 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Hydrazide-based reactive matrices for the sensitive detection of aldehydes and ketones by MALDI mass spectrometry imaging†
Henrik
Lodén‡
 a,
Luke S.
Schembri‡
a,
Luke S.
Schembri‡
 ab,
Anna
Nilsson
a,
Ibrahim
Kaya
a,
Reza
Shariatgorji
a,
Luke R.
Odell
ab,
Anna
Nilsson
a,
Ibrahim
Kaya
a,
Reza
Shariatgorji
a,
Luke R.
Odell
 *b and
Per E.
Andrén
*b and
Per E.
Andrén
 *a
*a
aDepartment of Pharmaceutical Biosciences, Spatial Mass Spectrometry, Science for Life Laboratory, Uppsala University, Uppsala, Sweden. E-mail: per.andren@uu.se
bDepartment of Medicinal Chemistry, Uppsala University, Uppsala, Sweden. E-mail: luke.odell@ilk.uu.se
First published on 8th August 2024
Abstract
A one-step, on-tissue chemical derivatisation method for MALDI mass spectrometry imaging was found to improve the detectability of aldehydes and ketones by charge-tagging. The developed reactive matrices, containing a UV-chromophore, ionisable moiety and hydrazide group, showed an equal or higher detection efficiency than Girard's reagent P, enabling improved imaging of brain metabolites without the need for additional co-matrices.
Carbonyl-containing compounds, notably aldehydes and ketones, represent a diverse group of endogenous biomolecules. They include fatty aldehydes and ketones, neurotransmitter metabolites and neurosteroids, which play pivotal roles in a wide array of biological functions, ranging from energy generation and metabolism to cellular signalling and the maintenance of homeostasis.1–3 Their involvement in lipid biosynthesis, neurotransmitter synthesis and antioxidant defence underscore their integral functions in sustaining cellular vitality and specialised physiological processes.4
The possibility of visualising carbonyl-containing compounds within the brain or other organs using imaging techniques has gained significant traction due to the intrinsic importance of these compounds.5–11 However, simultaneous imaging of numerous carbonyl-containing compounds in tissues can be challenging, primarily due to their diversity, which encompasses a wide range of molecules with varying structures, properties and concentrations.7,11,12 Therefore, development of an imaging method capable of providing precise spatial visualisation of multiple carbonyl compounds may enable deeper insights into their functional roles. Such an approach could be extended to practical applications for the detection, diagnosis and monitoring of disorders associated with disrupted carbonyl-related pathways. Additionally, it may allow the identification of biomarkers and examination of pharmaceutical interventions, further underscoring its practical significance.
Matrix-assisted laser desorption/ionisation mass spectrometry imaging (MALDI-MSI)13,14 enables in situ imaging of numerous analytes in tissue sections while preserving their spatial distribution. The technique requires minimal sample preparation and generates images (or heat maps) depicting the relative abundance of analytes within tissue sections by selecting individual peaks from mass spectra acquired from each spot in the raster ablated by the laser.
The properties of the matrix and optimisation of its application are crucial to obtain high sensitivity and spatial resolution. For use in MALDI-MSI, matrices must meet specific criteria, including an ability to absorb energy from the laser and form fine, uniform crystals when applied to a tissue section to maintain lateral resolution. Commonly used matrices often possess aromatic structures to satisfy these requirements. Furthermore, ionisation enhancement can be achieved by incorporating proton donating or accepting moieties (e.g. carboxylic acid or amine) into the matrix structure. The prevention of analyte cluster formation is also essential to minimise signal interference and facilitate mass spectra interpretation.15,16
The detection of aldehydes and ketones within tissue sections poses challenges due to their low abundance and low ionisation efficiencies, primarily because they lack inherent ionisable groups. Additionally, signal interference from both the matrix and other macromolecules, such as lipids, leads to reduced sensitivity. While methodological improvements, such as advancements in matrix deposition techniques, tissue washing and laser post-ionisation (MALDI-2),17–19 can enhance the detection of low-abundance and low-ionisation-efficiency compounds, more recent strategies have involved the use of permanently charged reactive matrices in MALDI-MSI. Such matrices serve the dual functions of chemically derivatising analytes (by reacting selectively with specific chemical functionalities, such as amines) and as light-absorbing matrices.20,21 Chemical derivatisation with this type of matrix increases the sensitivity by introducing a permanent charge in the final analyte-matrix adduct, thereby increasing the ionisation while simultaneously shifting the mass out of the low mass region. Hence, signal overlap, often encountered with standard matrices, is avoided.
Previously, we developed pyrylium-based matrices to covalently tag primary amine-bearing neurotransmitters and their metabolites.21 Further work on fluoromethylpyridinium-based (FMP) matrices allowed the tagging of primary and secondary amines, as well as phenolic hydroxyls, enabling the labelling and imaging of neurotransmitters and metabolites with high sensitivity and spatial resolution.20 An alternative two-step approach for covalent charge-tagging of carboxylic acids and aldehydes has recently been published,8 where 1-(4-(aminomethyl)phenyl)pyridin-1-ium chloride (AMPP) was applied to rat brain tissue sections together with triethylamine in a first step, followed by application of a norharmane matrix. Despite the utility of the aforementioned matrices, a reliable one-step application reactive matrix has not yet been developed for the detection of metabolites containing carbonyl groups in brain samples. Previous research has shown that derivatising reagents, such as Girard's reagents P and T,22 can be used for the detection of carbonyls. However, carbonyls have only been detected in tissues where they are abundant, such as adrenal glands and testes,10,23 and an additional co-matrix was required.
In the present study, we designed and synthesised several one-step application reactive matrices aimed at enhancing the ionisation efficiencies and signal intensities to enable the detection of endogenous carbonyl group-containing compounds for precise mapping of their locations (see ESI,† Experimental and Fig. S1–S9). We demonstrated the capabilities of the developed method in brain and lung tissue samples from animal models for the visualisation of endogenous carbonyls, such as fatty aldehydes and progesterone, as well as corticosteroid drugs, including budesonide and fluticasone propionate. The implications of this research extend to drug development applications, drug distribution studies24 and comparisons between regular and pathophysiological contexts.25–27
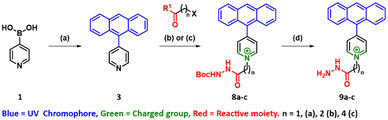
In our search for a tri-functional reactive MALDI matrix, we strategically designed candidate matrices 9a–c to encompass three essential functional domains (Fig. 1). These domains included a highly reactive yet selective group for facilitating covalent analyte capture, a permanently charged group to enhance mass sensitivity and a conjugated chromophore domain with high absorptivity at typical laser wavelengths to promote efficient laser desorption (Fig. 1). This approach has previously proven successful in the development of matrices based on FMP and pyrylium reactive groups, effectively overcoming limitations associated with current state-of-the-art matrices and derivatising agents.20 The target compounds were assembled by an initial Suzuki–Miyaura reaction, followed by pyridine alkylation and Boc-deprotection (9a) or an alkylation/hydrazide formation/deprotection sequence (9b, c). The structural difference between 9a–c lay in their spacer length (i.e. one carbon for 9a, and two and four carbons for 9b and 9c, respectively).
To evaluate the sensitivity enhancement of our novel reactive matrices compared to the well-established Girard's reagent P (Gir P), we conducted a comparative study. We selected three model compounds, i.e. budesonide, fluticasone propionate and progesterone (ESI,† Fig. S10), due to their steroid core structures and differing carbonyl functional groups (ketone, enone and dienone). These compounds served as a benchmark for assessing the sensitivity enhancement using different reactive matrices.
Our results demonstrated that all the reactive matrices (9a–c, Fig. 1) provided improved or equivalent signal intensities for the model compounds when applied to rat brain control tissue sections compared to Gir P in the presence of the co-matrix DHB (ESI,† Fig. S11). Unfortunately, a direct comparison with Gir P without the addition of co-matrix (DHB) was not feasible as Gir P alone yielded insufficient signal intensity. Nevertheless, the sensitivity obtained with reactive matrices 9a–c, even without co-matrix, was comparable to limits achieved with Gir P plus DHB (ESI,† Table S1).
As shown in ESI,† Table S1, the derivatisation efficiencies of 9a–c differed between the model compounds, with the highest sensitivities for budesonide, fluticasone propionate and progesterone obtained using matrices 9c, 9a and 9a, respectively. Following addition of DHB, the highest sensitivities for budesonide and fluticasone propionate were obtained using matrices 9c and 9b, respectively, while Gir P and 9a were superior for enhancing the detection of singly derivatised progesterone (I). The highest sensitivity of doubly derivatised progesterone (II) with co-matrix was obtained with matrix 9b. The performance of each matrix is a combination of its innate reactivity towards each substrate and the desorption efficiency of the derivatised analyte. These factors are expected to be inversely correlated for 9a–c, as spacer length increases the nucleophilicity of the hydrazide motif while also increasing flexibility and size, leading to lower desorption efficiency. In the latter case, the addition of DHB leads to a decrease in the limit of detection (LOD), and this effect is most pronounced for 9b and 9c. Interestingly, 9a performs exceptionally well as a reactive matrix for fluticasone propionate and progesterone, while it is the least efficient for budesonide. This is consistent with the lower electrophilicity of the carbonyl groups in budesonide and the lower nucleophilicity of 9a. Based on its overall performance, the 2-carbon spacer containing matrix 9b was selected for further evaluation.
A proof-of-principle quantification of budesonide in rat lung tissue following administration of 0.25 mg budesonide per kg bodyweight was conducted using reactive matrix 9b (Fig. 2). We employed the in-house-developed software msIQuant28 for the conversion of intensity data from spotted budesonide standards (Fig. 2A and B). After constructing a standard curve (Fig. 2C), the amounts of budesonide in selected regions of the dosed tissue (Fig. 2D and E) were calculated to be 59–87 pmol of budesonide per mg of tissue (Fig. 2F), equivalent to 25.3–37.6 mg per kg bodyweight, locally.
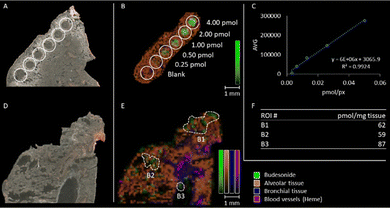 | ||
| Fig. 2 Quantification of budesonide in dosed rat lung tissue using reactive matrix 9b together with DHB as co-matrix. Spatial resolution: 120 μm (small laser focus) with 100 shots per raster position. RMS-normalised data. (A) Scanned image of budesonide standard spots (0.25–4.00 pmol, highlighted by white circles) on rat lung tissue. (B) Ion distribution image of budesonide (m/z 754.3856 [M + 9b-H2O]+). The intensity was set to visualise all spots. (C) Standard curve (R2 = 0.9924) obtained from the data using the msIQuant software28 calculated with a lung tissue density of 0.5 g cm−3.29 (D) Scanned image of tissue in vivo dosed with 0.25 mg per kg budesonide. (E) Green: ion distribution image of budesonide (m/z 754.3856 [M + 9b-H2O]+) on dosed tissue. Regions of interest (ROIs) selected for quantification (B1–B3) are highlighted with dashed lines. Orange: alveolar tissue marker at m/z 754.5363 [M + H]+. Blue: bronchial tissue marker at m/z 804.5525 [M + H]+. Purple: haem at m/z 616.1770 [M + H]+. (F) Calculated amounts of budesonide in the selected regions (B1–B3). For additional information, see the experimental section in the ESI.† | ||
Given the lower abundance of carbonyl-containing hormones and metabolites in brain tissue, we initially explored the derivatisation capabilities of the reactive matrices on rat adrenal cortex tissue (ESI,† Fig. S12). Similar to our findings with spotted standards on brain tissue (ESI,† Fig. S11), the tested reactive matrices (9a–c) exhibited improved or equivalent signal intensities for identified endocrine gland hormones and metabolites compared to Gir P in the presence of DHB. Again, a direct comparison with Gir P alone was not feasible due to insufficient ionisation, but the sensitivity obtained with reactive matrices 9a–9c without co-matrix was equivalent to that achieved with Gir P and DHB as co-matrix. Subsequently, we explored the use of reactive matrix 9b (without co-matrix) for imaging mouse brain tissue and demonstrated that DOPAL (3,4-dihydroxyphenylacetaldehyde) and several fatty aldehydes (decanals) could be successfully imaged (Fig. 3A). The same matrix (i.e.9b) also enabled the successful imaging of endogenous aldehydes and monosaccharides, such as hexose and octadecanal, in coronal rat brain sections (Fig. 3B). Compound identification was conducted either by comparison with the known distribution, distribution obtained using AMPP derivatisation reagent8 together with norharmane or by MSMS-fragmentation (ESI,† Table S2, Fig. S13).
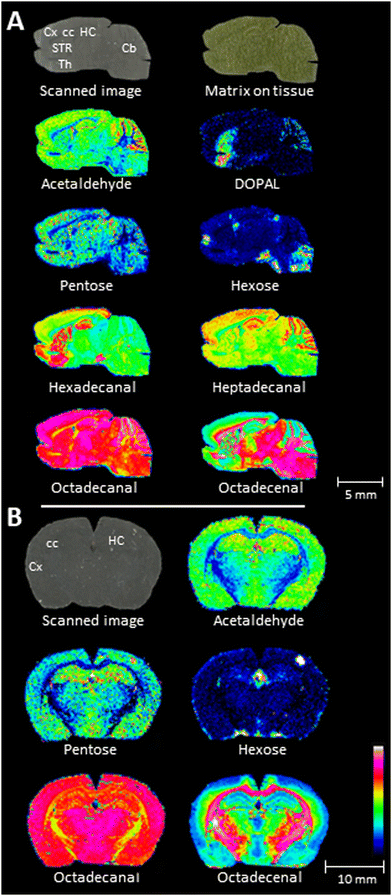 | ||
| Fig. 3 Imaging of endogenous aldehydes and monosaccharides using reactive matrix 9b (without co-matrix). (A) MSI of endogenous compounds, including DOPAL (3,4-dihydroxyphenylacetaldehyde) and fatty aldehydes, in mouse brain. Spatial resolution: 120 μm. (B) MSI of endogenous compounds in rat brain using the same matrix. Spatial resolution: 150 μm. 100 shots were collected per raster position (small laser focus). RMS-normalised data. The intensity was set individually to reveal the best possible distribution of the compounds. Cx: cortex; cc: corpus callosum; STR: striatum; HC: hippocampus. For additional information, see the experimental section in the ESI.† | ||
Finally, we explored the capabilities of reactive matrix 9b for the high-resolution imaging of the relatively abundant endogenous aldehyde heptadecanal in comparison with Gir P on coronal rat brain sections (ESI,† Fig. S14). In the presence of DHB as co-matrix, both reactive matrix 9b and Gir P produced identical distributions of heptadecanal (ESI,† Fig. S14). The matrix 9b alone also yielded the same distribution of heptadecanal. Notably, application of Gir P resulted in the formation of tissue holes, distorting the spatial information, which were not observed with matrix 9b (ESI,† Fig. S14). It is noteworthy that the same solvent and concentrations were used for both matrices which should have similar acidity (pKa). Hence, the tissue holes were likely explained by structural differences between Gir P and 9b.
In summary, our primary objective was to develop a single-step reactive matrix to enhance the selectivity for endogenous compounds containing carbonyl functional groups. MALDI-MSI of endogenous compounds was successfully demonstrated on adrenal cortex tissue. While brain tissue presented additional challenges, successful derivatisation and subsequent imaging of several endogenous compounds, such as DOPAL, was achieved using matrix 9b. Furthermore, the feasibility of using matrix 9b for high-resolution MALDI-MSI was confirmed with no apparent delocalisation effects at either 40 μm or 20 μm spatial resolution, highlighting its superiority compared to the standard carbonyl derivatising agent Gir P.
The authors acknowledge the Swedish Research Council (2021-03293 and 2022-04198 to P. E. A., 2018-05133 and 2022-04831 to L. R. O.), the Swedish Brain Foundation (FO2021-0318 and FO2023-0241 to P. E. A., FO2021-0234 and FO2023-0221 to L. R. O.) and the Science for Life Laboratory (to P. E. A.). Patrik Bjärterot (MSc) is acknowledged for valuable help with data handling. Current address Luke S. Schembri: Division of Biomolecular Sciences and Medicinal Chemistry, School of Pharmacy, University of Nottingham Biodiscovery Institute, Nottingham NG7 2RD, UK.
Data availability
The data supporting this article have been included as part of the ESI.† Mass spectrometry imaging raw data are available upon request.Conflicts of interest
There are no conflicts to declare.Notes and references
- E. E. Alexeev, J. M. Lanis, D. J. Kao, E. L. Campbell, C. J. Kelly, K. D. Battista, M. E. Gerich, B. R. Jenkins, S. T. Walk, D. J. Kominsky and S. P. Colgan, Am. J. Pathol., 2018, 188, 1183–1194 CrossRef CAS PubMed.
- D. L. Ebenezer, P. Fu, R. Ramchandran, A. W. Ha, V. Putherickal, T. Sudhadevi, A. Harijith, F. Schumacher, B. Kleuser and V. Natarajan, Biochim. Biophys. Acta Mol. Cell. Biol. Lipids, 2020, 1865, 158681 CrossRef CAS PubMed.
- M. Jové, I. Pradas, M. Dominguez-Gonzalez, I. Ferrer and R. Pamplona, Redox Biol., 2019, 23, 101082 CrossRef PubMed.
- F. di Michele, S. Luchetti, G. Bernardi, E. Romeo and P. Longone, Front. Neuroendocrinol., 2013, 34, 132–142 CrossRef CAS PubMed.
- D. F. Cobice, D. E. Livingstone, C. L. Mackay, R. J. A. Goodwin, L. B. Smith, B. R. Walker and R. Andrew, Anal. Chem., 2016, 88, 10362–10367 CrossRef CAS PubMed.
- E. Fridjonsdottir, R. Shariatgorji, A. Nilsson, T. Vallianatou, L. R. Odell, L. S. Schembri, P. Svenningsson, P. O. Fernagut, A. R. Crossman, E. Bezard and P. E. Andrén, Sci. Adv., 2021, 7, eabe5948 CrossRef CAS PubMed.
- C. Harkin, K. W. Smith, F. L. Cruickshank, C. Logan Mackay, B. Flinders, R. M. A. Heeren, T. Moore, S. Brockbank and D. F. Cobice, Mass Spectrom. Rev., 2022, 41, 662–694 CrossRef CAS PubMed.
- I. Kaya, L. S. Schembri, A. Nilsson, R. Shariatgorji, S. Baijnath, X. Zhang, E. Bezard, P. Svenningsson, L. R. Odell and P. E. Andrén, J. Am. Soc. Mass Spectrom., 2023, 34, 836–846 CrossRef CAS PubMed.
- M. Shariatgorji, N. Strittmatter, A. Nilsson, P. Källback, A. Alvarsson, X. Zhang, T. Vallianatou, P. Svenningsson, R. J. A. Goodwin and P. E. Andrén, NeuroImage, 2016, 136, 129–138 CrossRef CAS PubMed.
- S. Shimma, H. O. Kumada, H. Taniguchi, A. Konno, I. Yao, K. Furuta, T. Matsuda and S. Ito, Anal. Bioanal. Chem., 2016, 408, 7607–7615 CrossRef CAS PubMed.
- Q. Zhou, A. Fülöp and C. Hopf, Anal. Bioanal. Chem., 2021, 413, 2599–2617 CrossRef CAS PubMed.
- C. Tie, T. Hu, Z. X. Jia and J. L. Zhang, Anal. Chem., 2016, 88, 7762–7768 CrossRef CAS PubMed.
- R. M. Caprioli, T. B. Farmer and J. Gile, Anal. Chem., 1997, 69, 4751–4760 CrossRef CAS PubMed.
- J. L. Norris and R. M. Caprioli, Chem. Rev., 2013, 113, 2309–2342 CrossRef CAS PubMed.
- C. Esteve, E. A. Tolner, R. Shyti, A. M. J. M. van den Maagdenberg and L. A. McDonnell, Metabolomics, 2016, 12, 30 CrossRef PubMed.
- I. Kaya, S. M. Brulls, J. Dunevall, E. Jennische, S. Lange, J. Martensson, A. G. Ewing, P. Malmberg and J. S. Fletcher, Anal. Chem., 2018, 90, 13580–13590 CrossRef CAS PubMed.
- M. Niehaus, J. Soltwisch, M. E. Belov and K. Dreisewerd, Nat. Methods, 2019, 16, 925–931 CrossRef CAS PubMed.
- M. Shariatgorji, P. Källback, L. Gustavsson, N. Schintu, P. Svenningsson, R. J. A. Goodwin and P. E. Andrén, Anal. Chem., 2012, 84, 4603–4607 CrossRef CAS PubMed.
- M. Shariatgorji, A. Nilsson, R. J. A. Goodwin, P. Svenningsson, N. Schintu, Z. Banka, L. Kladni, T. Hasko, A. Szabo and P. E. Andrén, Anal. Chem., 2012, 84, 7152–7157 CrossRef CAS PubMed.
- M. Shariatgorji, A. Nilsson, E. Fridjonsdottir, T. Vallianatou, P. Källback, L. Katan, J. Sävmarker, I. Mantas, X. Zhang, E. Bezard, P. Svenningsson, L. R. Odell and P. E. Andrén, Nat. Methods, 2019, 16, 1021–1028 CrossRef CAS PubMed.
- M. Shariatgorji, A. Nilsson, R. J. A. Goodwin, P. Källback, N. Schintu, X. Zhang, A. R. Crossman, E. Bezard, P. Svenningsson and P. E. Andrén, Neuron, 2014, 84, 697–707 CrossRef CAS PubMed.
- A. Girard and G. Sandulesco, Helv. Chim. Acta, 1936, 19, 1095–1107 CrossRef CAS.
- Y. Sugiura, E. Takeo, S. Shimma, M. Yokota, T. Higashi, T. Seki, Y. Mizuno, M. Oya, T. Kosaka, M. Omura, T. Nishikawa, M. Suematsu and K. Nishimoto, Hypertension, 2018, 72, 1345–1354 CrossRef CAS PubMed.
- A. Nilsson, R. J. A. Goodwin, M. Shariatgorji, T. Vallianatou, P. J. Webborn and P. E. Andrén, Anal. Chem., 2015, 87, 1437–1455 CrossRef CAS PubMed.
- E. L. Gill, J. P. Koelmel, R. A. Yost, M. S. Okun, V. Vedam-Mai and T. J. Garrett, Anal. Chem., 2018, 90, 2979–2986 CrossRef CAS PubMed.
- G. J. McBean, M. Aslan, H. R. Griffiths and R. C. Torrão, Redox Biol., 2015, 5, 186–194 CrossRef CAS PubMed.
- M. Schumacher, S. Weill-Engerer, P. Liere, F. Robert, R. J. M. Franklin, L. M. Garcia-Segura, J. J. Lambert, W. Mayo, R. C. Melcangi, A. Parducz, U. Suter, C. Carelli, E. E. Baulieu and Y. Akwa, Prog. Neurobiol., 2003, 71, 3–29 CrossRef CAS PubMed.
- P. Källback, M. Shariatgorji, A. Nilsson and P. E. Andrén, J. Proteomics, 2012, 75, 4941–4951 CrossRef PubMed.
- W. F. Ward, P.-J. P. Lin, P. S. Wong, R. Behnia and N. Jalali, Radiat. Res., 1993, 135, 81–87 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4cc02475c |
| ‡ These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2024 |

![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
: