The relative impact of ligand flexibility and redox potential on the activity of Cu superoxide dismutase mimics†
Sharon
Signorella‡
a,
Micaela
Bruno‡
a,
Gianfranco
Frattini
a,
Claudia M.
Palopoli
a,
Diego M.
Moreno
 a,
Nora
Pellegri
b,
Verónica A.
Daier
a,
Nora
Pellegri
b,
Verónica A.
Daier
 *a and
Sandra R.
Signorella
*a and
Sandra R.
Signorella
 *a
*a
aIQUIR (Instituto de Química Rosario), Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario, Suipacha 531, 2000 Rosario, Argentina. E-mail: daier@iquir-conicet.gov.ar; signorella@iquir-conicet.gov.ar
bIFIR (Instituto de Física Rosario), Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Facultad de Ciencias Exactas, Ingeniería y Agrimensura, Universidad Nacional de Rosario, 27 de Febrero 210 bis, 2000 Rosario, Argentina
First published on 31st January 2025
Abstract
Two copper(II) complexes, [Cu(salbn)] and [Cu(py2bn)(OAc)]ClO4, formed with the Schiff-base ligands 1,4-bis(salicylidenamino)butane (H2salbn) and 1,4-bis(pyridin-2-ylmethyleneamino)butane (py2bn), have been prepared and characterized in solid state and in solution, and their ability to catalyse the dismutation of O2˙− has been evaluated in homogeneous medium and immobilized in a mesoporous matrix. The crystal structures show that [Cu(salbn)] possesses a distorted square-planar geometry, while [Cu(py2bn)(OAc)]ClO4 adopts a cis-distorted octahedral geometry. The two complexes experience structural changes in solution, and different spectroscopies were used to examine them. Moreover, their redox potentials are strongly affected by the solvent. In water, the complexes exist as [Cu(salbn)(H2O)] and [Cu(py2bn)(H2O)]2+ with Cu(II)/Cu(I) reduction potential at −361 mV and −229 mV, respectively, well different from redox potentials measured in acetonitrile. Although with a more unfavourable redox potential, [Cu(salbn)(H2O)] reacts with O2˙− faster than [Cu(py2bn)(H2O)]2+, with catalytic rate constants of 3.3 × 107 and 2.9 × 107 M−1 s−1, respectively, at pH = 7.8. Both complexes exhibit higher superoxide dismutase activity than the analogues with a shorter central alkyl chain. The observed catalytic rates essentially correlate with the ligand flexibility, rather than with the redox potential, which is also supported by the slower O2˙− dismutation rate when the complexes are immobilized by encapsulation into the channels of well-ordered mesoporous SBA-15 silica where the pore modifies the complex structures and restraints the ligand rearrangement.
Introduction
The superoxide radical (O2˙−) is a byproduct of respiration and a key component of the immune defence, which together with H2O2, constitute the primary reactive oxygen species (ROS) generated from the leakage of the electron transport chain during the oxidative phosphorylation in humans.1 Under normal conditions, the concentration of O2˙− is maintained at an optimal level by the release of the superoxide dismutase (SOD) enzyme.2 Even when superoxide is a crucial biological messenger and essential antibacterial agent, under certain pathological conditions, like Parkinson’ disease, arthritis, cancer, etc., the O2˙− production overcomes the endogenous defences and tissues become vulnerable to damage.3,4 The use of native SODs for the therapeutic treatment presents several drawbacks including their large size, poor stability under harsh reaction conditions, high cost and immunologic responses.5 To surmount these limitations, efforts have been directed towards the design of low molecular weight metal-based antioxidant catalysts.6–9Among SOD enzymes, copper–zinc SOD (CuZnSOD) catalyses O2˙− dismutation at a CuZn bimetallic centre through a ping-pong mechanism that involves the redox switch of the copper ion between Cu(II)/Cu(I) oxidation states, with Zn(II) engaged in a structural role contributing to the enzyme stability.10,11 Since only the Cu ion participates in the redox cycle of O2˙− dismutation, a number of mononuclear Cu(II) complexes with non-heme ligands bearing N/O donor sites, have been tested as SOD mimics.12–27 It is currently accepted that copper centres with more flexible environment can better accommodate the structural rearrangements that take place when the metal switches between Cu(II) and Cu(I) oxidation states during O2˙− dismutation.28–31 Also, it is known that the metal-centred redox potential is a crucial criterion for the activity of SOD mimics, and it indicates a thermodynamically favoured process when it falls between the one electron reduction and oxidation potentials of O2˙−.32 However, little is known about the relative importance of these two factors on the activity of Cu SOD mimics. Moreover, the redox potential of the copper(II) complexes has rarely been measured in aqueous medium, even when it is well known that the solvent can substantially modify it.33,34 In this work, we report the SOD activity, geometry and redox potential in aqueous solution of copper(II) complexes of two simple tetradentate Schiff-base ligands (Fig. 1), 1,4-bis(pyridin-2-ylmethyleneamino)butane (py2bn) and 1,4-bis(salicylidenamino)butane (H2salbn), bearing the same –(CH2)4– spacer between the two imino N-atoms and well different redox potentials in non-aqueous solvent, with the intention of ascertaining the relative influence of ligand flexibility and redox potential on the reactivity of the copper centre with O2˙−. Furthermore, in order to improve the complex stability and facilitate separation, recovery and handling, the complexes were immobilized by encapsulation into SBA-15 mesoporous silica. This strategy has been successfully employed to control the nuclearity, avoid undesired side reactions and replicate isolation and confinement of the metal centre as in the protein.7,35 SBA-15 silica was selected because of its large surface area, highly ordered mesostructure, biocompatibility and open porosity that allows good accessibility of the substrate to interact with the catalyst.36,37 The SOD activity of the heterogeneous catalysts was evaluated and differences in reactivity with respect to homogeneous complexes interpreted in terms of the geometrical changes that occur within the silica matrix.
Experimental
Materials
In this study, all the used reagents and solvents were commercial products of the highest available purity and, when necessary, were further purified by conventional methods.Synthetic procedures
Ligands N,N′-bis(2-pyridin-2-ylmethylene)butane-1,4-diamine (py2bn), and N,N‘-bis(salicylidene)butane-1,4-diamine (H2salbn) were prepared as previously described by J. Lange et al.24 and E. Kadwa et al.,38 respectively.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300), 273 (22
300), 273 (22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 926), 296 (sh), 368 (9318), 634 (190); in buffer phosphate pH 7.8: 271 (27
926), 296 (sh), 368 (9318), 634 (190); in buffer phosphate pH 7.8: 271 (27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 140), 295 (sh), 355 (10
140), 295 (sh), 355 (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600), 631 (175). Significant IR bands (KBr, ν, cm−1): 3048, 3012, 2931, 2911, 2852, 1620, 1607, 1534, 1471, 1325, 1148, 760. Dark blue crystals suitable for X-ray diffraction were obtained from the reaction mother liquor after 3 days, at 4 °C.
600), 631 (175). Significant IR bands (KBr, ν, cm−1): 3048, 3012, 2931, 2911, 2852, 1620, 1607, 1534, 1471, 1325, 1148, 760. Dark blue crystals suitable for X-ray diffraction were obtained from the reaction mother liquor after 3 days, at 4 °C.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 224), 656 (151); in buffer phosphate of pH 7.8: 281 (15
224), 656 (151); in buffer phosphate of pH 7.8: 281 (15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 360), 407 (sh), 720 (111), 865 (89). Significant IR bands (KBr, ν, cm−1): 3080, 2939, 1644, 1601, 1570, 1478, 1440, 1091, 625. Conductivity: 154 Ω−1 cm2 mol−1 in methanol. Green crystals suitable for X-ray diffraction were obtained from the reaction mother liquor after 3 days, at room temperature.
360), 407 (sh), 720 (111), 865 (89). Significant IR bands (KBr, ν, cm−1): 3080, 2939, 1644, 1601, 1570, 1478, 1440, 1091, 625. Conductivity: 154 Ω−1 cm2 mol−1 in methanol. Green crystals suitable for X-ray diffraction were obtained from the reaction mother liquor after 3 days, at room temperature.
Caution! The perchlorate salts used in this study are potentially explosive and should be handled with care.
Analytical and physical measurements
Infrared spectra were recorded in the 4000–400 cm−1 range on a PerkinElmer Spectrum One FT-IR spectrophotometer provided with a DTGS detector, resolution = 4 cm−1 and 10 accumulations. FT-IR spectra were registered from KBr sample pellets. Electronic spectra were recorded on a Jasco V-550 spectrophotometer. An Inductively coupled plasma mass spectrometer (ICP-MS) PerkinElmer NexION 350× was used to measure the metal content. For quantification of Cu in the hybrid materials, the samples (15 mg) were treated with 1 mL of H2SO4 and 0.5 mL of HNO3 and heated at 100 °C for 3 h. The solid was filtered and the filtrate diluted to a final volume of 25 mL. Cu in solution was analysed by ICP-MS. The copper content was verified by measuring the complex remaining in solution after the encapsulation. In this case, after filtering and washing the hybrid material, the filtrate was diluted to 250 mL. Then, 5 mL of this solution were diluted to 1 L with deionized water and Cu in solution was analysed by ICP-MS. Both methods afforded the same result. Each sample was prepared in triplicate. CHN analyses were performed on a PERKIN ELMER 2400 series II Analyzer. Electron Paramagnetic Resonance (EPR) spectra were obtained at 120 K on a Bruker EMX-Plus spectrometer with a microwave frequency of ∼9.3 GHz. Simulations were performed in MATLAB using the EasySpin 6.0.6 program package for EPR.391H NMR spectra were recorded on a Bruker AC 400 NMR spectrometer at ambient probe temperature (ca. 25 °C). Chemical shifts (in ppm) are referenced to tetramethylsilane and paramagnetic NMR spectra were acquired employing superWEFT sequence, with acquisition time of 80 ms. Conductivity measurements were performed on 1.0 mM solutions of the complexes using a Horiba F-54 BW conductivity meter. The electrochemical experiments were performed with a Princeton Applied Research potentiostat, VERSASTAT II model, with the 270/250 Research Electrochemistry Software. Studies were carried out with a standard three-electrode cell, under Ar, using 0.1 M Bu4NBF4 as a supporting electrolyte and ≈10−3 M of the complex. The working electrode was a glassy carbon disk, and the reference electrode was a saturated calomel electrode (SCE) isolated in a fritted bridge with a Pt wire as the auxiliary electrode. Under the employed conditions, E(ferrocene/ferrocenium) = 388 mV in MeCN, 474 mV in DMF, 409 mV in MeOH (vs. SCE), at room temperature. Studies in aqueous buffered solutions were carried out on ∼10−3 M of the complex in 50 mM buffer phosphate, pH = 7.8. Under these conditions, the reduction peak of the Zobell's solution = 185 mV vs. SCE. The size and morphology of the solid materials were analysed using an AMR 1000 Leitz scanning electron microscope (SEM) operated at variable accelerating voltages and with EDX detector NORAN System SIX NSS-200. Samples for SEM observation were prepared by dispersing a small amount of powder of dry silica and hybrid samples on double-sided conductive adhesive tabs on top of the SEM sample holders. Then, the samples were covered by a thin layer of gold deposited by sputtering to avoid charge accumulation on the surfaces. The selected accelerating voltage used in the shown images was 20 kV at high vacuum condition. Transmission electron microscopy (TEM) analysis was performed with a TEM/STEM JEM 2100 Plus with operational voltage of 200 kV (variable), with a LaB6 filament. The samples were prepared by placing a suspension of silica and hybrid samples in ethanol onto a square mesh copper grid (400 mesh), coated with a layer of Formvar and carbon. The suspension of material in ethanol was left to dry, evaporating the ethanol, leaving the dispersed particles to adhere to the Formvar/Carbon surface. SEM and TEM images were processed using public domain ImageJ program. N2 adsorption–desorption isotherms were obtained at 77 K on a Micrometric ASAP 2020 V4.02 (V4.02 G) apparatus. The samples were degassed at 10−3 Torr and 200 °C for 6 h prior to the adsorption experiment. Surface area (SBET) was calculated using the Brunauer–Emmett–Teller (BET)40 equation over the pressure range (p/p0) of 0.05–0.20. The volume of micropores and mesopores (VμP and VMP) was determined by the alpha-plot method using the standard Licrospher isotherm. The total pore volume (VTP) was determined with the Gurvich rule40 at 0.98p/p0. The pore size distributions were calculated using the Villarroel–Bezerra–Sapag (VBS) model,41 on the desorption branch of the N2 isotherms.Crystal data collection and refinement
Crystallographic data for compounds [Cu(salbn)] and [Cu(py2bn)(OAc)]ClO4 were collected at 298(2) K on a Bruker D8 QUEST ECO Photon II CPAD Diffractometer, using graphite monochromated Mo-Kα radiation (λ = 0.71073 Å). Data collection was carried out using the Bruker APEX342 and the Bruker APEX443 packages, cell refinement and data reduction were done with the program SAINT V8.40B.44 The structures were solved by direct methods with SHELXT V 2018/2.45and refined by full matrix least-squares on F2 data with SHELXL-2019/1.46 Crystal data collection and the results of the refined parameters for compounds [Cu(salbn)] and [Cu(py2bn)(OAc)]ClO4 are summarized in Table S1.† Molecular graphics were done with ORTEP-3,47 with 50% probability displacement ellipsoids. Crystallographic data for [Cu(salbn)] and [Cu(py2bn)(OAc)]ClO4 has been deposited at the CCDC under 2405330 and 2405337, respectively.†SOD activity measurement
The SOD activity of the free and immobilized complexes was determined by an indirect assay based on the inhibition of the photoreduction of nitro blue tetrazolium (NBT).48 In a typical experiment, the reaction mixture (3.2 mL) containing riboflavin (3.35 μM), methionine (9.52 mM), NBT (38.2 μM) and different concentrations of the complex either free (diluted aliquots (50 μL) of the complex in methanol added to the reaction mixture) or immobilized, was prepared in 50 mM phosphate buffer of pH 7.8. Riboflavin was added last and the mixture was illuminated for 15 min with a 20 W led lamp placed at a distance such that the rate of increase in absorbance was 0.017 min−1, at 25 °C. The reduction of NBT was measured at 560 nm. Control reactions were performed to verify that the complexes did not react with NBT or riboflavin directly. Inhibition percentage (IC) was calculated according to:The IC50 values were determined from plots of % inhibition vs. catalyst concentration and used to calculate the McCord–Fridovich second-order rate constant (kMcF).49 At 50% inhibition, O2˙− reacts with NBT and the mimic at the same rate, so kMcF IC50 = kNBT [NBT], with kNBT (pH = 7.8) = 5.94 × 104 M−1 s−1.
Computational calculations
All calculations were performed at B3LYP/6-31G** level50 using Gaussian 16 package.51 The solvent effects were computed by the polarizable continuum model (PCM)52 with the DMF, DMSO or water dielectric constants. All the optimized structures are proved to be local minimum using frequency calculations.Results and discussion
Geometrical and electronic properties of catalysts in solid state and solution
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching band at 1635 cm−1 in H2salbn shifts to 1627 cm−1 in the complex as a consequence of the Cu(II) coordination to the imino N-atom, while the peak at 1379 cm−1, assigned to O–H in plane bending vibration of the phenol groups in the free ligand, is absent in the spectrum of the complex, as expected for deprotonated phenol groups bound to copper.
N stretching band at 1635 cm−1 in H2salbn shifts to 1627 cm−1 in the complex as a consequence of the Cu(II) coordination to the imino N-atom, while the peak at 1379 cm−1, assigned to O–H in plane bending vibration of the phenol groups in the free ligand, is absent in the spectrum of the complex, as expected for deprotonated phenol groups bound to copper.
 | ||
| Fig. 2 (a) Molecular structure of [Cu(salbn)] at the 50% probability level with atom numbering. (b) FT-IR spectra of H2salbn and [Cu(salbn)]. | ||
The low-temperature EPR spectrum of a polycrystalline sample of [Cu(salbn)] (Fig. 3(a)) shows an anisotropic signal for which the principal components of the g-tensor gz = 2.20, and gy = 2.06, gx = 2.05 were obtained by simulation of the spectrum (Fig. S1(a)†). These values are consistent with the distorted square-planar geometry of the Cu(II) centre observed in the crystals. The lack of hyperfine structure in the powdered sample is the result of strong exchange interactions between neighbouring Cu(II) centres. In frozen DMSO glass, the EPR spectrum of [Cu(salbn)] shows axial anisotropy with the expected hyperfine splitting pattern for an I = 3/2 nucleus of naturally abundant Cu along the direction parallel to the magnetic field (Fig. 3(a)). The simulation of the spectrum (Fig. S1(b)†) afforded spectral parameters g∥ = 2.20, A∥ = 173 × 10−4 cm−1, g⊥ = 2.10, typical of Cu(II) complex possessing a dx2−y2 ground state. The calculated f-factor (g∥/A∥) = 127 cm for this complex is in the range of 105 to 135 cm expected for tetrahedrally distorted square-planar geometry.58 This tetrahedral distortion decreases as the central chain shortens. Therefore, f[Cu(salbn)] > f[Cu(salpn)] (119 cm in frozen DMSO)59 > f[Cu(salen)] (108 in frozen DMF),60 the last with the CuN2O2 chromophore arranged almost in the plane.
The absorption spectra of [Cu(salbn)], shown in Fig. 3(b), exhibit an intense band at 273 nm and a shoulder around 295 nm corresponding to π → π* and n → π* intraligand transitions. The band at 355–377 nm experiences a bathochromic shift as the solvent donor capacity increases and can be assigned to the phenolate-to-copper(II) charge transfer (LMCT) process. The d–d transitions are observed as a broad absorption band centred at 655 nm (ε = 264–269 M−1 cm−1) in DMF and MeCN solution. This band appears at wavelengths longer than observed for [Cu(salpn)] (605 nm in toluene or Cl3CH) or [Cu(salen)] (565 nm in toluene),54 consistent with a more distorted geometry around the copper ion in [Cu(salbn)], in agreement with the EPR results. The position of the d–d transitions varies with the solvent, and shifts to slightly shorter wavelengths (631–634 nm, ε = 175–190 M−1 cm−1) in water or methanol, as shown in Fig. 3(b). Both, the lower energy and higher molar intensity of the d–d transitions support tetrahedral perturbation of the tetragonal Cu(II) environment in both DMF and MeCN greater than in methanol or water.33,60,61
Further evidence of the coordination of the ligand to Cu(II) is provided by the 1H NMR spectrum of the complex (Fig. 3(c)) that shows the broadening and shift of the resonances of the aromatic ring protons as well as those from the azomethine protons as a result of the paramagnetic relaxation induced by the Cu(II) ion. Aromatic protons undergo differential isotropic shift depending on the distance to the metal centre. Therefore, the broad signal at 31 ppm can be assigned to the azomethine protons, H4/H4′ are observed at 10.5 ppm with peak half-width ω = 189 Hz, H5/H5′ and H6/H6′ protons appear overlapped at 5.6 ppm (ω = 95 Hz), and the resonance shifted up-field at −10 ppm can be attributed to H3/H3′ on the basis of comparison with reported spectra for related Cu(II) complexes.62 Given the large relaxation values of the phenolate proton signals, distortions from the square-planar geometry are not resolved and protons belonging to both aromatic rings cannot be distinguished. In addition, the coordination to the metal also affects the methylene resonances, displaying broad peaks that span in the 0.5–3 ppm range. The spectral assignment was verified recording 1H NMR spectra of H2salbn after addition of increasing amounts of Cu(ClO4)2. The intensity of the broad resonances grows with increasing amounts of salt as a result of metal coordination (shown in the inset of Fig. 3(c) for the peak of H4), while those of the free ligand decrease and disappear. For the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 metal-to-ligand mixture the spectrum is analogous to that of the solution of the complex.
1 metal-to-ligand mixture the spectrum is analogous to that of the solution of the complex.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N and pyridine ring stretching vibrations at 1644 and 1601 cm−1, respectively, two strong absorption bands at 1091 and 625 cm−1 corresponding to free perchlorate, and bands at 1652 (overlapped to the azomethine stretching band) and 1478 cm−1 assignable to the asymmetrical and symmetrical stretching modes of bound acetate (Fig. 4(a)). This spectral pattern is retained in the FT-IR spectrum of crystals of [Cu(py2bn)(OAc)]ClO4 formed from the mother liquors (Fig. S2†), evidencing that the crystal structure is analogous to that of the powdered compound.
N and pyridine ring stretching vibrations at 1644 and 1601 cm−1, respectively, two strong absorption bands at 1091 and 625 cm−1 corresponding to free perchlorate, and bands at 1652 (overlapped to the azomethine stretching band) and 1478 cm−1 assignable to the asymmetrical and symmetrical stretching modes of bound acetate (Fig. 4(a)). This spectral pattern is retained in the FT-IR spectrum of crystals of [Cu(py2bn)(OAc)]ClO4 formed from the mother liquors (Fig. S2†), evidencing that the crystal structure is analogous to that of the powdered compound.
 | ||
| Fig. 4 (a) FT-IR spectra of py2bn and [Cu(py2bn)(OAc)]ClO4. (b) Molecular structure of the complex cation [Cu(py2bn)(OAc)]+ at the 50% probability level. | ||
The molecular structure of the complex cation determined by X-ray diffraction analysis is shown in Fig. 4(b), and selected bond lengths and angles are listed in Table S3.† The compound crystallizes in the P21/n space group with the asymmetric unit containing a discrete Cu(II) complex cation and one non-coordinated ClO4− anion. The N4-tetradentate Schiff-base ligand is tightly bound to the Cu(II) ion via the two Nimine and two Npy atoms adopting a cis-β-configuration, in which the angle between the two aromatic ring planes is 102.6°. The two other coordination positions are occupied by acetate acting as a bidentate ligand bound to copper asymmetrically, with one short Cu1–O1 bond of 1.934(5) Å and one long Cu1–O2 distance of 2.754(5) Å. In line with this, the C11–O2 bond length is shorter than the C11–O1 one in the carboxylate group. The overall geometry may be described as cis-distorted octahedral63 with large deviations of bond angles around Cu(II), up to 24° from 90°.
The low temperature X-band EPR spectrum recorded on the powdered complex (Fig. 5(a)) shows an anisotropic signal with g1 = 2.05, g2 = 2.12 and g3 = 2.20 (obtained from the simulated spectrum, Fig. S3(a)†) consistent with the rhombically distorted octahedral geometry around the Cu(II) ion observed in the crystal structure and also found in other CuN4O2 chromophores with asymmetrically coordinated acetate groups.64,65 The EPR spectra of the powder show only slight variation with increasing temperature from 120 K to 290 K, compatible with the retention of the cis-distorted octahedral configuration. In DMSO and 120 K, the spectrum is still broad and anisotropic, with g components at g1 = 2.02, g2 = 2.05, and g3 = 2.11 (Fig. 5(a) and Fig. S3(b)†), suggesting that the rhombic distortion persists, probably with a less constrained geometry in solution. This was confirmed by optimizing the geometry of the complex using DFT calculations with DMF (as well as DMSO) as the implicit solvent, where the lowest-energy species corresponds to [Cu(py2bn)(OAc)]+ retaining essentially the same geometry observed in the crystal (Table S3 and Fig. S4(a)†). Electronic spectra of [Cu(py2bn)(OAc)]+ in different solvents are shown in Fig. 5(b). In the UV region the complex displays intense absorption bands around 280 nm originated from intraligand and charge transfer transitions, whereas in the visible region, the absorption features depend on the solvent. In DMF, the spectrum displays a weak broad band centred at 608 nm (ε = 206 M−1 cm−1), that can be assigned to d–d transitions in a distorted tetragonal environment. As observed in the inset of Fig. 5(b), the d–d transitions shift to the red in protic solvents. In methanol λmax = 656 nm with a shoulder at longer wavelengths, while in aqueous medium deconvolution of the d–d transitions afforded two bands at λmax = 720 and 865 nm. This trend suggests solvolysis of acetate in the protic solvents to form [Cu(py2bn)(solv)]2+. These two d–d bands are characteristic of the distortion of the Cu(II) centre towards trigonal bipyramidal geometry.66,67 This geometry is in line with the preference for the trigonal bipyramidal coordination reported for the related complexes [Cu(6-Me-py2bn)(H2O)]2+![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 25 and [Cu(py2bn)Cl]+,68 with water/chloride occupying an equatorial position. Acetate dissociation in the protic solvent was also confirmed by the molar conductivity of 154 Ω−1 cm2 mol−1 of the complex measured in methanol, a value expected for a 1
25 and [Cu(py2bn)Cl]+,68 with water/chloride occupying an equatorial position. Acetate dissociation in the protic solvent was also confirmed by the molar conductivity of 154 Ω−1 cm2 mol−1 of the complex measured in methanol, a value expected for a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 electrolyte in this solvent.69 The optimized structure of [Cu(py2bn)(HOAc)]2+ in water as implicit solvent, shows that the Cu–O1 bond lengthens when acetate protonates and that the preferred structure for the protonated complex is a trigonal bipyramid with the HOAc coligand occupying an equatorial position (shown in Fig. S4(b) and Table S3†). DFT calculations show that the labilised HOAc ligand can be substituted by water when the entering water molecule approaches the edge opposite to the equatorial pyridine of the trigonal bipyramidal complex (the attack on another position did not lead to the substituted product), as can be seen in Fig. S5.† [Cu(py2bn)(H2O)]2+ retains the geometry with water placed in the equatorial plane and Cu–OH2 bond distance of 2.085 Å. For comparative purposes, in [Cu(py2en)]2+ (py2en = N,N′-bis(2-pyridin-2-ylmethylene)etane-1,2-diamine), where the metal has a distinctively flat CuN4 plane as part of a tetragonally elongated octahedral geometry,70 the d–d bands are centred at 612 nm in MeCN, while for [Cu(py2pn)(DMF)]2+ (py2pn = N,N′-bis(2-pyridin-2-ylmethylene)etane-1,3-diamine) with a square-pyramidal structure in solution, the d–d transitions are centred at 626 nm in DMF.15
2 electrolyte in this solvent.69 The optimized structure of [Cu(py2bn)(HOAc)]2+ in water as implicit solvent, shows that the Cu–O1 bond lengthens when acetate protonates and that the preferred structure for the protonated complex is a trigonal bipyramid with the HOAc coligand occupying an equatorial position (shown in Fig. S4(b) and Table S3†). DFT calculations show that the labilised HOAc ligand can be substituted by water when the entering water molecule approaches the edge opposite to the equatorial pyridine of the trigonal bipyramidal complex (the attack on another position did not lead to the substituted product), as can be seen in Fig. S5.† [Cu(py2bn)(H2O)]2+ retains the geometry with water placed in the equatorial plane and Cu–OH2 bond distance of 2.085 Å. For comparative purposes, in [Cu(py2en)]2+ (py2en = N,N′-bis(2-pyridin-2-ylmethylene)etane-1,2-diamine), where the metal has a distinctively flat CuN4 plane as part of a tetragonally elongated octahedral geometry,70 the d–d bands are centred at 612 nm in MeCN, while for [Cu(py2pn)(DMF)]2+ (py2pn = N,N′-bis(2-pyridin-2-ylmethylene)etane-1,3-diamine) with a square-pyramidal structure in solution, the d–d transitions are centred at 626 nm in DMF.15
The 1H NMR spectrum of the paramagnetic [Cu(py2bn)(OAc)]+ complex was recorded in Cl3CD in the 50 to −50 ppm spectral range. Due to the copper(II) paramagnetic effect, all resonances are broadened to some extent, and broadening increases with the isotropic shift, so that a number of resonance lines become extremely wide precluding their detection. Fig. 5(c) shows the peaks that could be measured for this system using a superWEFT sequence with an acquisition time of 80 ms. The signals in the 0–2 ppm range can be assigned to the methylene protons of the alkyl chain of the ligand.71 The resonance of the –(CH2)2– moiety at δ = 1.75 is slightly shifted from its position in the free ligand and broadens (ω = 68 Hz). The two up-field peaks at 1.26 ppm (ω = 36 Hz) and 0.84 ppm (ω = 31 Hz) can correspond to diastereotopic methylene –CH2–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) , while the broad resonance at 14.7 ppm (ω = 650 Hz) is compatible with the methyl acetate shifted down-field as a result of spin density delocalization through contact shift.72 For the pyridine rings, it is expected that all protons exhibit a downfield shift from their respective positions in the free ligand arising from the positive spin density generated through the σ-metal-pyridine interactions, with downfield shifts for Hα-py larger than for Hβ,γ-py.73 The broad signal observed in the spectrum at 10.3 ppm (ω = 162 Hz) could arise from the Hβ/γ of one of the non-equivalent pyridine rings of the complex, the other resonances probably broaden beyond recognition under the experimental conditions used here or are outside the spectral sweep window.
, while the broad resonance at 14.7 ppm (ω = 650 Hz) is compatible with the methyl acetate shifted down-field as a result of spin density delocalization through contact shift.72 For the pyridine rings, it is expected that all protons exhibit a downfield shift from their respective positions in the free ligand arising from the positive spin density generated through the σ-metal-pyridine interactions, with downfield shifts for Hα-py larger than for Hβ,γ-py.73 The broad signal observed in the spectrum at 10.3 ppm (ω = 162 Hz) could arise from the Hβ/γ of one of the non-equivalent pyridine rings of the complex, the other resonances probably broaden beyond recognition under the experimental conditions used here or are outside the spectral sweep window.
Electrochemical studies
The potential of the Cu(II)/Cu(I) redox couple depends on the metal first coordination sphere and the flexibility of the chelate ring, which in turn control the reactivity of the metal centre in electron transfer reactions. Therefore, the electrochemical behaviour of the complexes was studied by cyclic voltammetry in different solvents (Fig. 6). In [Cu(salbn)], the N2O2 first coordination sphere of the metal stabilizes the Cu(II) oxidation state showing a quasi-reversible redox wave at −980 mV vs. SCE in MeCN, characteristic of the Cu(II)/Cu(I) couple in an elongated tetragonal environment.33 In DMF, the redox couple shifts to −901 mV vs. SCE, most likely with DMF at the apex of a slightly distorted square-pyramid. The adoption of a flatter conformation with lowering in the flexibility of the ligand as the length of the alkyl spacer decreases is reflected in the more negative redox potential values reported for the Cu(II)/Cu(I) redox couple of [Cu(salpn)] (EDMF1/2: −1016 mV vs. SCE) and [Cu(salen)] (EDMF1/2: −1176 mV vs. SCE).74 In agreement with the spectroscopic results described above, the anodic and cathodic waves of the Cu(II)/Cu(I) couple of [Cu(salbn)] shift to less negative potentials with the decreasing donor capacity of the solvent to bind copper at the apex of a square-pyramid (Fig. 6(a)).33 Therefore, EDMF1/2: −901 mV < EMeOH1/2: −618 mV <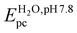 : −361 mV vs. SCE.
: −361 mV vs. SCE.
 | ||
| Fig. 6 Cyclic voltammograms of (a) [Cu(salbn)] and (b) [Cu(py2bn)(AcO)]+ in different media. Scan rate: 100 mV s−1. | ||
The absence of a redissolution peak in the anodic scan, indicates that, even in the aqueous buffer, no copper is released from the Cu(I) complex generated at the electrode. In the aqueous medium, the irreversibility of the process is probably related to the dissociation of the labile water molecule during reduction of [Cu(salbn)(OH2)] to form the negatively charged tetracoordinated Cu(I) complex [Cu(salbn)]− which is then re-oxidised at  in the anodic scan.
in the anodic scan.
In MeCN, [Cu(py2bn)(OAc)]+ exhibits a quasi-reversible redox wave at 93 mV corresponding to the Cu(II)/Cu(I) couple (Fig. 6(b)). The large peak to peak separation ΔE = 186 mV, suggests that geometrical changes occur during the redox process. Probably the long Cu–Npy bond breaks in the reduced form of the complex, retaining the bound acetate, so that Cu(I) can adopts a flattened tetrahedral geometry. Then, pyridine re-binds the metal in the oxidized complex to adopt the starting geometry. This electrochemical behaviour is different from that observed for [Cu(py2bn)]2+ in acetonitrile which shows a cathodic peak at 195 mV corresponding to the Cu(II)/Cu(I) reduction, and two oxidation peaks at 289 and 519 mV.68 It has been reported that in MeCN, when the complex concentration is >5 × 10−5 M, [Cu(I)(py2bn)]+ is in equilibrium with the dimer [Cu(I)2(py2bn)2]2+, and the two observed anodic waves were assigned to [Cu(I)(py2bn)]+ → [Cu(II)(py2bn)]2+ and [Cu(I)2(py2bn)2]2+ → 2 [Cu(II)(py2bn)]2+ oxidation processes, respectively.68 The oxidation of the metal in the second process is accompanied by partial dissociation of the ligand so that oxidation of the Cu(I) centre in the dimer should require more energy than in the monomer. In the present study, [Cu(py2bn)(OAc)]+ shows a similar electrochemical behaviour in methanol. As described before, in this protic solvent, acetate can protonate and dissociate affording [Cu(py2bn)(MeOH)]2+ which is reduced at Epc = 49 mV, while [Cu(I)(py2bn)]+ is oxidized at Epa1 = 183 mV and [Cu(I)2(py2bn)2]2+ at Epa2 = 319 mV. A behaviour that also agrees with the molar conductivity of the complex in methanol. Additionally, the formation of the solvolyzed complex was verified by the identical voltammogram obtained for [Cu(py2bn)]2+ generated in situ by mixing py2bn and Cu(ClO4)2 in methanol. In water, acetate hydrolyses, and, as described previously, the aquo complex adopts a trigonal–bipyramidal geometry, with water bound to Cu(II) in the equatorial plane. Therefore, the reduction peak at Epc = −229 mV must involve dissociation of the water molecule from the meridional plane and geometrical rearrangement of the ligand, while the re-oxidation process of the tetracoordinated Cu(I) to the aquo-Cu(II) complex occurs at Epa = 159 mV. In water, no dimerization was observed. The redox potentials of the two complexes in different solvents are summarized in Table S4.†
As expected, the N2O2-donor set of salbn stabilizes the Cu(II) oxidation state more than the N4-donor set of py2bn. Therefore, in aqueous buffer, [Cu(salbn)] is reduced at a more negative potential than [Cu(py2bn)]2+. Anyway, it must be emphasized that in the aqueous buffer the redox potentials of both complexes lay between the redox couples E(O2˙−/H2O2) = 642 mV and E(O2/O2˙−) = −404 mV vs. SCE, at pH 7,75 and thus, they are suitable to act as SOD mimics.
Characterization of the heterogeneous catalysts
Complexes [Cu(salbn)] and [Cu(py2bn)]2+ were successfully encapsulated inside the channels of well-ordered mesoporous SBA-15 silica particles through wet impregnation, yielding Cu-salbn@SBA-15 and Cu-py2bn@SBA-15. The loading of the divalent complex cation was more than 5-times higher than the neutral complex as the result of the strong electrostatic interactions with the negative silanolate groups of the pores surface that step-up the retention of the cationic complex through ionic exchange with the silanol protons. In the FT-IR spectra (Fig. S6 and S7†) of both hybrid materials, the strong bands of the Si–O–Si framework and Si–O/OH2 groups mask those from the complexes, even in the case of the material where the catalyst is present in higher proportion. However, a few weak bands are not completely hidden by those of the silica matrix and can be observed in the 3000–2800, 1600–1350 and 700–600 cm−1 spectral regions.The textural properties of SBA-15 and the hybrid materials were analysed by nitrogen adsorption–desorption measurements at 77 K. All the samples exhibit type IV isotherms with an H1 hysteresis loop at relative pressure p/p0 = 0.55–0.75 (Fig. 7(a and b)) typical of ordered mesoporous materials, and a steep step distinctive of the uniform mesopore size distribution. The plateau after the sharp step indicates the lack of significant secondary mesoporosity. Textural data of the mesoporous materials are summarized in Table 1. The prepared SBA-15 possesses a high specific surface area and pore width of 5.2 nm appropriate for hosting the complexes, which are ≈0.93–1.25 nm wide calculated from the crystal structures described above.
| S BET (m2 g−1) | V μP (cm3 g−1) | V MP (cm3 g−1) | V TP (cm3 g−1) | w P (nm) | mmol complex per 100 g material | |
|---|---|---|---|---|---|---|
| V TP = VμP + Vprimary MP + Vsecondary MP, MP = mesopore; μP = micropore; wp = pore diameter. | ||||||
| SBA-15 | 481 | 0.06 | 0.57 | 0.63 | 5.2 | — |
| Cu-salbn@SBA-15 | 375 | 0.03 | 0.47 | 0.50 | 5.2 | 5.2 |
| Cu-py2bn@SBA-15 | 261 | 0.01 | 0.37 | 0.43 | 5.4 | 28.1 |
The insertion of the catalyst causes the decrease of the BET surface area and total pore volume, indicating the compound is filling the pores, but leaves the overall shape of the BET curves almost unchanged, as well as the pore diameter, suggesting the ordered mesostructure of SBA-15 remains invariant, as confirmed through transmission electron microscopy (TEM) measurements described below. Cu-salbn@SBA-15 experiences a smaller decrease in the overall adsorption volume and surface area compared to Cu-py2bn@SBA-15, in line with the lower proportion of complex in the former, determined by ICP analysis.
The morphology and size of the particles of SBA-15 and the hybrid materials were analysed by scanning electron microscopy (SEM). SBA-15 particles possess elongated shapes of 1.1 ± 0.2 mm length with average aspect ratio of 1.8 ± 0.5, calculated from statistical analysis over 309 particles, that are lined up forming chains of 10–30 μm long (Fig. S8(a) and S9(a)†). Particles of Cu-salbn@SBA-15 are somewhat smaller than SBA-15 ones and retain their shape and arrangement (Fig. 7(c)), whereas Cu-py2bn@SBA-15 develops entangled assemblies of more irregular spindly particles (Fig. 7(d)). The average aspect ratios calculated from Image-J analysis of SEM images of selected particles are 2.0 ± 0.4 and 1.8 ± 0.5 for Cu-salbn@SBA-15 and Cu-py2bn@SBA-15, respectively, and the corresponding histograms are shown in Fig. S9(b and c).†
TEM images of the hybrid materials display a regular array of cylindrical channels along the long axis of the particles (Fig. 7(e and f)), confirming that incorporation of the complexes inside the pores preserve the highly ordered mesostructure of the SBA-15 (Fig. S8(b)†). Integration over the grey scales in carefully selected zones where the electron beam was perpendicular to the channels, afforded average pore diameter of 4.5 ± 0.6 and 4.3 ± 0.9 nm, and wall thickness of 3.5 ± 0.5 and 3.5 ± 0.6 nm, for Cu-salbn@SBA-15 and Cu-py2bn@SBA-15, respectively (Fig. S10†), which are in the order of pores size calculated from the adsorption isotherms. These channels reach the particle surface and adopt a hexagonal arrangement that can be observed in the images taken when the electron beam is parallel to the pores (Fig. S11†). This open porosity serves as an entry channel to the substrate to interact with the complex inside the pores, an essential aspect for catalysis.
SOD activity studies
As described previously, in water the two complexes possess labile coordination sites and display redox potentials appropriate to dismutate O2˙−, so that they are expected to act as SOD mimics. Before carrying out the SOD activity tests, the stability of the catalysts in phosphate buffer was verified. For this, the free and immobilized complexes were incubated in a phosphate buffer of pH 7.8. UV-vis spectra of the free complexes recorded during an interval of 2 h showed constant spectral features and band intensities (Fig. S12†), evidencing the stability of the complexes in the reaction medium of the SOD assay. For the hybrid materials, the UV-vis spectra were registered on the supernatant after centrifugation of the suspensions. Spectra taken at different incubation times were compared to those of the free complexes in solution, verifying negligible complex leaching (Fig. S13†), even in the case of the neutral complex. This behaviour highlights the efficiency of encapsulation that retains the complex inside the silica channels, and was confirmed by high angle X-ray diffractograms of Cu-salbn@SBA-15, which do not show peaks associated to crystallites of [Cu(salbn)] on the external surface of the silica particles (Fig. S14†).The SOD activity of free and encapsulated [Cu(salbn)] and [Cu(py2bn)]2+ was examined by the Beauchamps and Fridovich indirect assay using nitroblue tetrazolium (NBT) in phosphate buffer of pH 7.8.48 In this test, the catalyst competes with NBT for reacting with the photogenerated O2˙−, and the reduction of NBT by O2˙− leads to the formation of deep blue formazan determined at 560 nm after irradiating the reaction mixtures during a fixed time. In this way, the SOD activity of the catalysts is inversely related to the concentration of formazan, and the catalyst concentration required to decrease by 50% the reduction of NBT by O2˙− (IC50), can be determined from plots of % inhibition vs. [catalyst]. The IC50 values calculated for [Cu(salbn)] and [Cu(py2bn)]2+ are 0.068 μM and 0.078 μM, respectively. These values are dependent on the concentration and type of detector and cannot be contrasted with values reported for other synthetic catalysts. Therefore, the second-order rate constants kMcF = kNBT[NBT]/IC50, which are independent of the detector, were calculated and used to compare the present results with others from the literature.76 Both, [Cu(salbn)] and [Cu(py2bn)]2+, displayed excellent SOD activity (Fig. S15†), with kMcF (Table 2, entries 2 and 3) higher than most of the reported Cu(II) complexes with open-chain N2O2-, N3O-, and N4-tetradentate ligands, as shown in Table 2. Ligand structures of complexes listed in Table 2 are depicted in Chart S1. The selected Cu(II) catalysts are those for which kMcF was reported or could be calculated from the IC50 value.14,15,19–21,23,25–27,77,78
| Catalyst | Ligand donor sites | 106kMcF (M−1 s−1) | E (II/I) (mV vs. SCE), solvent | Ref. | |
|---|---|---|---|---|---|
| aptn = [N,N′-butylenebis(2-acetylpyridineiminato)]; HPBMPA = N-propanoate-N,N-bis-(2-pyridylmethyl)amine; HPClNOL = 1-[bis(pyridin-2-ylmethyl)amino]-3-chloropropan-2-ol; L1 = N1,N2-bis(phenyl(pyridine-2-yl)-methylene)-ethane-1,2-diamine; MPBMPA = N-methylpropanoate-N,N-bis-(2-pyridylmethyl)amine; Pu-6-MePy = N,N′-bis(2-(6-methyl-pyridyl)methylene)-1,4-butanediamine; PuPhePy = N,N′-bis(2-pyridyl-phenyl)methylene-l,4-butanediamine; pypapn = 1,3-bis[(pyridin-2-ylmethyl)(propargyl)amino]propane; py2pn = N,N′-bis(2-pyridinylmethylen)propane-1,3-diamine; 4-OMe-salen = N,N′-bis(4-methoxysalicylidene)cyclohexane-1,2-diamine; salpn = 1,3-bis(salicylidenamino)propane.a Previous values determined by the xanthine–xanthine oxidase method: kMcF = 26.4 M−1 s−1 at pH 7.4;79kMcF = 23.7 M−1 s−1 at pH 8.0.26b Using alkaline DMSP-NBT method, pH 8.6.c [Cu(salpn)] was prepared as described by Maurya et al.80 and synthetic details are given in ESI.†. | |||||
| 1 | [Cu(PuPhePy)]2+ | N4 | 52.8 | — | 26 |
| 2 | [Cu(salbn)(H 2 O)] | N2O2 | 33.3 | −361(Epc), H2O; −980, MeCN | This work |
| 3 | [Cu(py 2 bn)(H 2 O)] 2+ | N4 | 29.1a | −229 (Epc), H2O; 245, MeCN66 | This work |
| 4 | [Cu(MPBMPA)Cl2] | N3 | 21.2 | −471, MeCN | 21 |
| 5 | [Cu(pypapn)]2+ | N4 | 12.6 | −220 (Epc), MeCN | 14 |
| 6 | [Cu(PBMPA)Cl] | N3O | 12.5 | 213, MeCN | 21 |
| 7 | [Cu(Pu-6-MePy)(H2O)]2+ | N4 | 6.3 | — | 25 |
| 8 | [Cu(aptn)]2+ | N4 | 4.95 | — | 27 |
| 9 | [Cu(py2pn)]2+ | N4 | 4.05 | −44, MeCN | 15 |
| 10 | [Cu(PClNOL)Cl]+ | N3O | 3.3 | −413, MeCN | 19 and 77 |
| 11 | [CuL1]2+ | N4 | 1.67b | — | 20 |
| 12 | [Cu(salpn)]c | N2O2 | 1.24 | −1000, MeCN | This work |
| 13 | [Cu(4-OMe-salen)ZnCl2] | N2O2 | 0.87 | — | 23 |
| 14 | [salpnCuZnCl2] | N2O2 | 0.85 | −689, DMF | 78 |
| 15 | CuZnSOD | N4 | 2000 | 156 | 2 |
If the redox potential of the catalyst is one of the major features affecting the catalysed O2˙− dismutation, the compounds with E1/2 closer to 119 mV vs. SCE – the midpoint of the reduction/oxidation potentials of superoxide – should be among the most active ones. However, when the SOD activities of the complexes listed in Table 2 are plotted against their redox potentials in MeCN (the solvent used in the electrochemical studies of most of the listed complexes), no correlation is observed (Fig. 8). On the contrary, [Cu(salbn)], with E(Cu(II)/Cu(I)) well outside the potential range required for O2˙− dismutation, is the best SOD mimic of the set of {kMcF,E} data pairs plotted in the graph. As shown before for the two complexes studied in this work, the solvent affects the geometry and properties of species in solution, in particular the redox potentials, and the magnitude of this effect depends on the electronic structure of the complex, and, if coordinated, the position occupied by the solvent in the coordination sphere.33,34 In the present case, the redox potential of [Cu(salbn)] shifts by 600 mV to less negative potentials, while that of [Cu(py2bn)]2+ shifts 474 mV to more negative values, when going from MeCN to water, resulting close reduction potentials for the two complexes in water, in spite of the large difference observed in MeCN. This means that comparing SOD activity in terms of redox potentials registered in non-aqueous solvents could lead to erroneous conclusions.
 | ||
| Fig. 8 k McF vs. redox potentials of complexes in Table 2. Lit. values (blue and blue/green dots). Dashed lines correspond to O2˙− reduction and oxidation potentials. | ||
In addition to affecting the geometry or the distortion degree of the complex structure, the solvent may be involved in the solvolysis of a weakly bound ligand, as observed in the present work for [Cu(py2bn)(OAc)]2+ in the protic solvents. Therefore, given that the redox potential depends on the geometry of the complex, solvent parameters and type of the ligands bound to the metal ion,33,34 only the potentials measured in aqueous medium can be directly correlated to the observed SOD activity. In the present case, in aqueous medium [Cu(salbn)] reacts with O2˙− just 15% faster than [Cu(py2bn)]2+ despite having a redox potential more distant from the optimal value to disproportionate O2˙− (119 mV vs. SCE). This implies that in addition to the redox potential, there are other factors affecting the reaction of these complexes with O2˙−.
A feature that influences the SOD activity is the ability of the ligand to adopt different dispositions around the metal centre to best adapt to the preferred geometries of each oxidation state of copper during catalysis. For this reason, the more flexible ligands adapt better to the required conformational changes when going from Cu(II) to Cu(I). The flexibility of ligands used here depends largely on the length of the central chain. The two ligands with the –(CH2)4− linker between the imino N-atoms form flexible seven-membered chelate rings which can adopt different conformations with energy barriers lower than six-membered and five-membered ones. Therefore, [Cu(salbn)] reacts 27-times faster than [Cu(salpn)] (Table 2, entry 12) and 39-times higher than [CuZn(salpn (or salen))Cl2] (Table 2, entries 13 and14) formed with more rigid ligands with 3C- and 2C-central alkyl chains. Also, [Cu(py2bn)]2+ reacts with O2˙− 7.2-times faster than [Cu(py2pn)]2+ (Table 2, entry 9) and even more than twice as fast as [Cu(pypapn)]2+ (Table 2, entry 5), in which copper is bound to 2Npy2Namine donor set and possesses a 3C-central aliphatic chain. The methyl substituent on the pyridine ring in the trigonal–bipyramidal [Cu(Pu-6-MePy)(H2O)]2+ or on the imino C-atom in distorted square-planar [Cu(aptn)]2+ decreases the SOD activity about 5- and 6-times, respectively, compared to the unsubstituted [Cu(py2bn)]2+ (Table 2, entries 7 and 8), probably because the methyl induces a decrease of the positive charge on the metal to interact with the substrate. The opposite effect is observed for [Cu(PuPhePy)]2+ (Table 2, entry 1), a distorted square-pyramidal complex with solvent at the apex and the 4N-donor set at the base, where the phenyl substituent on the imino C-atom improves the activity compared to [Cu(py2bn)]2+, placing it as the most active Cu-based catalyst of open chain ligands known so far. Complex [CuL1]2+ is an analogue of [Cu(PuPhePy)]2+ with a shorter –(CH2)2– central alkyl chain. The shortening of the aliphatic chain increases the complex rigidity that adopts a square-planar geometry and reduces its SOD activity drastically (Table 2, entry 11) compared to [Cu(PuPhePy)]2+. [Cu(MPBMPA)Cl2], [Cu(PBMPA)Cl] and [Cu(PClNOL)Cl]+ (Table 2, entries 4, 6 and 10) are three square pyramidal complexes in which the Cu atom sits in the plane formed by the 3N-donor set of the ligand and one chloride anion, with the apical position occupied by a second chloride anion, an O atom from the carboxylate of the ligand, or an alcohol group, respectively. In [Cu(PBMPA)Cl], the carboxylate acts as a competing ligand lowering SOD activity by almost half of [Cu(MPBMPA)Cl2], for which the apical chloride is rapidly exchanged with the solvent leaving a labile position to interact with the substrate. This fact favours the reactivity of [Cu(MPBMPA)Cl]+ with O2˙− and places this complex among the more active ones. The apical alcohol group in [Cu(PClNOL)Cl]+ (Table 2, entry 10) further decreases the SOD activity, probably as a consequence of the lower labilising effect of the alcohol disfavouring ligand exchange.
Even when all the complexes listed in Table 2 are less reactive than the SOD enzyme (entry 15), it is evident that the ligand flexibility plays a critical role in the O2˙− dismutation catalysed by Cu-based SOD mimics, placing the two complexes of this work among the three most reactive catalysts with open-chain ligands.
In the enzyme, the protein environment isolate the active site and modulates its geometry so that the structural reorganization required to switch between Cu(II)/Cu(I) oxidation states is minimal, and the superoxide dismutation occurs efficiently at a rate close to the diffusion limit (≈109 M−1 s−1).2 Keeping in mind the confining effect of the protein matrix and with the intention of improving the stability and reuse of the catalysts, the hybrid materials prepared by encapsulation of the catalysts inside the SBA-15 silica channels were tested as SOD mimics. Besides, isolation of the catalyst within the pores would avoid undesired dimerization that can occur in homogeneous medium during the catalytic cycles competing with the target reaction.81 Cu-salbn@SBA-15 retains SOD activity but reacts at a lower rate than the homogeneous catalyst, affording IC50 = 0.11 μM, and kMcF = 20.6 × 106 M−1 s−1 (Fig. S15(a)†). In the case of Cu-py2bn@SBA-15, the decrease in rate is more evident (Fig. S15(b)†), giving IC50 = 0.28 μM, and kMcF = 8.1 × 106 M−1 s−1. The drop in activity might be related to a change in the geometry of the catalyst within the silica channels. This was confirmed by low-temperature X-band EPR spectra of the hybrids. The EPR spectrum of Cu-salbn@SBA-15 exhibits an axial signal (Fig. 9(a), left) with spectral parameters g∥ = 2.17 and g⊥ = 2.07 obtained from the simulated spectrum (Fig. S1(c)†). These features are diagnostic of a tetragonal geometry around the Cu(II) centre within the SBA-15 pores, although less distorted than the complex in frozen DMSO solution. The EPR spectrum is broad probably as a result of the overlap of species with slightly different geometries or orientations within the SBA-15 matrix. The low-temperature EPR spectrum of Cu-py2bn@SBA-15 (Fig. 9(a), right) displays the typical signature of Cu(II) in a tetragonal geometry with g∥ = 2.25, g⊥ = 2.07, A∥ = 164 × 10−4 cm−1, obtained from the simulated spectrum (Fig. S3(c)†), and an empirical distortion factor f(g∥/A∥) = 137 cm, denoting a tetrahedrally distorted N4-square-planar geometry. In this case, encapsulation constrains the ligand to dispose in the equatorial plane, instead to adopt the trigonal bipyramidal geometry observed in aqueous solution. The two hybrids react rapidly with KO2 at 25 °C in DMSO, where O2˙− is a powerful reductant and nucleophile, and rather stable towards self-dismutation. When Cu-salbn@SBA-15 was treated with excess of KO2 in DMSO, the EPR spectrum of the hybrid after reaction (Fig. 9(b), left) confirmed that the immobilized complex keeps the geometrical arrangement of the ligand around the Cu(II) ion. A different behaviour is observed for Cu-py2bn@SBA-15, for which the low-temperature EPR spectrum taken after reaction with O2˙− still shows axial symmetry (Fig. 9(b), right), but well different spectral parameters: g∥ = 2.36, g⊥ = 2.06, A∥ = 154 × 10−4 cm−1, and f = 153 cm, obtained from the simulated spectrum (Fig. S3(d)†). These spectral parameters correspond to a species much more tetrahedrally distorted than that of the starting material. Although with a distortion similar to the enzyme (for the enzyme, f = 160 cm),82 the species formed during the catalytic cycle is almost 3-times less reactive than the free complex probably because the silanolate-copper interaction restraints the ligand flexibility around the metal centre, resulting in a rate of reaction with O2˙− similar to that reported for Cu-py2pn@SBA-15 (kMcF = 6.9 × 106 M−1 s−1).14
 | ||
| Fig. 9 Low-temperature EPR spectra of Cu-salbn@SBA-15 (left) and Cu-py2bn@SBA −15 (right) before (a) and after (b) reaction with 10 equiv. of KO2 in DMSO. ν = 9.31 GHz, T = 120 K. | ||
The absence of the superoxide signal (g∥ = 2.1021, g⊥ = 2.003)83 in the EPR spectra recorded a few seconds after reaction of the hybrids with 10-times excess of KO2 in DMSO, confirms that these materials act as catalysts for O2˙− dismutation. Also, the lack of EPR signal in the supernatant solution after treatment of the materials with excess of KO2 in DMSO indicates the absence of complex leached during the reaction with O2˙−. In this way, encapsulation confers stability and protection to the catalyst emulating the protein matrix in the enzyme, both for the neutral and cationic catalyst. Furthermore, to determine if the activity of the free and encapsulated catalysts was retained after several cycles, the conversion of NBT was measured after successive illuminations of the reaction mixture with and without catalyst, adding after each new illumination the amount of NBT necessary to restore the initial concentration, so that the total NBT remained constant. When the homogeneous catalysts were used, the percentage of NBT converted to formazan decreased after each illumination period. In contrast, when the hybrid materials were used the NBT conversion remained almost invariant, indicating that immobilization prolongs the lifetime of the catalyst.
Conclusions
This comparative study highlights that ligand flexibility has a more significant impact on SOD activity than the redox potential of the copper centre. For copper complexes of open-chain diimino ligands with terminal phenolate or pyridine, the flexibility increases with the length of the aliphatic linker between the imino N-atoms and the size of the chelate rings. Therefore, [Cu(salbn)(H2O)] and [Cu(py2bn)(H2O)]2+, with –(CH2)4– alkyl chain, react with O2˙− much faster than the analogues with shorter central alkyl chains. In addition, the effect of the solvent on geometry and redox potential of these complexes denotes it is essential to characterize them in aqueous medium to correlate their structure, redox potential and SOD activity. The aqueous buffered medium levels the redox potential of the two complexes studied here to values within the range required to catalyse O2˙− dismutation. However, [Cu(salbn)(H2O)], with a more unfavourable redox potential, reacts with O2˙− faster than [Cu(py2bn)(H2O)]2+, pointing out the strong influence of the flexibility of the adjacent six-seven-six-membered chelate rings of [Cu(salbn)(H2O)] to accommodate the Cu(II)/Cu(I) geometric preferences. This is supported by the effect of immobilization of the complexes in SBA-15 on the catalytic activity. Encapsulation of the complexes in the silica matrix not only modifies the complex structures but, probably, the pore restrains the ligand flexibility to encompass the switch between Cu(II)/Cu(I) oxidation states, decreasing the O2˙− dismutation rate. Nevertheless, although the two hybrids react with O2˙− slower than the free complexes, the confinement was successful in isolating, protecting and extending the useful life of the catalyst. Whereby, there must be a compromise between isolation and protection of the catalyst and restriction of the geometrical arrangement within the pore, and these two features should be considered to delineate heterogeneous SOD mimics.Data availability
The data supporting this article have been included as part of the ESI.†Crystallographic data for [Cu(salbn)] and [Cu(py2bn)(OAc)]ClO4 have been deposited at the CCDC under 2405330 and 2405337.†
Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work was supported by the National University of Rosario (PIDs UNR 80020210200140UR, 8002019040023UR and 80020220700136UR), Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET, PIP 0852 and PUE 0068), and Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, PICT-2019-03276). We thank Gustavo Terrestre for solving the crystal structures. The Bruker D8 QUEST ECO Photon II CPAD Diffractometer was purchased with funds from ANPCyT (PME 2015-0022) and CONICET.References
- B. Yang, Y. Chen and J. Shi, Chem. Rev., 2019, 119, 4881–4985 CrossRef CAS PubMed.
- I. A. Abreu and D. E. Cabelli, Biochim. Biophys. Acta, 2010, 1804, 263–274 CrossRef CAS PubMed.
- B. Halliwell and J. M. Gutteridge, Free Radicals in Biology and Medicine, Oxford University Press, 2015 Search PubMed.
- H. Sies, Curr. Opin. Toxicol., 2018, 7, 122–126 CrossRef.
- R. Bonetta, Chem. – Eur. J., 2018, 24, 5032–5041 CrossRef CAS PubMed.
- G. Schanne, S. Demignot, C. Policar and N. Delsuc, Coord. Chem. Rev., 2024, 514, 215906 CrossRef CAS.
- C. Policar, J. Bouvet, H. C. Bertrand and N. Delsuc, Curr. Opin. Chem. Biol., 2022, 67, 102109 CrossRef CAS PubMed.
- S. Signorella, C. Palopoli and G. Ledesma, Coord. Chem. Rev., 2018, 365, 75–102 CrossRef CAS.
- I. Batinić-Haberle, J. S. Rebouças and I. Spasojević, Antioxid. Redox Signal., 2010, 13, 877–918 CrossRef PubMed.
- Y. Sheng, I. A. Abreu, D. E. Cabelli, M. J. Maroney, A.-F. Miller, M. Teixeira and J. S. Valentine, Chem. Rev., 2014, 114, 3854–3918 CrossRef CAS PubMed.
- J. A. Tainer, E. D. Getzoff, K. M. Beem, J. S. Richardson and D. C. Richardson, J. Mol. Biol., 1982, 160, 181–217 CrossRef CAS PubMed.
- S. Pramanik and S. Chattopadhyay, Inorg. Chim. Acta, 2023, 552, 121486 CrossRef CAS.
- M. A. Mekhail, K. J. Smith, D. M. Freire, K. Pota, N. Nguyen, M. E. Burnett and K. N. Green, Inorg. Chem., 2023, 62, 5415–5425 CrossRef CAS PubMed.
- M. Richezzi, J. Ferreyra, S. Signorella, C. Palopoli, G. Terrestre, N. Pellegri, C. Hureau and S. R. Signorella, Inorganics, 2023, 11, 425 CrossRef CAS.
- M. Richezzi, J. Ferreyra, J. Puzzolo, L. Milesi, C. M. Palopoli, D. M. Moreno, C. Hureau and S. R. Signorella, Eur. J. Inorg. Chem., 2022, e202101042 CrossRef CAS.
- Á. Martinez-Camarena, P. A. Sanchez-Murcia, S. Blasco, L. Gonzalez and E. Garcia-España, Chem. Commun., 2020, 56, 7511–7514 RSC.
- J. D. Siqueira, S. F. de Pellegrin, S. S. dos Santos, B. Almeida Iglesias, P. C. Piquini, L. P. Arantes, F. A. Soares, O. A. Chaves, A. Neves and D. F. Back, J. Inorg. Biochem., 2020, 204, 110950 CrossRef CAS PubMed.
- K. N. Green, K. Pota, G. Tircso, R. A. Gogolak, O. Kinsinger, C. Davda, K. Blain, S. M. Brewer, P. Gonzalez, H. M. Johnston and G. Akkaraju, Dalton Trans., 2019, 48, 12430–12439 RSC.
- T. P. Ribeiro, C. Fernandes, K. V. Melo, S. S. Ferreira, J. A. Lessa, R. W. A. Franco, G. Schenk, M. D. Pereira and A. Horn Jr, Free Radicals Biol. Med., 2015, 80, 67–76 CrossRef CAS PubMed.
- M. Choudhary, R. N. Patel and S. P. Rawat, J. Mol. Struct., 2014, 1070, 94–105 CrossRef CAS.
- J. S. Pap, B. Kripli, I. Bors, D. Bogáth, M. Giorgi, J. Kaizer and G. Speier, J. Inorg. Biochem., 2012, 117, 60–70 CrossRef CAS PubMed.
- C. Wang, S. Li, D.-J. Shang, X.-L. Wang, Z.-L. You and H.-B. Li, Bioorg. Med. Chem. Lett., 2011, 21, 4320–4324 CrossRef CAS PubMed.
- Z.-L. You, L.-L. Ni, P. Hou, J.-C. Zhang and C. Wang, J. Coord. Chem., 2010, 63, 515–523 CrossRef CAS.
- J. Lange, H. Elias, H. Paulus, J. Müller and U. Weser, Inorg. Chem., 2000, 39, 3342–3349 CrossRef CAS PubMed.
- J. Müller, D. Schübl, C. Maichle-Mössmer, J. Strähle and U. Weser, J. Inorg. Biochem., 1999, 75, 63–69 CrossRef.
- J. Müller, K. Felix, C. Maichle, E. Lengfelder, J. Strähle and U. Weser, Inorg. Chim. Acta, 1995, 233, 11–19 CrossRef.
- Q. Luo, Q. Lu, A. Dai and L. Huang, J. Inorg. Biochem., 1993, 51, 655–662 CAS.
- J. Stanek, A. Hoffmann and S. Herres-Pawlis, Renaissance of the entatic state principle, Coord. Chem. Rev., 2018, 365, 103–121 CrossRef CAS.
- B. A. Vaughn, A. M. Brown, S. H. Ahn, J. R. Robinson and E. Borosm, Inorg. Chem., 2020, 59, 16095–16108 CrossRef CAS PubMed.
- N. W. G. Smits, B. van Dijk, I. de Bruin, S. L. T. Groeneveld, M. A. Siegler and D. G. H. Hetterscheid, Inorg. Chem., 2020, 59, 16398–16409 CrossRef CAS PubMed.
- E. Falcone and C. Hureau, Chem. Soc. Rev., 2023, 52, 6595–6600 RSC.
- C. Policar, in Redox Active Therapeutics, ed. J. S. Reboucas, I. Batinic-Haberle, I. Spasojevic, D. S. Warner and D. St. Clair, Springer, 2016, ch. 17, pp. 125–164 Search PubMed.
- G. Tabbì, A. Giuffrida and R. P. Bonomo, J. Inorg. Biochem., 2013, 128, 137–145 CrossRef PubMed.
- P. A. Lay, N. S. McAlpine, J. T. Hupp, M. J. Weaver and A. M. Sargesons, Inorg. Chem., 1990, 29, 4322–4328 CrossRef CAS.
- M. Patriarca, V. Daier, G. Camí, N. Pellegri, E. Rivière, C. Hureau and S. Signorella, Microporous Mesoporous Mater., 2019, 279, 133–141 CrossRef CAS.
- A. Larki, S. J. Saghanezhad and M. Ghomi, Microchem. J., 2021, 169, 106601 CrossRef CAS.
- M. N. Iqbal, G. R. Nicoud, M. Ciurans-Oset, F. Akhtar, N. Hedin and T. Bengtsson, ACS Appl. Mater. Interfaces, 2023, 15, 9542–9553 CrossRef CAS PubMed.
- E. Kadwa, M. D. Bala and H. B. Friedrich, Appl. Clay Sci., 2014, 95, 340–347 CrossRef CAS.
- S. Stoll and A. Schweiger, J. Magn. Reson., 2006, 178, 42–55 CAS.
- M. Thommes, K. Kaneko, A. V. Neimark, J. P. Olivier, F. Rodriguez-Reinoso, J. Rouquerol and K. S. W. Sing, Pure Appl. Chem., 2015, 87, 1051–1069 CrossRef CAS.
- J. Villarroel Rocha, D. Barrera and K. Sapag, Top. Catal., 2011, 54, 121–134 CrossRef CAS.
- Bruker, APEX3 v2019.10-1, Bruker AXS Inc., Madison, WI, USA, 2019 Search PubMed.
- Bruker APEX4 v2022.10-1, Bruker AXS Inc., Madison, Wisconsin, USA, 2022 Search PubMed.
- Bruker SAINT V8.40B, Bruker AXS Inc., Madison, Wisconsin, USA, 2019 Search PubMed.
- G. M. Sheldrick, SHELXT-Integrated space-group and crystal-structure determination, Acta Crystallogr., Sect. A:Found. Adv., 2015, 71, 3–8 CrossRef PubMed.
- G. M. Sheldrick, Crystal structure refinement with SHELXL, Acta Crystallogr., Sect. B:Struct. Sci., Cryst. Eng. Mater., 2015, 71, 3–8 CrossRef PubMed.
- L. J. Farrugia, ORTEP3 for Windows, J. Appl. Crystallogr., 1997, 30, 565 CrossRef CAS.
- C. Beauchamps, Fridovich, Anal. Biochem., 1971, 44, 276–287 CrossRef PubMed.
- Z.-R. Liao, X.-F. Zheng, B.-S. Luo, L.-R. Shen, D.-F. Li, H.-L. Liu and W. Zhao, Polyhedron, 2001, 20, 2813–2821 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16, Revision C.01, Gaussian, Inc., Wallingford CT, 2016 Search PubMed.
- A. D. Becke, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- J. Tomasi, B. Mennucci and E. Cancès, J. Mol. Struct.:THEOCHEM, 1999, 464, 211–226 CrossRef CAS.
- H.-H. Yao, J.-M. Lo, B.-H. Chen and T.-H. Lu, Acta Crystallogr., Sect. C:Cryst. Struct. Commun., 1997, 53, 1012–1013 Search PubMed.
- L. C. Nathan, J. E. Koehne, J. M. Gilmore, K. A. Hannibal, W. E. Dewhirst and T. D. Mai, Polyhedron, 2003, 22, 887–894 CrossRef CAS.
- H. Keypour, M. Shayesteh, S. Salehzadeh, S. Dhers, F. Maleki, H. Ünverc and N. Dilek, New J. Chem., 2015, 39, 7429–7441 CAS.
- L. Yang, D. R. Powell and R. P. Houser, Dalton Trans., 2007, 955–964 RSC.
- H. Kanso, R. M. Clarke, A. Kochem, H. Arora, C. Philouze, O. Jarjayes, T. Storr and F. Thomas, Inorg. Chem., 2020, 59, 5133–5148 CAS.
- S. Muthuramalingam, K. Anandababu, M. Velusamy and R. Mayilmurugan, Inorg. Chem., 2020, 59, 5918–5928 CAS.
- K. Das, S. Goswami, B. B. Beyene, A. W. Yibeltal, E. Garribba, A. Frontera and A. Datta, Polyhedron, 2019, 159, 323–329 CAS.
- M. M. Bhadbhade and D. Srinivas, Inorg. Chem., 1993, 32, 5458–5466 CrossRef CAS.
- E. Movahedi and H. Golchoubian, J. Mol. Struct., 2006, 787, 167–171 CAS.
- M. Bühl, S. E. Ashbrook, D. M. Dawson, R. A. Doyle, P. Hrobárik, M. Kaupp and I. A. Smellie, Chem. – Eur. J., 2016, 15328–15339 Search PubMed.
- B. Murphy and B. Hathaway, Coord. Chem. Rev, 2003, 243, 237–262 CAS.
- A. Gubendran, J. Rajesh, K. Anitha and P. Athappan, J. Mol. Struct., 2014, 1075, 419–429 CAS.
- F. Clifford, E. Counihan, W. Fitzgerald, K. Seff, C. J. Simmons, S. Tyagi and B. J. Hathaway, J. Chem. Soc., Chem. Commun., 1982, 196–198 CAS.
- Y. Yao, M. W. Perkovic, D. P. Rillema and C. Woods, Inorg. Chem., 1992, 31, 3956–3962 CAS.
- A. Kunishita, J. D. Scanlon, H. Ishimaru, K. Honda, T. Ogura, M. Suzuki, C. J. Cramer and S. Itoh, Inorg. Chem., 2008, 47, 8222–8232 CAS.
- L. Llanos, C. Vera, A. Vega, D. Aravena and L. Lemus, Inorg. Chem., 2020, 59, 15061–15073 CAS.
- W. J. Geary, Coord. Chem. Rev., 1971, 7, 81–122 CrossRef CAS.
- J. N. Melville and P. V. Bernhardt, Inorg. Chem., 2021, 60, 9709–9719 CrossRef CAS PubMed.
- S. Odisitse and G. E. Jackson, Inorg. Chim. Acta, 2009, 362, 125–135 CrossRef CAS.
- I. Y. Ahmed and A. L. Abu-Hijleh, Inorg. Chim. Acta, 1982, 61, 241–246 CrossRef CAS.
- J. M. Brink, R. A. Rose and R. C. Holz, Inorg. Chem., 1996, 35, 2878–2885 CrossRef CAS.
- M. K. Taylor, J. Reglinski, L. E. A. Berlouis and A. R. Kennedy, Inorg. Chim. Acta, 2006, 359, 2455–2464 CrossRef CAS.
- P. M. Wood, Biochem. J., 1988, 253, 287–289 CrossRef CAS PubMed.
- S. Durot, C. Policar, F. Cisnetti, F. Lambert, J. P. Renault, G. Pelosi, G. Blain, H. Korri-Youssoufi and J. P. Mahy, Eur. J. Inorg. Chem., 2005, 3513–3523 CrossRef CAS.
- A. C. Fernandes, G. L. Parrilha, J. A. Lessa, L. J. M. Santiago, M. M. Kanashiro, F. S. Boniolo, A. J. Bortoluzzi, N. V. Vugman, M. H. Herbst and A. Horn Jr., Inorg. Chim. Acta, 2006, 359, 3167–3176 CrossRef.
- V. A. Daier, E. Rivière, S. Mallet-Ladeira, D. Moreno, C. Hureau and S. R. Signorella, J. Inorg. Biochem., 2016, 163, 162–175 CrossRef CAS PubMed.
- M. Linss and U. Weser, Inorg. Chim. Acta, 1986, 125, 117–121 CrossRef CAS.
- M. R. Maurya, S. J. J. Titinchi and S. Chand, Appl. Catal., A, 2002, 228, 177–187 CrossRef CAS.
- C. Palopoli, G. Gómez, A. Foi, F. Doctorovich, S. Mallet-Ladeira, C. Hureau and S. Signorella, J. Inorg. Biochem., 2017, 167, 49–59 CrossRef CAS PubMed.
- K. M. Beem, W. E. Rich and K. V. Rajagopa, J. Biol. Chem., 1974, 249, 7298–7305 CrossRef CAS PubMed.
- R. N. Bagchi, A. M. Bond, F. Scholz and R. Stoesser, J. Am. Chem. Soc., 1989, 111, 8270–8271 CrossRef CAS.
Footnotes |
| † Electronic supplementary information (ESI) available: X-ray crystallography summary tables, additional spectroscopic, kinetics, DRX and microscopic data, results from DFT calculations. CCDC 2405330 and 2405337. For ESI and crystallographic data in CIF or other electronic format see DOI: https://doi.org/10.1039/d4dt03403a |
| ‡ Equal contribution. |
| This journal is © The Royal Society of Chemistry 2025 |





