 Open Access Article
Open Access ArticleRed blood cells as probes for determining free QS21 in liposomal adjuvant formulations to support product safety and stability
Essie
Komla
ab,
Cierra
Rochelle
ab,
Adrian
Carreon
ab,
Oscar B.
Torres†
ab,
Gary R.
Matyas
 *b and
Erwin G.
Abucayon†
*ab
*b and
Erwin G.
Abucayon†
*ab
aHenry M. Jackson Foundation for the Advancement of Military Medicine, 6720A Rockledge Drive, Bethesda, Maryland 20817, USA. E-mail: erwin.abucayon@merck.com
bU.S. Military HIV Research Program, Center for Military Infectious Diseases Research, Walter Reed Army Institute of Research, 503 Robert Grant Avenue, Silver Spring, Maryland 20910, USA
First published on 6th November 2025
Abstract
Analytical method development is critical to establishing and supporting drug product quality. QS21 is a potent adjuvant, which is currently used in several clinical vaccine trials, and is utilized as an adjuvant in licensed vaccines against malaria, shingles and respiratory syncytial virus (RSV). Due to its toxic and hemolytic properties as a stand-alone or free adjuvant, QS21 has been incorporated into heterogeneous matrices containing cholesterol that serve as QS21 sinks to suppress its toxic activity. Hence, to address the safety attribute of the QS21-based adjuvant, it is imperative to ensure the absence of free QS21 in the drug formulation. To date, quantitative analytical methods that definitively discriminate free from bound QS21 in liposomal adjuvant formulations are not available in the literature. In view of this, we have developed and qualified a cell-based assay using human red blood cells (RBCs) as probes for detecting and estimating free QS21 in liposomal formulations. This method exhibits a medium throughput, and does not require any expensive and special instrumentation system. Qualification results showed that the method presents good specificity for free QS21 detection, accuracy and precision, with quantitation sensitivity in the micromolar range (lower limit of ∼25 μg mL−1), suitable for our intended purpose. The described method was successfully used as part of the regulatory specification tests for the product release and stability studies of Army Liposome Formulation containing QS21 (ALFQ).
Introduction
QS21 as a potent adjuvant has been established from multiple immunology studies, involving many preclinical and clinical studies.1–5 The chemistry of this compound has undergone extensive structural investigation, which resulted in the development of synthetic variants that provided insights into the contribution and function of each structural moiety in its total immunological profiles.1,3 The potency of QS21 as an adjuvant is evident in its presence in commercial malaria vaccine for children (RTS,S/AS01E, also known as Mosquirix),6 a herpes zoster vaccine for older adults (gE/AS01B, Shingrix),7 and a respiratory syncytial virus vaccine for adults (RSVPreF3-AS01E, AREXVY).8The majority of the commercially available QS21 has been produced from the bark extract of the Quillaja saponaria Molina tree, native to Chile,9–11 and recently from an aeroponic system, and cell cultures.12 Despite the advances of QS21 as an adjuvant, this molecule exhibits several liabilities and limitations. QS21 extracts contain at least four (4) isomers, which have been proven difficult to separate by chromatography or any separation techniques. Immunological profiles collected from preclinical and clinical studies indicated the total responses of the mixture of isomers in adjuvant suspension, which hinders the complete understanding of the QS21 adjuvant mechanism of action. In aqueous solution, QS21 undergoes a pH dependent hydrolytic cleavage of the acyl chain fragment, producing a less active hydrolytic product, QS21 HP.13 Further, standalone or free QS21 exhibits a dose-limiting toxicity.1,3 Like other saponins, QS21 intercalates into cholesterol of the RBCs, and its strong interactions with cholesterol result in the formation of pores in the lipid bilayer of cell membranes, leading to hemolysis.14,15 These complications challenge the safety and stability of adjuvant formulations containing QS21 during long-term storage. The described instability and hemolytic properties of free QS21 were addressed by incorporating QS21 in immune-stimulating complexes,16 liposomal formulations,14,15,17 or surfactant emulsions,11 which slow down its hydrolytic degradation, and eliminate its hemolytic activity.
Our laboratory at Walter Reed Army Institute of Research (WRAIR) has developed QS21-containing liposomes called Army Liposome Formulation with QS21 (ALFQ; Fig. 1A). This liposome formulation that acts as a QS21 sink contains saturated phospholipids, dimyristoyl phosphatidylcholine (DMPC) and dimyristoyl phosphatidylglycerol (DMPG), monophosphoryl lipid A (MPLA), and cholesterol, which eliminates its hemolytic activity.14,18,19 Vialed cGMP ALFQ contains 120 μg of QS21 (200 μg mL−1) in 0.6 mL. This adjuvant formulation has been tested in multiple phase 1 clinical trials for infectious diseases, most of which are ongoing.5,20 In order to establish the safety and stability of ALFQ, it is essential to monitor the presence of free QS21 in liposomal formulations. Currently, there is no available and established analytical method in the literature that can quantitatively discriminate free from bound QS21 in liposomal drug formulations. Hemolysis assay was crudely described in the literature to qualitatively determine the presence of free QS21 in the adjuvant formulation.14,21–23 Other methods were based on liquid chromatography coupled with MS, CAD and UV detection.24,25 Stand-alone MS together with 1H and 13C NMR have also been employed to investigate plausible fragmentation of QS21.26,27 All the described methods suffer from the inability to determine the amount of free QS21 in liposomal drug formulations and related products.
 | ||
| Fig. 1 (A) Proposed molecular assembly of ALFQ (adapted from ref. 24). (B) Drabkin's reaction, showing the derivatization of Hgb to cyanmethemoglobin. (C) Scheme for detecting free QS21 using hemolysis assay. Part of figure (C) was adapted from https://Biorender.com. | ||
In the present work, we have developed a method that utilizes RBCs as probes coupled with Drabkin's reagent (known as the cyanmethemoglobin reagent; CMHR) for detecting and estimating free QS21 in ALFQ suspensions (Fig. 1B and C). The interactions of free QS21 with RBCs resulted in the release of hemoglobin (Hgb), which was then derivatized by CMHR forming a more stable cyanmethemoglobin (CMH), followed by quantitative UV-vis spectroscopy (Fig. 1C).
The designed method was qualified following FDA Guidance for Industry: Q2(R1) Validation of Analytical Procedures. Qualification results demonstrated that the method has good analyte specificity and measurement sensitivity (micromolar range), accuracy, and precision, appropriate for determining free QS21 in the liposomal drug product. The method has been utilized for ALFQ product release and stability studies to ensure the quality of cGMP manufactured ALFQ at WRAIR for use in clinical studies.
Experimental
Chemicals and reagents
Drabkin's reagent also referred to as cyanmethemoglobin reagent (CMHR), Triton X-100 (Tx-100), lyophilized human hemoglobin, and sodium phosphate dibasic were purchased from Sigma-Aldrich (Saint Louis, MO). Sodium phosphate monobasic and sodium chloride were purchased from Fisher Scientific (Rockford, IL). Dulbecco's modified phosphate buffered saline (1× DPBS without Ca2+, Mg2+) was purchased from Quality Biological (Gaithersburg, MD). Human blood samples were purchased from Research Blood Component, LLC (Watertown, MA) and were classified as non-human subject research by the Walter Reed Army Institute of Research Institutional Review Board, Division of Human Subjects.Dimyristoyl phosphatidylcholine (DMPC), dimyristoyl phosphatidylglycerol (DMPG), synthetic MPLA (3D-PHAD), and cholesterol (plant derived) for liposomal preparation were purchased from Avanti Polar Lipids LLC (Alabaster AL). QS21 (cGMP-grade) was purchased from Desert King International (San Diego, CA).
Preparation of reagents
ALFQ was prepared as previously described.28–30 CMHR stock solution was prepared by reconstituting the commercial CMHR vial with 1000 mL of deionized water. This solution was stored at room temperature and protected from light using aluminum foil for 6 months. Sodium phosphate buffered saline (SPBS; pH = 6.2) was prepared by dissolving 53.36 mM sodium phosphate monobasic, 13.34 mM sodium phosphate dibasic, and 87.3 mM sodium chloride in 500 mL of deionized water, and was sterile filtered using a 0.22 μm Corning Filter System (Corning, AZ). 1% Triton X-100 (Tx-100) was prepared by dissolving 1 g of Tx-100 in 100 mL SPBS. This solution was stirred on a hot plate at 30 °C until completely dissolved. Tx-100 was stored at 4 °C for up to two weeks.Preparation of standard solutions
A standard Hgb stock solution of 32 mg mL−1 in SPBS was freshly prepared for each experiment. From this stock solution, Hgb calibration standards were prepared by series of dilutions to generate final concentrations of 16.0, 12.0, 8.0, 6.0, 4.0, 2.0, 1.0, and 0.5 mg mL−1. Hgb solutions were briefly vortexed for 30 s followed by ultrasonication in a water bath for 5–10 min at room temperature. 100 μL of each Hgb calibration standard was added in triplicate to a 96-well UV transparent plate containing 100 μL of CMHR. The plate was sealed using an adhesive film for microplates, and gently mixed on an orbital shaker for 15 min in the dark. These mixtures were then incubated at room temperature for 45 min in the dark, prior to absorbance reading of the resulting cyanmethemoglobin at 540 nm.Sample matrix evaluation
The sample matrix was evaluated using various test samples in SPBS as described. Four (4) test samples were prepared in separate microfuge tubes, to which were added 1 mL of 7 mg mL−1 Hgb (tube 1), 1 mL of 7 mg mL−1 Hgb (tube 2), 1 mL of 7 mg mL−1 Hgb with ALFQ containing 100 μg mL−1 QS21 (tube 3), and 1 mL of 7 mg mL−1 Hgb with 5 μg mL−1 (tube 4). These mixtures were incubated in a thermomixer at 15–25 °C for 10 min at 300 rpm, followed by centrifugation at 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 130g RCF. 100 μL of the supernatant from each tube was added in triplicate to a 96-well UV transparent plate. For the test sample from tube 1, 100 μL of SPBS was added. For test samples from tubes 2–4, 100 μL of CMHR solution was added. All test samples had a total volume of 200 μL, containing equal concentrations of Hgb and CMHR. Negative controls were also included in the experiment and prepared as described. SPBS and CHMR negative control samples were prepared by adding 200 μL of each solution in separate wells in triplicate.
130g RCF. 100 μL of the supernatant from each tube was added in triplicate to a 96-well UV transparent plate. For the test sample from tube 1, 100 μL of SPBS was added. For test samples from tubes 2–4, 100 μL of CMHR solution was added. All test samples had a total volume of 200 μL, containing equal concentrations of Hgb and CMHR. Negative controls were also included in the experiment and prepared as described. SPBS and CHMR negative control samples were prepared by adding 200 μL of each solution in separate wells in triplicate.
The plate was sealed using an adhesive film, and gently mixed on a plate rocker for 15 min in the absence of light. These mixtures were then incubated at room temperature for 45 min in the dark, prior to an absorbance scan at 440–700 nm.
Red blood cell (RBC) preparation
Fresh human blood (25 mL) samples were washed 3–5× with 25 mL 1× DPBS (without Ca2+ and Mg2+) by centrifugation at 469g RCF and 15–25 °C for 10 min to remove dead cells, plasma, and additive solution. The RBCs were then resuspended in 25 mL 1× DPBS and quantified using a hemocytometer (dilution factor = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400). RBC samples were diluted in an appropriate volume of 1× DPBS to generate the desired cell concentrations.
400). RBC samples were diluted in an appropriate volume of 1× DPBS to generate the desired cell concentrations.
To determine the optimal cell concentration for the hemolysis assay, 500 μL of 1% Tx-100 was added to RBC samples with a concentration range of 1 × 104 cells mL−1 to 5 × 108 cells mL−1 in microfuge tubes. These mixtures were incubated in a thermomixer at 15–25 °C at 300 rpm for 10 min, followed by centrifugation at 15–25 °C and 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 130g RCF for 10 min. 100 μL of the supernatant from each tube was added to a UV transparent plate in triplicate with the addition of 100 μL CMHR to each well. The plate was sealed and gently mixed on a plate rocker for 15 min, followed by incubation at room temperature for 45 min in the absence of light. The Hgb released from RBCs was detected based on the absorbance of cyanmethemoglobin measured at 540 nm.
130g RCF for 10 min. 100 μL of the supernatant from each tube was added to a UV transparent plate in triplicate with the addition of 100 μL CMHR to each well. The plate was sealed and gently mixed on a plate rocker for 15 min, followed by incubation at room temperature for 45 min in the absence of light. The Hgb released from RBCs was detected based on the absorbance of cyanmethemoglobin measured at 540 nm.
Hemolysis assay
The presence of free QS21 in ALFQ was detected using the described hemolysis assay. The schematic diagram of the optimized analytical workflow is presented in Fig. S1. Briefly, ALFQ containing 100 μg mL−1 QS21 was incubated with washed human RBC (2.5 × 108 cells mL−1) for 10 min. The mixture was then centrifuged at room temperature and 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 130g RCF for 10 min. An aliquot of the supernatant (100 μL) was added to a UV transparent plate containing 100 μL of CMHR. The plate was sealed and gently rocked for 15 min and left undisturbed for an additional 45 min in the dark. The presence of any free QS21 in ALFQ was detected by the release of Hgb from RBCs, which was detected as cyanmethemoglobin at 540 nm, using a UV-visible plate reader. The amount of “Hgb in test sample” was interpolated in the measured absorbance as a function of Hgb concentration in the standard Hgb calibration curve. In a similar manner, the RBC hemolysis by free QS21 at 1–5 μg mL−1 in SPBS was determined as described above. CMHR was used as a positive control for 100% hemolysis of RBCs and was performed in parallel. The Hgb determined from the positive control corresponds to the “Total blood Hgb” in the donor sample. The presence of free QS21 was detected from the hemolysis of RBCs, which was calculated using eqn (1).
130g RCF for 10 min. An aliquot of the supernatant (100 μL) was added to a UV transparent plate containing 100 μL of CMHR. The plate was sealed and gently rocked for 15 min and left undisturbed for an additional 45 min in the dark. The presence of any free QS21 in ALFQ was detected by the release of Hgb from RBCs, which was detected as cyanmethemoglobin at 540 nm, using a UV-visible plate reader. The amount of “Hgb in test sample” was interpolated in the measured absorbance as a function of Hgb concentration in the standard Hgb calibration curve. In a similar manner, the RBC hemolysis by free QS21 at 1–5 μg mL−1 in SPBS was determined as described above. CMHR was used as a positive control for 100% hemolysis of RBCs and was performed in parallel. The Hgb determined from the positive control corresponds to the “Total blood Hgb” in the donor sample. The presence of free QS21 was detected from the hemolysis of RBCs, which was calculated using eqn (1). | (1) |
The % hemolytic activity was calculated to account for the ALFQ light scattering using eqn (2). A negative percentage value was reported as 0%.
| % hemolytic activity = % hemolysis of (ALFQ + RBCs) − % hemolysis of (ALFQ + SPBS) − % hemolysis of (SPBS + RBCs) | (2) |
Method qualification
Method qualification was conducted to establish the selectivity/specificity, linearity, precision, and accuracy of the method to detect and semi-quantitate the presence of free QS21 in ALFQ adjuvant formulations, following ICH quality guidelines for validation.31Measurement of released Hgb
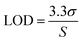 | (3) |
The LOQ on the other hand was calculated using eqn (4). The LOQ should exhibit a S/N ≥ 10. Both LOD and LOQ were established from 34 independent sets of calibration standards.
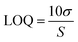 | (4) |
Detection of free QS21 by hemolysis
The purpose of this method was to detect free QS21 in ALFQ adjuvant formulations using RBCs as probes. The method was characterized by LOD and precision. Every experiment was accompanied by system suitability solutions to ensure the validity of the results during the time of the assay. Prior to actual hemolysis study to detect free QS21, donor selection was assessed to ensure the use of appropriate donor blood with sensitivity of 20–60% hemolysis using 1.5 μg mL−1 free QS21.Results and discussion
Hemolysis assay: concept
The described cell-based assay was developed to estimate the amount of free QS21 in ALFQ vaccine adjuvant via lysis of RBCs. As described above, the hemolytic activity of free QS21 has been documented in vitro.9,14,15 In this method, human RBCs were employed as probes for detecting free QS21 in adjuvant formulations, where its interactions with RBCs resulted in lysis, leading to the release of Hgb, which can be detected by UV-vis spectroscopy (Fig. 1). Direct determination of Hgb by UV-visible spectroscopy requires anaerobic conditions to avoid oxidation of the Fe heme metal center. To avoid the instability of the released Hgb-Fe(II), which has the propensity to undergo oxidation to Hgb-Fe(III) under aerobic conditions, this method involves a straightforward derivatization of Hgb with CMHR to form a stable cyanmethemoglobin with the Fe(III) metal center (Fig. 1B), which has a characteristic Q-band absorption at 540 nm (Fig. 2A). Derivatization of Hgb to cyanmethemoglobin by CMHR is well known and has been employed in the determination of Hgb in biological and non-biological samples.32–34 The amount of free QS21 in the ALFQ adjuvant formulation is correlated with the amount of released Hgb from RBCs.Detection of Hgb. Analytical conditions, matrices, and mode of detection were explored to reliably quantify the released Hgb. As shown in Fig. 2A, interaction of Hgb with CMHR resulted in the formation of a new band at 540 nm, characteristic of cyanmethemoglobin. Optimized conditions involve derivatization of Hgb in SPBS to cyanmethemoglobin using CMHR, followed by absorbance measurement at 540 nm using a UV-vis plate reader, with a linear range of 0.5–16 mg mL−1 Hgb in SPBS (Fig. S2). Since this method is designed to detect the released Hgb from the lysis of RBCs by free QS21 in ALFQ adjuvant, the effects of the sample matrix and the specificity of the signal were evaluated. Negative controls such as buffer only (SPBS) and CMHR only samples did not render an absorbance at 540 nm, suggesting the specificity of this band for cyanmethemoglobin from the interactions of Hgb and CMHR (Fig. 2A). However, inclusion of the particulate ALFQ in the sample containing only CMHR in SPBS resulted in a baseline shift (Fig. 2A), which can be ascribed in part to the light scattering effect of liposomes. This had an apparent influence on the absorbance measurement at a single wavelength (540 nm) using a plate reader. The contribution of the liposome light scattering effect was subtracted mathematically during the actual calculation of hemolysis (vide infra). We introduced the concept of hemolytic activity that can be described using eqn (2) (Experimental section), which isolates the lysis caused by free QS21 and removes the contribution of light scattering.
We have also explored several blood donor samples with different concentrations of free QS21 to establish the reliability of the method to determine released Hgb from RBC samples. As shown in Fig. 2B, the amount of released Hgb and extent of RBC hemolysis are directly proportional to the amount of free QS21 in a quadratic relation. The hemolysis profiles of 10 donor blood samples exhibit variations of ∼10% RSD, with R2 ranging from 0.91 to 0.99 (Fig. 2B and Table S1), which can be ascribed in part to donor variability. To address the variable sensitivity of blood donor samples, we have set criteria that were used for selecting appropriate donor blood in the assay. Those donor blood samples that exhibit hemolysis in the range 20–60% at 1.5 μg mL−1 free QS21 in SPBS (i.e., in the absence of ALFQ) will be used in the actual hemolysis assay of ALFQ samples to determine the concentration of free QS21. The data described in this work were obtained with donors that met the criteria to ensure appropriate and consistent sensitivity of the assay.
Detection of free QS21 in ALFQ by RBC hemolysis. As observed, the extent of hemolysis (%) is directly proportional to the amount of free QS21 in a quadratic relation (Fig. 2B). Thus, in this assay, the release of Hgb from RBC lysis in the ALFQ adjuvant expressed as % hemolysis or % hemolytic activity is indicative of the presence and quantitative measure of free QS21. To this effect, RBCs as probes should have a cell density that under 100% lysis conditions releases Hgb that is within the dynamic range of Hgb detection (0.5–16.0 mg mL−1). Triton X-100 (1%) and CMHR known to cause 100% RBC lysis were used to lyse varying concentrations of RBCs (1 × 104 to 5 × 108 cells mL−1; Table 1) and the released Hgb was quantified as described in the method section. A cell density of 2.5 × 108 cells mL−1 was selected as the optimal RBC concentration. The 100% lysis of this RBC density translates to an Hgb concentration of 9.28 mg mL−1 (Table 1), which is within the linear range of the Hgb standard curve (0.5–16.0 mg mL−1; Fig. S2).
| Cell conc. (cells mL−1) | Abs. at 540 nm (mean ± SD)a | Calc. Hgb conc. (mg mL−1) |
|---|---|---|
| a Background was subtracted from each mean value. SD = standard deviation. | ||
| 1.0 × 104 | 0.006 ± 0.001 | 0.453 |
| 1.0 × 105 | 0.010 ± 0.003 | 0.471 |
| 1.0 × 106 | 0.007 ± 0.002 | 0.458 |
| 1.0 × 107 | 0.069 ± 0.003 | 0.741 |
| 1.0 × 108 | 0.602 ± 0.021 | 3.181 |
| 2.5 × 108 | 1.738 ± 0.014 | 9.280 |
| 5.0 × 108 | 3.960 ± 0.009 | 20.971 |
The analytical capability of the method to estimate free QS21 concentration was evaluated using the ALFQ adjuvant spiked with increasing amounts of free QS21 (Fig. 3). The relationship of % hemolytic activity with the estimated amounts of free QS21 in SPBS exhibits a quadratic behavior with R2 > 0.99. Hence, from this relationship, the estimated amounts of free QS21 in ALFQ adjuvant can be extrapolated from the measured hemolytic activity of RBCs.
Selectivity/specificity. The specificity of the method for detecting Hgb using CMHR to generate a more stable cyanmethemoglobin was exemplified using RBCs with sample matrices such as SPBS, particulate ALFQ, and with or without CMHR. As shown in Table 2, free hemoglobin was only detected in samples containing both RBCs and CMHR. The observed minor baseline shift in samples containing ALFQ due to the light scattering of liposomes at 540 nm resulted in artifactual % hemolysis (eqn (1)). This contribution was corrected by calculating the % hemolytic activity using eqn (2), which accounts for this scattering effect (see the Experimental section).
| Analytec | Day | A 540 (mean ± SD) | Calc. Hgb conc. (mg mL−1) | Hemolysis (%) |
|---|---|---|---|---|
| a Background was subtracted from each mean value. b These values are due to the baseline shift from the light scattering effect liposomes. c All samples are in SPBS. | ||||
| RBCs | 1 | 0.028 ± 0.001 | 0.223 | 3 |
| 2 | 0.044 ± 0.003 | 0.138 | 2 | |
| 3 | 0.005 ± 0.000 | 0.000 | 0 | |
| 4 | 0.024 ± 0.001 | 0.183 | 3 | |
| CMHR | 1 | 0.000 ± 0.000 | — | — |
| 2 | 0.000 ± 0.000 | — | — | |
| 3 | 0.000 ± 0.000 | — | — | |
| 4 | 0.000 ± 0.000 | — | — | |
| RBCs + CMHR | 1 | 1.381 ± 0.0024 | 8.060 | 100 |
| 2 | 1.310 ± 0.016 | 7.230 | 100 | |
| 3 | 1.321 ± 0.029 | 7.605 | 100 | |
| 4 | 1.183 ± 0.001 | 6.516 | 100 | |
| RBCs (sup.) + CMHR | 1 | 0.010 ± 0.002 | 0.123 | 2 |
| 2 | 0.010 ± 0.003 | 0.000 | 0 | |
| 3 | 0.003 ± 0.001 | 0.000 | 0 | |
| 4 | 0.008 ± 0.000 | 0.094 | 1 | |
| ALFQ + CMHR | 1 | 0.086 ± 0.006 | 0.563 | 7b |
| 2 | 0.086 ± 0.008 | 0.372 | 5b | |
| 3 | 0.118 ± 0.010 | 0.652 | 9b | |
| 4 | 0.114 ± 0.008 | 0.673 | 10b | |
| ALFQ | 1 | 0.170 ± 0.028 | 1.047 | 13b |
| 2 | 0.111 ± 0.001 | 0.516 | 7b | |
| 3 | 0.196 ± 0.014 | 1.099 | 14b | |
| 4 | 0.220 ± 0.028 | 1.251 | 19b | |
| ALFQ + RBCs | 1 | 0.079 ± 0.006 | 0.518 | 6b |
| 2 | 0.113 ± 0.004 | 0.523 | 7b | |
| 3 | 0.099 ± 0.007 | 0.538 | 7b | |
| 4 | 0.112 ± 0.010 | 0.661 | 10b | |
Linearity/dynamic range and LOD/LOQ. The linear/dynamic range of the described method for detecting Hgb was observed in the range of 0.5–16.0 mg mL−1 (Fig. S2), with regression coefficient, R2 > 0.99, and % recoveries of 80 to 120% at each calibration point. The linear range was limited by the saturation of UV detection beyond the upper limit of quantification of Hgb. The method limit of detection (LOD) and limit of quantification (LOQ) were determined to be 0.413 ± 0.166 mg mL−1 and 1.252 ± 0.504 mg mL−1, respectively. These values were established from 34 independent assay runs, following eqn (3) and (4) (Experimental section).
Accuracy and precision. The accuracy and precision of the method were established at three levels of Hgb concentrations (4.0 mg mL−1, 8.0 mg mL−1, and 12.0 mg mL−1) in SPBS in the presence of ALFQ to mimic the matrix of the actual hemolysis samples. As shown in Table 3, the % recoveries were 85–115% for 4 mg mL−1 Hgb, 89–102% for 8 mg mL−1 Hgb, and 81–97% for 12 mg mL−1 Hgb. These findings showed that at these concentration levels, the observed baseline shift due to the light scattering of particulate liposomes was not substantial enough to influence the accurate measurement of Hgb. Overall, the described method exhibits good accuracy for measuring free Hgb in the presence of particulate ALFQ. Tighter variance was observed in the between-run variability with RSD < 7% at all concentration levels (Table 3).
| Analyte | Day | A 540 (mean ± SD) | Calc. Hgb mean (mg mL−1) ± SD | % recovery | % RSD |
|---|---|---|---|---|---|
| a Background was subtracted from each mean value. | |||||
| 4 mg mL–1 Hgb + ALFQ | 1 | 0.602 ± 0.021 | 3.633 ± 0.047 | 92 | 1 |
| 0.597 ± 0.004 | 91 | ||||
| 0.588 ± 0.010 | 90 | ||||
| 2 | 0.826 ± 0.027 | 4.450 ± 0.154 | 115 | 3 | |
| 0.773 ± 0.020 | 107 | ||||
| 0.806 ± 0.014 | 112 | ||||
| 3 | 0.575 ± 0.017 | 3.497 ± 0.087 | 85 | 2 | |
| 0.601 ± 0.013 | 89 | ||||
| 0.594 ± 0.018 | 88 | ||||
| 8 mg mL–1 Hgb + ALFQ | 1 | 1.146 ± 0.008 | 7.186 ± 0.055 | 91 | 1 |
| 1.136 ± 0.010 | 90 | ||||
| 1.129 ± 0.004 | 89 | ||||
| 2 | 1.385 ± 0.029 | 7.716 ± 0.135 | 98 | 2 | |
| 1.340 ± 0.010 | 95 | ||||
| 1.376 ± 0.040 | 97 | ||||
| 3 | 1.170 ± 0.025 | 7.811 ± 0.567 | 89 | 7 | |
| 1.325 ± 0.009 | 102 | ||||
| 1.327 ± 0.008 | 102 | ||||
| 12 mg mL–1 Hgb + ALFQ | 1 | 1.540 ± 0.015 | 9.764 ± 0.080 | 82 | 1 |
| 1.516 ± 0.020 | 81 | ||||
| 1.533 ± 0.012 | 82 | ||||
| 2 | 1.849 ± 0.012 | 10.700 ± 0.313 | 88 | 3 | |
| 1.854 ± 0.053 | 88 | ||||
| 1.945 ± 0.041 | 92 | ||||
| 3 | 1.883 ± 0.024 | 11.270 ± 0.363 | 97 | 3 | |
| 1.768 ± 0.033 | 91 | ||||
| 1.820 ± 0.028 | 94 | ||||
Dynamic range and limit of detection/quantification. The dynamic range of the method for detecting free QS21 was explored and established in the range of 1.0–64 μg mL−1. At this range, the % hemolytic activity as a function of free QS21 concentration exhibits a quadratic relationship, with R2 ∼ 0.997. The LOD and LOQ were not warranted, instead the LLOQ was considered, which shows method sensitivity in the micromolar range (Fig. S3).
Accuracy and precision. Laboratory grade ALFQ suspensions with spiked QS21, 5 μg mL−1 (low; n = 6), 10 μg mL−1 (mid; n = 6) and 25 μg mL−1 (high; n = 6), were used to establish the accuracy and precision expressed as the within-run repeatability of the described method. Table 4 shows that at lower spiked concentration levels (5.0 μg mL−1 and 10.0 μg mL−1), free QS21 measurements exhibit low % recoveries in the range of 20% to 59%, with % RSD > 90%. Although we expected high sensitivity of RBC lysis by free QS21, as observed this method shows non-quantitative recoveries at low spiked QS21 concentrations. These observed poor accuracy and precision at very low free QS21 concentrations are due to the combination of the ability of ALFQ particulates to act as a sink of free QS21 and its contribution to the light scattering effect, and blood donor variability. However, excellent accuracy with % recoveries of 97% to 111%, and precision characterized by % RSD ∼ 6% were observed in ALFQ samples containing 25 μg mL−1 spiked free QS21, which is within the acceptable limit. The results from the accuracy and precision studies showed that the described method is applicable for measuring free QS21 in ALFQ samples at the lowest nominal concentrations of ∼25 μg mL−1, characterized by % recoveries within the acceptable limit of 100 ± 20% with good measurement variability. The method limit of quantification, LOQ = ∼25 μg mL−1, translates to 15 μg in 0.6 mL ALFQ suspension. In a vialed cGMP ALFQ, the observed method LOQ (15 μg) is only ∼12% of the total 120 μg bound QS21. In the initial release of ALFQ, all QS21 is bound in the liposomes (i.e., no free QS21 detected; Table 5). However, during long-term storage, degradation of liposomes may result in a leakage of free QS21. This method can estimate free QS21 with a limit of 12% (15 μg) relative to the total QS21 in ALFQ, which is far below the toxicity limit of 50 μg described in the literature.1 In the case of ALFQ, the presence of 50 μg free QS21 translates to its severe degradation (i.e., 42% QS21 leakage relative to total QS21 content) during long-term storage. This method can detect the leakage of free QS21 far before reaching this toxicity limit. This observation suggests the relevance of the method to monitor free QS21 in ALFQ formulations to establish and support product safety during stability studies.
| Samples | % hemolytic activity (mean ± SD) | Free QS21 conc. (μg mL−1) | % recoveries (mean ± SD) | |
|---|---|---|---|---|
| Calculated (mean ± SD) | Theoretical (spiked) | |||
| ALFQ (n = 6) | 3.36 ± 1.45 | 1.15 ± 1.06 | 5.0 | 23.0 ± 21.2 |
| ALFQ (n = 6) | 7.21 ± 6.02 | 4.10 ± 5.88 | 10.0 | 41.0 ± 58.8 |
| ALFQ (n = 6) | 28.64 ± 1.69 | 28.64 ± 1.69 | 25.0 | 114 ± 6.8 |
The results consistently rendered virtually no hemolytic activities across blood donors (i.e., no free QS21 detected), suggesting consistency of the method for detecting free QS21. In these release samples, it is difficult to quantify free QS21, as the % hemolytic activities are below the LOQ of the method.
Conclusions
The described cell-based assay in a plate reader platform demonstrated promising analytical characteristics for detecting and estimating free QS21 in the ALFQ vaccine adjuvant. The method qualification studies showed that the designed assay for measuring released Hgb from RBC lysis by free QS21 exhibits good specificity/selectivity, and measurement accuracy and precision. Consequently, for the purpose of determining free QS21 in the ALFQ adjuvant, the method exhibits good accuracy and precision, and sensitivity in the micromolar range. This method exhibits a medium analytical throughput suitable for screening QS21-based adjuvant formulations, and most importantly, does not require the use of any expensive or complicated instrumentation system. Currently, cGMP manufactured ALFQ is released using a battery of tests, where the majority of the regulatory specifications focuses on physicochemical properties and microbial content. These tests were complemented with the described hemolysis assay that serves as a surrogate toxicity test during release and stability studies. The applicability of the method spans to particulate vaccine adjuvants containing QS21.Author contributions
The manuscript was written through contributions of all authors. Conceptualization and investigation, E. K., G. R. M., E. G. A., O. B. T.; data curation, E. K., C. R., A. C., E. G. A.; writing, review and editing, all authors; supervision and project administration, E. K., E. G. A., O. B. T., G. R. M. All authors have read and agreed to the current submission of the manuscript. All authors have given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: schematic diagram of hemolysis assay, representative calibration curve showing the linear relationship of absorbance as a function of Hgb concentration, % hemolytic activity as a function of spiked QS21 concentration in a quadratic relation, regression studies of the hemolysis as a function of free QS21 concentration, and initial system suitability results. See DOI: https://doi.org/10.1039/d5ay00854a.Acknowledgements
This research work was supported by the Military Infectious Disease Research Program (MI230009) and through Cooperative Agreement Awards (#W81XWH-07-2-067, #HT94252430004) between the Henry M. Jackson Foundation for the Advancement of Military Medicine and the U.S. Department of Defense (DoD). The authors would like to thank Alexander Anderson for outstanding technical assistance.References
- G. Ragupathi, J. R. Gardner, P. O. Livingston and D. Y. Gin, Expert Rev. Vaccines, 2011, 10, 463–470 CrossRef CAS PubMed.
- T. A. Gilewski, G. Ragupathi, M. Dickler, S. Powell, S. Bhuta, K. Panageas, R. R. Koganty, J. Chin-Eng, C. Hudis and L. Norton, et al. , Clin. Cancer Res., 2007, 13, 2977–2985 CrossRef CAS PubMed.
- A. Fernandez-Tejada, E. K. Chea, C. George, N. Pillarset-ty, J. R. Gardner, P. O. Livingston, G. Ragupathi, J. S. Lewis, D. S. Tan and D. Y. Gin, Nat. Chem., 2014, 6, 635–643 CrossRef CAS PubMed.
- M. A. Lacaille-Dubois, Phytomedicine, 2019, 60, 1–12 CrossRef PubMed.
- J. N. Hutter, P. Robben, C. Lee, M. Hamer, J. Moon, K. Merino, L. Zhu, H. Galli, X. Quinn, D. Brown, E. Dun-can, J. Bolton, X. Zou, E. Angov, D. Lanar, M. Rao, G. Matyas, Z. Beck, E. Bergmann-Leitner, L. Siosson, N. Waters, V. Ngauy, J. Regules and S. Dutta, Vacine, 2022, 40, 5781–5790 CrossRef CAS PubMed.
- P. Bejon, J. Lusingu, A. Olotu, A. Leach, M. Lievens, J. Vekemans, S. Mshamu, T. Lang, J. Gould, M. C. Dubois, M. A. Demoitié, J. F. Stallaert, P. Vansadia, T. Carter, P. Njuguna, K. O. Awuondo, A. Malabeja, O. Abdul, S. Gesase, N. Mturi, C. J. Drakeley, B. Savarese, T. Villafana, W. R. Ballou, J. Cohen, E. M. Riley, M. M. Lemnge, K. Marsh and L. von Seidlein, N. Engl. J. Med., 2008, 359, 2521–2532 CrossRef CAS PubMed.
- H. Lal, A. L. Cunningham, O. Godeaux, R. Chlibek, J. Diez-Domingo, S.-J. Hwang, M. J. Levin, J. E. McElhaney, A. Poder and J. Puig-Barberà, et al. , N. Engl. J. Med., 2015, 372, 2087–2096 CrossRef PubMed.
- T. F. Schwarz, S. J. Hwang, P. Ylisastigui, C. S. Liu, K. Takazawa, M. Yono, J. E. Ervin, C. P. Andrews, C. Fogarty, T. Eckermann, D. Collete, M. de Heusch, N. De Schrevel, B. Salaun, A. Lambert, C. Maréchal, A. Olivier, P. Nakanwagi, M. Lievens and V. Hulstrøm, J. Infect. Dis., 2024, 230, e102–e110 CrossRef CAS PubMed.
- C. R. Kensil, U. Patel, M. Lennick and D. Marciani, J. Immunol., 1991, 146, 431–437 CrossRef CAS.
- R. Richou, R. Jensen and C. Belin, Rev. Immunol. Ther. Antimicrob., 1964, 28, 49–62 CAS.
- N. Garcon, P. Chomez and M. Van Mechelen, Expert Rev. Vaccines, 2007, 6, 723–739 CrossRef CAS PubMed.
- M. Swart, J. Allen, B. Reed, A. Izquierdo Gil, J. Verspuij, S. Schmit-Tillemans, A. Chakkumkal, M. Findeis, A. V. Hafner, C. Harjivan, R. Kurnat, H. Kuipers, R. Zahn and B. Brandenburg, Vaccines, 2024, 12, 1435 CrossRef CAS PubMed.
- J. L. Cleland, C. R. Kensil, A. Lim, N. E. Jacobsen, L. Ba-sa, M. Spellman, D. A. Wheeler, J.-Y. Wu and M. F. Powell, J. Pharm. Sci., 1996, 85, 22–28 CrossRef CAS PubMed.
- Z. Beck, G. R. Matyas and C. R. Alving, Biochim. Biophys. Acta, 2015, 1848, 775–780 CrossRef CAS PubMed.
- N. Garcon and A. Di Pasquale, Hum. Vaccines Immunother., 2017, 13, 19–33 CrossRef PubMed.
- D. Drane, C. Gittleson, J. Boyle and E. Maraskovsky, Expert Rev. Vaccines, 2007, 6, 761–772 CrossRef CAS PubMed.
- N. Garcon, G. Leroux-Roels and W. F. Cheng, Perspectives in Vaccinology, 2011, 1, 89–113 CrossRef.
- M. Rao, K. K. Peachman and C. Alving, Curr. Top. Microbiol. Immunol., 2021, 433, 1–28 CAS.
- C. Alving, K. K. Peachman, G. Matyas, M. Rao and Z. Beck, Expert Rev. Vaccines, 2020, 19, 279–292 CrossRef CAS PubMed.
- B. L. Ober Shepherd, P. T. Scott, J. N. Hutter, C. Lee, M. D. McCauley, I. Guzman, C. Bryant, S. McGuire, J. Kennedy, W. H. Chen, A. Hajduczki, T. Mdluli, A. Valencia-Ruiz, M. F. Amare, G. R. Matyas, M. Rao, M. Rolland, J. R. Mascola, S. C. De Rosa, M. J. McElrath, D. C. Montefiori, L. Serebryannyy, A. B. McDermott, S. A. Peel, N. D. Collins, M. G. Joyce, M. L. Robb, N. L. Michael, S. Vasan and K. Modjarrad, Lancet Microbe, 2024, 5, e581–e593 CrossRef PubMed.
- N. Petrovsky, Drug Saf., 2015, 38, 1059–1074 CrossRef CAS PubMed.
- W. Yuan, Z. Wang, Y. Zou and G. Zheng, Biomedicines, 2024, 12, 1–12 Search PubMed.
- H. So, H. Yoon, D. Choi, Y. Kwon, J. Sung, T. G. Lee, E. Park, H. Cho, B. M. Lee, M. J. Cho and W. Ryu, Mol. Cells, 1997, 7, 178–186 CrossRef CAS PubMed.
- E. G. Abucayon, R. C. Barrientos, O. B. Torres, S. Sweeney, C. Whalen and G. R. Matyas, ACS Omega, 2023, 8, 21016–21025 CrossRef CAS PubMed.
- M. Hemida, R. C. Barrientos, C. Kinsey, N. Kuster, M. Bhavsar, A. G. Beck, H. Wang, A. Singh, P. Aggarwal, A. Arcinas, M. Mukherjee, E. Appiah-Amponsah and E. L. Regalado, ACS Pharmacol. Transl. Sci., 2024, 7, 2901–3278 CrossRef PubMed.
- N. E. Jacobsen, W. J. Fairbrother, C. R. Kensil, A. Lim, D. A. Wheeler and M. F. Powell, Carbohydr. Res., 1996, 280, 1–14 CrossRef CAS PubMed.
- F. Wallace, C. Fontana, F. Ferreira and C. Olivaro, Molecules, 2022, 27, 1–13 CrossRef PubMed.
- Z. Beck, G. R. Matyas, R. Jalah, M. Rao, V. R. Polonis and C. R. Alving, Vaccine, 2015, 33, 5578–5587 CrossRef CAS PubMed.
- G. R. Matyas, J. M. Muderhwa and C. R. Alving, Methods Enzymol., 2003, 562, 34–50 Search PubMed.
- P. Singh, Z. Beck, G. R. Matyas and C. R. Alving, J. Liposome Res., 2019, 29, 247–250 CrossRef CAS PubMed.
- FDA Guidance for Industry: Q2(R1) Validation of Analytical Procedures: Text and Methodology, 2005 Search PubMed.
- C. Coll-Satue, M. M. T. Jansman and L. Hosta-Rigau, Biomolecules, 2024, 14, 1–17 CrossRef PubMed.
- M. C. Knirsch, F. Dell'Anno, M. Salerno, C. Larosa, B. Polakiewicz, R. Eggenhoffner and A. Converti, Bioprocess Biosyst. Eng., 2017, 40, 431–438 CrossRef CAS PubMed.
- A. Dessy, A. M. Piras, G. Schirò, M. Levantino, A. Cupane and F. Chiellini, Eur. J. Pharm. Sci., 2011, 43, 57–64 CrossRef CAS PubMed.
Footnote |
| † Current address: Merck Sharp & Dohme LLC., 126 E. Lincoln Ave., Rahway, New Jersey 07065, USA. |
| This journal is © The Royal Society of Chemistry 2025 |


