Identification and characterization of binding thermodynamics and kinetics of inhibitors targeting FGFR1 via molecular modelling and ligand Gaussian accelerated molecular dynamics simulations†
Subhasmita
Mahapatra
 and
Parimal
Kar
and
Parimal
Kar
 *
*
Department of Biosciences and Biomedical Engineering, Indian Institute of Technology Indore, Khandwa Road, Indore 453552, Madhya Pradesh, India. E-mail: phd2101271009@iiti.ac.in; parimal@iiti.ac.in
First published on 16th April 2025
Abstract
Fibroblast growth factor receptor1 (FGFR1) kinase has a crucial role in cell proliferation, migration, and differentiation. Any imbalance in its level can cause cancer and many other illnesses. Despite the availability of numerous treatments, cytotoxicity, selectivity, and drug resistance issues demand the development of new FGFR1 inhibitors. Herein, we performed a high-throughput virtual screening of 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 624 compounds from NPASS and HMDB databases using the Schrodinger software suite. Compounds with a docking score cutoff of −11.0 kcal mol−1 were further screened for ADMET properties. Following the all-atom molecular dynamics simulation of selected molecules in replica, the binding free energy was calculated using the molecular mechanics Poisson Boltzmann surface area (MM-PBSA) scheme. We obtained two compounds, namely, bevantolol and 3-hydroxy glabrol, which exhibited higher binding affinities than the control drug ponatinib. Bevantolol was further optimized via virtual screening and simulation studies of its 100 structural analogues to obtain the best analogue. Subsequently, we investigated the binding thermodynamics and kinetics of the best analogue molecule via ligand Gaussian accelerated molecular dynamics (LiGaMD) simulation, performing independent replica runs of 4 μs each. The 1D and 2D potential of mean force and Kramer's rate theory determined the kinetic rate constants (kon/koff) associated with the FGFR1 complex. The binding constant was estimated to be 7.4 ± 0.27 nM, which was similar to the type II tyrosine kinase inhibitor ponatinib. Overall, this study highlights the dynamics of FGFR1–ligand interaction while proposing bevantolol and its analogue molecule ANLG-2 as promising drug candidates for FGFR1 therapeutic intervention.
624 compounds from NPASS and HMDB databases using the Schrodinger software suite. Compounds with a docking score cutoff of −11.0 kcal mol−1 were further screened for ADMET properties. Following the all-atom molecular dynamics simulation of selected molecules in replica, the binding free energy was calculated using the molecular mechanics Poisson Boltzmann surface area (MM-PBSA) scheme. We obtained two compounds, namely, bevantolol and 3-hydroxy glabrol, which exhibited higher binding affinities than the control drug ponatinib. Bevantolol was further optimized via virtual screening and simulation studies of its 100 structural analogues to obtain the best analogue. Subsequently, we investigated the binding thermodynamics and kinetics of the best analogue molecule via ligand Gaussian accelerated molecular dynamics (LiGaMD) simulation, performing independent replica runs of 4 μs each. The 1D and 2D potential of mean force and Kramer's rate theory determined the kinetic rate constants (kon/koff) associated with the FGFR1 complex. The binding constant was estimated to be 7.4 ± 0.27 nM, which was similar to the type II tyrosine kinase inhibitor ponatinib. Overall, this study highlights the dynamics of FGFR1–ligand interaction while proposing bevantolol and its analogue molecule ANLG-2 as promising drug candidates for FGFR1 therapeutic intervention.
1. Introduction
Human fibroblast growth factor receptors (FGFRs) exert important physiological roles in embryogenesis, wound repair, cellular behaviour and endocrine homeostasis by interacting with fibroblast growth factors (FGFs) and heparan sulfate proteoglycans (HSPGs).1 FGFRs are characterized as transmembrane receptors with one extracellular and one intracellular domain that act as a ligand binding domain and a tyrosine kinase domain, respectively. The extracellular region contains three immunoglobulin (Ig)-like components and an acidic box with eight sequential acidic residues. Ig-I participates in receptor auto-inhibition in conjunction with the acidic box. Meanwhile, Ig-II and Ig-III create FGF and HSPG binding domains2 (Fig. 1A). Similar to other receptor tyrosine kinases, FGFRs rely on tyrosine phosphorylation for activation. The ternary complex of FGFR-FGF-HSPG leads to dimerization, autophosphorylation and substrate phosphorylation in a coordinated manner. A cascade of signalling pathways, such as the phospholipase-γ (PLC-γ) pathway and the mitogen-activated protein kinase (MAPK) pathway, get activated by these phosphorylations to carry out various physiological processes.3 | ||
| Fig. 1 (A) Activation of FGFR. The extracellular ligand-binding domain of FGFR is made up of Ig-I, acid box, Ig-II, and Ig-III, followed by the transmembrane domain (TMD), Juxtra membrane domain (JMD), and the intracellular kinase domain. FGF ligand (blue) and heparin (violet) together make a ternary complex to activate FGFR. (B) Three-dimensional structure of the FGFR1-ponatinib complex (PDB ID: 4V04). The ligand interaction pocket was highlighted with mesh representation (C) workflow of processes associated with finding the promising inhibitor against FGFR1. | ||
The FGFR family is made up of four receptors: FGFR1–FGFR4. Although distinct genes encode these four receptors, they share a great deal of similarity and range in sequence identity from 56% to 71%.4 The fibroblast growth factor receptor-like 1 (FGFRL1 or FGFR5) does not have the kinase domain and acts as a bogus receptor to inhibit the signalling cascade.5,6 The FGFR signalling pathway plays a crucial role in cancer progression owing to its pleiotropic effects and involvement in key physiological processes. Its dysregulation leads to antiapoptotic, mutagenic, and angiogenic responses characteristic of cancer7 while serving as an escape mechanism for therapy resistance. A comprehensive analysis of 4853 solid tumours revealed FGFR1 aberrations in 3.5% of cases, compared to 1.5% for FGFR2, 2% for FGFR3, and 0.5% for FGFR4.8 Notably, FGFR1 amplifications were predominant, constituting 66% of FGFR alterations, and were especially prevalent in cancers such as breast cancer (10%)9 and squamous non-small cell lung cancer (20%).10 Although FGFR2, FGFR3, and FGFR4 also play roles in oncogenesis, their aberrations are less common or more tumour-specific.10
There is an arsenal of FGFR-targeted therapeutics, and the FDA has also approved two drugs: pemigatinib for FGFR2-fusion cholangiocarcinoma11 and erdafitinib for FGFR3-altered urothelial cancer.12 Although some patients respond to FGFR-targeted drugs at first, many later experiences acquired resistance, which causes the disease to worsen and therapy to be discontinued. Secondary chromosomal changes like gatekeeper mutation in FGFR1 kinase cause drug resistance while increasing autophosphorylation activity.13 The conserved amino acid sequences in key regions (A-loop, P-loop, and Hinge) share 65–78% identity,14 making kinase domain targeting a challenge for inhibitor development across malignancies. The higher frequency and broad distribution of FGFR1 alterations make it a compelling target for therapeutic intervention, which would benefit a larger patient population.
Analyzing the protein kinase conformational ensemble in their multiple functional states can aid in comprehending the receptor–ligand interactions.15,16 Current atomistic conventional molecular dynamics (cMD) simulation techniques are restricted to limited time scales and specific potential energy barriers between intermediate states.17,18 Multiple enhanced sampling techniques like metadynamics, hyperdynamics, and adaptive biasing approaches evolved to better assess the biological processes.19,20 However, these strategies are constrained by the necessity for predetermined parameters. The Gaussian accelerated molecular dynamics (GaMD) technique eliminates the need for specified collective variables or reaction coordinates, significantly lowering energy barriers and enhancing the speed of MD simulations exponentially.21 The GaMD technique has effectively elucidated molecular interactions that align with in vitro data and long-duration cMDs.22–24
Ligand GaMD (LiGaMD)21,25 enhance the sampling efficiency of molecular dynamics simulations, particularly for ligand binding and unbinding processes. Building upon the Gaussian accelerated molecular dynamics (GaMD) framework, LiGaMD selectively applies a boost potential to the ligand's non-bonded interaction energy, effectively accelerating ligand movements without the need for predefined reaction coordinates. This approach enables the capture of repetitive binding and unbinding events within feasible simulation timescales. In the initial development of LiGaMD, Miao et al. demonstrated its efficacy using simulations of guest molecules interacting with β-cyclodextrin and captured multiple binding and unbinding events within hundreds of nanoseconds. The calculated binding free energies closely matched experimental data, with errors less than 1.0 kcal mol−1.21 LiGaMD offers powerful tools for accurately characterizing ligand binding thermodynamics and kinetics, thereby facilitating computer-aided drug design and a deeper understanding of molecular interactions.15
Here, we used a thorough computational method to find a hit molecule that might be further refined into a potentially special inhibitor of FGFR1 kinase. This strategy includes structure-based virtual screening, ADMET (absorption, distribution, metabolism, excretion, and toxicity) analysis and molecular dynamics simulations (Fig. 1). We used the high throughput virtual screening (HTVS) method to test about 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 624 natural molecules against FGFR1 from the Natural Product Activity and Species Source Database, NPASS (30
624 natural molecules against FGFR1 from the Natural Product Activity and Species Source Database, NPASS (30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 926 compounds), and Human Metabolome Database, HMDB (23
926 compounds), and Human Metabolome Database, HMDB (23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 698 compounds). Following VS, every top lead compound from the ADMET analysis was filtered. With the aid of MD simulations, further binding poses, and structural refinement were performed on each complex. Then, we used the molecular mechanics Poisson–Boltzmann surface area (MM-PBSA) approach to calculate the binding free energies of each compound against FGFR1. The best-hit compound with high binding affinity was further optimized with the DELA-drug server by generating 100 analogues. These analogues underwent HTVS using the glide module, and two compounds with the best docking score were selected. We performed MD simulations of 2 × 200 ns followed by trajectory analysis. ANLG-2 showed better interaction with FGFR1 kinase. To gain deeper insights into the kinetics of the ANLG-2 and FGFR1 kinase complex, we performed LiGaMD simulations in replicates with each run extending to 4 μs. The kinetic dissociation constant of the complex was calculated using Kramer's rate theory.
698 compounds). Following VS, every top lead compound from the ADMET analysis was filtered. With the aid of MD simulations, further binding poses, and structural refinement were performed on each complex. Then, we used the molecular mechanics Poisson–Boltzmann surface area (MM-PBSA) approach to calculate the binding free energies of each compound against FGFR1. The best-hit compound with high binding affinity was further optimized with the DELA-drug server by generating 100 analogues. These analogues underwent HTVS using the glide module, and two compounds with the best docking score were selected. We performed MD simulations of 2 × 200 ns followed by trajectory analysis. ANLG-2 showed better interaction with FGFR1 kinase. To gain deeper insights into the kinetics of the ANLG-2 and FGFR1 kinase complex, we performed LiGaMD simulations in replicates with each run extending to 4 μs. The kinetic dissociation constant of the complex was calculated using Kramer's rate theory.
2. Methodology
2.1. Preparation of target protein
The crystal structure of FGFR1 kinase in association with ponatinib has been used for this study (PDB ID: 4V04, resolution – 2.12 Å).26 Chain A was considered, and apo conformation of FGFR1 kinase was obtained by removing the atomic coordinates of the sulfate ion (SO42−) and 1,2-ethandiol (EDO) using UCSF Chimera.27 All the missing residues were added using the modeller software.28 The Schrödinger software suite's protein creation wizard was used to prepare the receptor protein.29 The crystallographic waters beyond 5.0 Å from the kinase were eliminated, and all missing hydrogen atoms were added. With the aid of PROPKA, the structure was neutralized at pH 7.0 while considering the orientations of the retained water molecules.30 Finally, using the OPLS3 force field,31 the protein was minimized with default settings.2.2. Preparation of ligand
The primary emphasis of the current investigation is kinase inhibition. As a result, we have used natural compounds such as phytochemicals and metabolite databases. Phytochemical databases (30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 926 compounds) and the Metabolome Database (HMDB) (23
926 compounds) and the Metabolome Database (HMDB) (23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 698 compounds) were used to screen a total of 54
698 compounds) were used to screen a total of 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 624 compounds. The ligand compounds were downloaded in SDF file format. To create the compound conformations and their tautomeric combinations, the ligands were uploaded into the Maestro interface and submitted to the Ligprep module. After hydrogen atoms were added, the ligands were minimized and described using the OPLS3 force field.31 We used all the ligand conformers for virtual screening.
624 compounds. The ligand compounds were downloaded in SDF file format. To create the compound conformations and their tautomeric combinations, the ligands were uploaded into the Maestro interface and submitted to the Ligprep module. After hydrogen atoms were added, the ligands were minimized and described using the OPLS3 force field.31 We used all the ligand conformers for virtual screening.
2.3. Virtual screening
The virtual screening workflow (VSW) procedure in the Glide module of the Schrodinger suite was used to screen a total of 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 624 compounds against FGFR1.32 Initially, ponatinib from the complex crystal structure was used as the pocket of FGFR1 to construct the receptor grid using Glide, leaving 12 Å cubic space around it. The molecules from each database were initially filtered using Lipinski's rule of five and the QikProp module during the VSW method. Our method of docking was semi-flexible, making the ligands move freely inside the ligand pocket while the protein remains rigid. Thus, just one structure of the receptor was employed, but all configurations of each compound were utilized during the docking process. High-throughput virtual screening (HTVS), standard precession (SP), and extra precession (XP) were the three tiers in which the virtual screening was carried out progressively. For Glide-SP, the top 30% of HTVS were taken. Finally, 10% of Glide-XP was chosen once 20% of SP had been allocated for Glide-XP. All the leads were screened and chosen for further analysis based on the XP-docking score.
624 compounds against FGFR1.32 Initially, ponatinib from the complex crystal structure was used as the pocket of FGFR1 to construct the receptor grid using Glide, leaving 12 Å cubic space around it. The molecules from each database were initially filtered using Lipinski's rule of five and the QikProp module during the VSW method. Our method of docking was semi-flexible, making the ligands move freely inside the ligand pocket while the protein remains rigid. Thus, just one structure of the receptor was employed, but all configurations of each compound were utilized during the docking process. High-throughput virtual screening (HTVS), standard precession (SP), and extra precession (XP) were the three tiers in which the virtual screening was carried out progressively. For Glide-SP, the top 30% of HTVS were taken. Finally, 10% of Glide-XP was chosen once 20% of SP had been allocated for Glide-XP. All the leads were screened and chosen for further analysis based on the XP-docking score.
2.4. Pharmacokinetic profile
The Schrödinger QikProp module was used in the screening step to make predictions about the physical and therapeutic properties of organic molecules. This module forecasts 35 crucial traits that are used in the drug development programme as screening for ligand libraries. Following the virtual screening, the 27 top lead compounds were assessed for their ADMET characteristics using the ADMETlab 2.0 web server.33 Aspects including pharmacochemical characteristics, ease of production, drug-like characteristics, absorption, bioavailability and others were covered in our prediction of ADMET traits. Among different toxicological features, AMES toxicity, hepatotoxicity, carcinogenicity, and pulmonary toxicity were assessed. Four ligands were ultimately evaluated with additional MD simulations according to the docking score value and all the pharmacokinetic properties were determined.2.5. Molecular dynamic simulation
Atomistic MD simulations of 200 ns were conducted on the systems of interest, including apo, control (ponatinib/FGFR1 complex), and FGFR1/lead complex. The AMBER ff14SB34 force field was used for parameter assignment of the protein. For ligands, we employed the generalized AMBER force field, version 2 (GAFF2).35–37 Ions were added in the right quantity to balance the system. Each system was solvated in an octahedron water box using the TIP3P explicit water model38 and 10.0 Å box size. The SHAKE algorithm was utilized to describe all bonds, including hydrogen atoms.39 With a non-bonded cutoff of 10.0 Å, the particle–mesh Ewald (PME) technique40 was implemented to compute long-range interactions. During the MD simulation, the 2-fs time-step was fixed for each complex. All six systems were minimized using the 5000-step steepest descent followed by the 5000-step conjugate gradient, initially with a force constraint of 2.0 kcal mol−1 Å−2 and later without any constraint. With a steady pressure of 1 bar and a constant volume, each system's temperature was regulated for 1 ns between 0 and 300 K. The Langevin thermostat41 and Berendsen Barostat42 were used to maintain the temperature and pressure, respectively. Two replicates of 200 ns production run were carried out for each complex at the NPT level. Using AMBER18's Cpptraj43 module, 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 snapshots were created and examined altogether.
000 snapshots were created and examined altogether.
2.6. Binding free energy computation
The complexation free energy (ΔGbind) between receptor and ligand was computed utilizing the renowned Poisson–Boltzmann surface area (MM/PBSA) scheme. The MMPBSA.py tool of the AMBER suite was used to evaluate ΔGbind. We have discussed the specificities of the MMPBSA methods in earlier work.44,45According to the mentioned equation,46 the binding free energy has three terms: the conformational entropy term (TΔS), the solvation energy (ΔGsolv), and the internal energy of molecular mechanics (ΔEinternal).
| ΔGbind = ΔH − TΔS = ΔEinternal + ΔGsolv − TΔS | (1) |
Here, 5000 frames from the last 100 ns of the MD trajectory were utilized to evaluate the free energy. The normal mode analysis (NMA) technique was used to determine the entropy. Only 50 frames were considered for NMA because of the high computing expense. The respective contributions of each amino acid to the total binding free energy were determined using the MM-PBSA pair-wise decomposition approach.47
2.7. Production of conformational analogues
To optimize the best-selected molecule (bevantolol), 100 analogues were generated utilizing the DeLA-Drug48 software. This implements deep learning algorithms like sampling with substitutions (SWS) and a recurrent neural network (RNN). The Schrödinger suite was used to systematically screen the generated analogues against FGFR1 in the virtual screening workflow, utilizing a variety of previously mentioned parameters. Based on docking score data, 200 ns replica runs of atomistic molecular dynamics simulations were performed on the top two analogues (ANLG-1 and ANLG-2). Considering the conformational stability and distance from the pocket, we selected ANLG-2 for further LiGaMD simulation. The production simulation was run for 4.0 μs in replicate, followed by energy calculation using Kramer's rate theory.2.8. LiGaMD
When a ligand (L) attaches itself to a protein (P) in a physiological condition (E), potential energy V(r) is created. This potential energy can be estimated as follows:| V(r) = VP,b(rp) + VL,b(rL) + VE,b(rE) + VPP,nb(rp) + VLL,nb(rL) + VEE,nb(rE) + VPL,nb(rPL) + VPE,nb(rPE) + VLE,nb(rLE) | (2) |
The bound potential energies of protein P, ligand L, and environment E are denoted by the symbols VP,b, VL,b, and VE,b, respectively. The self-nonbonded potential energies of protein P, ligand L, and environment E are VPP,nb, VLL,nb, and VEE,nb, respectively. The non-bonded interaction energies of P–L, P–E, and L–E are VPL,nb, VPE,nb, and VLE,nb, respectively. The P–L and L–E nonbonded energies are the main mechanisms involved in the binding and unbinding of ligands. The expression for this interaction energy is as follows:
| VL,nb(r) = VLL,nb(rL) + VPL,nb(rPL) + VLE,nb(rLE) | (3) |
In the LiGaMD simulation, a boost potential energy DVL,nb(r), is introduced, which is stated as follows:
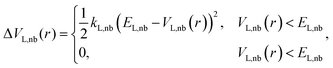 | (4) |
The harmonic constant in this case is kL,nb, while the boost potential's threshold energy is EL,nb. The parameters kL,nb and others are obtained in a manner similar to that of the GaMD algorithm.22
2.9. Free energy calculations for LiGaMD simulation using the Py reweighting method (1D- and 2D-PMF)
Given a reaction coordinate r, P*(A) represents the probability distribution of the chosen reaction coordinates A(r). P*(A) can be reweighted and represented as follows based on the LiGaMD boost energies of each reaction coordinate: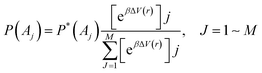 | (5) |
Here, M signifies the number of bins, β is equal to kBT, and [eβΔV(r)]j is the ensemble-average factor of the jth bin. The following is a definition of the ensemble-average factor, which is used to minimize energetic noise:
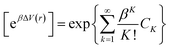 | (6) |
The first three cumulants of (6) can be explained as:
| C1 = [ΔV], C2 = [ΔV2] − [ΔV]2, C3 = [ΔV3] − 3[ΔV2] [ΔV] + 2[ΔV]2, | (7) |
The free energies from reweighting can be computed as:
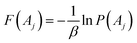 | (8) |
The 1D and 2D reweighted PMFs and associated ligand interaction-free energies were computed by reweighting LiGaMD simulations using the PyReweighting toolkit.21,49
2.10. Kinetic rate constants derived through Kramer's rate theory
Miao et al.’ used Kramer's' rate theory in combination with GaMD-related techniques to model the kinetic and thermodynamic aspects of the ligand binding interaction’.21 The time intervals and structural average of ligand-bound (τB) and ligand-unbound (τU) states can be obtained for a suitable sampling of ligand interaction during the simulation. The following is the expression for the ligand binding and dissociation rate constants (kon and koff):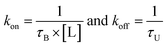 | (9) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 896 water molecules in the system along with one ligand molecule, the ligand concentration was calculated to be 5.1 mM. Initially estimating the system value guides towards [L] calculations. The following is an expression for a chemical reaction's rate in the large viscosity limit:
896 water molecules in the system along with one ligand molecule, the ligand concentration was calculated to be 5.1 mM. Initially estimating the system value guides towards [L] calculations. The following is an expression for a chemical reaction's rate in the large viscosity limit: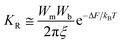 | (10) |
Here, Wm and Wb are the estimated harmonic oscillator frequencies close to the global energy minima, the energy barrier ξ specifies the fictional rate constant, and ΔF signifies the free energy difference of transversion. The fictional rate constant ξ is connected to the diffusion coefficient D as ξ = kBT/D. By dividing the kinetic rate computed using ‘he simulations’ transition time series by the kinetic rate computed utilizing the probability density of the Smoluchowski equation, one may determine the apparent diffusion coefficient D50,51 (dt = 0.00001 for kon and dt = 0.0001 for koff). The ratio of approximate diffusion coefficients (D) is computed based on simulation findings, and the free energy transitions of ligand binding and unbinding are computed based on PMF profiles. The computations are comparable for curvatures of PMF profiles close to the energy barrier (“Br”), ligand-bound (“B”) and unbound (“U”) low-energy wells. The original kinetic rate constants can then be recovered by using the numbers that were obtained in eqn (10) to compute the ligand association and dissociation rates throughout the LiGaMD simulations.
2.11. Residual motion and structure network analyses
We created a dynamic cross-correlation map (DCCM)52 in RStudio using the Bio3D53 package of R for analyzing the residual correlation dynamics of the FGFR1 kinase for all the systems. The calculations for the correlation coefficient were done according to the atoms of the protein, and the temporal average of the divergence of each residue was used from the mean value over the duration of production runs. A value of 0 indicates that there is no connection between the motions of the residues. Positive and negative values suggest correlated and anti-correlated motions, respectively. Following that, an FGFR1 structure-based model of the correlation motion was created, and PyMOL was used to visualize it. Additionally, the protein structure network (PSN)54 approach of the webPSN v2.055 portal was used to characterize the structural communications between the kinase residues. Many aspects of a PSN were examined and visualized using PyMOL, such as its nodes, linkages, hubs, and communities.3. Results and discussion
3.1. Virtual screening
The QikProp and Lipinski's rule of five filters was used to screen the large phytochemical dataset, and 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 624 chemicals were passed for the subsequent HTVS docking module. 16
624 chemicals were passed for the subsequent HTVS docking module. 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 387 phytochemicals were processed by the HTVS module for the SP docking protocol. Around 3278 compounds were sorted by the SP docking programme before moving on to the precise screening of the extra precision (XP) docking programme. Then, we selected 27 ligand molecules with a docking score cut off of −11.0 kcal mol−1 for further analysis (Table S1, ESI†).
387 phytochemicals were processed by the HTVS module for the SP docking protocol. Around 3278 compounds were sorted by the SP docking programme before moving on to the precise screening of the extra precision (XP) docking programme. Then, we selected 27 ligand molecules with a docking score cut off of −11.0 kcal mol−1 for further analysis (Table S1, ESI†).
3.2. ADMET analysis
Using the well-known web servers ADMETlab 2.0 and the QikProp module, we assessed their pharmacological profiles to evaluate the likelihood of these 27 hits being produced as human medicine in the future. The ESI,† Table provides a list of the anticipated pharmacological characteristics. According to Lipinski's rule of five, all 27 hits had molecular weights below 500 Da, a logarithm of the n-octanol/water distribution coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) p) below 5, and the number of hydrogen bond recipients and donors below 10 and 5, respectively (Tables S1 and S2, ESI†). The logp value of 13 compounds lies in the range of 2 to 3, while 7 compounds show a higher value of more than 4. A calculation of the aqueous permeability (log
p) below 5, and the number of hydrogen bond recipients and donors below 10 and 5, respectively (Tables S1 and S2, ESI†). The logp value of 13 compounds lies in the range of 2 to 3, while 7 compounds show a higher value of more than 4. A calculation of the aqueous permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) s) to check the cellular permeability suggested that all the compounds can move through the cell membrane except silibinin and NPC174512. An in vitro model of the Caco-2 (human colon adenocarcinoma) cell line is used to predict intestinal mucosal absorption, which gives an overview of oral absorption. Even though 11 out of the 27 hits were in the marginally unfavourable range of Caco-2 and MDCK (Madin–Darby canine kidney cells) permeability estimation, others produced satisfactory results (Table S2, ESI†). Moreover, we predicted each molecule's toxicity profile, which is listed in Table S3 (ESI†). It provided a variety of anticipated toxicity scores for several tests, including AMES toxicity, hepatotoxicity, respiratory toxicity and hERG-blocking potency. None of the molecules showed strong carcinogenicity and hepatotoxicity. After all the analysis, we considered bevantolol, 3-hydroxy glabrol, daurichromene A and pawhuskin B as the hit molecules for molecular dynamic simulation, based on their strong docking scores and overall projected ADMET properties.
s) to check the cellular permeability suggested that all the compounds can move through the cell membrane except silibinin and NPC174512. An in vitro model of the Caco-2 (human colon adenocarcinoma) cell line is used to predict intestinal mucosal absorption, which gives an overview of oral absorption. Even though 11 out of the 27 hits were in the marginally unfavourable range of Caco-2 and MDCK (Madin–Darby canine kidney cells) permeability estimation, others produced satisfactory results (Table S2, ESI†). Moreover, we predicted each molecule's toxicity profile, which is listed in Table S3 (ESI†). It provided a variety of anticipated toxicity scores for several tests, including AMES toxicity, hepatotoxicity, respiratory toxicity and hERG-blocking potency. None of the molecules showed strong carcinogenicity and hepatotoxicity. After all the analysis, we considered bevantolol, 3-hydroxy glabrol, daurichromene A and pawhuskin B as the hit molecules for molecular dynamic simulation, based on their strong docking scores and overall projected ADMET properties.
Further, these compounds were clustered by utilizing the multidimensional scaling (MDS56) algorithm in the ChemMine57 interface based on their structural and physical similarity. A similarity cutoff of 0.4 was used for the dissimilarity assessment. With the help of the Tanimoto coefficient,58 a similarity matrix was used to determine the distance matrices for each substructure based on the shared and distinctive characteristics seen across pairs. Lead molecules are divided into scatter plots (Fig. S4, ESI†), and all exhibit significant differences from the FDA-approved drug ponatinib. As a result, most investigated compounds differ structurally from current medications.
3.3. Conformational stability analysis
A thorough analysis was conducted following the 200 ns duplicate simulation run to ascertain the stability and dynamic characteristics of the studied systems. For every backbone Cα atom, we calculated the root mean square deviation (RMSD) with respect to their initial configuration (Fig. S1, ESI†). Simulations for every FGFR1/lead complex have shown convergence. To learn more about the conformational variation of the overall protein backbone, we implemented the kernel density estimation (KDE) technique and illustrated the probability distribution of RMSD during the duplicate simulations (Fig. 2A).The apo system shows a wider range of RMSD distribution (2.1–3.7 Å) compared with the complex systems. The control ponatinib (FDA-approved drug),59 bevantolol and pawhuskin B complex with FGFR1 show narrow peak distribution at 2 Å, 2.5 Å, and 2.8 Å, respectively. Bevantolol/FGFR1 shows a sharp peak with high probability and narrow distribution (1.5–3 Å), suggesting a single conformation with high structural stability. We also computed the potential of mean force (PMF) to understand the structural flexibility as energy functions (Fig. 2B). The apo and daurichromeneA/FGFR1 systems show a wider free energy basin. In contrast, others have a uniform global minimum with a smaller shoulder peak. The energy minima of ponatinib/FGFR1 have a range of 1.8–2.1 Å, with wider RMSD fluctuation until ∼3.8 Å. A narrow free energy minimum with a shoulder peak barrier of ∼1 kcal mol−1 is observed for the FGFR1/bevantolol system. Meanwhile, FGFR1/pawhuskin B suggests multiple conformational variations isolated by an energy barrier of ∼0.5 kcal mol−1.
The probability density map of the radius of gyration (ROG) in Fig. 2C depicts the structural compactness. Wider distributions with a ROG range of 19.8 Å to 21.2 Å were visible in the apo system. In contrast, the complex systems had a single unimodal distribution, indicating that the structure becomes more rigid when it is linked to the ligand. Each residue's RMSF were computed to study the residual variations of all the complexes. Fig. 2D shows that the P-loop, substrate binding region, and A-loop regions exhibit relatively higher fluctuations in all cases. It was observed that bevantolol had lesser fluctuation while daurichromene A showed elevated variation in the A-loop and substrate binding regions. The variation that was observed after ligand binding may have been caused by ligand–protein interactions at different regions.
3.4. Binding pocket dynamics
We determined the FGFR1–ligand distance and represented it as a probability density function to visualize the binding pocket stability (Fig. 3A). The minimal distances between the ligand and FGFR1 imply that the ligand remained in the binding pocket of FGFR1 during the simulation. The control ligand ponatinib showed a sharp peak at 0.5 Å with a high probability distribution, suggesting an efficient interaction with FGFR1. The other systems manifested a broader peak distribution in a range of ∼1 to 1.6 Å. The time evolution plot conveys a similar result as the KDE plot of ligand–protein distance (Fig. S2, ESI†). We additionally computed the RMSD of the backbone atoms of the interaction pocket of FGFR1, which comprises the residues in each system that are less than the 5 Å range. From all the FGFR1/ligand complexes, bevantolol showed a pronounced peak at 0.5 Å, with an elevated probability distribution. Ponatinib showed two peaks at 0.5 Å and 1.5 Å, while 3-hydroxy glabrol had a widespread variation from 1 Å to 1.5 Å, suggesting a high fluctuation of binding pocket throughout the simulation. Pawhuskin B and daurichromene A had broad distribution with shoulder peaks, implying multiple conformational possibilities. The time evolution of all the complexes was converged for duplicate simulations, and similar observations were inferred from the KDE plot (Fig. S3). | ||
| Fig. 3 Probability density function of (A) ligand protein distance and (B) binding pocket of lead molecules. (C) Hydrogen bond interaction profile of FGFR1 with respective compounds. | ||
The H-bond interactions between the ligand and protein were investigated to analyze the stability of the complex and comprehend the molecular interplay. Therefore, the intermolecular H-bonds between the ligands and FGFR1 were calculated. Fig. 3C shows the H-bond profile for each system during the simulations. Bevantolol had the most H-bonds between the inhibitor and FGFR1 compared to the other compounds. The average number of H-bonds ranges from 1 to 5, such as 3-hydroxy glabrol: 4; ponatinib: 2; pawhuskin B: 3; and daurichromene A: 2. These H-bonds remained steady throughout the entire simulation.
Additionally, the H-bond occupancy between the ligand and FGFR1 was computed and listed in Table S4 (ESI†) for each complex along the trajectory. Bevantolol strongly binds to FGFR1 primarily with the residues Glu537 (from αChelix, ∼72.2% occupancy) and Glu531 (from αChelix, ∼27.3% occupancy), indicating consistent H-bonding between them. Glu568 of FGFR1 (from Hinge, 64.5% occupancy) interacts strongly with 3-hydroxy glabrol. Likewise, the highest number of H-bonds from the other complexes include ponatinib, Glu531 (∼30.8%); daurichromene A, Asp647 (from DFG motif ∼51.4%); pawhuskin B, Glu568 (Hinge, ∼56.1%). Overall, the strong contacts formed by the ligands at the ATP-binding region seem to involve residues from the hinge region and the αChelix region. Our findings demonstrated that the phytochemical compound bevantolol and 3-hydroxy glabrol interact strongly with FGFR1 through H-bonding and hydrophobic contacts and may be useful for FGFR1-mediated therapeutics.
3.5. Binding free energy analysis
Free energy estimations combined with MD simulations are useful for examining the strength of interactions and the stability of ligand binding. To comprehend the ligand interaction strength with FGFR1, we evaluated several individual contributions of binding free energy and compared them with ponatinib (Table 1 and Fig. S6, ESI†). The protein/ligand complexation was influenced by electrostatic (ΔEelec), van der Waals (ΔEvdw), and non-polar solvation free energy (ΔGnon-polar) interactions. The polar solvation-free energy (ΔGpolar) and the entropic penalty (TΔS) oppose the complex formation. The four systems, along with the control drug, were used to calculate the average binding affinity.| FGFR1/ligand complex | ΔEvdw | ΔEelec | ΔGpol | ΔGnp | −TΔS | ΔGbind |
|---|---|---|---|---|---|---|
| Ponatinib | −70.98 (0.04) | −26.21 (0.05) | 56.56 (0.05) | −6.11 (0.00) | 30.30 (0.81) | −16.20 (1.05) |
| Bevantolol | −51.50 (0.05) | −149.95 (0.21) | 163.75 (0.16) | −4.85 (0.00) | 24.15 (0.98) | −21.20 (0.91) |
| 3-Hydroxy glabrol | −49.72 (0.05) | −27.85 (0.09) | 44.50 (0.05) | −4.86 (0.00) | 19.30 (1.11) | −18.63 (1.01) |
| Pawhuskin B | −52.29 (0.03) | −23.13 (0.06) | 45.15 (0.04) | −4.98 (0.00) | 22.56 (0.81) | −12.74 (0.97) |
| Daurichromene A | −46.97 (0.04) | −14.53 (0.06) | 34.44 (0.06) | −4.73 (0.00) | 21.26 (1.11) | −10.57 (0.93) |
All complexes have binding free energies that range from −10.57 to −21.20 kcal mol−1. Our findings revealed that human metabolite compounds had a higher binding affinity than the FDA-approved drug ponatinib. It shows a KD value of 7.7 nm, consistent with its binding free energy.60 The HMDB database compound bevantolol (−21.20 kcal mol−1) has the highest binding affinity against FGFR1. The other compound, 3-hydroxy glabrol, shows a binding affinity of −18.63 kcal mol−1. The binding energy of NPASS database compounds pawhuskin B and daurichromene A are −12.74 and −10.57 kcal mol−1, respectively, and are lower than the control compound.
The significant interaction between FGFR1 and bevantolol is promoted by ΔEelec, ΔEvdw, and lowered entropy (Table 1). Our earlier research on the FGFR1-ATP complex reveals that electrostatic interactions are crucial to robust protein–ligand bonds.45 Compared with 3-hydroxy glabrol, it seems that the reduced entropy plays an important role in increasing the binding affinity. For ponatinib, ΔEvdw favours complex formation, although it is opposed by the high entropic value. High polar contribution and low ΔEelec, ΔEvdw decrease the interaction strength of pawhuskin B and daurichromene A. Overall, it appears that bevantolol's strong binding affinity for FGFR1 is attributable to higher ΔEelec contributions than the others.
Bevantolol is a β1-adrenergic receptor antagonist primarily used for managing hypertension and angina.61 There is a lack of experimentation on bevantolol against FGFR1. However, a study has explored the combined effects of nebivolol, another β1-blocker, and FGFR inhibitors on breast cancer cells. The study demonstrated that nebivolol synergistically increased the sensitivity of BT-474 breast cancer cells to FGFR inhibitors like erdafitinib and AZD4547, primarily through the down-regulation of AKT activation.62 This suggests potential interactions between β1-blockers and FGFR pathways. Bevantolol could aid future therapeutic inventions against FGFR1 in cancer treatment. We have considered these two compounds for further analysis. We also calculated the MCS58 (Maximum Common Substructure) Tanimoto index to examine the structural similarity among our selected drugs. Tanimoto coefficient ranges from 0 to 1: a lower value implies less similarity, while a greater value implies high structural similarity between molecules. We compared bevantolol and 3-hydroxy glabrol with each other and the control ponatinib. The overall analysis signifies structural diversity among the selected compounds (Fig. S5, ESI†).
3.6. Per residue energy contribution
The MM/PBSA technique was used to calculate each residue's impact on the overall binding free energy to comprehend how the selected compounds interact with FGFR1. The per-residue breakdown of free energy reveals the critical residues responsible for ligand binding. Fig. 4 shows the per residue energy contribution of FGFR1–ligand complexes with binding free energy greater than the control drug. The residues from the αChelix, A-loop, and hinge region significantly influenced the interaction of the specified compounds. Glu537 and Asp647 from the activation loop region substantially impacted the interaction of bevantolol with a strong electrostatic value (Fig. 4B and Table S7, ESI†). | ||
| Fig. 4 Per-residue energy decomposition of (A) ponatinib, (B) bevantolol, and (C) 3-hydroxy glabrol. | ||
In the case of ponatinib, Asp641 and Glu531 work against complex formation, while Ile545 and Ala564 (from the Hinge region) promote it (Fig. 4A). Asp641 shows a strong positive free energy value supported by its high polar energy contribution (Table S7, ESI†). 3-Hydroxy glabrol has Val498 and Leu636, which encouraged the interaction with FGFR1 (Fig. 4C). This outcome is consistent with bevantolol's increased binding free energy compared to the others. All things considered, identifying the essential residues in complex formation might be beneficial for creating effective treatments against FGFR1.
Additionally, we extended the aforementioned findings by using Schrodinger Maestro software to analyze the final structure of each simulation. Consequently, Fig. 5 shows several H-bonds and hydrophobic interactions of the ligand. The 3D orientation of FGFR1/inhibitor complexes is shown in Fig. 5(A)–(C), emphasizing the binding residues. The cyan colour represents the ligand, and the pink colour shows its neighbouring residues in a range of 3 Å. Here, we observed that common residues like Ile545, Glu568, and Asp641 are present in the vicinity of the inhibitor for all the cases. A similar residual connection was observed from the 2D profile, suggesting they are both in sync. Dark lime-green residues indicate hydrophobic interactions, while a single pink arrow line indicates hydrogen bonding. We created 2D ligand–protein interaction profiles using Schrödinger Maestro for ponatinib and the two FGFR1-bound leads. This interaction profile displays the primary residues engaged in various interactions, such as polar contributions, hydrogen bonds, and hydrophobic contacts (Fig. 5(D)–(F)). There are nineteen hydrophobic contacts and two H-bond interactions between Asp641 and Glu531 (Fig. 5(D)). Similarly, Ala570 and Glu568 from the hinge region create H-bond interactions in the FGFR1/3-hydroxy glabrol complex (Fig. 5(E)). The ligand interaction in the FGFR1/bevantolol complex was supported by a Glu537 and Asp647 H-bond along with other hydrophobic interactions (Fig. 5(F)). Several important critical residues often appeared, such as Asp641, Glu568, Asp647, and Met541, which is consistent with their favourable effect on binding free enthalpy. These crucial residues could help choose novel FGFR1 inhibitors.
 | ||
| Fig. 5 Protein–ligand interaction profile of FGFR1 with lead compounds. (A) and (D) Ponatinib. (B) and (E) 3-Hydroxy glabrol. (C) and (F) Bevantolol. | ||
3.7. Residual correlation motion
Using the dynamics cross-correlation matrix (DCCM) approach, the effect of ligand binding on the kinetics of the FGFR1 residues was investigated. Consequently, a 3D structural illustration of residual motions was shown. Correlated and anti-correlated movements are shown by the red and blue lines, respectively. Four key areas demonstrating how the residual connection pattern varies across all systems are highlighted (Fig. 6). In contrast to the apo structure, the FGFR1/molecule complexes show more synchronized residual motion. | ||
| Fig. 6 Correlation and anti-correlation motion of residues displayed as DCCM matrix and 3D structures for (A) apo (B) ponatinib (C) bevantolol and (D) 3-hydroxy glabrol. | ||
The activation segment region (641–670), denoted by R1, exhibits varied correlation movements regarding R3. A similar observation was obtained for R2, where 3-hydroxy glabrol describes the strong positive movement with scattered pink colour. The R3 region highlights the coupled mobility between P-loop (481–491) and hinge (562–569), implying nearly identical residual motion in all systems. However, a decline in anti-correlation motion was seen for complex systems. The αChelix region has contributed efficiently to the binding of ligands, as discussed earlier (Fig. 4). This region displays elevated positive correlation motion in ponatinib and bevantolol, in contrast with the P-loop specified by R4. Due to their greater efficacy in ligand binding, 3-hydroxy glabrol and bevantolol showed a decrease in anti-correlation movement. Based on the different degrees of linked motions of the complex structures, the DCCM analysis shows that ligand binding stabilized the interacting regions of FGFR1.
3.8. Protein structure network analysis
To comprehend the changes brought about by FGFR1's complexation with the lead compounds, the PSN of the complexes and apo were examined using the webPSN v2.0 web server. The nodes in a protein structure network represent the amino acids, while the edges display their connections. Hubs are higher-degree nodes, and a node's degree is based on the number of direct links it creates. Regions with a larger concentration of intra-linked nodes are referred to as communities.Fig. 7(A)–(D) shows the communities arranged by rank, and Fig. 7(E)–(H) shows the hubs arranged by the strength of their interactions. Detailed information on the network features is provided in Table S4 (ESI†). Among the apo and complex systems, the total number of hubs and communities differed in an intriguing way. The inhibitor-bound structures for FGFR1/ponatinib, FGFR1/3-hydroxy glabrol and FGFR1/bevantolol have 32, 50, and 57 hubs, respectively, compared to the apo structure's 37 hubs. The complexes had greater stability and robustness in the top-ranked C1 community than the others. Following complexation, more nodes and linkages were provided to the communities, with bevantolol showing the highest number of nodes (298) and links within communities (146). These compared PSN data reveal structural dynamics that could be a key area for creating novel medicines targeting FGFR1.
3.9. Optimization of bevantolol through structural analogues
We used the DeLA-Drug48 web server to produce one hundred structural analogues of bevantolol, which were then assessed against FGFR1 in a high-throughput virtual screening method. The two best analogues (ANLG-1 and ANLG-2) demonstrated docking scores of −11.52 and −11.21 kcal mol−1 (Table S6, ESI†). Subsequently, these compounds were examined using a 200 ns atomistic MD simulation in a replica. Fig. 8(A)–(C) shows the conformations of the related analogues. The time series of the Cα backbone RMSD of complexes over the simulation is displayed in Fig. 8(D). The RMSD stabilized for ANLG-2 during the simulation demonstrating the simulations' convergence. ANLG-2 has a smaller ligand protein distance, indicating a stable interaction (Fig. 8(E)). Based on this analysis, we have employed ANLG-2 for ligand Gaussian accelerated molecular dynamics (LiGaMD) simulation for further understanding. | ||
| Fig. 8 2D structures of (A) bevantolol (B) ANLG-1 and (C) ANLG-2. Time series of (D) backbone Cα RMSD and (E) ligand–protein distance for FGFR1/ANLG complex throughout the 200 ns simulation. | ||
The ligand–protein distances and binding pocket RMSDs of the replica 4000 ns LiGaMD simulation trajectories were investigated using AmberTools CPPTRAJ software. To get the 1D and 2D PMF profiles, the trajectories were reweighted utilizing the PyReweighting method. The quantitative characterization of ligand binding to FGFR1 was done using the 1D PMF profiles (Fig. S7, ESI†). A unique free energy basin was obtained at a ligand–protein distance of ∼2.0 Å, signifying the bound state of ANLG-2. The ligand binding process was further explored by analyzing the complex system's conformational changes using the respective 2D free energy profile (Fig. 9A).
It displayed four significant local energy minima states: one complex bound state, one intermediate and two unbound states. The lowest energy signifying the bound state was found at a distance of ∼2 Å and a binding pocket RMSD of ∼1.25 Å. The lowest PMFs for the intermediate state exhibited CM distances of 8–10 Å and binding pocket RMSDs of 1.50–1.75 Å. The first unbound state was obtained at a distance of ∼14–16 Å and pocket RMSDs of ∼1.7–2.0 Å. The second unbound state was seen at a lig-prot distance of ∼38–40 Å and an RMSD range of 1.7–2.0 Å. The primary states of the 2D free energy profiles were largely compatible with the significant conformations of the 1D PMF when the two profiles were compared. The key states of 2D FES provided structural details on the unbinding and binding between the ligand and receptor. Structural and binding mode analyses were performed on the minimum conformations to identify the ligand binding process.
The K-means clustering method was used to identify the minimum conformations from the 2D-PMF plot (Fig. 9(B)–(E)). In the bound state, the ligand interacts with residues from the activation loop, P-loop, αC helix and αCβ4 loop of FGFR1 kinase. As it moved towards the intermediate state, the ligand slipped away from the binding pocket. It shows a residual connection with the αC helix and αCβ4 loop (Fig. 9(C)). The unbound states display no interaction between receptor and ANLG-2 (Fig. 9(D) and (E)).
To determine the kinetic rate constants of interaction with FGFR1 kinase, the LiGaMD (Dual) simulations were analyzed using Kramers’ rate theory and predicted values of 1D PMF. The binding (kon) and dissociation (koff) rate constants for ANLG-2 were 3.1 ± 0.41 × 106 M−1 s−1 and 2.3 ± 0.59 × 10−2 M−1 s−1, respectively. The dissociation binding constant (KD = koff/kon) was 7.4 ± 0.27 nM. As a type II tyrosine kinase inhibitor, ponatinib shows an IC50 value of 1.5 nM in hematologic cells63 and 40 nM in non-small-cell lung cancer.64 The experimentally determined dissociation constant (KD) value was is 7 nM to 8 nM.26 The predicted KD value has almost similar binding affinity as the experimental value. The predictions were sound and offered dependable insights into the interactions of ANLG-2 with FGFR1 kinase.
4. Conclusion
FGFR1 is linked to numerous downstream signalling cascades that are related to homeostasis and growth. Any imbalance can result in a variety of illnesses, including cancer. A number of FDA-approved medications have received approval as FGFR1 therapies. However, kinase specificity concerns and developing resistance challenge the treatment of several ailments. In an attempt to propose a better inhibitor against FGFR1, we performed a structure-based virtual screening of 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 624 compounds, taking FGFR1 as the target. Two databases, HMDB and NPASS, have been used for this study. We conducted a molecular docking-based screening of the small molecule structures using the virtual screening workflow (VSW) methodology of the Schrödinger suite. Then, we selected 27 lead molecules based on a docking score cutoff of −11 kcal mol−1. Four compounds were chosen after ADMET analysis and subjected to additional study using 2 × 200 ns atomistic molecular dynamics simulations and MM-PBSA-based free energy calculations. All the systems were stable in equilibrium throughout the 200 ns production simulation. By analysing the binding affinities of selected compounds, we found two lead molecules from the HMDB with higher free energy than the control drug ponatinib (ΔG = −16 kcal mol−1). Bevantolol has the highest binding efficacy, followed by 3-hydroxy glabrol, pawhuskin B and daurichromene A. This MD-MMPBSA analysis showed that electrostatic interactions are essential for the binding differences in bevantolol. Further, the hinge and αChelix region of FGFR1 played a crucial role in protein–ligand interaction as displayed by the per residue energy contribution and ligand interaction profile (Fig. 4 and 5). Based on all the analysis, we selected bevantolol for further optimization.
624 compounds, taking FGFR1 as the target. Two databases, HMDB and NPASS, have been used for this study. We conducted a molecular docking-based screening of the small molecule structures using the virtual screening workflow (VSW) methodology of the Schrödinger suite. Then, we selected 27 lead molecules based on a docking score cutoff of −11 kcal mol−1. Four compounds were chosen after ADMET analysis and subjected to additional study using 2 × 200 ns atomistic molecular dynamics simulations and MM-PBSA-based free energy calculations. All the systems were stable in equilibrium throughout the 200 ns production simulation. By analysing the binding affinities of selected compounds, we found two lead molecules from the HMDB with higher free energy than the control drug ponatinib (ΔG = −16 kcal mol−1). Bevantolol has the highest binding efficacy, followed by 3-hydroxy glabrol, pawhuskin B and daurichromene A. This MD-MMPBSA analysis showed that electrostatic interactions are essential for the binding differences in bevantolol. Further, the hinge and αChelix region of FGFR1 played a crucial role in protein–ligand interaction as displayed by the per residue energy contribution and ligand interaction profile (Fig. 4 and 5). Based on all the analysis, we selected bevantolol for further optimization.
Using the DeLA-Drug web server, 100 analogues of bevantolol were generated and assessed through the HTVS method of the Schrödinger suite. The top two compounds with high docking scores then underwent 2 × 200 ns MD simulation. Further, we selected ANLG-2 for LiGaMD (Dual) simulation based on the trajectory analysis. The bound, intermediate and unbound states were identified from the 2D-free energy plot. Using Kramer's rate theory, we computed the kinetic rate constant of receptor–ligand interactions. The KD value was 7.4 nM, which agrees with the experimental KD value of the control drug ponatinib. Overall, this study provides valuable insights into the kinetics of FGFR1–ligand interactions and highlights bevantolol along with its two analogues. ANLG-2 can be presented as a potential drug candidate for FGFR1 therapeutic intervention.
Author contributions
PK conceived and supervised the project. SM conducted simulations and data analysis and prepared the draft manuscript. PK edited the manuscript. All authors have approved the final version of the manuscript.Ethics statement
There are no human or animal experiments in this study.Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its ESI.† Two databases, namely, NPASS (https://bidd.group/NPASS/) and HMDB (https://hmdb.ca), were used for this study. Virtual screening and molecular docking data were obtained using Schrödinger (https://www.schrodinger.com/). The ADMET analysis was performed with the ADMETlab 2.0 server (https://admetmesh.scbdd.com). The conventional molecular dynamics simulation data were obtained using AMBER18, a licensed software for both academic and commercial use. It can be purchased on the https://ambermd.org website. The DeLA-Drug server was used to generate structural analogues (https://www.ba.ic.cnr.it/softwareic/deladrugportal/). The ligand Gaussian accelerated molecular dynamics (LiGaMD) data were obtained using the AMBER20 software. The GaMD reweighting was carried out with a toolkit of Python scripts generated by the Miao lab, which is freely available at https://www.med.unc.edu/pharm/miaolab/resources/pyreweighting/. Post-simulation analyses were performed using the cpptraj module associated with the AMBER18 and AmberTools19 packages. The AmberTools19 package used can be accessed freely upon registration at https://ambermd.org/GetAmber.php#ambertools, and its components are mostly released under the GNU General Public License (GPL). The in-house scripts for plotting were developed using Python3, and they are freely available at https://www.python.org/downloads. Various Python3 modules were used, such as Pandas, NumPy, Scipy, and Matplotlib. The molecular visualization software UCSF Chimera version 1.15 was used to analyze the structure of the protein. Chimera is available to non-commercial users under a distribution-specific license (https://www.cgl.ucsf.edu/chimera/). For protein structure network (PSN) analysis, the webPSN server was used, which is available free at https://webpsn.hpc.unimo.it/wpsn3.php.Conflicts of interest
The authors declare that there is no conflict of interest.Acknowledgements
This work was partially supported by the Department of Science and Technology, Govt. of India (grant number DST/NSM/R&D_HPC_Applications/2021/03.18 and SR/FST/LS-I/2020/621). SM thanks the Ministry of Education, Govt. of India, for providing a doctoral fellowship under the Prime Minister's Research Fellows (PMRF) scheme.References
- M. V. Dieci, M. Arnedos, F. Andre and J. C. Soria, Fibroblast growth factor receptor inhibitors as a cancer treatment: from a biologic rationale to medical perspectives, Cancer Discovery, 2013, 3, 264–279 CrossRef CAS PubMed.
- S. Mahapatra, N. A. Jonniya, S. Koirala, K. D. Ursal and P. Kar, The FGF/FGFR signalling mediated anti-cancer drug resistance and therapeutic intervention, J. Biomol. Struct. Dyn., 2023, 1–25 CAS.
- N. Zhou, Y. Xu, X. Liu, Y. Wang, J. Peng, X. Luo, M. Zheng, K. Chen and H. Jiang, Combinatorial Pharmacophore-Based 3D-QSAR Analysis and Virtual Screening of FGFR1 Inhibitors, Int. J. Mol. Sci., 2015, 16, 13407–13426 CrossRef CAS PubMed.
- S. Dai, Z. Zhou, Z. Chen, G. Xu and Y. Chen, Fibroblast Growth Factor Receptors (FGFRs): Structures and Small Molecule Inhibitors, Cells, 2019, 8, E614 CrossRef PubMed.
- M. Sleeman, J. Fraser, M. McDonald, S. Yuan, D. White, P. Grandison, K. Kumble, J. D. Watson and J. G. Murison, Identification of a new fibroblast growth factor receptor, FGFR5, Gene, 2001, 271, 171–182 CrossRef CAS PubMed.
- A.-M. Chioni and R. P. Grose, Biological Significance and Targeting of the FGFR Axis in Cancer, Cancers, 2021, 13, 5681 CrossRef CAS PubMed.
- I. S. Babina and N. C. Turner, Advances and challenges in targeting FGFR signalling in cancer, Nat. Rev. Cancer, 2017, 17, 318–332 CrossRef CAS PubMed.
- T. Helsten, S. Elkin, E. Arthur, B. N. Tomson, J. Carter and R. Kurzrock, The FGFR Landscape in Cancer: Analysis of 4853 Tumors by Next-Generation Sequencing, Clin. Cancer Res., 2016, 22, 259–267 CrossRef CAS PubMed.
- J. Drago, L. Formisano, D. Juric, A. Niemierko, A. Servetto, S. Wander, L. Spring, N. Vidula, J. Peppercorn, J. Younger, G. Malvarosa, M. Yuen, D. Sgroi, S. J. Isakoff, B. Moy, L. W. Ellisen, J. Iafrate, C. L. Arteaga and A. Bardia, FGFR1 gene amplification mediates endocrine resistance but retains TORC sensitivity in metastatic hormone receptor positive (HR+) breast cancer, Clin. Cancer Res., 2019, 25, 6443–6451 CrossRef CAS PubMed.
- A. Desai and A. A. Adjei, FGFR Signaling as a Target for Lung Cancer Therapy, J. Thorac. Oncol., 2016, 11, 9–20 CrossRef PubMed.
- Q. Lin, X. Chen, L. Qu, M. Guo, H. Wei, S. Dai, L. Jiang and Y. Chen, Characterization of the cholangiocarcinoma drug pemigatinib against FGFR gatekeeper mutants, Commun. Chem., 2022, 5, 100 CAS.
- Y. Loriot, A. Necchi, S. H. Park, J. Garcia-Donas, R. Huddart, E. Burgess, M. Fleming, A. Rezazadeh, B. Mellado, S. Varlamov, M. Joshi, I. Duran, S. T. Tagawa, Y. Zakharia, B. Zhong, K. Stuyckens, A. Santiago-Walker, P. De Porre, A. O'Hagan, A. Avadhani and A. O. Siefker-Radtke, and BLC2001 Study Group, Erdafitinib in Locally Advanced or Metastatic Urothelial Carcinoma, N. Engl. J. Med., 2019, 381, 338–348 CAS.
- C. D. Sohl, M. R. Ryan, B. Luo, K. M. Frey and K. S. Anderson, Illuminating the molecular mechanisms of tyrosine kinase inhibitor resistance for the FGFR1 gatekeeper mutation: the Achilles' heel of targeted therapy, ACS Chem. Biol., 2015, 10, 1319–1329 CAS.
- K. Sanachai, P. Mahalapbutr, K. Choowongkomon, R. P. Poo-arporn, P. Wolschann and T. Rungrotmongkol, Insights into the Binding Recognition and Susceptibility of Tofacitinib toward Janus Kinases, ACS Omega, 2020, 5, 369–377 CAS.
- Y.-T. Wang, J.-M. Liao, W.-W. Lin, C.-C. Li, B.-C. Huang, T.-L. Cheng and T.-C. Chen, Structural insights into Nirmatrelvir (PF-07321332)-3C-like SARS-CoV-2 protease complexation: a ligand Gaussian accelerated molecular dynamics study, Phys. Chem. Chem. Phys., 2022, 24, 22898–22904 CAS.
- N. A. Jonniya, S. Poddar, S. Mahapatra and P. Kar, Computer-aided Affinity Enhancement of a Cross-reactive Antibody against Dengue Virus Envelope Domain III, Cell Biochem. Biophys., 2023, 81, 737–755 CAS.
- D. Hamelberg, J. Mongan and J. A. McCammon, Accelerated molecular dynamics: a promising and efficient simulation method for biomolecules, J. Chem. Phys., 2004, 120, 11919–11929 CrossRef CAS PubMed.
- L. C. T. Pierce, R. Salomon-Ferrer, C. Augusto, F. de Oliveira, J. A. McCammon and R. C. Walker, Routine Access to Millisecond Time Scale Events with Accelerated Molecular Dynamics, J. Chem. Theory Comput., 2012, 8, 2997–3002 CrossRef CAS PubMed.
- R. D. Schaeffer, A. Fersht and V. Daggett, Combining experiment and simulation in protein folding: closing the gap for small model systems, Curr. Opin. Struct. Biol., 2008, 18, 4–9 CrossRef CAS PubMed.
- F. Khalili-Araghi, J. Gumbart, P.-C. Wen, M. Sotomayor, E. Tajkhorshid and K. Schulten, Molecular dynamics simulations of membrane channels and transporters, Curr. Opin. Struct. Biol., 2009, 19, 128–137 CrossRef CAS PubMed.
- Y. Miao, A. Bhattarai and J. Wang, Ligand Gaussian Accelerated Molecular Dynamics (LiGaMD): Characterization of Ligand Binding Thermodynamics and Kinetics, J. Chem. Theory Comput., 2020, 16, 5526–5547 CrossRef CAS PubMed.
- Y. Miao, V. A. Feher and J. A. McCammon, Gaussian Accelerated Molecular Dynamics: Unconstrained Enhanced Sampling and Free Energy Calculation, J. Chem. Theory Comput., 2015, 11, 3584–3595 CrossRef CAS PubMed.
- J. Chen, J. Wang, W. Yang, L. Zhao and G. Hu, Conformations of KRAS4B Affected by Its Partner Binding and G12C Mutation: Insights from GaMD Trajectory-Image Transformation-Based Deep Learning, J. Chem. Inf. Model., 2024, 64, 6880–6898 CrossRef CAS PubMed.
- J. Chen, J. Wang, W. Yang, L. Zhao, J. Zhao and G. Hu, Molecular Mechanism of Phosphorylation-Mediated Impacts on the Conformation Dynamics of GTP-Bound KRAS Probed by GaMD Trajectory-Based Deep Learning, Molecules, 2024, 29, 2317 CrossRef CAS PubMed.
- S. Pawnikar and Y. Miao, Pathway and mechanism of drug binding to chemokine receptors revealed by accelerated molecular simulations, Future Med. Chem., 2020, 12, 1213–1225 CrossRef CAS PubMed.
- J. A. Tucker, T. Klein, J. Breed, A. L. Breeze, R. Overman, C. Phillips and R. A. Norman, Structural insights into FGFR kinase isoform selectivity: diverse binding modes of AZD4547 and ponatinib in complex with FGFR1 and FGFR4, Struct. Lond. Engl., 2014, 22, 1764–1774 CAS.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng and T. E. Ferrin, UCSF Chimera-a visualization system for exploratory research and analysis, J. Comput. Chem., 2004, 25, 1605–1612 CrossRef CAS PubMed.
- N. Eswar, D. Eramian, B. Webb, M.-Y. Shen and A. Sali, Protein structure modeling with MODELLER, Methods Mol. Biol., 2008, 426, 145–159 CrossRef CAS PubMed.
- G. M. Sastry, M. Adzhigirey, T. Day, R. Annabhimoju and W. Sherman, Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments, J. Comput.-Aided Mol. Des., 2013, 27, 221–234 CrossRef PubMed.
- M. H. M. Olsson, C. R. Søndergaard, M. Rostkowski and J. H. Jensen, PROPKA3: Consistent Treatment of Internal and Surface Residues in Empirical pKa Predictions, J. Chem. Theory Comput., 2011, 7, 525–537 CrossRef CAS PubMed.
- E. Harder, W. Damm, J. Maple, C. Wu, M. Reboul, J. Y. Xiang, L. Wang, D. Lupyan, M. K. Dahlgren, J. L. Knight, J. W. Kaus, D. S. Cerutti, G. Krilov, W. L. Jorgensen, R. Abel and R. A. Friesner, OPLS3: A Force Field Providing Broad Coverage of Drug-like Small Molecules and Proteins, J. Chem. Theory Comput., 2016, 12, 281–296 CrossRef CAS PubMed.
- R. A. Friesner, J. L. Banks, R. B. Murphy, T. A. Halgren, J. J. Klicic, D. T. Mainz, M. P. Repasky, E. H. Knoll, M. Shelley, J. K. Perry, D. E. Shaw, P. Francis and P. S. Shenkin, Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy, J. Med. Chem., 2004, 47, 1739–1749 CrossRef CAS PubMed.
- G. Xiong, Z. Wu, J. Yi, L. Fu, Z. Yang, C. Hsieh, M. Yin, X. Zeng, C. Wu, A. Lu, X. Chen, T. Hou and D. Cao, ADMETlab 2.0: an integrated online platform for accurate and comprehensive predictions of ADMET properties, Nucleic Acids Res., 2021, 49, W5–W14 CrossRef CAS PubMed.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB, J. Chem. Theory Comput., 2015, 11, 3696–3713 CAS.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, Development and testing of a general amber force field, J. Comput. Chem., 2004, 25, 1157–1174 CAS.
- Z. Sun and P. Procacci, Methodological and force field effects in the molecular dynamics-based prediction of binding free energies of host–guest systems, Phys. Chem. Chem. Phys., 2024, 26, 19887–19899 RSC.
- X. He, V. H. Man, W. Yang, T.-S. Lee and J. Wang, A fast and high-quality charge model for the next generation general AMBER force field, J. Chem. Phys., 2020, 153, 114502 CrossRef CAS PubMed.
- D. J. Price and C. L. Brooks, A modified TIP3P water potential for simulation with Ewald summation, J. Chem. Phys., 2004, 121, 10096–10103 CrossRef CAS PubMed.
- A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations – Kräutler – 2001 – Journal of Computational Chemistry – Wiley Online Library, https://onlinelibrary.wiley.com/doi/10.1002/1096-<?pdb_no 987X?>987X<?pdb END?>(20010415)22:5%3C501::AID-JCC1021%3E3.0.CO;2-V, (accessed May 18, 2023).
- T. Darden, D. York and L. Pedersen, Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- R. J. Loncharich, B. R. Brooks and R. W. Pastor, Langevin dynamics of peptides: the frictional dependence of isomerization rates of N-acetylalanyl-N′-methylamide, Biopolymers, 1992, 32, 523–535 CrossRef CAS PubMed.
- H. J. C. Berendsen, J. P. M. Postma, W. F. van Gunsteren, A. DiNola and J. R. Haak, Molecular dynamics with coupling to an external bath, J. Chem. Phys., 1984, 81, 3684–3690 CrossRef CAS.
- D. R. Roe and T. E. I. Cheatham, PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data, J. Chem. Theory Comput., 2013, 9, 3084–3095 CrossRef CAS PubMed.
- S. Mahapatra and P. Kar, Computational biophysical characterization of the effect of gatekeeper mutations on the binding of ponatinib to the FGFR kinase, Arch. Biochem. Biophys., 2024, 758, 110070 CrossRef CAS PubMed.
- S. Mahapatra, N. A. Jonniya, S. Koirala and P. Kar, Molecular dynamics simulations reveal phosphorylation-induced conformational dynamics of the fibroblast growth factor receptor 1 kinase, J. Biomol. Struct. Dyn., 2024, 42, 2929–2941 CrossRef CAS PubMed.
- P. Kar, M. Seel, U. H. E. Hansmann and S. Höfinger, Dispersion terms and analysis of size- and charge dependence in an enhanced Poisson-Boltzmann approach, J. Phys. Chem. B, 2007, 111, 8910–8918 CrossRef CAS PubMed.
- H. Gohlke, C. Kiel and D. A. Case, Insights into protein–protein binding by binding free energy calculation and free energy decomposition for the Ras-Raf and Ras-RalGDS complexes, J. Mol. Biol., 2003, 330, 891–913 CrossRef CAS PubMed.
- T. M. Creanza, G. Lamanna, P. Delre, M. Contino, N. Corriero, M. Saviano, G. F. Mangiatordi and N. Ancona, DeLA-Drug: A Deep Learning Algorithm for Automated Design of Druglike Analogues, J. Chem. Inf. Model., 2022, 62, 1411–1424 CrossRef CAS PubMed.
- Y. Miao, W. Sinko, L. Pierce, D. Bucher, R. C. Walker and J. A. McCammon, Improved Reweighting of Accelerated Molecular Dynamics Simulations for Free Energy Calculation, J. Chem. Theory Comput., 2014, 10, 2677–2689 CrossRef CAS PubMed.
- D. Hamelberg, T. Shen and J. Andrew McCammon, Relating kinetic rates and local energetic roughness by accelerated molecular-dynamics simulations, J. Chem. Phys., 2005, 122, 241103 CrossRef PubMed.
- A. T. Frank and I. Andricioaei, Reaction Coordinate-Free Approach to Recovering Kinetics from Potential-Scaled Simulations: Application of Kramers' Rate Theory, J. Phys. Chem. B, 2016, 120, 8600–8605 CrossRef CAS PubMed.
- P. H. Hünenberger, A. E. Mark and W. F. van Gunsteren, Fluctuation and Cross-correlation Analysis of Protein Motions Observed in Nanosecond Molecular Dynamics Simulations, J. Mol. Biol., 1995, 252, 492–503 CrossRef PubMed.
- B. J. Grant, A. P. C. Rodrigues, K. M. ElSawy, J. A. McCammon and L. S. D. Caves, Bio3d: an R package for the comparative analysis of protein structures, Bioinf. Oxf. Engl., 2006, 22, 2695–2696 CAS.
- W. Sun, The Relationship Between Low-Frequency Motions and Community Structure of Residue Network in Protein Molecules, J. Comput. Biol., 2018, 25, 103–113 CrossRef CAS PubMed.
- A. Felline, M. Seeber and F. Fanelli, webPSN v2.0: a webserver to infer fingerprints of structural communication in biomacromolecules, Nucleic Acids Res., 2020, 48, W94–W103 CrossRef CAS PubMed.
- Y. Cao, A. Charisi, L.-C. Cheng, T. Jiang and T. Girke, ChemmineR: a compound mining framework for R, Bioinforma. Oxf. Engl., 2008, 24, 1733–1734 CrossRef CAS PubMed.
- T. W. H. Backman, Y. Cao and T. Girke, ChemMine tools: an online service for analyzing and clustering small molecules, Nucleic Acids Res., 2011, 39, W486–491 CrossRef CAS PubMed.
- Y. Cao, T. Jiang and T. Girke, A maximum common substructure-based algorithm for searching and predicting drug-like compounds, Bioinformatics, 2008, 24, i366–i374 CrossRef CAS PubMed.
- F. H. Tan, T. L. Putoczki, S. S. Stylli and R. B. Luwor, Ponatinib: a novel multi-tyrosine kinase inhibitor against human malignancies, OncoTargets Ther., 2019, 12, 635–645 CrossRef CAS PubMed.
- J. A. Tucker, T. Klein, J. Breed, A. L. Breeze, R. Overman, C. Phillips and R. A. Norman, Structural insights into FGFR kinase isoform selectivity: diverse binding modes of AZD4547 and ponatinib in complex with FGFR1 and FGFR4, Struct. Lond. Engl., 2014, 22, 1764–1774 CAS.
- Y. Yan, W. An, S. Mei, Q. Zhu, C. Li, L. Yang, Z. Zhao and J. Huo, Real-world research on beta-blocker usage trends in China and safety exploration based on the FDA Adverse Event Reporting System (FAERS), BMC Pharmacol. Toxicol., 2024, 25, 86 CrossRef PubMed.
- Y. J. Kim, S.-K. Jang, G. Kim, S.-E. Hong, C. S. Park, M.-K. Seong, H.-A. Kim, K. S. Kim, C.-H. Kim, K. S. Park, J. Hong, H.-O. Jin and I.-C. Park, Nebivolol Sensitizes BT-474 Breast Cancer Cells to FGFR Inhibitors, Anticancer Res., 2023, 43, 1973–1980 CrossRef CAS PubMed.
- T. O'Hare, W. C. Shakespeare, X. Zhu, C. A. Eide, V. M. Rivera, F. Wang, L. T. Adrian, T. Zhou, W.-S. Huang, Q. Xu, C. A. Metcalf, J. W. Tyner, M. M. Loriaux, A. S. Corbin, S. Wardwell, Y. Ning, J. A. Keats, Y. Wang, R. Sundaramoorthi, M. Thomas, D. Zhou, J. Snodgrass, L. Commodore, T. K. Sawyer, D. C. Dalgarno, M. W. N. Deininger, B. J. Druker and T. Clackson, AP24534, a Pan-BCR-ABL Inhibitor for Chronic Myeloid Leukemia, Potently Inhibits the T315I Mutant and Overcomes Mutation-Based Resistance, Cancer Cell, 2009, 16, 401–412 CrossRef PubMed.
- M. Ren, M. Hong, G. Liu, H. Wang, V. Patel, P. Biddinger, J. Silva, J. Cowell and Z. Hao, Novel FGFR inhibitor ponatinib suppresses the growth of non-small cell lung cancer cells overexpressing FGFR1, Oncol. Rep., 2013, 29, 2181–2190 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4cp04690k |
| This journal is © the Owner Societies 2025 |



