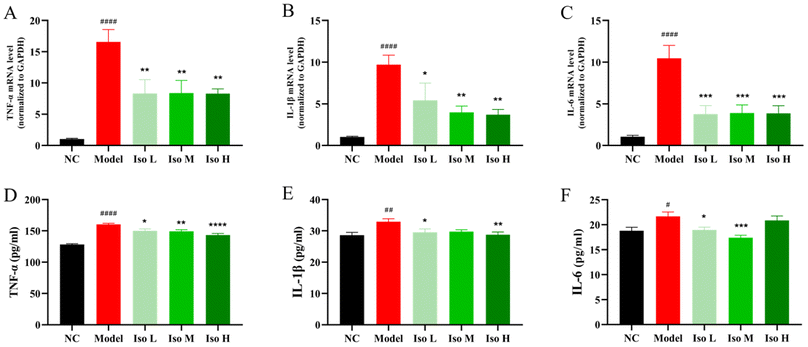
HUA development was often accompanied by renal inflammation. Persistent elevation of UA in the body might lead to mono-sodium urate crystal formation. The deposition of MSU crystals in the kidney could promote the expression of inflammatory cytokines and chemokines, including TNF-α, IL-1β, and IL-6, which would further induce inflammation and trigger kidney injury.25,49,50 Furthermore, increased levels of UA resulted in the generation of reactive oxygen species, leading to oxidative stress and cellular damage and exacerbating the inflammatory process in the kidney.40,51 TNF-α, IL-1β, and IL-6 are pivotal pro-inflammatory cytokines in the body involved in inflammation. TNF-α is produced by monocytes or macrophages. It can contribute to the systemic inflammatory response. IL-1β can activate neutrophils and guide them to the site of inflammation. IL-6 is implicated in early T-cell activation, serving as another pro-inflammatory factor in the inflammatory response.21,52 The renal inflammation ameliorating effect of isorhamnetin was evaluated by surveying the expression of inflammatory factors TNF-α, IL-1β and IL-6 in the renal tissues of mice using qRT-PCR and ELISA experiments. Fig. 6A–C demonstrate that the relative mRNA expression levels of all three inflammatory factors were abnormally elevated in the model group mice, with a 15.54-fold increase for TNF-α, a 9.54-fold increase for IL-1β, and a 9.95-fold increase for IL-6 compared to that in the NC group mice. After isorhamnetin gavage, the relative expression levels of TNF-α mRNA in the Iso-L, Iso-M and Iso-H group mice were dramatically reduced to 8.05, 8.1 and 8.03 folds of that in the NC group mice. IL-1β levels were down-regulated to 5.33, 3.91 and 3.63 folds of that in the NC group mice. Meanwhile, the mRNA expressions of IL-6 were also significantly restored to 3.59, 3.71 and 3.67 folds of that in the NC group mice.The expression of TNF-α, IL-1β and IL-6 at the transcriptional levels was further quantified by ELISA. The results depicted in Fig. 6D–F show a noticeable elevation in the levels of TNF-α (160.36 pg mL−1, P < 0.0001), IL-1β (32.93 pg mL−1, P < 0.01), and IL-6 (21.67 pg mL−1, P < 0.05) in the kidneys of HUA mice compared to the NC group mice (128.26, 28.64 and 18.78 pg mL−1). These indicated that HUA was capable of inducing severe renal inflammation in mice. Compared with the model group, TNF-α, IL-1β and IL-6 were significantly reduced in the Iso-L group. The Iso-M group was able to significantly reduce the levels of TNF-α and IL-6. The Iso-H group significantly decreased the contents of TNF-α and IL-1β compared to the model group. These results represented that isorhamnetin gavage could markedly reduce the elevated levels of pro-inflammatory factors and alleviate the renal inflammation induced by HUA in mice. ELISA coupled with qRT-PCR results further confirmed that the renal anti-inflammatory effect of isorhamnetin might be exerted through inhibiting the expression of TNF-α, IL-1β, and IL-6 at mRNA and protein levels. However, the specific mechanism by which isorhamnetin reduces the expression of inflammatory factors has not yet been elucidated and further exploration is needed.
3.6 Predicting potential mechanisms via network pharmacology
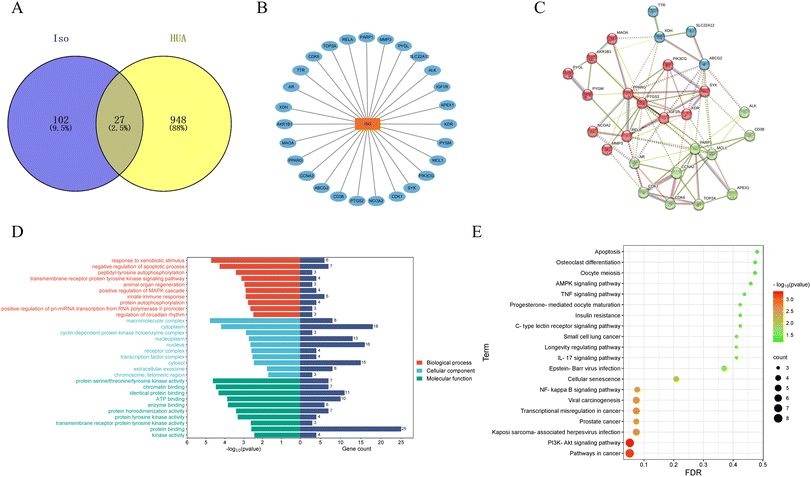
Isorhamnetin is a natural flavonoid found in various natural plants. After searching the TCMSP database, the oral bioavailability value of isorhamnetin was found to be 49.6% and the drug likeness value was found to be 0.31. These met the screening criteria of the active ingredients. The target genes of isorhamnetin were obtained from the Swiss and Uniprot databases. After deleting duplicates, a total of 129 potential target genes were obtained. Using “Hyperuricemia” as the search term, this study searched the GeneCards and DisGeNET databases for HUA disease target genes. After deleting the duplicates, a total of 973 target genes were identified. The 129 component targets of isorhamnetin and the 973 disease targets of HUA were imported into the online Venn diagram creation website, and 27 shared targets were obtained (Fig. 7A). These targets were identified as potential targets for isorhamnetin to ameliorate HUA (Fig. 7B). The shared targets were first imported into the STRING database and then imported into the Cytoscape 3.7.2 software to obtain the PPI diagram (Fig. 7C). There were a total of 94 edges between 27 nodes. The importance of the targets was evaluated according to betweenness centrality, closeness centrality and degree (Table 2). Among them, ABCG2 and XDH were known target proteins involved in UA production and excretion in HUA.| No | Target name | Betweenness centrality | Closeness centrality | Degree | Protein name |
|---|---|---|---|---|---|
| 1 | PTGS2 | 110.625 | 0.703 | 15 | Prostaglandin G/H synthase 2 |
| 2 | PARP1 | 108.087 | 0.667 | 15 | Poly [ADP-ribose] polymerase-1 |
| 3 | PPARG | 111.278 | 0.703 | 15 | Peroxisome proliferator activated receptor gamma |
| 4 | MCL1 | 62.131 | 0.634 | 13 | Induced myeloid leukemia cell differentiation protein Mcl-1 |
| 5 | RELA | 37.868 | 0.591 | 10 | Transcription factor p65 |
| 6 | KDR | 16.823 | 0.591 | 10 | Vascular endothelial growth factor receptor 2 |
| 7 | AR | 18.289 | 0.565 | 9 | Androgen receptor |
| 8 | IGF1R | 8.538 | 0.578 | 9 | Insulin-like growth factor I receptor |
| 9 | ABCG2 | 56.392 | 0.591 | 9 | ATP-binding cassette sub-family G member 2 |
| 10 | CDK1 | 14.277 | 0.500 | 8 | Cyclin-dependent kinase 1 |
| 11 | CCNA2 | 12.348 | 0.542 | 8 | Cyclin-A2 |
| 12 | SYK | 16.661 | 0.553 | 8 | Tyrosine-protein kinase SYK |
| 13 | XDH | 69.179 | 0.520 | 7 | Xanthine dehydrogenase/oxidase |
| 14 | CDK6 | 1.551 | 0.464 | 6 | Cyclin-dependent kinase 6 |
| 15 | AKR1B1 | 33.978 | 0.500 | 6 | Aldose reductase |
| 16 | PIK3CG | 0.286 | 0.473 | 5 | PI3-kinase p110-gamma subunit |
| 17 | TOP2A | 2.250 | 0.441 | 5 | DNA topoisomerase II alpha |
| 18 | MMP3 | 3.064 | 0.481 | 4 | Matrix metalloproteinase 3 |
| 19 | NCOA2 | 0.000 | 0.473 | 4 | Nuclear receptor coactivator 2 |
| 20 | MAOA | 0.000 | 0.481 | 4 | Monoamine oxidase A |
| 21 | PYGM | 19.404 | 0.464 | 4 | Glycogen phosphorylase, muscle form |
| 22 | CD38 | 0.429 | 0.433 | 3 | Lymphocyte differentiation antigen CD38 |
| 23 | APEX1 | 0.543 | 0.426 | 3 | DNA-(apurinic or apyrimidinic site) lyase |
| 24 | ALK | 0.000 | 0.448 | 3 | ALK tyrosine kinase receptor |
| 25 | PYGL | 0.000 | 0.351 | 2 | Liver glycogen phosphorylase |
| 26 | SLC22A12 | 0.000 | 0.413 | 2 | Solute carrier family 22 member 12 |
| 27 | TTR | 0.000 | 0.347 | 1 | Transthyretin |
GO and KEGG enrichment analyses of common targets of isorhamnetin and HUA were performed through the DAVID website. The results of GO enrichment analysis included 141 terms. They were categorized into three classes: 97 terms in the biological process (BP), 14 terms in the cell component (CC), and 30 terms in the molecular function (MF). Fig. 7D shows the top 10 terms in the three categories.
A total of 21 relevant signaling pathways were enriched in KEGG, and the top 20 are illustrated in Fig. 7F. These pathways included the PI3K-AKT signaling pathway, cancer-related pathways (pathways in cancer, transcriptional misregulation in cancer, viral carcinogenesis, etc.), and NF-κB signaling pathway. Notably, several previous studies proved that the PI3K-AKT and NF-κB signaling pathways are involved in the occurrence of HUA and renal inflammation.29,53–57 As can be seen in Table 2, seven targets are enriched in the PI3K/AKT pathway (CDK6, SYK, KDR, RELA, PIK3CG, IGF1R, and MCL1) and four targets are enriched in the NF-κB pathway (SYK, PARP1, PTGS2, and RELA). It could be speculated that isorhamnetin might have an effect on the development of HUA and renal inflammation through these two pathways.
3.7 Effects of isorhamnetin on the expression of key proteins of the PI3K/AKT/NF-κB signaling pathway
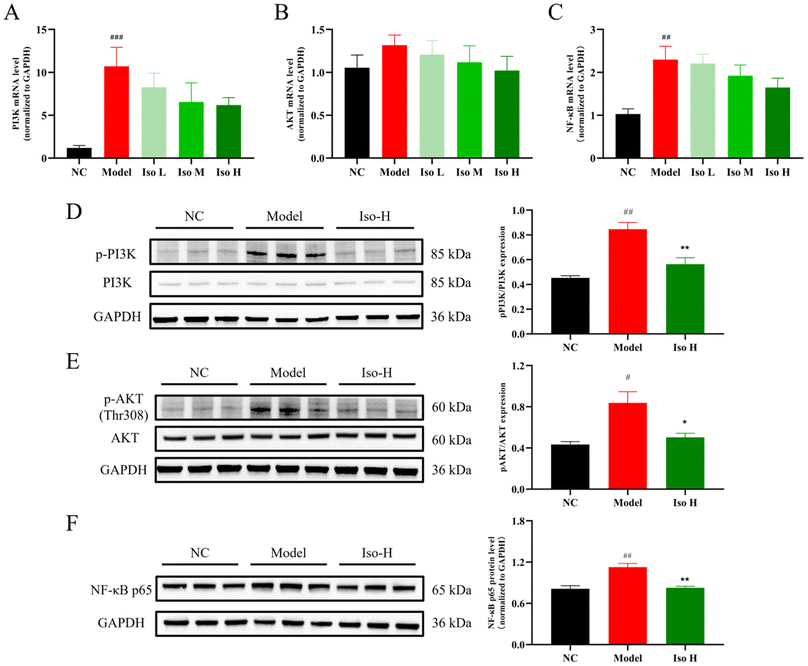
Further experiments were conducted to verify whether the PI3K/AKT/NF-κB signaling pathway was the potential pathway for isorhamnetin to alleviate the HUA-induced renal inflammation in the present study. The relative mRNA expression of PI3K, AKT and NF-κB in the mouse kidneys was first determined by qRT-PCR to explore the expression of the PI3K/AKT/NF-κB pathway at the gene level. As shown in Fig. 8A–C, PI3K and NF-κB mRNA levels were definitely elevated in the model group compared to the NC group (P < 0.001, P < 0.01). Isorhamnetin gavage had a tendency to decrease their expressions, but there was no significant difference. Meanwhile, AKT mRNA expressions also did not change noticeably between groups (P > 0.05). It suggested that the PI3K/AKT/NF-κB pathway might be activated in HUA mice, but isorhamnetin could not regulate the expression of this pathway at the gene level. Further western blot experiments were needed to investigate the expression of key proteins and their phosphorylation in the PI3K/AKT/NF-κB pathway.PI3K, AKT, and NF-κB protein expressions in the kidney tissues of mice in the NC, model, and Iso-H groups were determined using western blot experiments. PI3K is a dimeric complex consisting of a catalytic subunit and a regulatory subunit. It is one of the core proteins in the PI3K/AKT signaling pathway.58 P85, translated by the PI3KR gene, is the regulatory subunit of PI3K. Its phosphorylation level represents the degree of activation of PI3K.59 As shown in Fig. 8D, the p-PI3K/PI3K level in the model group was significantly increased (P < 0.01) compared with the NC group. It suggested that PI3K phosphorylation was activated in HUA mice. Compared with the model group, the p-PI3K/PI3K level in the Iso-H group was observably decreased (P < 0.01), indicating that isorhamnetin could inhibit the phosphorylation of the PI3K protein. The serine/threonine kinase AKT, also known as protein kinase B, is another core protein in the PI3K/AKT pathway.60 In this signaling pathway, the catalytic subunit of PI3K phosphorylated PIP2 to PIP3,59 and then the phosphoryl group at position 3 of PIP3 recruited both PDK1 and AKT proteins to the plasma membrane. Finally, PDK1 phosphorylated Thr308 of AKT proteins, leading to partial activation of them.61 The activated AKT further activated the downstream signaling pathways. The expression of AKT and p-AKT proteins is represented in Fig. 8E. There was no significant change in the total protein expression of AKT in all groups, whereas the expression of p-AKT in the model group was greatly elevated compared with the NC group (P < 0.05) and its level was remarkably decreased in the Iso-H group (P < 0.01). These indicated that AKT was activated in HUA mice and isorhamnetin could inhibit its phosphorylation. Based on the above-mentioned results, the PI3K/AKT pathway was significantly activated in HUA mice, and isorhamnetin could inhibit the expression of this pathway by inhibiting the phosphorylation of PI3K and AKT proteins.
The activated PI3K/AKT signaling would continue to activate downstream pathways, including the NF-κB pathway. Previous studies have demonstrated that the NF-κB pathway is closely related to the development of apoptosis, inflammation, etc.62 NF-κB protein could enter the nucleus and promote inflammation through activating the transcription and translation of TNF-α, IL-1β, and IL-6 genes.49,63 NF-κB could exist in the form of homo- or heterodimers. It consisted of p65 and p50 subunits.64 The expression of the NF-κB p65 subunit was measured in this study (Fig. 8F). Compared with the NC group, p65 expression of mouse kidneys was significantly higher in the model group (P < 0.01), whereas isorhamnetin gavage considerably reduced the level of p65 (P < 0.01). The results suggested that HUA could upregulate the expression of the NF-κB signaling pathway in the kidney, while isorhamnetin was able to downregulate the expression of this pathway. It had been shown that folic acid could improve gut-axis dysfunction and play an ameliorative role in HUA by inhibiting the TLR4/NF-κB signaling pathway.49 Meanwhile, naringenin could regulate renal UA excretion via the PI3K/AKT signaling pathway and relieve renal inflammation via the NF-κB signaling pathway, thus alleviating a series of symptoms of HUA.20 The above-mentioned results indicated that HUA could promote renal inflammation by activating the PI3K/AKT/NF-κB signaling pathway, whereas isorhamnetin could inhibit its activation through down-regulating the expression and phosphorylation of key proteins in this pathway, thus reducing pro-inflammatory cytokines levels (TNF-α, IL-1β, and IL-6) and ultimately alleviating renal inflammation in HUA mice and exerting a protective effect on the kidneys.
4. Conclusion
Isorhamnetin showed remarkable UA lowering effects. It could significantly alleviate HUA in mice through inhibiting the XOD activity to reduce UA production, regulating uric acid transporter proteins (GLUT9, ABCG2, OAT1, and OAT3) to inhibit UA reabsorption and promote UA excretion, and restoring purine metabolism and riboflavin metabolism disorders. Meanwhile, isorhamnetin also significantly alleviated HUA-induced renal inflammation by inhibiting the PI3K/AKT/NF-κB signaling pathway and decreasing the levels of renal inflammatory factors (TNF-α, IL-β, and IL-6) (Fig. 9). This study will lay solid theoretical foundations for the application of isorhamnetin in the field of functional foods or dietary supplements for improving HUA.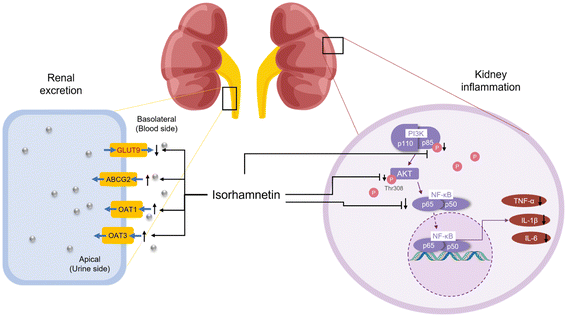 | ||
| Fig. 9 Mechanisms underlying the uric acid lowering and kidney inflammation relieving effects of isorhamnetin. | ||
Abbreviations
| HUA | Hyperuricemia |
| UA | Uric acid |
| XOD | Xanthine oxidase |
| URAT1 | Urate transport 1 |
| GLUT9 | Glucose transporter 9 |
| OAT1 | Organic anion transporter 1 |
| OAT3 | Organic anion transporter 3 |
| ABCG2 | ATP-binding cassette super-family G member 2 |
| All | Allopurinol |
| PO | Potassium oxonate |
| HX | Hypoxanthine |
| CRE | Creatinine |
| BUN | Blood urine nitrogen |
| TNF-α | Tumor necrosis factor-α |
| IL-1β | Interleukin-1β |
| IL-6 | Interleukin-6 |
| PI3K | Phosphoinositide 3-kinase |
| AKT | Protein kinase B |
| NF-κB | Nuclear factor kappa-B |
| GAPDH | Glyceraldehyde 3-phosphate dehydrogenase |
| CMC-Na | Carboxymethylcellulose sodium |
| ELISA | Enzyme-linked immunosorbent assay |
| H&E | Hematoxylin and eosin |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| qRT-PCR | Quantitative reverse transcription polymerase chain reaction |
| PCA | Principal component analysis |
| OPLS-DA | Orthogonal partial least squares discriminant analysis |
| VIP | Variable importance in projection |
| BP | Biological process |
| CC | Cell component |
| MF | Molecular function |
Author contributions
Jiaqing Zhu, Jike Lu, Qiaozhen Kang and Xiaoran Kong participated in the design of the research. Xiaoran Kong, Li Zhao and He Huang gave services to data acquisition and analysis. Xiaoran Kong, Jiaqing Zhu and Jike Lu wrote the rough draft of the article and revised the manuscript. All authors reviewed and endorsed the final manuscript.Data availability
The data supporting this article have been included as part of the ESI.†Conflicts of interest
The authors declare no conflict of interest.Acknowledgements
This work was financially supported by Zhongyuan Sci-Tech Innovation Leading Talents and the Program for Science & Technology Innovation Talents in Universities of Henan Province (22HASTIT037).References
- M. Skoczyńska, M. Chowaniec, A. Szymczak, A. Langner-Hetmańczuk, B. Maciążek-Chyra and P. Wiland, Pathophysiology of hyperuricemia and its clinical significance – a narrative review, Reumatologia, 2020, 58, 312–323 CrossRef PubMed.
- H. Wang, H. Zhang, L. Sun and W. Guo, Roles of hyperuricemia in metabolic syndrome and cardiac-kidney-vascular system diseases, Am. J. Transl. Res., 2018, 10, 2749–2763 CAS.
- C. Li, M.-C. Hsieh and S.-J. Chang, Metabolic syndrome, diabetes, and hyperuricemia, Curr. Opin. Rheumatol., 2013, 25, 210–216 CrossRef CAS PubMed.
- D. I. Feig, D.-H. Kang and R. J. Johnson, Uric acid and cardiovascular risk, N. Engl. J. Med., 2008, 359, 1811–1821 CrossRef CAS PubMed.
- G. Mancia, G. Grassi and C. Borghi, Hyperuricemia, urate deposition and the association with hypertension, Curr. Med. Res. Opin., 2015, 31, 15–19 CrossRef PubMed.
- O. S. P. Sah and Y. X. Qing, Associations between hyperuricemia and chronic kidney disease: A review, Nephro-Urol Mon., 2015, 7, e27233 Search PubMed.
- T. Zhang, R. Ye, Z. Shen, Q. Chang, Y. Zhao, L. Chen, L. Zhao and Y. Xia, Joint association of serum urate and healthy diet with chronic obstructive pulmonary disease incidence: Results from the UK Biobank Study, Food Funct., 2024, 15, 4642–4651 RSC.
- A. A. Alghamdi, J. S. Althumali, M. M. M. Almalki, A. S. Almasoudi, A. H. Almuntashiri, A. H. Almuntashiri, A. I. Mohammed, A. A. Alkinani, M. S. Almahdawi and M. A. H. Mahzari, An overview on the role of xanthine oxidase inhibitors in gout management, Arch. Pharm. Pract., 2021, 12, 94–99 CrossRef.
- L. Xu, Y. Shi, S. Zhuang and N. Liu, Recent advances on uric acid transporters, Oncotarget, 2017, 8, 100852–100862 CrossRef PubMed.
- M. Hosoyamada, K. Ichida, A. Enomoto, T. Hosoya and H. Endou, Function and localization of urate transporter 1 in mouse kidney, J. Am. Soc. Nephrol., 2004, 15, 261–218 CrossRef CAS PubMed.
- F. Preitner, O. Bonny, A. Laverrière, S. Rotman, D. Firsov, A. Da Costa, S. Metref and B. Thorens, Glut9 is a major regulator of urate homeostasis and its genetic inactivation induces hyperuricosuria and urate nephropathy, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 15501–15506 CrossRef CAS PubMed.
- G. Burckhardt, Drug transport by organic anion transporters (OATs), Pharmacol. Ther., 2012, 136, 106–130 CrossRef CAS PubMed.
- Z. Wang, T. Cui, X. Ci, F. Zhao, Y. Sun, Y. Li, R. Liu, W. Wu, X. Yi and C. Liu, The effect of polymorphism of uric acid transporters on uric acid transport, J. Nephrol., 2019, 32, 177–187 CrossRef PubMed.
- H. Matsuo, A. Nakayama, M. Sakiyama, T. Chiba, S. Shimizu, Y. Kawamura, H. Nakashima, T. Nakamura, Y. Takada, Y. Oikawa, T. Takada, H. Nakaoka, J. Abe, H. Inoue, K. Wakai, S. Kawai, Y. Guang, H. Nakagawa, T. Ito, K. Niwa, K. Yamamoto, Y. Sakurai, H. Suzuki, T. Hosoya, K. Ichida, T. Shimizu and N. Shinomiya, ABCG2 dysfunction causes hyperuricemia due to both renal urate underexcretion and renal urate overload, Sci. Rep., 2014, 4, 3755 CrossRef PubMed.
- R. Eckenstaler and R. A. Benndorf, The role of ABCG2 in the pathogenesis of primary hyperuricemia and gout—An update, Int. J. Mol. Sci., 2021, 22, 6678 CrossRef CAS PubMed.
- A. F. G. Cicero, F. Fogacci, M. Kuwabara and C. Borghi, Therapeutic strategies for the treatment of chronic hyperuricemia: An evidence-based update, Medicina, 2021, 57, 58 CrossRef PubMed.
- S. J. Lee and R. A. Terkeltaub, New developments in clinically relevant mechanisms and treatment of hyperuricemia, Curr. Rheumatol. Rep., 2006, 8, 224–230 CrossRef CAS PubMed.
- J. Graessler, A. Graessler, S. Unger, S. Kopprasch, A. Tausche, E. Kuhlisch and H. Schroeder, Association of the human urate transporter 1 with reduced renal uric acid excretion and hyperuricemia in a German Caucasian population, Arthritis Rheum., 2006, 54, 292–300 CrossRef CAS PubMed.
- A. Taniguchi, W. Urano, M. Yamanaka, H. Yamanaka, M. Hosoyamada, H. Endou and N. Kamatani, A common mutation in an organic anion transporter gene, SLC22A12, is a suppressing factor for the development of gout, Arthritis Rheum., 2005, 52, 2576–2577 CrossRef CAS PubMed.
- B. Yang, M. Xin, S. Liang, Y. Huang, J. Li, C. Wang, C. Liu, X. Song, J. Sun and W. Sun, Naringenin ameliorates hyperuricemia by regulating renal uric acid excretion via the PI3K/AKT signaling pathway and renal inflammation through the NF-κB signaling pathway, J. Agric. Food Chem., 2023, 71, 1434–1446 CrossRef CAS PubMed.
- T. Liu, H. Gao, Y. Zhang, S. Wang, M. Lu, X. Dai, Y. Liu, H. Shi, T. Xu, J. Yin, S. Gao, L. Wang and D. Zhang, Apigenin ameliorates hyperuricemia and renal injury through regulation of uric acid metabolism and JAK2/STAT3 signaling pathway, Pharmaceuticals, 2022, 15, 1442 CrossRef CAS PubMed.
- Y.-H. Chang, Y.-F. Chiang, H.-Y. Chen, Y.-J. Huang, K.-L. Wang, Y.-H. Hong, M. Ali, T.-M. Shieh and S.-M. Hsia, Anti-inflammatory and anti-hyperuricemic effects of chrysin on a high fructose corn syrup-induced hyperuricemia rat model via the amelioration of urate transporters and inhibition of NLRP3 inflammasome signaling pathway, Antioxidants, 2021, 10, 564 CrossRef CAS PubMed.
- T. Yong, D. Liang, S. Chen, C. Xiao, X. Gao, Q. Wu, Y. Xie, L. Huang, H. Hu, X. Li, Y. Liu and M. Cai, Caffeic acid phenethyl ester alleviated hypouricemia in hyperuricemic mice through inhibiting XOD and up-regulating OAT3, Phytomedicine, 2022, 103, 154256 CrossRef CAS PubMed.
- X. Zhou, B. Zhang, X. Zhao, Y. Lin, Y. Zhuang, J. Guo and S. Wang, Chlorogenic acid prevents hyperuricemia nephropathy via regulating TMAO-related gut microbes and inhibiting the PI3K/AKT/mTOR pathway, J. Agric. Food Chem., 2022, 70, 10182–10193 CrossRef CAS PubMed.
- Z.-R. Sun, H.-R. Liu, D. Hu, M.-S. Fan, M.-Y. Wang, M.-F. An, Y.-L. Zhao, Z.-M. Xiang and J. Sheng, Ellagic acid exerts beneficial effects on hyperuricemia by inhibiting xanthine oxidase and NLRP3 inflammasome activation, J. Agric. Food Chem., 2021, 69, 12741–12752 CrossRef CAS PubMed.
- H. Yu, Z. Lou, T. Wu, X. Wan, H. Huang, Y. Wu, B. Li, Y. Tu, P. He and J. Liu, Mechanisms of epigallocatechin gallate (EGCG) in ameliorating hyperuricemia: insights into gut microbiota and intestinal function in a mouse model, Food Funct., 2024, 15, 6068–6081 RSC.
- T. Yong, D. Liang, C. Xiao, L. Huang, S. Chen, Y. Xie, X. Gao, Q. Wu, H. Hu, X. Li, Y. Liu and M. Cai, Hypouricemic effect of 2,4-dihydroxybenzoic acid methyl ester in hyperuricemic mice through inhibiting XOD and down-regulating URAT1, Biomed. Pharmacother., 2022, 153, 113303 CrossRef CAS PubMed.
- H. Su, C. Yang, D. Liang and H. Liu, Research advances in the mechanisms of hyperuricemia-induced renal injury, BioMed Res. Int., 2020, 2020, 5817348 CrossRef PubMed.
- X. Wang, L. Dong, Y. Dong, Z. Bao and S. Lin, Corn silk flavonoids ameliorate hyperuricemia via PI3K/AKT/NF-κB pathway, J. Agric. Food Chem., 2023, 71, 9429–9440 CrossRef CAS PubMed.
- G. Gong, Y.-Y. Guan, Z.-L. Zhang, K. Rahman, S.-J. Wang, S. Zhou, X. Luan and H. Zhang, Isorhamnetin: A review of pharmacological effects, Biomed. Pharmacother., 2020, 128, 110301 CrossRef CAS PubMed.
- W.-Q. Li, J. Li, W.-X. Liu, L.-J. Wu, J.-Y. Qin, Z.-W. Lin, X.-Y. Liu, S.-Y. Luo, Q.-H. Wu, X.-F. Xie and C. Peng, Isorhamnetin: A novel natural product beneficial for cardiovascular disease, Curr. Pharm. Des., 2022, 28, 2569–2582 CrossRef CAS PubMed.
- S. Adachi, S. Kondo, Y. Sato, F. Yoshizawa and K. Yagasaki, Anti-hyperuricemic effect of isorhamnetin in cultured hepatocytes and model mice: Structure–activity relationships of methylquercetins as inhibitors of uric acid production, Cytotechnology, 2019, 71, 181–192 CrossRef CAS PubMed.
- F. Wang, X. Zhao, X. Su, D. Song, F. Zou and L. Fang, Isorhamnetin, the xanthine oxidase inhibitor from Sophora Japonica, ameliorates uric acid levels and renal function in hyperuricemic mice, Food Funct., 2021, 12, 12503–12512 RSC.
- L. Zhao, Y. Zhao, X. Kong, H. Huang, L. Hao, T. Wang, Y. Shi, J. Zhu and J. Lu, Deep insights into the mechanism of isorhamnetin's anti-motion sickness effect based on photoshoproteomics, Food Funct., 2024, 15, 10300–10315 RSC.
- Y. M. Roman, The role of uric acid in human health: Insights from the uricase gene, J. Pers. Med., 2023, 13, 1409 CrossRef PubMed.
- Z. Wang, Y. Huang, T. Yang, L. Song, Y. Xiao, Y. Chen, M. Chen, M. Li and Z. Ren, Lactococcus cremoris D2022 alleviates hyperuricemia and suppresses renal inflammation via potential gut-kidney axis, Food Funct., 2024, 15, 6015–6027 RSC.
- R. El Ridi and H. Tallima, Physiological functions and pathogenic potential of uric acid: A review, J. Adv. Res., 2017, 8, 487–493 CrossRef CAS PubMed.
- J. Li, Y. Gong, J. Li and L. Fan, In vitro xanthine oxidase inhibitory properties of Flos Sophorae Immaturus and potential mechanisms, Food Biosci., 2022, 47, 101711 CrossRef CAS.
- K. M. Hoque, E. E. Dixon, R. M. Lewis, J. Allan, G. D. Gamble, A. J. Phipps-Green, V. L. Halperin Kuhns, A. M. Horne, L. K. Stamp, T. R. Merriman, N. Dalbeth and O. M. Woodward, The ABCG2 Q141K hyperuricemia and gout associated variant illuminates the physiology of human urate excretion, Nat. Commun., 2020, 11, 2767 CrossRef CAS PubMed.
- L. Jiang, Y. Wu, C. Qu, Y. Lin, X. Yi, C. Gao, J. Cai, Z. Su and H. Zeng, Hypouricemic effect of gallic acid, a bioactive compound from Sonneratia apetala leaves and branches, on hyperuricemic Mice, Food Funct., 2022, 13, 10275–10290 RSC.
- X.-L. Guo, Y.-Y. Gao, Y.-X. Yang, Q.-F. Zhu, H.-Y. Guan, X. He, C.-L. Zhang, Y. Wang, G.-B. Xu, S.-H. Zou, M.-C. Wei, J. Zhang, J.-J. Zhang and S.-G. Liao, Amelioration effects of α-viniferin on hyperuricemia and hyperuricemia-induced kidney injury in mice, Phytomedicine, 2023, 116, 154868 CrossRef CAS PubMed.
- Z. Huang, W. Zhang, Q. An, Y. Lang, Y. Liu, H. Fan and H. Chen, Exploration of the anti-hyperuricemia effect of TongFengTangSan (TFTS) by UPLC-Q-TOF/MS-based non-targeted metabonomics, Chin. Med., 2023, 18, 17 CrossRef CAS PubMed.
- N. Qin, M. Qin, W. Shi, L. Kong, L. Wang, G. Xu, Y. Guo, J. Zhang and Q. Ma, Investigation of pathogenesis of hyperuricemia based on untargeted and targeted metabolomics, Sci. Rep., 2022, 12, 13980 CrossRef CAS PubMed.
- X. Pang, Z. Guo, L. Ao, Y. Yang, C. Liu, Z. Gu, Y. Xin, M. Li and L. Zhang, Integrated cell metabolomics and serum metabolomics to reveal the mechanism of hypouricemic effect of Inonotus hispidus, J. Funct. Foods, 2023, 105, 105572 CrossRef CAS.
- L. Wu, K. Yi, Z. Xiao, Q. Xia, Y. Cao, S. Chen and Y. Li, A metabolomics perspective reveals the mechanism of the uric acid-lowering effect of Prunus salicina Lindl. cv. “furong” polyphenols in hypoxanthine and potassium oxybate-induced hyperuricemic mice, Food Funct., 2024, 15, 8823–8834 Search PubMed.
- L. Xiang, Y. Huang, R. Li, Y. Tao, T. Wu, S. Pan and X. Xu, Artemisia Selengensis Turcz. leaves extract ameliorates hyperuricemia in mice by inhibiting hepatic xanthine oxidase activity, modulating renal uric acid transporters, and improving metabolic disorders, Food Biosci., 2023, 56, 102639 CrossRef CAS.
- H. Zhao, Z. Lu and Y. Lu, The potential of probiotics in the amelioration of hyperuricemia, Food Funct., 2022, 13, 2394–2414 RSC.
- K. Thakur, S. K. Tomar, A. K. Singh, S. Mandal and S. Arora, Riboflavin and health: A review of recent human research, Crit. Rev. Food Sci. Nutr., 2017, 57, 3650–3660 Search PubMed.
- P. Wang, X. Zhang, X. Zheng, J. Gao, M. Shang, J. Xu and H. Liang, Folic acid protects against hyperuricemia in C57BL/6J mice via ameliorating gut–kidney axis dysfunction, J. Agric. Food Chem., 2022, 70, 15787–15803 CrossRef CAS PubMed.
- X. Lin, Q. Zhou, L. Zhou, Y. Sun, X. Han, X. Cheng, M. Wu, W. Lv, J. Wang and W. Zhao, Quinoa (Chenopodium quinoa, Willd) bran saponins alleviate hyperuricemia and inhibit renal injury by regulating the PI3K/AKT/NFκB signaling pathway and uric acid transport, J. Agric. Food Chem., 2023, 71, 6635–6649 CrossRef CAS PubMed.
- B. Fang, L. Lu, M. Zhao, X. Luo, F. Jia, F. Feng and J. Wang, Mulberry (Fructus mori) extract alleviates hyperuricemia by regulating urate transporters and modulating the gut microbiota, Food Funct., 2024, 15, 12169–12179 RSC.
- L. Sun, Q. Liu, Y. Zhang, M. Xue, H. Yan, X. Qiu, Y. Tian, H. Zhang and H. Liang, Fucoidan from Saccharina japonica alleviates hyperuricemia-induced renal fibrosis through inhibiting the JAK2/STAT3 signaling pathway, J. Agric. Food Chem., 2023, 71, 11454–11465 CrossRef CAS PubMed.
- M.-Q. Zhang, K.-X. Sun, X. Guo, Y.-Y. Chen, C.-Y. Feng, J.-S. Chen, J. C. M. Barreira, M. A. Prieto, J.-Y. Sun, J.-D. Zhang, N.-Y. Li and C. Liu, The antihyperuricemia activity of Astragali Radix through regulating the expression of uric acid transporters via PI3K/Akt signalling pathway, J. Ethnopharmacol., 2023, 317, 116770 CrossRef CAS PubMed.
- L. Xu, J. Cheng, J. Lu, G. Lin, Q. Yu, Y. Li, J. Chen, J. Xie, Z. Su and Q. Zhou, Integrating network pharmacology and experimental validation to clarify the anti-hyperuricemia mechanism of cortex phellodendri in mice, Front. Pharmacol., 2022, 13, 964593 CrossRef CAS PubMed.
- L. Cao, T. Zhao, Y. Xue, L. Xue, Y. Chen, F. Quan, Y. Xiao, W. Wan, M. Han, Q. Jiang, L. Lu, H. Zou and X. Zhu, The anti-inflammatory and uric acid lowering effects of Si-Miao-San on gout, Front. Immunol., 2022, 12, 777522 CrossRef PubMed.
- Z. Liu, H. Xiang, Q. Deng, W. Fu, Y. Li, Z. Yu, Y. Qiu, Z. Mei and L. Xu, Baicalin and baicalein attenuate hyperuricemic nephropathy via inhibiting PI3K/AKT/NF–κB signalling pathway, Nephrology, 2023, 28, 315–327 CrossRef CAS PubMed.
- M.-H. Li, J. Guan, Z. Chen, J.-X. Mo, K.-R. Wu, X.-G. Hu, T. Lan and J. Guo, Fufang Zhenzhu Tiaozhi capsule ameliorates hyperuricemic nephropathy by inhibition of PI3K/AKT/NF-κB pathway, J. Ethnopharmacol., 2022, 298, 115644 CrossRef CAS PubMed.
- Y. Yuan, H. Long, Z. Zhou, Y. Fu and B. Jiang, PI3K-AKT-targeting breast cancer treatments: Natural products and synthetic compounds, Biomolecules, 2023, 13, 93 CrossRef CAS PubMed.
- P. T. Hawkins and L. R. Stephens, PI3K signalling in inflammation, Biochim. Biophys. Acta, 2015, 1851, 882–897 CrossRef CAS PubMed.
- B. D. Manning and A. Toker, AKT/PKB signaling: Navigating the network, Cell, 2017, 169, 381–405 CrossRef CAS PubMed.
- T. F. Franke, S.-I. Yang, T. O. Chan, K. Datta, A. Kazlauskas, D. K. Morrison, D. R. Kaplan and P. N. Tsichlis, The protein kinase encoded by the Akt proto-oncogene is a target of the PDGF-activated phosphatidylinositol 3-kinase, Cell, 1995, 81, 727–736 CrossRef CAS PubMed.
- M. Park and J. Hong, Roles of NF-κB in cancer and inflammatory diseases and their therapeutic approaches, Cells, 2016, 5, 15 CrossRef PubMed.
- Y. Zhou, L. Fang, L. Jiang, P. Wen, H. Cao, W. He, C. Dai and J. Yang, Uric acid induces renal inflammation via activating tubular NF-κB signaling pathway, PLoS One, 2012, 7, e39738 CrossRef CAS PubMed.
- S. Giridharan and M. Srinivasan, Mechanisms of NF-κB p65 and strategies for therapeutic manipulation, J. Inflammation Res., 2018, 11, 407–419 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4fo04867a |
| This journal is © The Royal Society of Chemistry 2025 |