Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Are the pKa values of free fatty acids in aqueous solution abnormally high? An NMR and computational perspective
Themistoklis
Venianakis
 a,
Tjaša
Goričan
a,
Tjaša
Goričan
 b,
Georgios
Papamokos
b,
Georgios
Papamokos
 a,
Simona
Golič Grdadolnik
a,
Simona
Golič Grdadolnik
 b,
Michael G.
Siskos
a and
Ioannis P.
Gerothanassis
b,
Michael G.
Siskos
a and
Ioannis P.
Gerothanassis
 *a
*a
aSection of Organic Chemistry and Biochemistry, Department of Chemistry, University of Ioannina, Ioannina, GR 45110, Greece. E-mail: igeroth@uoi.gr
bNational Institute of Chemistry, Laboratory for Molecular Structural Dynamics, Theory Department, Hajdrihova 19, SI-1000, Ljubljana, Slovenia
First published on 17th November 2025
Abstract
Accurate dissociation constants of the carboxylic groups of free fatty acids in aqueous solution are unknown, and a wide range of values extending over 5 pKa units have been suggested. We reinvestigated the problem with the use of 1H and 13C NMR and DFT methods and demonstrate that the pKa values fall within those of typical organic acids.
Free fatty acids (FFAs) are the main building units of complex lipids, which are a wide group of biological macromolecules with numerous biological properties and an important role in health and nutrition.1 Short, medium and long chain fatty acids are the digestion products of lipids, which participate in numerous protein-FFA interactions, in the regulation of energy metabolism, and in several inflammation and signaling processes.2 FFAs have also been widely used in lipid-based formulations which enhance drug absorption and membrane permeability.3
Free fatty acids due to their hydrophobicity, exist as self-assembled aggregates in aqueous solution which depend on concentration, pH and temperature. Several studies have shown that medium and long chain fatty acids display a broad polymorphism as a function of these parameters. Thus, they display mainly lamellar structures at low pH and/or low temperature and spherical micelles at high pH and/or temperatures and worm-like aggregates in an intermediate regime.4 pH controls the ionization state of the carboxylic groups and, thus, the competition between hydrogen bond interactions of the COOH groups and the electrostatic repulsions between the COO− groups.
Several studies have shown that the pKa values of medium and long chain fatty acids depend on the chain length and values much higher than those for short chain analogues and typical organic carboxylic acids have been reported.5a–c It has been claimed that the pKa of free fatty acids increases from 6.5 to about 9.0, as the chain length of the fatty acid increases from C8 to C16, due to strong ion-dipole interactions of the carboxylate and carboxylic groups. For palmitic (hexadecanoic C16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0) acid the values of 9.7,5d 8.55e and 8.6–8.8,5b by titration, were reported. For stearic (octadecanoic C18
0) acid the values of 9.7,5d 8.55e and 8.6–8.8,5b by titration, were reported. For stearic (octadecanoic C18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0) acid, the pKa values of 9.05f and 10.155g were estimated. The pKa values for elaidic (18
0) acid, the pKa values of 9.05f and 10.155g were estimated. The pKa values for elaidic (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, trans-9), oleic (18
1, trans-9), oleic (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, cis-9), linoleic (18
1, cis-9), linoleic (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, cis-9,12) and α-linolenic (18
2, cis-9,12) and α-linolenic (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, cis-9,12,15) acid were found to be 9.95, 9.85, 9.24 and 8.28, respectively.5g It has been suggested that the degree of unsaturation has a significant effect on the pKa values since it affects the intermolecular distance between the fatty acid molecules due to the significant kinks of the chain. Recent quantum chemical methods reported that the lengthening of the fatty acid chain by one CH2 fragment leads to an increase in the pKa by 0.43 units; thus, for stearic acid (C18
3, cis-9,12,15) acid were found to be 9.95, 9.85, 9.24 and 8.28, respectively.5g It has been suggested that the degree of unsaturation has a significant effect on the pKa values since it affects the intermolecular distance between the fatty acid molecules due to the significant kinks of the chain. Recent quantum chemical methods reported that the lengthening of the fatty acid chain by one CH2 fragment leads to an increase in the pKa by 0.43 units; thus, for stearic acid (C18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0), a surface pKa = 10.05 was calculated,5h in very good agreement with one of the experimental values (pKa = 10.15).5h
0), a surface pKa = 10.05 was calculated,5h in very good agreement with one of the experimental values (pKa = 10.15).5h
From the above, it is evident that large uncertainties exist in the literature regarding the pKa of short, medium and long-chain fatty acids. The precise estimation of the pKa values of the free fatty acids (FFAs), however, is of primary importance in protein–ligand interactions since the ionization state of the carboxylic group plays a significant role in complexes with serum albumin,6a–g retinoid X receptor in the mouse brain6h and human brain FA binding protein.6i
The continuing disagreement over the pKa values of medium and long-chain FFAs and the increasing use of high values in the chemical literature led us to investigate whether the pKa values of free fatty acids in aqueous solution are abnormally high with the combined use of NMR and DFT methods. 1H and 13C NMR is a rapid and sensitive technique, which allows direct observation of the carboxylic groups undergoing deprotonation.
For caproleic acid, 9-decenoic acid (H2C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH(CH2)7COOH), NMR experiments were performed at concentrations c = 2 mM and 150 mM, which are below and above the critical micelle concentration, respectively. Fig. S1A(a) (SI) presents the 1H NMR spectrum of 150 mM caproleic acid at pD = 9.3. The spectrum remains essentially the same for a period of over 10 days (SI: Fig. S1A(b)). For pD values 10 to 6, the C
CH(CH2)7COOH), NMR experiments were performed at concentrations c = 2 mM and 150 mM, which are below and above the critical micelle concentration, respectively. Fig. S1A(a) (SI) presents the 1H NMR spectrum of 150 mM caproleic acid at pD = 9.3. The spectrum remains essentially the same for a period of over 10 days (SI: Fig. S1A(b)). For pD values 10 to 6, the C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif)
![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) COO protons do not show any inflection point. At pD < 6 the 1H NMR spectra are more complex. Thus, at pD = 1.9, the H2, H3, H8 and H10a,b resonances show the presence of two species (SI: Fig. S1B(b)). A non-linear regression analysis of the experimental δobs, of the major species, vs. pD values resulted in the pKa = 4.30 ± 0.05 (SI: Table S1). On standing of the solution for approximately one week, the major aggregate with the α-CH2 protons at δobs = 2.26 ppm was transformed to the second one at δobs = 2.37 ppm. The chemical shifts of the new major species as a function of pD, are shown in Fig. 1 and resulted in a pKa value identical to the previous one. The pKa value of both species (SI: Fig. S2 and Table S1) is not significantly different from that for concentrations below the critical micelle concentration (pKa = 3.79, SI: Table S1). This value is within the expected range of typical organic carboxylic acids, and it is significantly lower than those reported in the literature for free fatty acids with ten carbon chains.
COO protons do not show any inflection point. At pD < 6 the 1H NMR spectra are more complex. Thus, at pD = 1.9, the H2, H3, H8 and H10a,b resonances show the presence of two species (SI: Fig. S1B(b)). A non-linear regression analysis of the experimental δobs, of the major species, vs. pD values resulted in the pKa = 4.30 ± 0.05 (SI: Table S1). On standing of the solution for approximately one week, the major aggregate with the α-CH2 protons at δobs = 2.26 ppm was transformed to the second one at δobs = 2.37 ppm. The chemical shifts of the new major species as a function of pD, are shown in Fig. 1 and resulted in a pKa value identical to the previous one. The pKa value of both species (SI: Fig. S2 and Table S1) is not significantly different from that for concentrations below the critical micelle concentration (pKa = 3.79, SI: Table S1). This value is within the expected range of typical organic carboxylic acids, and it is significantly lower than those reported in the literature for free fatty acids with ten carbon chains.
 | ||
| Fig. 1 Aliphatic region of the 1H NMR spectra (500 MHz) of 150 mM caproleic acid in D2O at various pD values (number of scans = 16, T = 298 K). Experimental time of each NMR spectrum is 2 min. | ||
The ionization state of the carboxylic group was further confirmed with 1H-13C HSQC and HMBC experiments. The 13C chemical shifts of the α-CH2 carbon are essentially the same at pD > 6.5 (δ = 40.2 ppm), which demonstrates that there is no inflection point in this pD range (Fig. 2A). At the low pD = 2.15 (Fig. 2B), the 13C chemical shift (δobs = 36.7 ppm) shows a shielding by 3.5 ppm, which is in excellent agreement with values of aliphatic carboxylic acids (3.5 ppm and 3.9 ppm for propionic and valeric acid, respectively).7 Similarly, the 13C shift of the COOH carbon (δobs = 182.6 ppm) shows a deshielding upon deprotonation (δobs = 187.1 ppm) by 4.6 ppm, which is in excellent agreement with the value of 4.7 ppm for propionic and valeric acid.7 It is of interest that the two self-assembled aggregated species at pD = 2.15 show identical 13C chemical shifts of the α-CH2 (δobs = 36.7 ppm, Fig. 2B) and carboxylic carbons (δobs = 182.6 ppm, SI: Fig. S3), which demonstrates non-ionic COOH groups.
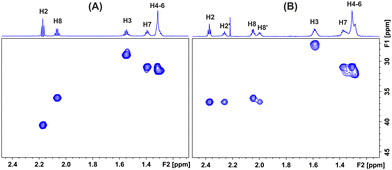 | ||
| Fig. 2 1H–13C HSQC NMR spectra (800 MHz) of 150 mM caproleic acid in D2O at: (A) pD 9.45 and (B) pD 2.15 (number of scans = 4, number of increments = 256, T = 298 K). Experimental time is 29 min. | ||
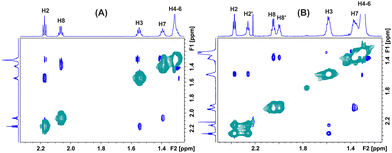
The 2D NOESY of caproleic acid at pD = 9.45 shows NOEs, which are anti-phase with respect to the diagonal (Fig. 3A); thus, the effective molecular weight of the self-aggregated species is within the extreme narrowing condition. Even with a long mixing time of 800 ms, no NOE cross-peaks were observed between the C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2–COO− and terminal C(10)HaHb/C(8)H2, which shows that the parallel arrangement of the ionic aggregated molecules is the most stable structural motif. At pD = 2.15, strong exchange cross-peaks between the two self-assembled aggregated species were observed (Fig. 3B). Again, NOE cross-peaks between the C
2–COO− and terminal C(10)HaHb/C(8)H2, which shows that the parallel arrangement of the ionic aggregated molecules is the most stable structural motif. At pD = 2.15, strong exchange cross-peaks between the two self-assembled aggregated species were observed (Fig. 3B). Again, NOE cross-peaks between the C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2–COOH and terminal C(10)HaHb/C(8)H2 were not observed, which confirms that the parallel arrangement of the non-ionic aggregated species is the most stable structural motif. It is of interest that the magnitudes of the NOE cross-connectivities between the H2 and H3 protons are different for the two aggregated species, presumably due to different effective molecular weights. This was verified with the use of DOSY experiments.
2–COOH and terminal C(10)HaHb/C(8)H2 were not observed, which confirms that the parallel arrangement of the non-ionic aggregated species is the most stable structural motif. It is of interest that the magnitudes of the NOE cross-connectivities between the H2 and H3 protons are different for the two aggregated species, presumably due to different effective molecular weights. This was verified with the use of DOSY experiments.
The DOSY experiment of caproleic acid at pD = 9.45 showed the presence of a unique self-aggregated species with diffusion constant D = 2.06 × 10−9 m2 s−1 (Fig. 4A). At pD = 2.10, the two self-aggregated species with α-C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2 resonances at δobs = 2.37 and 2.27 ppm showed significantly different diffusion constants with D = 5.2 × 10−10 m2 s−1 and 4 × 10−12 m2 s−1, respectively (Fig. 4B). Since both aggregated species show NOEs in the extreme narrowing condition (Fig. 3), it is evident that they exhibit significant backbone internal motions, despite the extremely slow diffusion mobility of the species with α-C
2 resonances at δobs = 2.37 and 2.27 ppm showed significantly different diffusion constants with D = 5.2 × 10−10 m2 s−1 and 4 × 10−12 m2 s−1, respectively (Fig. 4B). Since both aggregated species show NOEs in the extreme narrowing condition (Fig. 3), it is evident that they exhibit significant backbone internal motions, despite the extremely slow diffusion mobility of the species with α-C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2 at δobs = 2.27 ppm.
2 at δobs = 2.27 ppm.
 | ||
| Fig. 4 DOSY experiments of caproleic acid (150 mM in D2O) at: (A) pD = 9.45 and (B) pD = 2.15. Experimental time is 1 h, 40 min. | ||
NMR experiments for oleic acid were performed at a concentration of c = 2 mM, which is above the critical micelle concentration. Fig. 5 shows the 1H NMR chemical shifts of the α-CH2 group of oleic acid at different pD values. At intermediate pD values <6.5 different self-aggregated components were observed, which are slow on the NMR time scale. A well-defined titration curve with pKa = 3.95 (SI: Table S1) was observed, contrary to the literature value of pKa = 9.85.5g 2D 1H–13C HSQC experiments at pD ≥ 6.5 showed identical chemical shifts of the α-CH2 carbon (δobs = 40.6 ppm, SI: Fig. S4), which demonstrates the presence of the carboxylate group without any inflection point in this region. The 2D NOESY spectrum of oleic acid at pD = 10.9 shows that the NOEs are in-phase with respect to the diagonal (Fig. 6A), contrary to the case of caproleic acid. This demonstrates that the self-aggregated species are outside the extreme narrowing conditions. Of particular interest are the long-range NOE cross-connectivities between the H2 and H3 protons with the terminal CH3 group; this demonstrates that the cis-9 double bond induces a significant folding of the (CH2)n chains and, thus, through-space inter-molecular proximity of the H2 and H3 protons with the terminal CH3 group (see also the DFT calculations).
 | ||
Fig. 5 pD dependence of the α-C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2 protons of oleic acid (2 mM in D2O). Experimental time for each NMR spectrum is 32 min. 2 protons of oleic acid (2 mM in D2O). Experimental time for each NMR spectrum is 32 min. | ||
At pD = 2.15 the 1H NMR resonances become significantly broader than those at pD = 10.9 which is an indication of a larger MW of the self-aggregated species. The 2D 1H–13C HSQC experiment showed the non-ionic state of the carboxylic group (δobs = 36.7 ppm, SI: Fig. S4). The 2D NOESY spectra with mixing times 200, 400 and 800 ms did not show a long-range connectivity between the H2 and H3 protons with the terminal CH3 group, which demonstrates a different conformational state than that of pD = 10.9 (Fig. 6B).
Using density functional theory (DFT), the self-aggregation was simplified by considering dimers of caproleic acid and oleic acid in the neutral and anionic forms. Geometry optimizations and vibrational frequency calculations were carried out using Gaussian168 with the ωB97X-D9 density functional, which incorporates long-range correction and empirical dispersion.
An augmented correlation-consistent basis set, the aug-cc-pVDZ, was employed to model the more diffuse characteristics of the atomic anion.10 Frequency calculations were performed to confirm that all optimized geometries were true minima. For each system, the two molecules were initially arranged in a parallel configuration, and the total energy was minimized to determine whether this approach results in stabilization. For caproleic acid, a dimer was analyzed with an implicit solvent model (IEFPCM using water as the dielectric medium),11 starting from parallel structures, motivated by the NOE results. To reduce computational costs, the gas-phase optimized geometry of oleic acid was used as the initial guess for the subsequent re-optimization using the implicit solvation model. All reported structures and energies correspond to the final solvated minima. The results are presented in Fig. 7. For the caproleic acid dimer, the form of the carboxyl group produces a structure that deviates from parallelism (Fig. 7A and B) mainly due to the hydrogen bond formed between the carboxyl groups. In the anionic form, though, the parallelism is achieved primarily due to the hydrophobic effect (Fig. 7C and D).
 | ||
| Fig. 7 Energy minimized structures of caproleic acid (A) and (B), caproleic acid anion (C) and (D), and oleic acid anion (E)–(H) in two different configurations at the ω-B97X-D level of theory with the aug-cc-pVDZ basis set using the IEFPCM implicit solvation model.11 | ||
For oleic acid in its ionic form, a stable configuration was identified where one molecule adopts a folded conformation around the alkyl tail of another molecule, stabilized by hydrophobic interactions between the aliphatic chains (Fig. 7E and F). Motivated by the NOE data, we constructed an additional starting geometry in which the carboxylate group of one molecule was positioned to interact with the terminal methyl group of another. Geometry optimization of this setup produced a distinct, energetically minimized structure (Fig. 7G and H). Our calculations revealed a unique oleate dimer arrangement in which the carboxylate group approaches the terminal methyl of a second molecule (Fig. 7G and H), in excellent agreement with the NOE data. To our knowledge, such a carboxylate–methyl contact has not been explicitly reported in fatty acid dimers; related studies highlight head-to-head hydrogen bonding in neutral acids and mainly hydrophobic tail packing in carboxylates. Although CH⋯O interactions involving terminal methyl groups are inherently weak, their presence here supports the overall hydrophobic stabilization of the alkyl chains and shows a subtle geometric pattern consistent with the experimental data.
In summary, we have shown through the combined use of 1H/13C NMR and DFT calculations that the pKa values of caproleic acid and oleic acid in aqueous solution are not abnormally high and are within the pKa values of typical organic carboxylic acids.
This research was co-financed by the Hellenic Foundation for Research and Innovation (H.F.R.I.) under the “First Call for H.F.R.I. Research Projects to support faculty members and researchers and the procurement of high-cost research equipment grant” (project number 2050 to I. P. G.) and the Slovenian Research and Innovation Agency (ARIS) (research grants J1-4400 and P1-0010 to S. G. G. and T. G.).
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: NMR experiments, DFT calculations, and Cartesians of optimized structures. See DOI: https://doi.org/10.1039/d5cc05822h.Notes and references
- (a) D. E. Vance and J. E. Vance, Biochemistry of lipids, lipoproteins and membranes (new comprehensive biochemistry), Elsevier, Amsterdam, Netherlands, 5th edn, 2008 Search PubMed; (b) C. C. Akoh and D. B. MinFood lipids, chemistry, nutrition and biochemistry, Marcel Dekker Inc., New York, 2nd edn, 2002 Search PubMed; (c) C. Leray Dietary lipids for healthy brain function, CRC Press, USA, 2021 Search PubMed; (d) E. Alexandri, R. Ahmed, H. Siddiqui, M. I. Choudhary, C. G. Tsiafoulis and I. P. Gerothanassis, Molecules, 2017, 22, 1663 CrossRef PubMed.
- (a) O. Quehenberger and E. A. Dennis, N. Engl. J. Med., 2011, 365, 1812–1823 CrossRef CAS; (b) P. Yli-Jama, H. E. Meyer, J. Ringstad and J. I. Pedersen, J. Intern. Med., 2002, 251, 19–28 CrossRef CAS; (c) A. H. Huber and A. M. Kleinfeld, J. Lipid Res., 2017, 58, 578–585 CrossRef CAS.
- (a) C. J. Porter, N. L. Trevaskis and W. N. Charman, Nat. Rev. Drug Discovery, 2007, 6, 231 CrossRef CAS; (b) M. J. Hackett, J. L. Zang, W.-C. Shen, P. C. Guley and M. J. Cho, Adv. Drug Delivery Rev., 2013, 65, 1331 CrossRef CAS; (c) N. Fattachi, M.-A. Shahbazi, A. Maleki, M. Hamidi, A. Ramazani and H. A. Santos, J. Controlled Release, 2020, 326, 556 CrossRef.
- W. Ma, B. Yang, H. Zhang and D. Sun, J. Mol. Liq., 2023, 390, 123133 CrossRef CAS.
- (a) T. H. Haines, Proc. Natl. Acad. Sci. U. S. A., 1983, 80, 160–164 CrossRef CAS PubMed; (b) J. R. Kanicky, A. F. Poniatowski, N. R. Mehta and D. O. Shah, Langmuir, 2000, 16, 172–177 CrossRef CAS; (c) J. R. Kanicky and D. O. Shah, Langmuir, 2003, 19, 2034–2038 CrossRef CAS; (d) R. E. Heikkila, D. W. Deamer and D. G. Cornwel, J. Lipid. Res., 1970, 11, 195 CrossRef CAS PubMed; (e) R. A. Peters, Proc. R. Soc. London, Ser. A, 1931, 133, 140 CrossRef CAS; (f) A. P. Christodoulou and H. L. Rosano, Adv. Chem. Ser., 1968, 84, 210 CrossRef; (g) J. R. Kanicky and D. O. Shah, J. Colloid Interface Sci., 2002, 256, 201–207 CrossRef CAS; (h) Y. D. Vysotsky, E. S. Kartashynska, D. Vollhardt and V. D. Fainerman, J. Phys. Chem. C, 2020, 124, 13809 CrossRef CAS.
- (a) M. A. Kenyon and J. A. Hamilton, J. Lipid Res., 1994, 35, 458–467 CrossRef CAS; (b) I. Petitpas, T. Grüne, A. A. Bhattacharya and S. Curry, J. Mol. Biol., 2001, 314, 955–960 CrossRef CAS PubMed; (c) J. R. Simard, P. A. Zunszain, C.-E. Ha, J. S. Yang, N. V. Bhagavan, I. Petitpas, S. Curry and J. A. Hamilton, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 17958–17963 CrossRef CAS PubMed; (d) E. S. Krenzel, Z. Chen and J. A. Hamilton, Biochemistry, 2013, 52, 1559–1567 CrossRef CAS; (e) E. Alexandri, A. Primikyri, G. Papamokos, T. Venianakis, V. K. Gkalpinos, A. G. Tzakos, A. Karydis-Messinis, D. Moschovas, A. Avgeropoulos and I. P. Gerothanassis, FEBS J., 2022, 289, 5617–5636 CrossRef CAS; (f) E. Alexandri, T. Venianakis, A. Primikyri, G. Papamokos and I. P. Gerothanassis, Molecules, 2023, 28, 3724 CrossRef CAS PubMed; (g) T. Venianakis, A. Primikyri, T. Opatz, S. Petry, G. Papamokos and I. P. Gerothanassis, Molecules, 2023, 28, 7991 CrossRef CAS PubMed; (h) A. M. de Urquiza, S. Liu, M. Sjöberg, R. H. Zetterström, W. Griffiths, J. Sjövall and T. Perlman, Science, 2000, 290, 2140–2144 CrossRef CAS PubMed; (i) M. Rademacher, A. W. Zimmerman, H. Rüterjans, J. H. Veerkamp and C. Lücke, Mol. Cell. Biochem., 2002, 239, 61–68 CrossRef CAS PubMed.
- R. Hagen and J. D. Roberts, J. Am. Chem. Soc., 1969, 91, 4504 CrossRef CAS.
- M. J. Frisch et al. , Gaussian 16 Rev. B.01, Wallingford, CT, 2016 Search PubMed.
- J. D. Chai and M. Head-Gordon, J. Chem. Phys., 2008, 128(8), 084106 CrossRef.
- T. H. Dunning, J. Chem. Phys., 1989, 90(2), 1007–1023 CrossRef CAS.
- E. Cancès, B. Mennucci and J. Tomasi, J. Chem. Phys., 1997, 107, 3032–3041 CrossRef.
| This journal is © The Royal Society of Chemistry 2025 |