Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Exploring excited-state proton transfer reactions in isothiazologuanosine, an isofunctional fluorescent analogue of guanosine
Olha
Tkach
a,
Lara
Martinez-Fernandez
b,
Atzin Esmeralda
Ruiz-Lera
a,
Nicolas
Humbert
a,
Ludovic
Richert
a,
Dmytro
Dziuba
 a,
Elisa
Bombarda
a,
Elisa
Bombarda
 a,
Aurélie
Bourderioux
a,
Fabien
Hanser
a,
Pascal
Didier
a,
Aurélie
Bourderioux
a,
Fabien
Hanser
a,
Pascal
Didier
 a,
Yitzhak
Tor
a,
Yitzhak
Tor
 c,
Roberto
Improta
c,
Roberto
Improta
 *d,
Jérémie
Léonard
*d,
Jérémie
Léonard
 *e and
Yves
Mély
*e and
Yves
Mély
 *a
*a
aLaboratoire de Bioimagerie et Pathologies, UMR 7021 CNRS Université de Strasbourg, Faculté de pharmacie 74 route du Rhin, 67401 Illkirch, France. E-mail: yves.mely@unistra.fr
bDepartamento de Química Física de Materiales, Instituto de Química Física Blas Cabrera, CSIC, Calle Serrano 119, 28006, Madrid, Spain
cDepartment of Chemistry and Biochemistry, University of California, San Diego, La Jolla CA 92093-0358, USA
dConsiglio Nazionale delle Ricerche, Istituto Biostrutture e Bioimmagini via De Amicis 95, 80145 Napoli, Italy. E-mail: roberto.improta@cnr.it
eUniversité de Strasbourg, CNRS, Institut de Physique et Chimie des Matériaux de Strasbourg, France. E-mail: jeremie.leonard@ipcms.unistra.fr
First published on 25th August 2025
Abstract
To fully exploit the potential of isothiazologuanosine (tzG), an isomorphic and isofunctional fluorescent analogue of guanosine, as a probe for DNA and RNA, we characterized its photophysics and in particular its excited-state reactions over a wide pH range (–0.6 to 12) and time scale (100 fs–100 ns) by combining transient absorption and time-correlated single photon counting measurements with quantum mechanical calculations. At acidic pH, the dominant ground-state species tzG-H1–H3+, where the N atoms in positions 1 and 3 are protonated, rapidly converts to the more stable tautomer tzG-H1–H7+ in its excited state. The latter then deprotonates to form the tzG-H1 neutral species with an excited-state pKa* value that differs by three pH units from the ground-state pKa value. The rate constants governing the excited-state reactions and the fluorescence lifetime of each species were all determined. With the exception of intramolecular and solvent relaxations, no excited-state reactions in the femtosecond to nanosecond time scale were, however, observed between the dominant tzG-H1 and tzG-H3 tautomers in equilibrium at neutral pH or for tzG-H1 deprotonation at high pH. Because of the distinct spectra, fluorescence quantum yields and lifetimes of its different protonated and deprotonated forms, tzG is highly responsive over a wide range of acidic (0–5) and basic pH values (8–10). The mechanisms revealed herein will be instrumental for tzG-labelled oligonucleotides in order to interpret proton transfer reactions as well as interactions with specific protein domains, which, due to local electrostatic changes and water exclusion effects, may shift the pKa values of tzG to a more physiologically relevant range.
1. Introduction
Understanding the key roles played by nucleic acids largely relies on the mechanistic investigation of the global and local conformational changes accompanying their dynamic interactions with diverse ligands. Fluorescence-based techniques are highly useful for such tasks due to their unrivaled sensitivity that allows one to monitor such conformational changes down to the single molecule level in a large range of time scales. Unfortunately, DNAs and RNAs suffer from the low intrinsic emission of their natural neutral nucleosides, as a result of very efficient nonradiative pathways that effectively thermally dissipate the electronic excitation by UV light absorption.1–5 These nonradiative pathways are thought to have been evolutionary relevant, leading to purine and pyrimidine building blocks that can tolerate harmful effects of UV light energy. The photophysics of natural nucleobases, both in their respective nucleosides and when incorporated into oligonucleotides, has been extensively investigated by a combination of femtosecond spectroscopy and quantum mechanical (QM) calculations, revealing that nonradiative pathways rely on barrierless transitions to conical intersections as well as on charge transfer and proton transfer processes.5–7 As a result of these very efficient nonradiative decay pathways, natural nucleosides cannot typically yield quantitative structural information from their spectroscopic data. Additionally, they suffer from exhibiting complex fluorescence decay kinetics with lifetimes spanning from the sub-ps to the ns range. This is further compounded by the fact that nucleic acids contain multiple copies of the four natural nucleobases, precluding residue-specific probing of local structural and dynamical properties.To lift these practical experimental barriers, a large number of fluorescent nucleoside analogues (FNAs) have been implemented,8–10 including for instance the popular 2-aminopurine (2AP),11,12 a fluorescent constitutional isomer of adenine, the tricyclic cytosine family13–17 and pyrrolocytosine.18–20 FNAs can be selectively incorporated into DNA/RNA oligonucleotides by solid-phase synthesis or enzymatic approaches. FNAs typically exhibit a red-shifted absorption compared to proteins and natural nucleosides, and can thus be excited selectively. However, they are frequently quenched and induce local structural perturbations when incorporated into nucleic acids.10,21 A breakthrough has been achieved with the development of FNAs based on thieno[3,4-d]pyrimidine (thN) and isothiazolo[4,3-d]pyrimidine (tzN).22,23 In particular, the guanosine analogues thG and tzG (Fig. 1) perfectly replace G in DNA/RNA duplexes, being truly isomorphic, while keeping high fluorescence quantum yields (ϕ) and environmental sensitivity.23–26 The fluorescence properties of thG both in its free form and incorporated in nucleic acids have been largely explored.27–32 Free thG in buffer exists as two keto-amino tautomers (thG-H1 and thG-H3) in ground-state equilibrium with distinct spectral properties but similar quantum yield (ϕ =0.50).27,30 In DNA duplexes, the thG-H1 tautomer, which is favored by Watson–Crick base pairing, has a constrained orientation and distance from adjacent bases.29 As a result, its ϕ and lifetime values are nearly independent of the flanking bases. Another key advantage of thG is its exceptionally long fluorescence lifetime in DNA duplexes (9–29 ns), allowing selective monitoring in complex environments and measurement of the rotational diffusion of protein-complexed duplexes.33 However, thG has an important limitation, i.e. the lack of N7 atom (Fig. 1), which is instead present in tzG. Since the purines’ N7 is involved in forming higher structures (e.g., triple helices, G-quadruplexes),34,35 and in protein–nucleic acid complexes, tzG is not only isomorphic, but also isofunctional to G.23 This was notably demonstrated by its deamination by guanine deaminase36 or its incorporation into cyclic dinucleotides by a bacterial enzyme.37 Therefore, tzG appears as a unique FNA for characterizing the local conformations, dynamics and molecular interactions of unperturbed nucleic acids.
 | ||
| Fig. 1 (A) Structure of guanosine (G), thienoguanosine (thG) and isothiazologuanosine (tzG). The tzG structure shown corresponds to the tzG-H1 tautomer where the N atom at position 1 is protonated. At neutral pH, this tautomer is in equilibrium with the tzG-H3 tautomer where instead the N atom at position 3 is protonated. Absorption (B) and emission (C) spectra (black lines) of tzG in 20 mM Hepes buffer at pH 7 are deconvoluted into the sum of the spectra of tzG-H1 (red lines) and tzG-H3 (blue lines) in equilibrium.39 Inset: zoomed tzG-H3 emission spectrum. | ||
In contrast to thG, only limited spectroscopic characterization has been performed on free tzG or tzG-labeled oligonucleotides.23,38 In a preliminary study, we have recently explored the steady-state and time-resolved (in the ps–ns range) fluorescence features of free tzG in pH 7 buffer and various solvents and interpreted the resulting data by QM calculations.39 Although the tautomeric equilibrium of tzG is similar to that of thG (Fig. 1B), differences were observed: (a) tzG displays an additional nonradiative decay channel via a conical intersection involving the NS bond elongation, responsible for a greater environmental sensitivity of the ϕ value of tzG-H1 tautomer as compared to thG-H1, and (b) a highly efficient intersystem crossing (ISC) for tzG-H3, which contributes to reducing its ϕ value by two orders of magnitude as compared to thG-H3. These data highlight the significant effect of the N7 atom on the photophysics of tzG, and tzG-H3 in particular.
While the ultimate goal would be to investigate the photophysics of tzG in oligonucleotides, our aim in the present study is to fully disentangle the photophysical processes occurring in free tzG on a fs–ns time scale, with a special focus on the possible excited-state protonation/deprotonation reactions. This knowledge will be critical to properly interpret in a later step the excited-state reactions that likely modulate tzG emission in oligonucleotides, as it happens for canonical bases.5–7 Hence, we combine transient absorption spectroscopy (TA), time-resolved fluorescence spectroscopy using time-correlated single photon counting (TCSPC), steady-state spectroscopy and QM calculations, to investigate the photophysics of free tzG as a function of pH. Our data allow us to draw up a complete scheme of the species involved and their interdependence in the ground and excited states, illustrating the high potential of tzG as a pH probe and providing a solid foundation for interpreting proton-transfer reactions in oligonucleotides.
2. Materials and methods
2.1. Materials and sample preparation
The ribonucleoside isothiazologuanosine (tzG) was synthesized as previously described by Rovira et al.23 and stored at –20 °C as a powder. Stock solutions of tzG (3.46 mM) were prepared in spectroscopic grade DMSO (Sigma-Aldrich, 99.9% purity) and stored at –20 °C. For steady-state and TCSPC spectroscopic measurements at different pH values, the tzG stock solution was diluted in buffers at concentrations of 10–25 μM (<1% DMSO). The samples were prepared freshly for each measurement. HCl was used for pH ≤ 2, 20 mM sodium citrate buffer for pH 3, 20 mM sodium formate buffer for pH 4, 20 mM sodium acetate buffer for pH 5, 20 mM sodium 2-(N-morpholino)ethanesulfonate (MES) buffer for pH 6, 20 mM sodium (4-(2-hydroxyethyl)-1-piperazineethanesulfonate) (HEPES) buffer for pH 7, 20 mM sodium TRIS buffer for pH 8, 20 mM sodium N-cyclohexyl-2-aminoethanesulfonate (CHES) buffer for pH 9, 25 mM sodium borate buffer for pH 10, 20 mM sodium 3-(cyclohexylamino)-1-propanesulfonate (CAPS) for pH 11, and NaOH for pH 12. The reagents for buffer preparation were purchased from Sigma-Aldrich, except sodium acetate (Merck) and TRIS (Serva). The buffers were prepared in Milli-Q water. The pH was adjusted by adding concentrated HCl or NaOH solutions, and monitored using a pH meter. Steady-state measurements of tzG at 15 μM concentration were also performed in diluted HCl from pH 2 to −0.6 ([H+] = 10−2 − 4 M) with 0.3 pH step. For TCSPC measurements of tzG at pH –0.6 (4 M HCl), a tzG stock solution (4 mM) in 4 M HCl was used to prepare a sample at 0.18 mM. At the most acidic pH, the tzG solution proved to be stable, since after a 48 h incubation, restoring a neutral pH led to the absorption and emission spectra that overlapped entirely with those of a freshly prepared solution at the same concentration.For TA spectroscopy, we prepared volumes of 1.5 to 2.5 mL of tzG solutions to be circulated using a peristaltic pump within a flow cell with a 0.5 mm optical path. To collect TA with optimum signal-to-noise ratio in such conditions, a typical 2 mM concentration is ideal. This is however a challenge, because tzG is most soluble in water at acidic pH, but less at neutral or basic conditions. Consequently, we devised the following experimental protocol. We solubilised 1.9 mg of tzG powder in 100 μL of DMSO, to obtain a concentration of 63.3 mM. This initial tzG solution was then diluted with 2.5 mL of a freshly prepared stock solution of 50 mM HEPES/KOH buffer (pH 7.5). Then, 65 μL of 5 M HCl was added to produce the pH 2 sample, which was first investigated by TA spectroscopy. Then, we used the same sample to acquire sequentially the TA data at pH 7 and pH 12 after adding the required amount (a few μL) of 12 M KOH and reducing the tzG concentration, sequentially. By carefully measuring the absorption spectra (in 1 mm cuvettes) before and after each TA acquisition, we observed no detectable photodegradation along the successive TA acquisition runs, and determined the concentrations of each sample to be 2.3 mM, 1.2 mM and 0.2 mM for the TA data recorded at pH 2, 7.5 and 12, respectively.
2.2. Steady-state spectroscopy
Absorption spectra were measured on Cary 4000 HP (Agilent) and UV-2700i (Shimadzu) UV-visible spectrophotometers. All measurements were done in quartz cuvettes with 1 cm path length. Fluorescence spectra were collected at 20 °C with Fluoromax 4 and Fluoromax Plus spectrofluorometers (Jobin-Yvon) equipped with a thermostated cell compartment. Emission and excitation spectra were corrected from buffer contribution, lamp spectral intensity fluctuations, detector spectral sensitivity and instrumental wavelength-dependent bias. The fluorescence quantum yields (ϕ) of tzG at different pH were determined by comparison with quinine sulfate (QS) in 0.5 M H2SO4 (ϕr = 0.546), taken as a reference40via the following equation: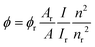 | (1) |
The pH dependence of tzG absorption maxima (λabs) and ϕ were fitted with the following modified Henderson–Hasselbach equation:31
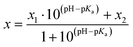 | (2) |
The radiative rate constant of the tzG-H1–H3+ acidic species was calculated using the Strickler–Berg equation:42
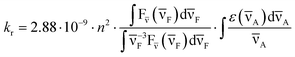 | (3) |
2.3. Time-resolved fluorescence measurements and data analysis
The time-correlated single-photon counting (TCSPC) technique was implemented in a home-made set-up with a pulse-picked supercontinuum white-light laser (EXR-20, NKT Photonics) equipped with an UV extension (SuperK Extent UV, NKT Photonics), which allows exciting tzG at λexc = 320 and 370 nm with 19.5 MHz repetition rate. tzG fluorescence decays were recorded over the whole emission spectrum using a cut-off filter KV 345 nm or KV 389 nm and a band-pass monochromator (Jobin-Yvon H10) with 16 nm slit. Photons were collected via a microchannel plate Hamamatsu R3809U photomultiplier coupled to a pulse preamplifier (HFAH 20, Becker & Hickl). The instrumental response function (IRF) recorded with a polished aluminium reflector had a full width at half maximum (FWHM) of 50–70 ps. Alexa 488 with lifetime of 4.1 ns and QS with lifetime of 19.5 ns were used as references for lifetime measurements.43,44 Fluorescence decays were analysed by the global analysis method with non-linear least-squares minimisation method and the use of the experimental IRF in the DecayFit software.45 The fits were performed with an increasing number of exponentials until the χ2 value reached a value close to 1 and a fully random distribution of the residuals around the zero value was achieved. The optimal number of components was further validated by the maximum entropy method (MEM).46 A component with negative amplitude was included in the fitting equation only when decay curves with initial rising part were observed. In these cases, the introduction of such a component with negative amplitude substantially improved the minimization of χ2 and the distribution of the residual at early time points. The decay-associated spectra (DAS)47 were calculated by: Ii(λ) = I(λ)·αi(λ)·τi/Σjαj(λ)·τj, where τi are the lifetimes and αi(λ), the associated amplitudes at emission wavelength λ, and I(λ), the steady-state emission spectrum.For the final description of the excited-state protonation/deprotonation reactions at acidic pH, the solution of the system of differential equations was calculated via IgorPro (WaveMetrics) using a Gaussian IRF with a FWHM of 50 ps.
2.4. Transient absorption data acquisition and treatment
The transient absorption (TA) spectroscopy setup is implemented using a 800 nm, 40 fs pulse generated by a Ti: sapphire regenerative amplifier operating at 5 kHz. A beam splitter is used to split a weak fraction of the energy (few μJ), which is focused in a vertically oscillating CaF2 plate to produce a broadband supercontinuum spanning 300–700 nm – i.e. a pulse of white light (WL) – and used as the probe beam. The remaining energy (∼0.4 mJ) is used in an optical parametric amplifier (TOPAS, Light Conversion) followed by subsequent sum frequency and/or second harmonic generation, to produce a 60–80 fs pump pulse at tunable wavelength. The pump wavelength λpump is adjusted to 300 nm or 370 nm, depending on the samples. Both pump and probe pulses are focused to ∼80 and 40 μm diameter respectively, and overlapped in a 0.5-mm-thick fused-silica flow cell. The pump power is adjusted to remain in the linear regime of excitation, i.e., we check that the TA signal scales linearly (at all time delays) with the pump power.For each pH value and/or pump wavelength, two TA datasets are acquired sequentially with two different spectral detection windows, i.e, UV (300–400 nm) and Vis (360–680 nm). After each acquisition, the TA data of the pure solvent (i.e. water buffered at various pH values) is systematically recorded. The solvent signal – also called “solvent artifact” - results from the pump and probe non-linear interaction in the solvent around the zero time delay corresponding to the temporal overlap of pump and probe pulses. The recorded solvent signal is subtracted from the raw data obtained for tzG in solution. In our case, the extinction coefficient and concentration of tzG are relatively low, such that the solvent artifact is comparatively strong. In most cases, the solvent signal subtraction is not perfect and a significant contribution of solvent artifact remains in a ±80 fs time range around time zero. Then, for a given pH value and pump wavelength, the ‘solvent-corrected’ UV and vis TA datasets are appended to produce a unified dataset covering the 300–680 nm probing spectral range. All UV-vis data presented here are then processed in order to correct the effect of the group velocity dispersion (GVD) in the WL probe pulse, as observed in the solvent artifact, such that the zero time delay is defined within ∼20 fs over the entire UV-vis probing window.
The above procedures for solvent subtraction, GVD correction and dataset appending constitute the usual data “pre-processing”, which is performed prior to any quantitative data analysis. The latter is done by global fitting of the entire two-dimensional maps, under the usual assumption that wavelength (λ) and time (t) are separable variables. The time evolution is modelled with a multiexponential function convolved with a normalized Gaussian function G(t) of standard deviation σ accounting for the instrument temporal response function (IRF). The values and number of time constants τi of the multiexponential model are shared across the entire dataset (i.e., at all wavelengths) and only the pre-exponential factors Ai(λ) depend on the wavelength and are thus referred to as ‘Decay-associated spectra’ (DAS):
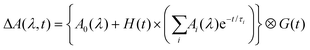 | (4) |
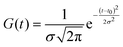 | (5) |
In addition, we account for a so-called “non-resolved component” that would correspond to time scales τi faster than the IRF and resulting (after convolution) in a Gaussian component G(t). Eventually, the fitting function can be expressed in the following way after analytically calculating the convolution for the “time-resolved” components:
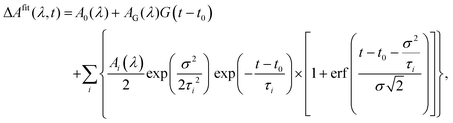 | (6) |
The global fit of entire datasets is performed by minimizing the residuals using an in-house Python implementation of the VARPRO algorithm,48,49 and seeking for one Gaussian component and 4 to 5 exponential components, depending on the datasets. In practice, neither the solvent signal subtraction nor the chirp correction are perfect. Consequently, even if no “non-resolved component” really exists in the photoreaction kinetics, the Gaussian component is always useful to improve the fit quality around time zero (t = t0). Hence, the corresponding amplitude – i.e. the “Gaussian-associated spectrum” also referred to as a non-resolved “Decay-associated spectrum” (DAS) – may not strictly reveal a non-resolved molecular dynamics but often accounts for unavoidable experimental artefacts around time zero. In the following, we display exclusively the DAS corresponding to time-resolved decay components, i.e., corresponding to time constants significantly larger than σ (the RMS width of the Gaussian component), i.e. τi ≳ 1.5 σ, typically. For some of the TA datasets recorded here on tzG, where the solvent artifact correction was unsatisfactory, we also tested a global fitting of the dataset restricted to times larger than ∼150 fs. We systematically obtained the same ‘time-resolved’ components.
2.5. Quantum mechanical calculations
We resorted to a computational strategy similar to the one successfully implemented in our previous studies on thG30,31 and canonical nucleobases.7 In our computational model, the sugar was mimicked by a simple methyl group, enabling to reduce the computational cost of the calculations, while including most of the electronic effect of the sugar on the excited states of tzG. However, it should be kept in mind that our model cannot treat conformational equilibria involving the sugar, such as the syn/anti conformational change.Our reference electronic method was density functional theory (DFT) for ground state calculations and its time-dependent extension (TD-DFT) for excited electronic states. We selected PBE0 and M052X functionals, which provide an accurate description of the tautomeric equilibria of tzG39 and the spectral properties of the main tautomers.39 Our reference functional will be M052X,50,51 which, despite a general overestimation of the excitation energies, provides a more solid basis for the extension of this study to DNA sequences containing tzG. Indeed, it can reliably treat stacking interactions and electronic transitions with significant charge transfer (CT) character and it has been profitably used to study the photophysics and photochemistry of NAs.7,50–52 We resort to PBE0 functional to double check the pKa predictions obtained with M052X, obtaining very similar trends (see SI).
We will mainly discuss the results obtained with the cost-effective 6-31+G(d,p) basis set.
Solvent effects were included by a mixed continuum/discrete model, using the polarizable continuum model (PCM)53,54 to describe solvent effects and explicitly considering 6 water molecules of the first solvation shell in the calculations. Absorption energies have been computed at the solvent non-equilibrium level, while emission energy at the equilibrium one. We have assumed that bulk solvent effects do not depend on the pH of the solution, in agreement with the procedure commonly adopted in the literature to compute acid/base pKa with mixed discrete continuum models.55 The excited-state absorption spectra were simulated by non-equilibrium TD-DFT calculations and using the multiwfn56 program to obtain the transition dipole moments between excited states. Using a protocol previously described,30 we provide an estimate of radiative lifetimes, knowing that absolute numbers are extremely dependent on the accuracy of the emission energies and the transition dipole moments.
All the calculations were performed with the Gaussian16 package.57
3. Results
3.1. tzG at neutral pH
Since the absorption and emission spectra of tzG in buffer at pH 7 represent the contribution of the equilibrating tzG-H1 and tzG-H3 tautomers (Fig. 1B and C),39 we measured TA spectra at pH 7 with two successive excitation wavelengths, at 370 nm, where only tzG-H1 absorbs, and at 300 nm, where both tautomers absorb and thus, both excited states (tzG-H1* and tzG-H3*) are populated. Fig. 2 compares the TA data recorded on tzG at pH 7 for both excitation wavelengths.Upon 370 nm excitation, the TA data (Fig. 2A and C) of tzG at pH 7 are characterized by a ground state bleach (GSB, negative signal) at wavelength λ < 350 nm. This is the signature of ground state tzG-H1 depopulation due to the formation of the excited state tzG-H1*. Hence, we expect to observe the spectral signatures of the excited state tzG-H1*, which are, a priori, stimulated emission (SE - negative ‘absorption’ expected to occur in the spectral range where tzG-H1* emits, i.e. around 460 nm, Fig. 1C) as well as excited state absorption (ESA, positive signal). Here, SE and ESA overlap spectrally, with ESA being more intense and, thus, resulting in an overall positive signal at λ > 380 nm. By 300 ps (Fig. 2C, green spectrum), the ESA spectrum is maximum at λmax ⋍ 410 nm and shows a local intensity minimum at 490 nm, attributed to the SE signal. On earlier time scales, i.e. from 200 fs to few tens of ps, the main ESA band slightly red-shifts and its intensity increases, while the local intensity minimum red-shifts from 450 nm to 490 nm (Fig. 2C). This is attributed to the red-shift of the overlapping SE resulting from solvent and intramolecular vibrational relaxations in the excited state. On time scales larger than 300 ps, the overall TA intensity decays with no further change in spectral shape. Global multiexponential fitting of the TA dataset (see Materials and methods) reveals the time scales associated with (i) the spectral relaxation indicating tzG-H1* vibrational and solvent relaxation and (ii) tzG-H1* decay on longer time scales. The corresponding decay-associated spectra (DAS) are plotted in Fig. 3A. Up to three time constants are needed to fit the early spectral relaxation. The three corresponding DAS have a similar dispersive shape with negative (resp. positive) amplitude in the range 400–420 nm (resp. 480–550 nm) supporting the interpretation of a red-shift of a negative SE signal with a distribution of time scales from 300 fs to 80 ps. The lifetimes τ1 and τ2 are typical for intramolecular and solvent relaxations, respectively. The origin of the τ3 component (83 ps) is less clear. Finally, a fourth time constant of 8 ns with a DAS showing a simultaneous decay to zero of the GSB, ESA and SE, describes the decay of tzG-H1* to the ground state tzG-H1 on this time scale. Since the largest pump–probe delay in the present TA experiment is no more than 5 ns, such a long lifetime is not accurately determined. In contrast, TSCPC experiments are much more accurate in this time range, revealing that tzG-H1* lifetime is 9.0 ± 0.2 ns, independent of the pH value between 5 and 7 (Fig. S2).
 | ||
| Fig. 3 Decay-associated spectra (DAS) resulting from the global analysis of TA data of tzG in HEPES buffer at pH 7 recorded with (A) 370-nm pump and (B) 300-nm-pump. (C) Simulated absorption spectra of the first excited S1 state of tzG-H1 and tzG-H3 tautomers based on PCM/TD-M052X/6-31+G(d,p) calculations. The grey area in panel A corresponds to the spectral window displaying enhanced noise due to 370 nm pump light scattering. Straight lines arbitrarily connect the short and long wavelength sides of the grey box in panel A as a guide to the eye. We note that the 84 ps DAS (Fig. 3B) shows a negative amplitude – i.e. a SE signature around 420 nm, while tzG-H3 emission maximum is at 382 nm (Fig. 1B, insert). This shift is probably due to the fact that the 84 ps DAS is the sum of the SE with a particularly intense ESA band around 350–380 nm (Fig. 3C). | ||
Upon excitation at 300 nm of tzG at pH 7 (Fig. 2B and D), we expect to observe both tzG-H1* and tzG-H3* signatures overlapping and decaying in parallel. While GSB is observed at λ < 335 nm, the TA signal is dominated by ESA signatures at λ > 335 nm which thus overlap less intense SE signals expected around 460 nm and 380 nm, respectively (Fig. 1C). As compared to the data recorded with the 370 nm pump, the ESA is globally blue-shifted during the first few picoseconds (blue and orange spectra in Fig. 2D), with λmax ⋍ 390 nm, a local intensity minimum around 440 nm and a secondary maximum around 480 nm. By 300 ps (Fig. 2D, green spectrum), the TA spectrum resembles more that of tzG-H1* with λmax ⋍ 410 nm, but its main positive band remains broader on its blue side. The DAS obtained from the global analysis of the 300 nm-pumped data are shown in Fig. 3B. Since long lifetimes are not accurately determined in the TA experiment, we actually force τ3 = 9 ns in the global fit and observe that the corresponding DAS is characterized by the 410 nm ESA band previously assigned to tzG-H1* (8 ns-DAS in Fig. 3A). While forcing τ3 value does not affect the shorter τ1 and τ2 values or DAS shapes, it clearly reveals that an even longer component, which we therefore force to τ4 = ∞, is required for a good fit of the TA data. Hence, unlike in the 370 nm-pumped data, a species, possibly a photoproduct, is observed to live longer than tzG-H1*. Its DAS is characterized by a residual GSB around 330 nm and a dominating absorption around 380 nm. A second absorption band is also observed around 460 nm, but we cannot rule out that it appears artificially to compensate for the negative amplitude observed at the same wavelength in the 9 ns DAS, but not in the 8 ns DAS obtained in the 370 nm-pumped data. Since no such ∞ DAS is needed to fit the TA data recorded upon excitation of tzG-H1 at 370 nm, this photoproduct likely results from tzG-H3 excitation at 300 nm.
In addition to the fixed τ3 and τ4 values, no more than two lifetimes, freely adjusted by the least-square minimization routine, are required to achieve an optimum fit of the 300-nm pumped TA data. The first one is τ1 = 600 fs with a DAS showing a dispersive shape, likely again indicative of fast excited state relaxation (possibly SE red-shifts for both tzG-H1* and tzG-H3*). The second is τ2 = 84 ps with a DAS dominated by two ESA bands: a broad one with λmax ∼ 390 nm and the other one with λmax ∼ 490 nm. The two bands are separated by a negative signal around 420 nm, indicative of SE decay on this time scale. We attribute this DAS to the TA signature of the tzG-H3* tautomer, thus characterized by a lifetime of 84 ps. To confirm this assignment, TCSPC experiments were performed to record the fluorescence decay kinetics of tzG at pH 7 upon excitation at 320 nm, where both tzG-H1 and tzG-H3 absorb equally. Global fitting of the decays in the 380–430 nm detection range (Fig. 4A and C) reveals two dominant time constants of 9.0 ns and 0.1 ns, close to the TA component of 84 ps (Fig. 3B). The corresponding DAS overlap very well with the steady-state emission spectra assigned to tzG-H1 and tzG-H3, respectively (Fig. 4D). These data justify the above analysis of the 300 nm-pumped TA data and in particular (i) the search for a fixed 9.0 ns component assigned to tzG-H1*, and (ii) the attribution of the 84 ps DAS to tzG-H3*.
Finally, we have computed the ESA spectra associated to the lowest energy excited state minima of tzG-H1* and tzG-H3*, the most stable tautomers of tzG (Fig. 3C). The predicted ESA spectra compare well with the experimental TA spectra, which are, in fact, the sum of GSB, ESA and SE spectra, assigned to tzG-H1* and tzG-H3*, confirming the attribution of these species at pH 7.
In conclusion, our data underline the complementarity of TA and TCSPC experiments with the 9 ns tzG-H1* lifetime being best determined with the latter, and the 84 ps lifetime of tzG-H3* with the former. In terms of excited-state reactions, only very fast solvent and intramolecular vibrational relaxations were detected. In contrast, there was no evidence of conversion between the tautomers in the excited state, indicating that this conversion occurs at a much slower time scale than their fluorescence lifetimes. This conclusion is supported by the absence of any tzG-H3 emission, when tzG is excited at 370 nm, where only tzG-H1 absorbs.
3.2. tzG in acidic solutions
While the spectroscopic properties at pH 5 and 6 were indistinguishable from those at pH 7 (Fig. S2D and data not shown), we observed a progressive blue shift of the tzG absorption spectrum from pH 4 to 1 (Fig. S3A–D), due to the conversion of the neutral species tzG-H1 and tzG-H3 into a protonated species denoted A, with a pKa = 3.5.23 At pH 1, the absorption spectrum of tzG shows a maximum at 315 nm (Fig. 5A), and an extinction coefficient of 5710 M−1 cm−1. The excitation spectra of tzG at pH 1 recorded at different emission wavelengths overlap well with the absorption spectrum (Fig. S3A), suggesting that only the A species exists in the ground state. Using the known absorption spectra of A, tzG-H1 and tzG-H3, their individual contributions at each pH value could be deconvoluted from the absorption spectra (Fig. S3). These deconvolutions confirm that the deprotonation of A to tzG-H1 is governed by a pKa value of 3.5 and that tzG-H1 equilibrates with tzG-H3 in a pH-independent manner, as reported for thG.31The emission spectra of tzG at pH 2, 3 and 4 (Fig. 5A) show a very large Stokes shift and overlap well with the emission of tzG* at pH 7, suggesting that in its excited state, A* deprotonates into tzG-H1*, the very dominant emissive species of tzG. At pH 1, the overlap with tzG-H1* emission was also high, but with a small broadening in the red part of the spectrum. The small blue-shifted shoulder of tzG-H3* (350–400 nm) was observed to decrease with pH when the pH value was decreased from 7 to 2, then to increase again at pH 1, suggesting that another tzG* species emitting in a wavelength range comparable to that of tzG-H3* probably replaces the latter at this pH value. The ϕ value of tzG at pH 1 is 0.03 ± 0.01, independently of the excitation wavelength λexc (Fig. 5B). The ϕ values at pH 2 (0.13 ± 0.01) and 3 (0.20 ± 0.01) were found constant at λexc < 350 nm. At λexc ≥ 350 nm, the ϕ values further increase, due to the direct excitation of tzG-H1, which is the major absorbing species at these wavelengths. At pH 4, tzG-H1 and tzG-H3 tautomers become dominant over A (Fig. S3D), and the ϕ values decrease for λexc < 320 nm, likely as the result of the increasing relative population of the poorly fluorescent tzG-H3 species in this λexc range.
The corresponding time-resolved fluorescence decays at pH 1–4 measured by TCSPC with λexc = 320 nm are well fitted with a two-exponential function for pH 2–4 (Fig. S4–S6). A three-exponential fit was used at pH 1, but with a 2 ns component of negligible amplitude, which was subsequently ignored (Fig. S7). The fastest decay component (0.1–0.2 ns) dominates at detection wavelengths <430 nm and its amplitude becomes negative at wavelengths >480 nm. Therefore, this component suggests an excited-state reaction, in agreement with the conversion of A* to tzG-H1* deduced from the steady-state data. In line with the decrease of ϕ with pH, the longest-lived component that dominates at longer detection wavelengths exhibits a progressive decrease of its value: 9.3 ns at pH 4, 8.5 ns at pH 3, 4.6 ns at pH 2 and 0.8 ns at pH 1, respectively.
TA data recorded for tzG at pH 2 with λpump = 300 nm are displayed in Fig. 6A. It is observed that the GSB at wavelengths <330 nm does not recover over time. At wavelengths >330 nm, ESA dominates over the entire visible spectral window. An overlapping SE around 430 nm is observed during the first few ps. At later times, it is replaced by an enhanced ESA while the SE red shifts toward ∼480 nm (Fig. 6B). The global, multiexponential fit (DAS in Fig. 6C) reveals the time scale associated to these spectral evolutions. The shortest time constant of 0.7 ps (DAS plotted in blue in Fig. 6C) is associated with the SE redshift accompanying early vibrational/solvent relaxation in the excited A* state. The following two time scales of 5.4 ps and 49 ps are characterized by almost identical DAS, showing the decay of the 430 nm SE and simultaneous rise of the ESA at 410 nm. The two DAS (especially 5.4 ps) are also slightly positive around 330 nm, which accounts for an apparent deepening of the GSB (Fig. 6B) but should actually be interpreted as a redshift of the high-energy side of the overlapping ESA. Moreover, Fig. 6D shows an excellent overlap of the 3.4 ns DAS at pH 2 with the 8 ns DAS assigned to tzG-H1* at pH 7. On this ground, we attribute the 3.4 ns species to tzG-H1*, produced by excited state proton transfer (ESPT) on earlier time scales (5.4 ps and/or 49 ps, see discussion below). This assignment, supported by QM calculations (see below), is confirmed by TCSPC data revealing a 4.6 ns tzG-H1* emission following the excitation of A at pH 2 (Fig. S6). Again, the 4.6 ns lifetime determined by TCSPC should be considered more accurate than the 3.5 ns component obtained from TAS. Finally, a very long-lived “∞” DAS is observed, which is composed of the GSB and a photoproduct absorbing at 350 nm, whose nature will be discussed below. The excellent overlap of the two DAS in Fig. 6D further reveals that following 300 nm excitation of tzG at pH 2, tzG-H1* decays directly to the ground state as observed above upon direct excitation of tzG-H1 at 370 nm at pH 7. Thus, the long-lived photoproduct detected here is formed already on an earlier time scale, in parallel to the formation of tzG-H1*, i.e. by 50 ps at maximum.
 | ||
| Fig. 6 TA data of tzG at pH 2 upon 300 nm excitation. (A) The 2D map represents the TA data (or pump-induced absorption change ΔA) coded in false colors, as a function of the pump–probe delay (in ps) and probe wavelength (nm). The time axis is displayed in a linear scale at early times (–0.5–1.0 ps) and logarithmic scale beyond. The short-lived signal around time zero (best seen at λprobe < 320 nm) is the solvent artifact. (B) Selection of TA spectra illustrating the growth of the 410 nm ESA band, and simultaneous deepening of the GSB at 330 nm. (C) DAS resulting from the multiexponential global fit of panel A. Only the time scales assigned to tzG spectral evolution are displayed (i.e. the solvent artifact is disregarded). (D) Comparison of the 3.4 ns DAS at pH 2 with the 8.0 ns DAS from Fig. 3A attributed to tzG-H1* at pH 7, normalized at their maximum. The quality of the fits of the data in panel A is illustrated in Fig. S1C. | ||
Since a small blue-shifted shoulder as well as a broadening in the red part of the tzG emission spectrum were perceived at pH 1 (Fig. 5A), additional species may emit at this pH value. To further characterize these species, we carefully investigated tzG absorption and emission spectra in the pH range –0.6 to 2 (Fig. 7). In line with the data at pH 1–4, the absorption spectra measured in the range pH –0.6 to 1.33 fully overlap, indicating that species A is also the only ground state species in this pH range (Fig. 7A). At pH 1.66 and 2, a small red-shifted shoulder due to minute amounts of the neutral species tzG-H1 is perceived. The normalized emission spectra of tzG indicate that the peak at 465 nm, attributed to tzG-H1* decreases in the pH range 2 to –0.6, revealing two spectra peaking at 394 nm and 519 nm (Fig. 7B). The two peaks, attributed to two excited-state acidic species, named blue-A* and red-A* species on the basis of their emission maximum, could be resolved at pH –0.6, where tzG-H1* emission was absent (Fig. 7C). These two spectra were then used to further deconvolute the tzG emission spectra in the pH range –0.6 – 2 (Fig. S8A–G), showing that the contribution of tzG-H1* peak increases strongly with pH, becoming dominant at pH ≥ 0.66. Interestingly, the ratio of the red-A* to blue-A* emission is constant (4.5 ± 0.4) over the pH range –0.3 to 1.66, indicating that their interconversion does not involve a protonation/deprotonation step. The ϕ value of tzG measured upon excitation at the absorption maximum was found to decrease progressively with pH down to 0.003 (Fig. S8H).
Time-resolved fluorescence measurements of tzG at pH –0.6 were then performed at λexc = 320 nm to record the fluorescence decays of blue-A* and red-A*. The fluorescence decay recorded close to the emission maximum of blue-A* (400 nm) was fitted with a single lifetime value of 0.033 ns (Fig. S9A). The decay recorded at 530 nm, where only red-A* emits, was fitted with a rise component of 0.037 ns and a 0.192 ns decay component (Fig. S9B). These data with a common short lifetime appearing as a rise component for red-A* and a long lifetime only perceived for red-A* are consistent with an excited-state reaction, which irreversibly converts blue-A* into red-A*.58 In this scenario, the long decay component should correspond to the lifetime of red-A*, while the short lifetime should correspond to the sum of the lifetime of blue-A* and the interconversion time between the two species.
To obtain a more detailed picture of tzG photophysics at acidic pH, we resorted to QM calculations. The most stable ground state species at acidic pH is predicted to be tzG-H1–H3+ and not the previously suggested tautomer tzG-H1–H7+ with a protonated N7 atom,23 which is more than 7 kcal mol−1 less stable (Table S1). This conclusion is also supported by the computed absorption energies of the different species (Table S2). tzG-H1–H3+ is predicted to be blue-shifted (4.52 eV) with respect to the neutral tzG-H1 (4.06 eV) in line with the experimental observations, while tzG-H1–H7+ is predicted to be red-shifted (3.66 eV), as it happens for the parent guanosine following N7 protonation. Protonation of N3 leads indeed to an increase of the HOMO/LUMO energy gap of tzG-H1, due to a larger relative stabilization of the HOMO. The pKa values calculated by the M052X (Table S3, pKa = 1.5) or PBE0 (Table S4, pKa = 3.0) functionals for the deprotonation of tzG-H1–H3+ are also consistent with the experimental pKa value (3.5), while in contrast, the pKa values were below – 4 for the deprotonation of tzG-H1–H7+. In addition, the shape of the calculated ESA tzG-H1–H3*+ (Fig. 8, purple line) is similar to the TA spectrum observed immediately after excitation (Fig. 6B, the 200-fs spectrum in blue), both displaying two bands with maxima close to 360 nm and 480 nm. Our computational results thus strongly substantiate our assignment of the A species to tzG-H1–H3+.
In contrast to the ground-state, the most stable protonated excited state species is tzG-H1–H7*+, which is >11 kcal/mol more stable than tzG-H1–H3*+ (Table S1) and characterized by a red-shifted emission (2.81 eV) with respect to that of tzG-H1* (3.11 eV) (Table S5), allowing us to identify the red-A* species as tzG-H1–H7*+. Given that tzG-H1–H3*+ emission is predicted to be blue-shifted relative to tzG-H1* (Table S5), we suggest that blue-A* is the initially excited tzG-H1–H3*+ species, which converts into tzG-H1–H7*+, assigned to red-A*. To determine which of these two species subsequently deprotonates into tzG-H1*, we calculated their excited-state pKa* values. The calculated pKa* values were –8.9 and –0.4 for tzG-H1–H3*+ and tzG-H1–H7*+, respectively (Table S6) indicating that only tzG-H1–H7*+ could deprotonate into tzG-H1* in our experimental pH range. Calculations therefore suggest that the initially excited tzG-H1–H3*+ species is converted into tzG-H1–H7*+, which then deprotonates to tzG-H1*. Of note, the predicted ESA spectra for tzG-H1–H7*+ (Fig. 8A) and tzG-H1* (Fig. 3C) are similar, with the same absorption maximum, and similar oscillator strength, which may explain the challenge in identifying tzG-H1–H7*+ as a putative short-lived species between tzG-H1–H3*+ and tzG-H1* in TA data. For tzG, the intramolecular ESPT reaction is likely mediated by solvent molecules and involves several elementary steps,59 which makes its accurate computational characterization challenging and out of the scope of the present study. tzG is not the case of an ESPT involving two close lying atoms, governed by a single reaction coordinate, as it happens in several other molecules,60,61 for which many theoretical studies exist in the literature.62–66
Finally, TA spectroscopy showed the formation of a long-lived photoproduct absorbing around 350 nm (the ∞DAS in Fig. 6C), via a channel parallel to (and not subsequent to) the formation of tzG-H1* from which emission is observed on the 4.6 ns time scale at pH 2. A likely candidate for the photoproduct could be the triplet electronic state of tzG-H1–H3+. This attribution is consistent with the predicted strong spin–orbit coupling that can efficiently produce a triplet state from tzG-H1–H3*+ (Table S7). Moreover, the computed absorption of the lowest energy triplet electronic state of tzG-H1–H3+ (Fig. 8B) shows a maximum at 310 nm, consistent with the experimental maximum at 350 nm, given the absence of vibrational effects and the errors inherent in our calculations. We therefore suggest that the long-lived photoproduct is the triplet state resulting from an intersystem crossing of the initially excited tzG-H1–H3*+ species.
3.3. tzG in basic solutions
As the second pKa value of tzG is 8.55,23 we characterized the basic form (B) of tzG and explored the possible excited-state deprotonation reactions at pH 12. The absorption spectrum of tzG at pH 12 shows a maximum at 350 nm (Fig. 9A), which is 4 nm red-shifted as compared to tzG-H1, and an extinction coefficient ε350 = 5670 M−1cm−1. The excitation spectra recorded at different emission wavelengths demonstrate a small but significant red-shift at increasing emission wavelengths, indicating inhomogeneity of tzG in the ground state (Fig. 9B). The emission spectra excited at λexc < 390 nm show a main emission band with a maximum at 458 nm, which is 4 nm blue-shifted as compared to tzG-H1 (Fig. 9A). However, for λexc ≥ 390 nm a gradual red-shift in the emission spectra was observed. The same behavior was observed at pH 10 and 11 (data not shown). These spectral shifts suggest that at least two B isomers (B1 and B2) with shifted absorption and emission properties populate the ground state. Since the emission spectrum starts to shift only at the very red edge of the absorption spectrum (Fig. 9A), the blue-shifted B1 species appears to be the dominant one. Due to the marginal contribution of B2, we could only determine the ϕ value of B1, which was found to be 0.05 (Fig. S10).Time-resolved fluorescence decays, triggered by excitation at 370 nm and recorded over the emission spectrum (Fig. S11A), were optimally fitted using a three-component decay function (Fig. S11 B and C). Including a negative component did not improve the χ2 value or distribution of residuals, suggesting the absence of excited-state reaction in the ps-ns time scale of TCSPC. The DAS revealed that tzG emission is largely dominated by the intermediate lifetime (τ2 = 0.94 ns), whose emission nearly overlaps with the steady-state emission and can therefore be attributed to the B1 isomer (Fig. S11D). The DAS of the short-lived lifetime (τ1 = 0.33 ns) and the long-lived lifetime (τ3 = 9.81 ns) are respectively, blue- and red-shifted compared to the DAS of the major species. Since the DAS of τ3 matches reasonably well with the red-shifted emission spectrum recorded at high excitation wavelength (Fig. S11D inset), we attribute the 9.81 ns lifetime to the minor B2 isomer.
The TA data recorded for tzG at pH 12 upon excitation with λpump = 370 nm are displayed in Fig. 10. The multiexponential global fit revealed a broad ESA with λmax = 410 nm, overlapped with a weaker SE (dip around 480 nm) characterized by a 0.93 ns lifetime (green DAS in Fig. 10B), in excellent agreement with the dominant decay component (τ2= 0.94 ns) observed with TSCPC. The 0.93 ns DAS is therefore assigned to the TA signature of the B1* species. The 5.8 ps DAS shape (blue DAS in Fig. 10B) is typical for SE red-shift induced by vibrational or solvent relaxation on this time scale. It should be mentioned that the minor short-lived (200 ps) and long-lived (8.5 ns) components seen in TCSPC are not observed with the less sensitive TA technique.
QM calculations revealed that the most stable species at high pH is tzG−, the deprotonated form of tzG-H1 at its N1 position (Table S1). In line with the experimental data, its predicted emission spectrum is similar to that of tzG-H1 (Table S2), so that it likely corresponds to the major B1 isomer. This assignment is strengthened by the good match of (i) the pKa values for the deprotonation of tzG-H1 to tzG− calculated by M052X (Table S3, pKa = 7.3) or PBE0 (Table S4, pKa = 9.7) functionals with the experimental pKa value (8.55) and (ii) the computed ESA of tzG− (Fig. 11) with the 0.93 ns experimental DAS (Fig. 10B), considering that the contribution of SE is not included in the calculated ESA.
4. Discussion
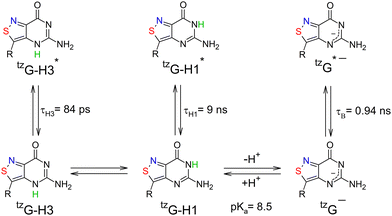
We used a combination of steady-state and time-resolved fluorescence spectroscopy, as well as transient absorption spectroscopy, and QM calculations to decipher the photophysics of tzG, a promising fluorescent analogue of G, as a function of pH. At neutral pH, tzG exists as a ground-state equilibrating mixture of a highly emissive tzG-H1 tautomer (τ = 9 ns) and a weakly emissive tzG-H3 tautomer (τ = 84 ps) (Scheme 1). With the exception of solvent and intramolecular vibrational relaxations, classically observed with polar fluorophores, no excited-state reaction as well as no conversion between tzG-H1 and tzG-H3 were observed in the measured time window (100 fs–100 ns). Based on the strong singlet/triplet spin–orbit coupling (SOC) in tzG-H3,39 we tentatively assign the long-lived species revealed by TA spectroscopy at pH 7 upon 300 nm excitation (∞ DAS in Fig. 3B) to the triplet T1 state of tzG-H3. At pH 12, the tzG-H1 tautomer was found to deprotonate in the ground state to produce tzG− (Scheme 1). This deprotonated form was largely dominant at pH > 10, being characterized by a 4 nm blue-shifted emission maximum and a much shorter fluorescence lifetime (0.94 ns), as compared to tzG-H1. As in neutral pH, with the exception of vibrational or solvent relaxation, no ultrafast excited-state dynamics could be observed.At acidic pH, a much more complex picture was observed with at least two different excited-state reactions. Based on the experimental data and QM calculations, Scheme 2 describes tzG's proposed photophysics at acidic pH. Both the time-resolved data at pH –0.6 (Fig. S9) and the large difference in stability between the two excited-state forms (Table S1) suggest that the back conversion of tzG-H1–H7*+ into tzG-H1–H3*+ is unlikely. In this model, kH1–H3, kH1–H7, kH1 and kH3 are the inverse of the fluorescence lifetimes of tzG-H1–H3*+, tzG-H1–H7*+, tzG-H1* and tzG-H3*, respectively, while k1 describes the conversion from tzG-H1–H3*+ to tzG-H1–H7*+. k+ and k− are the rate constants governing respectively, the protonation of tzG-H1* into tzG-H1–H7*+ and its back reaction in the excited state.
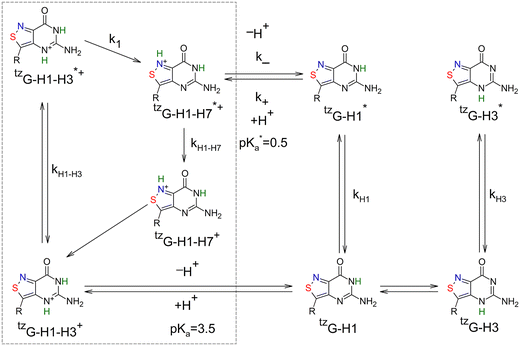 | ||
| Scheme 2 Proposed model to describe tzG photophysics at acidic pH. The dotted box shows the species involved at the most acidic pH (–0.6). | ||
To recover the full set of rate constants, the system of differential equations (eqn (S1)–(S4) in the SI) was solved numerically to globally fit the TCSPC decays recorded at λem = 430 nm and 500 nm in the pH range 0–4 (Fig. 12A). The kH1 and kH3 values were fixed respectively, at 0.1 ns−1 and 11.9 ns−1 from the inverse of the fluorescence lifetimes of tzG-H1* and tzG-H3* measured at pH 7.39 From the fit, we obtained the rate constant values given in Fig. 12C. The k+ value (37.2 L mol−1 ns−1) is close to the value of the corresponding diffusion-controlled rate constant (50 L mol−1 ns−1) previously reported for other photoinduced proton transfer reactions.58 Moreover, the k+ and k− values were found to properly fit the pH dependence of tzG-H1 quantum yield in the range −0.6 to 2 (Fig. S8I) and were used to calculate pKa* = −log(k−/k+) = 0.50, which governs the deprotonation of tzG-H1–H7*+ into tzG-H1* in the excited state. This pKa* value is significantly smaller than the ground-state pKa value (3.5),23 in agreement with the predictions made by QM calculations (Table S6, pKa* = –0.4). In line with our aforementioned expectations for an irreversible conversion of tzG-H1–H3*+ into tzG-H1–H7*+, the fitted kH1–H7 value (5.7 ns−1) is well consistent with the 190 ps lifetime measured for tzG-H1–H7 at pH –0.6 (Fig. S9B), while the sum k1 + kH1–H3 = 38.7 ns−1 is in good agreement with the 33 ps short-lived lifetime measured at pH −0.6 (Fig. S9) and the 49 ps lifetime measured from the TA data of tzG at pH 2 (Fig. 6C).
 | ||
| Fig. 12 Global fit of the TCSPC decay curves at pH 0, 1, 2, 3 and 4 using the system of differential eqn (S1)–(S4) associated to the model defined in Scheme 2. (A) The decays were excited at λexc = 320 nm and recorded at λem = 430 nm and 500 nm. The black solid lines describe the fitted curves to the experimental data (squares). (B) Distribution of residuals and χ2 value. (C) Values of the rate constants used for the global fit of the decays in panel A with eqn (S1)–(S4). (D) Comparison of ϕ values calculated by eqn (S6) with the experimental ϕ values measured by steady-state experiments. The measured values are expressed as means ± standard errors for 3 experiments. | ||
The obtained rate constant values (Fig. 12C) were further cross-checked by using them to recalculate the quantum yields at each pH value (eqn (S6)). The recalculated values were found to be in excellent agreement with the experimental ϕ values measured at an excitation wavelength of 320 nm (Fig. 12D). Altogether, our data give high confidence in the proposed Scheme 2 and highlight the importance of the proton transfer reactions in governing the photophysics of tzG at acidic pHs.
From TA data, a long-lived photoproduct was inferred and putatively attributed to tzG-H1–H3+ triplet state. The spin–orbit coupling of tzG-H1–H3+ triplet state is of the same magnitude as that for tzG-H3 (Table S7), for which ISC was predicted to be a major non-radiative pathway, explaining its low quantum yield39 and probably the ∞DAS observed at pH 7 upon excitation at 300 nm (Fig. 3B). Of note, strong spin–orbit coupling was also computed for tzG-H1–H7*+, but with a triplet state high in energy, making ISC from this species less likely.
Altogether, our results demonstrate tzG sensitive response to pH changes in a wide pH range. Because of the shift between the pKa and pKa* values, and the differences in the spectra and quantum yields of the different species involved at acid pH, monitoring the quantum yield and absorption maximum of tzG allows the pH value to be determined over a very wide pH range (0–5) (Fig. 13). Moreover, the spectral shift and difference in brightness between the neutral tzG-H1 and the basic tzG− also allow sensitive measurements of pH values in a more limited basic pH range (8–10).
 | ||
| Fig. 13 pH dependence of tzG quantum yield (blue squares) and absorption maximum (red squares). The solid lines are the fits of the data to eqn (2). The pH dependence of λabs was fitted with the ground-state pKa values given in the text. For the pH dependence of ϕ, the pKa* (8.7) in the basic pH range was found to be similar to the pKa (8.55) in the ground state, in line with the absence of ESPT reaction at basic pH. In the acidic region, we obtained an apparent pKa* value of 2.1, which differs from the pKa* value (0.5) calculated from the k+/k− ratio, due to the direct excitation of tzG-H1 at pH ≥ 2, which takes over from the conversion of tzG-H1–H7* to tzG-H1*. It should be noted that the consistency of the ϕ values in the pH range 5–8 and in the pH range 10–12, as well as the monotonic decrease of ϕ values in the pH range 8–10 and the fits of the ϕ values at acidic pH (Fig. 12D), strongly suggest that the ϕ values are independent of the nature of the buffer. | ||
5. Conclusion
In this work, the photophysical study of tzG as a function of pH revealed that at acidic pH the protonated acidic tzG-H1–H3+ species, which is the only species observed in the ground state at pH≤1, rapidly converts to the more stable species tzG-H1–H7*+ in the excited state. The tzG-H1–H3*+ and tzG-H1–H7*+ species have well separated emission bands but low quantum yields (ϕ ∼ 0.002–0.006) and the ratio of their emission bands is kinetically driven. Subsequently, a second excited-state reaction can occur, where tzG-H1–H7*+ deprotonates into the bright tzG-H1* species (ϕ = 0.36, τ = 9.0 ns) with a pKa* value of 0.50, shifted by nearly three pH units as compared to the ground-state pKa. These unique properties make tzG a suitable candidate for use as a sensitive pH sensor across a wide range of acidic pH values (0–5).Comparison with thG revealed several similarities but also significant differences. At acidic pH, the stable ground-state species of tzG and thG are similarly protonated at both N1 and N3 atoms. However, due to the absence of N7 atom, thG-H1–H3*+ converts into thG-H1*31 through a different intermediate thG-H1-OH*+.31 The latter, being characterized by a lifetime of 800 ps, appears significantly brighter than the tzG-H1–H7*+ intermediate (190 ps), but their pKa* values are comparable (1.0 vs. 0.5). The very low quantum yield of tzG-H3 as compared to thG-H3 is a distinctive advantage of tzG in the analysis of the photophysical properties.39 Indeed, as for both nucleosides, the spectra of the H3 tautomer and the weakly emissive acidic species largely overlap, the emission properties of the acidic species can only be characterized for tzG. Consequently, while we had to assume the values of several parameters in the photophysical model of thG, we have been able to determine all of them for tzG. Finally, at basic pHs, deprotonation of thG-H1 into thG− does not lead to any change in quantum yield and lifetime,31 which prevented thG from being used as a pH sensor at basic pH, unlike tzG. Interestingly, the shape of the dependence of tzG ϕ as a function of pH (Fig. 13) is analogous to that of the 2-aminopurine ribonucleoside.11 However, the latter showed a considerably steeper fluorescence change at acidic pH, allowing it to explore a much narrower pH range (pH 3–5) than tzG. Moreover, the high pKa value (12.0) of 2-aminopurine at basic pH limits its use as a pH sensor in the basic pH range.
Together with our previous study on the spectroscopic properties of tzG at neutral pH,39 the current study provides a comprehensive and in depth study of the photophysical properties of tzG in its free ribonucleoside form. This study constitutes a solid basis for discussing the photophysics of tzG included in oligonucleotides, as well as the spectroscopic changes of tzG-labelled oligonucleotides resulting from conformational changes and interaction with proteins. For a large number of applications with tzG-labelled oligonucleotides, the insensitivity of tzG fluorescence in the pH range 5–8 (Fig. 13) will be an asset, since any change in fluorescence could be directly interpreted in terms of conformational changes or interactions, without a bias due to a pH effect. By itself, the sensitivity of the free nucleoside to pH values below 5 is of limited interest in biological systems, where such low pH values are exceptional. In contrast, protonation of tzG and its accompanying fluorescence changes might be profitably used for specific cases of protein interactions with tzG-labelled oligonucleotides. Indeed, variations in local electrostatic environment caused by the presence of ionizable groups (such as amino, carboxyl, and phosphate groups) in combination with exclusion of water molecules from the interior of proteins and from nucleic acid/protein binding interfaces may significantly shift the pKa values of amino acid side chains and nucleobase residues.67–69 In the case of tzG-labelled oligonucleotides, the positively charged acidic form tzG-H1–H3+ is expected to be stabilized by the negatively charged phosphates, leading to an increase of its pKa to a value closer to physiological pH values. Therefore, conformational changes or interaction with specific protein domains, such as nucleotide binding pockets, could potentially impact this pKa value. This will in turn lead to a change in tzG protonation and fluorescence properties, therefore providing an additional channel of information on the system. At the same time, the negatively charged basic form thG− will likely be disfavored in nucleic acids for the same reasons, implying that tzG deprotonation will occur at even higher pH, extending above pH 8 the range in which tzG is pH insensitive. In addition, given that proton transfer reactions have been extensively reported for G residues in nucleic acids, where they play a major role in the photophysical properties of the natural nucleosides and in the promotion of mutations in DNA sequences,70–73 the current study should also be instrumental to monitor and interpret excited-state proton transfer reactions in tzG-labelled sequences.
Author contributions
O. T.: writing – original draft, methodology, investigation, formal analysis, data curation. L. M. F.: visualization, software, methodology, investigation, formal analysis, data curation. A. E. R-L.: investigation, formal analysis, N. H.: supervision, methodology, investigation. L. R.: methodology, investigation, formal analysis. D. D.; E. B.: writing – review & editing, methodology, investigation. A. B. and F. H.: methodology, investigation, P. D.: software, methodology, formal analysis, Y. T.: writing – review & editing, resources, methodology, conceptualization. R. I., J. L. and Y. M.: writing – review & editing, writing – original draft, validation, supervision, funding acquisition, conceptualization.Conflicts of interest
The authors declare no conflict of interest.Abbreviations
| DAS | Decay-associated spectra |
| ESA | Excited state absorption |
| ESPT | Excited-state proton transfer |
| FNA | Fluorescent nucleoside analogue |
| FWHM | Full width at half maximum |
| QS | Quinine sulfate |
| ϕ | Fluorescence quantum yield |
| WL | White light |
| GVD | Group velocity dispersion |
| GSB | Ground state bleach |
| IRF | Instrumental response function |
| ISC | Intersystem crossing |
| PCM | Polarizable continuum model |
| QM | Quantum mechanical calculations |
| TA | Transient absorption spectroscopy |
| TCSPC | Time-correlated single photon counting |
| TD-DFT | Time-dependent extension of Density Functional Theory |
| thG | Thienoguanosine |
| tzG | Isothiazologuanosine |
| SE | Stimulated emission |
| SOC | Spin–orbit coupling. |
Data availability
The data supporting this article have been included as part of the SI. SI contains the quality of TA data fits, TCSPC data, spectral deconvolution, and the proposed model that describes tzG photophysics at acidic pH, as well as the results of QM calculations of species stability, absorption and emission energies, pKa values, SOC, and energy barriers to T states. See DOI: https://doi.org/10.1039/d5cp02642cAcknowledgements
This work and OT were supported by the Agence Nationale de la Recherche (ANR GQFluodynint), the Fondation Jean-Marie Lehn, the Région Grand-Est (EpiRNA project) and the Centre National de la Recherche Scientifique (CNRS). YM is grateful to the Institut Universitaire de France (IUF) for support and providing additional time to be dedicated to research. RI thanks financial support from CN3, National Center for Gene Therapy and Drugs based on RNA technology, funded by the European Union-NextGenerationEU-PNRR. RI also thanks the CNR program “Progetti di Ricerca @cnr”, project UCATG4, MUR ‘PRIN 2022 2022K3AY2K’ and Nutrage FOE 2022 for financial support. LM-F acknowledges the grant PID2023-151719NA-I00 funded by MICIU/AEI/10.13039/501100011033 and FEDER, UE. This research project was made possible through the access granted by the Galician Supercomputing Center (CESGA) to its supercomputing infrastructure. YT thanks the National Institutes of Health for generous support (via grant number R35 GM139407). JL acknowledges support from the Interdisciplinary Thematic Institute QMat as part of the ITI 2021-2028 program of the University of Strasbourg, CNRS and Inserm via the IdEx Unistra (ANR 10 IDEX 0002), SFRI STRAT’US (ANR 20 SFRI 0012), and Labex NIE (ANR-11-LABX-0058-NIE) projects of the French Investments for the Future Program.Notes and references
- C. E. Crespo-Hernández, B. Cohen, P. M. Hare and B. Kohler, Chem. Rev., 2004, 104, 1977–2020 CrossRef PubMed.
- C. T. Middleton, K. de La Harpe, C. Su, Y. K. Law, C. E. Crespo-Hernández and B. Kohler, Annu. Rev. Phys. Chem., 2009, 60, 217–239 CrossRef CAS PubMed.
- K. Kleinermanns, D. Nachtigallová and M. S. de Vries, Int. Rev. Phys. Chem., 2013, 32, 308–342 Search PubMed.
- M. Barbatti, A. C. Borin and S. Ullrich, Photoinduced Phenomena in Nucleic Acids I: Nucleobases in the Gas Phase and in Solvents, Springer International Publishing, 2015, pp. 1–32 Search PubMed.
- R. Improta, F. Santoro and L. Blancafort, Chem. Rev., 2016, 116, 3540–3593 CrossRef CAS PubMed.
- T. Gustavsson and D. Markovitsi, Acc. Chem. Res., 2021, 54, 1226–1235 CrossRef CAS PubMed.
- L. Martínez Fernández, F. Santoro and R. Improta, Acc. Chem. Res., 2022, 55, 2077–2087 CrossRef PubMed.
- R. W. Sinkeldam, N. J. Greco and Y. Tor, Chem. Rev., 2010, 110, 2579–2619 CrossRef CAS PubMed.
- W. Xu, K. M. Chan and E. T. Kool, Nat. Chem., 2017, 9, 1043–1055 CrossRef CAS PubMed.
- D. Dziuba, P. Didier, S. Ciaco, A. Barth, C. A. M. Seidel and Y. Mély, Chem. Soc. Rev., 2021, 50, 7062–7107 RSC.
- D. C. Ward, E. Reich and L. Stryer, J. Biol. Chem., 1969, 244, 1228–1237 CrossRef CAS PubMed.
- A. C. Jones and R. K. Neely, Q. Rev. Biophys., 2015, 48, 244–279 CrossRef CAS PubMed.
- L. M. Wilhelmsson, A. Holmén, P. Lincoln, P. E. Nielsen and B. Nordén, J. Am. Chem. Soc., 2001, 123, 2434–2435 CrossRef CAS PubMed.
- P. Sandin, K. Börjesson, H. Li, J. Mårtensson, T. Brown, L. M. Wilhelmsson and B. Albinsson, Nucleic Acids Res., 2008, 36, 157–167 CrossRef CAS PubMed.
- K. Börjesson, S. Preus, A. H. El-Sagheer, T. Brown, B. Albinsson and L. M. Wilhelmsson, J. Am. Chem. Soc., 2009, 131, 4288–4293 CrossRef PubMed.
- M. B. Turner, J. M. Cizmic, D. B. Rosansky, J. Ceja, M. Patterson, S. Kilcoyne, K. Thurber, G. Kim, T. J. Dwyer and B. W. Purse, Bioconjugate Chem., 2023, 34, 1061–1071 CrossRef CAS PubMed.
- D. D. Burns, K. L. Teppang, R. W. Lee, M. E. Lokensgard and B. W. Purse, J. Am. Chem. Soc., 2017, 139, 1372–1375 CrossRef CAS PubMed.
- T. L. Coulson and J. R. Widom, Nucleic Acids Res., 2025, 53, gkaf262 CrossRef PubMed.
- Q. L. Nguyen, V. A. Spata and S. Matsika, Phys. Chem. Chem. Phys., 2016, 18, 20189–20198 RSC.
- K. C. Thompson and N. Miyake, J. Phys. Chem. B, 2005, 109, 6012–6019 CrossRef CAS PubMed.
- A. Dallmann, L. Dehmel, T. Peters, C. Mügge, C. Griesinger, J. Tuma and N. P. Ernsting, Angew. Chem., Int. Ed., 2010, 49, 5989–5992 CrossRef CAS PubMed.
- D. Shin, R. W. Sinkeldam and Y. Tor, J. Am. Chem. Soc., 2011, 133, 14912–14915 CrossRef CAS PubMed.
- A. R. Rovira, A. Fin and Y. Tor, J. Am. Chem. Soc., 2015, 137, 14602–14605 CrossRef CAS PubMed.
- S. Park, H. Otomo, L. Zheng and H. Sugiyama, Chem. Commun., 2014, 50, 1573–1575 RSC.
- M. Sholokh, R. Sharma, D. Shin, R. Das, O. A. Zaporozhets, Y. Tor and Y. Mely, J. Am. Chem. Soc., 2015, 137, 3185–3188 CrossRef CAS PubMed.
- Y. Tor, Acc. Chem. Res., 2024, 57, 1325–1335 CrossRef CAS PubMed.
- M. Sholokh, R. Improta, M. Mori, R. Sharma, C. Kenfack, D. Shin, K. Voltz, R. H. Stote, O. A. Zaporozhets, M. Botta, Y. Tor and Y. Mely, Angew. Chem., Int. Ed., 2016, 55, 7974–7978 CrossRef CAS PubMed.
- J. Kuchlyan, L. Martinez-Fernandez, M. Mori, K. Gavvala, S. Ciaco, C. Boudier, L. Richert, P. Didier, Y. Tor, R. Improta and Y. Mély, J. Am. Chem. Soc., 2020, 142, 16999–17014 CrossRef CAS PubMed.
- S. Ciaco, K. Gavvala, V. Greiner, V. Mazzoleni, P. Didier, M. Ruff, L. Martinez-Fernandez, R. Improta and Y. Mély, Methods Appl. Fluoresc., 2022, 10, 035003 CrossRef CAS PubMed.
- L. Martinez-Fernandez, K. Gavvala, R. Sharma, P. Didier, L. Richert, J. Segarra Martì, M. Mori, Y. Mely and R. Improta, Chem. – Eur. J., 2019, 25, 7375–7386 CrossRef CAS PubMed.
- P. Didier, J. Kuchlyan, L. Martinez-Fernandez, P. Gosset, J. Léonard, Y. Tor, R. Improta and Y. Mély, Phys. Chem. Chem. Phys., 2020, 22, 7381–7391 RSC.
- M. Sholokh, R. Sharma, N. Grytsyk, L. Zaghzi, V. Y. Postupalenko, D. Dziuba, N. P. Barthes, B. Y. Michel, C. Boudier, O. A. Zaporozhets, Y. Tor, A. Burger and Y. Mely, Chem. – Eur. J., 2018, 24, 13850–13861 CrossRef CAS PubMed.
- N. Grytsyk, L. Richert, P. Didier, D. Dziuba, S. Ciaco, V. Mazzoleni, T. Lequeu, M. Mori, Y. Tor, L. Martinez-Fernandez, R. Improta and Y. Mély, Int. J. Biol. Macromol., 2022, 213, 210–225 CrossRef CAS PubMed.
- J. A. Brown, Wiley Interdiscip. Rev.: RNA, 2020, 11, e1598 CrossRef CAS PubMed.
- J. Jana, S. Mohr, Y. M. Vianney and K. Weisz, RSC Chem. Biol., 2021, 2, 338–353 RSC.
- M. S. Bucardo, Y. Wu, P. T. Ludford, Y. Li, A. Fin and Y. Tor, ACS Chem. Biol., 2021, 16, 1208–1214 CrossRef CAS PubMed.
- Y. Li, P. T. Ludford III, A. Fin, A. R. Rovira and Y. Tor, Chem. – Eur. J., 2020, 26, 6076–6084 CrossRef CAS PubMed.
- D. Cong, Y. Li, P. T. Ludford III and Y. Tor, Chem. – Eur. J., 2022, 28, e202200994 CrossRef CAS PubMed.
- O. Tkach, L. Martinez-Fernandez, N. Humbert, L. Richert, D. Dziuba, P. Didier, Y. Tor, R. Improta and Y. Mély, J. Photochem. Photobiol., A, 2024, 446, 115075 CrossRef CAS.
- D. F. Eaton, Pure Appl. Chem., 1988, 60, 1107–1114 CrossRef CAS.
- S. Preus, a|e - UV-Vis-IR Spectral Software 1.2, https://www.fluortools.com, 2013 Search PubMed.
- B. Valeur and M. N. Berberan-Santos, Molecular fluorescence: principles and applications, John Wiley & Sons, 2012, pp. 53–74 Search PubMed.
- P.-A. Muller, Fluorescence Quantum Yields (QY) and Lifetimes (τ) for Alexa Fluor Dyes—Table 1.5, https://www.thermofisher.com/fr/fr/home/references/molecular-probes-the-handbook/tables/fluorescence-quantum-yields-and-lifetimes-for-alexa-fluor-dyes.html Search PubMed.
- D. A. Barrow and B. R. Lentz, Chem. Phys. Lett., 1984, 104, 163–167 CrossRef CAS.
- S. Preus, DecayFit 1.3, https://www.fluortools.com, 2013 Search PubMed.
- D. A. Smith, G. McKenzie, A. C. Jones and T. A. Smith, Methods Appl. Fluoresc., 2017, 5, 042001 CrossRef PubMed.
- J. R. Lakowicz, Principles of fluorescence spectroscopy, Springer, Boston, 2006, 581 Search PubMed.
- G. H. Golub and V. Pereyra, SIAM J. Numer. Anal., 1973, 10, 413–432 CrossRef.
- D. P. O’Leary and B. W. Rust, Comput. Optim. Appl., 2013, 54, 579–593 CrossRef.
- Y. Zhao and D. G. Truhlar, Acc. Chem. Res., 2008, 41, 157–167 CrossRef CAS PubMed.
- Y. Zhao, N. E. Schultz and D. G. Truhlar, J. Chem. Theory Comput., 2006, 2, 364–382 CrossRef PubMed.
- C. Zuluaga, V. A. Spata and S. Matsika, J. Chem. Theory Comput., 2021, 17, 376–387 CrossRef CAS PubMed.
- S. Miertuš, E. Scrocco and J. Tomasi, Chem. Phys., 1981, 55, 117–129 CrossRef.
- J. Tomasi, B. Mennucci and R. Cammi, Chem. Rev., 2005, 105, 2999–3094 CrossRef CAS PubMed.
- B. Thapa and H. B. Schlegel, J. Phys. Chem. A, 2015, 119, 5134–5144 CrossRef CAS PubMed.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, et al., Gaussian 16 Revision A.03, 2016 Search PubMed.
- B. Valeur and M. N. Berberan-Santos, Molecular fluorescence: principles and applications, John Wiley & Sons, 2012, pp. 141–179 Search PubMed.
- N. Agmon, J. Phys. Chem. A, 2005, 109, 13–35 CrossRef CAS PubMed.
- T. Kumpulainen, B. Lang, A. Rosspeintner and E. Vauthey, Chem. Rev., 2017, 117, 10826–10939 CrossRef CAS PubMed.
- A. C. Sedgwick, L. Wu, H.-H. Han, S. D. Bull, X.-P. He, T. D. James, J. L. Sessler, B. Z. Tang, H. Tian and J. Yoon, Chem. Soc. Rev., 2018, 47, 8842–8880 RSC.
- O. Louant, B. Champagne and V. Liégeois, J. Phys. Chem. A, 2018, 122, 972–984 CrossRef CAS PubMed.
- J. Jankowska and A. L. Sobolewski, Molecules, 2021, 26, 5140 CrossRef CAS PubMed.
- L. F. B. Fontes, J. Rocha, A. M. S. Silva and S. Guieu, Chem. – Eur. J., 2023, 29, e202301540 CrossRef CAS PubMed.
- T. Qiao, W. Shi, H. Zhuang, G. Zhao, X. Xin and Y. Li, Spectrochim. Acta, Part A, 2024, 309, 123802 CrossRef CAS PubMed.
- H. Zhuang, J. Wang, W. Shi, Y. Mei and Y. Li, Spectrochim. Acta, Part A, 2025, 333, 125896 CrossRef CAS PubMed.
- D. Bashford and M. Karplus, Biochemistry, 1990, 29, 10219–10225 CrossRef CAS PubMed.
- C. L. Tang, E. Alexov, A. M. Pyle and B. Honig, J. Mol. Biol., 2007, 366, 1475–1496 CrossRef CAS PubMed.
- G. M. Ullmann and E. Bombarda, Biol. Chem., 2013, 394, 611–619 CrossRef CAS PubMed.
- R. Srivastava, Front. Chem., 2019, 7, 536 CrossRef CAS PubMed.
- L. Slocombe, M. Winokan, J. Al-Khalili and M. Sacchi, Commun. Chem., 2022, 5, 144 CrossRef CAS PubMed.
- G. Villani, J. Chem. Phys., 2008, 128, 114306 CrossRef PubMed.
- Y. Lee, R. Roy, S. Gu, S. B. Shetty, A. K. Rangadurai and H. M. Al-Hashimi, J. Am. Chem. Soc., 2025, 147, 19643–19666 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2025 |