Importance of chirality in the self-organizing peptides – from single molecules to functional supramolecular structures
Agata
Chotera-Ouda
 ab,
Katarzyna
Trzeciak
ab,
Katarzyna
Trzeciak
 a and
Marek J.
Potrzebowski
a and
Marek J.
Potrzebowski
 *a
*a
aCentre of Molecular and Macromolecular Studies, Polish Academy of Sciences, Sienkiewicza 112, 90-363 Lodz, Poland. E-mail: marek.potrzebowski@cbmm.lodz.pl
bInternational Centre for Research on Innovative Biobased Materials (ICRI-BioM)-International Research Agenda, Lodz University of Technology, Zeromskiego 116, 90–924 Lodz, Poland
First published on 14th August 2025
Abstract
Chirality plays a key role in the self-assembly of peptides, influencing their structural and functional properties from the molecular to the supramolecular level. The stereochemistry of peptides determines their assembly pathways, leading to the formation of well-defined nanostructures such as tubes, wires, helices, fibers, sheets, and gels. This chiral self-assembly not only determines the mechanical and optical properties of the resulting materials but also influences their biological interactions and functionality. Understanding and controlling chirality in peptide systems is essential for the design of advanced biomaterials with tailored properties for applications in nanotechnology, medicine, and tissue engineering. This tutorial review article presents experimental techniques that are dedicated to the study of chiral compounds, allow to determine optical purity, and determine the absolute configuration of the studied species. Techniques for studying higher-order structures of self-assembled peptide systems are also discussed. The role of non-covalent intermolecular contacts in the formation of crystal structures made of heterochiral peptides is presented. The structure and morphology of peptide nanotubes (PNTs), peptide nanowires (PNWs), and other functional peptide devices that arise from the self-assembly and multiplication of chiral structural synthons (CSS), the smallest building blocks, are discussed. Finally, examples of systems belonging to the category of soft materials (peptide hydrogels, organogels) are presented.
1. Introduction
In the history of the world and in the development of civilization, there are phenomena that have always, from the very beginning, controlled fundamental processes, but the knowledge of their existence and significance appeared relatively late.1–3 One of such phenomena is “chirality”, which defines the key properties of amino acids and nucleic acids, the most important building blocks of living organisms.4 Chirality, derived from the Greek word “cheir”, meaning hand, is the property of asymmetry in molecules. Chiral molecules are non-superimposing mirror images of each other, just like our left and right hands. This property is crucial in various biological, chemical, and physical processes and has significant implications in nature.5 Although many decades have passed since Pasteur's breakthrough discoveries of chirality in the nineteenth century, this scientific platform is still an area of active research.6–8 Chiral (and non-chiral) building blocks formed and can form higher structures using different types of interactions, which in the first approach can be divided into two groups, covalent bonds and non-covalent interactions.9 The latter processes are a complex, multi-element phenomenon that is the basis for the development of a very broad strategy called “molecular self-organization”.10,11 In this approach, small molecular building blocks spontaneously interact to form hierarchical, functional chiral supramolecular materials. Common key interactions involved in the process include hydrogen bonds, π–π stacking, Coulomb, and van der Waals interactions; thus, in contrast with an approach based on covalent bonding, supramolecular assemblies offer greater responsiveness and tunability.12 A crucial role in self-organization is played by the hydrophobic effect, which minimizes interactions between nonpolar side chains and water, promoting the aggregation of peptides into ordered structures. The amino acids and short peptides made up of L and d units play a special role among chiral compounds.13,14 Thanks to their ability to adopt various secondary structures, biocompatibility, and relatively easy design and synthesis, short peptides provide versatile self-organizing soft-matter15 and have been widely studied in recent years for their implementation in the development of functional smart materials, for example by incorporating synthetic and biological16–18 or (opto-) electronic moieties,19,20 providing an effective interface with natural and artificial systems. Moreover, the fundamental knowledge and control over peptide self-assembly can offer crucial information about the formation principles of more complex protein aggregation.Despite the great progress achieved in recent years in the understanding of the molecular basis of homo- and hetero-chiral peptide organization processes, the rational design and regulation of their definite structure and function remain challenging. In the case of many sequences, the introduction of D-amino acids destroys their original organization, while for some others, the heterochirality has positive effects on the self-assembly and gelation. Hence, increasing efforts have been focused on stereochemical factors that can influence the properties of the final assemblies, and broaden the repertoire of inherent structural determinants, allowing the formation of more diverse, although controllable, architectures.21
The aim of this review is to summarize recent findings in the field and highlight the importance of chirality in the self-organization of short peptides. Special focus is given to its effects on the assemblies across different size scales – all the way from atomic to macroscopic characterization. In the following sections, we discuss how the chirality of amino acid residues influences crystal structure and molecular packing, leads to the formation of various self-assembling morphologies, and finally results in materials of different properties, with a particular focus on the supramolecular gels. Upon conclusion, a few crucial remaining questions and future perspectives are also delivered.
2. Chirality, basic definitions, and methods for distinguishing stereoisomers
Chirality is a fundamental geometric property of molecules and objects that cannot be superimposed on their mirror images, much like left and right hands. It is pervasive in nature, manifesting across scales from molecular structures like amino acids to macroscopic forms and even galaxies.22 In chemistry, chiral molecules exist as two non-superimposable mirror-image forms called enantiomers, often categorized as “right-handed” or “left-handed” (Fig. 1. The source of molecular chirality often lies in stereogenic elements, commonly a carbon atom bonded to four distinct groups in a tetrahedral geometry (Fig. 1B). | ||
| Fig. 1 Stereoisomers of amino acids. (A) Two enantiomers of alanine (L-Ala and D-Ala) that are the mirror reflection of each other. (B) The general structural formula for an α-amino acid. | ||
Specifically, in the case of amino acids, they exist in nature as either L- or D- enantiomers (except of a non-chiral glycine). The two forms are characterized by the different orientations of the four substituents at the α-carbon. In the convention of CIP (Cahn-Ingold-Prelog), the absolute configuration of L amino acids is defined as S, while for D amino acids absolute configuration is R. The presence of L-amino acids is clearly dominant in the living organisms, resulting in homochirality of the biological world. Taking into account this prevalent use of homochirality in nature, and its preference towards single-handedness of biopolymers, the chirality seems to be critical for a range of complex processes. Nonetheless, the utilization of non-proteogenic D-amino acids in biological molecules, even though rare, seems to play an important role, since naturally existing peptides that contain them include various bioactive peptides such as peptidoglycans, opioid peptides, antibiotics, and cytotoxins, hormones, and others. The incorporation of D-amino acids in peptides for biological applications has been predominantly investigated for increased enzymatic stability and antimicrobial effects.23 The abovementioned discoveries can be perceived as a motivation for employing D-amino acids in the design of building blocks for functional peptide-based materials, and for comparative analysis of homo- and hetero-chiral self-assembling peptide sequences, as well as their mixtures.
Currently, there are several experimental techniques that allow to determine the absolute configuration of amino acids and their enantiomeric purity (see Fig. 2, left panel). The right panel shows the techniques used to analyze higher-order peptide structures that are formed by short- and long-range intermolecular interactions (bottom panel). These approaches are briefly described below.
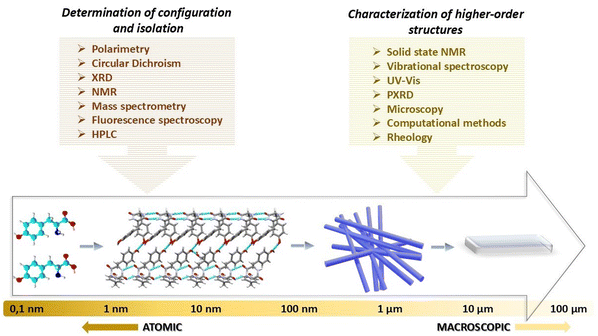 | ||
| Fig. 2 Schematic representation of the self-assembly of chiral molecules across scales and a list of techniques used for its characterization. | ||
2.1. Polarimetry
Polarimetry is a technique that measures the degree of rotation of plane-polarized light caused by chiral molecules. It is well known that the pair of enantiomers has identical physical properties except for how they interact with plane-polarized light. In normal monochromatic light, the electric field oscillates in all directions. When light hits a polarizer, only that part of the light that oscillates in the same plane in which the polarizing filter is positioned, can pass through. This plane is called the plane of polarization, and the light is called plane-polarized light.24 If the substance is optically inactive, the plane of polarized light will not change in orientation. Chiral molecules can rotate the plane of polarized light, and this property is called optical activity. Optical rotation is the degree to which the sample rotates the polarized light. Depending on the direction of rotation of the light, the enantiomer is defined as dextrorotatory (Latin: right, clockwise) and is labeled with the prefix (D) or (+) or levorotatory (Latin: left, counterclockwise) and is designated as (L) or (−).25 To compare the optical rotation of different compounds under constant conditions, specific rotation is used, a fundamental property of chiral substances expressed as the angle at which the material causes polarized light to rotate at a given temperature, wavelength and concentration.26–28 Polarimetry determines the ratio of two enantiomers and their purity and concentration. The measurement of the enantiomeric excess (EE), and thus the determination of optical purity, is crucial for pharmaceutical applications because different enantiomers may exhibit divergent physiological effects. Optical purity (expressed as a percentage) is a comparison of the optical rotation of a pure sample of unknown stereochemistry with the optical rotation of a sample of the pure enantiomer. The enantiomeric excess value can range from 0% to 100%. The absolute value of EE is 0% for a racemic mixture and 100% for an optically pure sample.26,29Polarimetry is generally used by pharmacopeias to determine the specific rotation in order to characterize optically pure L-amino acids.14 The more sensitive type – laser-based polarimetry has been applied in specific rotation measurements of not only amino acids but also di- and tri-peptides.30
2.2. Circular dichroism (CD) and vibrational circular dichroism (VCD)
Circular dichroism (CD) is a spectroscopic technique based on differential absorption of left-handed circularly polarized (L-CP) light and right-handed circularly polarized (R-CP) light. In the presence of chiral chromophores, one state of circularly polarized light will be absorbed to a greater extent than the other. CD spectroscopy is an essential analytical technique used to analyze chirality in molecules through their optical activity. Synchrotron radiation circular dichroism (SRCD) spectroscopy offers significant improvements to the well-established method of conventional circular dichroism (cCD) spectroscopy. SRCD takes advantage of the high photon flux available from the light source over a wide range of wavelengths, which results in higher signal-to-noise ratios and also enables the collection of data at lower wavelengths than is possible with cCD spectrometers. Fig. 3 shows the SRCD profiles for L and D alanine.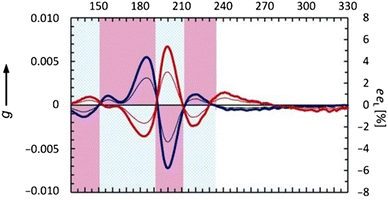 | ||
| Fig. 3 Anisotropy spectra (thick lines) of isotropic amorphous D-Ala (red) and L-Ala (blue) in the vacuum-UV and UV spectral region. Thin lines represent the corresponding eeL plots inducible by either left or right circularly polarized light at ξ = 0.9999. Amino acids were sublimated and condensed in the form of isotropic amorphous films. Reproduced and adapted from ref. 31 with permission from John Wiley and Sons, copyright 2012. | ||
The obtained CD spectrum does not constitute an intrinsic property of the molecule, but rather depends on the molecular conformation, meaning it is strongly related to the external environmental factors, such as temperature, solvent, etc.
Vibrational circular dichroism (VCD) is the extension of circular dichroism spectroscopy into the infrared and near-infrared ranges. The method is mostly applied for the absolute configuration of chiral organic molecules32 but also provides a possibility to determine the secondary structure of proteins and peptides, complemented by observation of conformational changes. For instance, it was proven to be a useful method for the structural analysis of naturally occurring D-amino acid-containing peptides.33 Importantly, the VCD spectrum of the chiral molecule can be calculated using an ab initio density functional theory (DFT) method.34 The method is currently gaining popularity due to the low amount of sample required for a measurement and its complementarity with X-ray and NMR methods.
2.3. X-ray diffraction (XRD)
X-ray crystallography is the major and routine analytical technique for determining the three-dimensional arrangement of atoms in a crystal, revealing the absolute configuration of chiral molecules.35–39 If the studied single crystal is of good quality and the diffraction data is unambiguous, the resulting crystallographic results provide accurate and reliable structural parameters that allow researchers to characterize and understand the behavior and function of chiral compounds. The absolute configuration of an optically active molecule, which possesses one or more stereogenic centers, becomes determinable through the effect of X-rays’ anomalous scattering (or anomalous dispersion).35,40 A widely utilized measure in determining absolute configuration is the Flack parameter.41 In the computation of this parameter, a crystal is treated as if it were twinned by inversion, and the occupancies of the two domains are calculated. This parameter has a physical meaning in the range from 0 to 1. However, experimental outcomes might occasionally deviate from this range due to statistical variations and systematic errors. The Flack parameter assumes a zero value when analyzing an enantiomerically pure crystal in the correct absolute configuration. However, if the crystal structure is inverted, the Flack parameter equals 1.38,42 Generally, determining the absolute configuration requires a heavy atom in the structure. However, improvements in methods may allow the absolute configuration to be determined from the “light atom” structures (i.e. C, O, and N).392.4. Nuclear magnetic resonance (NMR)
NMR spectroscopy is an effective technique in structural studies and chirality recognition of various chemical compounds,43–47 which uses the magnetic properties of nuclei.48 In the case of enantiomers, samples with a single stereogenic center, the active nuclei of NMR are isochronous in an optically inactive environment, so it is not possible to distinguish between them. However, these nuclei are anisochronous in a chiral medium, resulting in different NMR spectra for both enantiomers. There are two general strategies for determining the absolute configuration of compounds. These approaches involve forming a diastereomeric pair of chiral molecules with chiral discriminating agents,49–51 such as chiral derivatizing agents (CDA)52–54 and chiral solvating agents (CSA),55 to obtain two different chemical species with no symmetric relationship. CDAs form covalent bonds with the functional group of the substrate, while CSAs create non-covalent associations, such as hydrogen bonds, ion pairing, dipole–dipole or van der Waals interactions. Among many different chiral discriminating agents, Pirkle's alcohol56 and Mosher's acid57,58 are often applied to analyse chiral compounds. The most common NMR technique used to determine the stereochemistry of chiral centres in molecules is 1H NMR.46,59–63 This is undoubtedly due to the properties of 1H nuclei, such as natural abundance and great sensitivity to environmental modifications.51 Nevertheless, 1H NMR spectroscopy has certain limitations, and sometimes, analysis of 1H NMR spectra, even small compounds, is complicated. Therefore, 13C NMR is an interesting alternative or good combination with 1H NMR spectroscopy.64–67 Other nuclei used to determine the absolute configuration of organic compounds are 19F NMR, 31P NMR, and 77Se NMR.68–74 While applying multinuclear NMR spectroscopy for chiral discrimination and absolute configuration assignment holds promise, it is crucial to assess the reliability of such analyses for specific applications. This evaluation is imperative owing to each element's different nuclear properties and general behaviors.2.5. Ion mobility mass spectrometry
Ion mobility mass spectrometry (IM-MS) is a powerful analytical technique used to separate and characterize ions based on their size, shape, and charge. When applied to D- and L-amino acids, IM-MS provides valuable insights into their structural properties and stereochemical differences. Mass spectrometry (MS) is a great tool to support chiral analysis in amino acids and peptides. Even though MS is intrinsically “chirally blind”, because two enantiomers have the same mass and typically show identical mass spectra, some recent developments allow for enantiomer differentiation. The advancements include ion mobility mass spectrometry (IM-MS), photodissociation mass spectrometry, and mass-selected photoelectron circular dichroism (MS-PECD).75 Specifically, IM-MS has been shown as a promising new technique to achieve rapid isomer separation, and a wide range of IMS-based methods have been described over the years, including chiral ion mobility spectrometry (CIMS),76 high-field asymmetric waveform ion mobility spectrometry (FAIMS),77 traveling wave ion mobility and mass spectrometry (TWIM-MS),78 hybrid trapped ion mobility spectrometry-time-of-flight mass spectrometry (TIMS-TOFMS),79 and many others. In IM-MS, ions are first generated, usually by electrospray ionization (ESI), and then passed through a drift tube or another mobility separator, where they are subjected to a weak electric field. The ions’ drift time, which is influenced by their size and shape, is measured and used to determine their mobility. These measurements are then combined with mass spectrometry to provide high-resolution data on both the mass and structural characteristics of the ions.For D- and L-amino acids, IM-MS can help distinguish between enantiomers, as they can have slightly different ion mobilities due to subtle differences in their three-dimensional shapes. Although the mass of D- and L-amino acids is identical, their ion mobility can differ, allowing for the potential identification and differentiation of the two forms in complex mixtures. This technique is especially useful in proteomics and biomolecular research, where the precise identification of amino acid stereoisomers is crucial for understanding protein structure, folding, and function.
Recently, a novel automatic analysis was reported, where the measurement is divided into two stages - firstly, chiral derivatization of samples with an (S)-naproxen chloride, and secondly, Trapped ion-Mobility MS analysis (TIMS-MS) shown in Fig. 4.80 Developed protocol provides an interesting possibility for progress in enantioselective high-throughput screenings.
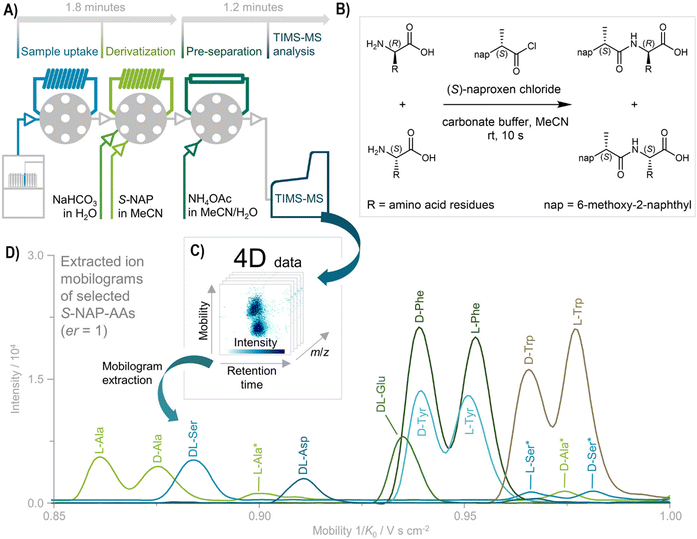 | ||
| Fig. 4 High-speed chiral analysis of amino acids (AAs); (A) the approach combines automated chiral derivatization with rapid TIMS–MS separation, achieving diastereomer resolution in under 3 minutes per sample. The derivatization process is automated through an autosampler with an integrated chromatography system. After ionization, the AA diastereomers are separated in the TIMS funnel based on their gas-phase mobility; (B) AA enantiomers undergo chiral derivatization using S-NAP; (C) the TIMS–MS system acquires 4D data, capturing retention time, mobility, m/z, and intensity; (D) extracted ion mobilograms of S-NAP-AAs from an AA standard mix (er = 1) confirm successful resolution of most D- and L-forms as deprotonated ion species. Ser is distinguished after forming a sodium acetate adduct (marked with *). Reproduced and adapted from ref. 80 with permission from American Chemical Society, copyright 2021. | ||
Another new approach, providing access to chirality information applicable to non-volatile molecular materials, such as amino acids, has been proposed by a combination of electrospray ionization (ESI) with the detection of photoelectron circular dichroism (PECD).81
2.6. Fluorescence spectroscopy
Fluorescence spectroscopy is fast, relatively inexpensive, and one of the most widely used techniques for studying numerous biological systems. The enantioselectivity, conformational changes, and binding properties of chiral compounds can be easily recognized due to the high selectivity, sensitivity, fast reaction time, non-destructive nature, and versatility of the method.82–87 Fluorescence measurements make it possible to detect compounds in the systems under study, even if they are present at very low concentrations. Fluorescence is the property of specific atoms and molecules absorbing light at a particular wavelength and then emitting light at a longer wavelength after a short delay, known as the fluorescence lifetime. The emission of electromagnetic radiation is associated with the transition of the molecule between electronic states of the same multiplicity, most often between singlet states S1 → S0.88 This transition is allowed by the quantum mechanical selection rule and is accompanied by the emission of a photon. Fluorescence emission occurs at a rate of 108 s−1, and the typical fluorescence lifetime is near 10 ns. After absorbing energy in the form of electromagnetic radiation, a molecule has several routes available to return to its ground state. The fluorescence process is illustrated by the Jablonski diagram,89 which is often used as a starting point for discussing molecular processes occurring in excited states involving the absorption and emission of light.2.7. Other methods (kinetic resolution, HPLC, electrochemistry)
Occasionally, other, more or less sophisticated methods are applied in chirality-related research, especially in cases where the isolation of the preferred enantiomer is necessary. This can be achieved not only by a typical diastereomeric or preferential crystallization, but also by kinetic resolution (chemical or enzymatic) or chromatographic resolution.90 Kinetic resolution is based on different reaction rates of two enantiomers with a chiral catalyst or reagent, resulting in an enantioenriched sample of the less reactive enantiomer. Various types of catalytic strategies might be employed, using either a single organic catalyst or enzyme, or a combination of two or more types of catalysis, such as enzyme-metal, enzyme-organo, enzyme-photoredox, heterobimetallic, and metal-organo.91 In peptide specifically, since the conversion of L/D-amino is not uniform, it is necessary to examine individual amino acids at each specific site susceptible to isomerization. It can be performed following one of the two chirality determination protocols: a sequence-dependent strategy or a sequence-independent strategy (see Fig. 5). The sequence-dependent strategy can be applied to identify the isomerized position of the amino acid on the sequence, where in the first step, an enzyme digestion is performed, followed by a chromatographic method, such as RP-HPLC, and a mass spectrometry analysis of collected fractions. Next, the sequences are hydrolyzed to amino acids, followed by a coupling to a standard protected amino acid, in order to obtain LL- and DL- dipeptides that are easy to identify due to different retention times. | ||
| Fig. 5 Flow charts for the determination of D-amino acid residues in peptides and proteins using (A) sequence-dependent and (B) sequence-independent analytical methods.92,93 Reproduced and adapted from ref. 93 with permission from Elsevier, copyright 2018. | ||
In a sequence-independent strategy, the analysis starts with a peptide or protein homogenization and separation of the water-soluble and water-insoluble fractions that undergo enzymatic digestion. The digested peptides are then separated and identified by an LC-MS analysis. Finally, their retention times are compared to that of standard synthesized peptides with several possibilities of enantiomer positions.92
Regarding the direct chromatographic separation of stereoisomers, the conventional HPLC method cannot be successfully applied, thus, chiral chromatography is used to separate compounds into their optical isomers. Nowadays, it is usually achieved by applying the chiral functionalized silica gel as a stationary phase. Some chiral stationary phases (CSPs) can separate a wide range of chiral compounds, while others are useful only for specific types of samples. Commonly used are normal-phase solvents; however, reversed-phase solvents are sporadically applied for specific columns. In different approaches, chiral additives in the mobile phase might be utilized, or derivatization of the analyte to form diastereometric products of the two enantiomers is performed, to allow a standard HPLC separation later on. In addition, capillary and microchip electrophoresis have recently emerged as powerful techniques for the analysis and separation of stereoisomers of various chiral compounds, including peptides.94
Out of other separation methods, electrochemistry is sporadically employed for the determination of the chirality of electroactive compounds. Such an approach often requires the development of novel selective and sensitive sensors for the recognition of L- and D- amino acids.95–97
Summary of common techniques used to distinguish stereoisomers, along with their typical limitations, presented as Table 1.
| Technique | Description | Limitations |
|---|---|---|
| Polarimetry | ✓ Measures the angle of rotation of plane-polarized light | ✓ Cannot distinguish diastereomers or racemic mixtures |
| ✓ Determines optical purity, distinguishes enantiomers | ✓ Varying conditions, including solvent, concentration, and temperature, can affect the results | |
| ✓ Monitors stereochemical changes in reactions | ✓ Requires precise experimental conditions | |
| Circular dichroism (CD) | ✓ Measures differences in absorption of left- and right-circularly polarized light | ✓ The results are strongly dependent on the solvent, temperature, or pH |
| ✓ Provides spectra with a shape and intensity characteristic of each enantiomer, enabling identification | ✓ Requires clean samples | |
| ✓ Enables the distinction of enantiomers | ✓ Spectra can be difficult to interpret for complex systems | |
| ✓ Sometimes needs support from other techniques | ||
| Synchrotron radiation circular dichroism (SRCD) | ✓ Enhanced CD technique using intense synchrotron light with a wide wavelength range | ✓ Requires synchrotron access |
| ✓ Provides a higher signal-to-noise ratio | ||
| ✓ Enables data collection at shorter wavelengths | ✓ Despite its higher sensitivity, it still has limitations in distinguishing stereoisomers with similar spectra | |
| ✓ Extends applications of conventional CD | ||
| Vibrational circular dichroism (VCD) | ✓ Measures differential absorption of circularly polarized light in the infrared region | ✓ Sensitive to the confirmation and environmental conditions |
| ✓ Determines the absolute configuration of organic compounds | ✓ Often needs DFT calculations | |
| ✓ Provides info on secondary structure and conformational changes in peptides and proteins | ✓ Low signal intensity compared to other spectroscopic methods | |
| X-ray diffraction (XRD) | ✓ Provides 3D atomic-level molecular structure | ✓ Requires high-quality crystals, which are difficult for large, complex, or unstable molecules |
| ✓ Determines absolute configuration | ||
| Nuclear magnetic resonance (NMR) | ✓ Analyzes chiral compounds by interaction with chiral shift reagents or by forming diastereomers | ✓ Requires relatively large amounts of a clean sample (at least a few milligrams) |
| ✓ Determines enantiomeric excess and absolute configuration | ✓ The analysis of complex spectra can be difficult and time-consuming | |
| ✓ Can study various nuclei (proton, fluorine, carbon, selenium) | ||
| Ion mobility mass spectrometry (IM-MS) | ✓ Separates ions by size, shape, and charge in a buffer gas under a weak electric field | ✓ Requires specialized equipment and expertise |
| ✓ Distinguishes enantiomers (e.g., D- vs.L-amino acids) via subtle 3D structural differences affecting mobility | ✓ Analyzing complex mixtures can be challenging, as the presence of many similar compounds often complicates spectrum interpretation | |
| ✓ Offers fast, precise, and sensitive chiral analysis with low sample consumption | ||
| Fluorescence spectroscopy | ✓ Fast, affordable, sensitive method for studying chiral fluorescent molecules | ✓ Not all compounds naturally fluoresce, which sometimes requires labeling or molecular modification |
| ✓ Enables chirality recognition and determination of enantiomeric excess | ✓ Sensitive to pH, ionic strength, and quenchers | |
| ✓ Detects compounds at very low concentrations | ✓ Difficult to distinguish molecules with similar fluorescence properties | |
| Kinetic resolution | ✓ Separates enantiomers by selectively reacting one to form a new product | ✓ Max. yield for one enantiomer is 50% without recycling |
| ✓ Yields enantiomerically pure compounds | ✓ Requires separation of product from unreacted substrate | |
| ✓ Used to determine absolute configuration | ✓ Often needs advanced variants (e.g., dynamic or parallel resolution). | |
| ✓ Common in synthesis and chiral analysis | ||
| High-performance liquid chromatography (HPLC) | ✓ Separates and quantifies enantiomers using chiral stationary phases | ✓ HPLC requires a properly selected methodology |
| ✓ Can use chiral additives or derivatization to form diastereomers for separation | ✓ Sometimes it is associated with high costs and the need for laborious optimization of conditions | |
| Electrochemistry | ✓ Distinguishes enantiomers and diastereomers by differences in redox potentials | ✓ Only for electrochemically active molecules |
| ✓ Differences may be subtle, which can make unambiguous identification difficult | ||
3. Experimental methods for studying higher-order structures of self-assembling chiral peptides.
High-resolution detection methods are largely limited by the nature of peptide self-assemblies, due to their poor water solubility and non-crystalline nature, as well as the formation of heterogeneous structures resulting in poly-dispersed samples of different sizes, shapes, and possibly different conformations. Moreover, treating solid-state constructs with solvents can destroy their subtle morphology, leading to misinterpretation of structural data. It is therefore not surprising that experimental methods designed for the study of condensed matter are preferred and give the most reliable results. These techniques can be divided into three groups: spectroscopic methods, diffraction techniques, and microscopic visualizations.3.1. Spectroscopic methods
During the MAS NMR measurement sample is spun rapidly at an angle of 54.74° relative to the applied magnetic field (B0) (See Fig. 6).103,104 This angle corresponds to the magic angle, where the second-order Legendre polynomial (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) cos2
cos2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ − 1) vanishes, reducing dipolar broadening and chemical shift anisotropy (CSA). Typically, the sample is packed in a rotor and spun at speeds ranging from a few kHz to over 100 kHz. Faster spinning leads to better averaging of anisotropic interactions, yielding sharper spectral lines. Fast sample spinning (over 60 kHz) is in particular important for recording the high-resolution 1H MAS spectra.105
θ − 1) vanishes, reducing dipolar broadening and chemical shift anisotropy (CSA). Typically, the sample is packed in a rotor and spun at speeds ranging from a few kHz to over 100 kHz. Faster spinning leads to better averaging of anisotropic interactions, yielding sharper spectral lines. Fast sample spinning (over 60 kHz) is in particular important for recording the high-resolution 1H MAS spectra.105
 | ||
| Fig. 6 Schematic representation of MAS NMR measurement and exemplary spectra. (A) Visualization of the idea of magic angle spinning NMR spectroscopy. Reproduced and adapted from ref. 104 with permission from Elsevier, copyright 2014. (B) Solid State NMR spectra of L-histidine HCl recorded with spinning rate 60 kHz (a) 1H MAS NMR (red), (b) 1H-13C CP MAS (blue), and (c) 1H-15N CP MAS (black). | ||
One of the greatest weaknesses of NMR spectroscopy is the low sensitivity of the measurements. This problem is especially acute when “rare” spins are studied. This group includes such nuclei as 13C and 15N important from the point of view of structural research of amino acids and peptides. The natural abundance of 13C is approx. 1.1%, for the 15N isotope approx. 0.36%. Sensitivity, which is a critical factor determining the time of measurement of one-dimensional spectra, becomes even more important with multi-dimensional NMR experiments, as experimental time increases with the introduction of additional indirect dimensions. Sensitivity enhancement results in the reduction of experimental time and hence can more efficiently lead to refined structural insight and complex information on molecular dynamics within the NMR time scales.106
In the solid state, the sensitivity enhancement for rare nuclei with low gyromagnetic ratio X is often achieved through Cross Polarization (CP).107 CP provides sensitivity enhancement by transferring polarization from highly abundant protons to X nuclei. Due to the excitation of protons and observation of X nucleus, an increase in S/N is expected, proportional to gyromagnetic ratio γe/γd (4 for 13C and 10 for 15N), but in real cases, this factor is always smaller. Another benefit of CP is the possibility of reducing a repetition delay due to favourable relaxation properties of 1H nuclei.
In the case when the samples are not fully solid but have some degree of motional restriction, e.g. peptides’ semi-solids, hydrogels, and slurries. High-resolution magic angle spinning (HR-MAS) NMR is a technique of choice. This technique, dedicated to the study of soft matter, enables high-resolution spectra similar to solution NMR while retaining solid-like sample conditions.108
Raman spectroscopy is another technique used to determine the vibrational modes of molecules, and it relies upon the inelastic scattering of photons, known as Raman scattering. While IR examines the wavenumber at which a functional group has a vibrational mode, Raman observes the shift in vibration from an incident source. Moreover, while IR absorption requires a dipole moment or change in charge distribution to be associated with the vibrational mode, Raman signals occur because of a molecule's polarizability. This results in the complementarity of both techniques. Even though Raman spectroscopy is not regarded as a direct chirality analysis tool, it can be effective in enantioselectivity studies.111 A relatively new method combining the variability of scattering experiments with the structural sensitivity of chiral spectroscopy is called Raman optical activity (ROA) spectroscopy. ROA measures a small difference in the intensity of Raman scattering from chiral molecules in right- and left-circularly polarized light, providing unique and detailed information about molecular structure and conformational dynamics.112 ROA has been applied for various studies of peptides and proteins, for example, the decomposition of ROA experimental spectra into calculated spectra of individual conformers, has led to the construction of the two-dimensional potential energy surface (“Ramachandran plot”) for the alanine dipeptide Ac-Ala-NHMe.113
Peptides that form amorphous or poorly crystalline aggregates present significant challenges for study through PXRD. In such cases, small-angle X-ray scattering (SAXS) and wide-angle X-ray scattering (WAXS) are particularly useful. SAXS analyzes structures at low scattering angles, delivering information about large-scale assemblies and their general shapes, such as bundles of fibrils or layered configurations. In contrast, WAXS examines higher scattering angles, providing insights into short-range molecular ordering and crystallinity, including the arrangement of peptide backbones, hydrogen bonding, and stacking interactions at the angstrom scale. A combination of SAXS and WAXS methods enable hierarchical structural elucidation from the molecular-level details up to the micron scale dimension of the nanotubes, thereby playing a crucial role in the understanding of multi-scale peptide assembly.119,120
3.2. Microscopic methods
One of the direct ways to observe the formation of self-assembling structures is through microscopic methods. Among them, transmission electron microscopy (TEM), scanning electron microscopy (SEM), atomic force microscopy (AFM), and scanning tunneling microscopy (STM) are the most popular (see Table 2).| Technique | STM | AFM | TEM | SEM |
|---|---|---|---|---|
| Resolution | >0.01 nm | >0.1 nm | >0.01 nm | >1 nm |
| Imaging principles | Tunneling electrons | Inter-atomic forces | Electrons | Electrons |
| Environment | Air/vacuum | All | Vacuum | Vacuum |
| Advantages | ✓ Excellent resolution | ✓ 3D imaging | ✓ Good resolution | ✓ Good resolution |
| ✓ 3D imaging | ✓ Applicable to conductors and insulators | ✓ Possible tomography | ✓ Possible tomography | |
| ✓ High temperature range | ✓ Low cost | ✓ Diffraction | ✓ Diffraction | |
| ✓ Height and phase imaging | ✓ Imaging in solution | ✓ Imaging in solution | ||
| Challenges | ✓ Applicable to conducting samples | ✓ Possible tip artefacts | ✓ Only 2D imaging | ✓ Low resolution |
| ✓ Fragile and very expensive | ✓ Limited vertical range and magnification range | ✓ Expensive | ✓ Only 2D imaging | |
| ✓ Time consuming | ✓ Radiation damage | ✓ Applicable to conducting samples | ||
| ✓ Difficult to operate | ✓ Difficult sample preparation | ✓ Radiation damage |
In the study of the influence of chirality on the organization of peptides, EM is and will undoubtedly remain the gold standard of imaging. The observed development of well-known new methods, as well as the use of completely new tools, provides increasingly broader possibilities for analyzing structures at both the molecular and supramolecular levels. Moreover, additional possibilities, such as the observation of dynamic processes or 3D imaging, open broader perspectives for studying changes in the created morphologies and even the slightest differences between them.
There are also promising results coming from the development of advanced techniques combining microscopy and spectroscopy methods, such as Raman and Fourier transform infrared (FTIR) imaging.142,143
3.3. Computational methods
In recent years, extensive research has focused on computational approaches to study the self-organization and self-assembly of peptides, with molecular dynamics (MD) methods being particularly powerful and frequently applied. Since self-assembling peptides can form various nanostructures, the ways to virtually screen and predict their behavior are of utmost importance. The predictive tool for self-assembly of homochiral dipeptides144 and tripeptides145 have been already explored, and involve mostly the coarse-grained MARTINI force field molecular simulations146,147 The described protocol offers a rapid determination of whether a given peptide sequence is likely to aggregate, whereas longer and larger simulations can be performed after initial screening to obtain more insights into the mechanism of formation of those systems identified as having a potential for self-assembly. The stable conformations accessible to 7- to 10-residues macrocyclic peptides composed of L- and D-amino acids were characterized through near-exhaustive backbone sampling, followed by sequence optimization and energy landscape analysis, yielding over 200 design predictions.148 The computational design of peptides and peptide macrocycles, including those of mixed chirality, has also been achieved by the Rosetta design software.149One of the novel approaches for conducting MD simulation of various molecular nanostructures, is the so-called molecular dynamics (MD) manipulator, which allows efficient assembly of simulated molecular structures, taking into account the chirality of molecular components, using external force actions.150,151 The replica exchange molecular dynamics (REMD) technique has also been utilized to determine the morphologies and diversity of self-assembled peptides, including the heteropeptides.152,153
In the last few years, novel methods based on artificial intelligence (AI) and deep learning are gaining increased attention for various peptide-related predictions,154–156 and they are expected to be a very helpful tool for studies of the heterochiral assemblies.
3.4. Rheology
Rheology is the study of deformation and flow behavior. Self-assembly of peptides often leads to the production of new soft matter, most often in the form of gels. Such materials can be characterized from various angles through rheological tests, describing and comparing their strength, viscosity, deformation, thixotropy, and others. Peptide-based materials are characterized by viscoelastic properties, which are measured in oscillation tests, where one of the most important parameters is the storage modulus G′ that represents the elastic portion of the viscoelasticity, thus represents the solid-state behavior of the sample, and the loss modulus G′′ that characterizes the viscous part, often seen as the liquid-state behavior. The two properties, monitored against time, frequency, and strain, give insight into gelation kinetics that, together with the general stiffness, directly impact the final application of the material.157 Rheology offers valuable insights into injectability or thixotropy—a material's ability to shear thin under high shear strain, followed by the recovery to a solid-like material upon removal of high shear.158 Investigating the kinetics of gel recovery and the underlying restoration mechanisms is an interesting aspect in rheological investigations and is especially crucial for the development of peptide-based injectable therapies.157In peptide-based gel research, the chirality of amino acids plays a pivotal role in determining the material's mechanical properties. Different gel types may be created with altered stiffness, distinct response to changes in concentration, alternated gelation dynamics, and in some cases, a single change in chirality may completely prevent gelation. Therefore, understanding the relationship between peptide hydrogel structure and bulk mechanical properties is one of the central challenges in this field. Thus, to tackle this, rheological studies – essential for characterizing gel behavior – should always be supplemented by structural studies using microscopy and spectroscopy. For example, techniques that integrate rheology with microscopy or scattering have been employed to explore the behavior of shear-thinning gels during and after shear flow.157 As research advances, there is a growing focus on understanding the interplay between mechanical and morphological properties, especially in the context of chirality-related studies. Specifically, rheological measurements are used in combination with other techniques to draw conclusions about the atomic-level changes in chirality and their influence on the final properties on a macroscopic level. A multidisciplinary strategy combining rheological, structural, computational, and biological methods is essential to uncover the full impact of chirality on peptide-based gels.
4. Self-assembly of homo- and heterochiral peptides
4.1. Molecular crystals of peptides
The crystal structure of a material is one of the highest forms of organization of condensed matter with a well-defined nature of nonbonding interactions. In many cases, the structural information obtained from X-ray studies of single crystals for amino acids and small peptides is taken as reference data that can be extrapolated to the interpretation of higher-order peptide assemblies. Solid state structural studies of amino acid chirality began at the end of the 19th century when O. Wallach tried to answer the question of why DL-alanine crystals are denser than L-alanine crystals. Systematic studies of chiral crystals have led to the conclusion that heterochiral molecules pack together more efficiently than homochiral molecules.159 This statement, known in the literature as Wallach's rule, has been critically evaluated several times.160 An attempt to explain the correlation between crystal packing and the stability of chiral and racemic proteogenic amino acids on the basis of experimental results and theoretical calculations was made by J. D. Dunitz and A. Gavezzotti.161It is well known that crystals are organized according to specific rules with a hierarchy of intermolecular interactions.162 The list of these interactions includes: electrostatic interactions between charged moieties, hydrogen bond interactions, van der Waals interactions, and π–π arrangement. These interactions play different roles in the hierarchical process of self-organization, moving at various scales, from elementary molecules to higher-order structures (see Fig. 7). The presence of stereogenic peptide centers can significantly interfere with the nature of these interactions. Below are some examples showing the importance of cumulative effects (non-covalent interactions, chirality) responsible for peptide self-assembly.
The most spectacular observations on the effect of chirality on the morphology of peptide crystals are related to structural studies of the phenylalanine-phenylalanine (Phe-Phe) dipeptide. Self-assembling of Phe-Phe first discovered and described by Gorbitz et al.,163 was intensively studied by Gazit164–166 and many other research groups over the years.167–171 This model has become a reference sample and a “workhorse” used to understand the fundamental processes that define the final form of matter. Fig. 8 shows the molecular packing for the dipeptides L-Phe-L-Phe and D-Phe-D-Phe constructed from experimental data of their crystallographic structures obtained by X-ray methods and deposited in the crystallographic database of the Cambridge Crystallographic Data Center (CCDC). (These data correspond to No. CCDC 163340 for L-Phe-L-Phe and No. CCDC 1853771 for D-Phe-D-Phe).172,173 In both cases the crystal structures contain six Phe molecules, and it is formed by a total of 258 atoms, while the space groups of these enantiomers are different: P61 space group for L-Phe-L-Phe and P65 for D-Phe-D-Phe. The basic structural information are collected in Table 3.
 | ||
| Fig. 8 (A) Molecular packing of L-Phe-L-Phe (CCDC 163340) and (B) D-Phe-D-Phe (CCDC 1853771) crystals. | ||
| L-Phe-L-Phe | d-Phe-d-Phe | |
|---|---|---|
| Space group | P61 | P65 |
| a, Å | 24.0709(13) | 23.9468(14) |
| b, Å | 24.0709(13) | 23.9468(14) |
| c, Å | 5.4560(4) | 5.4411(2) |
| V, Å3 | 2737.7(3) | 2702.2(2) |
It is worth noting that both crystals are organized according to the hierarchical rules described above. The zwitterionic nature of the dipeptide is correlated with long-range electrostatic interactions. Aromatic–aromatic interactions force a synclinal arrangement of the main skeleton, which opens up the possibility of creating a higher-order structure by means of hydrogen bonds. External stimuli and environmental factors can significantly affect molecular packing and disrupt the hierarchy of non-covalent interactions. Fig. 9 shows the molecular packing of L-Phe-L-Phe dipeptide crystalized in methanol.175 The differences between hydrate and methanol solvate are apparent. The latter system crystallizes in orthorhombic space group P212121 with Z′ equal 1 and Z = 4. The cell volume is 1937.25 Å3, much less compared to the hydrate. The size of the unit cell is a = 4.866 Å, b = 17.275 Å, c = 23.046 Å. The most important difference is the conformation of the dipeptide. In the case of methanol solvate, the main skeleton of the dipeptide is present in the form of a stretched chain with phenyl groups on opposite sides. In the crystalline unit, only intermolecular, weak aromatic–aromatic interactions are observed. Such a “stretched” conformation defines further possibilities for self-assembly and virtually eliminates the formation of nanotube peptides.
The intriguing example of a correlation between nonbonding interactions and chirality are crystals of racemic mixture D-Phe-D-Phe and L-Phe-L-Phe. The unit cell contains eight molecules (Z = 8) organized in homochiral pairs (see Fig. 10). Crystal crystallizes in the P21 space group. The size of the unit cell is a = 8.9648(15) Å, b = 30.515(5) Å, c = 11.9680(19) Å, volume is 3273.86 Å3. The subtle structure of this crystal is controlled by electrostatic interaction, hydrogen bonding, and chirality. These effects have an impact on further self-assembly. The specific character of hydrogen bonding interactions is the reason why the creation of nanowires rather than nanotubes is preferred.
 | ||
| Fig. 10 Molecular packing of racemic mixture D-Phe-D-Phe and L-Phe-L-Phe (CDCC 205941). Reproduced and adapted from ref. 176 with permission from John Wiley and Sons, copyright 2021. | ||
Expanding research on Phe-Phe analogs, the group of Marchesan studied the supramolecular packing modes of heterochiral, aliphatic dipeptides L-Leu-D-Xaa (Xaa = Ala, Val, Ile, Leu).177 The XRD analyses were performed using synchrotron radiation. L-Leu-D-Ala (CCDC: 2300851) crystallizes as a hydrate with water molecules localized in layers parallel to the ab plane, while the previously reported homochiral analog (L-Leu-L-Ala) forms water channels.178 The second dipeptide – L-Leu-D-Val (CCDC: 2300852) also crystallizes as a hydrate with a simple solid-state assembly, characterized by the presence of amphipathic layers and water molecules occupying non-interconnected cavities. Interestingly, a homochiral L-Leu-L-Val reported by Görbitz,179 showed a peculiar crystal packing, being the first linear peptide with Z = 24. The next two dipeptides L-Leu-D-Leu and L-Leu-D-Ile (CCDC: 2300853 and 2300854, respectively), both crystallize as DMSO solvate crystal structures with the guest molecules which occupy channels parallel to the a axis, whereas analyzed earlier L-Leu-L-Leu involves head-to-tail interaction of peptide molecules and segregation of polar and nonpolar regions.180 All the basic structural information for the abovementioned peptides is summarized in Table 4.
| La | LA | Lv | LV | Ll | LL | Li | |
|---|---|---|---|---|---|---|---|
| Space group | C2 (5) | P212121 (19) | C2 (5) | P62 (171) | P21 | P21 | P21 |
| a, Å | 16.704(3) | 6.186(2) | 19.250(4) | 29.212(16) | 5.362(11) | 5.434(4) | 5.583(11) |
| b, Å | 4.766(10) | 12.548(4) | 6.476(13) | 6.4760(13) | 14.938(3) | 15.712(7) | 14.434(3) |
| c, Å | 16.102(3) | 19.883(6) | 13.049(3) | 11.723(9) | 11.345(2) | 11.275(2) | 22.161(4) |
| α | 90° | 90° | 90° | 90° | 90° | 90° | 90° |
| β | 99.05(3)° | 90° | 103.72(3)° | 90° | 91.61(3)° | 100.41(1)° | 92.76(3)° |
| γ | 90° | 90° | 90° | 119.99° | 90° | 90° | 90° |
To further broaden the scope of research, Mandal and colleagues published a comprehensive X-ray diffraction study of single-crystal X-ray (SC-XRD) heterochiral Boc-Val-Phe-Phe-OMe tripeptides.181 In contrast to the previously discussed examples, tripeptide models are protected at both the C and N ends, which means that they are not zwitterionic systems. It has a significant impact on the organization of the crystal lattice because it eliminates electrostatic interactions as a controlling factor for self-assembly.
Table 5 shows the basic structural information and lattice cell parameters for Boc-Val-Phe-Phe-OMe crystals. Boc-L-Val-D-Phe-D-Phe-OMe crystallized in the monoclinic (P21) space group and exhibited one molecule in the asymmetric unit. Each subunit was interconnected through an intermolecular H-bond to adopt a parallel beta-sheet structure along the c-axis in the crystalline state. The NH group and oxygen of L-Val of one subunit were intermolecularly H-bonded to the next subunit of the oxygen of urethane and NH of central D-Phe, respectively. Both peptide Boc-L-Val-D-Phe-L-Phe-OMe and Boc-D-Val-L-Phe-D-Phe-OMe crystallized in the triclinic (P1) space group and showed one molecule in the asymmetric unit. Each molecule of both Boc-L-Val-D-Phe-L-Phe-OMe and Boc-D-Val-L-Phe-D-Phe-OMe was interlinked through three intermolecular H-bonds to create a parallel beta-sheet structure along b-direction. Peptides Boc-L-Val-L-Phe-D-Phe-OMe and Boc-D-Val-D-Phe-L-Phe-OMe crystallized in orthorhombic (P212121) space group and exhibited one molecule) in the asymmetric unit. Each molecule of both Boc-L-Val-L-Phe-D-Phe-OMe and Boc-D-Val-D-Phe-L-Phe-OMe was intermolecularly interconnected through three H-bonds to form a parallel beta-sheet structure along c-direction. The oxygen of urethane, NH, and oxygen of central Phe of one molecule was intermolecularly H-bonded to the next molecule of the NH and oxygen of Val and the NH of C-terminal Phe, respectively in both Boc-L-Val-L-Phe-D-Phe-OMe and Boc-D-Val-D-Phe-L-Phe-OMe. Next, molecules were self-associated to build a helical sheet-like structure in higher-order packing along a-axis in both enantiomeric peptides Boc-L-Val-L-Phe-D-Phe-OMe and Boc-D-Val-D-Phe-L-Phe-OMe. Fig. 11 shows the molecular structure of models refined by means of X-ray crystallography.
| Vff | VfF | vFf | VFf, | vfF | |
|---|---|---|---|---|---|
| Space group | P21 | P1 | P1 | P212121 | P212121 |
| a, Å | 5.0081(2) | 4.9397(5) | 4.9548(4) | 5.0710(2) | 5.0543(9) |
| b, Å | 26.9990(17) | 10.8307(10) | 10.8351(12) | 15.1956(6) | 15.267(3) |
| c, Å | 11.0196(8) | 13.6935(13) | 13.6908(16) | 37.6927(14) | 39.355(7) |
| V, Å3 | 1488.98 | 724.469 | 726.851 | 2904.48 | 3036.79 |
 | ||
| Fig. 11 Crystal structures of different tripeptides: (A) Boc-L-Val-D-Phe-D-Phe-OMe (Vff), (B) Boc-L-Val-D-Phe-L-Phe-OMe (VfF) and Boc-D-Val-L-Phe-D-Phe-OMe (vFf), (C) Boc-L-Val-L-Phe-D-Phe-OMe (VFf) and Boc-D-Val-D-Phe-L-Phe-OMe (vfF). Reproduced and adapted from ref. 181 with permission from John Wiley and Sons, copyright 2021. | ||
Fig. 12 shows how chirality inversion can dramatically change the arrangement of molecules in the crystal lattice for peptides with different stereochemistry. Fig. 12A displays the molecular packing for Boc-L-Val-D-Phe-L-Phe-OMe, while Fig. 12B shows the molecular packing for Boc-L-Val-L-Phe-D-Phe-OMe sample.
 | ||
| Fig. 12 Molecular packing of (A) Boc-L-Val-D-Phe-L-Phe-OMe (CCDC 2015774) and (B) Boc-L-Val-L-Phe-D-Phe-OMe (CCDC 2015776). | ||
The same group has investigated L/L and L/D amino acid-containing dipeptides, Boc-L-Ile-L-Phe-OMe, Boc-L-Ile-D-Phe-OMe, Boc-L-Ile-L-Phg-OMe (Phg = phenylglycine), and Boc-L-Ile-D-Phg-OMe (Fig. 13A).182Fig. 13B shows how stereochemistry and chemical modification affect the solid-state peptide backbone arrangement. The presence of extra methylene (–CH2–) group on the side-chain of C-terminal Phe of Boc-L-Ile-L-Phe-OMe causes a significant deviation of molecular arrangements from that of Boc-L-Ile-L-Phg-OMe. While the molecules in Boc-L-Ile-L-Phe-OMe self-assemble to form a single helix-like architecture, the molecules in Boc-L-Ile-L-Phg-OMe self-associate around a water molecule to form a cylinder-like structure in the crystalline state. Alternatively, the observed supramolecular arrangements of Boc-D-Val-L-Ile-OMe and Boc-L-Ile-D-Val-OMe are similar, but deviate from that of Boc-L-Val-L-Ile-OMe and Boc-L-Ile-L-Val-OMe, due to chirality difference.183
 | ||
| Fig. 13 The X-ray crystal structures of N- and C-protected L/L and D/L dipeptides containing such amino acids as Boc-L-Val-L-Ile-OMe, Boc-D-Val-L-Ile-OMe, Boc-L-Ile-L-Val-OMe, and Boc-L-Ile-D-Val-OMe. Reproduced and adapted from ref. 183 with permission from Elsevier, copyright 2020. | ||
Other interesting cases might be found in studies focused on cyclic dipeptides. Investigations have been conducted on the cyclic compounds of Tyr-Ala and Tyr-D-Ala noted as cyclo(L-Tyr-L-Ala) and cyclo(L-Tyr-D-Ala), respectively, and structural differences between them have been noticed (see Fig. 14A–D). In particular, slight differences were observed in the arrangement of water molecules in the hydrophilic channels and the geometry of the diketopiperazine rings.184 Very recently, structural data for cyclo(L-Tyr-L-Tyr) and cyclo(L-Tyr-D-Tyr) were published. While the cyclo(L-Tyr-L-Tyr) exists in few solid forms that display a network of parallel hydrogen-bonded stripes, promoting a unilateral growth, its D-Tyr containing analog shows a propensity towards a two-dimensional growth (see Fig. 14E and F).185
 | ||
| Fig. 14 Molecular and crystal structures for cyclo(L-Tyr-L-Ala) (panels (A) and (B)) and cyclo(L-Tyr-D-Ala) (panels (C) and (D)); (E) hydrogen bonding network present in the crystal structure of cyclo(L-Tyr-L-Tyr) cY(D)Y; (F) hydrogen bonds network observed in the crystal structure of cyclo(L-Tyr-L-Tyr) methanol solvate. Reproduced and adapted from ref. 184 with permission from American Chemical Society, copyright 2015, and from ref. 185 with permission from John Wiley and Sons, copyright 2022. | ||
4.2. Higher order structures of self-assembling homo and heterochiral peptides
Self-assembly of peptides refers to the spontaneous organization of peptide molecules into ordered nanostructures or supramolecular architectures driven by non-covalent interactions shortly discussed in Section 4.1. This process results in the formation of well-defined structures with specific functions and properties. The spatial distribution of the residues around the stereogenic center determines the morphology of the final product. Thus, we can conclude that the chiral structural synthon (CSS), which is the smallest basic building block, is responsible for the formation of complex higher-order structures in the process of proliferation (self-organization). This hypothesis can be verified taking as an example molecular structure in the crystal lattice of Phe-Phe dipeptide (see Section 4.1). For L-Phe-L-Phe and D-Phe-D-Phe, where the arrangement of phenyl groups is synclinal, then CSS creates the possibility of formation of peptide nanotubes because the hydrophobic and hydrophilic parts are located in different spaces (Fig. 15). | ||
| Fig. 15 (A) Supramolecular structure formed by multiplication of chiral structural synthon (CSS) of L-Phe-L-Phe or D-Phe-D-Phe; (B) SEM picture of peptide nanotube (PNT) formed by L-Phe-L-Phe or D-Phe-D-Phe. Reproduced and adapted from ref. 186 with permission from American Chemical Society, copyright 2012. | ||
When such an arrangement is not possible because the solvent forces a different spatial arrangement (e.g.L-Phe-L-Phe crystallized from methanol) and the arrangement of phenyl groups is anti-planar, then there is no chance to form nanotubes. In this case, the peptide nanowires might be formed (see Fig. 16A and B). The influence of solvent type on morphology, assembly kinetics, and orientation of Phe-Phe peptides has been explored both experimentally and computationally, revealing that its lower solvophobicity in methanol results in much weaker self-assembly.175,187
 | ||
| Fig. 16 Supramolecular structure formed by multiplication of chiral structural synthon (A) L-Phe-L-Phe crystallized from methanol; (B) racemic mixture of D-Phe-D-Phe and L-Phe-L-Phe. | ||
The chemically modified di- and tri-peptides offer a great variety of self-assembling structures and thus have been largely studied in recent years, especially for biomedical and material applications.10,167,188 Recently, it has been discovered that halogenation affects supramolecular organization, in particular, fluorination leads to the preservation of dipeptide packing in the water channels, while iodination completely changes the packing from the water channels to the amphipathic layers of the stacked peptides.189 Marchesan and coworkers for the analysis of the self-organization of heterochiral and halogenated dipeptides have employed modifications of fluorine or iodide substituted Phe–Phe. In particular, fluorination maintained the arrangement of dipeptides within aqueous channels, similar to what is observed with non-halogenated dipeptides. This modification provided an intermediate level of intra- versus inter-molecular hydrophobic interactions, leading to a moderate level of fibril bundling (Fig. 17).189 SC-XRD structures revealed that despite the different stereo-configurations of homochiral and heterochiral diphenylalanine, they displayed a similar supramolecular packing, although the N-terminal amino acid dictated the screw-sense handedness from N- to C-terminus to be left-handed for L-Phe-L-Phe and right-handed for D-Phe-L-Phe (see Fig. 18). Moreover, the heterochiral peptide displayed an increased intramolecular hydrophobic contact area between Phe side chains.
 | ||
| Fig. 17 (A) Single-crystal XRD structure (CCDC 2016373) of D-(4-F)-Phe-L-Phe; (B) and (C) powder XRD on microcrystals grown on hydrogels of D-(2-F)-Phe-L-Phe and D-(3-F)-Phe-L-Phe confirmed packing analogous to sample D-(4-F)-Phe-L-Phe. Reproduced and adapted from ref. 189 under the terms of the Creative Commons CC BY 4.0 license (https://creativecommons.org/licenses/by/4.0). | ||
 | ||
| Fig. 18 Single-crystal XRD structures of homochiral L-Phe-L-Phe36 (silver) and heterochiral D-Phe-L-Phe (cyan, CCDC 2016375) peptide nanotubes.37 (A) Side-view of homochiral and heterochiral peptides revealed opposite screw-sense, dictated by the N-terminal amino acids. (B) Top-view of nanotube inner cavities defined by the projection six peptide molecules arranged head-to-tail. (C) Top-view of the nanotubes identified by the projection of 18 peptide molecules showing hierarchical bundling stabilized by aromatic zippers for L-Phe-L-Phe (silver) and a smoother outer surface, with an outer diameter matching well the fibril size in a case of D-Phe-L-Phe (cyan). Reproduced and adapted from ref. 189 under the terms of the Creative Commons CC BY 4.0 license (https://creativecommons.org/licenses/by/4.0). | ||
Of the twenty amino acids known, proline is a very special case due to its unique structure. Unlike most amino acids, proline contains a cyclic side chain in which the nitrogen atom is part of a five-membered ring. The change in conformation and/or puckering of the five-membered ring may be another factor (in addition to chirality) that can have a strong effect on the self-assembly of short peptides. To explain the role of proline in the formation of functional nanostructures, Garcia et al. studied D- and L-amino acids in the Pro-Phe-Phe sequence (see Fig. 19) to resolve the conflict between the contradictory assembly directing effects of Pro as β-breaker and Phe-Phe as a β-sheet-associated motif.190
 | ||
| Fig. 19 (A) Pro-Phe-Phe Stereoisomers by Enantiomer Pairs (a/b) and Their Cartoon Keycode (Each Amino Acid is Represented as Either a Blue (L) or Gold (D) Sphere); (B) TEM (left) and AFM (right) images: (a) 1a aggregates, 2a bundles of nanofibrils, (c) 3a nanoparticles, (d) 4a nanotapes. Reproduced and adapted from ref. 190 under CC BY 4.0 license (https://creativecommons.org/licenses/by/4.0). | ||
Based on the obtained results, the authors concluded that the stereo-configuration determines the ability of each of the eight possible Pro-Phe-Phe stereoisomers to self-associate into diverse nanostructures, including nanoparticles, nanotapes, or fibrils, which yielded hydrogels with gel-to-sol transition at a physiologically relevant temperature.
Going further with understanding the role of proline chirality in the self-assembly of peptides Chatterjee et al. have studied the nature of the conformational preferences of four distinct classes of model diproline peptides, by crystallographic and NMR studies.191 The analysis of pyrrolidine conformations suggested a preferred proline puckering geometry favored only in the case of heterochiral diproline segments. Moreover, a detailed view of the possible conformational states of heterochiral and homochiral Pro-Pro sequences was additionally supported by the NMR studies in solution. Of the fourteen systems studied, the crystal and molecular structures of the two heterochiral tripeptides Piv-L-Pro-D-Pro-L-Ala-OMe and Piv-D-Pro-L-Pro-D-Ala-OMe were also discussed. The first system crystallizes in P212121 (19) space group and Z′ = 1 (Fig. 20A) while the second one in P21 (4) space group and two independent molecules in the crystal lattice (Z′ = 2) (Fig. 20B and C).
 | ||
| Fig. 20 Molecular structure of tripeptides: (A) Piv-L-Pro-D-Pro-L-Ala-OMe; (B) and (C) Piv-D-Pro-L-Pro-D-Ala-OMe. | ||
The molecular structure of each molecule displayed in Fig. 20 shows subtle differences in the conformation of the proline ring. On the other hand, the D-Pro-Pro motif is known to serve as an inducing element of β-torsion, while homochiral Pro-Pro induces local helical folding.192,193 The localization of D-Pro in the peptide sequence may also be an important factor in determining supramolecular organization. Fig. 21 shows the supramolecular structure for Piv-L-Pro-D-Pro-L-Ala-OMe and Piv-D-Pro-L-Pro-D-Ala-OMe tripeptides. A strong tendency to form fibers is evident in the second case.
 | ||
| Fig. 21 Supramolecular structure formed by multiplication of chiral structural synthon (CSS): (A) Piv-L-Pro-D-Pro-L-Ala-OMe and (B) Piv-D-Pro-L-Pro-D-Ala-OMe. | ||
It is interesting to note that even single aromatic amino acids can self-assemble into amyloid-like fibers. Specifically, phenylalanine (Phe) and tryptophan (Trp) were found to be important in nanostructure formation in many different studies.194–197 Recently, Bera et al. decided to explore the effect of D-enantiomers of these amino acids on the self-assembly pattern and the molecular arrangement of the resultant structural organization and demonstrated the critical role of chirality in the process. The authors applied a wide range of different experimental techniques to probe the self-assembly mechanism and the involvement of various interactions in the structure formation. The DL-composites of aromatic amino acids have a high aggregation propensity and present a favorable knob-to-hole packing of aromatic rings, which induced the easy growth of racemic crystals and the fabrication of self-assembled rigid materials (see Fig. 22).198 The potential of D- and L-enantiomers of phenylalanine was later further explored towards the development of functional metal–organic materials.199,200 Taking into account the association of some self-assembling peptides with pathological conditions, such as the amyloid-β in Alzheimer's disease, there is a tremendous interest in studying the short amyloidogenic sequences.
 | ||
| Fig. 22 Single-crystal structural analysis of (A)–(C) L-Trp and (D)–(F) DL-Trp. L-Trp: (A) molecular chain formation (pink) and tape-like arrangement (blue); (B) supramolecular β-sheet structure; (C) twisted angle between two adjacent aromatic rings highlighted by a yellow circle in (B). DL-Trp: (D) 3D network arrangement through H-bonding interactions of molecular chains (pink); (E) supramolecular β-sheet structure; (F)“Knobs-in-holes” fitting of adjacent aromatic rings highlighted by a yellow circle in (E). Reproduced and adapted from ref. 198 under CC BY 4.0 license (https://creativecommons.org/licenses/by/4.0). | ||
The influence of stereochemistry on self-organization in the solid state of cyclic dipeptides (CDP) employing two diastereomeric samples; cyclo(L-Tyr-L-Ala) and cyclo(L-Tyr-D-Ala) as models was investigated by Jeziorna et al.184,186 It has been found that distinction in chirality of alanine residue causes a significant difference in self-assembling and formation of higher order structures. Sample cyclo(L-Tyr-L-Ala) forms peptide nanotubes (PNT) and nanowires (PNW) (see Fig. 23A and B), while for sample cyclo(L-Tyr-D-Ala) only formation of peptide microtubes (PMT) was observed (see Fig. 23C and D). Crystal and molecular structures for both samples were refined using PXRD due to failure in attempts to grow crystals with quality suitable for single crystal studies. Both compounds crystallize in the P21 space group and monoclinic system. The size of the unit cell is highly similar; however, small differences in alignment of water molecules in the hydrophilic channels and geometry of diketopiperazine rings were observed. The water molecules can be thermally removed from the lattice without destroying the subtle crystal structures of nano- and microdevices. This reversible process observed for cyclo(L-Tyr-D-Ala) is a unique feature, rarely occurring for the linear dipeptide devices.
 | ||
| Fig. 23 SEM images of cyclo(L-Tyr-L-Ala) (A) and cyclo(L-Tyr-D-Ala) (C); supramolecular array of cyclo(L-Tyr-L-Ala) (B) and cyclo(L-Tyr-D-Ala) (D). Reproduced and adapted from ref. 184 with permission from American Chemical Society, copyright 2015. | ||
Finally, it is worthy to stress that the self-organization of heterochiral peptides can be controlled by chemical modification of the protecting group. For example, peptides related to the Ala-Ala building block have been investigated by Lin et al.201 In their work, a special focus has been given to the position of the substituent attached to the benzoyl residue at the N-terminus, on the molecular packing, and the self-assembly behavior. Specifically, three peptide isomers with an ortho-, meta-, and para-dodecyloxy group on the benzoyl residue at the N-terminus were designed and synthesized (see Fig. 24). It was demonstrated that the ortho- substituent position presents a strong steric hindrance and the occurrence of intramolecular hydrogen bonding, resulting in different molecular packing compared with the two other isomers.
 | ||
| Fig. 24 FE-SEM images of the three dipeptide self-assemblies. All samples were prepared in a CH3OH/H2O mixed solvent (6/4, v/v) at a concentration of 2.0 g L−1; ortho (A)–(E), meta (F) and (G) and para (H) and (I). Reproduced and adapted from ref. 201 with permission from Royal Society of Chemistry, copyright 2021. | ||
5. Formation of gels
Many self-assembling peptides can create new functional soft matter, such as hydrogels or organogels. Hydrogels are characterized by a high amount of water inside the network, making them biocompatible and biodegradable, thus making them auspicious materials for biological and biomedical applications. On the other hand, organogels are created by the entrapment of large amounts of organic solvents into the peptide assemblies. Peptide-based gels are stimuli-responsive and sensitive to the type of solvent, presence of salts, pH, temperature, or ultrasounds, thus, they can be easily affected by environmental chemical and physical stimuli. Many other internal factors, including chirality, influence the gelation properties of peptides and might be used as a valuable tool for driving self-assembly. Effects observed at molecular and supramolecular levels, described in detail in the previous chapters, are amplified through increasing size scale to macroscopic changes and finally affect the properties of peptide-based materials.The influence of molecular packing and structural features on the properties of peptide-based hydrogels has been extensively studied, offering valuable insights into their design. Chirality effects play a crucial role in the self-assembly and macroscopic properties of peptide hydrogels. A great number of insights into the effects of chirality on peptide-based soft matter have been delivered in works published by Marchesan and co-workers. Studies on dipeptides like Phe-Val and Val-Phe demonstrated that heterochirality enables fibrillation and gelation from otherwise non-gelling homochiral sequences (see Fig. 25).202
 | ||
| Fig. 25 Photographs and optical microscopy images of (A) D-Phe–L-Val and (B) D-Val–L-Phe in PBS at concentrations above their solubility. Scale bar = 100 μm. Reproduced and adapted from ref. 202 with permission from Royal Society of Chemistry, copyright 2022. | ||
Likewise, the four stereoisomers of Leu-Phe dipeptides revealed that heterochirality promotes hydrogelation, with all heterochiral dipeptides forming stable gels and one specific configuration (D-Phe-L-Leu) leading to metastable hydrogel converting into crystals.203 Investigations of tripeptides, such as Val-Phe-Phe and Phe-Phe-Val, highlighted the influence of D- and L-amino acid positions on supramolecular order and macroscopic properties. For instance, configurations like DLL or LDD facilitated the formation of hydrophobic channels that excluded water, resulting in durable hydrogels with high thermal stability. Conversely, other configurations formed lower-order assemblies, leading to weaker gels.204
Steric hindrance is another important factor in hydrogel formation and stability. For example, investigations into tripeptides such as D-Leu-L-Phe-L-Phe revealed that heterochirality promotes hydrogelation in an aqueous solution following a pH switch through extended 3D molecular structures. This arrangement supports the formation of elongated fibers, high fiber density, and large fiber bundles, resulting in immediate macroscopic gelation. In addition, a phenylalanine zipper has been identified as an important interaction motif between interdigitated β-sheets of the D-Leu-L-Phe-L-Phe. In contrast, the sterically hindered L-epimer did not gel, highlighting the significance of steric considerations.205 Similarly, studies showed that peptides with β-branched amino acids (such as valine or isoleucine) formed dense networks with higher resistance to applied stress, while in contrast the linear side chains negatively affected hydrogel durability and rheological properties.206
Also, the high supramolecular order was shown to significantly influence the mechanical and thermal properties of peptide-based hydrogels. The presence of tightly packed β-sheets and hydrophobic channels contributed to enhanced gel durability. Notably, stereoisomers such as D-Val-L-Phe-L-Phe and L-Val-D-Phe-D-Phe exhibited contrasting behaviors in water exclusion and stability, which were directly linked to their ability to form high-order supramolecular structures. Specifically, in the mentioned configuration the dry hydrophobic channels are formed, resulting in the exclusion of water molecules, giving rise to a higher supramolecular order.207 In case of some tri-peptides, the formation of a racemic mixture can lead to novel or improved properties of the gel. It has been described that the co-assembly of tripeptides: L-His-D-Phe-D-Phe and D-His-L-Phe-L-Phe forms anisotropic nanostructures that form gel with better viscoelastic properties and enhanced catalytic activity.208
Finally, the influence of external stimuli, such as ultrasound, was studied to optimize nanostructure alignment towards anisotropic organo- and hydrogels consisting of oriented tripeptide structures based on D-Phe-L-Phe-L-Asp and D-Phe-L-Phe-L-Ile. The former peptide formed an anisotropic organogel in methanol, while the latter formed an anisotropic hydrogel, showing the relationship between the anisotropic arrangement of supramolecular interactions with peptide sequence and solvent environment.209
In addition to frequently studied linear peptides, the gelling properties of their cyclic analogues are sporadically explored. It was found that while cyclo(L-Tyr-L-Tyr) easily assembles into hydrogels, conformation of cyclo(D-Tyr-L-Tyr) promotes rather two-directional growth, resulting in a tendency towards crystallization.185 Interestingly, heterochiral cyclodipeptides cyclo(D-Leu-L-Phe) and cyclo(L-Leu-D-Phe) are able to form gels in a soybean oil.210
5.1. Heterochirality in biomedical approaches
In many cases, biological applications and biocompatibility of the peptide-based materials are explored alongside structural insights. Especially, since peptides containing D-amino acids are considered a great tool in biomedical research due to their resistance to enzymatic degradation. Since human proteases exclusively recognize and cleave L-amino acids, replacing them with their D-enantiomers makes peptides more resistant to proteolysis, significantly increasing their biostability. Consequently, the improved stability may translate into better in vivo circulation half-lives, improving delivery yield and efficacy, and potentially reducing dosing frequency.Many naturally occurring antibiotics, such as dactinomycin, bacitracin, gramicidin, and others, contain D-amino acid residues in their sequences. In many cases, the pattern formed by the alternation of D- and L-amino acids is crucial for the formation of the peptide's proper structure, with gramicidin helices being one of the most important examples.211 Thus, a lot of scientific attention in organic chemistry has been given to the total synthesis of peptide antibiotics, highlighting the role of heterochirality in their structural and functional properties.212 Over the years, heterochiral peptides have been used not only as antimicrobial agents.213,214 but also as potential cancer therapeutics215,216 and vaccine candidates.217 The recent progress and challenges in the exploration of D-amino acids at the interface of chemistry and life science have been nicely reviewed elsewhere,218,219 as well as their role in antimicrobial resistance,220,221 and in numerous physiological processes in the human body.222–224 Therefore, in this chapter, we would like to briefly introduce some recent examples focusing specifically on the biomedical applications of hydrogels based on heterochiral peptides.
Most of the gelling peptides, regardless of chirality, demonstrated excellent performance in cell culture experiments, with no significant differences observed between enantiomers in supporting cell viability and proliferation. For example, both D-Val-L-Phe-L-Phe and L-Val-D-Phe-D-Phe gelling peptides performed well in cell culture experiments, with no cytotoxicity observed.207 Such results support the potential future development of cost-effective biomaterial gels from heterochiral assembling peptides. One of the envisioned biomedical applications of peptide hydrogels is their possible role as cell scaffolds. Unfortunately, the native amino acids can be easily decomposed by proteolytic enzymes, making them mostly unsuitable for this application. Thus, sequences containing D-amino acids might offer a reasonable alternative. For instance, a hydrogel formed by a D-P1 peptide (Ac–Phe–Phe–Phe–Gly–Lys) has gelation properties like its L-form,225 but is resistant to a protease, stable in a cell-culture medium, and can serve as a scaffold for cell cultures.226 Few relevant studies on other peptide sequences explored further different aspects of chirality influence the hydrogel properties and degradation, as well as its impact in biological context, especially towards novel immunotherapies.227–229 Co-assembly and stereocomplexation are promising strategies for improved rigidity and generation of hybrid biomaterials, applicable, for example, in regenerative medicine.230–232 They can dramatically affect hydrogel stiffness and degradation, allowing control over novel shapes, network morphologies, and material properties, and offer proteolytic stability for the material's lifetime control.
Much less is known about the gelation process in longer peptides, and especially how it is affected by the changes in chirality. One interesting recent example describes a systematic D-scan of natural hexapeptide Phe-Ile-Asn-Tyr-Val-Lys that was performed, including the biophysical characterization of the obtained hydrogels and their biocompatibility. Despite the differences in the conformational intermediates during aggregation, all sequences resulted in hydrogels compatible with a cellular environment.233
6. Self-assembly and gel formation with chemically modified heterochiral peptides
In recent years, many groups have focused their research not only on the native peptide sequences but also on modified peptides and peptide-based conjugates, expanding the scope of native peptide studies to explore new ways of influencing self-assembly processes. This concept is based on the idea that the gelling properties of peptides can be affected by both chirality and the presence of chemical modifications. Even the mere presence of molecular chirality in a peptide sequence has a crucial impact on its supramolecular assembly and gelation, as has been shown for 1,8-naphthalimide (NMI)-conjugated hybrid dipeptides. While chiral NMI derivatives (NMI-βAla-L-Val-OMe and NMI-βAla-D-Val-OMe) formed self-supported gels, the achiral NMI derivative (NMI-βAla-Aib-OMe) failed to form any kind of gel.234 In many examples, peptide gelators were believed to require a single enantiomer for gelation, however, some recent studies demonstrate that heterochiral sequences can yield gels with unique properties. McAulay et al. explored the dipeptide 2Nap-Phe-Phe containing naphthyl (Nap) group, which is a popular building block for assisting peptide self-assembly, developing immune adjuvants, and adding specific electrical properties. In their investigations, all enantiomers, diastereomers, and even racemates of this dipeptide could form gels at high pH.235 These findings were further supported by a cryo-EM near-atomic resolution approach. The pH-dependent gelation mechanism has been explained by the tubes' carbonyl group protonation that allows them to come together and form a gel.236The possibility of gelation in racemic mixtures was also explored in the case of modified cyclic dipeptides – specifically fluorenyl (OFm) mono-substituted cyclo(L-Glu-L-Glu). Their racemate enables gelation in a large variety of organic solvents and water and displays a tunable thixotropic behavior.237 This ability of peptide racemates to gel broadens the design possibilities for novel gel-based materials.
The effects of subtle changes in chirality can significantly influence biofunctionality and mechanical properties, for instance as described for Nap-containing peptides. Specifically, Nap-L-Gly-L-Phe-L-Phe-L-Tyr, have been used to create vaccine adjuvant hydrogels.228 Its heterochiral variant, Nap-L-Gly-L-Phe-L-Phe-D-Tyr, showed improved mechanical properties and resistance to external forces, due to their dense nanofiber network (see Fig. 26).238
 | ||
| Fig. 26 The chemical structures and optical images of (A) Nap-L-Gly-D-Phe-L-Phe-L-Tyr (5) (B) Nap-L-Gly-L-Phe-D-Phe-L-Tyr (6), and (C) Nap-L-Gly-L-Phe-L-Phe-D-Tyr (7) hydrogels and their mechanical properties (D). Reproduced and adapted from ref. 238 under the terms of the Creative Commons CC BY-NC 3.0 license. | ||
Ferrocene (Fc)-based peptide conjugates offer another example of modified systems with tunable self-assembly and gelation behaviors, especially due to the interesting structural properties of the Fc group. Studies on monosubstituted Fc conjugates (FcCO–Phe-Phe-Ala-OMe) demonstrated the dramatic effects of diastereomeric interactions on gelation, with enantiomers showing no significant impact.239 A ferrocene-diphenylalanine (Fc-Phe-Phe) peptide derivative forms stable chiral nanostructures in a narrow pH range (5.7–5.9) (Fig. 27). Compared to the uniformly chiral Fc-L-Phe-L-Phe, a racemic mixture of the L and D enantiomers resulted in formation of weaker and less stable gels, demonstrating the crucial role of molecular chirality.240
 | ||
| Fig. 27 Schematic illustration for the ultra-pH-sensitive of Fc-L-Phe-L-Phe hydrogel (A) and photographic images showing the self-assembly behavior of 2 mg mL−1 Fc-L-Phe-L-Phe in 10% 2-propanol/90% H2O at phosphate buffer (0.1 M) with different pH values (B). Reproduced and adapted from ref. 240 with permission from Elsevier, copyright 2020. | ||
The impact of chirality on modulation of photoresponsive behaviors was analyzed in a case of azobenzene-modified dipeptides. Chiral gelators have been built on LL-dipeptides or DD-dipeptides with an azobenzene group functioning as a bridge between the peptide units. These gelators, built on LL- or DD-dipeptides demonstrated distinct gel–sol transition speeds during the trans-to-cis (E/Z) isomerization of the azobenzene group.241 The self-assembly of azobenzene-linked chiral dipeptide gelators can also be modulated by circularly polarized light (CPL). In such case right-handed CPL causes the gel to collapse, whereas left-handed CPL promotes new self-assembly, maintaining the macroscopic gel structure (see Fig. 28).242
 | ||
| Fig. 28 (A) Illustration of different supramolecular self-assembly modulated by CPL handedness, right-CPL promotes the collapse of ordered structure, but left-CPL triggers the formation of new helical structure; (B) chemical structure of 4,4′-azobenzene-linked dipeptide gelator, abbreviated to Azo-DF. Reproduced and adapted from ref. 242 under the terms of the Creative Commons CC BY-NC 3.0 license. | ||
Chirality has also been explored in metal-peptide assemblies, where it influences both structure and function. These effect have been understood more deeply in studies focused on single amino acid residues243,244 but still quite little is known about the effect of chirality on the architectures and properties of metal-peptide assemblies. Zhang et al. investigated Phe-Phe dipeptides coordinated with Cu2+ ions, forming a metalloenzyme mimic (see Fig. 29). Both the homochiral L-Phe-L-Phe-Cu and their racemate (L/D) Phe-Phe-Cu assemblies demonstrated laccase-like catalytic activity, but the heterochiral assemblies exhibited significantly higher catalytic efficiency and reusability. These results highlight the potential of chirality in tuning the physicochemical properties of functional materials.245
 | ||
| Fig. 29 Schematic illustration of the chirality-dependent coordination assembly process between Phe-Phe molecules and Cu2+, in which the peptide heterochirality leads to enhanced properties of a laccase mimic. Reproduced and adapted from ref. 245 with permission from American Chemical Society, copyright 2022. | ||
Peptide gels also serve as effective media for protein crystallization, with their inherent chirality significantly influencing the process. Conejero-Muriel et al. examined the influence of the chirality of the gel fibers in protein crystallogenesis. Two cysteine-based peptides – N,N′-di(benzoyl)-L-cysteine diamide and its corresponding D-enantiomer have been employed to grow protein crystals, and differences in crystal quality and packing have been observed, suggesting a significant influence of the chirality in this process.246 Protein crystallization in gels based on the Fmoc-dipeptides family resulted in formation of high-quality protein crystals with enantiomeric hydrogels influencing mostly the nucleation and growth of the same protein.247
Amino acid chirality was also employed in the formation of chiral gelators for the self-assembly driven enantioselective recognition. The development of chiral supramolecular gels for enantioselective recognition through gel formation or collapse has been nicely summarized in a review by Gambhir et al.248 Some most notable recent examples involving amino acids, peptides, or their derivatives have been applied to enantioselective recognition of carboxylates,249 chiral recognition of mandelic acid and N-tosyl α-amino acids,250 or enantioselective gelation of tartaric acid enantiomers.251 Future investigations are likely to explore more in-depth the effects of chirality on chiral recognition in other peptide sequences, towards their sensing applications.
7. Conclusion and future perspectives
In recent years, remarkable progress has been made in the field of peptide self-assembly, including studies on the heterochiral peptide systems that combine both L- and D-amino acids. These advanced biomaterials offer unique structural and functional properties that homochiral systems cannot achieve. In this tutorial review, we show analytical tools that allow us to distinguish stereoisomers, describe the problem of chirality and its impact on the organization of higher-order structures, and understand at the molecular level the mechanism of self-assembly and the formation of different hetero-chiral materials with tailored functions. Existing literature highlights chirality as one of the vital factors that can easily and profoundly influence the formation of various peptide morphologies. We believe that one of the most important achievements in the field will be the ability to successfully predict and dictate the formation of heterochiral supramolecular structures. Firstly, it may provide important insights for protein aggregation disorders and subsequently lead to the discovery of amyloid formation inhibitors and new therapeutic approaches. Secondly, it will offer the possibility to create novel soft materials with pre-designed properties for various applications in biomedicine, catalysis, nanotechnology, etc. Finally, it will broaden our basic understanding of self-organization significantly, improving our knowledge about one of the fundamental bases for the bottom-up creation of artificial life-like systems. Even though a lot of progress has been made in recent years, it remains challenging to draw a pattern allowing for a straightforward prediction of the chirality switching influence on the self-assembly of a particular peptide. The current progress in machine learning (ML) and deep learning (DL) approaches will inevitably result in a more rationalized design of heterochiral peptide sequences with predictable properties.The spectrum of future applications is vast, demanding, and far from routine solutions. The structure of this review and the choice of the presented material reflect our belief that overcoming these challenges will require investigations combining a wide range of novel experimental and computational methods, often including hybrid techniques. A lot of progress is anticipated especially in studies of peptide assembly dynamics, with the use of the latest techniques, such as advanced microscopic imaging via fluorescence lifetime imaging microscopy (FLIM), single-molecule fluorescence lifetime correlation spectroscopy (FLCS),252 stochastic optical reconstruction microscopy (STORM),253 and others, that might be particularly interesting in studies of self-assembly. Further improvements are also expected from state-of-the-art spectroscopic methods, such as time-resolved synchrotron radiation circular dichroism (tr-SRCD),254 surface-enhanced Raman spectroscopy (SERS),255 new NMR approaches,256 and many more. Undoubtedly, such a cutting-edge approach will require interdisciplinary collaboration across chemistry, materials science, biology, and engineering.
Finally, based on current research trends and technological advancements, several exciting prospects are emerging for biomedical functional materials formed by the self-assembly of heterochiral peptides. Future bio-applications include work to improve the mechanical properties of peptide gels through topological control, as well as advanced antimicrobial and therapeutic applications, personalized medicine, and targeted drug delivery systems. Additionally, advances in the development of biomimetic materials and tissue engineering are anticipated. However, since the incorporation of D-amino acids has significant biological importance, altering stability, solubility, cytotoxicity, assembly, and therapeutic effects, biological studies must explore deeper cell interactions with peptides of altered chirality, their bioavailability, and biodegradability. Moreover, manufacturing heterochiral peptide compounds presents additional challenges, related to the complex synthesis, scalability, high costs, and difficulties with purification and analysis. The regulatory concerns and GMP considerations must also be addressed to enable the successful production of such materials for clinical use.
In summary, to realize the full potential of D-amino acid-containing peptides as self-assembling materials, future research must incorporate interdisciplinary strategies that combine advanced characterization techniques with AI-driven design. At the same time, deeper biological investigations will be crucial for safe and effective biomedical applications, towards clinical use. Based on the remarkable advancement in recent years, we expect to observe even faster progress in the field soon, with great promise in the use of engineered heterochiral peptides as a versatile and functional platform for next-generation therapies and technologies.
Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This work was supported by a Polish National Science Centre (NCN) [Grant No. UMO-2022/47/I/ST4/01629].References
- L. D. Barron, Chem. Soc. Rev., 1986, 15, 189–223 RSC.
- L. D. Barron, Space Sci. Rev., 2008, 135, 187–201 CrossRef CAS.
- Stereochemistry. A Three-Dimensional Insight, ed. A. V. Karnik and M. Hasan, Elsevier, 2021 Search PubMed.
- Q. Sallembien, L. Bouteiller, J. Crassous and M. Raynal, Chem. Soc. Rev., 2022, 51, 3436–3476 RSC.
- V. A. Pavlov and E. I. Klabunovskii, Curr. Org. Chem., 2014, 18, 93–114 CrossRef CAS.
- J.-M. Cavaillon and S. Legout, Biomolecules, 2022, 12, 596 CrossRef CAS PubMed.
- H. D. Flack, Acta Crystallogr., Sect. A:Found. Crystallogr., 2009, 65, 371–389 CrossRef CAS PubMed.
- A. B. Buda, T. A. der Heyde and K. Mislow, Angew. Chem., Int. Ed. Engl., 1992, 31, 989–1007 CrossRef.
- J. Wang, K. Liu, R. Xing and X. Yan, Chem. Soc. Rev., 2016, 45, 5589–5604 RSC.
- G. Fichman and E. Gazit, Acta Biomater., 2014, 10, 1671–1682 CrossRef CAS PubMed.
- E. Mattia and S. Otto, Nat. Nanotechnol., 2015, 10, 111–119 CrossRef CAS PubMed.
- D. C. Sherrington and K. A. Taskinen, Chem. Soc. Rev., 2001, 30, 83–93 RSC.
- Y. Shen, Y. Wang, I. W. Hamley, W. Qi, R. Su and Z. He, Prog. Polym. Sci., 2021, 123, 101469 CrossRef CAS.
- Y. Zheng, K. Mao, S. Chen and H. Zhu, Front. Bioeng. Biotechnol., 2021, 9, 703004 CrossRef PubMed.
- V. Apostolopoulos, J. Bojarska, T.-T. Chai, S. Elnagdy, K. Kaczmarek, J. Matsoukas, R. New, K. Parang, O. P. Lopez, H. Parhiz, C. O. Perera, M. Pickholz, M. Remko, M. Saviano, M. Skwarczynski, Y. Tang, W. M. Wolf, T. Yoshiya, J. Zabrocki, P. Zielenkiewicz, M. AlKhazindar, V. Barriga, K. Kelaidonis, E. M. Sarasia and I. Toth, Molecules, 2021, 26, 430 CrossRef CAS PubMed.
- R. Perlikowska, Peptides, 2021, 140, 170528 CrossRef CAS PubMed.
- L. Schnaider, L. Shimonov, T. Kreiser, D. Zaguri, D. Bychenko, I. Brickner, S. Kolusheva, A. Lichtenstein, J. Kost and E. Gazit, ACS Appl. Bio Mater., 2020, 3, 8395–8401 CrossRef CAS PubMed.
- A. Arul, S. Sivagnanam, A. Dey, O. Mukherjee, S. Ghosh and P. Das, RSC Adv., 2020, 10, 13420–13429 RSC.
- B. Gilboa, C. Lafargue, A. Handelman, L. J. W. Shimon, G. Rosenman, J. Zyss and T. Ellenbogen, Adv. Sci., 2017, 4, 1700052 CrossRef PubMed.
- B. Xue, M. Qin, T. Wang, J. Wu, D. Luo, Q. Jiang, Y. Li, Y. Cao and W. Wang, Adv. Funct. Mater., 2016, 26, 9053–9062 CrossRef CAS.
- Y. Shen, Y. Wang, I. W. Hamley, W. Qi, R. Su and Z. He, Prog. Polym. Sci., 2021, 123, 101469 CrossRef CAS.
- U. J. Meierhenrich, Eur. Rev., 2013, 21, 190–199 CrossRef.
- V. S. Lamzin, Z. Dauter and K. S. Wilson, Curr. Opin. Struct. Biol., 1995, 5, 830–836 CrossRef CAS PubMed.
- H. G. Brittain, J. Pharm. Biomed. Anal., 1998, 17, 933–940 CrossRef CAS PubMed.
- A. Gogoi, S. Konwer and G.-Y. Zhuo, Front. Chem., 2020, 8, 611833 CrossRef CAS PubMed.
- D. Kondepudi, in Chiral Analysis, ed. P. L. Polavarapu, Elsevier, 2nd edn, 2018, pp. 3–28 Search PubMed.
- P. L. Polavarapu, Tetrahedron: Asymmetry, 1997, 8, 3397–3401 CrossRef CAS.
- P. L. Polavarapu, Molecules, 2016, 21, 1056 CrossRef PubMed.
- L. A. Nafie, Vibrational Optical Activity: Principles and Applications, Wiley, 2011 Search PubMed.
- K. Ng, T. J. Edkins and D. R. Bobbitt, Chirality, 1999, 11, 187–194 CrossRef CAS.
- C. Meinert, J. H. Bredehöft, J.-J. Filippi, Y. Baraud, L. Nahon, F. Wien, N. C. Jones, S. V. Hoffmann and U. J. Meierhenrich, Angew. Chem., Int. Ed., 2012, 51, 4484–4487 CrossRef CAS PubMed.
- P. J. Stephens, F. J. Devlin and J.-J. Pan, Chirality, 2008, 20, 643–663 CrossRef CAS PubMed.
- H. Yamagishi, H. Sato and I. Kawamura, Chirality, 2021, 33, 652–659 CrossRef CAS PubMed.
- D. Kurouski, Anal. Chim. Acta, 2017, 990, 54–66 CrossRef CAS PubMed.
- J. M. Bijvoet, A. F. Peerdeman and A. J. van Bommel, Nature, 1951, 168, 271–272 CrossRef CAS.
- A. L. Thompson and D. J. Watkin, Tetrahedron: Asymmetry, 2009, 20, 712–717 CrossRef CAS.
- H. D. Flack, Chimia, 2014, 68, 26 CrossRef CAS PubMed.
- H. D. Flack and G. Bernardinelli, Chirality, 2008, 20, 681–690 CrossRef CAS PubMed.
- S. Parsons, Tetrahedron: Asymmetry, 2017, 28, 1304–1313 CrossRef CAS.
- Determination of Absolute and Relative Configuration: 4.2. Determination of Absolute and Relative Configuration by X-ray and Neutron Diffraction Methods, Thieme Verlag, 4th edn, 1995.
- H. D. Flack, Acta Crystallogr., Sect. A:Found. Crystallogr., 1983, 39, 876–881 CrossRef.
- J. R. Deschamps, Life Sci., 2010, 86, 585–589 CrossRef CAS PubMed.
- V. Kumar Vashistha, Asian J. Org. Chem., 2022, 11, e202200544 CrossRef CAS.
- C. Aroulanda and P. Lesot, Chirality, 2022, 34, 182–244 CrossRef CAS PubMed.
- D. Parker, Chem. Rev., 1991, 91, 1441–1457 CrossRef CAS.
- T. J. Wenzel, Differentiation of Chiral Compounds Using NMR Spectroscopy, John Wiley & Sons, Ltd, 2018, pp. i–xxi Search PubMed.
- D. Leung, S. O. Kang and E. V. Anslyn, Chem. Soc. Rev., 2011, 41, 448–479 RSC.
- M. P. Williamson and J. P. Waltho, Chem. Soc. Rev., 1992, 21, 227–236 RSC.
- V. Kumar Vashistha, Asian J. Org. Chem., 2022, 11, e202200544 CrossRef CAS.
- P. L. Rinaldi, Prog. Nucl. Magn. Reson. Spectrosc., 1982, 15, 291–352 CrossRef CAS.
- T. J. Wenzel and C. D. Chisholm, Chirality, 2011, 23, 190–214 CrossRef CAS PubMed.
- R. R. Groleau, M. E. Powell, C. D. Evans, P. S. Fordred, S. D. Bull and T. D. James, in Comprehensive Chirality, ed. J. Cossy, Academic Press, Oxford, 2nd edn, 2024, pp. 593–631 Search PubMed.
- J. M. Seco, E. Quiñoá and R. Riguera, Chem. Rev., 2004, 104, 17–118 CrossRef CAS.
- J. M. Seco, E. Quiñoá and R. Riguera, Chem. Rev., 2012, 112, 4603–4641 CrossRef CAS PubMed.
- F. Balzano, A. Iuliano, G. Uccello-Barretta and V. Zullo, J. Org. Chem., 2022, 87, 12698–12709 CrossRef CAS PubMed.
- W. H. Pirkle, D. L. Sikkenga and M. S. Pavlin, J. Org. Chem., 1977, 42, 384–387 CrossRef CAS.
- J. A. Dale, D. L. Dull and H. S. Mosher, J. Org. Chem., 1969, 34, 2543–2549 CrossRef CAS.
- J. M. Seco, E. Quiñoá and R. Riguera, Tetrahedron: Asymmetry, 2001, 12, 2915–2925 CrossRef CAS.
- M. Malinowska, S. Jarzyński, A. Pieczonka, M. Rachwalski, S. Leśniak and A. Zawisza, J. Org. Chem., 2020, 85, 11794–11801 CrossRef CAS PubMed.
- J. Wen, L. Feng, H. Zhao, L. Zheng, P. Stavropoulos, L. Ai and J. Zhang, J. Org. Chem., 2022, 87, 7934–7944 CrossRef CAS PubMed.
- Z. Sun, Z. Chen, Y. Wang, X. Zhang, J. Xu, G. Bian and L. Song, Org. Lett., 2020, 22, 589–593 CrossRef CAS PubMed.
- J. Yang, B. Chatelet, V. Dufaud, D. Hérault, M. Jean, N. Vanthuyne, J.-C. Mulatier, D. Pitrat, L. Guy, J.-P. Dutasta and A. Martinez, Org. Lett., 2020, 22, 891–895 CrossRef CAS PubMed.
- S. Jang and H. Kim, Asian J. Org. Chem., 2021, 10, 886–890 CrossRef CAS.
- I. Louzao, J. M. Seco, E. Quiñoá and R. Riguera, Chem. Commun., 2010, 46, 7903–7905 RSC.
- M. Pérez-Trujillo, E. Monteagudo and T. Parella, Anal. Chem., 2013, 85, 10887–10894 CrossRef PubMed.
- D. Prasad, S. Mogurampelly and S. R. Chaudhari, RSC Adv., 2020, 10, 2303–2312 RSC.
- M. Palomino-Schätzlein, A. Virgili, S. Gil and C. Jaime, J. Org. Chem., 2006, 71, 8114–8120 CrossRef PubMed.
- M. S. Silva, Molecules, 2017, 22, 247 CrossRef PubMed.
- Y. Jia, L. Wen, W. Bao, Z. Xu, J. Wu and Y. Zhao, Anal. Chem., 2023, 95, 10362–10367 CrossRef CAS PubMed.
- H. Li, Z. Xu, S. Zhang, Y. Jia and Y. Zhao, Anal. Chem., 2022, 94, 2023–2031 CrossRef CAS PubMed.
- J. Liang, Z. Xu, J. Wu and Y. Zhao, Anal. Chem., 2023, 95, 7569–7574 CrossRef CAS PubMed.
- O. N. Gorunova, I. M. Novitskiy, Y. K. Grishin, I. P. Gloriozov, V. A. Roznyatovsky, V. N. Khrustalev, K. A. Kochetkov and V. V. Dunina, Organometallics, 2016, 35, 75–90 CrossRef CAS.
- Y. Zhao and T. M. Swager, J. Am. Chem. Soc., 2015, 137, 3221–3224 CrossRef CAS PubMed.
- Z. Xu and Y. Zhao, J. Fluorine Chem., 2023, 266, 110089 CrossRef CAS.
- D.-Q. Han and Z.-P. Yao, TrAC, Trends Anal. Chem., 2020, 123, 115763 CrossRef CAS.
- P. Dwivedi, C. Wu, L. M. Matz, B. H. Clowers, W. F. Siems and H. H. Hill, Anal. Chem., 2006, 78, 8200–8206 CrossRef CAS PubMed.
- A. Mie, M. Jörntén-Karlsson, B.-O. Axelsson, A. Ray and C. T. Reimann, Anal. Chem., 2007, 79, 2850–2858 CrossRef CAS.
- V. Domalain, M. Hubert-Roux, V. Tognetti, L. Joubert, C. M. Lange, J. Rouden and C. Afonso, Chem. Sci., 2014, 5, 3234–3239 RSC.
- R. Pérez-Míguez, B. Bruyneel, M. Castro-Puyana, M. L. Marina, G. W. Somsen and E. Domínguez-Vega, Anal. Chem., 2019, 91, 3277–3285 CrossRef PubMed.
- J. M. Will, A. Behrens, M. Macke, C. D. Quarles and U. Karst, Anal. Chem., 2021, 93, 878–885 CrossRef CAS PubMed.
- P. Krüger and K.-M. Weitzel, Angew. Chem., Int. Ed., 2021, 60, 17861–17865 CrossRef PubMed.
- S. O. Fakayode, A. A. Williams, M. A. Busch, K. W. Busch and I. M. Warner, J. Fluoresc., 2006, 16, 659–670 CrossRef CAS.
- O. Rusin, O. Alpturk, M. He, J. O. Escobedo, S. Jiang, F. Dawan, K. Lian, M. E. McCarroll, I. M. Warner and R. M. Strongin, J. Fluoresc., 2004, 14, 611–615 CrossRef CAS PubMed.
- O. Rusin, V. Král, J. O. Escobedo and R. M. Strongin, Org. Lett., 2004, 6, 1373–1376 CrossRef CAS PubMed.
- A. Belardini, E. Petronijevic, R. Ghahri, D. Rocco, F. Pandolfi, C. Sibilia and L. Mattiello, Appl. Sci., 2021, 11, 11375 CrossRef CAS.
- L. Pu, Acc. Chem. Res., 2017, 50, 1032–1040 CrossRef CAS PubMed.
- Y.-Y. Zhu, X.-D. Wu, S.-X. Gu and L. Pu, J. Am. Chem. Soc., 2019, 141, 175–181 CrossRef CAS PubMed.
- B. Valeur and M. N. Berberan-Santos, Molecular Fluorescence: Principles and Applications, John Wiley & Sons, Ltd, 2012 Search PubMed.
- D. Frackowiak, J. Photochem. Photobiol., B, 1988, 2, 399 CrossRef.
- I. P. Silvestri and P. J. J. Colbon, ACS Med. Chem. Lett., 2021, 12, 1220–1229 CrossRef CAS PubMed.
- G. L. Thejashree, E. Doris, E. Gravel and I. N. N. Namboothiri, Eur. J. Org. Chem., 2022, e202201035 CrossRef CAS.
- M. Morvan and I. Mikšík, Separations, 2021, 8, 112 CrossRef CAS.
- T. Miyamoto and H. Homma, Biochim. Biophys. Acta, Proteins Proteomics, 2018, 1866, 775–782 CrossRef CAS PubMed.
- V. Kasicka, Electrophoresis, 2022, 43, 82–108 CrossRef CAS PubMed.
- R. Dominguez, B. Serra, A. J. Reviejo and J. M. Pingarron, Anal. Biochem., 2001, 298, 275–282 CrossRef CAS PubMed.
- L. Zhang, G. Wang, C. Xiong, L. Zheng, J. He, Y. Ding, H. Lu, G. Zhang, K. Cho and L. Qiu, Biosens. Bioelectron., 2018, 105, 121–128 CrossRef.
- Y. Zhou, B. Yu and K. Levon, Chem. Mater., 2003, 15, 2774–2779 CrossRef CAS.
- T. Polenova, R. Gupta and A. Goldbourt, Anal. Chem., 2015, 87, 5458–5469 CrossRef CAS PubMed.
- R. Tycko, Annu. Rev. Phys. Chem., 2001, 52, 575–606 CrossRef CAS PubMed.
- M. Bonaccorsi, T. Le Marchand and G. Pintacuda, Curr. Opin. Struct. Biol., 2021, 70, 34–43 CrossRef CAS PubMed.
- P. J. Chu, M. J. Potrzebowski, A. I. Scott and Y. Gao, J. Am. Chem. Soc., 1990, 112, 881–883 CrossRef CAS.
- M. M. Słabicki, M. J. Potrzebowski, G. Bujacz, S. Olejniczak and J. Olczak, J. Phys. Chem. B, 2004, 108, 4535–4545 CrossRef.
- B. Reif, S. E. Ashbrook, L. Emsley and M. Hong, Nat. Rev. Methods Primers, 2021, 1, 1–23 CrossRef.
- M. Deschamps, in Annual Reports on NMR Spectroscopy, ed. G. A. Webb, Academic Press, 2014, vol. 81, pp. 109–144 Search PubMed.
- Y. Nishiyama, G. Hou, V. Agarwal, Y. Su and A. Ramamoorthy, Chem. Rev., 2023, 123, 918–988 CrossRef CAS PubMed.
- M. K. Dudek, S. Kaźmierski and M. J. Potrzebowski, in Annual Reports on NMR Spectroscopy, ed. G. A. Webb, Academic Press, 2021, vol. 103, pp. 97–189 Search PubMed.
- E. O. Stejskal, J. Schaefer and J. S. Waugh, J. Magn. Reson., 1977, 28, 105–112 CAS.
- H. Farooq, D. Courtier-Murias, R. Soong, W. Bermel, W. M. Kingery and A. J. Simpson, Curr. Org. Chem., 2013, 17, 3013–3031 CrossRef CAS.
- N. Sengupta, H. Maekawa, W. Zhuang, C. Toniolo, S. Mukamel, D. J. Tobias and N.-H. Ge, J. Phys. Chem. B, 2009, 113, 12037–12049 CrossRef CAS PubMed.
- V. Cervetto, P. Hamm and J. Helbing, J. Phys. Chem. B, 2008, 112, 8398–8405 CrossRef CAS PubMed.
- H. Kong, X. Sun, L. Yang, X. Liu, H. Yang and R.-H. Jin, Anal. Chem., 2020, 92, 14292–14296 CrossRef CAS PubMed.
- V. Parchaňský, J. Kapitán and P. Bouř, RSC Adv., 2014, 4, 57125–57136 RSC.
- V. Parchaňský, J. Kapitán, J. Kaminský, J. Šebestík and P. Bouř, J. Phys. Chem. Lett., 2013, 4, 2763–2768 CrossRef PubMed.
- G. H. Beaven and E. R. Holiday, in Advances in Protein Chemistry, ed. M. L. Anson, K. Bailey and J. T. Edsall, Academic Press, 1952, vol. 7, pp. 319–386 Search PubMed.
- L. Yang, F. Luo and W. Wei, Anal. Methods, 2021, 13, 1905–1910 RSC.
- L. Huang, X. Zhang, C. Xu, J. Wang, A. Zhang, Y. Liu and H. Zhu, Adv. Opt. Mater., 2025, 2500053 CrossRef CAS.
- K. Makabe, D. McElheny, V. Tereshko, A. Hilyard, G. Gawlak, S. Yan, A. Koide and S. Koide, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 17753–17758 CrossRef CAS PubMed.
- I. Bak-Sypien, T. Pawlak, P. Paluch, A. Wroblewska, R. Dolot, A. Pawlowicz, M. Szczesio, E. Wielgus, S. Kaźmierski, M. Górecki, R. Pawlowska, A. Chworos and M. J. Potrzebowski, Sci. Rep., 2024, 14, 12825 CrossRef CAS PubMed.
- I. W. Hamley and V. Castelletto, Adv. Colloid Interface Sci., 2023, 318, 102959 CrossRef CAS PubMed.
- T. Narayanan, A. Rüter and U. Olsson, Front. Bioeng. Biotechnol., 2021, 9, 654339 CrossRef PubMed.
- T. Gulik-Krzywicki, C. Fouquey and J. Lehn, Proc. Natl. Acad. Sci. U. S. A., 1993, 90, 163–167 CrossRef CAS PubMed.
- F. Schulz, I. Lokteva, W. J. Parak and F. Lehmkühler, Part. Part. Syst. Charact., 2021, 38, 2100087 CrossRef.
- R. Kubota, W. Tanaka and I. Hamachi, Chem. Rev., 2021, 121, 14281–14347 CrossRef CAS PubMed.
- J. W. Smith and Q. Chen, J. Mater. Chem. B, 2020, 8, 8490–8506 RSC.
- K. M. Yip, N. Fischer, E. Paknia, A. Chari and H. Stark, Nature, 2020, 587, 157–161 CrossRef CAS PubMed.
- T. Nakane, A. Kotecha, A. Sente, G. McMullan, S. Masiulis, P. M. G. E. Brown, I. T. Grigoras, L. Malinauskaite, T. Malinauskas, J. Miehling, T. Uchański, L. Yu, D. Karia, E. V. Pechnikova, E. de Jong, J. Keizer, M. Bischoff, J. McCormack, P. Tiemeijer, S. W. Hardwick, D. Y. Chirgadze, G. Murshudov, A. R. Aricescu and S. H. W. Scheres, Nature, 2020, 587, 152–156 CrossRef CAS PubMed.
- J. Harapin, M. Eibauer and O. Medalia, Structure, 2013, 21, 1522–1530 CrossRef CAS PubMed.
- K. C. Bustillo, S. E. Zeltmann, M. Chen, J. Donohue, J. Ciston, C. Ophus and A. M. Minor, Acc. Chem. Res., 2021, 54, 2543–2551 CrossRef CAS PubMed.
- K. Bian, C. Gerber, A. J. Heinrich, D. J. Müller, S. Scheuring and Y. Jiang, Nat. Rev. Methods Primers, 2021, 1, 1–29 CrossRef.
- P. K. Hansma, V. B. Elings, O. Marti and C. E. Bracker, Science, 1988, 242, 209–216 CrossRef CAS PubMed.
- K. Skorupska, D. J. Smith, D. S. A. Campbell, H. Jungblut and J. J. L. Hans, ECS Trans., 2007, 2, 63 CrossRef.
- L.-P. Xu, Y. Liu and X. Zhang, Nanoscale, 2011, 3, 4901–4915 RSC.
- B. Hou, T. Zhang, H. Yang, X. Han, L. Liu, L. Li, C. Grazioli, X. Wu, N. Jiang and Y. Wang, Interdiscip. Mater., 2023, 2, 511–528 Search PubMed.
- J. A. A. W. Elemans, I. D. Cat, H. Xu and S. D. Feyter, Chem. Soc. Rev., 2009, 38, 722–736 RSC.
- M. Mura, F. Silly, G. A. D. Briggs, M. R. Castell and L. N. Kantorovich, J. Phys. Chem. C, 2009, 113, 21840–21848 CrossRef CAS.
- T. Kudernac, S. Lei, J. A. A. W. Elemans and S. D. Feyter, Chem. Soc. Rev., 2009, 38, 402–421 RSC.
- A. Rodríguez-Galván and F. F. Contreras-Torres, Nanomaterials, 2022, 12, 3013 CrossRef PubMed.
- M. Nasrollahzadeh, M. Atarod, M. Sajjadi, S. M. Sajadi and Z. Issaabadi, in Interface Science and Technology, ed. M. Nasrollahzadeh, S. M. Sajadi, M. Sajjadi, Z. Issaabadi and M. Atarod, Elsevier, 2019, vol. 28, pp. 199–322 Search PubMed.
- S. Maity, J. Ottelé, G. M. Santiago, P. W. J. M. Frederix, P. Kroon, O. Markovitch, M. C. A. Stuart, S. J. Marrink, S. Otto and W. H. Roos, J. Am. Chem. Soc., 2020, 142, 13709–13717 CrossRef CAS PubMed.
- M. Manrho, S. R. Krishnaswamy, B. Kriete, I. Patmanidis, A. H. de Vries, S. J. Marrink, T. L. C. Jansen, J. Knoester and M. S. Pshenichnikov, J. Am. Chem. Soc., 2023, 145, 22494–22503 CrossRef CAS PubMed.
- V. G. Gisbert, S. Benaglia, M. R. Uhlig, R. Proksch and R. Garcia, ACS Nano, 2021, 15, 1850–1857 CrossRef CAS PubMed.
- C.-C. Hsu, J. Xu, B. Brinkhof, H. Wang, Z. Cui, W. E. Huang and H. Ye, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 18412–18423 CrossRef CAS PubMed.
- F. Palombo, F. Tamagnini, J. C. G. Jeynes, S. Mattana, I. Swift, J. Nallala, J. Hancock, J. T. Brown, A. D. Randall and N. Stone, Analyst, 2018, 143, 850–857 RSC.
- P. W. J. M. Frederix, R. V. Ulijn, N. T. Hunt and T. Tuttle, J. Phys. Chem. Lett., 2011, 2, 2380–2384 CrossRef CAS PubMed.
- P. W. J. M. Frederix, G. G. Scott, Y. M. Abul-Haija, D. Kalafatovic, C. G. Pappas, N. Javid, N. T. Hunt, R. V. Ulijn and T. Tuttle, Nat. Chem., 2015, 7, 30–37 CrossRef CAS PubMed.
- S. J. Marrink, H. J. Risselada, S. Yefimov, D. P. Tieleman and A. H. de Vries, J. Phys. Chem. B, 2007, 111, 7812–7824 CrossRef CAS PubMed.
- L. Monticelli, S. K. Kandasamy, X. Periole, R. G. Larson, D. P. Tieleman and S.-J. Marrink, J. Chem. Theory Comput., 2008, 4, 819–834 CrossRef CAS PubMed.
- P. Hosseinzadeh, G. Bhardwaj, V. K. Mulligan, M. D. Shortridge, T. W. Craven, F. Pardo-Avila, S. A. Rettie, D. E. Kim, D.-A. Silva, Y. M. Ibrahim, I. K. Webb, J. R. Cort, J. N. Adkins, G. Varani and D. Baker, Science, 2017, 358, 1461–1466 CrossRef CAS PubMed.
- V. K. Mulligan, C. S. Kang, M. R. Sawaya, S. Rettie, X. Li, I. Antselovich, T. W. Craven, A. M. Watkins, J. W. Labonte, F. DiMaio, T. O. Yeates and D. Baker, Protein Sci., 2020, 29, 2433–2445 CrossRef CAS PubMed.
- I. V. Likhachev and V. S. Bystrov, Math. Biol. Bioinf., 2021, 16, 244–255 CrossRef CAS.
- V. Bystrov, I. Likhachev, A. Sidorova, S. Filippov, A. Lutsenko, D. Shpigun and E. Belova, Nanomaterials, 2022, 12, 861 CrossRef CAS PubMed.
- P. Zhou, L. Deng, Y. Wang, J. R. Lu and H. Xu, Langmuir, 2016, 32, 4662–4672 CrossRef CAS PubMed.
- P. Zhou, X. Hu, J. Li, Y. Wang, H. Yu, Z. Chen, D. Wang, Y. Zhao, S. M. King, S. E. Rogers, J. Wang, J. R. Lu and H. Xu, J. Am. Chem. Soc., 2022, 144, 21544–21554 CrossRef CAS PubMed.
- Z. Liu, J. Wang, Y. Luo, S. Zhao, W. Li and S. Z. Li, Briefings Bioinf., 2023, 24, bbad409 CrossRef PubMed.
- Z. Yang, S. K. Yorke, T. P. J. Knowles and M. J. Buehler, Sci. Adv., 2025, 11, eadv1971 CrossRef CAS PubMed.
- E. Dražić, D. Jelušić, P. Janković Bevandić, G. Mauša and D. Kalafatovic, ACS Nano, 2025, 19, 20295–20320 CrossRef PubMed.
- C. Yan, A. Altunbas, T. Yucel, R. P. Nagarkar, J. P. Schneider and D. J. Pochan, Soft Matter, 2010, 6, 5143–5156 RSC.
- J. K. Wychowaniec, A. M. Smith, C. Ligorio, O. O. Mykhaylyk, A. F. Miller and A. Saiani, Biomacromolecules, 2020, 21, 2285–2297 CrossRef CAS PubMed.
- K.-H. Ernst, Chimia, 2018, 72, 399 CrossRef CAS PubMed.
- C. P. Brock, W. B. Schweizer and J. D. Dunitz, J. Am. Chem. Soc., 1991, 113, 9811–9820 CrossRef CAS.
- J. D. Dunitz and A. Gavezzotti, J. Phys. Chem. B, 2012, 116, 6740–6750 CrossRef CAS PubMed.
- C. Yuan, W. Ji, R. Xing, J. Li, E. Gazit and X. Yan, Nat. Rev. Chem., 2019, 3, 567–588 CrossRef CAS PubMed.
- C. H. Görbitz, Chem. - Eur. J., 2001, 7, 5153–5159 CrossRef.
- M. Reches and E. Gazit, Science, 2003, 300, 625–627 CrossRef CAS PubMed.
- M. Reches and E. Gazit, Nano Lett., 2004, 4, 581–585 CrossRef CAS.
- N. Kol, L. Adler-Abramovich, D. Barlam, R. Z. Shneck, E. Gazit and I. Rousso, Nano Lett., 2005, 5, 1343–1346 CrossRef CAS PubMed.
- S. Marchesan, A. V. Vargiu and K. E. Styan, Molecules, 2015, 20, 19775–19788 CrossRef CAS PubMed.
- J.-H. Lee, K. Heo, K. Schulz-Schönhagen, J. H. Lee, M. S. Desai, H.-E. Jin and S.-W. Lee, ACS Nano, 2018, 12, 8138–8144 CrossRef CAS PubMed.
- E. Mayans and C. Alemán, Molecules, 2020, 25, 6037 CrossRef CAS PubMed.
- H. Erdoğan, Soft Matter, 2021, 17, 5221–5230 RSC.
- P. M. Rodger, C. Montgomery, G. Costantini and A. Rodger, Phys. Chem. Chem. Phys., 2021, 23, 4597–4604 RSC.
- P. S. Zelenovskiy, A. S. Nuraeva, S. Kopyl, S. G. Arkhipov, S. G. Vasilev, V. S. Bystrov, D. A. Gruzdev, M. Waliczek, V. Svitlyk, V. Ya. Shur, L. Mafra and A. L. Kholkin, Cryst. Growth Des., 2019, 19, 6414–6421 CrossRef CAS.
- V. S. Bystrov, P. S. Zelenovskiy, A. S. Nuraeva, S. Kopyl, O. A. Zhulyabina and V. A. Tverdislov, J. Mol. Model., 2019, 25, 199 CrossRef PubMed.
- Advancing Structural Science | CCDC, https://www.ccdc.cam.ac.uk/, (accessed 21 March 2025).
- T. O. Mason, D. Y. Chirgadze, A. Levin, L. Adler-Abramovich, E. Gazit, T. P. J. Knowles and A. K. Buell, ACS Nano, 2014, 8, 1243–1253 CrossRef CAS PubMed.
- P. S. Zelenovskii, K. Romanyuk, M. S. Liberato, P. Brandão, F. F. Ferreira, S. Kopyl, L. M. Mafra, W. A. Alves and A. L. Kholkin, Adv. Funct. Mater., 2021, 31, 2102524 CrossRef CAS.
- E. Scarel, M. De Corti, M. Polentarutti, G. Pierri, C. Tedesco and S. Marchesan, J. Pept. Sci., 2024, 30, e3559 CrossRef CAS PubMed.
- C. H. Görbitz, Acta Crystallogr., Sect. C:Cryst. Struct. Commun., 1997, 53, 736–739 CrossRef.
- C. H. Görbitz, E. Gundersen, F. Teixidor, C. Viñas, M. M. Abad, R. I. Nielsen, C. E. Olsen, C. N. Rosendahl, M. Haugg, N. Trabesinger-Rüf and E. G. Weinhold, Acta Chem. Scand., 1996, 50, 537–543 CrossRef.
- S. N. Mitra and E. Subramanian, Biopolymers, 1994, 34, 1139–1143 CrossRef CAS PubMed.
- G. Dolai, R. S. Giri, S. Roy and B. Mandal, Pept. Sci., 2021, 113, e24229 CrossRef CAS.
- S. Roy, R. S. Giri, G. Dolai and B. Mandal, J. Mol. Struct., 2020, 1221, 128877 CrossRef CAS.
- R. S. Giri, S. Pal, S. Roy, G. Dolai, S. R. Manne, S. Paul and B. Mandal, Pept. Sci., 2021, 113, e24176 CrossRef CAS.
- A. Jeziorna, K. Stopczyk, E. Skorupska, K. Luberda-Durnas, M. Oszajca, W. Lasocha, M. Górecki, J. Frelek and M. J. Potrzebowski, Cryst. Growth Des., 2015, 15, 5138–5148 CrossRef CAS.
- A. Chotera-Ouda, A. Jeziorna, S. Kaźmierski, R. Dolot, M. K. Dudek and M. J. Potrzebowski, Chem. – Eur. J., 2022, 28, e202202005 CrossRef CAS PubMed.
- M. Jaworska, A. Jeziorna, E. Drabik and M. J. Potrzebowski, J. Phys. Chem. C, 2012, 116, 12330–12338 CrossRef CAS.
- A. N. Rissanou, E. Georgilis, E. Kasotakis, A. Mitraki and V. Harmandaris, J. Phys. Chem. B, 2013, 117, 3962–3975 CrossRef CAS PubMed.
- P. Makam and E. Gazit, Chem. Soc. Rev., 2018, 47, 3406–3420 RSC.
- S. Kralj, O. Bellotto, E. Parisi, A. M. Garcia, D. Iglesias, S. Semeraro, C. Deganutti, P. D’Andrea, A. V. Vargiu, S. Geremia, R. De Zorzi and S. Marchesan, ACS Nano, 2020, 14, 16951–16961 CrossRef CAS PubMed.
- A. M. Garcia, M. Melchionna, O. Bellotto, S. Kralj, S. Semeraro, E. Parisi, D. Iglesias, P. D’Andrea, R. De Zorzi, A. V. Vargiu and S. Marchesan, ACS Nano, 2021, 15, 3015–3025 CrossRef CAS PubMed.
- B. Chatterjee, I. Saha, S. Raghothama, S. Aravinda, R. Rai, N. Shamala and P. Balaram, Chem. - Eur. J., 2008, 14, 6192–6204 CrossRef CAS PubMed.
- I. Saha and N. Shamala, Biopolymers, 2012, 97, 54–64 CrossRef CAS PubMed.
- R. Rai, S. Aravinda, K. Kanagarajadurai, S. Raghothama, N. Shamala and P. Balaram, J. Am. Chem. Soc., 2006, 128, 7916–7928 CrossRef CAS PubMed.
- S. Shaham-Niv, L. Adler-Abramovich, L. Schnaider and E. Gazit, Sci. Adv., 2015, 1, e1500137 CrossRef PubMed.
- P. Singh, S. K. Brar, M. Bajaj, N. Narang, V. S. Mithu, O. P. Katare, N. Wangoo and R. K. Sharma, Mater. Sci. Eng., C, 2017, 72, 590–600 CrossRef CAS PubMed.
- S. Shaham-Niv, P. Rehak, L. Vuković, L. Adler-Abramovich, P. Král and E. Gazit, Isr. J. Chem., 2017, 57, 729–737 CrossRef CAS.
- D. Banik, S. Kundu, P. Banerjee, R. Dutta and N. Sarkar, J. Phys. Chem. B, 2017, 121, 1533–1543 CrossRef CAS PubMed.
- S. Bera, B. Xue, P. Rehak, G. Jacoby, W. Ji, L. J. W. Shimon, R. Beck, P. Král, Y. Cao and E. Gazit, ACS Nano, 2020, 14, 1694–1706 CrossRef CAS PubMed.
- A. K. Mondal, N. Brown, S. Mishra, P. Makam, D. Wing, S. Gilead, Y. Wiesenfeld, G. Leitus, L. J. W. Shimon, R. Carmieli, D. Ehre, G. Kamieniarz, J. Fransson, O. Hod, L. Kronik, E. Gazit and R. Naaman, ACS Nano, 2020, 14, 16624–16633 CrossRef CAS PubMed.
- B. P. Bloom, Y. Paltiel, R. Naaman and D. H. Waldeck, Chem. Rev., 2024, 124, 1950–1991 CrossRef CAS PubMed.
- S. Lin, Q. Tong, P. Jiang, B. Li, Y. Li and Y. Yang, New J. Chem., 2021, 45, 12585–12592 RSC.
- O. Bellotto, G. Pierri, P. Rozhin, M. Polentarutti, S. Kralj, P. D’Andrea, C. Tedesco and S. Marchesan, Org. Biomol. Chem., 2022, 20, 6211–6218 RSC.
- O. Bellotto, S. Kralj, R. D. Zorzi, S. Geremia and S. Marchesan, Soft Matter, 2020, 16, 10151–10157 RSC.
- S. Marchesan, C. D. Easton, K. E. Styan, L. J. Waddington, F. Kushkaki, L. Goodall, K. M. McLean, J. S. Forsythe and P. G. Hartley, Nanoscale, 2014, 6, 5172–5180 RSC.
- S. Marchesan, L. Waddington, C. D. Easton, D. A. Winkler, L. Goodall, J. Forsythe and P. G. Hartley, Nanoscale, 2012, 4, 6752–6760 RSC.
- A. M. Garcia, D. Iglesias, E. Parisi, K. E. Styan, L. J. Waddington, C. Deganutti, R. De Zorzi, M. Grassi, M. Melchionna, A. V. Vargiu and S. Marchesan, Chem, 2018, 4, 1862–1876 CAS.
- S. Marchesan, K. E. Styan, C. D. Easton, L. Waddington and A. V. Vargiu, J. Mater. Chem. B, 2015, 3, 8123–8132 RSC.
- M. C. Mañas-Torres, P. Alletto, S. Adorinni, A. V. Vargiu, L. Á. de Cienfuegos and S. Marchesan, Org. Biomol. Chem., 2025, 23, 2797–2801 RSC.
- C. G. Pappas, P. W. J. M. Frederix, T. Mutasa, S. Fleming, Y. M. Abul-Haija, S. M. Kelly, A. Gachagan, D. Kalafatovic, J. Trevino, R. V. Ulijn and S. Bai, Chem. Commun., 2015, 51, 8465–8468 RSC.
- B. Rosetti, E. Scarel, L. Colomina-Alfaro, S. Adorinni, G. Pierri, O. Bellotto, K. Mamprin, M. Polentarutti, A. Bandiera, C. Tedesco and S. Marchesan, Polymers, 2022, 14, 4554 CrossRef CAS PubMed.
- T. G. Meikle, C. E. Conn, F. Separovic and C. J. Drummond, RSC Adv., 2016, 6, 68685–68694 RSC.
- K. Tatsuta, J. Antibiot., 2013, 66, 107–129 CrossRef CAS PubMed.
- K. Hamamoto, Y. Kida, Y. Zhang, T. Shimizu and K. Kuwano, Microbiol. Immunol., 2002, 46, 741–749 CrossRef CAS PubMed.
- C. Zhong, S. Gou, T. Liu, Y. Zhu, N. Zhu, H. Liu, Y. Zhang, J. Xie, X. Guo and J. Ni, Microb. Pathog., 2020, 139, 103871 CrossRef CAS PubMed.
- H. Wang, Z. Feng and B. Xu, Adv. Drug Delivery Rev., 2017, 110–111, 102–111 CrossRef CAS PubMed.
- Y. Zhou, Y. Zou, M. Yang, S. Mei, X. Liu, H. Han, C.-D. Zhang and M.-M. Niu, J. Am. Chem. Soc., 2022, 144, 7117–7128 CrossRef CAS PubMed.
- S. Muller Benkirane, N. Guichard, G. Regenmortel, M. H. V. Van and F. Brown, Expert Opin. Invest. Drugs, 1998, 7, 1429–1438 CrossRef PubMed.
- Z. Feng and B. Xu, Biomol. Concepts, 2016, 7, 179–187 CrossRef CAS PubMed.
- L. Zhao and W. Lu, Curr. Opin. Chem. Biol., 2014, 22, 56–61 CrossRef CAS PubMed.
- S. Kapil and V. Sharma, Can. J. Microbiol., 2021, 67, 119–137 CrossRef CAS PubMed.
- O. Bellotto, S. Semeraro, A. Bandiera, F. Tramer, N. Pavan and S. Marchesan, Pharmaceutics, 2022, 14, 446 CrossRef CAS PubMed.
- M. Seia and E. Zisman, FASEB J., 1997, 11, 449–456 CrossRef PubMed.
- S. A. Fuchs, R. Berger, L. W. J. Klomp and T. J. de Koning, Mol. Genet. Metab., 2005, 85, 168–180 CrossRef CAS PubMed.
- J. J. A. J. Bastings, H. M. van Eijk, S. W. Olde Damink and S. S. Rensen, Nutrients, 2019, 11, 2205 CrossRef CAS PubMed.
- W. K. Restu, Y. Nishida, S. Yamamoto, J. Ishii and T. Maruyama, Langmuir, 2018, 34, 8065–8074 CrossRef CAS PubMed.
- W. K. Restu, S. Yamamoto, Y. Nishida, H. Ienaga, T. Aoi and T. Maruyama, Mater. Sci. Eng., C, 2020, 111, 110746 CrossRef CAS PubMed.
- A. Agrawal, E. M. Euliano, B. H. Pogostin, M. H. Yu, J. W. R. Swain, J. D. Hartgerink and K. J. McHugh, Cel. Mol. Bioeng., 2024, 17, 441–451 CrossRef CAS PubMed.
- Z. Luo, Q. Wu, C. Yang, H. Wang, T. He, Y. Wang, Z. Wang, H. Chen, X. Li, C. Gong and Z. Yang, Adv. Mater., 2017, 29, 1601776 CrossRef PubMed.
- J. Ding, T. Wang, Z. Lin, Z. Li, J. Yang, F. Li, Y. Rong, X. Chen and C. He, Nat. Commun., 2025, 16, 1222 CrossRef CAS PubMed.
- K. J. Nagy, M. C. Giano, A. Jin, D. J. Pochan and J. P. Schneider, J. Am. Chem. Soc., 2011, 133, 14975–14977 CrossRef CAS PubMed.
- T. Tan, Y. Hou, J. Shi, B. Wang and Y. Zhang, Mater. Today Bio, 2024, 25, 100961 CrossRef CAS PubMed.
- I. J. Duti, J. R. Florian, A. R. Kittel, C. D. Amelung, V. P. Gray, K. J. Lampe and R. A. Letteri, J. Am. Chem. Soc., 2023, 145, 18468–18476 CrossRef CAS PubMed.
- D. Florio, C. Di Natale, P. L. Scognamiglio, M. Leone, S. La Manna, S. Di Somma, P. A. Netti, A. M. Malfitano and D. Marasco, Bioorg. Chem., 2021, 114, 105047 CrossRef CAS PubMed.
- S. Kuila, A. K. Singh, A. Shrivastava, S. Dey, T. Singha, L. Roy, B. Satpati and J. Nanda, J. Phys. Chem. B, 2023, 127, 4808–4819 CrossRef CAS PubMed.
- K. McAulay, B. Dietrich, H. Su, M. T. Scott, S. Rogers, Y. K. Al-Hilaly, H. Cui, L. C. Serpell, A. M. Seddon, E. R. Draper and D. J. Adams, Chem. Sci., 2019, 10, 7801–7806 RSC.
- R. R. Sonani, S. Bianco, B. Dietrich, J. Doutch, E. R. Draper, D. J. Adams and E. H. Egelman, Cell Rep. Phys. Sci., 2024, 5, 101812 CrossRef CAS PubMed.
- L. Wang, X. Jin, L. Ye, A. Zhang, D. Bezuidenhout and Z. Feng, Langmuir, 2017, 33, 13821–13827 CrossRef CAS PubMed.
- M. Li, M. Liu, Y. Shang, C. Ren, J. Liu, H. Jin and Z. Wang, RSC Adv., 2020, 10, 13900–13906 RSC.
- B. Adhikari, C. Singh, A. Shah, A. J. Lough and H.-B. Kraatz, Chem. - Eur. J., 2015, 21, 11560–11572 CrossRef CAS PubMed.
- G. Zhang, L. Zhang, H. Rao, Y. Wang, Q. Li, W. Qi, X. Yang, R. Su and Z. He, J. Colloid Interface Sci., 2020, 577, 388–396 CrossRef CAS PubMed.
- Z. Chen, Z. Lv, G. Qing and T. Sun, J. Mater. Chem. B, 2017, 5, 3163–3171 RSC.
- K. Shao, Z. Lv, Y. Xiong, G. Li, D. Wang, H. Zhang and G. Qing, RSC Adv., 2019, 9, 10360–10363 RSC.
- X. Wang, C. Wei, J.-H. Su, B. He, G.-B. Wen, Y.-W. Lin and Y. Zhang, Angew. Chem., Int. Ed., 2018, 57, 3504–3508 CrossRef CAS PubMed.
- L. Ruckthong, A. F. A. Peacock, C. E. Pascoe, L. Hemmingsen, J. A. Stuckey and V. L. Pecoraro, Chem. - Eur. J., 2017, 23, 8232–8243 CrossRef CAS PubMed.
- G. Zhang, Y. Liang, Y. Wang, Q. Li, W. Qi, W. Zhang, R. Su and Z. He, ACS Nano, 2022, 16, 6866–6877 CrossRef CAS PubMed.
- M. Conejero-Muriel, J. A. Gavira, E. Pineda-Molina, A. Belsom, M. Bradley, M. Moral, J. de, D. G.-L. Durán, A. L. González, J. J. Díaz-Mochón, R. Contreras-Montoya, Á. Martínez-Peragón, J. M. Cuerva and L. Á. de Cienfuegos, Chem. Commun., 2015, 51, 3862–3865 RSC.
- M. Conejero-Muriel, R. Contreras-Montoya, J. J. Díaz-Mochón, L. Á. de Cienfuegos and J. A. Gavira, CrystEngComm, 2015, 17, 8072–8078 RSC.
- D. Gambhir, S. Kumar and R. R. Koner, Soft Matter, 2022, 18, 3624–3637 RSC.
- D. Yang, X. Li, Y.-F. Fan and D.-W. Zhang, J. Am. Chem. Soc., 2005, 127, 7996–7997 CrossRef CAS PubMed.
- A. Tripathi, A. Kumar and P. S. Pandey, Tetrahedron Lett., 2012, 53, 5745–5748 CrossRef CAS.
- F. Zhang, Z. Xu, S. Dong, L. Feng, A. Song, C.-H. Tung and J. Hao, Soft Matter, 2014, 10, 4855–4862 RSC.
- M. C. Mañas-Torres, C. Gila-Vilchez, J. A. González-Vera, F. Conejero-Lara, V. Blanco, J. Manuel Cuerva, M. T. Lopez-Lopez, A. Orte and L. Á. de Cienfuegos, Mater. Chem. Front., 2021, 5, 5452–5462 RSC.
- X. Zhang, Z. Yang, J. Lin, W. Zhou, N. Sun and Y. Jia, Int. J. Mol. Sci., 2025, 26, 3998 CrossRef CAS PubMed.
- F. Auvray, D. Dennetiere, A. Giuliani, F. Jamme, F. Wien, B. Nay, S. Zirah, F. Polack, C. Menneglier, B. Lagarde, J. D. Hirst and M. Réfrégiers, Struct. Dynam., 2019, 6, 054307 CrossRef PubMed.
- Z. Zhang, K. Chen, K. Tang, K. Chen, R. Li, X. Sun, Y. Hu, Q. Liu, M. Chen, H. Yang and X. Chen, Anal. Chem., 2023, 95, 4923–4931 CrossRef CAS PubMed.
- S. Wadhwa, D. Buyens and J. G. Korvink, Adv. Mater., 2024, 36, 2408547 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2025 |





