Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Highly cytotoxic Cu(II) terpyridine complexes as chemotherapeutic agents†
Kaixin
Ni‡
,
Nicolás
Montesdeoca‡
and
Johannes
Karges
 *
*
Faculty of Chemistry and Biochemistry, Ruhr-University Bochum, Universitätsstrasse 150, 44780 Bochum, Germany. E-mail: johannes.karges@ruhr-uni-bochum.de; Web: https://www.kargesgroup.ruhr-uni-bochum.de Tel: +49 2343224187
First published on 19th April 2024
Abstract
Cancer is considered as the biggest medicinal challenge worldwide. During a typical treatment, the tumorous tissue is removed in a surgical procedure and the patient further treated by chemotherapy. One of the most frequently applied drugs are platinum complexes. Despite their clinical success, these compounds are associated with severe side effects and low therapeutic efficiency. To overcome these limitations, herein, the synthesis and biological evaluation of Cu(II) terpyridine complexes as chemotherapeutic drug candidates is suggested. The compounds were found to be highly cytotoxic in the nanomolar range against various cancer cell lines. Mechanistic insights revealed that the compounds primarily accumulated in the cytoplasm and generated reactive oxygen species in this organelle, triggering cell death by apoptosis. Based on their high therapeutic effect, these metal complexes could serve as a starting point for further drug development.
Introduction
The last decades, cancer remains the leading cause of death worldwide.1 Statistics from the World Health Organization indicated that nearly 10 million people died from cancer in 2020, and it is predicted that the number of new cancer diagnoses per year is expected to rise to 29.5 million and the number of cancer-related deaths to 16.4 million by 2040.2 To date, platinum complexes (i.e., cisplatin, oxaliplatin, carboplatin) are among the most commonly used chemotherapeutic agents worldwide.3 Despite their frequent clinical use, these compounds are associated with severe side effect such as nerve and kidney damage, nausea, vomiting, or bone marrow suppression.4 Insights into the mechanism of action revealed that the metal complexes coordinate to the nucleobases of the DNA, triggering cell death of the cancer cells.5 However, due to various biological barriers only a small fraction of the metal complex reaches its target. A clinical study has reported that approximately 95% of the administered dose binds to human serum albumin and is therefore therapeutically deactivated.6 Further analyses indicated that less than 1% of the administered doses accumulated in the tumor and less than 0.01% binds to the nuclear DNA, causing the desired therapeutic effect.7 These results strongly suggest that the clinically used platinum complexes are associated with a poor therapeutic efficiency and that there is a need for the development of new compounds with an improved therapeutic efficiency.Copper complexes as anticancer agents have received increasing attraction due their high bioavailability and biocompatibility.8 The ability of copper to change its redox state between copper(I) and copper(II) under physiological conditions allows for electron transfer reactions inside the cancer cells and ultimately the generation of reactive oxygen species.9 Reactive oxygen species are able to induce oxidative stress inside the cancer cells, causing damage to biomolecules such as lipids, proteins, and DNA and therefore triggering cell death.9b,10 Capitalizing on this, considerable research effort has been focused on developing copper-containing compounds with anticancer properties.8–11 Several copper(II) polypyridine complexes have demonstrated to localize in the nucleus of cancerous cells and interact with DNA. In this localization, these compounds are able to generate reactive oxygen species, causing oxidative cleavage of the phosphodiester bond of the DNA and ultimately triggering cell death.12 Our own group has recently reported on terminally functionalized copper(II) terpyridine complexes with a high cytotoxic effect against drug resistant colon or lung cancer cells.11b,13 Recently variously functionalized copper(II) 4′-phenyl-2,2′:6′,2′′-terpyridine complexes have been reported as highly active anticancer agents that interact through a multimodal mechanism including cell cycle arrest, mitochondria misfunctioning and cell apoptosis. Promisingly, the compounds demonstrated a strong tumor growth inhibition effect in a tumor-bearing mouse model.14 Based on the generally promising properties of copper(II) complexes, the Casiopeinas® complexes with the molecular formula [Cu(NN)(OO)]NO3 or [Cu(NN)(NO)]NO3 (NN = 2,2′-bipyridines or 1,10-phenanthrolines, OO = acetylacetonate or salicylaldehyde, NO = amino acid or peptide) are currently studied in clinical trials in Mexico.15 These specific metal complexes were found with a high cytotoxic effect against various cancer cell lines, inducing cell death through apoptosis by catalytic redox cycling.16
Based on these promising findings, herein, a structure–activity relationship study of differently methoxy functionalized copper(II) terpyridine complexes including Cu(4′-(4′-ortho-methoxy-phenyl)-2,2′:6′,2′′-terpyridine)(Cl)2 (Cu1), Cu(4′-(4′-meta-methoxy-phenyl)-2,2′:6′,2′′-terpyridine)(Cl)2 (Cu2), and Cu(4′-(4′-para-methoxy-phenyl)-2,2′:6′,2′′-terpyridine)(Cl)2 (Cu3) was performed. The compounds exhibited a high cytotoxicity at nanomolar concentrations against diverse cancer cell lines. Mechanistic investigations revealed that these compounds predominantly accumulated in the cytoplasm, inducing the generation of reactive oxygen species within this organelle. The reactive oxygen species production led to cell death through apoptosis. Based on their high therapeutic effects, these metal complexes could be considered as a promising starting point for subsequent drug development.
Results and discussion
The terpyridine ligands 4′-(4′-ortho-methoxy-phenyl)-2,2′:6′,2′′-terpyridine,17 4′-(4′-meta-methoxy-phenyl)-2,2′:6′,2′′-terpyridine,18 and 4′-(4′-para-methoxy-phenyl)-2,2′:6′,2′′-terpyridine were synthesized according to the previous published reports.19 The differently methoxy functionalized benzaldehydes, 2-acetylpyridine, and potassium hydroxide were dissolved in ethanol and stirred at room temperature for several hours. After this time, ammonia was added and the mixture stirred at room temperature overnight to generate the tridentate terpyridine ligand scaffold.19c The desired 4′-(4′-substituted-phenyl)-2,2′:6′,2′′-terpyridine copper(II) complexes Cu(4′-(4′-ortho-methoxy-phenyl)-2,2′:6′,2′′-terpyridine)(Cl)2 (Cu1), Cu(4′-(4′-meta-methoxy-phenyl)-2,2′:6′,2′′-terpyridine)(Cl)2 (Cu2), and Cu(4′-(4′-para-methoxy-phenyl)-2,2′:6′,2′′-terpyridine)(Cl)2 (Cu3) (Fig. 1) were synthesized upon stirring of equimolar amounts of the terpyridine ligand and copper(II) chloride in a mixture of dry methanol and dichloromethane at room temperature overnight. The generated precipitate was collected, washed, and recrystallized from methanol. The identity of the compound was verified by high resolution electrospray ionization mass spectroscopy (Fig. S1†) and the purity was confirmed by elemental analysis.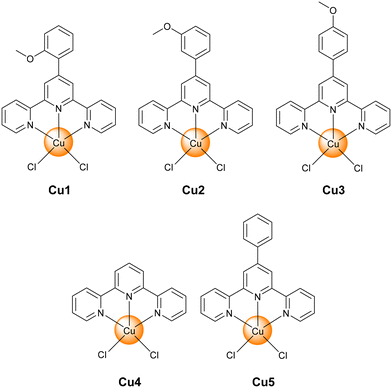 | ||
Fig. 1 Chemical structures of Cu(II) terpyridine complexes Cu1–Cu3 investigated in this study. The metal complexes Cu4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 11b and Cu5 11b and Cu5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 13 have been previously reported and are used here for comparison with previous studies. 13 have been previously reported and are used here for comparison with previous studies. | ||
The cytotoxicity of the metal complexes Cu1–Cu3 against cancerous human breast adenocarcinoma (MCF-7), colon adenocarcinoma (HT-29), mouse colorectal carcinoma (CT-26) as well as and non-cancerous human fibroblasts (GM-5657) cells was assessed using a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. Promisingly, all compounds exhibited a highly cytotoxicity in the low micromolar down to nanomolar range (IC50 = 236–3620 nM) against all cell lines. Importantly, the organic ligands 4′-(4′-ortho-methoxy-phenyl)-2,2′:6′,2′′-terpyridine, 4′-(4′-meta-methoxy-phenyl)-2,2′:6′,2′′-terpyridine, and 4′-(4′-para-methoxy-phenyl)-2,2′:6′,2′′-terpyridine were found to be non-toxic (>20 μM) in all tested cell lines. The metal complexes were not able to differentiate between cancerous and non-cancerous cells. Cu1 was 63-times more cytotoxic against MCF-7 cells, Cu3 was 29-times more cytotoxic against MCF-7 cells, and Cu2 was 116-times more cytotoxic against CT-26 cells than cisplatin (overview: Table 1, drug response curves: Fig. S2–S4†). These findings suggest that the cytotoxic effect of the metal complex is dependent on the position of the functionalization of the methoxy substituent. The observed cytotoxic effect was in the same range as for previously reported copper(II) terpyridine complexes against (cisplatin-resistant) cervix adenocarcinoma cells.12a The herein reported Cu(II) complexes were found to be significantly more cytotoxic than the previously reported metal complexes Cu4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 11b,d and Cu5.13 Within the herein studied series of compounds, Cu2 was found with the highest cytotoxicity against MCF-7 (0.687 ± 0.125 μM), HT-29 (0.537 ± 0.117 μM), or CT-26 (0.236 ± 0.059 μM) cells. Based on the strongest therapeutic effect, further experiments were performed with Cu2 against CT-26 cells. Besides the treatment for 48 h, the cytotoxicity of the metal complex was further studied upon treatment for 4 h. With this incubation time, Cu2 was found to be cytotoxic in the very low micromolar range against CT-26 (1.04 ± 0.08 μM) cells.
11b,d and Cu5.13 Within the herein studied series of compounds, Cu2 was found with the highest cytotoxicity against MCF-7 (0.687 ± 0.125 μM), HT-29 (0.537 ± 0.117 μM), or CT-26 (0.236 ± 0.059 μM) cells. Based on the strongest therapeutic effect, further experiments were performed with Cu2 against CT-26 cells. Besides the treatment for 48 h, the cytotoxicity of the metal complex was further studied upon treatment for 4 h. With this incubation time, Cu2 was found to be cytotoxic in the very low micromolar range against CT-26 (1.04 ± 0.08 μM) cells.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 11b,d and Cu5
11b,d and Cu5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 13 have been previously reported and are used here for comparison with previous studies
13 have been previously reported and are used here for comparison with previous studies
| MCF-7 | HT-29 | CT-26 | GM-5657 | |
|---|---|---|---|---|
| Cu1 | 0.542 ± 0.136 | 0.964 ± 0.214 | 0.630 ± 0.113 | 0.841 ± 0.127 |
| Cu2 | 0.687 ± 0.125 | 0.537 ± 0.117 | 0.236 ± 0.059 | 0.740 ± 0.089 |
| Cu3 | 1.159 ± 0.206 | 3.620 ± 0.472 | 1.247 ± 0.358 | 2.761 ± 0.492 |
| Terpy-ortho-OMe | >20 | >20 | >20 | >20 |
| Terpy-meta-OMe | >20 | >20 | >20 | >20 |
| Terpy-para-OMe | >20 | >20 | >20 | >20 |
| Cisplatin | 34.5 ± 1.2 | 54.3 ± 8.6 | 27.5 ± 2.4 | 42.7 ± 9.2 |
| Cu4 | 3.2 ± 0.6 | 2.7 ± 1.1 | 2.3 ± 0.7 | 3.4 ± 0.5 |
| Cu5 | 1.1 ± 0.3 | 1.5 ± 0.6 | 1.3 ± 0.4 | 2.1 ± 0.4 |
To understand the difference in cytotoxicity of the metal complexes, the cellular uptake of Cu1–Cu3 upon incubation for 4 h was investigated. The results showed no statistically significant differences (Fig. S5†), ruling out cellular uptake efficiency as the primary reason. Based on its superior therapeutic effect, further studies were performed with Cu2.
The stability of a compound is essential for its use in a biological setting. The stability of Cu2 upon incubation in Dulbecco's modified Eagle's cell media was studied upon monitoring of the absorption profile over a period of 48 h (Fig. S6†). No significant changes were observed, indicative of the stability of the metal complex under physiological conditions.
Previous studies have indicated that copper(II) terpyridine complexes could undergo redox cycling within a biological setting.20 The ability of Cu2 to interact with the cellular reducing agents glutathione or ascorbate was studied upon constant monitoring of the absorption spectra of the metal complex in an aqueous solution. As no significant changes of the absorption spectrum were observed upon titration of glutathione (Fig. S7†), the change in oxidation state of the metal complex was ruled out. Contrary, the titration of ascorbate to Cu2 showed drastic change in the intensity of the peak centred at approximately 270 nm (Fig. S8†), indicative of the reduction of the Cu(II) centre to Cu(I).
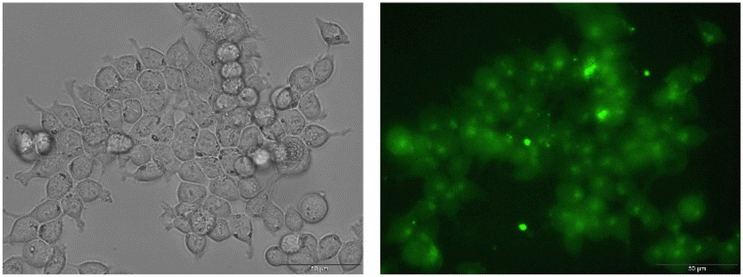
Previous studies have indicated that copper(II) terpyridine complexes could generate reactive oxygen species.20 The ability of the metal complex to generate reactive oxygen species in CT-26 cells was studied by fluorescence microscopy using the reactive oxygen species specific dye 2′,7′-dichlorodihydrofluorescein diacetate. This dye is non-emissive in human cells but gets readily oxidized in the presence of reactive oxygen species, generating a strongly green fluorescent signal. As expected, no fluorescence signals were observed in the cancerous cell incubated with 2′,7′-dichlorodihydrofluorescein diacetate (5 μM) for 30 min (Fig. S7†). The cancer cells were treated with Cu2 (1 μM) for 4 h and incubated with 2′,7′-dichlorodihydrofluorescein diacetate (5 μM) for 30 min. The cancer cells treated with the metal complex showed a strong green emission, indicative of generation of reactive oxygen species inside the cancer cells (Fig. 2).
For an understanding of the observed cytotoxicity, the biodistribution of Cu2 was determined. The cancer cells were incubated with the metal complex (10 μM) for 4 h. After this time, the cell organelles were separately extracted using a specific nucleus extraction kit, a mitochondria extraction kit, a lysosome extraction kit, and a cytoplasm extraction kit, and digested in concentrated nitric acid. The metal content in each organelle was determined by inductively coupled plasma optical emission spectroscopy and the naturally occurring copper content subtracted. While small amounts of Cu2 were found in the nucleus, mitochondria, and the lysosomes, the metal complex primarily accumulated in the cytoplasm of the cancer cells (Fig. 3). Notably, small quantities of a compound inside a specific cell organelle could cause a significant therapeutic effect and therefore cannot be neglected in the assessment of the biological mechanism of action.
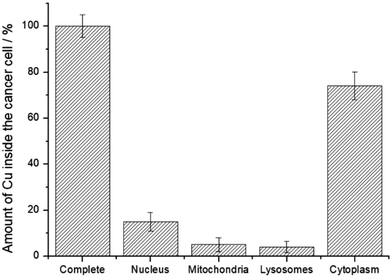 | ||
| Fig. 3 Subcellular localization of Cu2 (10 μM) upon incubation in CT-26 cells for 4 h and determination of the metal content by inductively coupled plasma optical emission spectroscopy. | ||
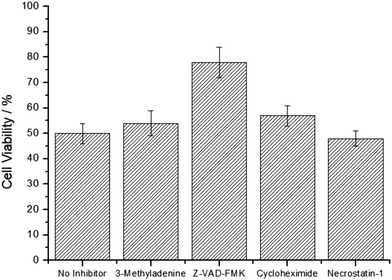
For an understanding of the therapeutic effect, the cell death mechanism of the metal complex was studied upon pre-incubation of the CT-26 cells with specific cell death inhibitors 3-methyladenine (autophagy inhibitor), Z-VAD-FMK (apoptosis inhibitor), cycloheximide (paraptosis inhibitor), and necrostatin-1 (necrosis inhibitor), and afterwards treated with Cu2. As the pre-incubation with 3-methyladenine, cycloheximide, and necrostatin-1 did not significantly influence the cell survival, these cell death mechanisms were ruled out. Contrary, the incubation with Z-VAD-FMK enhanced the survival of the cells (Fig. 4), indicative that the cell death mechanism of Cu2 is primarily driven by apoptosis.
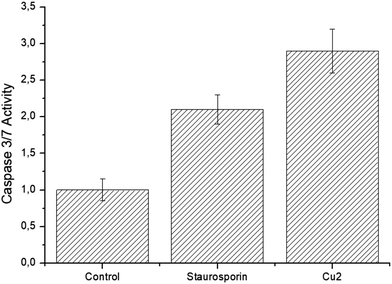
For a deeper insight into the cell death mechanism, the dependency of the cell death of Cu2 on the caspase 3/7 pathway was assessed using a caspase 3/7 glo assay. These caspases are well characterized executes of apoptotic cell death. Staurosporine was used as a well-known positive control substance. Upon treatment of the cancerous cells with Cu2, the caspase 3/7 activity was strongly elevated (Fig. 5). These findings suggest that the metal complex triggers cell death by apoptosis using the caspase 3/7 pathway.
Conclusions
In summary, this study reports on the chemical synthesis and biological evaluation of differently methoxy functionalized copper(II) terpyridine complexes as potential chemotherapeutic agents for anticancer therapy. The compounds were synthesized upon coordination of copper(II) ions with a tridentate terpyridine ligand and characterized by high resolution electrospray ionization mass spectroscopy and elemental analysis. The biological evaluation against human breast adenocarcinoma, colon adenocarcinoma, and mouse colorectal carcinoma cells demonstrated a high cytotoxic effect in the nanomolar range. Mechanistic insights of the lead compound revealed that the metal complex primarily accumulated in the cytoplasm. Within this organelle, the copper(II) complex efficiently generated reactive oxygen species, that induced cell death by apoptosis. Based on their high cytotoxic effect, these metal complexes could represent a starting point for further drug development. Previous studies have indicated that anticancer agents that specifically localize in a cell organelle are associated with an enhanced therapeutic effect. Future work could focus on chemically modifying the lead compound with cell organelle targeting moieties. The reported metal complexes were unable to differentiate between cancerous and healthy cells. Future studies will focus on the chemical modification of the structural scaffold to generate a conjugate with tumor selectivity.Conflicts of interest
The author declares no competing interests.Acknowledgements
J. K. gratefully acknowledges the financial support with the Liebig fellowship from the Chemical Industry Fund of the German Chemical Industry Association and the Life Sciences Bridge Award from the Aventis Foundation.References
- (a) F. Bray, A. Jemal, N. Grey, J. Ferlay and D. Forman, Lancet Oncol., 2012, 13, 790–801 CrossRef PubMed; (b) H. Zhou, D. Tang, Y. Yu, L. Zhang, B. Wang, J. Karges and H. Xiao, Nat. Commun., 2023, 14, 5350 CrossRef CAS PubMed.
- H. Sung, J. Ferlay, R. L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal and F. Bray, CA Cancer J. Clin., 2021, 71, 209–249 CrossRef PubMed.
- (a) S. Dilruba and G. V. Kalayda, Cancer Chemother. Pharmacol., 2016, 77, 1103–1124 CrossRef CAS PubMed; (b) L. Kelland, Nat. Rev. Cancer, 2007, 7, 573–584 CrossRef CAS PubMed; (c) G. Kim, H. L. Tan, R. Sundar, B. Lieske, C. E. Chee, J. Ho, A. Shabbir, M. V. Babak, W. H. Ang, B. C. Goh, W. P. Yong, L. Wang and J. B. Y. So, Clin. Cancer Res., 2021, 27, 1875–1881 CrossRef CAS PubMed; (d) L. Zhang, N. Montesdeoca, J. Karges and H. Xiao, Angew. Chem., Int. Ed., 2023, 62, e202300662 CrossRef CAS PubMed; (e) M. S. Galanski, M. A. Jakupec and B. K. Keppler, Curr. Med. Chem., 2005, 12, 2075–2094 CrossRef CAS PubMed; (f) H. Wang, Y. Lai, D. Li, J. Karges, P. Zhang and H. Huang, J. Med. Chem., 2024, 67, 1336–1346 CrossRef CAS PubMed.
- (a) D. Wang and S. J. Lippard, Nat. Rev. Drug Discovery, 2005, 4, 307–320 CrossRef CAS PubMed; (b) A. Eskandari, A. Kundu, S. Ghosh and K. Suntharalingam, Angew. Chem., 2019, 131, 12187–12192 CrossRef; (c) X. Gao, G. Lei, B. Wang, Z. Deng, J. Karges, H. Xiao and D. Tan, Adv. Sci., 2023, 10, 2205241 CrossRef CAS PubMed; (d) M. J. R. Tham, M. V. Babak and W. H. Ang, Angew. Chem., Int. Ed., 2020, 59, 19070–19078 CrossRef CAS PubMed; (e) C. A. Rabik and M. E. Dolan, Cancer Treat. Rev., 2007, 33, 9–23 CrossRef CAS PubMed; (f) M. Shahlaei, S. M. Asl, A. Derakhshani, L. Kurek, J. Karges, R. Macgregor, M. Saeidifar, I. Kostova and A. A. Saboury, J. Mol. Struct., 2024, 1301, 137366 CrossRef CAS.
- (a) T. C. Johnstone, K. Suntharalingam and S. J. Lippard, Chem. Rev., 2016, 116, 3436–3486 CrossRef CAS PubMed; (b) X. Wang, X. Wang, S. Jin, N. Muhammad and Z. Guo, Chem. Rev., 2019, 119, 1138–1192 CrossRef CAS PubMed; (c) M. V. Babak, Y. Zhi, B. Czarny, T. B. Toh, L. Hooi, E. K.-H. Chow, W. H. Ang, D. Gibson and G. Pastorin, Angew. Chem., Int. Ed., 2019, 58, 8109–8114 CrossRef CAS PubMed; (d) G. Liang, T. Sadhukhan, S. Banerjee, D. Tang, H. Zhang, M. Cui, N. Montesdeoca, J. Karges and H. Xiao, Angew. Chem., Int. Ed., 2023, 62, e202301074 CrossRef CAS PubMed; (e) Y.-R. Zheng, K. Suntharalingam, T. C. Johnstone, H. Yoo, W. Lin, J. G. Brooks and S. J. Lippard, J. Am. Chem. Soc., 2014, 136, 8790–8798 CrossRef CAS PubMed; (f) C. Schmidt, T. Babu, H. Kostrhunova, A. Timm, U. Basu, I. Ott, V. Gandin, V. Brabec and D. Gibson, J. Med. Chem., 2021, 64, 11364–11378 CrossRef CAS PubMed; (g) D. Wei, Y. Huang, B. Wang, L. Ma, J. Karges and H. Xiao, Angew. Chem., Int. Ed., 2022, 61, e202201486 CrossRef CAS PubMed; (h) X. Wang, X. Wang and Z. Guo, Acc. Chem. Res., 2015, 48, 2622–2631 CrossRef CAS PubMed; (i) J. Karges, ChemNanoMat, 2023, 9, e202300295 CrossRef CAS.
- A. I. Ivanov, J. Christodoulou, J. A. Parkinson, K. J. Barnham, A. Tucker, J. Woodrow and P. J. Sadler, J. Biol. Chem., 1998, 273, 14721–14730 CrossRef CAS PubMed.
- A. Yimit, O. Adebali, A. Sancar and Y. Jiang, Nat. Commun., 2019, 10 Search PubMed.
- (a) T. J. P. McGivern, S. Afsharpour and C. J. Marmion, Inorg. Chim. Acta, 2018, 472, 12–39 CrossRef CAS; (b) C. Marzano, M. Pellei, F. Tisato and C. Santini, Anti-Cancer Agents Med. Chem., 2009, 9, 185–211 CrossRef CAS PubMed; (c) L. Ruiz-Azuara and M. E. Bravo-Gómez, Curr. Med. Chem., 2010, 17, 3606–3615 CrossRef CAS PubMed; (d) K. D. Mjos and C. Orvig, Chem. Rev., 2014, 114, 4540–4563 CrossRef CAS PubMed; (e) F. Tisato, C. Marzano, M. Porchia, M. Pellei and C. Santini, Med. Res. Rev., 2010, 30, 708–749 CrossRef CAS PubMed; (f) S. Tardito and L. Marchiò, Curr. Med. Chem., 2009, 16, 1325–1348 CrossRef CAS PubMed; (g) N. K. Singh, A. A. Kumbhar, Y. R. Pokharel and P. N. Yadav, J. Inorg. Biochem., 2020, 210, 111134 CrossRef CAS PubMed; (h) C. Wende, C. Lüdtke and N. Kulak, Eur. J. Inorg. Chem., 2014, 2014, 2597–2612 CrossRef CAS.
- (a) C. Marzano, M. Pellei, F. Tisato and C. Santini, Anti-Cancer Agents Med. Chem., 2009, 9, 185–211 CrossRef CAS PubMed; (b) C. Santini, M. Pellei, V. Gandin, M. Porchia, F. Tisato and C. Marzano, Chem. Rev., 2014, 114, 815–862 CrossRef CAS PubMed.
- (a) C. Marzano, M. Pellei, F. Tisato and C. Santini, Anti-Cancer Agents Med. Chem., 2009, 9, 185–211 CrossRef CAS PubMed; (b) S. Zehra, S. Tabassum and F. Arjmand, Drug Discovery Today, 2021, 26, 1086–1096 CrossRef CAS PubMed; (c) B. D'Autréaux and M. B. Toledano, Nat. Rev. Mol. Cell Biol., 2007, 8, 813–824 CrossRef PubMed; (d) J. P. Fruehauf and F. L. Meyskens Jr., Clin. Cancer Res., 2007, 13, 789–794 CrossRef CAS PubMed; (e) J. Wang and J. Yi, Cancer Biol. Ther., 2008, 7, 1875–1884 CrossRef CAS PubMed; (f) E. C. Cheung and K. H. Vousden, Nat. Rev. Cancer, 2022, 22, 280–297 CrossRef CAS PubMed.
- (a) P. Ji, P. Wang, H. Chen, Y. Xu, J. Ge, Z. Tian and Z. Yan, Pharmaceuticals, 2023, 16, 234 CrossRef CAS PubMed; (b) J. Karges, K. Xiong, O. Blacque, H. Chao and G. Gasser, Inorg. Chim. Acta, 2021, 516, 120137 CrossRef CAS; (c) X.-L. Wang, H. Chao, H. Li, X.-L. Hong, L.-N. Ji and X.-Y. Li, J. Inorg. Biochem., 2004, 98, 423–429 CrossRef CAS PubMed; (d) J. Grau, A. Caubet, O. Roubeau, D. Montpeyó, J. Lorenzo and P. Gamez, ChemBioChem, 2020, 21, 2348–2355 CrossRef CAS PubMed; (e) A. Kumar, J. P. Chinta, A. K. Ajay, M. K. Bhat and C. P. Rao, Dalton Trans., 2011, 40, 10865–10872 RSC; (f) J. Rani and S. Roy, ChemMedChem, 2023, 18, e202200652 CrossRef PubMed; (g) S. Xu, D. Liu, T. Chang, X. Wen, S. Ma, G. Sun, L. Wang, S. Chen, Y. Xu and H. Zhang, Front. Genet., 2022, 13, 938259 CrossRef CAS PubMed; (h) K. Greish, V. Pittalà, S. Taurin, S. Taha, F. Bahman, A. Mathur, A. Jasim, F. Mohammed, I. M. El-Deeb, S. Fredericks and F. Rashid-Doubell, Nanomaterials, 2018, 8 Search PubMed; (i) Y. Hu, Y. Qian, J. Wei, T. Jin, X. Kong, H. Cao and K. Ding, Front. Pharmacol., 2021, 12, 752825 CrossRef CAS PubMed; (j) H. Li, J. Wang, C. Wu, L. Wang, Z. S. Chen and W. Cui, Drug Discovery Today, 2020, 25, 1099–1108 CrossRef CAS PubMed; (k) W. Wang, J. Cassidy, V. O'Brien, K. M. Ryan and E. Collie-Duguid, Cancer Res., 2004, 64, 8167–8176 CrossRef CAS PubMed; (l) B. Xu, P. Shi, I. S. Fombon, Y. Zhang, F. Huang, W. Wang and S. Zhou, Blood Cells, Mol., Dis., 2011, 47, 264–269 CrossRef CAS PubMed; (m) W. J. Guo, S. S. Ye, N. Cao, J. Huang, J. Gao and Q. Y. Chen, Exp. Toxicol. Pathol., 2010, 62, 577–582 CrossRef CAS PubMed; (n) J. Liu, Y. Yuan, Y. Cheng, D. Fu, Z. Chen, Y. Wang, L. Zhang, C. Yao, L. Shi, M. Li, C. Zhou, M. Zou, G. Wang, L. Wang and Z. Wang, J. Am. Chem. Soc., 2022, 144, 4799–4809 CrossRef CAS PubMed.
- (a) H. Nakamura and K. Takada, Cancer Sci., 2021, 112, 3945–3952 CrossRef CAS PubMed; (b) A. De Vizcaya-Ruiz, A. Rivero-Müller, L. Ruiz-Ramirez, J. A. Howarth and M. Dobrota, Toxicology, 2003, 194, 103–113 CrossRef CAS PubMed.
- K. Xiong, Y. Zhou, J. Karges, K. Du, J. Shen, M. Lin, F. Wei, J. Kou, Y. Chen, L. Ji and H. Chao, ACS Appl. Mater. Interfaces, 2021, 13, 38959–38968 CrossRef CAS PubMed.
- Y. Yang, C.-F. Chen, F.-F. Guo, Y.-Q. Gu, H. Liang and Z.-F. Chen, J. Inorg. Biochem., 2023, 246, 112284 CrossRef CAS PubMed.
- (a) R. Galindo-Murillo, J. C. García-Ramos, L. Ruiz-Azuara, T. E. Cheatham 3rd and F. Cortés-Guzmán, Nucleic Acids Res., 2015, 43, 5364–5376 CrossRef CAS PubMed; (b) Y. Figueroa-DePaz, J. Pérez-Villanueva, O. Soria-Arteche, D. Martínez-Otero, V. Gómez-Vidales, L. Ortiz-Frade and L. Ruiz-Azuara, Molecules, 2022, 27, 3504 CrossRef CAS PubMed.
- (a) F. Carvallo-Chaigneau, C. Trejo-Solís, C. Gómez-Ruiz, E. Rodríguez-Aguilera, L. Macías-Rosales, E. Cortés-Barberena, C. Cedillo-Peláez, I. Gracia-Mora, L. Ruiz-Azuara, V. Madrid-Marina and F. Constantino-Casas, BioMetals, 2008, 21, 17–28 CrossRef CAS PubMed; (b) C. Trejo-Solís, G. Palencia, S. Zuñiga, A. Rodríguez-Ropon, L. Osorio-Rico, S. Torres Luvia, I. Gracia-Mora, L. Marquez-Rosado, A. Sánchez, M. E. Moreno-García, A. Cruz, M. E. Bravo-Gómez, L. Ruiz-Ramírez, S. Rodríquez-Enriquez and J. Sotelo, Neoplasia, 2005, 7, 563–574 CrossRef PubMed; (c) R. Galindo-Murillo, J. C. García-Ramos, L. Ruiz-Azuara, T. E. Cheatham III and F. Cortés-Guzmán, Nucleic Acids Res., 2015, 43, 5364–5376 CrossRef CAS PubMed.
- Q.-P. Qin, T. Meng, M.-X. Tan, Y.-C. Liu, X.-J. Luo, B.-Q. Zou and H. Liang, Eur. J. Med. Chem., 2018, 143, 1387–1395 CrossRef CAS PubMed.
- J.-W. Liang, Y. Wang, K.-J. Du, G.-Y. Li, R.-L. Guan, L.-N. Ji and H. Chao, J. Inorg. Biochem., 2014, 141, 17–27 CrossRef CAS PubMed.
- (a) Q. P. Qin, T. Meng, M. X. Tan, Y. C. Liu, X. J. Luo, B. Q. Zou and H. Liang, Eur. J. Med. Chem., 2018, 143, 1387–1395 CrossRef CAS PubMed; (b) J. Li, H. Yan, Z. Wang, R. Liu, B. Luo, D. Yang, H. Chen, L. Pan and Z. J. D. T. Ma, Dalton Trans., 2021, 50, 8243–8257 RSC; (c) B. N. Mongal and S. Naskar, J. Coord. Chem., 2017, 70, 451–462 CrossRef CAS.
- (a) Z. Ma, L. Wei, E. C. B. A. Alegria, L. M. D. R. S. Martins, M. F. C. Guedes de Silva and A. J. L. Pombeiro, Dalton Trans., 2014, 43, 4048–4058 RSC; (b) V. Uma, M. Elango and B. U. Nair, Eur. J. Inorg. Chem., 2007, 2007, 3484–3490 CrossRef.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4dt00759j |
| ‡ The authors have contributed equally to this study. |
| This journal is © The Royal Society of Chemistry 2024 |