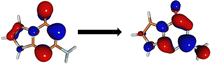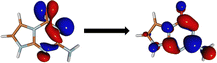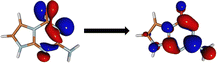Efficient photodeactivation mechanism in an unnatural nucleic acid base†
Somsuta
Ray
 and
Debashree
Ghosh
and
Debashree
Ghosh
 *
*
School of Chemical Sciences, Indian Association for the Cultivation of Science, Kolkata 700032, India. E-mail: pcdg@iacs.res.in; Fax: +91 (0)33 2473 2805; Tel: +91 (0)33 2473 4971
First published on 12th May 2025
Abstract
5-aza-7-deazaguanine (5N7C-G), also known as P, is an unnatural nucleic acid base (NAB) closely related in structure to the natural NAB guanine. It forms a Watson–Crick base pair with the unnatural NAB Z synthesized by Benner and co-workers. We study the ultrafast decay pathways for this modified NAB with high-level multireference methods. We observe from both static and dynamic studies that there are multiple deactivation channels with significant differences in energetics and, therefore, timescales. The nonradiative deactivation mechanisms at sub-picosecond timescales are remarkably similar to those of the natural NAB guanine. These findings explain the experimental observations of Krul et al. [Krul et al., Photochem. Photobiol., 2023, 99, 693–705].
1. Introduction
Deoxyribonucleic acid (DNA) contains all the genetic information in an organism. The genetic alphabet in an organism is written with the help of 4 letters or nucleic acid bases, namely-adenine (A), thymine (T), cytosine (C), and guanine (G).1 In RNA, thymine is replaced by uracil. Intermolecular hydrogen bonding between these nucleobases contributes to the stability of the double helical structure of DNA.2,3 Adenine forms a base pair with thymine via two hydrogen bonds, and cytosine pairs with guanine via three hydrogen bonds. To understand the carcinogenic effects of UV radiation in DNA, the photophysics of nucleic acid bases (NABs) have been studied. The mechanism of self-repair and ultrafast non-radiative decay in these molecules make them resistant to damage by solar radiation.4–7However, in recent decades, significant effort has been made to expand the genetic alphabet by artificially synthesizing nucleobases that are stable in the DNA structure and can be replicated efficiently with minimal mispairing.8–10 The idea was initially started by Alexander Rich in 1962.11 It was proposed that by incorporating small changes in the structures of the natural nucleobases, the genetic alphabet could be increased. However, the creation and incorporation of artificial or unnatural base pairs into DNA can present several challenges, such as chemical stability, replication fidelity, etc. Despite these challenges, the expansion of the genetic code is a promising field of research.12 One can expect the synthesis of new amino acids and proteins13 leading to the creation of semi-synthetic organisms.14,15 Of late, DNA based approaches are also being used to store data and expansion of the genetic code will increase the ability to store large amount of data more affordably. In 1989, Steven A. Benner and co-workers synthesized the IsoG-IsoC base pair.16 Due to keto–enol tautomerization in isoG (which led to the formation of enol isoG, which eventually mispaired with thymine17), it showed 98% selectivity per replication in PCR,18 and further improvements were required in this base pair. A thio-derivative of thymine was synthesized, and it was expected to reduce the stability of the enol IsoG-T mispair.19 In 2007, they successfully synthesized the P–Z base pair.20 The structure of the P-base is shown in Fig. 1. Its IUPAC nomenclature is 2-aminoimidazo[1,2-a][1,3,5]triazin-4(1H)-one. It has significant structural similarity with guanine and can be found by replacing the 5-C with N and 7-N with C. Therefore, this base is also referred to as 5-aza-7-deazaguanine (5N7C-G). 5N7C-G does not show mispairing due to the absence of keto–enol tautomerization, and the base pair showed improved selectivity (>99%).21
The stability of the nucleobase pair comes from not only hydrogen bonding but also other non-covalent interactions such as van der Waals forces, shape fitting, etc.9,22 This led to the development of Ds-Pa and 5SICS-NaM unnatural base pairs by the Hirao and Romesberg groups, respectively.14,23 Theoretical studies have been conducted on these non-covalently stabilized unnatural base pairs.12,24,25 However, for the best performance of these unnatural NABs (uNABs), along with these ground state properties, they should also be stable upon irradiation.
Here, it should be mentioned that the photoprocesses in natural NABs have been extensively studied over the last few decades. Efficient ultrafast nonradiative decay channels have been identified for all the NABs from both experimental and theoretical studies. Ring puckering and nonplanarity induced conical intersections between the ground and lowest excited states form the basis of these nonradiative channels.26–28 However, similar studies on unnatural NABs are few, and therefore, the photoactivities of these molecules are relatively unknown. Understanding these processes is expected to be crucial to estimate their photostability and provide design principles to improve their robustness.
In a recent study from our group, we found that the excited state processes in nucleobase Z (the complementary base of P) are governed by the rotation of the electron-withdrawing nitro group,29 contradictory to its natural counterpart cytosine30 or any natural NAB that decays to the ground state non-radiatively via ring puckering.26 In another study, the P–Z base pair was investigated, and it was concluded that photo-decay depends significantly on the nitro group.31 These results give us a hint that the P base has less contribution toward the photo-decay of the base pair as a whole. Also, since it has a structure very similar to that of its natural analog, guanine (P is also known as 5-aza-7-deazaguanine), its deactivation pathway is not expected to be much different from that of guanine.32–36 Hence, we have elucidated the entire photo-deactivation pathway of the P molecule, involving the low-lying singlet and triplet states.
2. Computational details
The geometry of the ground state molecule is optimized using the MP2/cc-pVDZ, ωB97x-D/cc-pVDZ and ωB97x-D/6-31+G(d) levels of theory. The optimized geometries are similar (given in the ESI†), and we have used the MP2/cc-pVDZ optimized geometry for further calculations.The vertical excitation energies of the optimized geometry were calculated using EOM-CCSD/cc-pVDZ,37,38 TD-CAM-B3LYP/cc-pVDZ,39 SOS-CIS(D)/cc-pVDZ40 and CASPT2/cc-pVDZ.41,42 The effect of the basis set is also determined at the EOM-CCSD level by comparing the excited states at the basis sets cc-pVDZ, 6-311++G(d,p), 6-31G(d,p) and 6-31G. Calculations are done with the Q-Chem 5.1 quantum chemistry package.43
To understand the molecule away from the Franck–Condon (FC) region, one needs to employ multi-reference approaches. Complete active space self-consistent field (CASSCF) was used to incorporate the static correlation and second-order perturbation at the CASPT2 level was used to include dynamic correlation. For CASSCF calculations, a four state averaged calculation was performed for singlet and triplet states. The dynamic correlation was incorporated by CASPT2 (rs2c) calculation with 0.2 a.u. level shift. An active space of 12 orbitals and 16 electrons at cc-pVDZ basis was used. The active space comprises five π orbitals, four π* orbitals, one lone pair on the N of the triazine ring, and two nonbonding orbitals of σ symmetry (given in the ESI†). The singlet and triplet-optimized CASSCF orbitals in the active space are also shown in the ESI.† The active space-based calculations are performed with Molpro-201544 and Molpro-2024 software.45
The optimized geometries at the excited states (singlet and triplet) are calculated along with conical intersections, minimum energy crossing points, and singlet–triplet crossing points at the SA(4)-CASSCF(12o,16e)/cc-pVDZ level of theory. With the help of these stationary points, the potential energy surface is reconstructed with linearly interpolated internal coordinates (LIIC). The minimum energy pathways are then constructed to estimate the energy barriers along those pathways. The pathways are calculated at state-specific CASPT2(12o,16e)/cc-pVDZ. At the singlet–triplet crossing points, spin orbit couplings (SOCs) are calculated at CAM-B3LYP/cc-pVDZ.
Dynamic calculations with surface hopping46 and the CASSCF level of theory are performed at the interface of SHARC 3.047 and Molpro-201248 to ascertain the fate of the excited state molecule and calculate the timescales of the nonradiative pathways. Fewest switches surface hopping (FSSH) dynamics is performed for 37 trajectories with the CASSCF/6-31G level of theory. Initial conditions are obtained using the Wigner distribution along the normal modes calculated at the MP2/6-31G level of theory. A reduced active space (6o,8e) was used for SA-CASSCF/surface hopping. In this reduced active space, there are three π orbitals, two π* orbitals, and one non-bonding orbital of σ symmetry. These orbitals were retained since the static calculations showed that they were predominantly responsible for the lowest-lying excited states. The orbital transitions involved in the first bright state (S1) and the lowest four triplet states (T1–T4) are included in the ESI.† The time step used for generating the trajectories was 0.5 fs, while the trajectories were run for a total time of 500 fs.
3. Results and discussion
3.1. Vertical excitation energies
The ground state optimized geometries at the MP2/cc-pVDZ, wB97x-D/cc-pVDZ, and wB97x-D/6-31+g(d) levels of theory are quite similar (shown in ESI†) and the MP2/cc-pVDZ optimized geometry that is used for the later calculations is shown in Fig. 2. The molecule is mostly planar with non-planarity at the amine group.The vertical excitation energies (VEEs) of the singlet and triplet states at different levels of theory are given in Table 1. The nature of the excited states and the most important orbitals involved are also shown, along with the oscillator strengths.
The lowest singlet state is an optically bright π–π* state, which involves a HOMO → LUMO excitation. This is akin to the La excited state in guanine.33,35 The experimental absorption of 5N7C-G shows a maximum at 255 nm or 4.86 eV.49 This is in good agreement with our calculated value of S1 (4.62 eV or 268 nm) at the CASPT2/cc-pVDZ level of theory. However, unlike in the case of guanine, the next two excited states are both optically dark and predominantly n–π* in nature. It should be noted that at the SOS-CIS(D) level of theory, the S3 state is also π–π* in nature with significant oscillator strength. The effect of the basis set is also determined at the EOM-CCSD level of theory (shown in ESI†). It is observed that while different levels of theory and basis sets can show quantitative differences in excitation energies, the qualitative ordering and nature of the excited states are similar at these different levels of theory.
There are several low-lying triplet states below or near the S1 state. They are all formed by excitation in the π and π* manifold. The T1 state has similar orbitals that are involved in the formation of the S1 state and lies significantly below the S1 state energy due to large exchange correlation. The T2, T3 and T4 states are very closely spaced, lying close to the S1 state and are near degenerate at the CASPT2 level of theory.
3.2. Stationary points and excited state manifold
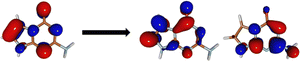
The minima at the low-lying singlet and triplet excited states are shown in Fig. 3a and b. The S1 minimum undergoes a half-chair type ring puckering with a significant angle between the fused imidazole and triazine rings (34.15°). There are more changes to the bond lengths in the triazine ring as opposed to the imidazole ring (shown in ESI†). The planarity of the rings is broken mainly due to the reduced aromaticity of the triazine ring, which leads to its puckering and movement out of plane. The T1 minimum, on the other hand, is very close in geometry to the Franck–Condon region and is almost planar.We obtained two CIs between the S1 and S0 states, referred to as CI-1 and CI-2 (shown in Fig. 3c and d). CI-1 is energetically much more favorable than CI-2 (61.68 kcal mol−1 lower in energy). The CI-1 geometry is similar to the S1 minimum with a chair-type ring puckering. Here, the angle between the imidazole and triazine rings is 42.16°. The CI-2 shows a small chair-type distortion and, therefore, a small amount of non-planarity in the rings. However, here, the major difference from the Franck–Condon region is the large out-of-plane degree of freedom of the amine group (62.0°).
Two STCs are obtained between the S1 and T1 states (shown in Fig. 3e and f), referred to as S1/T1 STC-1 and STC-2. The STC-2 is lower in energy than the STC-1 by 2.33 kcal mol−1. The former is similar in geometry to the CI-1 geometry. The half-chair conformation leads to a non-planarity of 39.16° between the two fused rings. The STC-2 geometry is a twist boat-type ring puckered structure. At the Franck–Condon region, the S1 state is in near degeneracy with the T2, T3 and T4 states, and therefore, the STCs between these states were not explicitly calculated. The FC geometry is considered as the STC between the S1 and these triplet states.
Two STC points between the S0 and T1 geometry are obtained (shown in Fig. 3g and h). The STC-1 has a chair-type conformation; however, here, the angle between the fused rings is even more (43.30°). On the other hand, STC-2 shows both chair and twist boat-type deformation. The energy difference between these STCs is 4.34 kcal mol−1.
LIIC is employed along the twist angles with these stationary points to recreate the excited state (singlet and triplet) manifold along the important degrees of freedom. Here, it is important to note that STC-1 and CI-1 have marked similarities in their geometries. Minimum energy paths along these coordinates are constructed to estimate the energy barriers. The excited state manifold in this reaction coordinate-1 is shown in the right half of Fig. 4a. As can be noticed, CI-1 is energetically favorable and has a minimum energy path with minimal barriers with respect to the FC energy. The S2 and S3 states are significantly higher in energy than the S1 state at the FC region and continue to be so for the reaction coordinate-1, i.e., the CI with a chair-type deformation. The same profile along the reaction coordinate-2 is shown in the left half of Fig. 4a. CI-2 between S0–S1 is energetically unfavorable and, therefore, involves a significant energy barrier. On the other hand, there is a crossover between the S1 and S2 states (along reaction coordinate-2), and it is the S2 state that can lead to a conical intersection or near degeneracy at lower energies than the previously mentioned CI-2. It should be noted that the energy of this crossing is also quite higher than the FC region, and therefore, this pathway is expected to be the minor channel.
 | ||
| Fig. 4 Minimum energy pathways along the excited state manifold for the nonradiative decay channels: (a) singlet pathway, and (b) singlet–triplet pathway. | ||
Here, we should note some similarities and significant differences between our results and those obtained by Crespo-Hernandez and co-workers.49 The S1 minimum geometry that we obtain is in good agreement with their results. However, unlike this previous study we have obtained two CIs that are significantly different from each other.
The CI-2 in our study is similar to the CI obtained in ref. 49. The small differences in the geometry between these two CIs are in the chair-like deformation that we observe. Both the studies have found that the amine group is out of plane. There are also differences in the energetics. The barrier from the S1 minimum that Crespo-Hernandez and co-workers obtain is 19 kcal mol−1 while we notice a barrier of 48 kcal mol−1 for the same. The conical intersection calculated by Crespo-Hernandez and co-workers is a planar ring structure with the amine group perpendicular to it. It shows a barrier height of 19 kcal mol−1. The CI-2 we have calculated also shows out-of-plane movement of the amine group but some amount of ring puckering is also present. This makes CI-2 a more distorted geometry compared to their conical intersection. So CI-2 shows a larger barrier height (48 kcal mol−1). When compared to the energy at the FC region, their CI is almost similar in energy while our CI-2 is quite high in energy. However, due to the barriers from the S1 minimum, our CI-2 (and the CI obtained in ref. 49) are energetically unfavorable. Furthermore, in their study on continuum and explicit solvent effects this barrier does not reduce significantly. This points to this CI being energetically unfavorable in realistic environments. We, therefore, do not expect this to be a major nonradiative decay channel.
We have, however, obtained another CI, denoted as CI-1, which is significantly different in geometry and quite energetically favorable. In this CI, there is puckering of the imidazole ring and this CI along with the pathway from the FC region is barrierless. Given the difference in the energetics in the two CIs, we expect this CI to be the major pathway in nonradiative decay processes.
The triplet-mediated pathways are shown in Fig. 4b. The right panel shows the manifold along the half-chair configuration STC-1. The S1/T1 STC-1 is 12.57 kcal mol−1 higher in energy than the FC region. The T1/S0 STC-1 is an STC at the CASSCF level of theory but shows an energy gap between the states at the CASPT2 level of theory, as shown in Fig. 4b. However, this pathway is expected to be unfeasible due to a large barrier to the initial STC point (S1/T1 STC-1). Furthermore, the SOC values calculated at these STCs are small (16 cm−1 and 18 cm−1, respectively). In the left panel, the triplet pathway towards the twist boat puckered structure is shown. This shows that the T1/S0 STC-2 is similar in energy to the FC region, but S1/T1 STC-2 remains energetically unfavorable (10.24 kcal mol−1 higher than the FC region).
We further note that the S1 state is near degenerate with the T2, T3 and T4 states near the FC region. Therefore, SOC values are calculated between these states, and for all cases, small values are obtained. Details of the SOC values are included in the ESI.† The topology near the CIs is also calculated by estimating the tilt, pitch, etc. measures50 and is given in the ESI.†
3.3. Dynamic studies
Fewest switches surface hopping (FSSH) dynamics is performed for 37 trajectories with the CASSCF/6-31G level of theory. Wigner distribution was used to generate the starting geometries of these structures. For the trajectories, a 0.5 fs time step was used, and it was propagated for 500 fs as described in the previous section. 35% of the trajectories deactivated into the ground S0 state within 500 fs. The population of the states S0 and S1 only for singlet-mediated trajectories is shown in Fig. 5a. Fig. 5b shows the population when both singlet and triplet mediated trajectories are allowed. As expected from static potential energy calculations, triplet-mediated pathways are relatively less probable and are not important. | ||
| Fig. 5 Surface hopping dynamics: population vs. time (fs) plots involving: (a) singlet states, and (b) singlet and triplet states. | ||
The geometries from which surface hopping is observed can be classified into 2 types. The CI-1 type of geometry is the predominant hopping geometry (70%), while the CI-2 type is observed for only 3 trajectories. Representative trajectories of these two types are shown in the ESI.† Furthermore, the timescales for these hopping geometries are quite different – ≈120 fs is the average time taken for the hops with the CI-1 type of geometry and ≈285 fs is the average time taken to reach the CI-2 type geometry for a trajectory to hop from the S1 to S0 state. Here, it should be noted that experimental results also show two or more different timescales for these deactivation processes,51 and they might correspond to these different classes of events.
3.4. Discussions
As expected, guanine32–36 and 5N7C-G show significant similarities in their excited state processes. The most noteworthy of these similarities is that both these molecules show multiple low-lying CIs and at least one of these CIs provides a barrierless or near barrierless pathway. The two CIs obtained by Borin and co-workers for guanine33 show significant similarities with 5N7C-G – one CI with significant out of plane movement of the amine group, and the other CI with ring puckering. The difference is in the kind of puckering observed in guanine versus 5N7C-G. In guanine, an envelope-like puckering of both the rings is observed33,34,36 while in 5N7C-G there is a chair-like puckering where the imidazole ring moves out of plane.Another difference between these two molecules is that in guanine the CI with out-of-plane amine is significantly lower in energy than the ring puckered CI. It is important to note that in 5N7C-G, the out-of-plane amine CI is higher in energy than the ring puckered CI. However, this does not affect the final pathway of guanine as noted in ref. 32. They have shown that the two CIs fall in the same seam, but when the MEP is constructed between the FC and the CI regions, the path of steepest descent takes the molecule to the ring puckered CI. In the case of the out-of-plane amine CI (although the CI is lower in energy), there is a barrier that must be climbed to reach this CI. Therefore, the nonradiative deactivation in guanine is expected to follow the ring puckered CI and here the pathway is completely downhill. The degree of downhill nature of the MEP in guanine is probably steeper than what we observed in 5N7C-G.
From the existing literature on spectroscopic measurements on guanine and 5N7C-G, there are some similarities in their behavior. Both these species have been known to exhibit multiple timescales of nonradiative decay. While the timescales measured for guanine are 0.6, 0.7, and 2.7 ps,52 those for 5N7C-G are 0.3, 0.8, and 18.1 ps in phosphate buffer.49 Therefore, at sub-ps, these species have comparable timescales of deactivation processes. The long timescale process might be markedly different. In our theoretical work, we can mainly comment on the shorter timescale phenomena and therefore, we try to compare those with the experimental observations.
The average timescales for surface hops observed in 5N7C-G are 0.1 ps and 0.3 ps, respectively, which are in good agreement with the experimental observations.49 It is important to note that since we can only attempt to run a few trajectories, the correct statistical estimate is difficult to obtain. Furthermore, we observe that these different timescales are due to entirely different pathways taken by the trajectories. The major pathway is via CI-1, and the minor pathway is via the almost planar (with out-of-plane amine) CI-2. Along the latter pathway, there might be degeneracies with higher singlet states, which is similar to the observations in guanine. It is important to note that Thiel and co-workers have observed two different types of geometries for surface hops in guanine.34 They observe a low timescale (150 fs) surface hop from puckered geometry and higher timescale (>300 fs) for out of plane amine geometry. The longer timescales could not be observed in most of these studies as well as ours due to computational limitations, i.e., inability to run more than ps time scale trajectories.
We have also observed from both static and dynamic calculations that triplet-mediated pathways are rarely feasible for the nonradiative deactivation process of this molecule. This is due to the somewhat unfavorable energetics and also quite low values of SOC. Here it is important to note that the triplet state, especially T1 is due to orbital transitions between the HOMO and LUMO, which is similar to S1. In such a situation, it is expected to have low SOC values, obeying El-Sayed's rule. Our observations are in line with these expectations.
4. Conclusions
In this work, we have studied the nonradiative deactivation mechanism of the uNAB 5N7C-G, which is also referred to as P. Due to its similarities with the guanine molecule we have also compared our results with that of guanine.We have observed that ultrafast deactivation pathways of 5N7C-G are feasible, especially via a CI that shows a half chair configuration. Another CI that shows a small chair-type puckering with an out-of-plane amine group is also obtained. These CIs and similar geometries are also obtained in the dynamical studies as surface hopping geometries. They form the major and minor deactivation channels with markedly different timescales, and we conjecture that these pathways are indeed responsible for the experimentally obtained sub-picosecond timescales in these molecules. Our claim is also based on the fact that the ratios of these two timescales from experiment (0.38) and theory (0.43) are in excellent agreement. We further ascertain that the triplet-mediated pathways for these deactivation processes are quite unlikely in the molecules, and this is also quite unsurprising from El-Sayed's rule.
In summary, the theoretical observations are in good agreement with the experimental findings, and the pathways of nonradiative deactivation in 5N7C-G that we observe are similar to guanine. The timescales for the early events are in the same orders of magnitude as that of the natural NAB guanine. This points towards the uNAB 5N7C-G as a suitable candidate for expanding the genetic alphabet from the standpoint of photostability.
Author contributions
Somsuta Ray: data curation, investigation and visualization (lead), writing and formal analysis (equal). Debashree Ghosh: conceptualization, funding acquisition, supervision and resources (lead), writing and formal analysis (equal).Data availability
The authors confirm that the data supporting the findings of this study are available within the article and/or its ESI.†Conflicts of interest
There are no conflicts to declare.Acknowledgements
DG thanks ANRF and SERB for generous funding (SPF/2021/000194 and CRG/2023/001806). SR thanks UGC for the Junior Research Fellowship and IACS for computational facilities.References
- E. Szathmáry, Nat. Rev. Genet., 2003, 4, 995–1001 CrossRef PubMed.
- W. Gilbert, Nature, 1986, 319, 618 CrossRef.
- S. A. Benner, A. D. Ellington and A. Tauer, Proc. Natl. Acad. Sci. U. S. A., 1989, 86, 7054–7058 CrossRef CAS PubMed.
- S. Faraji and A. Dreuw, Annu. Rev. Phys. Chem., 2014, 65, 275–292 CrossRef CAS PubMed.
- L. Lapinski, I. Reva, M. J. Nowak and R. Fausto, Phys. Chem. Chem. Phys., 2011, 13, 9676–9684 RSC.
- C. R. Kozak, K. A. Kistler, Z. Lu and S. Matsika, J. Phys. Chem. B, 2010, 114, 1674–1683 CrossRef CAS PubMed.
- W. J. Schreier, J. Kubon, N. Regner, K. Haiser, T. E. Schrader, W. Zinth, P. Clivio and P. Gilch, J. Am. Chem. Soc., 2009, 131, 5038–5039 CrossRef CAS PubMed.
- A. A. Henry and F. E. Romesberg, Curr. Opin. Chem. Biol., 2003, 7, 727–733 CrossRef CAS PubMed.
- B. A. Schweitzer and E. T. Kool, J. Am. Chem. Soc., 1995, 117, 1863–1872 CrossRef CAS PubMed.
- E. T. Kool, Curr. Opin. Chem. Biol., 2000, 4, 602–608 CrossRef CAS PubMed.
- X.-R. Chen, W.-J. Jiang, Q.-H. Guo, X.-Y. Liu, G. Cui and L. Li, Photochem. Photobiol., 2024, 100, 380–392 CrossRef CAS PubMed.
- P. Ghosh, J. Phys. Chem. A, 2024, 128, 8065–8071 CrossRef CAS PubMed.
- I. Hirao, T. Ohtsuki, T. Fujiwara, T. Mitsui, T. Yokogawa, T. Okuni, H. Nakayama, K. Takio, T. Yabuki and T. Kigawa, et al. , Nat. Biotechnol., 2002, 20, 177–182 CrossRef CAS PubMed.
- D. A. Malyshev, K. Dhami, T. Lavergne, T. Chen, N. Dai, J. M. Foster, I. R. Corrêa and F. E. Romesberg, Nature, 2014, 509, 385–388 CrossRef CAS PubMed.
- V. T. Dien, M. Holcomb, A. W. Feldman, E. C. Fischer, T. J. Dwyer and F. E. Romesberg, J. Am. Chem. Soc., 2018, 140, 16115–16123 CrossRef CAS PubMed.
- C. Switzer, S. E. Moroney and S. A. Benner, J. Am. Chem. Soc., 1989, 111, 8322–8323 CrossRef CAS.
- C. Y. Switzer, S. E. Moroney and S. A. Benner, Biochemistry, 1993, 32, 10489–10496 CrossRef CAS PubMed.
- I. Hirao and M. Kimoto, TCI Mail, 2012, 2–11 Search PubMed.
- A. M. Sismour and S. A. Benner, Nucleic Acids Res., 2005, 33, 5640–5646 CrossRef CAS PubMed.
- Z. Yang, A. M. Sismour, P. Sheng, N. L. Puskar and S. A. Benner, Nucleic Acids Res., 2007, 35, 4238–4249 CrossRef CAS PubMed.
- Z. Yang, F. Chen, J. B. Alvarado and S. A. Benner, J. Am. Chem. Soc., 2011, 133, 15105–15112 CrossRef CAS PubMed.
- S. Jahiruddin and A. Datta, J. Phys. Chem. B, 2015, 119, 5839–5845 CrossRef CAS PubMed.
- I. Hirao, M. Kimoto, T. Mitsui, T. Fujiwara, R. Kawai, A. Sato, Y. Harada and S. Yokoyama, Nat. Methods, 2006, 3, 729–735 CrossRef CAS PubMed.
- P. Ghosh, A. Ghosh and D. Ghosh, J. Phys. Chem. A, 2021, 125, 5556–5561 CrossRef CAS PubMed.
- K. Bhattacharyya and A. Datta, Chem. – Eur. J., 2017, 23, 11494–11498 CrossRef CAS PubMed.
- R. Improta, F. Santoro and L. Blancafort, Chem. Rev., 2016, 116, 3540–3593 CrossRef CAS PubMed.
- C. E. Crespo-Hernández, B. Cohen, P. M. Hare and B. Kohler, Chem. Rev., 2004, 104, 1977–2020 CrossRef PubMed.
- A. Giussani, J. Segarra-Marti, D. Roca-Sanjuán and M. Merchan, Photo. Pheno. Nucl. Acids I, 2015, 57–97 CAS.
- A. K. Pradeep Kumar, S. Santra and D. Ghosh, J. Phys. Chem. A, 2024, 128, 9551–9558 CrossRef CAS PubMed.
- K. A. Kistler and S. Matsika, J. Phys. Chem. A, 2007, 111, 2650–2661 CrossRef CAS PubMed.
- T.-h Fu, M.-y Lin, C.-b Fu, X.-f Yu, B. Xiao, J.-b Cheng and Q. Li, Spectrochim. Acta, Part A, 2022, 267, 120549 CrossRef CAS PubMed.
- H. Chen and S. Li, J. Chem. Phys., 2006, 124, 154315 CrossRef PubMed.
- L. Serrano-Andres, M. Merchan and A. C. Borin, J. Am. Chem. Soc., 2008, 130, 2473–2484 CrossRef CAS PubMed.
- Z. Lan, E. Fabiano and W. Thiel, Chem. Phys. Chem., 2009, 10, 1225–1229 CrossRef CAS PubMed.
- R. González-Luque, T. Climent, I. Gonzalez-Ramirez, M. Merchán and L. Serrano-Andrés, J. Chem. Theory Comput., 2010, 6, 2103–2114 CrossRef PubMed.
- M. Barbatti, J. J. Szymczak, A. J. Aquino, D. Nachtigallová and H. Lischka, J. Chem. Phys., 2011, 134, 14304 CrossRef PubMed.
- R. J. Bartlett, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2012, 2, 126–138 CAS.
- J. F. Stanton and R. J. Bartlett, J. Chem. Phys., 1993, 98, 7029–7039 CrossRef CAS.
- E. Runge and E. K. Gross, Phys. Rev. Lett., 1984, 52, 997 CrossRef CAS.
- Y. M. Rhee and M. Head-Gordon, J. Phys. Chem. A, 2007, 111, 5314–5326 CrossRef CAS PubMed.
- P. Pulay, Int. J. Quantum Chem., 2011, 111, 3273–3279 CrossRef CAS.
- J. Finley, P.-Å. Malmqvist, B. O. Roos and L. Serrano-Andrés, Chem. Phys. Lett., 1998, 288, 299–306 CrossRef CAS.
- Y. Shao, Z. Gan, E. Epifanovsky, A. T. Gilbert, M. Wormit, J. Kussmann, A. W. Lange, A. Behn, J. Deng and X. Feng, et al. , Mol. Phys., 2015, 113, 184–215 CrossRef CAS.
- H. Werner, P. Knowles, G. Knizia, F. Manby, M. Schütz, P. Celani, W. Györffy, D. Kats, T. Korona, R. Lindh, et al., MOLPRO, version 2015.1, a package of ab initio programs, University of Cardiff Chemistry Consultants, Cardiff, Wales, UK, 2015 Search PubMed.
- H.-J. Werner, P. J. Knowles, F. R. Manby, J. A. Black, K. Doll, A. Heßelmann, D. Kats, A. Köhn, T. Korona and D. A. Kreplin, et al. , J. Chem. Phys., 2020, 152, 1–24 CrossRef PubMed.
- J. C. Tully, J. Chem. Phys., 1990, 93, 1061–1071 CrossRef CAS.
- S. Mai, P. Marquetand and L. González, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2018, 8, e1370 Search PubMed.
- H. Werner, P. Knowles, G. Knizia, F. Manby, M. Schuitz, P. Celani, T. Korona, R. Lindh, A. Mitrushenkov, G. Rauhut, et al., Molpro: a general–purpose quantum chemistry program package, University of Stuttgart/Cardiff University, 2012 Search PubMed.
- S. E. Krul, G. J. Costa, S. J. Hoehn, D. Valverde, L. M. Oliveira, A. C. Borin and C. E. Crespo-Hernández, Photochem. Photobiol., 2023, 99, 693–705 CrossRef CAS PubMed.
- M. Barbatti, A. J. Aquino and H. Lischka, J. Phys. Chem. A, 2005, 109, 5168–5175 CrossRef CAS PubMed.
- C. Canuel, M. Mons, F. Piuzzi, B. Tardivel, I. Dimicoli and M. Elhanine, J. Chem. Phys., 2005, 122, 074316 CrossRef PubMed.
- S. E. Krul, S. J. Hoehn, K. J. Feierabend and C. E. Crespo-Hernández, J. Chem. Phys., 2021, 154, 075103 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d5cp00986c |
| This journal is © the Owner Societies 2025 |