Open Access Article
Open Access ArticleCombining aptamers for thiamphenicol and chloramphenicol for detecting both antibiotics
Yun
Shu
ab,
Shiyuan
Liu
b and
Juewen
Liu
 *b
*b
aSchool of Chemistry and Chemical Engineering, Yangzhou University, Yangzhou, 225005, China
bDepartment of Chemistry, Waterloo Institute for Nanotechnology, University of Waterloo, Waterloo, Ontario N2L 3G1, Canada. E-mail: liujw@uwaterloo.ca
First published on 16th October 2025
Abstract
A DNA aptamer named TAP-1 for thiamphenicol (TAP) was selected using the capture-systematic evolution of ligands by exponential enrichment (SELEX) technique, exhibiting a dissociation constant (Kd) of 6.4 μM as determined by isothermal titration calorimetry (ITC). When combined with a previously selected chloramphenicol (CAP) aptamer, they collectively can differentiate TAP and CAP.
Chloramphenicol (CAP) is a key member of the amphenicol antibacterial family and has been used for a long time. Due to its severe side effects in humans and the rapid emergence of bacterial resistance to CAP, its clinical application has been restricted since the 1980 s in most countries.1 To overcome these limitations, many chloramphenicol derivatives, such as thiamphenicol (TAP) and fluorinated derivatives, have been synthesized, and some have demonstrated promising antibacterial potential.2 Currently, TAP is used as an alternative to CAP for animal treatment.3 Given its structural and physicochemical similarity to CAP, and the potential of non-compliance of withdrawal regulations, concerns have been raised regarding residue contamination of TAP in food and in the environment. Consequently, the detection of TAP and CAP simultaneously and individually is essential to ensure an accurate monitoring of CAP-class antibiotics, safeguard public health, and meet international food safety standards.4
In addition to instrument-based analytical techniques, such as liquid chromatography,5,6 immunoassays7,8 have been employed for the detection of TAP. For small-molecule targets, immunoassays typically employ a competitive format. In recent years, aptamer-based detection methods have gained increasing popularity.9–16 Aptamers are single-stranded oligonucleotides capable of selectively binding to target molecules. For small-molecule targets, aptamers offer multiple non-competitive detection strategies.13,17 Aptamers are typically identified through a process known as systematic evolution of ligands by exponential enrichment (SELEX).17–21 Although a few studies on the selection of CAP aptamers have been reported,22–24 no reports are available on TAP aptamers yet. Our group previously reported a CAP aptamer with a Kd value of 9.8 μM, but its binding to TAP was not characterized.23 In this work, we used TAP as a target for aptamer selection. The obtained aptamer, when combined with the previous CAP aptamer, can detect both analytes as a group or individually.
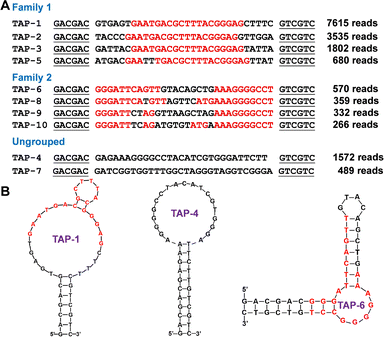
To select aptamers for TAP, the capture-SELEX method was used. In this method, a DNA library comprising a 30-nucleotide randomized region was immobilized on streptavidin-coated beads via hybridization to a short biotinylated complementary strand.25–27 TAP-binding sequences were selectively eluted after incubation with TAP, recovered, and PCR-amplified to seed the next round. Selection was performed in buffer (50 mM Tris–HCl, pH 7.5, 300 mM NaCl, 10 mM MgCl2) using 5 mM TAP for a total of 15 rounds. The selection progress was monitored by real-time PCR (RT-PCR) to quantify the TAP-eluted DNA in each round. A notable cycle threshold (Ct) shift was observed in round 14 (Fig. S1), suggesting aptamer enrichment. Deep sequencing of the round 15 library was performed. Out of the obtained 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 455 reads, two predominant sequence families were identified, with TAP-1 (32.5%), TAP-6 (2.4%) being the dominating sequence in each family (Fig. 1A). Among the top 10 sequences, TAP-4 and TAP-7 cannot be assigned to a family. Since TAP-4 is also an abundant sequence, it was included in our binding studies as well. The predicted secondary structures of these three aptamers are shown in Fig. 1B.
455 reads, two predominant sequence families were identified, with TAP-1 (32.5%), TAP-6 (2.4%) being the dominating sequence in each family (Fig. 1A). Among the top 10 sequences, TAP-4 and TAP-7 cannot be assigned to a family. Since TAP-4 is also an abundant sequence, it was included in our binding studies as well. The predicted secondary structures of these three aptamers are shown in Fig. 1B.
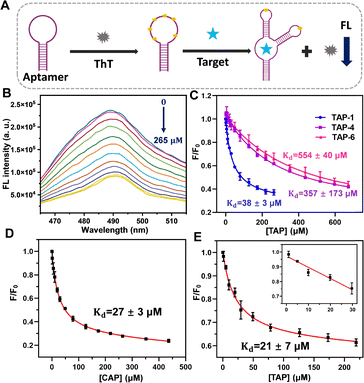
ThT fluorescence-based binding assays were then employed to evaluate the aptamers.19,28 ThT is a water-soluble fluorogenic dye that exhibits negligible intrinsic fluorescence as a free dye but undergoes marked enhancement in fluorescence upon binding to a single-stranded DNA, particularly G-quadruplexes.29 ThT binding to an aptamer produces strong fluorescence, whereas target binding can sometimes displace associated ThT, leading to fluorescence quenching (Fig. 2A).30 When TAP was titrated into a TAP-1/ThT mixture, the saturated fluorescence decrease reached more than 60% (Fig. 2B), yielding an apparent dissociation constant (Kd) of 38 ± 3 μM (Fig. 2C). If using this method as a sensor to detect TAP, a limit of detection (LOD) of 8.0 μM was obtained (Fig. S2). European Union (EU) has stipulated that the maximum residue of TAP in animal food should be less than 50 μg kg−1 (0.14 μM).31 While this ThT-based method cannot directly meet this requirement, enhanced sensitivity can be achieved via various signal amplification methods and pre-concentration of samples.32,33 For the TAP-4 and TAP-6 aptamers, TAP also induced a ∼60% fluorescence drop, although their apparent Kd values were much larger (Kd = 357 ± 173 μM and 554 ± 40 μM, respectively). The different apparent Kd values of these aptamers indicated that ThT was able to give useful information on binding, and TAP-1 appeared to be our best aptamer candidate.
Since TAP and CAP have similar structures, we also tested the binding of the TAP aptamers to CAP. The ThT fluorescence assay revealed strong binding of the TAP-1 aptamer to CAP, with >70% quenching (Kd = 27 ± 3 μM), indicating a similarly high binding affinity of the TAP-1 aptamer to CAP (Fig. 2D). Moreover, the binding of TAP-4 and TAP-6 to CAP was also determined (Kd = 373 ± 31 μM and 475 ± 46 μM, respectively), and these affinities were similar to their binding to TAP (Fig. S3). Therefore, the way of binding of the TAP aptamers to both TAP and CAP should be similar, where the sulfonate group in TAP contributed less to binding.
The ThT-based sensing platform was further evaluated using filtered lake water collected from an area near a steel plant (Burlington Bay, Lake Ontario). The water samples were buffered to contain 20 mM Tris-HCl (pH 7.5, 300 mM NaCl, 10 mM MgCl2), after which TAP-1 (500 nM) and ThT (1 μM) were added. Subsequently, varying concentrations of TAP were spiked into the buffered matrix, and the fluorescence response was recorded. The fluorescence decreased by 47%, yielding an apparent Kd of 21 ± 7 μM (Fig. 2E). A linear decrease in the fluorescence intensity was observed at low TAP concentrations, yielding a LOD of 8.5 μM (Fig. 2E). This performance was comparable to that obtained in clean buffer, validating the robustness of the sensing system in complex matrices.
Isothermal titration calorimetry (ITC) was subsequently used to confirm aptamer binding and to obtain thermodynamic parameters. Titration of TAP into the TAP-1 aptamer produced an exothermic reaction (Fig. 3A), with a Kd of 6.4 μM, which is 5.9-fold lower with the value obtained from the ThT assay, suggesting that ThT inhibited aptamer binding. The negative enthalpy change (ΔH = −9.5 kcal mol−1) and negative entropy change (ΔS = −8.1 cal K−1 mol−1) suggest that the binding is enthalpy-driven. In contrast, titration of CAP into the TAP-1 aptamer yielded a Kd of 13.8 μM (Fig. 3B), which is closer to the ThT-based measurement. All of the above results indicated that TAP-1 binds TAP and CAP with a comparable affinity. Titration of TAP into the TAP-4 (Kd = 16.6 μM) and TAP-6 (Kd = 27.4 μM) aptamers was also performed (Fig. S4). Thus, the relative affinities of the three aptamers for TAP, as determined by ThT fluorescence and ITC, were consistent. Given that TAP-1 has the best affinity, we focused on this aptamer in subsequent studies.
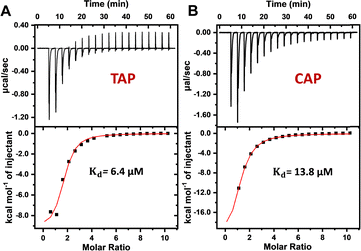 | ||
| Fig. 3 ITC results of aptamer binding by titrating 2 mM TAP (A) and CAP (B) into 40 μM TAP-1 aptamer. | ||
To further elucidate the binding mechanism of the TAP-1 aptamer, circular dichroism (CD) spectroscopy was employed to monitor its structural changes. The TAP-1 aptamer exhibited a positive peak near 280 nm and a negative peak around 240 nm, characteristic of a typical B-form DNA structure (Fig. S5). Upon the addition of TAP, the positive and negative peaks both underwent a blue shift. These spectral changes also support aptamer binding to TAP.
With reliable aptamers in hand, many signal transduction methods are available to develop biosensors, each having their advantages and disadvantages. For example, the strand-displacement reaction allows ‘signal-on’ detection but it requires costly covalent modification of DNA.19,34 While label-free gold nanoparticle (AuNP)-based colorimetric assays are simple to perform, the unintended interactions between target molecules and AuNPs often lead to artifacts.35 DNA staining dyes offer a simple, label-free, and cost-effective platform with high sensitivity, and simplicity, but it is more susceptible to interference.36,37 Nevertheless, it serves as a good method for proof-of-concept work. We validated thioflavin T (ThT) to be a general probe for aptamer binding assays and biosensors.30
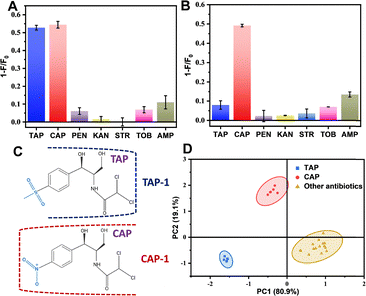
The TAP-1 aptamer was mixed with ThT to construct a label-free biosensor, and the sensor responses were evaluated in the presence of various antibiotics. As described above, this aptamer responded similarly to TAP and CAP, but it did not show signal changes to the other tested antibiotics (Fig. 4A). We then tried our previously published CAP-1 aptamer (isolated using CAP as a target molecule).23 Interestingly, this aptamer responded only to CAP (Fig. 4B). Therefore, we can deduce that the CAP-1 aptamer binds to CAP in the region that is different in TAP. Likely, binding of the CAP-1 aptamer to CAP involved the nitro group (Fig. 4C). Based on the different selectivity profile, we can use the TAP-1 and CAP-1 aptamers to form a small sensor array, which should be able to differentiate both analytes using principal component analysis (PCA).38 To construct the training dataset, the two target antibiotics (TAP and CAP) were measured in six replicate runs, while all other antibiotics were measured in triplicate. The training data were analyzed using the PCA analysis module from Origin software, and a representative score plot was generated (Fig. 4D). This score plot revealed clear separation of TAP and CAP from each other and from other antibiotics. Although this is only a training data, this distinct clustering highlights the feasibility of the array-based sensing strategy for selective antibiotics discrimination, despite the moderate selectivity of individual sensing elements.
In this work, TAP aptamers were obtained using the capture-SELEX approach, and the best TAP-1 aptamer has a Kd value of 6.4 μM from ITC and it can bind to CAP with a similar affinity. A sensor array composed of TAP-1 and CAP-1 was tested using ThT fluorescence, which successfully distinguished TAP and CAP from each other and from other antibiotics. Finally, the developed label-free aptasensor was successfully applied to the detection of TAP in lake water samples, achieving a LOD of 8.5 μM. When combined with various signal transduction methods,17,39–42 the aptamers for TAP and CAP can be used to construct sensitive and robust biosensors for environmental and food monitoring applications.
Funding for this work was from the Natural Sciences and Engineering Research Council of Canada (NSERC) and Canada Research Chairs program. Prof. Y. Shu received a China Scholarship Council Scholarship to visit the University of Waterloo.
Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the findings of this study are openly available in The Federated Research Data Repository (FRDR) at: https://doi.org/10.20383/103.01469.The data supporting this article, including RT-PCR results, ThT assay, ITC and CD spectra results have been included as part of the supplementary information (SI). See DOI: https://doi.org/10.1039/d5cc04872a.
Notes and references
- J. C. Hanekamp and E. J. Calabrese, Dose-Response, 2006, 5, 91–93 Search PubMed.
- J. V. Samsonova, A. Cannavan and C. T. Elliott, Crit. Rev. Anal. Chem., 2012, 42, 50–78 CrossRef CAS.
- M. Cannon, S. Harford and J. Davies, J. Antimicrob. Chemother., 1990, 26, 307–317 CrossRef CAS PubMed.
- H. N. Jung, D. H. Park, Y. J. Choi, S. H. Kang, H. J. Cho, J. M. Choi, J. H. Shim, A. A. Zaky, A. M. Abd El-Aty and H. C. Shin, Front. Nutr., 2021, 8, 812803 CrossRef PubMed.
- E. Guo, L. Zhao, K. Wu, W. Huang, K. Zhao, J. Li and A. Deng, Food Agric. Immunol., 2021, 32, 283–297 CrossRef CAS.
- K. Xie, L. Jia, Y. Yao, D. Xu, S. Chen, X. Xie, Y. Pei, W. Bao, G. Dai and J. Wang, J. Chromatogr. B:Biomed. Sci. Appl., 2011, 879, 2351–2354 CrossRef CAS PubMed.
- L. Guo, S. Song, L. Liu, J. Peng, H. Kuang and C. Xu, Biomed. Chromatogr., 2015, 29, 1432–1439 CrossRef CAS PubMed.
- J. Wang, Q. Wang, Y. Zheng, T. Peng, K. Yao, S. Xie, X. Zhang, X. Xia, J. Li and H. Jiang, Food Agric. Immunol., 2018, 29, 56–66 CrossRef CAS.
- L. Wu, Y. Wang, X. Xu, Y. Liu, B. Lin, M. Zhang, J. Zhang, S. Wan, C. Yang and W. Tan, Chem. Rev., 2021, 121, 12035–12105 CrossRef CAS PubMed.
- J. Liu, Z. Cao and Y. Lu, Chem. Rev., 2009, 109, 1948–1998 CrossRef CAS PubMed.
- E. M. McConnell, J. Nguyen and Y. Li, Front. Chem., 2020, 8, 434 CrossRef CAS PubMed.
- P.-J. J. Huang and J. Liu, Angew. Chem., Int. Ed., 2023, 62, e202212879 CrossRef CAS PubMed.
- A. Ruscito and M. C. DeRosa, Front. Chem., 2016, 4, 14 Search PubMed.
- Y. Wu, I. Belmonte, K. S. Sykes, Y. Xiao and R. J. White, Anal. Chem., 2019, 91, 15335–15344 CrossRef CAS PubMed.
- Y. Zhao, K. Yavari and J. Liu, TrAC-Trend Anal. Chem., 2022, 146, 116480 CrossRef CAS.
- N. Nakatsuka, K. A. Yang, J. M. Abendroth, K. M. Cheung, X. Xu, H. Yang, C. Zhao, B. Zhu, Y. S. Rim, Y. Yang, P. S. Weiss, M. N. Stojanović and A. M. Andrews, Science, 2018, 362, 319–324 CrossRef CAS PubMed.
- H. Yu, O. Alkhamis, J. Canoura, Y. Liu and Y. Xiao, Angew. Chem., Int. Ed., 2021, 60, 16800–16823 CrossRef CAS PubMed.
- S. Stangherlin, N. Lui, J. H. Lee and J. Liu, TrAC-Trend Anal. Chem., 2025, 191, 118349 CrossRef CAS.
- K. Yang, N. M. Mitchell, S. Banerjee, Z. Cheng, S. Taylor, A. M. Kostic, I. Wong, S. Sajjath, Y. Zhang, J. Stevens, S. Mohan, D. W. Landry, T. S. Worgall, A. M. Andrews and M. N. Stojanovic, Science, 2023, 380, 942–948 CrossRef CAS PubMed.
- C. Lyu, I. M. Khan and Z. Wang, Talanta, 2021, 229, 122274 CrossRef PubMed.
- H. Yu, Y. Luo, O. Alkhamis, J. Canoura, B. Yu and Y. Xiao, Anal. Chem., 2021, 93, 3172–3180 CrossRef CAS PubMed.
- Y. Duan, Z. Gao, L. Wang, H. Wang, H. Zhang and H. Li, Appl. Biochem. Biotechnol., 2016, 180, 1644–1656 CrossRef CAS PubMed.
- Y. Zhao, A. Z. Li and J. Liu, Environ. Health, 2023, 1, 102–109 CAS.
- D. H. Burke, D. C. Hoffman, A. Brown, M. Hansen, A. Pardi and L. Gold, Chem. Biol., 1997, 4, 833–843 CrossRef CAS PubMed.
- J. Wang, Y. A. Kaiyum, X. Li, H. Lei, P. E. Johnson and J. Liu, J. Am. Chem. Soc., 2025, 147, 1831–1839 CrossRef CAS PubMed.
- X. Li, X. Zhang, J. Wang, X. Wu, Z. Zhang and J. Liu, Anal. Chem., 2025, 97, 17242–17250 CrossRef CAS PubMed.
- K.-A. Yang, R. Pei and M. N. Stojanovic, Methods, 2016, 106, 58–65 CrossRef CAS PubMed.
- Y. Ding, Y. Xie, A. Z. Li, P.-J. J. Huang and J. Liu, Biochemistry, 2023, 62, 2280–2288 CrossRef CAS PubMed.
- A. Renaud de la Faverie, A. Guédin, A. Bedrat, L. A. Yatsunyk and J. L. Mergny, Nucleic Acids Res., 2014, 42, e65 CrossRef CAS PubMed.
- Y. Shu, S. Liu and J. Liu, Anal. Chem., 2025, 97, 19767–19774 CrossRef CAS PubMed.
- H.-R. Zhang, X.-H. Ren, D.-W. Wang, X.-W. He, W.-Y. Li and Y.-K. Zhang, Talanta, 2023, 259, 124506 CrossRef CAS PubMed.
- J. Zhao, X. Li, D. Zhang and S. Wang, Biomed. Technol., 2024, 5, 14–25 CrossRef CAS.
- S. Shen, Z. Deng, X. Lin, N. Duan, Z. Wang and S. Wu, Crit. Rev. Food Sci., 2025, 1–25, DOI:10.1080/10408398.2025.2525502.
- R. Nutiu and Y. Li, J. Am. Chem. Soc., 2003, 125, 4771–4778 CrossRef CAS PubMed.
- X. Liu, F. He, F. Zhang, Z. Zhang, Z. Huang and J. Liu, Anal. Chem., 2020, 92, 9370–9378 CrossRef CAS PubMed.
- J. Wang and B. Liu, Chem. Commun., 2008, 4759–4761 RSC.
- P. J. Huang and J. Liu, Angew. Chem., Int. Ed., 2023, 62, e202212879 CrossRef CAS PubMed.
- X. Chen, C. Yao and Z. Li, TrAC-Trend Anal. Chem., 2023, 158, 116785 CrossRef CAS.
- H. Pei, J. Li, M. Lv, J. Wang, J. Gao, J. Lu, Y. Li, Q. Huang, J. Hu and C. Fan, J. Am. Chem. Soc., 2012, 134, 13843–13849 CrossRef CAS PubMed.
- R. Nutiu and Y. Li, Angew. Chem., Int. Ed., 2005, 44, 1061–1065 CrossRef CAS PubMed.
- M. Zandieh, J. H. Lee and J. Liu, Chem. Sci., 2025, 16, 14865–14883 RSC.
- H. Song, D. H. Jung, Y. Cho, H. H. Cho, V. G. Panferov, J. Liu, J. H. Heo and J. H. Lee, Coord. Chem. Rev., 2025, 541, 216835 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2025 |