Open Access Article
Open Access ArticleApplication of surface enhanced Raman spectroscopy (SERS) for the rapid detection and enumeration of probiotics in yogurt
Kaitlyn
Allen
a,
Gene
Ahlborn
b,
Julia
Wangsgard
b and
Shintaro
Pang
 *b
*b
aDepartment of Microbiology and Molecular Biology, Brigham Young University, UT, USA
bDepartment of Nutrition, Dietetics & Food Science, Brigham Young University, UT, USA. E-mail: shintaro_pang@byu.edu
First published on 1st October 2025
Abstract
Probiotics are a critical component in fermented dairy products such as yogurt. An efficient method to enumerate probiotics in commercially produced yogurt is important, but traditional methods are time-consuming, sometimes taking up to 72 hours. The objective of this study was to develop a rapid method using surface-enhanced Raman spectroscopy (SERS) to identify and enumerate probiotic bacterial cells in yogurt. The identification and enumeration of three different probiotic bacterial strains, Lactobacillus acidophilus, Lactobacillus bulgaricus, and Streptococcus thermophilus were successfully conducted using a developed SERS protocol. L. bulgaricus and S. thermophilus were also identified from a yogurt matrix using a simple extraction procedure utilizing a Percoll gradient. Statistical spectral analysis including principal component analysis and partial least squares regression analysis allowed for the differentiation, characterization and quantification of probiotic strains. The limits of detection were 105 CFU mL−1 for L. acidophilus, 104 CFU mL−1 for L. bulgaricus, and 106 CFU mL−1 for S. thermophilus. Quantification capability was tested up to 108 cfu mL−1 and showed strong correlation coefficients (0.94, 0.98 and 0.95 respectively) between predicted and actual concentrations. The presence of probiotic mixtures were clearly detectable from a yogurt food matrix starting at 105 CFU mL−1. Sample analysis took less than three hours per sample set. Our results demonstrate the capability of SERS to detect and enumerate probiotics from a yogurt matrix rapidly.
1 Introduction
Yogurt is a fermented dairy product characterized by its distinct flavor and texture, primarily due to lactic acid fermentation conducted by probiotic lactic acid bacteria (LAB).1 These LAB metabolize lactose to lactic acid, leading to milk coagulation and the formation of yogurt's unique sensory and textural qualities.2,3 Beyond sensory attributes, these probiotics can confer numerous health benefits such as improved gut microbiota, enhanced lactose tolerance, cholesterol reduction and overall immune system support.4–7 To achieve these health benefits, probiotics must remain viable and be present at adequate concentrations throughout processing and storage.8 Consequently, regulatory standards mandate that yogurt must contain specific minimum levels of viable probiotics, typically around 107 CFU g−1, especially when health-related claims are made.9Ensuring compliance with these standards necessitates reliable, rapid methods for identifying and enumerating probiotic strains. Traditional culture-based enumeration techniques, although accurate, are labor-intensive and time-consuming, often taking up to 72 hours to yield results.10 Culture-based methods also vary depending on the strain of bacteria, further complicating and lengthening the process.11 Alternative methods have been developed to identify and enumerate probiotics, such as immunology-based methods, like antigen/antibody interactions, or nucleic acid analyses using polymerase chain reaction (PCR),10 qualitative polymerase chain reaction (qPCR), and random amplified polymorphic DNA (RAPD) analysis.12,13 These methods require molecule specific probes, nucleic acid detection, and different enzymes and substrates to obtain results.14 While these methods are capable of producing test results at a more rapid turnaround than colony counting, they can suffer other limitations, such as cross-reactivity of biocapture receptors resulting in false positive results, and contamination of nucleic acid extraction resulting in false negative readings. Another limitation is the inability to discern between live and dead bacteria, which differentiation is critical for the quality control of yogurt and other probiotic products.10,12,14
Surface-enhanced Raman spectroscopy (SERS) has gained popularity in recent years as a sensitive, accurate, and rapid analytical technique for the analysis of biochemical and microbiological targets.15–18 Raman scattering is the inelastic scattering of light that occurs when a laser shines on a target analyte.19,20 The energy from the scattering laser is detected by the Raman microscope and correlates with spectral peaks.21 Every molecule has its own distinct Raman spectra, or molecular ‘fingerprint’ which allows for determination of a target analyte.22–24 While Raman scattering signals are often very weak and can be hard to detect, SERS enhances the Raman scattering of target analytes that are in close proximity to SERS-active metal nanoparticles to induce a resonance that enhances the field strength of an oscillatory electro-magnetic field on the surface of the target analyte. This increases the intensity of the Raman scattered light which allows for increased sensitivity of Raman spectroscopy, typically as much as 106 to 107 times.25 By leveraging this phenomenon, researchers have been able to detect low concentrations of various target analytes, including cells.26–28
An added advantage of SERS is that it is capable of identifying targets based on their intrinsic biomolecular make-up, thus producing biochemical fingerprints that are unique to its identity.23,29 This enables label-free detection without the need for a reporter molecule, reducing the probability for false positives due to potential non-specific binding with the receptor. SERS is advantageous by quickly and precisely detecting a wide range of bacteria. It has been shown to distinguish between closely related bacteria, even within a genus, and is a rapid detection method which can reduce detection times by more than half, compared to other methods.23,25 SERS equipment can also be made smaller and more portable than instruments required for many other detection techniques. As a result, it has the potential to act as an accessible, accurate, and rapid detection technology in the food industry.30,31
SERS has been used to detect and identify food-borne pathogens in different food matrices including salmon, eggs, powdered infant formula milk, mixed herbs, fruits, and vegetables,25 but limited studies have been performed to explore probiotics detection using SERS.32,33 In this study, a rapid method was developed to identify and enumerate probiotics using SERS from a yogurt matrix, which included the development of a yogurt extraction method. As far as we know, this is the first study that focuses on using SERS to identify and enumerate probiotics in food. In this study, we aim to distinguish three different strains of probiotics using SERS spectral data and to determine the method's quantification capabilities. We then aim to develop a yogurt extraction method that would separate probiotics from yogurt samples. Lastly, we aim to detect the extracted probiotics using the developed SERS method and quantify their concentration in yogurt.
2 Materials and methods
2.1 Strains and growth conditions
Three probiotic bacterial strains, L. acidophilus (LA-5 CHR Hansen), L. bulgaricus (LB-12 CHR Hansen), and S. thermophilus (i420 CHR Hansen), were grown from frozen pellets in broth. The liquid culture was plated to isolate a single colony. Once grown, a colony was selected and grown overnight in liquid broth to create a pure culture. L. acidophilus and L. bulgaricus were grown in deMan, Rogosa, and Sharpe (MRS) broth and agar plates. S. thermophilus was grown in Brain Heart Infusion (BHI) broth and agar plates. Broths and agars were made according to manufacturers' instructions. All bacterial cultures were grown, shaking overnight until turbid, and plated bacteria were grown for 1–2 days or until colonies were visible. Unless otherwise stated. L. acidophilus and L. bulgaricus were grown anaerobically at 37 °C, and S. thermophilus was grown aerobically at 42 °C.Concentrations of bacteria were determined by serially diluting cultures in their growth media followed by plating. Once growth was visible (24–36 hours), colonies were enumerated by counting plates that had between 30–300 colonies and adjusting for the appropriate dilution factors. This process was also used to evaluate the enumeration capability of this SERS method by confirming bacteria concentrations.
2.2 Fabrication of silver (Ag) dendrites to be used as SERS substrates
Silver (Ag) dendrites were made to enhance the Raman signal of the bacteria and act as the SERS substrate. They were made in accordance with previously published methods for the purpose of SERS enhancement.34 In summary, a zinc metal plate was rinsed with 1 M HCl then double distilled water removing any metal oxides that formed on the outer layer of the plate. The plate was dried with cold air and immersed in 200 mM AgNO3 (aq) for exactly 60 seconds to form approximately 50 nm diameter size silver dendritic nanoparticles. Once formed the dendrites were peeled off the zinc plate using a glass rod and washed with double distilled water three times. The dendrites were left in water overnight, then rinsed one more time before being stored in double distilled water in the dark for up to six months.2.3 Yogurt extraction
To detect probiotics in yogurt, an extraction method was developed to isolate the bacterial cells from the fermented dairy product. A previously developed Percoll based extraction method was modified to function using commercial yogurt (Oikos Plain Nonfat Greek Yogurt) as the extraction matrix.35 The extraction was done by first diluting the yogurt, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 in 1× PBS (Fisher bioreagents). 400 μL of the dilution was placed on top of a Percoll working solution (300 μL Percoll, 700 μL PBS). This suspension was then centrifuged at 649×g for 30 minutes. After centrifugation, the supernatant was discarded and the pellet was resuspended in 50 mM ammonium bicarbonate (Fig. 1a). The bacteria sample preparation protocol was executed as illustrated in Fig. 1b. After the final washing, the pellet was resuspended in 400 μL 50 mM ammonium bicarbonate. Then 4 μL Ag dendrites were added and incubated for 45 minutes with shaking before being plated on a glass slide and viewed under the DXR3 Raman microscope (Thermo Scientific). Standard plate count was used as the traditional method to determine extraction efficiency.
10 in 1× PBS (Fisher bioreagents). 400 μL of the dilution was placed on top of a Percoll working solution (300 μL Percoll, 700 μL PBS). This suspension was then centrifuged at 649×g for 30 minutes. After centrifugation, the supernatant was discarded and the pellet was resuspended in 50 mM ammonium bicarbonate (Fig. 1a). The bacteria sample preparation protocol was executed as illustrated in Fig. 1b. After the final washing, the pellet was resuspended in 400 μL 50 mM ammonium bicarbonate. Then 4 μL Ag dendrites were added and incubated for 45 minutes with shaking before being plated on a glass slide and viewed under the DXR3 Raman microscope (Thermo Scientific). Standard plate count was used as the traditional method to determine extraction efficiency.
 | ||
| Fig. 1 Schematic for (a) extraction of probiotics from yogurt matrix, and (b) sample preparation with SERS active silver dendrites. | ||
2.4 Probiotic sample preparation
Probiotic samples were made up of the extracted probiotics from yogurt or made from pure culture of the three bacteria strains tested. Pure culture samples were grown to 108 CFU mL−1 for L. acidophilus and S. thermophilus and 107 CFU mL−1 for L. bulgaricus. 1 mL of the probiotic sample was aliquoted and centrifuged at 6070×g for 3 minutes to form a pellet and the supernatant was discarded. The bacteria pellet was resuspended in 1 mL 50 mM ammonium bicarbonate (Sigma-Aldrich) then centrifuged at 6070×g for 3 minutes to form a pellet again. This was repeated three times. Once washed, the bacteria were resuspended in 1 mL of 50 mM ammonium bicarbonate (Fig. 1b). If needed, the washed bacteria were serially diluted in 50 mM ammonium bicarbonate. For SERS measurements, 10 μL Ag dendrites were dropped into each 1 mL bacterial sample. The sample was shaken briefly to incorporate the dendrites then placed on a fixed speed (20 rpm) nutating mixer and incubated for 45 minutes. After incubation, 6 μL of the dendrite bacteria suspension was pipetted onto a glass microscope slide, ensuring that dendrites were picked up. The drops were allowed to air dry completely before being observed under a DXR3 Raman microscope (Thermo Scientific). Each trial was performed in triplicate.2.5 SERS measurements and statistical data analysis
All samples were analyzed using a DXR3 Raman microscope (Thermo Scientific) using the following parameters: 10× microscope objective lens, 785 nm laser excitation wavelength, with 5 mW laser power, 1 s exposure time, and 50 μm slit aperture. 10–20 data points were selected for each sample. Points were selected where the Ag dendrites were clearly visible for SERS enhancement.OMNIC™ software (Thermo Scientific) was used to control the Raman instrument and to obtain Raman spectra. The SERS spectra was then analyzed using TQ Analyst (version 9.0) software (Thermo Scientific) using chemometric tools. Second derivative transformation and smoothing were applied at times to reduce spectral noise and to separate overlapping bands. To assess the variability both within and between sample variables, principal component analysis (PCA) was used. This method focuses a multidimensional data set to the most dominant features while removing random variation so that principal components can be used to capture the variation between spectra. In general, non-over-lapping clusters (or classes) indicate statistical significant differences at a confidence level of p = 0.05.36 By using PCA, the limit of detection was determined as the lowest concentration at which the variable data cluster could be distinguished from the negative control data cluster. Partial least squares (PLS), a multivariate analysis model, was also employed to evaluate the linear relationship between calculated (predicted) and actual concentrations. The PLS model was calibrated and validated using a portion of the dataset, and the predictive performance was evaluated using correlation coefficient values.
3 Results and discussion
3.1 Detection and identification of probiotics
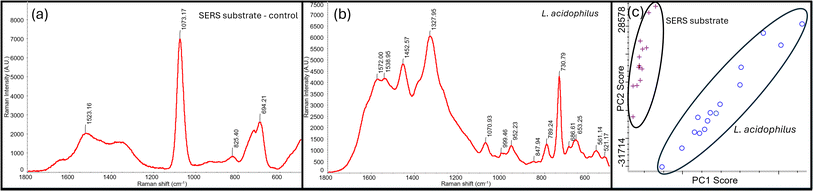
To ensure the developed method detected probiotics, it was first tested with L. acidophilus in pure culture. The surface-enhanced Raman spectra of target samples provided distinct Raman peaks that were specific to L. acidophilus and clearly distinguishable from the SERS substrate (Ag dendrites). These results are shown through spectral data and PCA plot in Fig. 2. L. bulgaricus and S. thermophilus were also analyzed using the same method and found to have distinct peaks from the SERS substrate control (Fig. S1).The three probiotic strains were compared to each other using PCA to determine the differentiation capability of SERS (Fig. 3). The SERS spectra were found to be specific to each probiotic, including differentiation between the two Lactobacillus strains that are of the same genus. This result was similar to a previous study demonstrated through similar enhancement methods.37 The PCA shows the difference between the three strains and the SERS spectra shows distinct peaks that correlate to each bacterial strain.31,38,39 Although there were Raman peaks that all three bacterial strains had in common, there were enough distinct peaks to allow for differentiation of the bacteria from each other from the bacterial cocktail (Fig. S2). This insight makes SERS a valuable resource for probiotic detection in yogurt and other food products as it allows manufacturers and others to quickly identify the strains of bacteria present in their product. Additional spectral data can be found in the SI figures.
 | ||
| Fig. 3 SERS capability to distinguish between L. acidophilus, L. bulgaricus, and S. thermophilus strains tested, and analyzed through principal component analysis (PCA). | ||
Table 1 lists the most prevalent peaks of each probiotic bacterial strain, with the assignment representation of each peak. Of notable mention is the peak on the control spectra around 1070 cm −1, which is often observed on the bacterial spectra. This is due to the NO3− stretching residues in the Ag dendrites.44,47 Changes in the intensity of this peak can be used as an indicator of the sample interacting with the SERS substrate. As the concentration of sample bacteria increases, the intensity of the peak around 1070 cm−1 decreases.47 One of the most prominent SERS peaks found in bacteria is found between 720–735 cm−1.43,48 This region has had many different assignments over the years, but the leading idea is that it is due to adenine or adenine containing molecules.43 The peak around 650 cm −1 can be attributed to C–S stretching, C–C twisting of proteins, or COO− deformation in amino acids.41 There are two peaks that are distinct to L. bulgaricus. These are potential landmarks to distinguish L. bulgaricus from the other probiotics tested. These peaks are 1091 cm−1 and 1461 cm−1 which correlate with chains of 1,3-β-D-glucan and lipids respectively.41L. acidophilus and S. thermophilus also had distinct peaks at 1539 cm−1 and 1560 cm−1 respectively, which correlate with Amide II of proteins.46 These results demonstrate the method's capability to identify different bacteria based on the SERS spectra.
| L. acidophilus peaks (cm−1) | L. bulgaricus peaks (cm−1) | S. thermophilus peaks (cm−1) | Assignment | Ref. |
|---|---|---|---|---|
| 521 | — | 523 | Flavin | 40 |
| 561 | 562 | 565 | Carbohydrates in cell wall | 41 and 42 |
| 653 | 650 | 656 | C–S stretch model of cysteine | 41 |
| 731 | 731 | 731 | Adenine or adenine related compounds | 42 and 43 |
| 952 | 955 | 957 | D2O or CN | 42 and 43 |
| 1071 | — | 1074 | NO3− | 44 |
| — | 1091 | — | Branched chains of 1,3-β-D-glucan | 41 |
| 1328 | 1329 | 1329 | Proteins or aromatic ring structure of cytosine and uracil nucleic acids | 41 and 45 |
| 1453 | — | 1451 | C–H deformation bond of cell wall carbohydrate | 45 |
| — | 1461 | — | Lipids | 41 |
| 1539 | — | — | Amide II, CN, NH | 42 |
| — | — | 1560 | Amide II of proteins | 46 |
| 1572 | 1579 | — | Guanine, adenine, tryptophane (proteins) | 46 |
Some studies have demonstrated SERS capability to differentiate between different strains of Gram-negative bacteria,49,50 while other studies have shown SERS capability of differentiating between Gram-positive and Gram-negative bacterial strains when compared with each other.51,52 There have been limited studies comparing Gram-positive bacteria to each other.53 All three of the probiotic strains tested were Gram-positive bacteria which showed that SERS can differentiate between distinct Gram-positive bacteria. These results taken together demonstrate that SERS can differentiate between closely related Gram-positive probiotic strains. This knowledge expands the reach of Raman spectroscopy as it can rapidly distinguish between different strains of both Gram-positive and Gram-negative bacteria. Furthermore, establishing the specificity of SERS for differentiating among Gram-positive strains significantly broadens its application potential in quality assurance and microbial analysis in food. Future studies can expand upon this research by exploring more different strains, enhancing the versatility of SERS as a rapid diagnostic tool.
3.2 Limit of detection of probiotics
The detection capabilities were further analyzed by determining the lowest detectable concentration of bacteria using this method. This was done by serial diluting the washed bacteria before mixing them with the SERS substrate and observing them under the Raman microscope. The limit of detection was found to be dependent on the probiotics tested, but all were 106 CFU mL−1 or lower. This demonstrates the usability of this method for detection of probiotics in yogurt as the necessary initial concentration of probiotics in yogurt is 106 to 107 CFU per gram or milliliter, or higher in some countries.9,54 While 106 CFU mL−1 may not be considered a low concentration to detect in traditional microbial methods, it may be sufficient as a rapid method for yogurt as the mandated minimum storage threshold is 106 CFU mL−1.1,55 As illustrated in Fig. 4, the limit of detection for L. acidophilus was 105 CFU mL−1. While a few spectral data points for 105 CFU mL−1 overlapped with 106 CFU mL−1, they were both separate from the negative control (i.e. SERS substrate). The limit of detection for L. bulgaricus was 104 CFU mL−1 (Fig. S3). Lastly, the limit of detection of S. thermophilus was 106 CFU mL−1 (Fig. S3). The observed differences in the limits of detection may be attributed to variations in bacterial surface composition, such as differences in cell-wall structure, extracellular polymeric substances, or expression of Raman-active biomolecules.40,56 It should also be noted that there is inherent variability when using SERS depending on the SERS substrate and how the bacteria interacts with it during a given trial which can impact the uniformity of detection.57 Thus, further optimization with SERS substrate types has the potential to achieve lower limits of detection. Nevertheless, these results demonstrate the capability of SERS to detect probiotic bacteria at various targeted concentrations. More importantly, the results show the potential of SERS as an effective and efficient tool for enumerating probiotics at concentrations below the regulatory minimum of some countries for yogurt.3.3 Quantification capability of probiotic strains
The SERS spectral data of L. acidophilus, L. bulgaricus, and S. thermophilus at various concentrations were evaluated using partial least squares (PLS) regression analysis to predict the concentration of these probiotic strains (Fig. 5). Results showed a strong linear correlation between the actual and predicted concentrations, with a correlation coefficient of 0.94 for L. acidophilus (Fig. 5a), 0.98 for L. bulgaricus (Fig. 5b) and 0.95 for S. thermophilus (Fig. 5c). Five distinct bacterial concentrations were used for both L. acidophilus (from 104 to 108 CFU mL−1) and L. bulgaricus (from 103 to 107 CFU mL−1), whereas four concentrations were used for S. thermophilus (from 105 to 108 CFU mL−1) due to its comparatively higher detection limit which prevented the inclusion of a lower concentration. It is worth noting that the limit of quantification was the same as the limit of detection for this method as lower concentrations were less distinct from the control. Nevertheless, these results suggest the potential to quantify the concentrations of distinct bacteria in a solution. This is important for the quality assurance testing of yogurt because specific strains of Gram-positive bacteria, namely L. bulgaricus and S. thermophilus, must reach at least 107 CFU g−1 for some product claims.9One of the challenges in the yogurt and probiotic markets worldwide is to ensure that the probiotic count is maintained at a high enough concentration during its shelf life. This is essential for the claims of health impacts from probiotics in the food to be viable. Although most countries seem to agree on 106 CFU mL−1 as a minimum concentration that still provides the claimed health benefits to the consumer, some impose even higher restrictions because it is difficult to ensure the viability of the claims without rapid accurate analysis of bacterial concentrations.54,55,58 Consequently, inconsistent international standards complicate global trade of probiotic products. The demonstrated quantification capability of SERS in this study highlights its potential as an efficient and accurate tool for verifying probiotic concentrations, thereby aiding regulatory compliance, ensuring product consistency, and enhancing consumer confidence in probiotic products.
3.4 Analysis of extraction method
To determine the practical application of this method on probiotics from a yogurt product, a commercially made yogurt containing S. thermophilus and L. bulgaricus was used. This yogurt was specifically selected as it only contained strains we used to develop this method, though it is missing L. acidophilus. This was because a yogurt could not be found that included all three probiotic strains tested in this study without the addition of other probiotics. A Percoll based extraction method was developed to extract probiotics from the milk proteins and casein micelles found in yogurt. The extracted bacteria were analyzed using the determined method outlined previously. The extracted bacteria were compared against a bacterial cocktail containing 107 CFU mL−1 of S. thermophilus and L. bulgaricus grown in pure culture. The spectral data obtained shows similarities in the extracted bacteria spectra with the bacterial cocktail spectra indicating that we successfully isolated bacteria from the milk proteins found in yogurt. The spectral data was analyzed using principal component analysis (PCA) which confirmed what was observed in the Raman spectra. The PCA showed an overlap between the bacterial cocktail and extracted probiotics indicating similar spectra and therefore isolation of the bacteria from the yogurt (Fig. 6). In total, the extraction and SERS detection took less than 3 hours, which is significantly shorter than the current culture-based method.12 Being able to accurately detect probiotics from a yogurt matrix in under 3 hours shows the usability of this method in the yogurt industry. This signifies that the extraction method was effective in extracting and isolating bacteria from a yogurt matrix.It is also worth noting that approximately a one-log reduction was consistently observed between the probiotic concentrations in the original yogurt and the extraction sample. Because this loss was consistent, it was readily accounted for in our calculations and results. Optimization could be performed to further limit the bacterial loss seen in the extraction method. Testing could also be done on different yogurt types containing different bacterial strains as well as yogurts that contain whole food particles. This would require a wide range of probiotics being analyzed and mixed in different ratios to mimic commercial yogurt types. Further analysis of more probiotics could lead to the creation of a probiotic SERS library. As such, the determination of a bacteria given the spectra would be easier and more uniform. Taken together, our study shows that SERS is a potential method to rapidly detect and enumerate probiotics in yogurt, providing a faster yet accurate method for quality control of yogurt.
3.5 Quantification of probiotics from a yogurt matrix
Once the extraction method was successful, a limit of detection was performed on the extracted probiotics. The yogurt was serially diluted, then the probiotics were extracted from each dilution before being mixed with the SERS substrate, resulting in a detectable difference between various concentrations of extracted bacteria. It was found that all concentrations of the extraction were different than the control, and they follow the expected trend with 105 CFU mL−1 being closest to the control and 107 CFU mL−1 correlating with the bacteria cocktail control and being furthest from the SERS substrate control (Fig. 7). To enumerate the bacteria extracted from yogurt, the dilutions were analyzed using a partial least square (PLS) regression curve which had a correlation coefficient of 0.9996. This result demonstrates the high level of accuracy and precision in the quantification of probiotics in yogurt using a Percoll extraction method and SERS. The high correlation between the actual and calculated concentrations of probiotics in the yogurt samples highlight SERS as a potential method of analyzing probiotics in food.One limitation of this study, however, is that live and dead probiotic bacteria cells were not explored in detail. When detecting probiotics in yogurt it is important that only the live bacteria are counted in the concentration because only the live probiotics provide health benefits. Some previous SERS studies have shown the capacity to differentiate between live and dead bacteria.59–61 Similarly, we performed preliminary testing and found that our SERS method can differentiate between live and dead bacteria (Fig. S4), but further optimization of the method needs to be done to determine if SERS can accurately distinguish and enumerate live versus dead bacteria from a yogurt matrix. Hence, future studies should focus on optimizing and testing this method to determine if SERS is able to differentiate between live and dead bacteria from a yogurt matrix.
4 Conclusion
In summary, this study successfully developed a rapid method to detect and enumerate probiotics in yogurt using surface enhanced Raman spectroscopy (SERS). A Percoll density gradient extraction method was used to isolate probiotics, thereby minimizing interference from complex food matrices found in yogurt. SERS demonstrated significant potential to allow for the specific detection of multiple probiotics, clearly differentiating three common Gram-positive probiotic strains, including closely related strains within the same genus (Lactobacillus). The limit of detection varied depending on the bacterial strain but was determined to be 105 CFU mL−1 for L. acidophilus and for the yogurt extraction combination of L. bulgaricus and S. thermophilus. The method also achieved a limit of detection of 104 CFU mL−1 for L. bulgaricus, and 106 CFU mL−1 for S. thermophilus. Overall, the integration of the extraction protocol with SERS detection represents a promising and practical advancement for the rapid, accurate quantification of probiotics in yogurt.Author contributions
Conceptualization: K. Allen, G. Ahlborn, S. Pang. Methodology: K. Allen, S. Pang. Formal analysis: K. Allen, S. Pang. Investigation: K. Allen, S. Pang. Resources: G. Ahlborn, S. Pang. Data curation: K. Allen, S. Pang. Writing: K. Allen, G. Ahlborn, J. Wangsgard, S. Pang. All authors have read and agreed to the submitted/published version of the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article has been included as part of the SI. See DOI: https://doi.org/10.1039/d5ay00931f.Acknowledgements
This study was financially supported by the BYU College of Life Sciences faculty research startup fund.References
- R. Nyanzi, P. J. Jooste and E. M. Buys, J. Dairy Sci., 2021, 104, 1–19 CrossRef CAS PubMed.
- K. Bostan, A. Unver Alcay, S. Yalcin, U. Eren Vapur and M. Nizamlioglu, Food Sci. Biotechnol., 2017, 26, 1625–1632 CrossRef CAS PubMed.
- C. Chen, S. Zhao, G. Hao, H. Yu, H. Tian and G. Zhao, Int. J. Food Prop., 2017, 20, S316–S330 CrossRef CAS.
- H. Nakao and Y. Magariyama, Anal. Sci., 2022, 38, 191–197 CrossRef CAS PubMed.
- H. Nakao and J. D. Kim, Anal. Sci., 2019, 35, 1065–1067 CrossRef CAS PubMed.
- C. Mazziotta, M. Tognon, F. Martini, E. Torreggiani and J. C. Rotondo, Cells, 2023, 12, 184 CrossRef CAS PubMed.
- S. Paul, T. J. Hossain, F. Ali and J. Ferdouse, Ann. Agri. Bio. Res., 2024, 30, 17–24 Search PubMed.
- J.-W. Chon, K.-H. Seo, T.-J. Kim, H.-Y. Youn, S.-H. Kang, W.-U. Hwang, H. Jeong, D. Jeong and K.-Y. Song, J. Dairy Sci. Biotechnol., 2023, 41, 9–25 CrossRef.
- CFR - Code of Federal Regulations Title 21; 131.200, 2025, online, available: https://www.ecfr.gov/current/title-21/chapter-I/subchapter-B/part-131/subpart-B/section-131.200, accessed 3 June, 2025.
- X. Meng, R. Pang, C. Wang and L. Wang, J. Dairy Res., 2010, 77, 498–504 CrossRef CAS PubMed.
- T. Sibanda, T. A. Marole, U. L. Thomashoff, M. S. Thantsha and E. M. Buys, Front. Microbiol., 2024, 15, 1327010 CrossRef PubMed.
- R. Najafi, S. Mukherjee, J. Hudson, Jr., A. Sharma and P. Banerjee, Int. J. Food Microbiol., 2014, 189, 89–97 CrossRef CAS PubMed.
- G. Saxami, O. S. Papadopoulou, N. Chorianopoulos, Y. Kourkoutas, C. C. Tassou and A. Galanis, Int. J. Mol. Sci., 2016, 17, 668 CrossRef PubMed.
- R. T. Noble and S. B. Weisberg, J. Water Health, 2005, 3, 381–392 CrossRef PubMed.
- L. Jiang, M. M. Hassan, S. Ali, H. Li, R. Sheng and Q. Chen, Trends Food Sci. Technol., 2021, 112, 225–240 CrossRef CAS.
- H. Ma, S.-Q. Pan, W.-L. Wang, X. Yue, X.-H. Xi, S. Yan, D.-Y. Wu, X. Wang, G. Liu and B. Ren, ACS Nano, 2024, 18, 14000–14019 CrossRef CAS PubMed.
- L. L. Lin, R. Alvarez-Puebla, L. M. Liz-Marzán, M. Trau, J. Wang, L. Fabris, X. Wang, G. Liu, S. Xu and X. X. Han, ACS Appl. Mater. Interfaces, 2025, 17, 16287–16379 CrossRef CAS PubMed.
- Y. Xu, Q. Dong, S. Cong and Z. Zhao, Analysis Sensing, 2024, 4, e202300067 CrossRef CAS.
- G. S. Bumbrah and R. M. Sharma, Egypt. J. Forensic Sci., 2016, 6, 209–215 CrossRef.
- R. R. Jones, D. C. Hooper, L. Zhang, D. Wolverson and V. K. Valev, Nanoscale Res. Lett., 2019, 14, 231 CrossRef PubMed.
- P. A. Mosier-Boss, Biosensors, 2017, 7, 51 CrossRef PubMed.
- Z. Lin and L. He, Curr. Opin. Food Sci., 2019, 28, 82–87 CrossRef.
- Z. Guo, X. Wu, H. Jayan, L. Yin, S. Xue, H. R. El-Seedi and X. Zou, Food Chem., 2024, 434, 137469 CrossRef CAS PubMed.
- Z. Liu, R. Yang, H. Chen and X. Zhang, Foods, 2025, 14, 1115 CrossRef CAS PubMed.
- E. Witkowska, D. Korsak, A. Kowalska, M. Ksiezopolska-Gocalska, J. Niedziolka-Jonsson, E. Rozniecka, W. Michalowicz, P. Albrycht, M. Podrazka, R. Holyst, J. Waluk and A. Kaminska, Anal. Bioanal. Chem., 2017, 409, 1555–1567 CrossRef CAS PubMed.
- B. Z. Xiao Xia Han and Y. Ozaki, ScienceDirect, 2012, 38, 67–78 Search PubMed.
- P. Wang, S. Pang, J. Chen, L. McLandsborough, S. R. Nugen, M. Fan and L. He, Analyst, 2016, 141, 1356–1362 RSC.
- Y. Yan, Y. Nie, L. An, Y.-Q. Tang, Z. Xu and X.-L. Wu, Front. Bioeng. Biotechnol., 2020, 8, 573777 CrossRef PubMed.
- D. Cialla, A. Marz, R. Bohme, F. Theil, K. Weber, M. Schmitt and J. Popp, Anal. Bioanal. Chem., 2012, 403, 27–54 CrossRef CAS PubMed.
- H. Pu, Q. Ouyang and D.-W. Sun, Trends Food Sci. Technol., 2024, 104416 CrossRef CAS.
- M. H.-U. Rahman, R. Sikder, M. Tripathi, M. Zahan, T. Ye, E. Gnimpieba Z, B. K. Jasthi, A. B. Dalton and V. Gadhamshetty, Chemosensors, 2024, 12, 140 CrossRef CAS.
- G. Velázquez, Doctoral dissertation, 2018, pp. 1–45.
- S. Liu, L. Zhao, M. Li, J.-H. Lee, Y. Zhu, Y. Liu, L. Sun, Y. Ma, Q. Tu and G. Zhao, Food Res. Int., 2024, 197, 115249 CrossRef CAS PubMed.
- L. He, M. Lin, H. Li and N. J. Kim, J. Raman Spectrosc., 2010, 41, 739–744 CrossRef CAS.
- T. Nishino, Y. Matsuda and Y. Yamazaki, Heliyon, 2018, 4, e00597 CrossRef PubMed.
- L. He, T. Chen and T. P. Labuza, Food Chem., 2014, 148, 42–46 CrossRef CAS PubMed.
- K. E. Stephen, D. Homrighausen, G. DePalma, C. H. Nakatsu and J. Irudayaraj, Analyst, 2012, 137, 4280–4286 RSC.
- Y. Liu, Y. Gao, R. Niu, Z. Zhang, G.-W. Lu, H. Hu, T. Liu and Z. Cheng, Anal. Chim. Acta, 2024, 1332, 343376 CrossRef CAS PubMed.
- D. Li, Y. Zhu, A. Mehmood, Y. Liu, X. Qin and Q. Dong, Talanta, 2025, 285, 127268 CrossRef CAS PubMed.
- S. Efrima and L. Zeiri, J. Raman Spectrosc., 2009, 40, 277–288 CrossRef CAS.
- S. M. Berus, M. Adamczyk-Popławska, K. Goździk, G. Przedpełska, T. R. Szymborski, Y. Stepanenko and A. Kamińska, Int. J. Mol. Sci., 2022, 23, 12576 CrossRef CAS PubMed.
- H. Zhou, D. Yang, N. P. Ivleva, N. E. Mircescu, R. Niessner and C. Haisch, Anal. Chem., 2014, 86, 1525–1533 CrossRef CAS PubMed.
- P. Kubryk, R. Niessner and N. P. Ivleva, Analyst, 2016, 141, 2874–2878 RSC.
- P. Wang, S. Pang, B. Pearson, Y. Chujo, L. McLandsborough, M. Fan and L. He, Anal. Bioanal. Chem., 2017, 409, 2229–2238 CrossRef CAS PubMed.
- J. Sundaram, B. Park, A. Hinton, K. C. Lawrence and Y. Kwon, J. Food Meas. Charact., 2013, 7, 1–12 CrossRef.
- N. Paccotti, F. Boschetto, S. Horiguchi, E. Marin, A. Chiadò, C. Novara, F. Geobaldo, F. Giorgis and G. Pezzotti, Biosensors, 2018, 8, 131 CrossRef CAS PubMed.
- L. He, T. Rodda, C. L. Haynes, T. Deschaines, T. Strother, F. Diez-Gonzalez and T. P. Labuza, Anal. Chem., 2011, 83, 1510–1513 CrossRef CAS PubMed.
- W. R. Premasiri, Y. Gebregziabher and L. D. Ziegler, Appl. Spectrosc., 2011, 65, 493–499 CrossRef CAS PubMed.
- D. Vaitiekūnaitė and V. Snitka, Microorganisms, 2021, 9, 1969 CrossRef PubMed.
- R. M. Jarvis and R. Goodacre, Anal. Chem., 2004, 76, 40–47 CrossRef CAS PubMed.
- X. Wu, Y.-W. Huang, B. Park, R. A. Tripp and Y. Zhao, Talanta, 2015, 139, 96–103 CrossRef CAS PubMed.
- Y. Dong, Y. Hu, J. Jin, H. Zhou, S. Jin and D. Yang, TrAC, Trends Anal. Chem., 2024, 117974 CrossRef CAS.
- A. Colniţă, N. E. Dina, N. Leopold, D. C. Vodnar, D. Bogdan, S. A. Porav and L. David, Nanomaterials, 2017, 7, 248 CrossRef PubMed.
- V. Garg, D. Velumani, Y.-C. Lin and A. Haye, PharmaNutrition, 2024, 100423 CrossRef CAS.
- D. Liang, F. Wu, D. Zhou, B. Tan and T. Chen, Crit. Rev. Food Sci. Nutr., 2024, 64, 6455–6476 CrossRef PubMed.
- S. Dastgir, S. Rauf, H. Nawaz, M. I. Majeed, M. R. Javed, N. Alwadie, R. Seemab, A. Fatima, A. B. Salfi and M. Usman, Plasmonics, 2025, 1–13 Search PubMed.
- D. B. Grys, R. Chikkaraddy, M. Kamp, O. A. Scherman, J. J. Baumberg and B. de Nijs, J. Raman Spectrosc., 2021, 52, 412–419 CrossRef CAS.
- A. Ranjan, J. Arora, A. Chauhan, R. K. Basniwal, A. Kumari, V. D. Rajput, E. V. Prazdnova, A. Ghosh, N. Mukerjee and S. S. Mandzhieva, Biotechnol. Genet. Eng. Rev., 2024, 40, 3226–3269 CrossRef PubMed.
- J. Zhang, J. Zhang, J. Ding, Q. Lin, G. M. Young and C. Jiang, Vib. Spectrosc., 2022, 118, 103332 CrossRef CAS.
- R. Li, D. Dhankhar, J. Chen, A. Krishnamoorthi, T. C. Cesario and P. M. Rentzepis, IEEE Access, 2019, 7, 23549–23559 Search PubMed.
- H. Zhou, D. Yang, N. P. Ivleva, N. E. Mircescu, S. r. Schubert, R. Niessner, A. Wieser and C. Haisch, Anal. Chem., 2015, 87, 6553–6561 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2025 |