Preserving structural integrity: fold reproducibility in computational design of proteins non-homologous to wild-type sequences†
Bondeepa
Saikia
 and
Anupaul
Baruah
and
Anupaul
Baruah
 *
*
Department of Chemistry, Dibrugarh University, Dibrugarh-786004, India. E-mail: anupaulbaruah@dibru.ac.in
First published on 12th May 2025
Abstract
Even with remarkable accomplishments, designing a protein with a given structure is still a challenging task. There is no general approach that works for all challenges. Protein sequences with higher sequence similarity are usually shown to have similar three dimensional structures. This work is focused on designing non-homologous protein sequences with low sequence similarity to the wild-type sequence while maintaining secondary structure integrity. Basically, the aim of the present study is to check whether or not dissimilar sequences tend to encode a similar structure. In this work, we employ a negative design approach to design protein sequences by optimizing non-native conformational ensembles. Three non-native conformational ensembles are created for each of the three chosen target structures. During the design of protein sequences using the Monte Carlo simulation method and developed Cα distance-based statistical potentials, these ensembles are destabilized along with stabilization of the targets. The structures of the designed sequences are determined using AlphaFold2. Interestingly, the results suggest that secondary structure elements like alpha helices and beta sheets can be conserved even for non-homologous sequences with low sequence similarity. It is also observed that the designed sequences have the ability to reproduce the three target protein's fold viz. all-α, all-β and mixed αβ despite very low sequence similarity to the wild-type sequences. This indicates that the employed design strategy is effective in preserving structural integrity despite low sequence similarity.
1 Introduction
Computational protein design (CPD) aims to create energetically optimal novel sequences that reliably fold into predefined target structures. CPD is useful in determining the sequence–structure relationships.1–3 Due to its potential to create novel biomolecules with specialized functions like catalysts,4,5 therapeutic proteins,6–9 biosensors10–12etc., this field has gained significant attention. Most structure-based protein design methods aim at achieving a higher level of accuracy in sequence as well as structure recovery rates. However, this remains challenging despite many computational advancements in the field of protein design. However, different graph-based deep-learning methods such as ProteinMPNN,13 AlphaDesign,14 PiFold (protein inverse folding)15etc. have shown to improve the accuracy in sequence recovery.Proteins can be designed employing many approaches viz. positive design, negative design, and machine learning based sequence design. Positive design approaches involve stabilization of the native state only. In the positive design approach the competing non-native states are not explicitly disfavored during the design of protein sequences for the desired target fold. Negative design, on the other hand, chooses sequences that destabilize the competing non-native conformations, guaranteeing that only the target fold is energetically favorable. Machine learning based sequence design methods such as AlphaFold2 based protein design approaches16–18 and diffusion models19 predict sequences for given target structures. However, unlike negative design, these approaches do not specifically account for competing non-native states and instead they primarily rely on evolutionary data. Moreover, a lower success rate in protein design can be attributed to the reliance on the positive design approaches in designing protein sequences. Thus, in order to design protein sequences that are non-homologous to wild-type sequences and are less prone to misfolding, negative design approaches are more effective in designing such sequences.
Most experimentally characterized proteins with similar sequences encode a similar type of structure.20 However, the literature provides proof of many structurally similar proteins with low sequence similarities. Thus, ensuring the designed sequence exhibits low sequence similarity to the wild-type protein sequence while precisely reproducing the desired three dimensional structure is a quite challenging task. This challenge is specifically very crucial for de novo protein design, because de novo protein design21,22 aims to design protein sequences with low resemblance to any existing protein databases. Therefore, this can be accomplished by carefully balancing structural integrity with sequence variation, so that the designed sequences maintain the desired three dimensional structure even while their amino acid composition diverges dramatically.
The ability to reliably reproduce a desired protein fold in novel sequences despite significant sequence variations can be termed as fold reproducibility.23,24 Computational protein design methodologies combine in-depth understanding of protein energetics with specific scoring systems and statistical potentials in designing protein sequences for a given target structure. Over the past two decades, tremendous efforts have been made to develop efficient potentials that can capture the actual protein energetics.25,26 In order to maintain protein fold stability, most protein design methods rely on evaluating inter-residue interactions, side-chain packing and backbone geometry.27,28 Utilization of knowledge-based statistical potentials derived from large datasets of experimentally determined protein structures is a commonly employed approach in the protein design procedure.29–31 Fold reproducibility in design workflows has been improved by recent developments in computational tools, such as machine learning-based techniques32 and models based on AlphaFold.33 These tools have been found to be incorporated into design loops while designing protein sequences.16,17 Despite the advances in machine learning, deep learning-based methods for protein design, it still remains challenging to develop one computational method that can be used to design any protein.
The design of protein sequences that have low sequence similarity to the wild-type sequences while maintaining fold integrity has wide-ranging applications in synthetic biology, drug design, and biotechnology. Additionally, de novo design of proteins broadens the range of strategies available for developing novel biomolecules with specific characteristics not seen in nature. Advances in protein design approaches are opening the door to more robust and adaptable protein designs by solving the combined issues of sequence divergence and structural preservation. Thus, the ability to design protein sequences non-homologous to wild-type sequences will pave the way for a better understanding of sequence–structure relationships and will push the boundaries that can be achieved in the field of protein design.
In our previous work,34,35 the foldability criterion Δ,36 which incorporates negative design features, was found to be an optimal criterion for designing more stable and target compatible protein sequences. Moreover, several other studies37,38 assert that negative design is important for protein design. Inspired by the previous work's results, this study investigates the fold reproducibility ability of the sequences designed using negative design approaches for three structurally classified proteins (an all-α, an all-β and a mixed αβ). Monte Carlo simulation in sequence space is applied to design sequences utilizing the foldability criterion Δ for three selected target structures. The threading method is applied to generate an ensemble of non-native conformations for each selected target protein. A one body and different two body statistical potentials are developed as a function of Cα distances of amino acid residues. These developed potentials are used to assess the stability of a designed sequence in a given conformation. The three dimensional structures of the designed sequences are predicted using AlphaFold2.33 The secondary structures of proteins are typically assigned using the DSSP (define secondary structure of proteins) algorithm.39 Here, we used DSSP3 (3-states-DSSP) to assign the secondary structures of the designed as well as the target proteins. The major goal of this work is to examine whether or not the designed sequences with low sequence similarity to the wild-type sequences encode a similar fold to the target structure. Therefore, designing protein sequences having significant sequence variation will help to assess how sequence mutations affect the overall fold of the protein. Since the method gives non-homologous sequences, it provides a scope to study a sequence space, which may be non-existent in nature and may also fold to existing structures. This opens up a whole new possibility of novel sequences. The results suggest that the designed sequences encode a similar fold to the target protein despite significant sequence variation. The secondary structure comparison between the designed and the target proteins shows that the secondary structure content found in the target proteins is well maintained in all the designed proteins. It is also found that all designed proteins preserve the residue–residue contacts found in the targets to a great extent. Despite significant sequence variations, the ability to design protein sequences that encode folds similar to the target proteins demonstrates the effectiveness of the developed statistical potentials in capturing protein energetics. Again, preservation of secondary structure content found in the target proteins highlights the effectiveness of the potentials in modeling the stabilizing forces that govern local structural motifs. Moreover, residue–residue contact conservation suggests that the developed statistical potentials capture important interactions that are essential for preserving the target proteins' overall fold. The developed statistical potentials capture protein energetics because they are derived from observed frequencies of structural features i.e. residue's Cα distances in 500 known protein structures. Thus, the findings of this work suggest that the developed statistical potentials are able to capture the protein energetics to a certain extent. Therefore, this work suggests that the protein design method such as this one will be quite helpful in understanding the protein sequence–structure relationship that will ultimately lead to the de novo design of protein sequences.
2 Methods
Protein design refers to searching of an amino acid sequence compatible with a desired target structure. The foremost step of any protein design procedure is to chose the target structure, followed by choosing of foldability criteria and most importantly an energy function or potential to quantify the sequence energy in a given conformation.2.1 Target structures
Three experimentally resolved protein structures with PDB ID 6FM8 (chain length = 50, X-ray resolution = 1.78 Å, R-value = 0.270, an all-α protein) (Fig. 1a), 2IGD (chain length = 61, X-ray resolution = 1.10 Å, R-value = 0.125, a mixed αβ protein) (Fig. 1b) and 1HOE (chain length = 74, X-ray resolution = 2.00 Å, R-value = 0.199, an all-β protein) (Fig. 1c) are chosen from the RCSB PDB (Research Collaboratory for Structural Bioinformatics Protein Data Bank). The Cα backbone of these proteins is taken as the target structure for the protein design procedure. Selection of the proteins is based on their secondary structure content, availability of high resolution structure and moderate sequence length.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 881 real-like non-native conformations is generated. Similarly, 18
881 real-like non-native conformations is generated. Similarly, 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 319 non-native conformations of chain length 61 are generated for the target 2IGD and 19
319 non-native conformations of chain length 61 are generated for the target 2IGD and 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 536 non-native conformations of chain length 50 are generated for the target 6FM8. These three sets of conformational ensembles of the proteins 6FM8, 2IGD and 1HOE are considered for protein design procedures. Root mean square deviations (RMSDs) of these three sets of non-native conformational ensembles are determined with respect to respective targets and the mean RMSDs of these three ensembles of non-native conformations are tabulated in Table 1.
536 non-native conformations of chain length 50 are generated for the target 6FM8. These three sets of conformational ensembles of the proteins 6FM8, 2IGD and 1HOE are considered for protein design procedures. Root mean square deviations (RMSDs) of these three sets of non-native conformational ensembles are determined with respect to respective targets and the mean RMSDs of these three ensembles of non-native conformations are tabulated in Table 1.
2.2 Foldability criteria
A well defined foldability criterion is a prerequisite for any protein design procedure. A foldability criterion quantifies the fitness of a sequence to a given conformation in terms of optimized potential or energy function.42–44 In protein design, preventing a protein from adopting non-native or undesirable conformations is the major goal of negative design approaches. With this approach, the native conformation is maintained as the most stable conformation while protein sequences are designed to be energetically unfavorable for alternative conformations. Negative design in terms of stability gap, Δ,36 has proven to be a more effective criterion for designing protein sequences.34,35 Therefore, in this work, we have selected the foldability criterion Δ to design protein sequences. This foldability criterion can be expressed mathematically as| Δ = Ef − 〈E〉u | (1) |
Here, the first term Ef represents energy at the target state and the second term 〈E〉u represents average energy over all non-native conformations.
2.3 Potentials for design
Potentials or energy functions are used in protein design methods to distinguish between sequences that fit the target structure and those that do not. The energy of a sequence in a given conformation can be quantified by using potentials.29 Two statistical potentials are developed in this work and employed in the design of protein sequences. is calculated. Then, the probabilities of a bin of amino acid α for each NNi are calculated as
is calculated. Then, the probabilities of a bin of amino acid α for each NNi are calculated as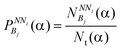 | (2) |
Then, for each NNi and for each bin, Bj, the probabilities 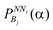 for all 20 amino acids are summed as given in eqn (3) and this is followed by calculation of their averages as given in eqn (4)
for all 20 amino acids are summed as given in eqn (3) and this is followed by calculation of their averages as given in eqn (4)
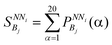 | (3) |
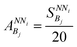 | (4) |
Then the propensity of an amino acid α to be in a particular bin Bj in each NNi can be calculated as
 | (5) |
Finally, this propensity 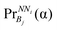 is converted to potential by applying the Boltzmann inversion method45 as follows
is converted to potential by applying the Boltzmann inversion method45 as follows
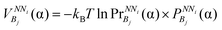 | (6) |
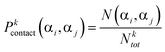 | (7) |
Then, we have determined the unbiased probability of contact formation by unique residue pairs. This is done by first calculating the probability that each amino acid in the data set is in contact. In order to accomplish this, we determine the number of times each amino acid αi is involved in contact formation within the data set by calculating Ncontact(αi). Afterwards, the following formula is used to determine their respective probabilities of contact formation:
 | (8) |
Therefore, the unbiased probability of residues αi,αj forming a contact is computed as
| Pkunbias(αi,αj) = Pkcontact(αi) × Pkcontact(αj) | (9) |
Finally, the 11 two body statistical potentials are obtained by applying the Boltzmann inversion method45 as follows
 | (10) |
2.4 Design of protein sequences
Protein sequences are designed for the target structures 6FM8, 2IGD and 1HOE using Monte Carlo simulation in sequence space. Δ is used as the foldability criterion with a combination of the developed one body and two body statistical potentials for the purpose of protein sequence design. Three Monte Carlo simulations, each of 3 × 104 steps, are performed for the three selected target structures. Each simulation is initiated by generating a random sequence, which is considered as an input for the simulation. For this input sequence the value of foldability criterion Δold is calculated. Then, a random residue site is selected and at that site a point mutation is incorporated randomly. Then, the value of Δnew of this mutated sequence is calculated. The metropolis criterion is utilized to accept or reject the mutated sequence. Using this we have designed 120 sequences for each of the target structures 6FM8, 2IGD and 1HOE.3 Results and discussion
3.1 Potential: distinction between hydrophobic and hydrophilic amino acids
As discussed in Section 2.3.1, a one body statistical potential is developed as a function of residue's Cα distances from neighboring residues for the design of protein sequences. Based on the size, charge, and hydrophobicity an amino acid exhibits distinct structural and energetic preferences. Similarly, different amino acids show specific preferences for either the surface or core environment. These specific preferences of amino acids can be ascertained from their corresponding potential values at specific Cα distances. The lower the value of the potential, the higher the tendency of the amino acid to occupy that specific environment. In soluble, globular proteins hydrophobic amino acids tend to occupy the core of a protein while the hydrophilic amino acids tend to occupy the surface of a protein. When it comes to membrane-integral proteins, this trend differs as hydrophobic residues are frequently surface-exposed to interact with the lipid bilayer. In the context of potential values, hydrophobic amino acid residues exhibit favorable potentials (lower potential value) for shorter Cα distances. This is because amino acid residues found in the core of a protein tend to pack tightly in that environment, while hydrophilic amino acid residues exhibit favorable potentials for longer Cα distances, as they are typically solvent-exposed. In Fig. 2a–f, one body potentials are plotted for some hydrophilic amino acid residues (ARG, LYS, ASN, ASP, GLU and GLN) and some hydrophobic amino acid residues (ALA, CYS, ILE, LEU, MET, PHE, TRP and VAL) for three different nearest number distances represented by NNi, i = 1, 5, and 10. In Fig. 2a–c, it can be observed that the hydrophilic amino acid residues exhibit favorable potentials for longer Cα distances. In Fig. 2d–f the hydrophobic amino acid residues exhibit favorable potentials for shorter Cα distances. A similar trend is observed for all remaining nearest neighbor distances and their corresponding plots of potential vs. bins of Cα distances are incorporated in the ESI† as Fig. S1(a)–(n). It is observed that the most favorable potential values (the steep regions of the curve) for the CYS residue gradually decrease from NN1 to NN4 and then again increase to NN10, with NN4 having the lowest potential value of −0.1136. Since the potential is derived from statistical observations of residue–residue distances in a dataset of proteins compiled from the RCSB PDB, the NN4 might represent the most favorable distance for interactions involving CYS residues. The rise in potential values beyond the NN4 could indicate decreasing interaction frequency or less favorable energetic contributions at larger distances. From these observations, it can be inferred that the developed potential allows the distinction between hydrophobic and hydrophilic amino acid residues.3.2 Determination of sequence similarity
Protein sequences are designed by applying Monte Carlo simulation in sequence space and the developed potentials. As discussed in Section 2.4, 120 sequences are designed for each of the three selected target proteins 6FM8, 2IGD and 1HOE respectively. These designed sequences are taken for determination of sequence similarity with their respective wild-type sequences. This is done by aligning the entire length of two sequences, i.e. the wild-type and the designed sequence, and the total number of exact matches between two aligned sequences is determined. The corresponding percentages of sequence similarity of all the designed sequences are also determined.3.3 Fold reproducibility: three-dimensional structure determination
All 360 sequences (120 sequences for each target) having different sequence similarities designed for the proteins 6FM8, 2IGD and 1HOE are considered for determination of their three-dimensional structures. AlphaFold233 (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/batch/AlphaFold2_batch.ipynb), a artificial intelligence method developed by DeepMind, is used to predict the three-dimensional structure of the designed sequences. Root mean squared deviation (RMSD) is used to compare the target protein structure and the predicted structure of the designed sequence. The RMSDs are calculated using the command “super PDB1, PDB2” in PyMOL.46 The total number of aligned atoms is also determined. A sequence is selected for each target protein 6FM8, 2IGD and 1HOE based on certain criteria as given in Table 2. After subjecting these criteria on the 120 sequences designed for each of the three proteins, the predicted structures of the designed sequences having the highest number of aligned atoms are selected for further analyses. The selected designed proteins with their foldability values are given in Table 2.In Fig. 3a–c, the top-ranked structures of these selected designed sequences predicted by AlphaFold2 are compared with their respective target structures. Here, the top-ranked structure refers to the highest-confidence model having the highest pLDDT values among the five predicted structures generated during the protein structure prediction by AlphaFold2. The full term for pLDDT is “predicted local distance difference test”. This is a per-residue measure of local confidence used by AlphaFold and it assesses the confidence in the local structure by measuring how well the prediction agrees with an experimental structure. The RMSDs and pLDDTs (predicted local distance difference test) are also presented. In Fig. 3a, it is observed that despite very low sequence similarity to the wild-type sequence (8.00%), the sequence having a foldability value of Δ = −8.0582 designed for the target 6FM8 encodes a similar fold to the target 6FM8, which can be confirmed by its RMSD value of 3.59 Å. In Fig. 3b and c, although the RMSDs are observed to be slightly greater, however, we have seen that the predicted structures of the sequences designed for 2IGD and 1HOE assume a similar type of fold i.e. αβ mixed and all-β as the target structures respectively. The sequence similarities of these designed sequences are also very low (Table 2). These results indicate that our developed statistical potential is able to capture the target protein characteristics in the designed sequences despite significant sequence variation. A sequence's ability to reproduce the target protein folds infers that structural stability of a protein is more dependent on the fold than on the sequence. This inference is of utmost value in protein design, where structural or functional characteristics are desired without the need for significant sequence conservation.
 | ||
| Fig. 3 The three-dimensional structures of the target protein (colored green) and the top-ranked predicted structure of the designed protein (colored cyan) and their corresponding RMSDs and pLDDTs. | ||
3.4 Secondary structure comparison
To get a better picture of the fold reproducibility ability of the designed sequences, the predicted structures of the designed sequences are compared with their respective target protein structure in the context of secondary structure contents. This is done to check whether or not the predicted structures of the designed sequences maintain the structural integrity in terms of secondary structure similarity. The same set of sequences having foldability values Δ = −8.0582, Δ = −10.8426 and Δ = −16.5262 designed for proteins 6FM8, 2IGD and 1HOE are considered for this analysis. The secondary structures of the top-ranked predicted structures of the designed sequences as well as the target proteins are assigned using DSSP3 (3-states-DSSP). DSSP3 is a reduced representation of DSSP8 where eight types of secondary structures are assigned to amino acids. While in DSSP3, three types of secondary structures are assigned to amino acids: helix (H), strand (E), and loop (C). We have used the general reduction of 8 types of secondary structures to 3 types: (H/G/I → H, E/B → E, S/T/C → C). Fig. 4a–c are the secondary structure comparison maps between the predicted structures of the designed sequences and their target structures 6FM8, 2IGD and 1HOE respectively. The percentages of secondary structure similarities are calculated for each case and are tabulated in Table 3. We have observed that the secondary structures found in the target proteins are well maintained in all the three designed sequences. It is also observed that the percentage of secondary structure similarity obtained for the sequence designed for all-α protein 6FM8 is the highest (78%). This indicates that the design approach used in this work is well-suited for all-α proteins. Nevertheless, it is also observed that the designed protein precisely captures the helical and most of the β portions of the αβ mixed protein 2IGD, and the percentage of secondary structure similarity is found to be 68.85%. This result highlights the versatility of the design approach. The versatility of the design approach is further strengthened by the percentage of secondary structure similarity of 43.24% for the sequence designed for the all-β protein 1HOE. Overall, it is seen that our employed design approach is effective in designing protein sequences that retain structural integrity despite having very low sequence similarity to the wild-type sequences. The relatively low value of secondary structure similarity for the sequence designed for all-β protein 1HOE suggests that beta-sheet-dominated structures are less tolerant to sequence changes than all-α or α dominant proteins. | ||
| Fig. 4 Secondary structure comparison maps of the designed proteins with their respective target proteins, (a) 6FM8, (b) 2IGD, and (c) 1HOE. | ||
 | (11) |
 | ||
| Fig. 5 Contact maps of the target protein, designed protein (sequence_1) and their comparison within a distance of 7.5 Å: (a) 6FM8, (b) 2IGD, and (c) 1HOE. In the comparison map, the contacts found in the target are colored red, the contacts found only in the designed protein are colored blue and the rose colored contacts represent the contacts present in both structures. | ||
 | ||
| Fig. 6 Contact maps of the target protein, designed protein (sequence_1) and their comparison within a range of distance of 7.5 Å to 15 Å, (a) 6FM8, (b) 2IGD, and (c) 1HOE. In the comparison map, the contacts found in the target are colored red, the contacts found only in the designed protein are colored blue and the rose colored contacts represent the contacts present in both structures. | ||
A local contact map comparison is also done to check to what extent the designed proteins preserve local contacts found in the target proteins. Local contacts are related to the stability and formation of secondary structures like α-helices and β-sheets. Thus, a local contact map comparison helps to determine if designed proteins maintain the secondary structure elements of the target proteins, which is important for fold stability. For this purpose, contacts between residues that are 3 to 7 residues apart along the sequence are calculated within a cut off distance of 7.5 Å and their corresponding sensitivities (Table 5) are also determined using eqn (11). This is done for all the target and the designed proteins. Then the contact maps for the target proteins 6FM8, 2IGD and 1HOE and designed proteins and their comparison for all the designed and their target structures are plotted and presented in Fig. 7a–c. Since sensitivity measures the true positive rate, the sensitivity percentages are calculated for comparing the local contact maps of the target proteins with the designed proteins and are tabulated in Table 5. Sufficiently high sensitivity percentages are observed for the all-α protein 6FM8 (81.70%) and αβ mixed protein 2IGD (70.45%). This indicates that the designed sequences also preserve the local contacts found in their respective target proteins. However, it is found to be relatively low for the sequence designed for all-β protein 1HOE (33.87%).
 | ||
| Fig. 7 Local contact maps of the target protein, designed protein (sequence_1) and their comparison within a distance of 7.5 Å, (a) 6FM8, (b) 2IGD, and (c) 1HOE. In the comparison map, the contacts found in the target are colored red, the contacts found only in the designed protein are colored blue and the rose colored contacts represent the contacts present in both structures. | ||
High sensitivity percentages are observed in the contact map comparisons for the all-α protein 6FM8 and αβ mixed protein 2IGD in contrast to the relatively low sensitivity for all-β protein 1HOE, whether it is short-range or long-range or local residue–residue contacts. This suggests that different folds exhibit varying degrees of designability and structural flexibility. All-α and α dominant αβ mixed proteins are able to reproduce most of the residue–residue contacts because of their ability to tolerate sequence variations while maintaining structural integrity.
3.5 Ramachandran plot analysis
A Ramachandran plot is a two dimensional graphical representation of backbone dihedral angles, ϕ and ψ, of amino acid residues in proteins. It can be used to analyze whether ϕ and ψ angles in the designed protein are realistic and consistent with the target proteins. Similar ϕ and ψ angle distribution in the designed protein to that of the target protein suggests that the backbone structure is well preserved in the designed protein. Thus, Ramachandran plots are constructed for all designed as well as target proteins using the data obtained from the MolProbity web server47 and are presented in Fig. 8a–f. Fig. 8d and f exhibit a similar pattern of Ramachandran plot occupancy as in Fig. 8a–c. This means all the designed proteins exhibit similar ϕ and ψ dihedral angle distribution to that of the target proteins. This ensures that the target protein's α-helical and β-sheet portions are well-captured by all the designed proteins. All the designed proteins have most of the amino acid residues in the Ramachandran favored and allowed regions. This result assures that the backbone dihedral angles are well preserved and consistent with those of stable target protein structures. | ||
| Fig. 8 Ramachandran plots for the target and the designed proteins (sequence_1) along with the percentage of residues in the favored and allowed regions. | ||
4 Conclusions
Protein sequences with very low sequence similarity to the wild-type sequences are designed for three target structures: an all-α protein 6FM8, a mixed αβ protein 2IGD, and an all-β protein 1HOE using Monte Carlo simulation in sequence space and negative design approaches. A one body and different two body statistical potentials are developed as a function of Cα distances of amino acid residues and are used to assess the stability of the designed sequence in a given conformation. AlphaFold2 is used to predict the three-dimensional structures of the designed sequences. The results suggest that despite being non-homologous (low sequence similarity) to wild-type protein sequences, the designed protein sequences encode a similar fold to the target. The fold reproducibility ability of the designed sequences can be evaluated by analyzing how well the designed sequences preserve the secondary structure elements like α-helix, β-sheet, coil etc. Using DSSP3, the secondary structures of the target and AlphaFold2 top-ranked predicted structures of the designed sequences are determined. The results ensure that the structural integrity in terms of secondary structure similarity is well preserved in all the designed proteins. Contact maps are plotted for short-range and long-range contacts and are compared based on sensitivity and specificity. The results obtained from comparison of contact maps ensure that the sequence designed for the all-α protein captures both short-range and long-range residue–residue contacts found in the target protein. In the case of the sequence designed all-β protein, the sensitivity for short-range contacts is found to be relatively low as compared to sensitivities of all-α and αβ mixed proteins. However, the sensitivity for long-range contacts is found to be high for all designed proteins. This indicates that the design approach might be more promising in designing protein sequences for all-α and for α dominant αβ mixed proteins as compared to β dominant protein folds. However, the design approach efficiently captures the residue–residue short-range and long-range contacts that are absent in both the designed and target proteins very well. Comparison of local contact maps produces similar results. A comparison of our method with RosettaDesign48 is also carried out by designing three sequences for each target structure and is incorporated in the ESI† as Table S1 and Fig. S2–S4.Although the structures of the designed proteins show good performance, the RMSDs and other results indicate some deviations. This deviation seems to be notable in the case of the all-β protein 1HOE. It is evident that all-β proteins frequently possess complex topologies with long loops and are structurally less regular.26,49,50 This makes it challenging to design all-β proteins accurately than all-α and mixed αβ proteins, leading to deviations in RMSD and pLDDT values. Again, the sensitivity percentage for the residue–residue contacts within the distance of 7.5 Å for the designed all-β protein is found to be relatively less than all-α and mixed αβ proteins. In all-β proteins, short-range interactions mostly rely on exact hydrogen bonding patterns and are challenging to reproduce without atomic-level precision in design.51 Since the design method utilized here is a coarse-grained method, there are some deviations in sensitivity for short range contacts despite the good overall performance of the designed proteins. These observed deviations may arise from methodological limitations in developed statistical potentials derived from the data set of proteins that may not fully capture the intricate physics of protein folding. The dependence on only the Cα model restricts the depiction of atomic interactions and side-chain specificity, even though the statistical potential is good at capturing coarse-grained energetic preferences. Thus, a design approach that incorporates all-atom modeling with improved potential that is capable of accurately modeling alpha helices, beta sheets, loop regions will enhance the designability and reliability of designed protein structures.
The comparison of Ramachandran plots of the designed and the target proteins exhibits a similar pattern of Ramachandran plot occupancy. This reflects that the target protein's α-helical and β-sheet portions are well captured by all the designed proteins. It is also observed that, in all the designed proteins, most of the backbone dihedral angles are well preserved and consistent with those of stable target protein structures. Thus, the findings of this work suggest that the secondary structural elements like alpha helices and beta sheets can be conserved even for non-homologous sequences with very low sequence similarity. Since the method gives non-homologous sequences, it provides a scope to study a sequence space, which may be non-existent in nature and may also fold to existing structures. This opens up a whole new possibility of novel sequences. The ability to design protein sequences with very low sequence similarity to wild-type sequences will pave the way of a better understanding of sequence–structure relationships. Therefore, a protein design method such as this one will be quite helpful in de novo design of protein sequences in the future.
Author contributions
Bondeepa Saikia: writing – original draft, conceptualization, methodology, software, formal analysis and Anupaul Baruah: writing – reviewing and editing, supervision, funding acquisition, conceptualization.Data availability
The data that support the findings of this study are available within this article and its ESI.† The 3989 proteins are compiled from the RCSB PDB. The 556 proteins that are taken for generation of non-native conformations are provided in the article's ESI† as Table S2. The in-house codes used in this study are available at https://github.com/ABLab2018/Fortran-codes.Conflicts of interest
There are no conflicts to declare.Acknowledgements
The authors gratefully acknowledge DST, India PURSE (Project No. SR/PURSE/2022/143). The authors acknowledge the financial support from DST, India (Project No. CRG/2023/008126). The financial assistance of the DST-FIST and UGC-SAP program to the Department of Chemistry, Dibrugarh University is also gratefully acknowledged.References
- J. G. Saven, Curr. Opin. Colloid Interface Sci., 2010, 15, 13–17 CrossRef CAS PubMed.
- I. Samish, C. M. MacDermaid, J. M. Perez-Aguilar and J. G. Saven, Annu. Rev. Phys. Chem., 2011, 62, 129–149 CrossRef CAS PubMed.
- E. Michael, S. Polydorides, T. Simonson and G. Archontis, J. Chem. Phys., 2020, 153, 054113 CrossRef CAS PubMed.
- D. N. Bolon and S. L. Mayo, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 14274–14279 CrossRef CAS PubMed.
- J. Kaplan and W. DeGrado, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, 11566–11570 CrossRef CAS PubMed.
- B. Leader, Q. J. Baca and D. E. Golan, Nat. Rev. Drug Discovery, 2008, 7, 21–39 CrossRef CAS PubMed.
- A. Chevalier, D.-A. Silva, G. J. Rocklin, D. R. Hicks, R. Vergara, P. Murapa, S. M. Bernard, L. Zhang, K.-H. Lam and G. Yao, et al. , Nature, 2017, 550, 74–79 CrossRef CAS PubMed.
- L. Cao, I. Goreshnik, B. Coventry, J. B. Case, L. Miller, L. Kozodoy, R. E. Chen, L. Carter, A. C. Walls and Y.-J. Park, et al. , Science, 2020, 370, 426–431 CrossRef CAS PubMed.
- S. B. Ebrahimi and D. Samanta, Nat. Commun., 2023, 14, 2411 CrossRef CAS PubMed.
- L. L. Looger, M. A. Dwyer, J. J. Smith and H. W. Hellinga, Nature, 2003, 423, 185–190 CrossRef CAS PubMed.
- C. E. Tinberg, S. D. Khare, J. Dou, L. Doyle, J. W. Nelson, A. Schena, W. Jankowski, C. G. Kalodimos, K. Johnsson and B. L. Stoddard, et al. , Nature, 2013, 501, 212–216 CrossRef CAS PubMed.
- M. J. Bick, P. J. Greisen, K. J. Morey, M. S. Antunes, D. La, B. Sankaran, L. Reymond, K. Johnsson, J. I. Medford and D. Baker, eLife, 2017, 6, e28909 CrossRef PubMed.
- J. Dauparas, I. Anishchenko, N. Bennett, H. Bai, R. J. Ragotte, L. F. Milles, B. I. Wicky, A. Courbet, R. J. de Haas and N. Bethel, et al. , Science, 2022, 378, 49–56 CrossRef CAS PubMed.
- Z. Gao, C. Tan and S. Z. Li, arXiv, 2022, preprint, arXiv:2202.01079 DOI:10.48550/arXiv.2202.01079.
- Z. Gao, C. Tan, P. Chacón and S. Z. Li, arXiv, 2022, preprint, arXiv:2209.12643 DOI:10.48550/arXiv.2209.12643.
- C. A. Goverde, B. Wolf, H. Khakzad, S. Rosset and B. E. Correia, Protein Sci., 2023, 32, e4653 CrossRef CAS PubMed.
- L. Moffat, J. G. Greener and D. T. Jones, bioRxiv, 2021, preprint DOI:10.1101/2021.08.24.457549.
- M. Jendrusch, J. O. Korbel and S. K. Sadiq, bioRxiv, 2021, preprint DOI:10.1101/2021.10.11.463937.
- J. L. Watson, D. Juergens, N. R. Bennett, B. L. Trippe, J. Yim, H. E. Eisenach, W. Ahern, A. J. Borst, R. J. Ragotte and L. F. Milles, et al. , Nature, 2023, 620, 1089–1100 CrossRef CAS PubMed.
- A. Šali, J. Overington, M. Johnson and T. Blundell, Trends Biochem. Sci., 1990, 15, 235–240 CrossRef PubMed.
- P.-S. Huang, S. E. Boyken and D. Baker, Nature, 2016, 537, 320–327 CrossRef CAS PubMed.
- I. V. Korendovych and W. F. DeGrado, Q. Rev. Biophys., 2020, 53, e3 CrossRef CAS PubMed.
- B. Rost, R. Schneider and C. Sander, J. Mol. Biol., 1997, 270, 471–480 CrossRef CAS PubMed.
- D. J. Rigden, From protein structure to function with bioinformatics, Springer, 2009, vol. 355 Search PubMed.
- A. Goldenzweig and S. J. Fleishman, Annu. Rev. Biochem., 2018, 87, 105–129 CrossRef CAS PubMed.
- X. Pan and T. Kortemme, J. Biol. Chem., 2021, 296, 100558 CrossRef CAS PubMed.
- M. M. Gromiha and S. Selvaraj, Prog. Biophys. Mol. Biol., 2004, 86, 235–277 CrossRef CAS PubMed.
- B. Kuhlman and P. Bradley, Nat. Rev. Mol. Cell Biol., 2019, 20, 681–697 CrossRef CAS PubMed.
- J. U. Bowie, R. Lüthy and D. Eisenberg, Science, 1991, 253, 164–170 CrossRef CAS PubMed.
- M. J. Sippl, Curr. Opin. Struct. Biol., 1995, 5, 229–235 CrossRef CAS PubMed.
- S. Kmiecik, D. Gront, M. Kolinski, L. Wieteska, A. E. Dawid and A. Kolinski, Chem. Rev., 2016, 116, 7898–7936 CrossRef CAS PubMed.
- L. Wei and Q. Zou, Int. J. Mol. Sci., 2016, 17, 2118 CrossRef PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Ždek and A. Potapenko, et al. , Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- B. Saikia, C. R. Gogoi, A. Rahman and A. Baruah, J. Chem. Phys., 2021, 155, 144102 CrossRef CAS PubMed.
- B. Saikia and A. Baruah, Soft Matter, 2024, 20, 3283–3298 RSC.
- N. V. Dokholyan and E. I. Shakhnovich, J. Mol. Biol., 2001, 312, 289–307 CrossRef CAS PubMed.
- I. N. Berezovsky, K. B. Zeldovich and E. I. Shakhnovich, PLoS Comput. Biol., 2007, 3, e52 CrossRef PubMed.
- A. Rossi, C. Micheletti, F. Seno and A. Maritan, Biophys. J., 2001, 80, 480–490 CrossRef CAS PubMed.
- W. Kabsch and C. Sander, Biopolymers, 1983, 22, 2577–2637 CrossRef CAS PubMed.
- C. M. Summa, M. M. Rosenblatt, J.-K. Hong, J. D. Lear and W. F. DeGrado, J. Mol. Biol., 2002, 321, 923–938 CrossRef CAS PubMed.
- W. Jin, O. Kambara, H. Sasakawa, A. Tamura and S. Takada, Structure, 2003, 11, 581–590 CrossRef CAS PubMed.
- J. G. Saven, Chem. Rev., 2001, 101, 3113–3130 CrossRef CAS PubMed.
- P. Biswas, J. Zou and J. G. Saven, J. Chem. Phys., 2005, 123, 154908 CrossRef PubMed.
- A. Baruah and P. Biswas, J. Chem. Phys., 2015, 142, 05B608_1 CrossRef PubMed.
- D. Reith, M. Pütz and F. Müller-Plathe, J. Comput. Chem., 2003, 24, 1624–1636 CrossRef CAS PubMed.
- L. Schrödinger, The PyMOL Molecular Graphics System, Version 1.8, Schrödinger, LLC, 2015.
- I. W. Davis, A. Leaver-Fay, V. B. Chen, J. N. Block, G. J. Kapral, X. Wang, L. W. Murray, W. B. Arendall III, J. Snoeyink and J. S. Richardson, et al. , Nucleic Acids Res., 2007, 35, W375–W383 CrossRef PubMed.
- Y. Liu and B. Kuhlman, Nucleic Acids Res., 2006, 34, W235–W238 CrossRef CAS PubMed.
- E. Marcos and D.-A. Silva, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2018, 8, e1374 Search PubMed.
- Y. Zhang, Y. Liu, Z. Ma, M. Li, C. Xu and H. Gong, bioRxiv, 2024, preprint DOI:10.1101/2024.10.05.616664.
- D. L. Minor Jr and P. S. Kim, Nature, 1994, 367, 660–663 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d5cp01373a |
| This journal is © the Owner Societies 2025 |


