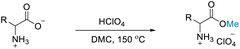
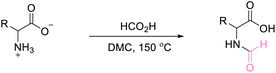
A novel approach to amino acid synthesis: acid-assisted reactions with dimethyl carbonate for efficient O-methylated, N,O-methylated and N-formylated derivatives†
Pattamabhorn
Worsawat
a,
Pakin
Noppawan
 b,
Nontipa
Supanchaiyamat
b,
Nontipa
Supanchaiyamat
 a,
Himmy
Wu
c,
Con R.
McElroy
a,
Himmy
Wu
c,
Con R.
McElroy
 cd and
Andrew J.
Hunt
cd and
Andrew J.
Hunt
 *a
*a
aMaterials Chemistry Research Center (MCRC), Department of Chemistry and Centre of Excellence for Innovation in Chemistry, Faculty of Science, Khon Kaen University, Khon Kaen, 40002, Thailand. E-mail: andrew@kku.ac.th
bDepartment of Chemistry, Faculty of Science, Mahasarakham University, Maha Sarakham, 44150, Thailand
cDepartment of Chemistry, University of York, York, YO10 5DD, UK
dSchool of Chemistry, University of Lincoln, Brayford Pool Campus, Lincoln, LN6 7TS, UK
First published on 20th February 2024
Abstract
A novel and efficient method for the modification of amino acids using a dimethyl carbonate (DMC) and acid system has been developed. N-Methylation, N,O-dimethylation, and N-formylation of various amino acids have been successfully achieved using this simple acid-assisted method (>99% conversions and >99% yields) and the modified amino acids were fully characterized using NMR spectroscopy. This method offers several advantages, including the use of a sustainable and cost-effective reagent, high selectivity, efficiency, eco-friendliness, broad applicability to a range of amino acids with different side chain functionalities, and being secure in the knowledge that there is no risk of racemization and epimerization. This study provides a new, sustainable, and practical approach for the modification of amino acids, which has potential applications in drug discovery and chemical biology.
Introduction
Amino acids are fundamental components of proteins, and their modification has gained significant interest in a number of scientific sectors.1 Amino acids and peptide-based drugs play a crucial role in the current landscape of the health industry.2–5 Within the structures of modern pharmaceuticals, one frequently encounters residues of individual amino acids and their various chemical modifications. This underscores the pervasive influence and integral contribution of amino acids to the development and formulation of pharmaceutical compounds,2,3,6,7 especially derivatives of amino acid methyl esters, N-methyl amino acid methyl esters, and N-formyl amino acid, which are important intermediates utilized in a variety of fields, including medical chemistry,8–11 chiral auxiliary,12–16 peptide synthesis,17–20 and polymer materials.21–23 The modification of amino acids is an important aspect of organic chemistry, with applications ranging from the synthesis of peptides and proteins to the development of pharmaceuticals and other high-value chemicals.24–28 A key challenge in this area is the development of selective synthetic routes for the modification of specific functional groups within the amino acid molecule.Numerous reagents have been reported for the conversion of amino acids into amino acid methyl esters, including methanol/trimethylchlorosilane,29 methanol/hydrochloric acid,30 butanol/thionyl chloride,31 dichloromethane/thionyl chloride,32 dimethyl sulfate,33,34 2,2-dimethoxypropane/hydrochloric acid,35 ion-exchange resins (amberlyst™-15),36 and diazomethane.37 Frequently, procedures require multiple complicated and potentially wasteful synthetic steps to achieve the desired product, such as N-protection, esterification, and deprotection. While some of these procedures are used regularly, they have several drawbacks, including arduous workups and harsh reaction conditions, as well as having safety and waste disposal issues.
N-Methylamino acids are one of the naturally occurring biologically active compounds,38–40 such as cyclosporine,41,42 dolastatins,43–45 and didemnins,46,47 which have fascinating therapeutic profiles. The inclusion of N-methylamino acids into physiologically active peptides have been shown to regulate backbone conformation and enhance pharmacological characteristics.24,26,27 There have been a few studies on the synthesis of N-methyl amino acid methyl esters. The use of conventional methylating agents such as diazomethane,9 methyl iodide,48 and trimethyloxonium tetrafluoroborates49 has limited interest in N-methyl amino acid methyl ester compounds due to reagent handling issues and their chronic health risks.50–54
In chemical synthesis and medicinal chemistry, N-formylation of amines is essential. Several medicinally active compounds are formed from formamides. N-Formyl protection is widely used in peptide synthesis because it can be easily deprotected without damaging peptide bonds and used as a precursor for isocyanide and formamidine synthesis.55 Reported literature methods for the preparation of N-formyl amino acid derivatives use reagents such as anhydrous ammonium formate,20 formyloxyacetoxyphenyl-methane,56 acetic formic anhydride,57 trimethylorthoformate,58 triethylorthoformate,59N-(diethylcarbamoyl)-N-methoxyform-amide,60 pivalaldehyde,61 cyanomethylformate,62 and formic acid/EDCI/glyceroacetonide-oxyma.63 Most of these procedures involve the use of hazardous volatile organic solvents, particularly chlorinated hydrocarbons, which have presented a threat to the environment and human health.64,65 Such procedures require multistep reactions that are extremely intricate, difficult to prepare, and require costly reagents, inert atmospheres and dry solvents, and use poisonous or unstable chemicals to yield a vast array of products.
These traditional procedures for methylation and formylation of amino acids rely on hazardous and costly chemicals, limiting their practical application in large-scale synthesis. Consequently, it is necessary to develop a novel, convenient, and simple approach for the modification of amino acids, which employs sustainable solvents/reagents and is cost effective. Dimethyl carbonate (DMC) is non-toxic and has found application as a solvent or reagent in agrochemicals, pharmaceuticals, polymers, solvents for coatings and adhesives, green oxidants, lithium-ion battery electrolytes and gasoline additives.66–72 DMC has long been viewed as a sustainable alternative to highly toxic and hazardous reagents such as dimethyl sulphate and methyl halides, used for synthetic methylation reactions. Owing to its low toxicity and biodegradability, DMC has been utilized as a green solvent73 and a green substitute for hazardous intermediates such as phosgene.74,75 DMC can be commercially produced from waste carbon dioxide, making this a truly sustainable solvent and chemical.76,77 The by-products of the methylation process are carbon dioxide and methanol, which are easily separated and can be recycled into the DMC synthesis process.78 Previous research has shown that under acid-catalysed conditions, DMC is able to perform highly efficient carboxymethylation and methylation of amines.79 Ji et al.80 reported the use of DMC to O-methylate a few examples of N-Boc amino acids under basic conditions with moderate to high yields. Until now, no publication has studied amino acid modification with DMC under acidic conditions.
To the best of our knowledge, this work reports the first modification of amino acids with DMC under acidic conditions to date. Herein, a sustainable, single step, highly selective and high-yielding alternative method for the modification of O-methylation, N,O-dimethylation, and N-formylation of amino acids based on treatment with DMC in acid-assisted reactions is reported. In addition, these methods are facile and do not require a dry solvent or an inert atmosphere. Green chemistry metrics have been assessed and to better comprehend the reaction's selectivity, the reaction mechanism has been investigated.
Results and discussion
Our initial studies focused on the selectivity of DMC modification of DL-alanine catalysed by homogenous systems. Various Brønsted acids (H2SO4, HClO4, HCl, NHO3, and formic acid) were investigated under reflux conditions (90 °C, 24 h) or in a high-pressure reactor (150 °C, 6 h). Under reflux conditions, DL-alanine is slightly soluble in DMC. Analysis by 1H NMR reveals no product since only the characteristic signals of the starting material were detected. Thus, no reaction occurs under reflux conditions (Fig. S49–S53†). The solubility of DL-alanine is enhanced when the reaction is carried out in a high-pressure reactor (150 °C, 6 hours) and the crude product is a homogeneous phase, which agreed with the previously reported literature.81 Under some conditions, more than 99% of DL-alanine was converted with excellent selectivity (>99%) into alanine methyl ester (Table 1, entry 6, and Scheme 1) with DMC acting as a reagent and N-formyl alanine (Table 1, entry 10, and Scheme 1), with DMC acting as a solvent. Although only 59.68% was converted into N-methyl alanine methyl ester, the selectivity was also high (>99%) (Table 1, entry 7, and Scheme 1).| Entry | Brønsted acid | Temp (°C) | Time (h) | Conversionb (%) | Selectivityb (%) |
|---|---|---|---|---|---|
| a Conditions: DL-alanine (1 mmol), Brønsted acid (1 mmol), DMC (1 mmol). b Calculated by 1H NMR and structure confirmed by HMBC. | |||||
| 1 | HClO4 | 90 | 24 | — | — |
| 2 | H2SO4 | 90 | 24 | — | — |
| 3 | HCl | 90 | 24 | — | — |
| 4 | HNO3 | 90 | 24 | — | — |
| 5 | HCO2H | 90 | 24 | — | — |
| 6 | HClO4 | 150 | 6 | >99 | >99 (O-methylation) |
| 7 | H2SO4 | 150 | 6 | 59.68 | >99 (N,O-dimethylation) |
| 8 | HCl | 150 | 6 | >99 | — |
| 9 | HNO3 | 150 | 6 | >99 | — |
| 10 | HCO2H | 150 | 6 | >99 | >99 (N-formylation) |
HMBC method was used to confirm the structure of the reaction products (ESI†). Thus, highlighting the function of DMC at elevated temperatures (>150 °C) in the presence of H2SO4 and HClO4 as methylating agents is consistent with the previous research,82 while a combination of formic acid and DMC functions as a formylating agent.
Nonetheless, HCl and HNO3 exhibited non-selective conversion under the same conditions, the products of which cannot be identified based on the 1H NMR spectra, (Fig. S54 and S55†). According to the literature, a mixture of HCl and H2O can induce the decomposition of amino acids,83 whereas HNO3 can be decomposed under high-temperature conditions to form NO2,84,85 which can cause amino acid damage.86 In addition, heterogeneous acid systems (FeCl3, AlCl3, H2SO4–SiO2 and HClO4–SiO2) were employed for methylation or carboxymethylation but were unsuccessful due to the lack of selectivity towards the product. This is in contrast with previous results for the methylation and carboxymethylation of amines, which provided positive results with these heterogeneous acids.79 The 1H NMR spectra of the DMC reactions of amino acids involving AlCl3, H2SO4–SiO2, and HClO4–SiO2 as acid indicate the presence of unidentified products, whereas FeCl3 results in a significantly low conversion (Fig. S56–S59†).
Furthermore, we attempt to enhance the production of N-methyl alanine methyl ester by adjusting the ratio of DMC to 20, 40, and 80 equiv., respectively (Table 2). When stoichiometric through to 40 equiv. of DMC is used, excellent conversion and yield of more than 99% are observed (ESI†). However, for HClO4 and formic acid systems, increasing DMC loading results in a decrease in conversion. It was discovered that the conversion and yield of the reaction may be affected by the type of acid catalyst and the DMC-to-acid ratio. Both formic acid and HClO4 are monoprotic acids, demonstrating that 1 equiv. acid![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 equiv. DMC allowed significant conversions (>99%). In contrast, H2SO4 is a diprotic acid with a high quantitative yield of 1 equiv. acid
20 equiv. DMC allowed significant conversions (>99%). In contrast, H2SO4 is a diprotic acid with a high quantitative yield of 1 equiv. acid![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 equiv. DMC (Table 2).
40 equiv. DMC (Table 2).
| Entry | Brønsted acid | Product | %Conversion (%selectivity)b (DMC mmol) | ||
|---|---|---|---|---|---|
| 20 | 40 | 80 | |||
| a Conditions: DL-alanine (1 mmol), Brønsted acid (1 mmol), 150 °C, 6 h. b Calculated by 1H NMR and structure confirmed by HMBC. | |||||
| 1 | HClO4 |
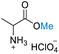
|
>99 (>99) | 79.38 (>99) | 53.70 (>99) |
| 2 | H2SO4 |
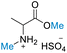
|
59.68 (>99) | >99 (>99) | 73.04 (>99) |
| 3 | HCO2H |

|
>99 (>99) | 92.44 (>99) | 85.59 (>99) |
In addition to the dilution effect, pressure may also influence the reaction's conversion and yield. Increasing the DMC ratio leads to increases in the pressure of the hydrothermal reactor. During the reaction, an increase in pressure decreased the rate of conversion, due to an increase the viscosity of the reaction mixture, thus restricting molecular mobility, both of which inhibit mass transfer.87,88
Table 3 presents the results of reacting DL-alanine, DL-phenylalanine, DL-leucine, and DL-methionine with HClO4 in stoichiometric amounts of 20, 40, and 80 equiv. of DMC in a high-pressure reactor. In the case of DL-alanine, DL-phenylalanine, and DL-leucine, 20 equiv. of DMC with relation to the amino acid displayed more than 99% conversion, >99% selectivity to O-methylation with no evidence of competitive side reactions such as N-methylation. When an excess loading of DMC (20 equiv.) was used, the 1H-NMR spectra revealed a significant decrease in conversion (ESI†). These results may be attributable to the low concentration of amino acids and HClO4. The 1H and 13C NMR spectra revealed the addition of a singlet methoxy proton for ester –CO2Me of amino acids in the range of δH 3.75 to 3.82 ppm and a methoxy carbon in the range of δC 51.70 to 53.45 ppm. In addition, the HMBC spectra also exhibited correlations between the –OMe group and the carbonyl of the amino acid, confirming that the products were amino acid methyl ester derivatives (ESI†). Selective O-methylation of amino acids by nucleophilic substitution on DMC at temperatures up to 150 °C is consistent with previously reported literature.75 Ji and co-workers80 investigated the use of DMC as a methylating agent for two amino acids, including isoleucine and phenylalanine, under basic conditions with a moderate to high yield (83% and 68%, respectively). Their approach demonstrates various limitations, including the need for N-protection with Boc, reaction under an inert atmosphere, long reaction times (18 h), multiple workup steps, chromatographic separation, and the use of an additional solvent (DMSO) despite the use of 20 equiv. of DMC. As such, this current procedure provides a highly efficient single step reaction, which is significantly easier, more convenient and potentially selective than previous methods for producing DL-alanine, DL-phenylalanine, and DL-leucine methyl ester with an excellent yield (>99%), little work up, no column purification, using non-toxic compounds without the use of volatile chlorinated hydrocarbons, and with reduced chemical wastes.29,31–34,37,89,90 Unfortunately, these DMC conditions were not successful for the reaction of DL-methionine. Under our conditions, decomposition of methionine appears to take place as evidenced by a strong smell upon opening the reactor (under ventilation) and by the disappearance of the SMe chemical shift in 1H NMR (Fig. S60†). Furthermore, the preparative-scale reaction was successfully conducted using 12 mmol of DL-alanine, representing a 12-fold increase. This resulted in an impressive 88.21% conversion with a product selectivity exceeding 99%. Thereby demonstrating the potential applicability of this method at a preparative scale.
| Entry | Substrate | Product | %Conversion (%selectivity)b (DMC mmol) | ||
|---|---|---|---|---|---|
| 20 | 40 | 80 | |||
| a Conditions: amino acid (1 mmol), HClO4 (1 mmol), 150 °C, 6 h. b Calculated by 1H NMR and structure confirmed by HMBC. | |||||
| 1 | DL-Alanine |
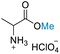
|
>99 (>99) | 79.38 (>99) | 53.70 (>99) |
| 2 | DL-Phenylalanine |
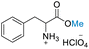
|
>99 (>99) | 80.15 (>99) | 49.26 (>99) |
| 3 | DL-Leucine |
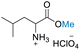
|
>99 (>99) | 87.71 (>99) | 53.70 (>99) |
| 4 | DL-Methionine | Decomposition | — | — | — |
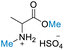
Subsequently, H2SO4 mediated reactions of DMC with different amino acids were investigated. It was discovered that, except for methionine (which decomposed, Fig. S61†),91,92 almost all amino acids were selective for N,O-dimethylation, resulting in good to high conversion and yields of N-methyl amino acid methyl esters (Table 4). Comparing 1H and 13C NMR spectra of the product and substrate demonstrated the presence of one methoxy and one methyl ammonium proton signals within the range of δH 3.60–3.82 ppm, as well as one methoxy and one methyl ammonium carbons within the range of δC 52.90–55.03 ppm, respectively. HMBC demonstrated correlation of one methoxy group to carbonyl carbon and the methine proton (H-2) of amino acids to methylammonium carbon. This confirmed the structure of the product (ESI†). Dimethylation in the reaction may be caused by the dissociation of H2SO4, which may contribute to the formation of two hydrogen ions or protons (H+) in the reaction. Diprotic H2SO4 with 40 equiv. of DMC, a double stoichiometric loading in comparison to HClO4, resulted in excellent conversions and yields. De Marco et al.49 reported the preparation of N-methyl amino acid methyl esters through the N-methylation of N-arylsulfonyl amino acid methyl esters of four amino acids (alanine, valine, isoleucine, and phenylalanine) using diazomethane and trimethyloxonium tetrafluoroborate as methylating agents in the presence of an organic base. When using N-nosyl-protected amino acids, the diazomethane reactions gave quantitative yields, but if using less reactive N-protecting groups, low yields were reported. Although their reactions using both methylating agents produced high-yields at room temperature, they have several limitations, including the need for an N-protecting group, the use of explosive and toxic diazomethane, a toxic solvent (dichloromethane), the need for purifying and drying solvents, the use of flame-dried glassware, and reactions under an inert atmosphere (dry N2), plus multiple workup steps. The current methodology presented in this work offers significant advantages that are superior to those of the previously published methods for preparing N-methyl amino acid methyl esters.
| Entry | Substrate | Product | %Conversion (%selectivity)b (DMC mmol) | ||
|---|---|---|---|---|---|
| 20 | 40 | 80 | |||
| a Conditions: amino acid (1 mmol), H2SO4 (1 mmol), 150 °C, 6h. b Calculated by 1H NMR and structure confirmed by HMBC. | |||||
| 1 | DL-Alanine |
|
59.68 (>99) | >99 (>99) | 73.04 (>99) |
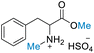
| 2 | DL-Phenylalanine |
|
50.00 (>99) | >99 (>99) | 68.15 (>99) |
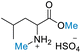
| 3 | DL-Leucine |
|
53.92 (>99) | >99 (>99) | 71.26 (>99) |
| 4 | DL-Methionine | Decomposition | — | — | — |
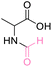
Finally, the optimal conditions for N-formylation with various amino acids were determined. All amino acids were investigated in stoichiometric loading of formic acid and 20 equiv. of DMC which acted as solvent and potentially as a promotor but not as reagent. All reactions produced N-formyl amino acid in considerable yields (>99%) as shown in Table 5.
| Entry | Substrate | Product | %Conversion (%selectivity)b (DMC mmol) | ||
|---|---|---|---|---|---|
| 20 | 40 | 80 | |||
| a Conditions: amino acid (1 mmol), HCO2H (1 mmol), 150 °C, 6h. b Calculated by 1H NMR and structure confirmed by HMBC. | |||||
| 1 | DL-Alanine |
|
>99 (>99) | 92.44 (>99) | 86.59 (>99) |
| 2 | DL-Phenylalanine |
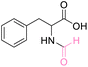
|
>99 (>99) | 88.97 (>99) | 79.29 (>99) |
| 3 | DL-Leucine |
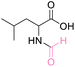
|
>99 (>99) | 92.68 (>99) | 86.34 (>99) |
| 4 | DL-Methionine |
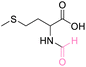
|
>99 (>99) | 87.40 (>99) | 82.45 (>99) |
The NMR spectra of the reactions exhibited singlet proton additions in the range of δH 8.08–8.14 ppm for aldehyde and carbonyl carbon additions in the range of δC 162.03–162.36 ppm for the N-formyl group. The lone pair of the amino groups may delocalize to the aldehyde carbon to form an ammonium ion, resulting in a proton deshielding effect at H-2 of amino acids (δH 4.50 to 4.50 ppm). In addition, the HMBC spectra of all products revealed correlations between H-2 of amino acids and both carboxyl and formyl group as well as the aldehyde proton of formyl group exhibiting a correlation with C-2 of amino acids. Hence, NMR data helps to identify the product as N-formyl amino acid derivatives (ESI†). Due to the stability of methionine in weak acids such as formic acid, there is no decomposition of methionine, like in the case of HClO4 and H2SO4. It is vital to highlight that this method did not result in O-formylation side products. A recent study describes the N-formylation of amino acids using formyloxyacetoxy-phenylmethane (FAPM) as the N-formylating reagent. Six different amino acids were treated for 16 hours with 3 equiv. FAPM (an initial charge of 1.5 equiv. and a second addition of 1.5 equiv. after 8 h), resulting in 71–89% conversion to their corresponding formamides.56 This method has the limitations of requiring the preparation and use of a formylating agent in considerable volumes, a prolonged reaction time, and the necessity for an extraction and purification procedure. Therefore, the current method in this work can be viewed as an efficient approach for the synthesis of N-formylation amino acid esters in a single step reaction, resulting in a higher yield than those previously reported.20,56,57,60–63
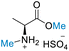
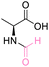
An investigation of a broader substrate scope and assessment of the limitations of the process including both the risk of racemization and epimerization were undertaken. Various functionalised amino acids, along with enantiopure substrates, dipeptides, and tripeptides were investigated using the optimal conditions as outlined in Table 6. In instances where amino acids lacked additional functional groups (L-alanine, L-leucine, L-proline, and glycine), utilizing 20 equivalents of DMC relative to the amino acid in HClO4-assisted conditions resulted in more than 99% conversion, with a selectivity exceeding 99% toward O-methylation (Table 6, entries 1–4). Similarly, amino acids featuring functional groups like L-histidine and L-tyrosine displayed comparable conversions and selectivity under the same conditions (Table 6, entries 5 and 6). However, L-glutamine and L-tryptophan exhibited lower conversions of 77.5% and 50%, respectively, while still maintaining over 99% selectivity to O-methylation (Table 6, entries 7 and 8). The lack of interference from these functional groups in HClO4-assisted conditions might be attributed to the delocalization of their lone pair electrons into aromatic rings. Conversely, amino acids with specific functional groups such as L-lysine, D-serine, and the dipeptide aspartame did not exhibit selectivity for O-methylation under the tested conditions and was attributed to the interference of their free lone pair electrons, resulting in the formation of unselective products (Table 6, entries 9, 10 and 12). Unfortunately, the DMC conditions were not effective for the reaction involving sulfur-containing amino acids like L-cystine and L-glutathione. Additionally, under the specified conditions, the decomposition of L-cystine and L-glutathione were observed, evident from a strong smell upon opening the reactor (under ventilation) (Table 6, entries 11 and 13). In addition, optical rotation experiments have substantiated that the specific rotation of the compounds remained unchanged throughout the successful reactions, as indicated in Table S6.† This observation underscores that our established conditions effectively mitigate the risk of racemization and epimerization during these reactions.
| Entry | Substrate | HClO4 conditionsa | H2SO4 conditionsb | HCO2H conditionsc | |||
|---|---|---|---|---|---|---|---|
| a Conditions: amino acid (1 mmol), HClO4 (1 mmol), DMC (20 mmol), 150 °C, 6 h. b Conditions: amino acid (1 mmol), H2SO4 (1 mmol), DMC (40 mmol), 150 °C, 6 h. c Conditions: amino acid (1 mmol), HCO2H (1 mmol), DMC (20 mmol), 150 °C, 6 h. | |||||||
| 1 | L-Alanine |
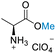
|
|
|
|||
| >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | ||
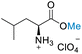
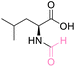
| 2 | L-Leucine |
|

|
|
|||
| >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | ||
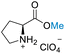
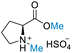
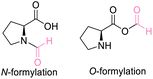
| 3 | L-Proline |
|
|
|
|||
| >99% conversion | >99% selectivity | 65% conversion | >99% selectivity | >99% conversion | 50% selectivity for N-formylation 50% selectivity for O-formylation | ||
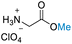
| 4 | Glycine |
|
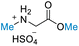
|
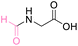
|
|||
| >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | ||
| 5 | L-Histidine |
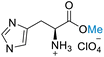
|
Unknown solid |
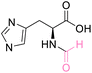
|
|||
| >99% conversion | >99% selectivity | >99% conversion | No selectivity | >99% conversion | >99% selectivity | ||
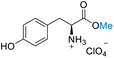
| 6 | L-Tyrosine |
|
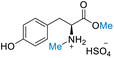
|
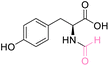
|
|||
| >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | >99% conversion | >99% selectivity | ||
| 7 | L-Glutamine |
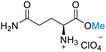
|
Unknown liquid |
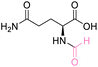
|
|||
| 78% conversion | >99% selectivity | >99% conversion | No selectivity | >99% conversion | 50% selectivity | ||
| 8 | L-Tryptophan |
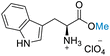
|
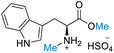
|

|
|||
| 50% conversion | >99% selectivity | 50% conversion | >99% selectivity | >99% conversion | >99% selectivity | ||
| 9 | L-Lysine | >99% conversion | No selectivity | >99% conversion | No selectivity | >99% conversion | No selectivity |
| 10 | D-Serine | >99% conversion | No selectivity | >99% conversion | No selectivity | >99% conversion | No selectivity |
| 11 | L-Cystine | Decomposition | Decomposition | Decomposition | |||
| 12 | Aspartame | >99% conversion | No selectivity | >99% conversion | No selectivity | >99% conversion | No selectivity |
| 13 | L-Glutathione | Decomposition | Decomposition | Decomposition | |||
Reactions in the presence of H2SO4 and formic acid, were consistent with the outcomes of HClO4-assisted conditions (Table 6). Amino acids without additional functional groups (L-alanine, L-leucine, L-proline, and glycine) exhibited remarkable performance. Employing 40 equivalents of DMC relative to the amino acid in H2SO4 or 20 equivalents of DMC relative to the amino acid formic acid conditions yielded conversions surpassing 99%, with selectivity exceeding 99% toward N,O-dimethylation, except for proline, which achieved a 64.9% conversion and >99% selectivity in H2SO4 conditions, or N-formylation, except for proline, which attained >99% conversion and 50% selectivity in formic acid conditions (Table 6, entries 1–4). Similarly, amino acids with functional groups such as L-histidine and L-tyrosine demonstrated comparable conversions and selectivity under the same conditions (Table 6, entries 5 and 6). An exception was noted for L-histidine in H2SO4 conditions, where an unknown solid formed, potentially contributing to the generation of two hydrogen ions (H+) or protons from H2SO4 in the reaction. This solid likely a result of protonation of one nitrogen atom of the imidazole side chain, leading to salt formation (Table 6, entry 5). L-Tryptophan exhibited >99% conversions with >99% selectivity to N-formylation in formic acid conditions, while displaying 50% conversion with >99% selectivity to N,O-dimethylation in H2SO4 conditions (Table 6, entry 8). In the case of L-glutamine, it exhibited low selectivity (50%) for N-formylation in formic acid conditions and was unsuccessful for N,O-dimethylation in H2SO4 conditions (Table 6, entry 7). Other functionalised amino acids, including L-lysine, D-serine, the dipeptide aspartame, and sulfur-containing amino acids (L-cystine and L-glutathione), presented high conversions but were unselective to desired products, these results were comparable to the outcomes under HClO4 conditions (Table 6, entries 9–13). Intriguingly, despite these varied conditions, the prevention of racemization and epimerization risks during the reactions remained consistent (Table S6†).
The comparison of the qualitative and quantitative green metrics93–95 between the current reactions [A (HClO4–DMC, O-methylation), B (H2SO4–DMC, N,O-dimethylation), and C (HCO2H–DMC, N-formylation)] and recent literature methods [A′ (KHCO3–DMC, O-methylation),80 B′ (DIPEA–Me3OBF4, N-methylation),49 and C′ (FAPM, N-formylation)56] (Table 7, see ESI† for details) clearly shows the advantages of the current methods over previous methods except for the high-temperature reaction. The previous processes necessitate workup and/or chromatographic processes that consume a considerable amount of solvent, some of which are hazardous, resulting in a high process mass intensity (PMI), E-factor, solvent intensity (SI), and water intensity (WI). In the current procedures, high conversion and yield, no workup, no chromatographic process, and the use of a green solvent/reagent (DMC) result in significantly improved PMI, E-factor, SI, and WI compared to earlier studies. Although this study demonstrates solvent elimination by evaporation, acid removal would require a minimal additional washing procedure. As such, the methods developed in this current work are preferred for organic chemists and industrial applications.
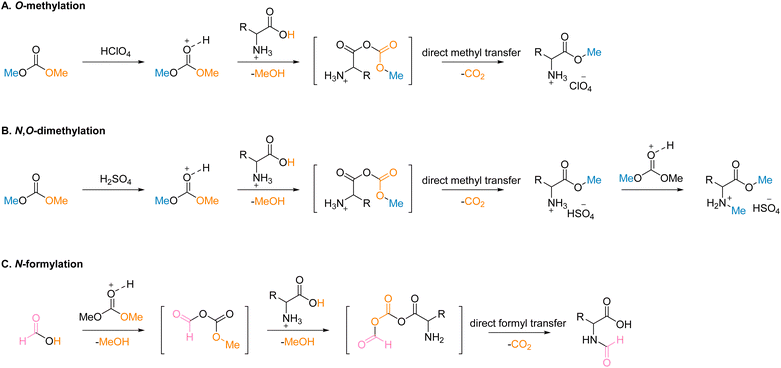
The mechanism of the reactions was proposed to better comprehend the high selectivity observed. The basis of the proposed mechanism for the esterification of a carboxylic acid with DMC was obtained from isotope-labelling experiments.80 For the monomethylation reaction employing monoprotic acid, HClO4 (Scheme 2A), acid-activated DMC reacts with the substance to generate carbonic carboxylic anhydride. The O-methylation product is then formed by the direct transfer of a methyl group from DMC to the carboxylate oxygen.
In the case of diprotic acid, H2SO4 (Scheme 2B), subsequent N-methylation of the O-methylation product from the same pathway in Scheme 2A with another acid-activated DMC to yield the N,O-dimethylation product. DMC is suspected to be involved in the formylation reaction with the use of formic acid. When water was substituted for DMC as the solvent in a chemical reaction, conversion was poor (Fig. S62†).
It is hypothesized that DMC participates catalytically in the formylation reaction. 1H NMR of the reaction mixture containing only formic acid and DMC (1 mmol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 mmol) indicates disappearance of the hydroxyl proton of formic acid (Fig. S63–S65†), due to the formation of a formic acid and DMC species, as presented in Scheme 2A to generate carbonic formic anhydride. To corroborate the formation of this formic acid and DMC species, a 1
20 mmol) indicates disappearance of the hydroxyl proton of formic acid (Fig. S63–S65†), due to the formation of a formic acid and DMC species, as presented in Scheme 2A to generate carbonic formic anhydride. To corroborate the formation of this formic acid and DMC species, a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio of formic acid and DMC was used, and the 1H NMR reveals the presence of methanol in the reaction mixture, confirming that DMC reforms (Fig. S66†). The substrate then reacts with one side of carbonic formic anhydride, followed by formyl group transfer to the nitrogen of the substrate to yield an N-formylation product (Scheme 2C). DMC serves as the linker to transfer the methyl or formyl groups to the substrate, consequently enhancing the selectivity of the product.
1 ratio of formic acid and DMC was used, and the 1H NMR reveals the presence of methanol in the reaction mixture, confirming that DMC reforms (Fig. S66†). The substrate then reacts with one side of carbonic formic anhydride, followed by formyl group transfer to the nitrogen of the substrate to yield an N-formylation product (Scheme 2C). DMC serves as the linker to transfer the methyl or formyl groups to the substrate, consequently enhancing the selectivity of the product.
Conclusions
A novel, efficient, and eco-friendly strategy for the synthesis of amino acid derivatives using an acid assisted approach with dimethyl carbonate as a green methylating agent has been investigated. The application of simple acid systems, including perchloric acid, sulfuric acid, and formic acid demonstrated significant promise for selective modification of amino acids. The reactions achieved excellent yields and high selectivity for the synthesis of amino acid methyl esters, N-methylated amino acid methyl esters, and N-formylated amino acids. This approach offers several advantages over traditional methods, including reduced toxicity, lower cost, reduced chemical waste, fewer synthetic steps, excellent yields of the desired products in high purity, and mitigate the risk of racemization and epimerization. Furthermore, this efficient single step reaction used a sustainable solvent, readily available acid reagents/reactants, simple reaction setup, and the products were easily isolated from the solvent by removal under reduced pressure, however, removing acid would necessitate a minimally additional washing procedure. Overall, the results demonstrate the importance of acid selection and reaction optimization for achieving high yields and selectivity in the synthesis of amino acid derivatives. These findings will contribute to the development of sustainable and efficient synthesis of amino acid derivatives, important building blocks for the synthesis of biologically active compounds.Experimental
Materials
DL-Alanine >99%, DL-phenylalanine >99%, DL-leucine >99%, DL-methionine >99%, chloroform-d (CDCl3, >99.8% D), methanol-d4 (CD3OD, >99.68%), dimethylsulfoxide dimethyl carbonate (DMC) >99% and anhydrous sodium sulphate >99% were purchased from Sigma-Aldrich. Formic acid (AR grade) 98%, acetic acid (AR grade) >99.8%, sulphuric acid 98% and perchloric acid 70% were purchased from Qrec. Ethyl acetate (AR grade) >99.8%, methanol (AR grade) >99.8%, ethanol (AR grade) >99.8% and 4 Å molecular sieves (4–8 mesh) were purchased from RCI labscan.General experimental procedure for reaction of amino acids with dimethyl carbonate at 90 °C
In a 50 mL round-bottom flask, dimethyl carbonate DMC (20, 40, or 80 mmol) and stoichiometric acid catalysts (1.00 mmol) such as 1.685 mL for 20 mmol DMC and 88, 54, and 39 μL for 1 mmol of HClO4, H2SO4, and formic acid, respectively, were added (H2SO4, HClO4, HCl, HNO3, and HCO2H). 1 mmol of each amino acid (DL-alanine, DL-phenylalanine, DL-leucine, and DL-methionine, represented by 89.1 mg of alanine) was then added to the solution mixture 30 minutes later. 24 hours were used for heating the solution at 90 °C under reflux. Once the reaction was complete, the mixture was cooled to room temperature. With reduced pressure, the solution solvent was removed to provide the crude product for NMR analysis. 1H and 13C-NMR were used to compute conversion (%) and selectivity (%).General experimental procedure for reaction of amino acids with dimethyl carbonate at 150 °C
In a 25 mL hydrothermal reactor, dimethyl carbonate DMC (20, 40, or 80 mmol) and stoichiometric acid catalysts (1.00 mmol) such as 1.685 mL for 20 mmol DMC and 88, 54, and 39 μL for 1 mmol of HClO4, H2SO4, and formic acid, respectively, were added (H2SO4, HClO4, HCl, HNO3, and HCO2H). 1 mmol of each amino acid (DL-alanine, DL-phenylalanine, DL-leucine, and DL-methionine, represented by 89.1 mg of alanine) was then added to the solution mixture 30 minutes later. The solution was heated in an oven at 150 °C for 6 h. After the reaction was completed, the reaction was allowed to cool in the oven until 50 °C. With reduced pressure, the solution solvent was removed to provide the crude product for NMR analysis. 1H and 13C-NMR were used to compute conversion (%) and selectivity (%).1H and 13CNMR analysis
The structural was determined by based on the analysis 1H and 13CNMR spectra, which were recorded in CDCl3, and CD3OD as solvents on a Bruker 400 spectrometer using the residue of those solvents as the internal standards (16 scans for 1H NMR analysis, and 256 scans for 13C NMR analysis). The NMR signals were reported in terms of chemical shift as delta values are in pm, and J values are in Hz. The HMBC technique was used confirm structures.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This project is funded by National Research Council of Thailand (NRCT) (grant number: N42A650240). This work received financial supported from Materials Chemistry Research Center, Department of Chemistry, Faculty of Science, Khon Kaen University. The financial support of the Center of Excellence for Innovation in Chemistry (PERCH-CIC), Ministry of Higher Education, Science, Research, and Innovation is gratefully acknowledged. Dr Chadamas Sakonsinsiri, Khon Kaen University, is personally thanked for her assistance in providing amino acids samples.References
- N. Naowarojna, R. Cheng, J. Lopez, C. Wong, L. Qiao and P. Liu, Synth. Syst. Biotechnol., 2021, 6, 32–49 CrossRef CAS PubMed.
- J. Han, H. Konno, T. Sato, V. A. Soloshonok and K. Izawa, Eur. J. Med. Chem., 2021, 220, 113448 CrossRef CAS PubMed.
- J. Han, H. Konno, T. Sato, K. Izawa and V. A. Soloshonok, Curr. Org. Chem., 2021, 25, 1627–1658 CrossRef CAS.
- G. B. Santos, A. Ganesan and F. S. Emery, ChemMedChem, 2016, 11, 2245–2251 CrossRef CAS PubMed.
- X. Luo, H. Chen, Y. Song, Z. Qin, L. Xu, N. He, Y. Tan and W. Dessie, Eur. J. Pharm. Sci., 2023, 181, 106363 CrossRef CAS PubMed.
- M. S. Kang, T. W. S. Kong, J. Y. X. Khoo and T.-P. Loh, Chem. Sci., 2021, 12, 13613–13647 RSC.
- K. Sharma, K. K. Sharma, A. Sharma and R. Jain, Drug Discovery Today, 2023, 28, 103464 CrossRef CAS PubMed.
- W. M. Long, B. H. Chua, N. Lautensack and H. E. Morgan, Am. J. Physiol., 1983, 245, C101–C112 CrossRef CAS PubMed.
- M. L. Di Gioia, A. Leggio, A. Le Pera, A. Liguori, A. Napoli, C. Siciliano and G. Sindona, J. Org. Chem., 2003, 68, 7416–7421 CrossRef CAS PubMed.
- H. Fu, B. Han and Y.-F. Zhao, Chem. Commun., 2003, 134–135 RSC.
- A. I. Vicente, J. M. Caio, J. Sardinha, C. Moiteiro, R. Delgado and V. Félix, Tetrahedron, 2012, 68, 670–680 CrossRef CAS.
- T. Zieliński, M. Achmatowicz and J. Jurczak, Tetrahedron: Asymmetry, 2002, 13, 2053–2059 CrossRef.
- J.-K. Kim, J. Kim, S. Song, O.-S. Jung and H. Suh, J. Inclusion Phenom. Macrocyclic Chem., 2007, 58, 187–192 CrossRef CAS.
- C. Somlai, A. Péter, P. Forgó and B. Penke, Synth. Commun., 2003, 33, 1815–1820 CrossRef CAS.
- G. P. Pollini, N. Baricordi, S. Benetti, C. De Risi and V. Zanirato, Tetrahedron Lett., 2005, 46, 3699–3701 CrossRef CAS.
- L. Eberhardt, D. Armspach, D. Matt, L. Toupet and B. Oswald, Eur. J. Inorg. Chem., 2007, 2007, 4153–4161 CrossRef.
- H. Bruns, L. Ziesche, N. K. Taniwal, L. Wolter, T. Brinkhoff, J. Herrmann, R. Müller and S. Schulz, Beilstein J. Org. Chem., 2018, 14, 2964–2973 CrossRef CAS PubMed.
- J. C. Sheehan and D.-D. H. Yang, J. Am. Chem. Soc., 1958, 80, 1154–1158 CrossRef CAS.
- F. Boulay, M. Tardif, L. Brouchon and P. Vignais, Biochem. Biophys. Res. Commun., 1990, 168, 1103–1109 CrossRef CAS PubMed.
- S. Kotha, M. Behera and P. Khedkar, Tetrahedron Lett., 2004, 45, 7589–7590 CrossRef CAS.
- S. Anantharaj and M. Jayakannan, Biomacromolecules, 2012, 13, 2446–2455 CrossRef CAS PubMed.
- A. Nagai, T. Miyagawa, H. Kudo and T. Endo, Macromolecules, 2003, 36, 9335–9339 CrossRef CAS.
- F. Sanda and T. Endo, Macromol. Chem. Phys., 1999, 200, 2651–2661 CrossRef CAS.
- A. Sharma, A. Kumar, S. A. H. Abdel Monaim, Y. E. Jad, A. El-Faham, B. G. de la Torre and F. Albericio, Biopolymers, 2018, 109, e23110 CrossRef PubMed.
- S. M. A. Rauf, P. I. Arvidsson, F. Albericio, T. Govender, G. E. M. Maguire, H. G. Kruger and B. Honarparvar, Org. Biomol. Chem., 2015, 13, 9993–10006 RSC.
- J. Chatterjee, F. Rechenmacher and H. Kessler, Angew. Chem., Int. Ed., 2013, 52, 254–269 CrossRef CAS PubMed.
- J. Chatterjee, C. Gilon, A. Hoffman and H. Kessler, Acc. Chem. Res., 2008, 41, 1331–1342 CrossRef CAS PubMed.
- A. V. Malkov, K. Vranková, M. Černý and P. Kočovský, J. Org. Chem., 2009, 74, 8425–8427 CrossRef CAS PubMed.
- J. Li and Y. Sha, Molecules, 2008, 13, 1111–1119 CrossRef CAS PubMed.
- P. G. Andersson, D. Guijarro and D. Tanner, J. Org. Chem., 1997, 62, 7364–7375 CrossRef CAS PubMed.
- L. Molnár-Perl, M. Pintér-Szakács and V. Fábián-Vonsik, J. Chromatogr. A, 1987, 390, 434–438 CrossRef PubMed.
- S. M. Jones, J. E. Urch, M. Kaiser, R. Brun, J. L. Harwood, C. Berry and I. H. Gilbert, J. Med. Chem., 2005, 48, 5932–5941 CrossRef CAS PubMed.
- S. McGhie, Synth. Commun., 2002, 32, 1275–1278 CrossRef CAS.
- D. J. Wardrop and A. Basak, Org. Lett., 2001, 3, 1053–1056 CrossRef CAS PubMed.
- J. R. Rachele, J. Org. Chem., 1963, 28, 2898–2898 CrossRef CAS.
- R. C. Anand and Vimal, Synth. Commun., 1998, 28, 1963–1965 CrossRef CAS.
- H. H. Weinstock Jr. and E. L. May Jr., J. Am. Chem. Soc., 1940, 62, 3266–3266 CrossRef.
- J. M. Humphrey and A. R. Chamberlin, Chem. Rev., 1997, 97, 2243–2266 CrossRef CAS PubMed.
- P. Wipf, Chem. Rev., 1995, 95, 2115–2134 CrossRef CAS.
- N. Fusetani and S. Matsunaga, Chem. Rev., 1993, 93, 1793–1806 CrossRef CAS.
- R. M. Wenger, Helv. Chim. Acta, 1984, 67, 502–525 CrossRef CAS.
- R. M. Wenger, Angew. Chem., Int. Ed. Engl., 1985, 24, 77–85 CrossRef.
- G. R. Pettit, Y. Kamano, C. L. Herald, C. Dufresne, R. B. Bates, J. M. Schmidt, R. L. Cerny and H. Kizu, J. Org. Chem., 1990, 55, 2989–2990 CrossRef CAS.
- G. R. Pettit, Y. Kamano, C. L. Herald, Y. Fujii, H. Kizu, M. R. Boyd, F. E. Boettner, D. L. Doubek, J. M. Schmidt, J.-C. Chapuis and C. Michel, Tetrahedron, 1993, 49, 9151–9170 CrossRef CAS.
- R. B. Bates, K. G. Brusoe, J. J. Burns, S. Caldera, W. Cui, S. Gangwar, M. R. Gramme, K. J. McClure, G. P. Rouen, H. Schadow, C. C. Stessman, S. R. Taylor, V. H. Vu, G. V. Yarick, J. Zhang, G. R. Pettit and R. Bontems, J. Am. Chem. Soc., 1997, 119, 2111–2113 CrossRef CAS.
- P. Jouin, J. Poncet, M. N. Dufour, A. Pantaloni and B. Castro, J. Org. Chem., 1989, 54, 617–627 CrossRef CAS.
- J. M. Ramanjulu, X. Ding, M. M. Joullié and W.-R. Li, J. Org. Chem., 1997, 62, 4961–4969 CrossRef CAS.
- J. R. Coggins and N. L. Benoiton, Can. J. Chem., 1971, 49, 1968–1971 CrossRef CAS.
- R. De Marco, M. L. Di Gioia, A. Liguori, F. Perri, C. Siciliano and M. Spinella, Tetrahedron, 2011, 67, 9708–9714 CrossRef CAS.
- A. Leggio, A. Liguori, F. Perri, C. Siciliano and M. C. Viscomi, Chem. Biol. Drug Des., 2009, 73, 287–291 CrossRef CAS PubMed.
- B. E. Mileson, L. M. Sweeney, M. L. Gargas and J. Kinzell, Inhalation Toxicol., 2009, 21, 583–605 CrossRef CAS PubMed.
- C. A. Valdez, E. P. Salazar and R. N. Leif, Sci. Rep., 2022, 12, 1401 CrossRef CAS PubMed.
- C. A. Valdez, R. N. Leif and A. Alcaraz, Anal. Chim. Acta, 2016, 933, 134–143 CrossRef CAS PubMed.
- G. Lamoureux and C. Agüero, ARKIVOC, 2009, 2009, 251–264 Search PubMed.
- S. Joseph, P. Das, B. Srivastava, H. Nizar and M. Prasad, Tetrahedron Lett., 2013, 54, 929–931 CrossRef CAS.
- R. S. L. Chapman, R. Lawrence, J. M. J. Williams and S. D. Bull, Org. Lett., 2017, 19, 4908–4911 CrossRef CAS PubMed.
- I. Muramatsu, M. Murakami, T. Yoneda and A. Hagitani, Bull. Chem. Soc. Jpn., 1965, 38, 244–246 CrossRef CAS PubMed.
- T. Meinnel, L. Patiny, S. Ragusa and S. Blanquet, Biochemistry, 1999, 38, 4287–4295 CrossRef CAS PubMed.
- T. Chancellor and C. Morton, Synthesis, 1994, 1023–1025 CrossRef CAS.
- N. Akikusa, K. Mitsui, T. Sakamoto and Y. Kikugawa, Synthesis, 1992, 1058–1060 CrossRef CAS.
- T. Giard, D. Bénard and J.-C. Plaquevent, Synthesis, 1998, 297–300 CrossRef CAS.
- W. Duczek, J. Deutsch, S. Vieth and H.-J. Niclas, Synthesis, 1996, 37–38 CrossRef CAS.
- B. A. Aleiwi, K. Mitachi and M. Kurosu, Tetrahedron Lett., 2013, 54, 2077–2081 CrossRef CAS PubMed.
- A. M. Ruder, Ann. N. Y. Acad. Sci., 2006, 1076, 207–227 CrossRef CAS PubMed.
- R. E. Jackson, Technol. Cult., 2004, 45, 55–79 CrossRef.
- A.-A. G. Shaikh and S. Sivaram, Chem. Rev., 1996, 96, 951–976 CrossRef CAS PubMed.
- E. Foy, J. B. Farrell and C. L. Higginbotham, J. Appl. Polym. Sci., 2009, 111, 217–227 CrossRef CAS.
- G. Fiorani, A. Perosa and M. Selva, Green Chem., 2018, 20, 288–322 RSC.
- D. Delledonne, F. Rivetti and U. Romano, Appl. Catal., A, 2001, 221, 241–251 CrossRef CAS.
- P. Tundo, Pure Appl. Chem., 2001, 73, 1117–1124 CAS.
- C. Peschel, F. Horsthemke, M. Leißing, S. Wiemers-Meyer, J. Henschel, M. Winter and S. Nowak, Batteries Supercaps, 2020, 3, 1183–1192 CrossRef CAS.
- M. A. Pacheco and C. L. Marshall, Energy Fuels, 1997, 11, 2–29 CrossRef CAS.
- B. Schäffner, F. Schäffner, S. P. Verevkin and A. Börner, Chem. Rev., 2010, 110, 4554–4581 CrossRef PubMed.
- P. Tundo, M. Selva and S. Memoli, in Green Chemical Syntheses and Processes, American Chemical Society, 2000, vol. 767, pp. 87–99 Search PubMed.
- P. Tundo and M. Selva, Acc. Chem. Res., 2002, 35, 706–716 CrossRef CAS PubMed.
- S. Huang, B. Yan, S. Wang and X. Ma, Chem. Soc. Rev., 2015, 44, 3079–3116 RSC.
- Yashentech Corporation, US Pat., US2017057904A1, 2017 Search PubMed.
- M. Bilde, T. E. Møgelberg, J. Sehested, O. J. Nielsen, T. J. Wallington, M. D. Hurley, S. M. Japar, M. Dill, V. L. Orkin, T. J. Buckley, R. E. Huie and M. J. Kurylo, J. Phys. Chem. A, 1997, 101, 3514–3525 CrossRef CAS.
- P. Worsawat, P. Noppawan, C. Croise, N. Supanchaiyamat, C. R. McElroy and A. J. Hunt, Org. Biomol. Chem., 2023, 21, 1070–1081 RSC.
- Y. Ji, J. Sweeney, J. Zoglio and D. J. Gorin, J. Org. Chem., 2013, 78, 11606–11611 CrossRef CAS PubMed.
- J. P. Amend and H. C. Helgeson, Pure Appl. Chem., 1997, 69, 935–942 CrossRef CAS.
- P. Tundo, M. Musolino and F. Aricò, Green Chem., 2018, 20, 28–85 RSC.
- Z.-Y. Tian, Z. Zhang, S. Wang and H. Lu, Nat. Commun., 2021, 12, 5810 CrossRef CAS PubMed.
- G. D. Robertson Jr., D. M. Mason and W. H. Corcoran, J. Phys. Chem., 1955, 59, 683–690 CrossRef.
- H. S. Johnston, L. Foering, Y.-S. Tao and G. H. Messerly, J. Am. Chem. Soc., 1951, 73, 2319–2321 CrossRef CAS.
- K. Kikugawa, T. Kato and Y. Okamoto, Free Radicals Biol. Med., 1994, 16, 373–382 CrossRef CAS PubMed.
- S. P′eng, R. H. Sapiro, R. P. Linstead and D. M. Newitt, J. Chem. Soc., 1938, 784–791 RSC.
- P. Prasertpong, C. Jaroenkhasemmeesuk, J. R. Regalbuto, J. Lipp and N. Tippayawong, Energy Rep., 2020, 6, 1–9 CrossRef.
- B. D. White, J. Mallen, K. A. Arnold, F. R. Fronczek, R. D. Gandour, L. M. B. Gehrig and G. W. Gokel, J. Org. Chem., 1989, 54, 937–947 CrossRef CAS.
- Y. H. Jung and J. D. Kim, Arch. Pharmacal Res., 2000, 23, 574–578 CrossRef CAS PubMed.
- T. F. Lavine and N. F. Floyd, J. Biol. Chem., 1954, 207, 97–105 CrossRef CAS PubMed.
- L. W. Butz and V. du Vigneaud, J. Biol. Chem., 1932, 99, 135–142 CrossRef CAS.
- D. Prat, A. Wells, J. Hayler, H. Sneddon, C. R. McElroy, S. Abou-Shehada and P. J. Dunn, Green Chem., 2015, 18, 288–296 RSC.
- R. A. Sheldon, ACS Sustainable Chem. Eng., 2018, 6, 32–48 CrossRef CAS.
- S. Michałek, L. Gurba-Bryśkiewicz, W. Maruszak, M. Zagozda, A. M. Maj, Z. Ochal, K. Dubiel and M. Wieczorek, RSC Adv., 2022, 12, 33605–33611 RSC.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d3gc03455k |
| This journal is © The Royal Society of Chemistry 2024 |