Energetic and structural dynamic drivers of transcription factor MycMax, Omomyc homodimer, and MaxMax recognition on DNA
Carmen Al
Masri
 a and
Jin
Yu
a and
Jin
Yu
 *b
*b
aDepartment of Physics and Astronomy, University of California, Irvine, California, USA
bDepartment of Physics and Astronomy, Department of Chemistry, NSF-Simons Center for Multiscale Cell Fate Research, University of California, Irvine, California, USA. E-mail: jin.yu@uci.edu
First published on 12th November 2025
Abstract
Basic helix–loop–helix leucine zipper (bHLHLZ) transcription factors (TFs), such as MycMax, MaxMax, and the engineered mutant Omomyc dimers, regulate gene expression by binding to the E-box (5′-CACGTG-3′) motif in DNA. Here, we used microsecond molecular dynamics (MD) simulations along with MMGBSA energetic and hydrogen bonding (HB) analyses to investigate how MycMax, Omomyc, and MaxMax dimers interact with both canonical E-box and homogeneous polyA reference sequences. Our MD results indicate that protein–DNA van der Waals (VDW) interactions and surface complementarity dominate the binding affinity measurements. Per-residue decomposition underscores the central role of conserved arginine residues at the protein–DNA binding interface, while HB information readout (or KL divergence) analyses reveal a biased DNA strand association that reinforces protein sequence recognition for specificity. Although MaxMax binds to the canonical E-box energetically stably, it loses the stability on polyA DNA more than the other two TFs, suggesting an essential reliance of MaxMax on the E-box motif for TF–DNA interaction coordination. MycMax exhibits moderate stability energetically and shows more variable dynamical responses to local DNA sequences. In contrast, the Omomyc dimer retains slightly higher binding affinity or energetic stability than MycMax on both E-box and polyA DNA, suggesting an enhanced ability to sequester MycMax in oncogenic settings. These findings provide new physical insights into dimeric bHLHLZ–TFs on DNA recognition and reveal potential strategies for targeting Myc-driven transcription or overexpression in cancer.
Introduction
Basic helix–loop–helix leucine zipper (bHLHLZ) transcription factors (TFs) (Fig. 1A) govern key cellular processes such as cell proliferation, differentiation, and apoptosis by binding to specific DNA sequences.1,2 Dysregulation of bHLHLZ proteins is implicated in numerous diseases; for example, overexpression of the Myc family proteins is observed in approximately 70% of human tumors and is a key driver of oncogenesis.3–5 Myc heterodimerizes with Max via their bHLHLZ domains to bind to a canonical E-box motif (5′-CACGTG-3′) in gene promoters, driving a broad transcriptional program that influences cell growth, metabolism, and the cell cycle.2,4,6 In contrast, Max can homodimerize and repress transcription by competing for the same E-box sites, thus balancing Myc's oncogenic effects.1,7 Other bHLHLZ proteins, such as the Mxd family, further modulate Myc activity by forming repressive complexes.1,2 | ||
| Fig. 1 Structural and sequence comparison of MycMax, Omomyc dimer, and MaxMax proteins. The structures were obtained for MycMax (PDB: 1NKP19), the Omomyc dimer (PDB: 5I509), and MaxMax (PDB: 1AN221) from the Protein Data Bank (PDB). (A) Schematic representation of the DNA-bound MycMax bHLHLZ domain, highlighting the basic region (BR, yellow), helix 1 (H1, orange), loop (L, green), helix 2 (H2, red), and leucine zipper (LZ, blue). The E-box on the DNA is colored in red. (B) Molecular view of the HB interactions within the LZ regions of the MycMax (left), Omomyc dimer (center), and MaxMax (right) complexes. The four mutations (N156/N247, Q155/Q247, I149/I241, and T142/T234) in Omomyc with respect to Myc are highlighted in magenta. (C) Sequence alignment of the bHLHLZ domains from Max, Myc, and Omomyc. Conserved residues are highlighted in blue. The four mutated residues in the Omomyc LZ are marked with a star and colored in red. Omomyc residues from mutations that align with Max are highlighted in pink. The percentages indicate Myc and Omomyc sequence identity compared to Max. (D) DNA sequence utilized in modelling protein binding, including the central core (E-box, highlighted in red) and flanking sequence (highlighted in blue) regions. | ||
A promising strategy to inhibit Myc in cancer is via Omomyc, a synthetic 91-amino-acid protein mutant derived from Myc's bHLHLZ domain.8 Omomyc was engineered by substituting four key charged residues (two Asp and two Arg) in the leucine zipper (LZ), which normally prevent Myc homodimerization, with those found mainly in Max8 (see the mutated residues T, I, Q, and N in Omomyc in Fig. 1B and C). This design enables Omomyc to efficiently homodimerize as well as to heterodimerize with both Myc and Max.9–11 As a result, Omomyc can compete with or prevent Myc from both dimerizing with Max and binding to E-box sequences, thereby reducing or disrupting Myc-driven gene expression. Preclinical studies have shown that Omomyc causes significant tumor regression with minimal and reversible side effects on normal tissues,5,11 suggesting its potential as an effective Myc inhibitor. Notably, Omomyc has advanced to clinical evaluation; OMO-103, the clinical formulation of Omomyc, successfully completed a phase I trial in patients with advanced solid tumors, demonstrating safety and preliminary signs of anti-tumor activity.12
Although Myc, Max, and Omomyc share a common bHLHLZ architecture, differences in their LZ and basic DNA-binding regions (BRs) critically influence their dimerization strength and DNA-binding specificity, respectively. For example, the four point mutations (E142T, E149I, R155Q, and R156N) in Omomyc not only promote homodimerization by mimicking key Max residues but also enhance dimerization stability through additional hydrophobic and hydrogen-bond (HB) interactions (Fig. 1B, C and Fig. S1).8,9 However, while crystallographic and biochemical studies have detailed the static interactions, the dynamic and energetic determinants that govern how these dimers selectively recognize specific (E-box) versus nonspecific DNA remain incompletely understood. It is also unclear how bHLHLZ TF–DNA interfacial contacts modulate binding affinity or specificity in different DNA contexts and which of the dimers is the most stabilized energetically bound on E-box or nonspecific DNA.
To address these questions, we performed microsecond all-atom MD simulations of MycMax, MaxMax, and Omomyc dimers in complex with both a canonical E-box (CACGTG) and a homogeneous polyadenine (polyA) 40-base pair (bp) sequence. PolyA was chosen as a control or reference sequence with minimal information entropy, and it has also been confirmed from protein binding microarray (PBM) experiments that polyA is among the most weakly bound by bHLHLZ proteins.13 Additionally, its relatively uniform DNA shape removes bias from sequence-dependent shape recognition, making it a useful control highlighting nonspecific interactions or comparing with specific interactions. By analyzing protein–DNA conformational dynamics, interfacial contacts and HB patterns, and binding free energies or components computed via the MMGBSA method,14 we characterized how each dimer adapts to E-box versus polyA DNA context. Our simulations reveal that although MaxMax binds the most stably energetically on E-box DNA, it undergoes the most energetic destabilization on polyA, whereas Omomyc and MycMax maintain their relative stabilities on DNA, with Omomyc slightly more stabilized than MycMax no matter on E-box or polyA. These observations can be compatible with early experimental findings on relatively high DNA binding affinity (with a low dissociation constant) of Max monomer and dimer to the E-box15 as well as later studies showing overall low DNA binding stability of MaxMax, likely due to its notable dimerization instabilities.9,15,16 Additionally, it has been found that MycMax binds to weaker motifs at high concentrations,17,18 and Omomyc effectively occupies low-affinity promoter regions, thereby displacing Myc from DNA.10
Overall, our findings provide new physical insights into how the protein sequence variations within the bHLHLZ BR or DNA binding domain (DBD) influence DNA binding stability and specificity, deepen our understanding of Myc-mediated transcriptional regulation, and offer a framework for improving rational design of next-generation therapeutics to target Myc-driven tumors.
Materials and methods
All systems underwent initial processing, minimization, and equilibration using GROMACS. Each system was placed in a periodic box with a minimum distance of 1.5 nm to the box edge, solvated, and ionized with Na+ and Cl− ions to a final ionic strength of 0.15 M. A rectangular box was used for E-box simulations, and a cubic box was used for the polyA simulations to be compatible with Anton2. The total number of atoms per system was approximately 140![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 for E-box systems and about 490
000 for E-box systems and about 490![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 for polyA systems. Short-range electrostatic and van der Waals (VDW) interactions were treated with a 1.2 nm cutoff, while long-range electrostatics were computed using the particle-mesh Ewald31 (PME) method.
000 for polyA systems. Short-range electrostatic and van der Waals (VDW) interactions were treated with a 1.2 nm cutoff, while long-range electrostatics were computed using the particle-mesh Ewald31 (PME) method.
Energy minimization was performed using the steepest-descent algorithm. The systems were then equilibrated in two stages: first for 4 ns under the NVT ensemble, followed by another 4 ns under the NPT ensemble, using a time step of 2 fs. Heavy-atom position restraints were applied with a force constant of 1000 kJ mol−1 nm−2. Temperature was maintained at 300 K using the leap-frog stochastic dynamics integrator with an inverse friction constant of 2 ps, while pressure was controlled at 1 bar using the Parrinello–Rahman barostat.32
For E-box systems, production simulations were performed for 1 µs in GROMACS, applying restraints only to the first and last heavy atoms of DNA to maintain alignment along the x-axis. For polyA systems, an additional 30 ns equilibration with DNA end restraints was carried out in GROMACS, after which the systems were transferred to Anton2 for a 10-µs production run. Only the first 1 µs of this trajectory is reported in the current study.
| ΔGbind = ΔGbind,vacuum + ΔGCsolvation − (ΔGPsolvation + ΔGDsolvation) | (1) |
| ΔGbind,vacuum = ΔEMM − TΔS | (2) |
Entropic contributions were incorporated using the interaction entropy method,36 which estimates the relative entropy or entropy change before and after protein–ligand (or DNA nucleotides here) binding −TΔS from fluctuations in the gas-phase protein–DNA interactions (ΔEMM) over the trajectory as 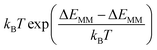 with 〈…〉 denoting an ensemble average. In the calculations, the protein–DNA interaction energy ΔEMM is regarded as the difference of gas-phase energy between that of the protein–DNA complex and those of the separate protein and DNA, which can be directly evaluated in the simulation of the protein–DNA complex. This interaction entropy method offers an efficient alternative to conventional normal mode analysis, providing improved accuracy in capturing interaction conformational entropy effects.37
with 〈…〉 denoting an ensemble average. In the calculations, the protein–DNA interaction energy ΔEMM is regarded as the difference of gas-phase energy between that of the protein–DNA complex and those of the separate protein and DNA, which can be directly evaluated in the simulation of the protein–DNA complex. This interaction entropy method offers an efficient alternative to conventional normal mode analysis, providing improved accuracy in capturing interaction conformational entropy effects.37
The solvation free energy, ΔGsolvation, is partitioned into polar and nonpolar components for the complex (C), the protein (P) and the DNA (D):
| ΔGC/P/Dsolvation = ΔGC/P/Dpolar + ΔGC/P/Dnonpolar | (3) |
All MMGBSA calculations were performed on the entirety of 1-µs production runs, with one frame sampled every nanosecond (1000 frames in total). To appropriately capture the high charge density effect inherent in DNA, the internal dielectric constant intdiel was set to 10 while the external dielectric was fixed at 78.5. The choice of internal dielectric was based on our benchmarks (Fig. S4), which showed that the total binding free energy convergence was achieved starting intdiel = 10.
Results and discussion
Structural dynamics stability of dimeric TFs on E-box and polyA DNA
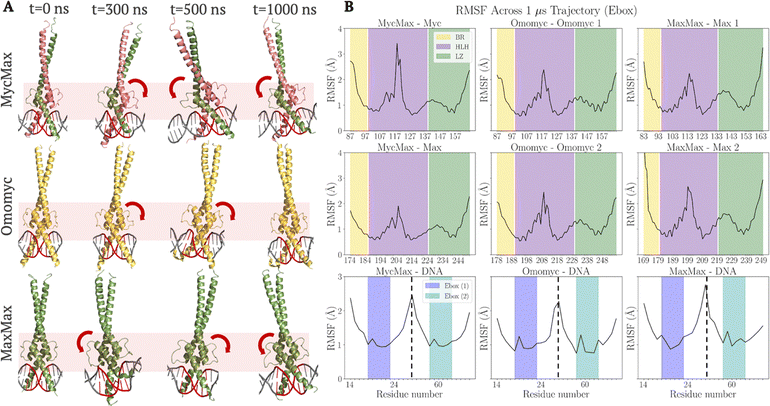
We first examined how MycMax, Omomyc, and MaxMax bHLHLZ dimers adapt to binding either a canonical E-box (5′-CACGTG-3′) or a polyA (5′-AAAAAA-3′) DNA sequence at the center of a 40-bp length DNA construct. Despite sharing the same core bHLHLZ fold, the three complexes displayed both common and distinct structural dynamics over the microsecond-scale simulations. Note that the structural dynamics stability can be directly monitored by root-mean square deviation (RMSD) of protein and/or DNA backbone structures (shown in Fig. S5).Note that there is a lysine residue (K57) on a loop from HLH identified in the MaxMax crystal structure (PDB: 1AN221) contacting the DNA backbone in the minor groove that is also involved in the rocking motion, which corresponds to K207 in Max and K121 in Myc of MycMax modelled, or K121 and K212 in the Omomyc dimer, and K116 and K202 in MaxMax (see numbering convention in Fig. S3).
Overall, the structural dynamics along with stability and flexibilities of three complexes underscore how the dimeric bHLHLZ domain architectures are modulated by protein–DNA interactions. In particular, MycMax as a heterodimer displays notable asymmetry of flexibilities between its HLH domains on the E-box and very large deviations of Myc–BR occasionally on poly-A, while overall it shows significant LZ rocking motions on both DNA types. The pronounced rocking motion of MycMax on the E-box and occasional large BR deviation on polyA together indicate some structural instabilities and consequent energetic instabilities of MycMax on DNA, as we analysed below (by ΔGbind). Note that current simulations were conducted with only the ordered structural region of Myc modeled. The structural dynamics properties may however be modified if Myc becomes full-length including the disordered region. Additionally, we observed that the LZ region of the MaxMax homodimer exhibits greater flexibility on E-box DNA compared to poly-A DNA. This increased LZ flexibility may correlate with increased TF–DNA binding affinity, as MaxMax is found energetically more stable (by ΔGbind) on E-box DNA compared to MycMax and Omomyc, with the trend reversed on poly-A DNA (see the next subsection). These results thus set the stage for further investigations into the energetic basis of TF–DNA binding and the particular roles of key residues and flanking sequences in modulating the binding affinity and specificity.
TF–DNA binding energetic stability analysis via MMGBSA
MMGBSA provides a fast method to estimate binding free energies, although it is also known to overestimate the energies33 and yield large standard deviations (in our case, on the order of tens of KBT for ΔGbind). In the current study, we calculated TF–DNA binding free energy components via MMGBSA (eqn (1)) over the microsecond-long MD trajectories. We compared averages computed over the entire trajectory with those from earlier segments (100–300 ns, i.e., before major conformational changes in the BR occurred) and found that the overall energy trends remained unchanged (data not shown). From Fig. S10 and S11, we can see that around 100–200 ns, the total energy trends become consistent. Consequently, we consider our MMGBSA estimates to be significantly converged and report the averages over the entire trajectory. | ||
| Fig. 3 TF–DNA binding (free) energetics and correlation analyses. (A) and (B) Box plots showing MMGBSA energy decomposition for MycMax (blue), the Omomyc dimer (orange), and MaxMax (green) bound to E-box (A) or polyA (B) across six energy terms: ESURF (non-polar solvation energy), EEL (electrostatic energy), EGB (polar solvation energy), EGB + EEL (sum of polar solvation and electrostatic energies), VDW (van der Waals energy), and TOTAL (total binding energy ΔGbind from eqn (1)). Insets on the top right provide a closer view of VDW and TOTAL energy terms (with values labelled) for each complex. The violin plot insets represent the buried surface areas (BSA) between each TF dimer and either E-box or polyA DNA. (C) Cross-correlations between MMGBSA energy terms for MycMax on E-box (left) and polyA (right). (D) Minor (top) and major (bottom) groove widths measured along DNA position in the MycMax–DNA complex, with error bars showing the standard deviation across the trajectory. DNA alone width (without protein) is also shown in grey with a light grey band indicating its standard deviation. DNA numbering is re-scaled to have the central C in the E-box CACGTG as index 0. | ||
In contrast, electrostatic (EEL) and polar solvation (EGB) terms largely offset each other, resulting in small net electrostatic contributions similarly in three complexes. Consequently, the overall ranking of binding free energies closely follows the VDW strength and correlates particularly strongly with the buried surface area (BSA) at the protein–DNA interface. A correlation analysis (Fig. 3C on MycMax, and Fig. S12 on the Omomyc dimer and MaxMax) supported this observation, showing near-perfect correlation (r ≈ 0.98) between VDW and total binding energy ΔGbind, along with substantial correlation (r ≈ 0.87–0.89) between total binding energy and nonpolar solvation (ESURF). By contrast, EEL and EGB exhibited near-complete anti-correlation. Such patterns thus persist from MycMax (Fig. 3C) to the Omomyc dimer and MaxMax (Fig. S12). Interestingly, the correlation between the electrostatic/polar solvation terms and the other energy components varies significantly between systems (with the highest overall correlations in MaxMax and the lowest in the Omomyc dimer), however, with no clear trend directly related to overall binding stability.
On polyA DNA, where the unbound groove profile is uniform, MycMax induced the largest groove distortions (Fig. 3D, right). In contrast, Omomyc and MaxMax maintained groove widths similar to those of unbound DNA. Notably, these distortions did not correlate directly with changes in BSA or binding affinity. These observations again indicate that TF–DNA interactions not only reflect the DNA sequences but also reveal protein active actions and responses to modulate and fit to the DNA conformation. In essence, the TF–DNA interface is dynamic, with the TFs reshaping the DNA and supplying surface complementarity. This adaptive modulation at the TF–DNA interface likely contributes to the distinct binding stabilities observed among the complexes.
Conserved arginines provide key energetic stabilization at the bHLHLZ–DNA interfaces
It is also noteworthy that the energy decomposition for Omo1 bound to E-box shows an intermediate pattern between Myc and Max (Fig. 4A and C). This suggests that the charged residue mutations introduced to generate Omo1 not only promote homodimerization but also allosterically influence BR–DNA interactions. Finally, examination of the electrostatic (EEL) and polar solvation (EGB) contributions (Fig. S14B) confirms that these terms largely cancel at the E-box center, reinforcing that VDW interactions and overall interface complementarity are the primary determinants of TF–DNA binding energetics related stability.
HB patterns at the TF–DNA interface and KL divergence on sequence readout
In particular, one can compare HB patterns between Myc (in MycMax) and DNA with those of the Omomyc monomer and DNA, as the BRs presumably in association with DNA are identical in Myc and Omomyc. Interestingly, one can see that although Myc–DNA HB patterns are formed similarly to those of Omomyc1–DNA (Fig. 5A), Omomyc1 forms additional HBs with flank DNA on the preferred strand 2 (on 66DA) while Myc forms more HBs with flank DNA on the non-preferred strand 1 (e.g., on 15DT to 16DA).
These observations highlight a common recognition mechanism centered on a stable E-box core and a system-dependent use of flanking base contacts. MycMax and MaxMax both show more flanking HBs than Omomyc on the E-box (Fig. S17). We also noted a preference for single-strand binding within the E-box core observed as in recently studied monomeric TF domain protein WRKY on W-box,47,48 which may enhance sequence specificity. This idea was further supported by Kullback–Leibler (KL) divergence analyses (see below).
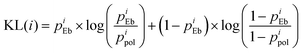 | (4) |
![[T with combining low line]](https://www.rsc.org/images/entities/char_0054_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) motif, indicating strong sequence specificity or information readout at these two nucleotide positions. In contrast, the non-preferred strand (i.e., with very few HB contacts) at the same positions showed near-zero KL divergence, suggesting minimal differences in HB contacts or not much DNA sequence information readout there. The central CA
motif, indicating strong sequence specificity or information readout at these two nucleotide positions. In contrast, the non-preferred strand (i.e., with very few HB contacts) at the same positions showed near-zero KL divergence, suggesting minimal differences in HB contacts or not much DNA sequence information readout there. The central CA![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) TG nucleotides (positions 0C and 1G) had lower KL values than the right-edge or outer (including the left-edge −2C and −1A) nucleotides, although the base of the central G is a key contributor to high-affinity E-box binding. The prominence of the outer edge nucleotides in sequence readout can be explained by the CANNTG definition of the E-box,17 where CA and TG steps are recognized as important for specificity, and NN at the center are less constrained for specific recognition.
TG nucleotides (positions 0C and 1G) had lower KL values than the right-edge or outer (including the left-edge −2C and −1A) nucleotides, although the base of the central G is a key contributor to high-affinity E-box binding. The prominence of the outer edge nucleotides in sequence readout can be explained by the CANNTG definition of the E-box,17 where CA and TG steps are recognized as important for specificity, and NN at the center are less constrained for specific recognition.
Outside the 6-bp E-box, certain flanking nucleotides also showed moderate to large KL peaks (KL values ∼1–2), particularly on strand 1, which is predominantly contacted by Myc, Omo1, and Max1 on the left flanking site (Fig. 6, top left).
Overall, the KL divergence analysis confirms that E-box versus polyA (Fig. S19–S20) binding sequence specificity or constraint is most pronounced at the edge of the E-box, and the DNA information readout is notable on the DNA strand each monomer prefers in association. Meanwhile, flanking DNA sites also contribute to sequence sensitivity, almost comparable to the specific DNA bound region.
Conclusions
In this study, we used microsecond-scale MD simulations and MMGBSA energy analyses to compare how the three bHLHLZ TFs MycMax, the Omomyc dimer, and MaxMax bind to a canonical E-box (5′-CACGTG-3′) versus reference polyA (5′-AAAAAA-3′) DNA. Despite sharing a common bHLHLZ structural architecture, our results show distinct structural dynamic behaviors and binding energetic stabilities among the three TF–DNA complexes.Our MMGBSA calculations reveal that TF–DNA VDW contacts and the BSA (buried surface area) at the interface dominate the binding free energy in all three dimers. This observation aligns with reports that shape complementarity at the protein–DNA interface can achieve binding specificity.41,43 Notably, MaxMax emerges as the most stabilized complex energetically on the E-box motif but undergoes the most destabilization on polyA. By contrast, the Omomyc dimer consistently exhibits slightly higher stability energetically than MycMax on both the E-box and polyA sequences.
These trends can be compatible with previous experimental studies on Myc and Max.9,10,15,16 For example, the Max monomer's high intrinsic affinity and its dimer's high affinity to E-box DNA15 can explain why MaxMax outperforms MycMax stabilizing on the E-box under certain conditions. Meanwhile, some other experiments also show MycMax outcompeting MaxMax on certain E-box variants.9 We reason that some discrepancies in experimental findings can be attributed to the construct of Myc (full-length or not) or DNA sequences flanking the E-box that contribute to the binding affinity measurements (Fig. 4C). Various flanking sequences may impact TF dimer stability on DNA differently, allowing MaxMax to outcompete MycMax on some occasions,15,17,18 or MycMax can emerge as the stronger binder.9 More importantly, MycMax's dimerization is much stronger than MaxMax's,9,15 which can translate into higher overall binding affinity or stability of MycMax to E-box DNA than that of MaxMax. To test this, experiments at very high concentrations of TF monomers (to prevent dimer dissociation) can be conducted. Besides, MaxMax can bind less favorably than MycMax to nonspecific DNA, as we show on polyA DNA in the current study. Hence, MaxMax can be overall less stable than MycMax on DNA, particularly with nonspecific binding contexts.
Our simulations also show that the Omomyc dimer retains slightly stronger binding than MycMax on DNA of both the E-box to polyA sequences, consistent with earlier EMSA data.9 Given that Omomyc and Myc share the same bHLH and BR domains, this enhanced binding appears to be allosteric, conferred from LZ mutations to the bHLH and BR–DNA interface. Interestingly, Omomyc dimer stability on DNA is least affected by the switch from E-box to polyA, and it may exhibit the highest affinity to nonspecific DNA, assuming similarly to polyA in the current study. This robust binding may allow Omomyc to effectively compete with and displace Myc from low-affinity DNA sites. In cancer cells, overexpressed Myc can abnormally occupy weak binding motifs, a phenomenon known as “promoter invasion”.10 By maintaining strong binding even on nonspecific DNA, Omomyc may help prevent this aberrant promoter occupancy. Prior studies10 similarly demonstrated that Omomyc maintains strong binding to noncanonical (“weak”) E-boxes, supporting our observation that Omomyc's robust binding range may underlie its therapeutic efficacy in disrupting Myc-mediated transcription.9–11
Our HB analyses confirm that each monomer BR in the dimer predominantly binds to one DNA strand, consistent with the biased DNA strand association reported for the monomeric TF domain protein.47,48 Notably, the interfacial TF–DNA HB contacts are highly conserved across the three systems, suggesting an evolutionarily preserved recognition mechanism, centered mostly at the arginine residues in each BR. These arginines exhibit multivalent HBs, particularly with guanine bases. Our KL divergence analysis further highlights how each monomer's preferred DNA strand shows the greatest difference on HB patterns between E-box and polyA DNA, i.e., to facilitate DNA sequence readout on one DNA strand, particularly on the E-box edge, though flanking bases also exhibit moderate divergence, underscoring the role of peripheral sequences in refining specificity.
In future studies, systematically varying flanking DNA elements could clarify the molecular basis of TF–DNA sequence sensitivity across diverse genomic contexts. Overall, our findings provide new avenues for rationally modulating bHLHLZ TF–DNA interactions and inform improved strategies to target Myc in cancer, potentially exploiting MaxMax's central role in binding specifically to E-box DNA and Omomyc's ability to outcompete Myc across both canonical and noncanonical sites.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: PDF of additional plots and measurements (.pdf), PDB files of all the systems studied (.zip), and movies from MD simulations (.zip). See DOI: https://doi.org/10.1039/d5cp03629a.Acknowledgements
CM and JY have been supported by UC Cancer Research Coordinating Committee (CRCC) award #C23CR5636. JY had been supported by the CMCF of UCI via the NSF grant DMS1763272 and the Simons Foundation grant #594598, and start-up fund from UCI School of Physical Science. CM has been supported by CMCF fellowship via the NSF grant DMS1763272, and the Simons Foundation grant #594598. This work utilized the high-performance computing, data storage, and software tools provided by the Research Cyberinfrastructure Center (RCIC) at the University of California, Irvine (UCI). Anton 2 computer time was provided by the Pittsburgh Supercomputing Center (PSC) on award MCB220008P. The Anton 2 machine at PSC was generously made available by D.E. Shaw Research.References
- P. J. Hurlin and J. Huang, The MAX-interacting transcription factor network, Semin. Cancer Biol., 2006, 16, 265–274 CrossRef CAS.
- B. Lüscher and J. Vervoorts, Regulation of gene transcription by the oncoprotein MYC, Gene, 2012, 494, 145–160 CrossRef.
- C. V. Dang, MYC on the path to cancer, Cell, 2012, 149, 22–35 CrossRef PubMed.
- M. Eilers and R. N. Eisenman, Myc's broad reach, Genes Dev., 2008, 22, 2755–2766 CrossRef CAS PubMed.
- S. K. Madden, A. D. de Araujo, M. Gerhardt, D. P. Fairlie and J. M. Mason, Taking the Myc out of cancer: toward therapeutic strategies to directly inhibit c-Myc, Mol. Cancer, 2021, 20, 3 CrossRef PubMed.
- B. Amati, M. W. Brooks, N. Levy, T. D. Littlewood, G. I. Evan and H. Land, Oncogenic activity of the c-Myc protein requires dimerization with Max, Cell, 1993, 72, 233–245 CrossRef CAS PubMed.
- Y. Pan, P. J. van der Watt and S. A. Kay, E-box binding transcription factors in cancer, Front. Oncol., 2023, 13, 1223208 CrossRef CAS PubMed.
- L. Soucek, M. Helmer-Citterich, A. Sacco, R. Jucker, G. Cesareni and S. Nasi, Design and properties of a Myc derivative that efficiently homodimerizes, Oncogene, 1998, 17, 2463–2472 CrossRef CAS PubMed.
- L. A. Jung, A. Gebhardt, W. Koelmel, C. P. Ade, S. Walz, J. Kuper, B. Von Eyss, S. Letschert, C. Redel, L. D’Artista, A. Biankin, L. Zender, M. Sauer, E. Wolf, G. Evan, C. Kisker and M. Eilers, OmoMYC blunts promoter invasion by oncogenic MYC to inhibit gene expression characteristic of MYC-dependent tumors, Oncogene, 2017, 36, 1911–1924 CrossRef CAS PubMed.
- M. J. Demma, C. Mapelli, A. Sun, S. Bodea, B. Ruprecht, S. Javaid, D. Wiswell, E. Muise, S. Chen, J. Zelina, F. Orvieto, A. Santoprete, S. Altezza, F. Tucci, E. Escandon, B. Hall, K. Ray, A. Walji and J. O’Neil, Omomyc Reveals New Mechanisms To Inhibit the MYC Oncogene, Mol. Cell. Biol., 2019, 39(22), e00248-19 CrossRef.
- D. Massó-Vallés and L. Soucek, Blocking Myc to Treat Cancer: Reflecting on Two Decades of Omomyc, Cells, 2020, 9(4), 883 CrossRef.
- E. Garralda, M.-E. Beaulieu, V. Moreno, S. Casacuberta-Serra, S. Martínez-Martín, L. Foradada, G. Alonso, D. Massó-Vallés, S. López-Estévez, T. Jauset, E. Corral de la Fuente, B. Doger, T. Hernández, R. Perez-Lopez, O. Arqués, V. Castillo Cano, J. Morales, J. R. Whitfield, M. Niewel, L. Soucek and E. Calvo, MYC targeting by OMO-103 in solid tumors: a phase 1 trial, Nat. Med., 2024, 30, 762–771 CrossRef CAS.
- M. Allevato, E. Bolotin, M. Grossman, D. Mane-Padros, F. M. Sladek and E. Martinez, Sequence-specific DNA binding by MYC/MAX to low-affinity non-E-box motifs, PLoS One, 2017, 12(7), e0180147 CrossRef PubMed.
- P. A. Kollman, I. Massova, C. Reyes, B. Kuhn, S. Huo, L. Chong, M. Lee, T. Lee, Y. Duan, W. Wang, O. Donini, P. Cieplak, J. Srinivasan, D. A. Case and T. E. Cheatham, Calculating Structures and Free Energies of Complex Molecules:
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Combining Molecular Mechanics and Continuum Models, Acc. Chem. Res., 2000, 33, 889–897 CrossRef CAS.
Combining Molecular Mechanics and Continuum Models, Acc. Chem. Res., 2000, 33, 889–897 CrossRef CAS. - J. Hu, A. Banerjee and D. J. Goss, Assembly of b/HLH/z Proteins c-Myc, Max, and Mad1 with Cognate DNA:
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Importance of Protein−Protein and Protein−DNA Interactions, Biochemistry, 2005, 44, 11855–11863 CrossRef CAS.
Importance of Protein−Protein and Protein−DNA Interactions, Biochemistry, 2005, 44, 11855–11863 CrossRef CAS. - O. Ecevit, M. A. Khan and D. J. Goss, Kinetic Analysis of the Interaction of b/HLH/Z Transcription Factors Myc, Max, and Mad with Cognate DNA, Biochemistry, 2010, 49, 2627–2635 CrossRef CAS PubMed.
- M. Allevato, E. Bolotin, M. Grossman, D. Mane-Padros, F. M. Sladek and E. Martinez, Sequence-specific DNA binding by MYC/MAX to low-affinity non-E-box motifs, PLoS One, 2017, 12, e0180147 CrossRef PubMed.
- J. Guo, T. Li, J. Schipper, K. A. Nilson, F. K. Fordjour, J. J. Cooper, R. Gordân and D. H. Price, Sequence specificity incompletely defines the genome-wide occupancy of Myc, Genome Biol., 2014, 15, 482 CrossRef.
- S. K. Nair and S. K. Burley, X-ray structures of Myc-Max and Mad-Max recognizing DNA: Molecular bases of regulation by proto-oncogenic transcription factors, Cell, 2003, 112, 193–205 CrossRef CAS PubMed.
- H. M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T. N. Bhat, H. Weissig, I. N. Shindyalov and P. E. Bourne, The Protein Data Bank, Nucleic Acids Res., 2000, 28(1), 235–242 CrossRef CAS PubMed.
- A. R. Ferré-D’Amaré, G. C. Prendergast, E. B. Ziff and S. K. Burley, Recognition by Max of its cognate DNA through a dimeric b/HLH/Z domain, Nature, 1993, 363, 38–45 CrossRef.
- UniProt: the Universal Protein Knowledgebase in 2025, Nucleic Acids Research, Oxford Academic, https://academic.oup.com/nar/article/53/D1/D609/7902999, (accessed 4 March 2025).
- L. Schrödinger and W. DeLano, PyMOL, 2020, Available at: http://www.pymol.org/pymol.
- S. Li, W. K. Olson and X.-J. Lu, Web 3DNA 2.0 for the analysis, visualization, and modeling of 3D nucleic acid structures, Nucleic Acids Res., 2019, 47, W26–W34 CrossRef CAS.
- T. J. Dolinsky, J. E. Nielsen, J. A. McCammon and N. A. Baker, PDB2PQR: an automated pipeline for the setup of Poisson–Boltzmann electrostatics calculations, Nucleic Acids Res., 2004, 32, W665–W667 CrossRef CAS PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers, SoftwareX, 2015, 1–2, 19–25 CrossRef.
- P. Bauer, B. Hess and E. Lindahl, GROMACS 2022 Source Code, 2022, https://doi.org/10.5281/zenodo.6103835.
- D. E. Shaw, J. P. Grossman, J. A. Bank, B. Batson, J. A. Butts, J. C. Chao, M. M. Deneroff, R. O. Dror, A. Even, C. H. Fenton, A. Forte, J. Gagliardo, G. Gill, B. Greskamp, C. R. Ho, D. J. Ierardi, L. Iserovich, J. S. Kuskin, R. H. Larson, T. Layman, L.-S. Lee, A. K. Lerer, C. Li, D. Killebrew, K. M. Mackenzie, S. Y.-H. Mok, M. A. Moraes, R. Mueller, L. J. Nociolo, J. L. Peticolas, T. Quan, D. Ramot, J. K. Salmon, D. P. Scarpazza, U. B. Schafer, N. Siddique, C. W. Snyder, J. Spengler, P. T. P. Tang, M. Theobald, H. Toma, B. Towles, B. Vitale, S. C. Wang and C. Young, in SC ’14: Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, 2014, pp. 41–53.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB, J. Chem. Theory Comput., 2015, 11, 3696–3713 CrossRef CAS.
- Refinement of the Sugar–Phosphate Backbone Torsion Beta for AMBER Force Fields Improves the Description of Z- and B-DNA, J. Chem. Theory Comput., https://pubs.acs.org/doi/10.1021/acs.jctc.5b00716, (accessed 3 March 2025).
- T. Darden, D. York and L. Pedersen, Particle mesh Ewald: An N
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log(N) method for Ewald sums in large systems, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
log(N) method for Ewald sums in large systems, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS. - M. Parrinello and A. Rahman, Crystal Structure and Pair Potentials: A Molecular-Dynamics Study, Phys. Rev. Lett., 1980, 45, 1196–1199 CrossRef CAS.
- S. Genheden and U. Ryde, The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities, Expert Opin. Drug Discovery, 2015, 10, 449–461 CrossRef CAS PubMed.
- B. R. I. Miller, T. D. McGee, J. M. Swails, N. Homeyer, H. Gohlke and A. E. Roitberg, MMPBSA.py: An Efficient Program for End-State Free Energy Calculations, J. Chem. Theory Comput., 2012, 8, 3314–3321 CrossRef CAS PubMed.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, Gmx_MMPBSA: A New Tool to Perform End-State Free Energy Calculations with GROMACS, J. Chem. Theory Comput., 2021, 17, 6281–6291 CrossRef PubMed.
- Interaction Entropy: A New Paradigm for Highly Efficient and Reliable Computation of Protein–Ligand Binding Free Energy, J. Am. Chem. Soc., https://pubs.acs.org/doi/10.1021/jacs.6b02682, (accessed 20 February 2025).
- L. Duan, G. Feng, X. Wang, L. Wang and Q. Zhang, Effect of electrostatic polarization and bridging water on CDK2–ligand binding affinities calculated using a highly efficient interaction entropy method, Phys. Chem. Chem. Phys., 2017, 19, 10140–10152 RSC.
- G. D. Hawkins, C. J. Cramer and D. G. Truhlar, Pairwise solute descreening of solute charges from a dielectric medium, Chem. Phys. Lett., 1995, 246, 122–129 CrossRef CAS.
- V. Tsui and D. A. Case, Theory and applications of the generalized born solvation model in macromolecular simulations, Biopolymers, 2000, 56, 275–291 CrossRef CAS.
- T. Tuccinardi, What is the current value of MM/PBSA and MM/GBSA methods in drug discovery?, Expert Opin. Drug Discovery, 2021, 16, 1233–1237 CrossRef CAS PubMed.
- R. Rohs, X. Jin, S. M. West, R. Joshi, B. Honig and R. S. Mann, Origins of specificity in protein–DNA recognition, Annu. Rev. Biochem., 2010, 79, 233–269 CrossRef CAS PubMed.
- B. Jayaram, K. McConnell, S. B. Dixit, A. Das and D. L. Beveridge, Free-energy component analysis of 40 protein–DNA complexes: A consensus view on the thermodynamics of binding at the molecular level, J. Comput. Chem., 2002, 23, 1–14 CrossRef CAS PubMed.
- R. Rohs, S. M. West, A. Sosinsky, P. Liu, R. S. Mann and B. Honig, The role of DNA shape in protein–DNA recognition, Nature, 2009, 461, 1248–1253 CrossRef CAS PubMed.
- R. Gordân, N. Shen, I. Dror, T. Zhou, J. Horton, R. Rohs and M. L. Bulyk, Genomic Regions Flanking E-Box Binding Sites Influence DNA Binding Specificity of bHLH Transcription Factors through DNA Shape, Cell Rep., 2013, 3, 1093–1104 CrossRef.
- N. M. Luscombe, R. A. Laskowski and J. M. Thornton, Amino acid–base interactions: a three-dimensional analysis of protein–DNA interactions at an atomic level, Nucleic Acids Res., 2001, 29, 2860–2874 CrossRef CAS PubMed.
- J. DeRouchey, B. Hoover and D. C. Rau, A Comparison of DNA Compaction by Arginine and Lysine Peptides: A Physical Basis for Arginine Rich Protamines, Biochemistry, 2013, 52, 3000–3009 CrossRef CAS PubMed.
- Revealing atomic-scale molecular diffusion of a plant-transcription factor WRKY domain protein along DNA, PNAS, https://www.pnas.org/doi/abs/10.1073/pnas.2102621118?url_ver=Z39.88-2003&rfr_id=ori%3Arid%3Acrossref.org&rfr_dat=cr_pub++0pubmed, (accessed 20 February 2025).
- Nonspecific vs. specific DNA binding free energetics of a transcription factor domain protein, Biophys. J., https://www.cell.com/biophysj/fulltext/S0006-3495(23)00661-6, (accessed 20 February 2025).
- S. Kullback and R. A. Leibler, On Information and Sufficiency, Ann. Math. Stat., 1951, 22, 79–86 CrossRef.
| This journal is © the Owner Societies 2025 |