Open Access Article
Open Access ArticleMetal and metal oxide nanoparticle toxicity: moving towards a more holistic structure–activity approach†
G. P.
Gakis
,
I. G.
Aviziotis
and
C. A.
Charitidis
 *
*
Research Lab of Advanced, Composite, Nano-Materials and Nanotechnology, Materials Science and Engineering Department, School of Chemical Engineering, National Technical University of Athens, 9 Heroon Polytechneiou Street, Zografos, Athens 15780, Greece. E-mail: charitidis@chemeng.ntua.gr; Tel: +30 2107724046
First published on 30th January 2023
Abstract
The recent emergence of nanotechnology has led to the rapid increase of intentional and unintentional exposure to engineered nanoparticles (NPs), raising concerns over their impact on humans, animals and ecosystems. The demanding experimental assessment of toxicity, compared with NP innovation and time to market, has led to the extensive development of in silico methods, such as SAR models, aiming at providing a more rapid toxicity screening of such NPs. However, such models are usually built upon a limited number of data, making the different approaches case-sensitive. Furthermore, the focus on the predictive capabilities of the models, deem the extraction of scientific knowledge secondary, hindering the mechanistic understanding of toxicity mechanisms. In this paper, we instead shift the focus by using the models as a first step towards induction and extraction of valuable mechanistic information, once the predictive ability of the model has been validated. For this reason, we use a large dataset consisting of 935 toxicity measurements for 45 metal and metal oxide NPs, to build classification nano-SAR models. To the best of the authors' knowledge, this is the largest dataset of individual toxicity measurements for such NPs. Although the dataset is heterogeneous, the models developed are able to accurately classify the NPs based on their toxicity towards a variety of cells and organisms, using the same descriptors. Based on the quality of the results, the potential mechanisms of toxicity are identified and discussed in depth, providing a more holistic approach towards metal and metal oxide NP toxicity. The presented approach aims to trigger a discussion regarding information that could be derived from nano-SAR models, that could pave the way towards a more knowledge-based risk assessment of NPs and guide researchers towards the synthesis of safe-by-design NPs.
Environmental significanceThe emerging applications of nanotechnology have increased the intentional and unintentional exposure to engineered nanoparticles (NPs), raising concerns over their impact on humans, animals and ecosystems. Furthermore, in vitro and in vivo toxicity assessment methods are very demanding compared with innovation and time-to-market of NPs. In this work, an in silico classification approach is presented for the toxicity of metal and metal oxide NPs towards a variety of cells and organisms, namely human and mammalian cells, eukaryotes, crustaceans, bacteria, fish and plants. This holistic approach provides a general understanding regarding the toxicity mechanisms of metal and metal oxide NPs, assisting the production of safe-by-design NPs, thus minimizing their hazard towards humans, animals and organisms in the environment. |
1. Introduction
During the last decades, the field of nanotechnology has emerged as an increasingly important part of today's technology and economy.1–3 The plethora of applications in electronics,4 medicine,5 food,6 energy7 and catalysis,8 have made engineered nanomaterials (ENMs) an indispensable part of everyday life. As a consequence, scientific research has focused towards the synthesis and investigation of the properties of such ENMs in the form of nanofilms,9–11 nanoparticles (NPs)12 and nanomaterials in other promising forms, such as nanotubes.13,14The increased use and production of ENMs has led to an increasing exposure to such materials, especially in the form of NPs. Such exposure can be intentional, for example through the use of nanoparticles as carriers or radiosensitizers,15,16 or through vaccines, as demonstrated in the case of COVID-19 vaccines.17–19 The exposure to NPs can also be non-intentional, occurring in workplaces,20 households,21 atmospheric air,22 as well as through the use of common consumer products.23 It is evident that this increased production, use and exposure has raised concerns regarding the safety of such NPs, not only for humans, but also for ecosystems.24
The hazard and risk of NPs exposure is usually addressed by assessing the NP toxicity. The toxicity of a given NP can be investigated using in vitro or in vivo assessments. In vitro toxicity methods are widely used for the toxicity screening of NPs, as they provide a lower cost, less time consuming, and include less ethical concerns.25 Using such methods, the acute toxicity of NPs can be estimated using different methods and assays, towards different cell lines.25 On the other hand, in vivo methods constitute the most reliable methods to determine toxicity, as such a study includes the absorption, distribution, metabolism and excretion (ADME), that play a significant role for the human, animal and organism exposure to NPs.26 However, in vivo assessments have a high cost, are more time consuming and include ethical concerns regarding animal testing. For this reason, in vitro studies are usually the first step of the toxicity assessment, by conducting dose–response studies to find the NP concentrations for which toxic effects arise, followed by in vivo assessments under controlled exposure.26 In any case, the toxicity assessment of a large number of NPs, under a wide range of concentrations, towards various cell lines and organisms, is very time consuming and experimentally demanding compared with innovation and time-to-market of nanomaterials.27
In this context, in silico approaches have emerged as an alternative toxicity assessment method, during the last decade.27–29 Within those approaches, structure–activity relationship (SAR) models have been widely developed for NP toxicity assessment,30 especially for the case of metal oxide NPs. Being data-driven models, they use correlations between physicochemical or structural properties of the NPs, known as descriptors, and toxicity endpoint data, in order to build predictive models for the NPs toxicity.31 These models can be either quantitative, predicting the value of the toxicity endpoint for a given NP,32 or classification models, where the toxic class of a NP is predicted.27,29 These approaches offer the potential of a low-cost and time-efficient way to obtain a rapid toxicity screening of NPs, which is a promising tool in the effort to reduce the time and cost of experiments, as well as to minimize animal testing. Although still limited in regulatory use, the potential of nano-SAR is a key subject of present research, and a potential future technology and tool for the risk assessment of NPs.1
The toxicity of metal oxide nanoparticles has been extensively studied, with series of different NPs being tested towards various cells and organisms.33–35 This has led to the extensive development of nano-SAR models for metal oxides, built using several descriptors and toxicological endpoints.32,36,37 The interested reader is referred to the reviews of Buglak et al.30 and Li et al.,38 concerning developed SAR models for various metal and metal oxide NPs. Using such models, important insights regarding the mechanisms of action of metal oxide NPs can be provided.39,40
However, the majority of the developed models are built on individual datasets from a certain study, dealing with the toxicity of a series of metal oxide NPs, on a specific cell line, under the same experimental conditions. This has led to models being built on a limited number of data points (usually less than 30), dealing with a limited number of different NPs.32,36,41,42 Nonetheless, attempts have been made to build nano-SAR models using toxicity data for a wider number of NPs, towards various cell lines and organisms under different experimental conditions. In particular, Kleandrova et al., developed classification models for a dataset of 84 measurements, regarding the ecotoxicity of 18 different metal and metal oxide NPs towards 20 different organisms,43 showing that the toxicity of multiple NPs could be predicted towards these organisms with a high accuracy. The same group of researchers also used a similar approach for the prediction of cytotoxic effects of NPs, using a dataset of 40 measurements for the cytotoxicity towards mammalian cell lines.44 The simultaneous prediction of toxicity towards various organisms and cells of different kind under different experimental conditions was also realized, providing high accuracy models, using datasets consisting of up to 260 measurements.45,46 However, the combination of descriptors used in larger datasets is often complex,47 rendering the mechanistic interpretation of the model results a rather challenging task.
Although the predictive ability of SAR models is a primary goal of such model development, the mechanistic interpretation of the model results should also be of high importance, as stated in the OECD standards for the validation of (Q)SAR models.48 Many QSAR models, built for limited datasets, have used several combinations of descriptors to predict toxicity of metal oxide NPs,32,37,42 providing different information regarding the toxicity mechanisms. Furthermore, other works have used complex descriptors, that can render the understanding and extraction of toxicity mechanisms challenging.41,43,45,47,49–52 All the above have rendered the interpretation of QSARs case-sensitive, often failing to extrapolate the models to other datasets, even in a qualitative way. On the other hand, Burello and Worth suggested a framework to approach the metal oxide NP toxicity in a more mechanistic way.39,53 This approach has since been used to explain experimental results regarding the toxicity of different sets of metal oxide NPs.27,35,36,40,54 Although these different toxicity measurements have a somewhat limited number of NPs and are applied in each case separately, they deal with metal oxide toxicity towards a variety of cell lines and organisms, and the results could consistently be explained by the framework proposed by Burello and Worth.53 This could hint that (Q)SAR models could indeed be used successfully to extract information regarding the toxicity mechanisms in a more holistic way, and not in a case-dependent manner. This is something that is often overlooked in (Q)SAR models, where the intent towards a more global mechanistic explanation is deemed secondary to a better predictive ability towards the dataset of the specific study. However, in order to proceed with a more holistic approach towards metal oxide NP toxicity, extensive datasets for a wide number of NPs with toxicity measurements towards different cell lines/organisms under different experimental conditions need to be built, in order to test more global hypotheses.
In the present manuscript, we aim at investigating whether mechanistic information regarding NPs toxicity can be drawn using large datasets for a wide number of NPs and measurements towards different cell lines and organisms. For this reason, a dataset that consists of 935 individual measurements for half maximal concentrations of 45 different metal and metal oxide NPs, towards various cell lines and organisms, is built. This dataset is, to the best of the authors' knowledge, the largest set of individual half maximal concentration measurements used for the development of a metal oxide nano-SAR model. Using this dataset, toxicity classification models are built for individual cell lines and organisms, as well as cell and organism groups, namely human cells, mammalian cells, bacteria, crustaceans, fish, plants and eukaryotes. A predictive model is also built for the complete dataset, providing a toxicity assessment for all 45 NPs.
The novelty of the approach lies in the size and heterogeneous nature of the dataset which is used to build the models. Furthermore, the fact that the same combination of descriptors is used for the models built for all the different cell lines, as well as the cell and organism groups, allows a more general description of the key NP properties that induce toxicity, along with an insight on the toxicity mechanism of metal oxide NPs. Although such mechanisms have been previously addressed for the toxicity of certain metal oxides towards a limited number of cell lines, the large and heterogeneous dataset of the present work allows a more holistic understanding of the metal oxide toxicity towards various cells and organisms. It is also shown that the approach can be extended to account for the case of metal NPs. To this end, the presented approach aims at triggering a discussion regarding information that could be derived from nano-(Q)SAR modelling methods and it may pave the way towards a more holistic understanding of metal and metal oxide NPs toxicity, as well as their safe application to the different fields of daily life.
2. Methods
2.1 Dataset
The dataset used in this work consists of 935 measurements for half maximal effective (EC50), inhibitory (IC50), and lethal (LC50) concentration toxicity endpoints, obtained from 70 published research papers.32–37,40–42,55–115 Although most SAR models are built using a specific endpoint, the inclusion of these different half maximal concentrations within the same dataset has been previously addressed.43,46 The data incorporates measurements for 35 different metal/semimetal oxide and 10 metal nanoparticles (NPs). The endpoints were measured after the exposure of different cells and organisms to NPs, using different assays. The cell types in the dataset measurements are divided into cell groups: crustaceans, eukaryotes, plants, fish, bacteria, mammalian cells, and human cells. The measurements as well as the corresponding experimental exposure times, can be found in the ESI† of the paper (ESI† file 2). It is noted here that SAR models are usually built using a given endpoint, towards a specific cell line, under the same experimental conditions. However, the inclusion of different half maximal concentration endpoints, towards different cells and organisms, under different experimental conditions within the same dataset for the development of SAR models has been previously addressed.43–472.2 Toxicity classification
The classification of each NP measurement as toxic or non-toxic was based on previous published works. In particular, the measurement was classified as toxic if the logarithm of the half maximal concentration endpoint, in molar concentration units (mol L−1) was lower than −2.5 (log(C50) ≤ −2.5), based on the work of Simeone and Costa.27 Usually, the reported values for the half maximal concentrations are reported in mass concentrations and a conversion to molar units needs to be conducted. The threshold mass concentration value for each NP (leading to a molar log(C50) = −2.5) is presented in the ESI† of the paper (ESI† file 3). However, in many cases, the range of experimental concentrations tested is limited, and the half maximal concentrations are reported as just being higher than the maximum tested concentrations (C50 > Cmax,tested) (e.g. EC50 > 100 mg L−1). Hence, a framework for the classification of such kind of measurements is needed.In the present work, this is done as follows: If the maximum concentration (in molar units) tested yields a logarithm value higher than the threshold (log(Cmax) > −2.5), then the half maximal concentration was set to the maximum tested concentration, and the NP is classified as non-toxic. Otherwise, if the logarithm value is lower than the threshold (log(Cmax) ≤ −2.5), then the following procedure is adopted: If the maximum concentration tested is more than 40% of the threshold concentration, the NP is classified as non-toxic. For example, the EC50 TiO2 was estimated to be >200 mg L−1 against BEAS-2B and RAW 264.7 cells in,35 which is below the threshold value for TiO2 (∼253 mg L−1). However, the maximum concentration tested is more than 40% the threshold value. Hence, the measurement of EC50 > 200 mg L−1 for TiO2 is classified as non-toxic. On the other hand, if the maximum concentration tested is less than 40% of the threshold value, the measurement is omitted from the dataset. The value of 40% was arbitrarily chosen, so that a significant amount of NP has been exposed experimentally in order to classify the NP as non-toxic, as well as to reduce the number of data omitted from the dataset. The effect of this arbitrary value is presented in the Results section (section 3.7).
2.3 Descriptors
The initial descriptor matrix consisted from a list of 41 descriptors for each NP. The initial descriptors used were the metal cation charge (z) of the NP as well as the ionic radius (r), taken from ref. 116. The hydration energy (HE) of the metal ion was computed using Latimer's equation:40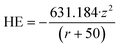 | (1) |
| Dbio = abs(Ec − (−4.48) eV) | (2) |
2.4 Descriptor selection
Once the complete set of descriptors is obtained, feature selection methods are employed in order to reduce the number of descriptors to be tested by the model, by discarding redundant descriptors. This is done to reduce the complexity of the model and increase the interpretability of the results. First, intercorrelated descriptors are identified using Pearson's correlation coefficient,124 so that only one of intercorrelated descriptors is used. Then, ReliefF,125 minimum redundancy maximum relevance (MRMR)126 and chi-square127 were used as feature selection methods to rank the different descriptors based on their relevance for the prediction of the endpoints. Using the above methods, the list of 4 most relevant descriptors were obtained. Finally, combinations of these descriptors were tested to develop the optimal models for the toxicity classification of NPs.2.5 Model implementation, training and validation
The total dataset is divided in subsets based on the cell or organism on which the NP toxicity was tested. A minimum of 30 measurements was set as a criterion for the creation of each dataset. From the above data division, seven different subsets of data were built for individual cell lines or organisms. Then, datasets were built for cells that belong to the same type of organisms. This resulted to seven different groups, namely mammalian cells, human (and human cancer) cells, fish, plants, bacteria, crustacean, and eukaryotes. Finally, the total set of measurements was also used as a dataset. The details for each dataset are given in Table 1.| Cell type/cell type | No of measurements | No of NPs | % of toxic measurements | % of non-toxic measurements |
|---|---|---|---|---|
| Individual cell lines and organisms | ||||
| A549 | 168 | 31 | 55.9 | 44.1 |
| BEAS-2B | 39 | 13 | 54.5 | 45.5 |
| Caco-2 | 39 | 8 | 61.5 | 38.5 |
| Danio rerio | 62 | 18 | 46.8 | 53.2 |
| E. coli | 135 | 30 | 60.7 | 39.3 |
| Daphnia magna | 41 | 8 | 78 | 22 |
| HaCaT | 42 | 25 | 47.6 | 52.4 |
| Cell line and organism groups | ||||
|---|---|---|---|---|
| Human cells | 408 | 35 | 58.6 | 41.4 |
| Mammalian cells | 58 | 18 | 53.4 | 46.6 |
| Fish | 73 | 18 | 47.9 | 52.1 |
| Plants | 63 | 14 | 54 | 46 |
| Crustaceans | 64 | 12 | 84.4 | 15.6 |
| Bacteria | 190 | 35 | 63.1 | 36.9 |
| Eukaryotes | 68 | 20 | 39.7 | 60.3 |
| Complete dataset | 935 | 45 | 58.6 | 41.4 |
For datasets with a lower number of measurements (n < 100), a five-fold cross validation was used. For datasets with a higher number of observations (n ≥ 100), a hold-out validation was performed by using 80% of the data as a training set and 20% as a validation set. The data splitting was random and performed using MATLAB®. The resulting sets and folds of data can be found in the ESI† of the paper (ESI† files 4–6). The model training was performed in MATLAB®, with the classification learner toolkit, using different classification algorithms, such as support vector machines (SVM), k-nearest-neighbors (kNN), and random forests (RF). The optimal models were selected for each dataset using different statistical metrics, based on the confusion matrix of the model results over the validation set. The metrics used for the model performance are shown below:
Accuracy (ACC):
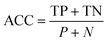 | (3) |
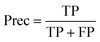 | (4) |
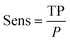 | (5) |
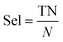 | (6) |
2.6 Applicability domain
According to the OECD standards, an applicability domain should be defined for a developed SAR model.48 The applicability domain defines the descriptor space within which the developed model is applicable. Any NP tested with the model should therefore be within the applicability domain defined for the particular model. Many different techniques and methods have been developed and used for the definition of the applicability domain in quantitative and classification SAR models.128 In this work, the applicability domain is defined using the bounding box PCA, convex hull and Euclidean distance from centroid methods.128 The above analysis is performed using the Applicability Domain toolbox developed for MATLAB by the Milano Chemometrics and QSAR Research Group.128,1293. Results
3.1 Descriptor selection
The first step of the descriptor selection is the identification of intercorrelated descriptors. As previously discussed, this analysis is performed using Pearson's correlation coefficient.124 The absolute Pearson correlation coefficient matrix is presented in Fig. S1, in ESI† (ESI† file 1). The threshold value for a set of descriptors to be deemed as correlated was set to 0.95, based on the findings of Rácz et al.130 From this process, 29 descriptors are eliminated from the initial set, presented in Table S1 of the ESI† (ESI† file 1).Following the intercorrelation analysis, the remaining set of descriptors are analyzed using the ReliefF,125 MRMR126 and chi-square127 methods. These methods rank the descriptors based on their relevance towards the response variable (i.e. the toxicity classification). Using these methods, the four most relevant descriptors are identified. The resulting descriptors and their impact scores, as identified by the applied methods, are presented in Fig. 1, below.
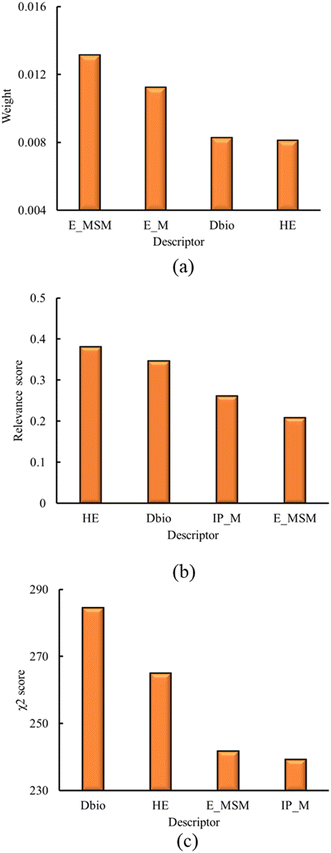 | ||
| Fig. 1 Results for the four most relevant descriptors, as derived from a) ReliefF, b) MRMR, c) chi-square methods. | ||
Results of Fig. 1 show that HE, Dbio and the sum of electrons of the metals and semimetals (E_MSM) are ranked among the four most relevant descriptors, using all of the methods mentioned above. The electrons of the metals (E_M), and the metal ionization potential (IP_M) are also identified within the most relevant descriptors, using ReliefF, MRMR and chi-square methods, respectively. However, from the correlation analysis, E_M and E_MSM showed a correlation of 0.89. Although this correlation value is below the threshold of 0.95, E_M and E_MSM are very similar in their physical meaning, and differ only for the semimetal oxides. For this reason, only E_MSM is chosen for the subsequent analysis, while E_M is omitted.
3.2 Individual cell lines and organisms
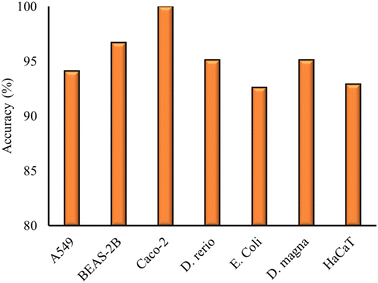
The first part of this investigation presents the results of the models built on datasets consisting of measurements on the NP toxicity on one particular cell line or organism. The models are built by using different combinations of the four NP descriptors obtained from the results of the previous section to identify the optimal descriptor combination. Descriptors were omitted from the model development when the statistical metrics were not significantly affected by their inclusion. The results of the different models built with various descriptor combinations are presented in the ESI† of the paper, for the representative case of A549 cells, which is the most populous dataset (ESI† file 1). The optimal classification models derived for the individual cell lines and organisms are presented in Table 2, along with the model details and statistical metrics. The model accuracy for the different models is also plotted in Fig. 2.| Cell line/organism | Model | Descriptors | Validation scheme | Acc (%) | Prec (%) | Sens (%) | Sel (%) |
|---|---|---|---|---|---|---|---|
| A549 | SVM | HE, Dbio | Training (n = 134) | 94.1 | 94.1 | 94.1 | 94.1 |
| Validation (n = 34) | |||||||
| BEAS-2B | kNN, RF | HE, Dbio | 5-fold cross validation | 96.7 | 100 | 94.4 | 100 |
| Caco-2 | SVM | HE, Dbio | 5-fold cross validation | 100 | 100 | 100 | 100 |
| Danio rerio | SVM, RF | HE, Dbio | 5-fold cross validation | 95.1 | 93.3 | 96.5 | 93.4 |
| E. coli | SVM, RF | HE, Dbio | Training (n = 108) | 92.6 | 100 | 90.9 | 100 |
| Validation (n = 27) | |||||||
| Daphnia magna | SVM, kNN, RF | HE, Dbio | 5-fold cross validation | 95.1 | 100 | 93.7 | 100 |
| HaCaT | kNN, RF | HE, Dbio | 5-fold cross validation | 92.9 | 100 | 85 | 100 |
As presented in Table 2, for all datasets, the optimal combination of descriptors was found to be the hydration enthalpy (HE) and the energy difference between the conduction band and the average value of the redox potential in biological media pairs (Dbio). The statistical metrics computed had acceptable values, with an accuracy percentage higher than 92% being computed for all models (Fig. 2). The Caco-2 model produced a 100% accuracy due to the different measurements of NP toxicity within the dataset, which are well described and separated by the two descriptors used. The high accuracy of the model could be mainly assigned to the nature of the initial dataset and not only the model predictive ability. The developed models showed acceptable values for the rest of the statistical metrics. The model for HaCaT cells showed a slightly lower selectivity (85%) than the other models, nevertheless it still exhibited acceptable values for those metrics.
For the two datasets that were validated using hold-out validation, the model was able to accurately predict the toxicity of NPs that did not have a measurement for toxicity within the training set of the respective model. In particular, for the A549 model, Yb2O3 and Bi2O3 were accurately classified as toxic, while MoO3 and HfO2 as non-toxic. For the E. coli model, Co2O3, Ag2O and Er2O3 were correctly classified as toxic.
The above classification model results show that the metal/semimetal oxide and metal NPs toxicity towards the different cell lines and organisms can be predicted using two descriptors. Also, the high accuracy of the developed models shows that the hydration energy and the energy difference between the conduction band of the NPs and the redox potential of biological pairs can describe the toxicity of the NPs towards various cells, which can assist towards mechanistic interpretations of the model results regarding the NP toxicity pathways.
3.3 Cell and organism groups
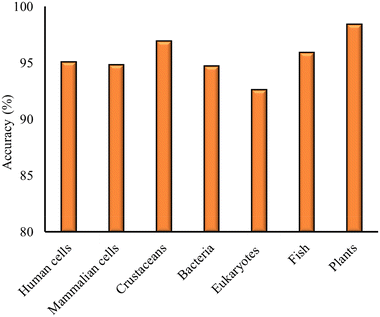
Once the models for the individual cell lines and organisms have been built, the focus is shifted towards the development of models with the datasets consisting of measurements for the different cell and organism groups. The same procedure was followed as in the case of models built for individual cell lines and organisms. The optimal classification models developed for the different cell and organism groups are presented in Table 3, along with the model details and statistical metrics. The model accuracy for the different models is also plotted in Fig. 3.| Cell/organism group | Model | Descriptors | Validation scheme | ACC (%) | Prec (%) | Sens (%) | Sel (%) |
|---|---|---|---|---|---|---|---|
| Human cells | SVM | HE, Dbio | Train. (n = 327) | 95.06 | 100 | 90.1 | 100 |
| Val. (n = 81) | |||||||
| Mammalian cells | SVM | HE, Dbio | 5-fold cross validation | 94.8 | 91.2 | 100 | 88.9 |
| Crustaceans | RF | HE, Dbio | 5-fold cross validation | 96.9 | 100 | 96.3 | 100 |
| Bacteria | SVM | HE, Dbio | Training (n = 152) | 94.7 | 92.6 | 100 | 84.6 |
| Validation (n = 38) | |||||||
| Eukaryotes | RF | HE, Dbio | 5-fold cross validation | 92.6 | 92.3 | 88.9 | 95.1 |
| Fish | SVM | HE, Dbio | 5-fold cross validation | 95.9 | 94.4 | 97.1 | 94.7 |
| Plants | SVM, kNN | HE, Dbio | 5-fold cross validation | 98.4 | 97.1 | 100 | 96.6 |
As in the case of models built for individual cell lines and organisms, HE and Dbio were found to be the optimal combination of descriptors. The optimal models were developed using RF, SVM and kNN methods, as shown in Table 3. The accuracy of all models was found to be above 92% (Fig. 3). Precision, sensitivity and selectivity were also above 90%, except from the eukaryotes data, where the classifier showed an 88.9% sensitivity, the selectivity for the mammalian cells model, which was 88.9%, and the selectivity of the bacteria model, which was 84.6%.
For the two datasets that were validated using hold-out validation, namely for human cells and bacteria, the model was able to accurately classify the NPs that did not have a measurement for toxicity within the training set of the respective model. Specifically, the human cell model classified HfO2 as non-toxic, consistent with the measurements of HfO2 toxicity in the validation set. The bacteria model classified Bi2O3 and Ni2O3 as toxic, consistent with the measurements for these NPs in the validation set.
Results of Table 3 show that a classification model can be built to predict the toxicity class of various metal oxide and metal NPs towards cell lines and organisms belonging to the same group. The hydration energy and conduction bang energy of the NPs can classify the NPs to toxic or not toxic with a high accuracy, for the cell and organism groups of Table 3. The fact that the same descriptors were found to be the most relevant to the toxicity class, as in the case of individual cell lines, along with the high accuracy of the models, show that these descriptors could be indicative of at least some of the toxicity mechanisms of the NPs. This will be further elaborated in the Discussion section.
3.4 Complete dataset
The complete dataset of 935 measurements of toxicity for 45 different NPs was also used to build a toxicity classification model with the aim to investigate whether or not the toxic class can be predicted using toxicity measurements of several NPs towards different cell lines and organisms under different toxicity assays. The same procedure was followed as in the case of models built for individual cell lines and organisms. The optimal classification model developed for the complete NP toxicity dataset is presented in Table 4, along with the model details and statistical metrics.| Cell/organism group | Model | Descriptors | Validation scheme | ACC (%) | Prec (%) | Sens (%) | Sel (%) |
|---|---|---|---|---|---|---|---|
| Complete dataset | SVM, RF | HE, Dbio | Train. (n = 748) | 92.5 | 90.2 | 97.1 | 86.6 |
| Val. (n = 187) |
As for the previous cases, the optimal model for the complete dataset was built using the combination of HE and Dbio as descriptors. The models built using kNN and SVM methods showed the same class predictions for the validation set of data. The accuracy, precision, sensitivity were all above 90%, while selectivity is 86.6%.
The results presented in Tables 2–4 show that the hydration enthalpy and the descriptor Dbio can classify the NPs in the datasets with a high accuracy for individual cell lines and organisms, cell and organism groups, as well as the complete toxicity dataset of all NP toxicity measurements, irrespective of the cell or organism on which the toxicity was tested. This shows that the two descriptors used to build the model can be used for a mechanistic interpretation of the model results thus, assisting in the understanding of the mode of action of the different metal and metal/semimetal oxide NPs towards different type of cells.
3.5 Model extrapolation
Based on the results of the previous sections, the developed models for the individual cell lines and organisms, as well as the different cell and organism groups showed a good accuracy for the toxicity classification. Furthermore, the same set of descriptors, HE and Dbio, were used to classify the NPs as toxic or non-toxic, for all datasets. Based on these findings, the possibility of a model developed for a specific cell line being extrapolated to classify the NPs toxicity class towards other individual cells and organisms, cell and organism groups, as well as the complete set of data was investigated. This was done by using the model built for the A549 cells to predict the toxicity class of the NPs in the rest of the datasets presented in Table 1. The choice of the A549 model is due to the fact that it is the individual cell model built with the largest dataset, both in terms of training data points (134 toxicity measurements) and number of NPs (27 NPs from the total of 45 from the complete set of data). The results of the A549 model extrapolation are presented in Table 5.| Cell/organism dataset | ACC (%) | Prec (%) | Sens (%) | Sel (%) |
|---|---|---|---|---|
| Individual cells and organisms | ||||
| BEAS-2B | 93.9 | 90 | 100 | 86.7 |
| Caco-2 | 100 | 100 | 100 | 100 |
| Danio rerio | 88.7 | 82.4 | 96.6 | 81.8 |
| E. coli | 89.6 | 90.5 | 92.7 | 84.9 |
| Daphnia magna | 92.7 | 100 | 90.6 | 100 |
| HaCaT | 88.1 | 85.7 | 90 | 86.4 |
| Cell and organism groups | ||||
|---|---|---|---|---|
| Human cells | 92.4 | 92.6 | 94.6 | 89.4 |
| Mammalian cells | 98.3 | 96.9 | 100 | 96.2 |
| Fish | 90.4 | 85 | 97.1 | 84.2 |
| Plants | 91.9 | 88.9 | 97 | 86.2 |
| Crustaceans | 93.8 | 100 | 92.6 | 100 |
| Bacteria | 90.5 | 91.8 | 93.3 | 85.7 |
| Eukaryotes | 92.6 | 92.3 | 88.9 | 95.1 |
| Complete dataset | ||||
|---|---|---|---|---|
| Complete dataset | 92.5 | 92.8 | 94.5 | 89.7 |
The results of Table 5 show that the predictions of the model built for A549 cells can classify the NPs based on their toxicity towards different cell lines and groups with an acceptable accuracy, for all datasets. Overall, the good accuracy towards the datasets, as well as the good accuracy towards the complete dataset, allows the conclusion that the two descriptors (HE, Dbio) can classify the NPs based on their toxicity against a large variety of cell and organism groups. Furthermore, the A549 model was trained over 27 different NPs. The model extrapolation to predict the measurements in the complete dataset (45 NPs) was used to assess the predictive capacity of the model for the classification of 18 NPs that were not part of the training set. The results are shown in Table 6.
| NP | Model classification | No of toxic measurements | No of non-toxic measurements |
|---|---|---|---|
| Ag2O | Toxic | 7 | 0 |
| Al | Non-toxic | 1 | 3 |
| Au | Toxic | 8 | 0 |
| Bi2O3 | Toxic | 12 | 2 |
| CaO | Non-toxic | 0 | 2 |
| Co2O3 | Toxic | 1 | 0 |
| Er2O3 | Toxic | 1 | 0 |
| Fe3O4 | Non-toxic | 1 | 12 |
| Ga2O3 | Non-toxic | 0 | 2 |
| HfO2 | Non-toxic | 0 | 5 |
| MoO3 | Non-toxic | 0 | 2 |
| Ni2O3 | Toxic | 5 | 0 |
| Pd | Toxic | 3 | 0 |
| Sn | Non-toxic | 0 | 2 |
| V2O3 | Toxic | 3 | 1 |
| W | Non-toxic | 0 | 2 |
| Yb2O3 | Toxic | 3 | 1 |
| Zn | Toxic | 4 | 0 |
Table 6 shows that all 18 NPs that were classified by the A549 model agree with the majority of the measurement classifications. The toxicity class is consistent for the majority of toxicity measurements in the complete dataset. The good predictive ability of the A549 model for the NP toxicity classification towards different cells and organisms, even for NPs that were not in the model training set, allows us to conclude that the two descriptors used for the model training can classify the NPs based on their toxicity. Hence the model can be used to predict the toxicity class for a variety of different NPs, as well as to provide mechanistic interpretations regarding metal and metal oxide NPs toxicity towards cells and organisms. Furthermore, the extrapolation of the model developed for a specific cell line to predict the toxicity class towards other cell lines and organisms, shows the possibility of SAR analyses to unravel more holistic trends, as well as to suggest dominant mechanistic pathways in metal and metal oxide toxicity. This will be addressed in the discussion section.
3.6 Applicability domain
In order to explore the descriptor space over which the developed models are applicable, the applicability domain is determined. As discussed in section 2.6, the applicability domain was assessed using bounding box PCA, convex hull, and centroid distance methods. The NPs present in the validation sets for each model were tested in order to see if any NP is outside the applicability domain. The NPs that were deemed to be outside the applicability domain of the respective model, using the different methods for the applicability domain definition, are shown in Table 7.| Dataset | Bounding box PCA | Convex hull | Centroid distance |
|---|---|---|---|
| A549 | MoO3 | MoO3 | SiO2, MoO3, WO3 |
| BEAS-2B | — | — | SiO2 |
| Caco-2 | — | — | SiO2 |
| Danio rerio | — | — | — |
| E. coli | — | — | — |
| Daphnia magna | — | — | SiO2 |
| HaCaT | MoO3 | MoO3 | MoO3 |
| Human cells | MoO3, WO3 | — | MoO3, WO3 |
| Mammalian cells | — | — | — |
| Fish | — | — | WO3 |
| Plants | — | — | — |
| Crustaceans | — | — | SiO2 |
| Bacteria | — | — | — |
| Eukaryotes | — | — | — |
| Complete dataset | MoO3, WO3 | SiO2, MoO3, WO3 | SiO2, MoO3, WO3 |
The bounding box PCA and convex hull methods yielded applicability domains that included all the NPs in the validation set of each model, as seen in Table 7, except from the A549 and HaCaT datasets, where MoO3 was found to be outside the applicability domain. However, the centroid distance showed that WO3 and MoO3 were deemed to be outside the applicability domain of most datasets, when present in the validation set. SiO2 was also deemed to be outside the applicability domain for certain datasets. Hence, the applicability domain of a developed model depends on the method by which it is defined. The different methods have advantages and shortcomings and the issue of applicability domain definition has been a subject in relevant scientific works.128 In our approach, we used three methods to assess the applicability domain. From the results of Table 7, it is seen that oxides with a high metal oxidation number (+6) are situated mostly outside the applicability domain of the models. This is due to the more negative value of the HE descriptor value for these oxides, as will be shown in the next sections. Such applicability domains have been presented in works that used classification SARs built using similar descriptor combinations, with WO3 and SiO2 being deemed outside the applicability domain.54 This should be taken into account when the model results are discussed.
3.7 NP toxicity classification
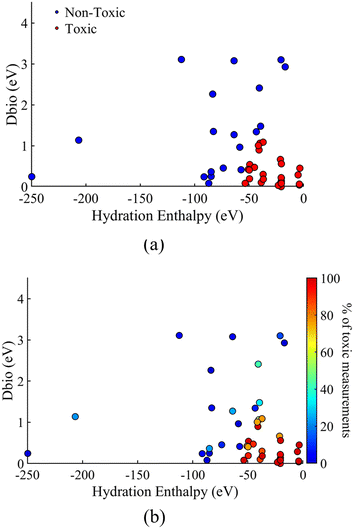
As all the developed models in sections 3.2–3.4 classified the NPs with good accuracy over the different datasets using the same descriptors, the complete dataset model was used to present the toxicity classification of the 45 different NPs over the descriptor space. The different NP classification, as a functions of the two descriptors used for the model development (HE, Dbio), is presented in Fig. 4a. The percentage of toxic measurements for each NP in the complete dataset is also plotted as a function of the two descriptors, in Fig. 4b, for comparison.Fig. 4 shows that the two descriptors are successful in classifying the different NPs in the toxic or non-toxic classes based on the mapping of the NPs over the two descriptors space. In particular, toxic NPs are characterized by a lower Dbio value as well as a low absolute value of the hydration enthalpy (less negative). According to the definition of the two descriptors in section 2.3, this shows that NPs that have a conduction band energy (Ec) close or within the range of the redox potential of couples in biological media along with a low energy release from the respective metal cation hydration, are more probable to be toxic.
The limiting values of the descriptor space for the toxicity classification, as derived from the model, are a hydration enthalpy of ≃55 eV, and a value of Dbio ∼1.1 eV. A less negative hydration enthalpy and a smaller Dbio value lead to a higher toxic probability of the NPs. On the other hand, more negative hydration enthalpies and higher Dbio values, lead to the NP being less probable to be toxic. From the results in Fig. 4b, it is seen that the percentage of toxic measurements for each NP follows the same trend over the descriptor space of the HE and Dbio descriptors. This means that for a lower absolute value of HE and Dbio for a given NP, there is a higher probability of the measurement being toxic. This validates the model classification of NPs based on their toxicity by applying the same descriptors.
As seen in the results of Fig. 4, the majority of the studied NPs exhibit a hydration enthalpy of ≥100 eV. The vast majority of NPs are situated in the region of 0 > HE ≥ 100, except from SiO2 (≃112 eV), WO3 (≃206 eV) and MoO3 (≃250 eV), which show a more negative hydration enthalpy. Similarly, most of the NPs have a value of Dbio < 2.5 eV, with only 5 NPs showing a higher value. Finally, Fig. 4 shows that MoO3, with a hydration energy of ≃250 eV, has the highest distance from the rest of the NPs, followed by WO3 and SiO2. This explains the results of the previous section, where MoO3, WO3, and SiO2 were predicted to be outside the applicability domain of most models, when distance from centroid was used to estimate the applicability domain.
Regarding the metallic NPs, using the complete dataset model, Ag, Au, Cu, Pd, Zn were classified as toxic NPs, validated by the vast majority of the measurements in the dataset. Co and Ni NPs have also been classified as toxic from the model, which is also consistent with the measurements. On the other hand, W, Sn and Al NPs were classified as non-toxic, consistent with the majority of measurements.
For the metal oxide NPs, Ag2O, Co2O3, CoO, CuO, Cu2O, Mn2O3, Mn3O4, Er2O3, Ni2O3, Yb2O3 and ZnO were all classified as toxic by the complete dataset model and validated by the vast majority of measurements. On the other hand, Al2O3, CaO, CeO2, Fe2O3, Ga2O3, Gd2O3, HfO2, MgO, MoO3, SiO2, SnO2, TiO2, WO3, Y2O3 and ZrO2 were all correctly classified as non-toxic, as shown from the vast majority of measurements. Bi2O3 and Cr2O3 have been classified as toxic, and agrees with the majority of measurements. However they have been shown to be non-toxic towards Danio rerio.82 Co3O4 has also been classified as toxic, consistently with most measurements, but it has been also found to be non-toxic towards plants.106 Fe3O4 has been found to be non-toxic towards bacteria,40 as well as mammalian and human cells35 and was characterized non-toxic by the model. Even though NiO and Sb2O3 have been deemed as non-toxic in some measurements towards BEAS-2B cells,35 the majority of measurements show that they are toxic towards human cells41,72,84,102 and classified as toxic by the model. The models classified V2O3 as toxic, although it has a hydration enthalpy value close to the limit for its classification as toxic. It has been reported to be toxic against E. coli.32,37 It was found non-toxic towards HaCaT cells in one study,36 however, other studies have found it to be toxic towards other human cells.131 Finally, La2O3, and In2O3 have a more balanced percentage of toxic measurements, with their Dbio values being close to the threshold between the toxicity classes. Nonetheless, for the model trained over the complete dataset, La2O3, and In2O3 were classified as toxic, consistent with the majority of the measurements.
This classification of the total 45 NPs on the basis of those two descriptors shows that the overall approach can be used not only as a predictive tool for the assessment of the metal and metal oxide NPs toxic class, but also as a starting point for the understanding of the mechanism of action of the different NPs towards different cells and/or organisms.
3.8 Effect of data classification scheme
In section 2.2, we presented the framework used to characterized the measurements in the datasets as toxic or non-toxic. The measurements that did not present an absolute value for a half maximal concentration (C50 > Cmax, tested) were characterized as non-toxic only if the maximum concentration tested was 40% of the threshold value that yields log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) C50 = −2.5. The 40% value is arbitrarily selected so that a significant amount of NP has been exposed experimentally to classify the NP as non-toxic. In this section, the effect of this arbitrary approach is investigated, by removing all non-absolute values of half maximal concentration endpoints. Hence, only endpoints with a reported value for C50 or for maximum tested concentrations that yield log
C50 = −2.5. The 40% value is arbitrarily selected so that a significant amount of NP has been exposed experimentally to classify the NP as non-toxic. In this section, the effect of this arbitrary approach is investigated, by removing all non-absolute values of half maximal concentration endpoints. Hence, only endpoints with a reported value for C50 or for maximum tested concentrations that yield log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Cmax > −2.5 are kept. Using this approach, the complete dataset consists of 783 measurements. The dataset is also more imbalanced, with 69.8% of measurements being toxic, as all the measurements that were removed, were non-toxic measurements. The model developed for the dataset consisting of only absolute endpoint values is developed and the results are presented in Table 8.
Cmax > −2.5 are kept. Using this approach, the complete dataset consists of 783 measurements. The dataset is also more imbalanced, with 69.8% of measurements being toxic, as all the measurements that were removed, were non-toxic measurements. The model developed for the dataset consisting of only absolute endpoint values is developed and the results are presented in Table 8.
| Cell/organism group | Model | Descriptors | Validation scheme | ACC (%) | Prec (%) | Sens (%) | Sel (%) |
|---|---|---|---|---|---|---|---|
| Complete dataset (absolute endpoints) | RF | HE, Dbio | Train. (n = 627) | 91.7 | 93.5 | 95.3 | 86 |
| Val. (n = 156) |
Results of Table 8 show that acceptable statistical metrics are obtained for the classification model built for the dataset consisting of only absolute endpoint values, with accuracy, precision and sensitivity all being above 91%, and selectivity being computed equal to 86%. The fact that the model showed HE and Dbio as the optimal descriptor combination, with good accuracy, precision sensitivity and selectivity, shows that the approach we used for the data characterization in section 2.2 did not affect the optimal descriptor combination or the model prediction statistical metrics qualitatively. However, the incorporation of the approach of section 2.2 allowed the addition of measurements, making the dataset larger and more balanced as the measurements of the type C50 > Cmax,tested are classified as non-toxic.
4. Discussion
The results presented in section 3 allow a mechanistic interpretation of the toxicity classification of the 45 NPs that are included in the different model datasets. All the models exhibited an acceptable accuracy for the individual models built for the respective datasets (section 3.2–3.4), while the model built with the data for A549 cells could be extrapolated to predict the toxicity class of NPs against other cells and organism groups, with an acceptable accuracy (section 3.5). All of the developed models used HE and Dbio as descriptors. The good overall accuracy of the models thus, allows to drive the deduction that these two descriptors can be applied to explain the NPs toxicity and predict their toxicity class.The descriptor Dbio represents the absolute energy difference between the conduction band of the NP and the mean value of standard redox potential of couples in biological media, as described in eqn (2). Hence, the descriptor Dbio indirectly represents the conduction band energy of the NPs. From the results of the presented models, NPs that are classified as toxic show a lower value for Dbio (Fig. 4), hence a conduction band energy that is within or close to the range of the redox potential of couples in biological media. This energy difference determines whether an electron transfer occurs between the NP and the redox pairs in biological media. NPs that show an overlapping conduction band energy and redox pair potential can transfer electrons from biological media, thus unbalancing the reducing capacity of a cell and resulting to increased oxidative stress in the cells.53
The energy difference between the conduction band and the redox potential of biological redox pairs has been previously used to describe the toxicity of metal oxide NPs. Burello and Worth developed a theoretical framework to predict the oxidative stress potential of metal oxide NPs.39,53 They correlated this energy difference with toxicity studies from literature for 6 such nanomaterials. This approach was then used from Zhang et al., to explain the different oxidative stress of 24 metal oxide NPs towards BEAS-2B and RAW 264.7 cells,35 showing a correlation between the experimental findings and the developed theoretical findings for the energy difference between the conduction band and the redox potential of the pairs in the cell cultures. Liu et al., subsequently used this data to build a nano-SAR model, predicting the probability of a metal oxide NP to be toxic.54 The developed model adopted the conduction band energy as one of the descriptors and it showed that the NPs were more probable to be toxic when the conduction band energy was within or close to the range of the redox potential of biological couples. Kaweeteerawat applied a similar approach to build a classification nano-SAR model for the toxicity class prediction of the same series of metal oxide NPs towards Escherichia coli.40 The developed model was able to reproduce the experimental findings regarding the membrane damage and generation of biotic and abiotic reactive oxygen species (ROS).40 Gajewicz et al., used the conduction band energy to explain a possible mechanism for metal oxide toxicity, following the experimental toxicity assessment of 18 metal oxide NPs toward HaCaT cell lines.36 Finally, a similar descriptor of reduction potential was implemented for the metal oxide NP grouping based on their redox activity.27
The exposure to metal oxide NPs has been previously reported to induce oxidative stress in cells.132 The generation of various ROS under UV irradiation has been investigated towards E. coli and was correlated with the relative position of the conduction band and redox potential of biological pairs.133 The main ROS produced were OH radicals and hydrogen peroxides. Similar results have been observed for metal oxides without irradiation,35,76,132,134–138 towards various cell lines. Several mechanisms have been presented for the production of ROS species upon exposure to metal oxide NPs, either by direct production of radicals and superoxides,133 or by catalyzing their production.53 Furthermore, the transfer of electrons between the metal oxide and the biological redox pair can result to a decreased efficiency of the reducing potential of antioxidants in the cells, leading to increased oxidative stress.53 This increase of oxidative stress through ROS generation or decrease of the reducing capacity of redox pairs in biological media can be expressed by the descriptor Dbio. The good accuracy of the model regarding the toxicity classification of 45 NPs, using 935 toxicity measurements towards various cells and organisms as well as the correlation between similar descriptors and ROS generation in previous works for a more limited number of measurements35,133 allows this interpretation.
Beside the conduction band relative energy to the biological redox potential, the hydration enthalpy of the metal cation was also deemed as an optimal descriptor for the toxicity classification of metal and metal oxide NPs. The hydration enthalpy descriptor expresses the energy released form the hydration of a metal ion, or the affinity of water molecules to the metal ion.40 A higher absolute value (more negative hydration enthalpy) for a given metal cation shows that the water molecules have more affinity towards this cation thus, making the hydration more probable. The value of the hydration enthalpy also reflects the residence time of water molecules near a cation and it is inversely proportional to the water substitution rate.139 Hence, the hydration enthalpy can also be used to express the permeation of a cation through the cell membrane, as ions with a higher hydration enthalpy attract more water molecules, increasing their hydration shell and hindering the adsorption and transport of ions through membranes.139–143
The oxidation number or cation charge (z) and ionic potential (z/r) have been previously used to classify and group metal oxide NPs into toxicity classes.27 The oxidation number has been correlated with the metal oxide NP solubility and metal ion release, with lower oxidation numbers leading to more soluble oxides.27,38,144 In turn, the cation charge has been correlated with toxicity.71 The ionic potential expresses the charge potential or electronegativity of the metal cation released.27,38 It has been previously correlated with the toxicity of metal oxide NPs.145 The product of oxidation number and ionic potential, z2/r, has been successfully used by Liu et al. as a descriptor to classify 24 NPs based on their toxicity54 against RAW264.7 and BEAS-2B cells, showing that both these mechanisms attribute to NP toxicity. The hydration enthalpy computed with Latimer's equation (eqn (1)) in our work is computed using a similar term. This hydration enthalpy has been used by Kaweeteerawat et al., for the same 24 metal oxide NPs toxicity classification towards E. coli, yielding accurate results.40 In the present work, the hydration enthalpy was able to classify 45 different NPs regarding their toxicity towards various cells and organisms, with an acceptable accuracy. Based on the above discussion, the HE descriptor indirectly expresses the contribution from the NP solubility and release of metal ions, as well as their ionic potential. Moreover, it directly describes the affinity of water molecules to the released metal ions. Hence this descriptor highlights the effect of the release of metal cations and their interaction with aqueous media on the toxicity of metal and metal oxide NPs.
The toxicity of metallic and metal oxide NPs has been widely attributed to the release of metal ions.132 Metal ions released in biological media can induce oxidative stress,146–150 lead to enzyme inactivation,146,149 replace essential elements in proteins and enzymes,146,147 affect the cell membrane integrity,35,151 or even lead to DNA damage.146,152,153 The initial release of metal ions from the NPs can be expressed by the solubility of the metal/metal oxide, or by surface defects of the metal oxide structure.27 More soluble NPs lead to more released ions and thus, are more probable to be toxic towards cells. The solubility has been also correlated with the metal oxide toxicity, in previous works.27,154,155 However, the solubility of NPs per se is not possible to explain the toxicity of all the NPs, in previously reported works.33–35,70 For example, although the toxicity of CuO and ZnO NPs has been attributed to their solubility,33–35,155 MgO (ref. 33, 70 and 155) NPs have been found to be soluble, but non-toxic. In this work, the toxicity of metal ions is expressed by the hydration enthalpy, which indirectly incorporates the contributions of solubility and charge potential of the released ions, while it directly reflects the affinity of water molecules to the metal ion and thus, the hindering of the ion transport through the cell membrane.40,139
The combination of the two descriptors (Dbio and HE) used in the present study were able to classify the 45 NPs based on their toxicity, as shown in the results of Tables 1–4, towards various cells and organisms. The model built for the most populous individual cell dataset (A549 cells) was also capable of predicting the toxicity class of NPs towards the different datasets, showing the robustness of the approach. As discussed above, these two descriptors reflect different mechanisms of action, with Dbio expressing direct electron transfer between the NP and the biological media, and HE showing metal ion release, reactivity, and transport. These can result to various mechanisms of toxicity, through the increase of oxidative stress, decrease of the membrane integrity, enzyme inactivation and DNA damage, amongst others. From the model results, it is not obvious which mechanism prevails regarding the NP toxicity, as can be seen in Fig. 4. NPs that have a higher Dbio are more probable to be non-toxic even with a less negative HE, while NPs with a more negative HE are more probable to be non-toxic even for a low Dbio. Hence, the combination of the two mechanisms yields the toxicity of the corresponding NP: the NP needs to be able to undergo electron exchange with the biological pairs and to release ions that are able to permeate the cell before undergoing hydration. The above analysis of these mechanisms hints that electron transfer and metal ion release are both necessary, but not sufficient conditions for the toxicity pathway.
Identifying which mechanism prevails over the other has been the topic of discussion in relevant works35,36,156 and should take into account the morphology of the cell, and the size and shape of the NP.36 The present model yielded acceptable predictions towards various cells of different kind, leading to similar conclusions. Hence, from the results of present work, the effect of the cell morphology seems to be less significant than the two descriptors (Dbio, HE) for the toxicity classification. It should be noted, however, that this could be due to the method and concentration thresholds used to classify the NP measurements as toxic and non-toxic.
The size and shape of the NPs were not taken into account in the final models of the present work. The NPs in the datasets varied from 5–200 nm, and their inclusion as descriptors in the models did not improve the results, showing that the size effect was not that significant. Various predictive models have been built for NPs with an extended size variation, that yielded the same conclusion.27,32,35–37,40 However, this does not mean that size does not play a role in the toxicity of NPs. The size and shape of the NPs define their surface area, which in turn is the area that interacts with the biological media and cells. Hence, although size may have a less significant effect on the mechanisms qualitatively (determining which mechanisms take place), it could significantly affect them quantitatively, by affecting their kinetics.27 This has led some published works to suggest the expression of NP doses in terms surface area instead of molar or mass concentrations.157
It should be noted here that in the present work, the influence of the exposure time is not included in the modelling. This does not mean, however, that the exposure time does not have an influence on the toxicity endpoints. As the half maximal concentration endpoints used in this work are obtained by studies that test different NP concentrations under the same conditions (including exposure time), the final endpoint differs under different exposure times. However, in our analysis, the endpoints used and predicted by the developed models are not directly the half maximal concentration endpoints, but rather a toxic class for the different NPs. The toxic class is, in turn, assigned to each measurement in the dataset based on a threshold value for the half maximal concentration endpoints, based on a previous work.27
The good accuracy of the models for the prediction of the NP toxic class, despite data heterogeneity in terms of the exposure time, can be assigned in part to the classification scheme. Given this classification scheme, the influence of the exposure time may not affect the half maximal concentration endpoints enough, so that the toxic class of NPs changes from toxic to non-toxic or vice versa. However, the fact that the model can predict the toxic class (under the same classification scheme) based on the values of some physicochemical properties of the NPs, shows that these properties can significantly affect the toxicological behavior of a metal or metal oxide NP. Whether or not this particular set of descriptors can directly predict quantitative experimental results (i.e. the half maximal concentration endpoint) can be the subject of future research. In that case, however, the data should be more homogenous in terms of tested cell lines and organisms and experimental conditions, or be included as a direct or indirect input to the model.
From the above discussion, it is clear that the described mechanisms for metal oxide NP toxicity have been studied individually and have been recognized as a pathway for metal oxide toxicity. The combination of these mechanisms has also been addressed, however, in a more limited number of works and concerning the toxicity towards a specific kind of cells. Nonetheless, in this work, it is shown that such mechanistic explanation can arise when dealing with data for NP toxicity towards an extensive variety of cell lines and organisms. Furthermore, the approach can be extended to include metal NPs, by using the Fermi level energy of the metal compared to the redox potential of biological pairs (Dbio for metals) providing accurate predictions.
The accuracy of the classification models built for all the different cell lines and organisms, illuminates the possibility of classification SAR models to possibly explain NPs toxicity in a more holistic way and brings the question of whether such models could and should assist in the understanding of toxicity mechanisms. This means that besides building SAR models as predicting tools for toxicity screening, such models could also focus on their inductive capabilities, with their results being the first step towards a mechanistic explanation of toxicity pathways. Such an approach could lead to a more knowledge-based development of NP toxicity models and could, in turn, guide scientific research towards the safe-by-design NP synthesis and production. Indicatively, Feng et al., used the previously mentioned observations for the correlation between conduction bands and toxicity to design multi-component NPs, with control over their toxicity.158 Such aspects of SAR models for nanomaterials are often undermined, with studies focusing more on the predictive capabilities of such models. This leads to models often incorporating complex descriptors, which makes the extraction of scientific information challenging. The results presented in this work can serve as a basis to guide SAR model development towards incorporating descriptors describing certain toxicity pathways when possible, paving the way for the understanding of metal oxide NPs toxicity in a more global way.
Finally, it is important to note the significance of the dose metrics when the toxicity on NPs is tested towards cells and/or organisms. In most of the published works used to build the datasets presented here, the dose–response studies were mostly conducted in terms of mass concentration of NPs, with the dose–response study being conducted over a certain range of mass concentrations in each study. However, when testing multiple NPs, the molar concentration dose metric is more adequate, as it expresses the dose in terms of active species per volume.27 As different NPs have a different molar mass, a given mass concentration could mean that a very significant difference between the quantity of active species could be exposed to the cells or organisms. For example, an upper concentration of 100 mg L−1 tested for Bi2O3, Sb2O3, and ZnO are equivalent of 0.215 mM of Bi2O3, 0.342 mM of Sb2O3, and 1.235 mM of ZnO. This means that the tested concentration range of ZnO is around six times larger than the concentration range of Bi2O3, rendering measurements comparison out of discussion. Hence, when assessing or comparing the toxicity of a series of NPs, the range of concentrations tested should be adjusted in a way that expresses the same range of concentration of active species, based on the molecular weights of the species constituting the NPs. In this work, we used a threshold value for the molar half maximal concentration as a criterion for toxicity classification, based on an approach presented in a previous published scientific work.27 Based on this threshold value, we calculated the threshold value for each NP in terms of mass concentration (ESI,† file 3), for the 45 NPs investigated in this work, which could help researchers adjust the experimental range of tested concentrations and/or interpret the results of published dose–response studies for metal oxide NPs.
5. Conclusions
In this work, classification nano-SAR models are developed to investigate the possible extension of such approaches towards larger and more heterogeneous datasets to extract mechanistic information regarding the toxicity pathways of metal and metal oxide NPs. For this reason, an extensive dataset consisting of half maximal concentration endpoints for 45 metal and metal oxide NPs is built, using published data from scientific papers. The dataset consists of 935 individual measurements, which at the best of the author's knowledge, is the largest dataset consisting of such kind of measurements. Classification models are then built for the data consisting of measurements for specific cell lines, cell and organism groups, as well as the whole dataset.The different models developed showed a high accuracy (>90%), while the same combination of descriptors led to the optimal classification, for all the different subsets of data, allowing the extraction of important mechanistic knowledge from such an approach. Furthermore, the model built for the most populous of the individual cell line dataset could be extrapolated to accurately classify the complete dataset.
The analysis of the classification results showed that the energy difference between the conduction band of the NP and the redox potential of the biological pairs (Dbio) as well as the hydration enthalpy of the released metal cation (HE) were the optimal descriptors, revealing potential dominant mechanistic pathways for metal and metal oxide NP toxicity. In particular, Dbio reflects the electron transfer between biological media and NPs, affecting the reducing capability of cells thus, increasing the oxidative stress from reactive oxygen species. On the other hand, HE indirectly reflects the release and charge potential of metal ions, while it directly expresses the water affinity towards the released metal cation, which affects the transport of cations through cells' membrane.
The results presented in this work highlight the potential of nano-SAR approaches to be used not only as predictive tools, as is often the case, but also as inductive ones, as a first step towards the understanding of the mechanisms for the biological activity of NPs, in a more holistic way. Such studies can assist in the synthesis of NPs for targeted biological applications. Furthermore, the incorporation of such approaches can pave the way towards a more knowledge-based risk assessment of nanomaterials using computational tools along with experiments and guide researchers towards the synthesis of safe-by-design NPs.
Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work was funded by the EU H2020 Project ‘Development and Implementation of a Sustainable Modelling Platform for NanoInformatics’ “NanoinformaTIX” (grant number 814426). The publication of the article in OA mode was financially supported by HEAL-LinkReferences
- A. Haase, F. Klaessig, P. Nymark, K. Paul and D. Greco, EU US Roadmap Nanoinformatics 2030, 2018 Search PubMed.
- Z. Guo and L. Tan, Fundamentals and applications of nanomaterials, Artech House, 2009 Search PubMed.
- A. McWilliams, The Maturing Nanotechnology Market: Products and Applications, BBC Research Report, 2016 Search PubMed.
- C.-Z. Ning, L. Dou and P. Yang, Bandgap engineering in semiconductor alloy nanomaterials with widely tunable compositions, Nat. Rev. Mater., 2017, 2, 17070 CrossRef CAS.
- X. Hu, Y. Zhang, T. Ding, J. Liu and H. Zhao, Multifunctional Gold Nanoparticles: A Novel Nanomaterial for Various Medical Applications and Biological Activities, Front. Bioeng. Biotechnol., 2020, 8, 990 CrossRef PubMed.
- P. Barua, A. H. Khan and N. Hossain, in Nanotechnology for Electronic Applications, ed. N. M. Mubarak, S. Gopi and P. Balakrishnan, Springer Nature Singapore, Singapore, 2022, DOI:10.1007/978-981-16-6022-1_12, pp. 253–267.
- F. Christian, E. Selly, D. Adityawarman and A. Indarto, Application of nanotechnologies in the energy sector: A brief and short review, Front. Energy, 2013, 7, 6–18 CrossRef.
- S. Mitchell, R. Qin, N. Zheng and J. Pérez-Ramírez, Nanoscale engineering of catalytic materials for sustainable technologies, Nat. Nanotechnol., 2021, 16, 129–139 CrossRef CAS PubMed.
- G. P. Gakis, H. Vergnes, F. Cristiano, Y. Tison, C. Vahlas, B. Caussat, A. G. Boudouvis and E. Scheid, In situ N2-NH3 plasma pre-treatment of silicon substrate enhances the initial growth and restricts the substrate oxidation during alumina ALD, J. Appl. Phys., 2019, 126, 125305 CrossRef.
- G. P. Gakis, C. Vahlas, H. Vergnes, S. Dourdain, Y. Tison, H. Martinez, J. Bour, D. Ruch, A. G. Boudouvis, B. Caussat and E. Scheid, Investigation of the initial deposition steps and the interfacial layer of Atomic Layer Deposited (ALD) Al2O3 on Si, Appl. Surf. Sci., 2019, 492, 245–254 CrossRef CAS.
- G. P. Gakis, H. Vergnes, E. Scheid, C. Vahlas, A. G. Boudouvis and B. Caussat, Detailed investigation of the surface mechanisms and their interplay with transport phenomena in alumina atomic layer deposition from TMA and water, Chem. Eng. Sci., 2019, 195, 399–412 CrossRef CAS.
- R. Sharma, Newer Methods of Nanoparticle Synthesis: Nitroimidazole properties with Nanometal oxides in Polymer Cages as Drug-Biomarker Monitors, Nat. Preced., 2009 DOI:10.1038/npre.2009.3952.1.
- G. P. Gakis, S. Termine, A.-F. A. Trompeta, I. G. Aviziotis and C. A. Charitidis, Unraveling the mechanisms of carbon nanotube growth by chemical vapor deposition, Chem. Eng. J., 2022, 445, 136807 CrossRef CAS.
- V. Jourdain and C. Bichara, Current understanding of the growth of carbon nanotubes in catalytic chemical vapour deposition, Carbon, 2013, 58, 2–39 CrossRef CAS.
- M. A. Hamzawy, A. M. Abo-youssef, H. F. Salem and S. A. Mohammed, Antitumor activity of intratracheal inhalation of temozolomide (TMZ) loaded into gold nanoparticles and/or liposomes against urethane-induced lung cancer in BALB/c mice, Drug Delivery, 2017, 24, 599–607 CrossRef CAS PubMed.
- Y. Hao, Y. Altundal, M. Moreau, E. Sajo, R. Kumar and W. Ngwa, Potential for enhancing external beam radiotherapy for lung cancer using high-Z nanoparticles administered via inhalation, Phys. Med. Biol., 2015, 60, 7035–7043 CrossRef CAS PubMed.
- A. K. Verma and S. Perlman, Lipid nanoparticle-mRNA: another step in the fight against COVID-19, Cell Res., 2022, 32, 421–422 CrossRef CAS PubMed.
- L. Schoenmaker, D. Witzigmann, J. A. Kulkarni, R. Verbeke, G. Kersten, W. Jiskoot and D. J. A. Crommelin, mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability, Int. J. Pharm., 2021, 601, 120586 CrossRef CAS PubMed.
- T. T. H. Thi, E. J. A. Suys, J. S. Lee, D. H. Nguyen, K. D. Park and N. P. Truong, Lipid-based nanoparticles in the clinic and clinical trials: From cancer nanomedicine to COVID-19 vaccines, Vaccines, 2021, 9, 359 CrossRef CAS PubMed.
- M. J. Bessa, F. Brandão, M. Viana, J. F. Gomes, E. Monfort, F. R. Cassee, S. Fraga and J. P. Teixeira, Nanoparticle exposure and hazard in the ceramic industry: an overview of potential sources, toxicity and health effects, Environ. Res., 2020, 184, 109297 CrossRef CAS PubMed.
- M. Manigrasso, C. Protano, M. L. Astolfi, L. Massimi, P. Avino, M. Vitali and S. Canepari, Evidences of copper nanoparticle exposure in indoor environments: Long-term assessment, high-resolution field emission scanning electron microscopy evaluation, in silico respiratory dosimetry study and possible health implications, Sci. Total Environ., 2019, 653, 1192–1203 CrossRef CAS PubMed.
- S. Sonwani, S. Madaan, J. Arora, S. Suryanarayan, D. Rangra, N. Mongia, T. Vats and P. Saxena, Inhalation Exposure to Atmospheric Nanoparticles and Its Associated Impacts on Human Health: A Review, Front. Sustain. Cities, 2021, 3, 690444 CrossRef.
- Y. Nazarenko, T. W. Han, P. J. Lioy and G. Mainelis, Potential for exposure to engineered nanoparticles from nanotechnology-based consumer spray products, J. Exposure Sci. Environ. Epidemiol., 2011, 21, 515–528 CrossRef CAS PubMed.
- J. Blasco, I. Corsi and V. Matranga, Particles in the oceans: Implication for a safe marine environment, Mar. Environ. Res., 2015, 111, 1–4 CrossRef CAS PubMed.
- V. Kumar, N. Sharma and S. S. Maitra, In vitro and in vivo toxicity assessment of nanoparticles, Int. Nano Lett., 2017, 7, 243–256 CrossRef CAS.
- D. T. Savage, J. Z. Hilt and T. D. Dziubla, in Nanotoxicity: Methods and Protocols, ed. Q. Zhang, Springer New York, New York, NY, 2019, DOI:10.1007/978-1-4939-8916-4_1, pp. 1–29.
- F. C. Simeone and A. L. Costa, Assessment of cytotoxicity of metal oxide nanoparticles on the basis of fundamental physical–chemical parameters: a robust approach to grouping, Environ. Sci.: Nano, 2019, 6, 3102–3112 RSC.
- I. Xiarchos, A. K. Morozinis, P. Kavouras and C. A. Charitidis, Nanocharacterization, Materials Modeling, and Research Integrity as Enablers of Sound Risk Assessment: Designing Responsible Nanotechnology, Small, 2020, 16, 2001590 CrossRef CAS PubMed.
- M. Kotzabasaki, I. Sotiropoulos, C. Charitidis and H. Sarimveis, Machine learning methods for multi-walled carbon nanotubes (MWCNT) genotoxicity prediction, Nanoscale Adv., 2021, 3, 3167–3176 RSC.
- A. A. Buglak, A. V. Zherdev and B. B. Dzantiev, Nano-(Q)SAR for Cytotoxicity Prediction of Engineered Nanomaterials, Molecules, 2019, 24, 4537 CrossRef CAS PubMed.
- L. Lamon, D. Asturiol, A. Vilchez, R. Ruperez-Illescas, J. Cabellos, A. Richarz and A. Worth, Computational models for the assessment of manufactured nanomaterials: Development of model reporting standards and mapping of the model landscape, Comput. Toxicol., 2019, 9, 143–151 CrossRef CAS PubMed.
- T. Puzyn, B. Rasulev, A. Gajewicz, X. Hu, T. P. Dasari, A. Michalkova, H. M. Hwang, A. Toropov, D. Leszczynska and J. Leszczynski, Using nano-QSAR to predict the cytotoxicity of metal oxide nanoparticles, Nat. Nanotechnol., 2011, 6, 175–178 CrossRef CAS PubMed.
- V. Aruoja, S. Pokhrel, M. Sihtmäe, M. Mortimer, L. Mädler and A. Kahru, Toxicity of 12 metal-based nanoparticles to algae, bacteria and protozoa, Environ. Sci.: Nano, 2015, 2, 630–644 RSC.
- A. Ivask, T. Titma, M. Visnapuu, H. Vija, A. Kakinen, M. Sihtmae, S. Pokhrel, L. Madler, M. Heinlaan, V. Kisand, R. Shimmo and A. Kahru, Toxicity of 11 Metal Oxide Nanoparticles to Three Mammalian Cell Types <i>In Vitro</i>, Curr. Top. Med. Chem., 2015, 15, 1914–1929 CrossRef CAS PubMed.
- H. Zhang, Z. Ji, T. Xia, H. Meng, C. Low-Kam, R. Liu, S. Pokhrel, S. Lin, X. Wang, Y.-P. Liao, M. Wang, L. Li, R. Rallo, R. Damoiseaux, D. Telesca, L. Mädler, Y. Cohen, J. I. Zink and A. E. Nel, Use of Metal Oxide Nanoparticle Band Gap To Develop a Predictive Paradigm for Oxidative Stress and Acute Pulmonary Inflammation, ACS Nano, 2012, 6, 4349–4368 CrossRef CAS PubMed.
- A. Gajewicz, N. Schaeublin, B. Rasulev, S. Hussain, D. Leszczynska, T. Puzyn and J. Leszczynski, Towards understanding mechanisms governing cytotoxicity of metal oxides nanoparticles: Hints from nano-QSAR studies, Nanotoxicology, 2015, 9, 313–325 CrossRef CAS PubMed.
- K. Pathakoti, M.-J. Huang, J. D. Watts, X. He and H.-M. Hwang, Using experimental data of Escherichia coli to develop a QSAR model for predicting the photo-induced cytotoxicity of metal oxide nanoparticles, J. Photochem. Photobiol., B, 2014, 130, 234–240 CrossRef CAS PubMed.
- J. Li, C. Wang, L. Yue, F. Chen, X. Cao and Z. Wang, Nano-QSAR modeling for predicting the cytotoxicity of metallic and metal oxide nanoparticles: A review, Ecotoxicol. Environ. Saf., 2022, 243, 113955 CrossRef CAS PubMed.
- E. Burello and A. Worth, in Towards Efficient Designing of Safe Nanomaterials: Innovative Merge of Computational Approaches and Experimental Techniques, The Royal Society of Chemistry, 2012, pp. 257–283, 10.1039/9781849735476-00257.
- C. Kaweeteerawat, A. Ivask, R. Liu, H. Zhang, C. H. Chang, C. Low-Kam, H. Fischer, Z. Ji, S. Pokhrel, Y. Cohen, D. Telesca, J. Zink, L. Madler, P. A. Holden, A. Nel and H. Godwin, Toxicity of metal oxide nanoparticles in Escherichia coli correlates with conduction band and hydration energies, Environ. Sci. Technol., 2015, 49, 1105–1112 CrossRef CAS PubMed.
- J. Cao, Y. Pan, Y. Jiang, R. Qi, B. Yuan, Z. Jia, J. Jiang and Q. Wang, Computer-aided nanotoxicology: risk assessment of metal oxide nanoparticles via nano-QSAR, Green Chem., 2020, 22, 3512–3521 RSC.
- S. Kar, K. Pathakoti, P. B. Tchounwou, D. Leszczynska and J. Leszczynski, Evaluating the cytotoxicity of a large pool of metal oxide nanoparticles to Escherichia coli: Mechanistic understanding through In Vitro and In Silico studies, Chemosphere, 2021, 264, 128428 CrossRef CAS PubMed.
- V. V. Kleandrova, F. Luan, H. González-Díaz, J. M. Ruso, A. Melo, A. Speck-Planche and M. N. D. S. Cordeiro, Computational ecotoxicology: Simultaneous prediction of ecotoxic effects of nanoparticles under different experimental conditions, Environ. Int., 2014, 73, 288–294 CrossRef CAS PubMed.
- F. Luan, V. V. Kleandrova, H. González-Díaz, J. M. Ruso, A. Melo, A. Speck-Planche and M. N. D. S. Cordeiro, Computer-aided nanotoxicology: assessing cytotoxicity of nanoparticles under diverse experimental conditions by using a novel QSTR-perturbation approach, Nanoscale, 2014, 6, 10623–10630 RSC.
- V. V. Kleandrova, F. Luan, H. González-Díaz, J. M. Ruso, A. Speck-Planche and M. N. D. S. Cordeiro, Computational Tool for Risk Assessment of Nanomaterials: Novel QSTR-Perturbation Model for Simultaneous Prediction of Ecotoxicity and Cytotoxicity of Uncoated and Coated Nanoparticles under Multiple Experimental Conditions, Environ. Sci. Technol., 2014, 48, 14686–14694 CrossRef CAS PubMed.
- R. Concu, V. V. Kleandrova, A. Speck-Planche and M. N. D. S. Cordeiro, Probing the toxicity of nanoparticles: a unified in silico machine learning model based on perturbation theory, Nanotoxicology, 2017, 11, 891–906 CrossRef CAS PubMed.
- M. K. Ha, T. X. Trinh, J. S. Choi, D. Maulina, H. G. Byun and T. H. Yoon, Toxicity Classification of Oxide Nanomaterials: Effects of Data Gap Filling and PChem Score-based Screening Approaches, Sci. Rep., 2018, 8, 3141 CrossRef PubMed.
- OECD, Guidance Document on the Validation of (Quantitative) Structure-Activity Relationship [(Q)SAR] Models, 2014 Search PubMed.
- A. P. Toropova, A. A. Toropov, R. Rallo, D. Leszczynska and J. Leszczynski, Optimal descriptor as a translator of eclectic data into prediction of cytotoxicity for metal oxide nanoparticles under different conditions, Ecotoxicol. Environ. Saf., 2015, 112, 39–45 CrossRef CAS PubMed.
- A. P. Toropova, A. A. Toropov, S. Manganelli, C. Leone, D. Baderna, E. Benfenati and R. Fanelli, Quasi-SMILES as a tool to utilize eclectic data for predicting the behavior of nanomaterials, NanoImpact, 2016, 1, 60–64 CrossRef.
- K. P. Singh and S. Gupta, Nano-QSAR modeling for predicting biological activity of diverse nanomaterials, RSC Adv., 2014, 4, 13215–13230 RSC.
- A. A. Toropov, A. P. Toropova, E. Benfenati, G. Gini, T. Puzyn, D. Leszczynska and J. Leszczynski, Novel application of the CORAL software to model cytotoxicity of metal oxide nanoparticles to bacteria Escherichia coli, Chemosphere, 2012, 89, 1098–1102 CrossRef CAS PubMed.
- E. Burello and A. P. Worth, A theoretical framework for predicting the oxidative stress potential of oxide nanoparticles, Nanotoxicology, 2011, 5, 228–235 CrossRef CAS PubMed.
- R. Liu, H. Y. Zhang, Z. X. Ji, R. Rallo, T. Xia, C. H. Chang, A. Nel and Y. Cohen, Development of structure–activity relationship for metal oxide nanoparticles, Nanoscale, 2013, 5, 5644–5653 RSC.
- M. Abudayyak, E. Öztaş, M. Arici and G. Özhan, Investigation of the toxicity of bismuth oxide nanoparticles in various cell lines, Chemosphere, 2017, 169, 117–123 CrossRef CAS PubMed.
- L. Otero-González, C. García-Saucedo, J. A. Field and R. Sierra-Álvarez, Toxicity of TiO2, ZrO2, Fe0, Fe2O3, and Mn2O3 nanoparticles to the yeast, Saccharomyces cerevisiae, Chemosphere, 2013, 93, 1201–1206 CrossRef PubMed.
- S. Areecheewakul, A. Adamcakova-Dodd, B. E. Givens, B. R. Steines, Y. Wang, D. K. Meyerholz, N. J. Parizek, R. Altmaier, E. Haque, P. T. O'Shaughnessy, A. K. Salem and P. S. Thorne, Toxicity assessment of metal oxide nanomaterials using in vitro screening and murine acute inhalation studies, NanoImpact, 2020, 18, 100214 CrossRef PubMed.
- B. Balusamy, B. E. Taştan, S. F. Ergen, T. Uyar and T. Tekinay, Toxicity of lanthanum oxide (La2O3) nanoparticles in aquatic environments, Environ. Sci.: Processes Impacts, 2015, 17, 1265–1270 RSC.
- O. Bar-Ilan, R. M. Albrecht, V. E. Fako and D. Y. Furgeson, Toxicity Assessments of Multisized Gold and Silver Nanoparticles in Zebrafish Embryos, Small, 2009, 5, 1897–1910 CrossRef CAS PubMed.
- I. Blinova, A. Ivask, M. Heinlaan, M. Mortimer and A. Kahru, Ecotoxicity of nanoparticles of CuO and ZnO in natural water, Environ. Pollut., 2010, 158, 41–47 CrossRef CAS PubMed.
- O. M. Bondarenko, M. Heinlaan, M. Sihtmäe, A. Ivask, I. Kurvet, E. Joonas, A. Jemec, M. Mannerström, T. Heinonen, R. Rekulapelly, S. Singh, J. Zou, I. Pyykkö, D. Drobne and A. Kahru, Multilaboratory evaluation of 15 bioassays for (eco)toxicity screening and hazard ranking of engineered nanomaterials: FP7 project NANOVALID, Nanotoxicology, 2016, 10, 1229–1242 CrossRef CAS PubMed.
- K. K. Comfort, E. I. Maurer and S. M. Hussain, Slow release of ions from internalized silver nanoparticles modifies the epidermal growth factor signaling response, Colloids Surf., B, 2014, 123, 136–142 CrossRef CAS PubMed.
- T. P. Dasari, K. Pathakoti and H.-M. Hwang, Determination of the mechanism of photoinduced toxicity of selected metal oxide nanoparticles (ZnO, CuO, Co3O4 and TiO2) to E. coli bacteria, J. Environ. Sci., 2013, 25, 882–888 CrossRef CAS PubMed.
- P. Dua, K. Chaudhari, H. Lee, N. Chaudhari, W. Sun, J.-S. Hong, J.-S. Yu, S. Kim and D.-K. Lee, Gene Expression Changes of Oxide Nanoparticles Evaluation of Toxicity and Gene Expression Changes Triggered by Oxide Nanoparticles, Bull. Korean Chem. Soc., 2011, 32326, 2051–2057 CrossRef.
- C. García-Saucedo, J. A. Field, L. Otero-Gonzalez and R. Sierra-Álvarez, Low toxicity of HfO2, SiO2, Al2O3 and CeO2 nanoparticles to the yeast, Saccharomyces cerevisiae, J. Hazard. Mater., 2011, 192, 1572–1579 CrossRef PubMed.
- M. Geppert, L. Sigg and K. Schirmer, Toxicity and translocation of Ag, CuO, ZnO and TiO2 nanoparticles upon exposure to fish intestinal epithelial cells, Environ. Sci.: Nano, 2021, 8, 2249–2260 RSC.
- R. J. Griffitt, J. Luo, J. Gao, J.-C. Bonzongo and D. S. Barber, Effects of particle composition and species on toxicity of metallic nanomaterials in aquatic organisms, Environ. Toxicol. Chem., 2008, 27, 1972–1978 CrossRef CAS PubMed.
- M. Heinlaan, A. Ivask, I. Blinova, H.-C. Dubourguier and A. Kahru, Toxicity of nanosized and bulk ZnO, CuO and TiO2 to bacteria Vibrio fischeri and crustaceans Daphnia magna and Thamnocephalus platyurus, Chemosphere, 2008, 71, 1308–1316 CrossRef CAS PubMed.
- S. M. Hoheisel, S. Diamond and D. Mount, Comparison of nanosilver and ionic silver toxicity in Daphnia magna and Pimephales promelas, Environ. Toxicol. Chem., 2012, 31, 2557–2563 CrossRef CAS PubMed.
- M. Horie, K. Fujita, H. Kato, S. Endoh, K. Nishio, L. K. Komaba, A. Nakamura, A. Miyauchi, S. Kinugasa, Y. Hagihara, E. Niki, Y. Yoshida and H. Iwahashi, Association of the physical and chemical properties and the cytotoxicity of metal oxide nanoparticles: metal ion release, adsorption ability and specific surface area, Metallomics, 2012, 4, 350–360 CrossRef CAS PubMed.
- X. Hu, S. Cook, P. Wang and H.-M. Hwang, In vitro evaluation of cytotoxicity of engineered metal oxide nanoparticles, Sci. Total Environ., 2009, 407, 3070–3072 CrossRef CAS PubMed.
- Y. W. Huang, C. H. Wu and R. S. Aronstam, Toxicity of Transition Metal Oxide Nanoparticles: Recent Insights from in vitro Studies, Materials, 2010, 3, 4842–4859 CrossRef CAS PubMed.
- A. Ivask, O. Bondarenko, N. Jepihhina and A. Kahru, Profiling of the reactive oxygen species-related ecotoxicity of CuO, ZnO, TiO2, silver and fullerene nanoparticles using a set of recombinant luminescent Escherichia coli strains: differentiating the impact of particles and solubilised metals, Anal. Bioanal. Chem., 2010, 398, 701–716 CrossRef CAS PubMed.
- H. A. Jeng and J. Swanson, Toxicity of Metal Oxide Nanoparticles in Mammalian Cells, J. Environ. Sci. Health, Part A: Toxic/Hazard. Subst. Environ. Eng., 2006, 41, 2699–2711 CrossRef CAS PubMed.
- J. Jeong, S.-H. Kim, S. Lee, D.-K. Lee, Y. Han, S. Jeon and W.-S. Cho, Differential Contribution of Constituent Metal Ions to the Cytotoxic Effects of Fast-Dissolving Metal-Oxide Nanoparticles, Front. Pharmacol., 2018, 9, 15 CrossRef PubMed.
- H. L. Karlsson, P. Cronholm, J. Gustafsson and L. Möller, Copper Oxide Nanoparticles Are Highly Toxic: A Comparison between Metal Oxide Nanoparticles and Carbon Nanotubes, Chem. Res. Toxicol., 2008, 21, 1726–1732 Search PubMed.
- K. Kasemets, A. Ivask, H.-C. Dubourguier and A. Kahru, Toxicity of nanoparticles of ZnO, CuO and TiO2 to yeast Saccharomyces cerevisiae, Toxicol. In Vitro, 2009, 23, 1116–1122 CrossRef CAS PubMed.
- A. Katsumiti, I. Arostegui, M. Oron, D. Gilliland, E. Valsami-Jones and M. P. Cajaraville, Cytotoxicity of Au, ZnO and SiO2 NPs using in vitro assays with mussel hemocytes and gill cells: Relevance of size, shape and additives, Nanotoxicology, 2016, 10, 185–193 CAS.
- Y.-S. Kim, K. K. Kim, S.-M. Shin, S. M. Park and S. S. Hah, Comparative Toxicity Studies of Ultra-Pure Ag, Au, Co, and Cu Nanoparticles Generated by Laser Ablation in Biocompatible Aqueous Solution, Bull. Korean Chem. Soc., 2012, 33, 3265–3268 CrossRef CAS.
- K.-S. Ko and I. C. Kong, Toxic effects of nanoparticles on bioluminescence activity, seed germination, and gene mutation, Appl. Microbiol. Biotechnol., 2014, 98, 3295–3303 CrossRef CAS PubMed.
- I. C. Kong, K.-S. Ko and D.-C. Koh, Comparisons of the Effect of Different Metal Oxide Nanoparticles on the Root and Shoot Growth under Shaking and Non-Shaking Incubation, Different Plants, and Binary Mixture Conditions, Nanomaterials, 2021, 11, 1653 CrossRef CAS PubMed.
- J. A. Kovriznych, R. Sotnikova, D. Zeljenkova, E. Rollerova, E. Szabova and S. Wimmerova, Acute toxicity of 31 different nanoparticles to zebrafish (Danio rerio) tested in adulthood and in early life stages - comparative study, Interdiscip. Toxicol., 2013, 6, 67–73 Search PubMed.
- J. M. Lacave, A. Retuerto, U. Vicario-Parés, D. Gilliland, M. Oron, M. P. Cajaraville and A. Orbea, Effects of metal-bearing nanoparticles (Ag, Au, CdS, ZnO, SiO2) on developing zebrafish embryos, Nanotechnology, 2016, 27, 325102 CrossRef PubMed.
- S. Lanone, F. Rogerieux, J. Geys, A. Dupont, E. Maillot-Marechal, J. Boczkowski, G. Lacroix and P. Hoet, Comparative toxicity of 24 manufactured nanoparticles in human alveolar epithelial and macrophage cell lines, Part. Fibre Toxicol., 2009, 6, 14 CrossRef PubMed.
- K. Li, Y. Chen, W. Zhang, Z. Pu, L. Jiang and Y. Chen, Surface Interactions Affect the Toxicity of Engineered Metal Oxide Nanoparticles toward Paramecium, Chem. Res. Toxicol., 2012, 25, 1675–1681 Search PubMed.
- L. Cheol-Hong, Toxicity of Two Different Sized Lanthanum Oxides in Cultured Cells and Sprague-Dawley Rats, Toxicol. Res., 2015, 31, 181–189 CrossRef PubMed.
- D. Lin and B. Xing, Phytotoxicity of nanoparticles: Inhibition of seed germination and root growth, Environ. Pollut., 2007, 150, 243–250 CrossRef CAS PubMed.
- W. Liu, Y. Wu, C. Wang, H. C. Li, T. Wang, C. Y. Liao, L. Cui, Q. F. Zhou, B. Yan and G. B. Jiang, Impact of silver nanoparticles on human cells: Effect of particle size, Nanotoxicology, 2010, 4, 319–330 CrossRef CAS PubMed.
- T. Lozano, M. Rey, E. Rojas, S. Moya, J. Fleddermann, I. Estrela-Lopis, E. Donath, B. Wang, Z. Mao, C. Gao and Á. González-Fernández, Cytotoxicity effects of metal oxide nanoparticles in human tumor cell lines, J. Phys.: Conf. Ser., 2011, 304, 012046 CrossRef.
- Y. Ma, L. Kuang, X. He, W. Bai, Y. Ding, Z. Zhang, Y. Zhao and Z. Chai, Effects of rare earth oxide nanoparticles on root elongation of plants, Chemosphere, 2010, 78, 273–279 CrossRef CAS PubMed.
- Y. Ma, X. He, P. Zhang, Z. Zhang, Z. Guo, R. Tai, Z. Xu, L. Zhang, Y. Ding, Y. Zhao and Z. Chai, Phytotoxicity and biotransformation of La2O3 nanoparticles in a terrestrial plant cucumber (Cucumis sativus), Nanotoxicology, 2011, 5, 743–753 CrossRef CAS PubMed.
- R. Eshaghi Malekshah, B. Fahimirad, M. Aallaei and A. Khaleghian, Synthesis and toxicity assessment of Fe3O4 NPs grafted by ∼ NH2-Schiff base as anticancer drug: modeling and proposed molecular mechanism through docking and molecular dynamic simulation, Drug Delivery, 2020, 27, 1201–1217 CrossRef CAS PubMed.
- M. Mauro, M. Crosera, M. Pelin, C. Florio, F. Bellomo, G. Adami, P. Apostoli, G. De Palma, M. Bovenzi, M. Campanini and F. L. Filon, Cobalt Oxide Nanoparticles: Behavior towards Intact and Impaired Human Skin and Keratinocytes Toxicity, Int. J. Environ. Res. Public Health, 2015, 12, 8263–8280 CrossRef CAS PubMed.
- M. Mortimer, K. Kasemets, M. Heinlaan, I. Kurvet and A. Kahru, High throughput kinetic Vibrio fischeri bioluminescence inhibition assay for study of toxic effects of nanoparticles, Toxicol. In Vitro, 2008, 22, 1412–1417 CrossRef CAS PubMed.
- H. Negi, P. Rathinavelu Saravanan, T. Agarwal, M. Ghulam Haider Zaidi and R. Goel, In vitro assessment of Ag<sub>2</sub>O nanoparticles toxicity against Gram-positive and Gram-negative bacteria, J. Gen. Appl. Microbiol., 2013, 59, 83–88 CrossRef CAS PubMed.
- C. H. Nguyen, J. A. Field and R. Sierra-Alvarez, Microbial toxicity of gallium- and indium-based oxide and arsenide nanoparticles, J. Environ. Sci. Health, Part A: Toxic/Hazard. Subst. Environ. Eng., 2020, 55, 168–178 CrossRef CAS PubMed.
- Q. Peng, D. Huo, H. Li, B. Zhang, Y. Li, A. Liang, H. Wang, Q. Yu and M. Li, ROS-independent toxicity of Fe3O4 nanoparticles to yeast cells: Involvement of mitochondrial dysfunction, Chem.-Biol. Interact., 2018, 287, 20–26 CrossRef CAS PubMed.
- J. M. Seiffert, M.-O. Baradez, V. Nischwitz, T. Lekishvili, H. Goenaga-Infante and D. Marshall, Dynamic Monitoring of Metal Oxide Nanoparticle Toxicity by Label Free Impedance Sensing, Chem. Res. Toxicol., 2012, 25, 140–152 Search PubMed.
- A. Sharan and S. Nara, Exposure of synthesized Co3O4 nanoparticles to Chlorella minutissima: An ecotoxic evaluation in freshwater microalgae, Aquat. Toxicol., 2020, 224, 105498 CrossRef CAS PubMed.
- D. Singh and A. Kumar, Effects of Nano Silver Oxide and Silver Ions on Growth of Vigna radiata, Bull. Environ. Contam. Toxicol., 2015, 95, 379–384 CrossRef CAS PubMed.
- T. Sovová, V. Kocí and L. Kochánková, Ecotoxicity of nano and bulk forms of metal oxides, 2009 Search PubMed.
- T. Titma, R. Shimmo, J. Siigur and A. Kahru, Toxicity of antimony, copper, cobalt, manganese, titanium and zinc oxide nanoparticles for the alveolar and intestinal epithelial barrier cells in vitro, Cytotechnology, 2016, 68, 2363–2377 CrossRef CAS PubMed.
- A. Ud-Daula, G. Pfister and K.-W. Schramm, Method for toxicity test of titanium dioxide nanoparticles in ciliate protozoan Tetrahymena, J. Environ. Sci. Health, Part A: Toxic/Hazard. Subst. Environ. Eng., 2013, 48, 1343–1348 CrossRef CAS PubMed.
- L. C. Wehmas, C. Anders, J. Chess, A. Punnoose, C. B. Pereira, J. A. Greenwood and R. L. Tanguay, Comparative metal oxide nanoparticle toxicity using embryonic zebrafish, Toxicol. Rep., 2015, 2, 702–715 CrossRef CAS PubMed.
- J. M. Wörle-Knirsch, K. Kern, C. Schleh, C. Adelhelm, C. Feldmann and H. F. Krug, Nanoparticulate Vanadium Oxide Potentiated Vanadium Toxicity in Human Lung Cells, Environ. Sci. Technol., 2007, 41, 331–336 CrossRef PubMed.
- S. G. Wu, L. Huang, J. Head, D. R. Chen, I. C. Kong and Y. Tang, Phytotoxicity of Metal Oxide Nanoparticles is Related to Both Dissolved Metals Ions and Adsorption of Particles on Seed Surfaces, J. Pet. Environ. Biotechnol., 2012, 3, 1000126 Search PubMed.
- X. Chen, C. Zhang, L. Tan and J. Wang, Toxicity of Co nanoparticles on three species of marine microalgae, Environ. Pollut., 2018, 236, 454–461 CrossRef CAS PubMed.
- Y. Xie, D. Liu, C. Cai, X. Chen, Y. Zhou, L. Wu, Y. Sun, H. Dai, X. Kong and P. Liu, Size-dependent cytotoxicity of Fe3O4 nanoparticles induced by biphasic regulation of oxidative stress in different human hepatoma cells, Int. J. Nanomed., 2016, 11, 3557–3570 CrossRef CAS PubMed.
- Y. Zhai, E. R. Hunting, M. Wouterse, W. J. G. M. Peijnenburg and M. G. Vijver, Importance of exposure dynamics of metal-based nano-ZnO, -Cu and -Pb governing the metabolic potential of soil bacterial communities, Ecotoxicol. Environ. Saf., 2017, 145, 349–358 CrossRef CAS PubMed.
- H. Zhang, J. Shi, Y. Su, W. Li, K. J. Wilkinson and B. Xie, Acute toxicity evaluation of nanoparticles mixtures using luminescent bacteria, Environ. Monit. Assess., 2020, 192, 484 CrossRef CAS PubMed.
- P. Zhang, Y. Ma, Z. Zhang, X. He, Z. Guo, R. Tai, Y. Ding, Y. Zhao and Z. Chai, Comparative toxicity of nanoparticulate/bulk Yb2O3 and YbCl3 to cucumber (Cucumis sativus), Environ. Sci. Technol., 2012, 46, 1834–1841 CrossRef CAS PubMed.
- W. Zhang, X. Liu, S. Bao, B. Xiao and T. Fang, Evaluation of nano-specific toxicity of zinc oxide, copper oxide, and silver nanoparticles through toxic ratio, J. Nanopart. Res., 2016, 18, 372 CrossRef.
- X. Zhu, Y. Chang and Y. Chen, Toxicity and bioaccumulation of TiO2 nanoparticle aggregates in Daphnia magna, Chemosphere, 2010, 78, 209–215 CrossRef CAS PubMed.
- X. Zhu, L. Zhu, Z. Duan, R. Qi, Y. Li and Y. Lang, Comparative toxicity of several metal oxide nanoparticle aqueous suspensions to Zebrafish (Danio rerio) early developmental stage, J. Environ. Sci. Health, Part A: Toxic/Hazard. Subst. Environ. Eng., 2008, 43, 278–284 CrossRef CAS PubMed.
- X. Zhu, L. Zhu, Y. Chen and S. Tian, Acute toxicities of six manufactured nanomaterial suspensions to Daphnia magna, J. Nanopart. Res., 2009, 11, 67–75 CrossRef CAS.
- R. D. Shannon, Revised effective ionic radii and systematic studies of interatomic distances in halides and chalcogenides, Acta Crystallogr., Sect. A: Cryst. Phys., Diffr., Theor. Gen. Crystallogr., 1976, 32, 751–767 CrossRef.
- J. Portier, G. Campet, J. Etourneau and B. Tanguy, A simple model for the estimation of electronegativities of cations in different electronic states and coordinations, J. Alloys Compd., 1994, 209, 285–289 CrossRef CAS.
- J. Portier, H. S. Hilal, I. Saadeddin, S. J. Hwang, M. A. Subramanian and G. Campet, Thermodynamic correlations and band gap calculations in metal oxides, Prog. Solid State Chem., 2004, 32, 207–217 CrossRef CAS.
- M. Kosmulski, Isoelectric points and points of zero charge of metal (hydr)oxides: 50years after Parks' review, Adv. Colloid Interface Sci., 2016, 238, 1–61 CrossRef CAS PubMed.
- C. Tantardini and A. R. Oganov, Thermochemical electronegativities of the elements, Nat. Commun., 2021, 12, 2087 CrossRef CAS PubMed.
- M. Auffan, J. Rose, M. R. Wiesner and J. Y. Bottero, Chemical stability of metallic nanoparticles: a parameter controlling their potential cellular toxicity in vitro, Environ. Pollut., 2009, 157, 1127–1133 CrossRef CAS PubMed.
- G. S. Plumlee, S. A. Morman and T. L. Ziegler, The Toxicological Geochemistry of Earth Materials: An Overview of Processes and the Interdisciplinary Methods Used to Understand Them, Rev. Mineral. Geochem., 2006, 64, 5–57 CrossRef CAS.
- P. Ambure, R. B. Aher, A. Gajewicz, T. Puzyn and K. Roy, “NanoBRIDGES” software: Open access tools to perform QSAR and nano-QSAR modeling, Chemom. Intell. Lab. Syst., 2015, 147, 1–13 CrossRef CAS.
- I. Guyon and A. Elisseeff, An introduction to variable and feature selection, J. Mach. Learn. Res., 2003, 3, 1157–1182 Search PubMed.
- M. Eklund, U. Norinder, S. Boyer and L. Carlsson, Choosing Feature Selection and Learning Algorithms in QSAR, J. Chem. Inf. Model., 2014, 54, 837–843 CrossRef CAS PubMed.
- P. Hanchuan, L. Fuhui and C. Ding, Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy, IEEE Trans. Pattern Anal. Mach. Intell., 2005, 27, 1226–1238 Search PubMed.
- Y. Li, Z. Dai, D. Cao, F. Luo, Y. Chen and Z. Yuan, Chi-MIC-share: a new feature selection algorithm for quantitative structure–activity relationship models, RSC Adv., 2020, 10, 19852–19860 RSC.
- F. Sahigara, K. Mansouri, D. Ballabio, A. Mauri, V. Consonni and R. Todeschini, Comparison of Different Approaches to Define the Applicability Domain of QSAR Models, Molecules, 2012, 17, 4791–4810 CrossRef CAS PubMed.
- F. Sahigara, D. Ballabio, R. Todeschini and V. Consonni, Assessing the Validity of QSARs for Ready Biodegradability of Chemicals: An Applicability Domain Perspective, Curr. Comput.-Aided Drug Des., 2014, 10, 137–147 CrossRef CAS PubMed.
- A. Rácz, D. Bajusz and K. Héberger, Intercorrelation Limits in Molecular Descriptor Preselection for QSAR/QSPR, Mol. Inf., 2019, 38, 1800154 CrossRef PubMed.
- J. M. Worle-Knirsch, K. Kern, C. Schleh, C. Adelhelm, C. Feldmann and H. F. Krug, Nanoparticulate vanadium oxide potentiated vanadium toxicity in human lung cells, Environ. Sci. Technol., 2007, 41, 331–336 CrossRef PubMed.
- A. B. Djurisic, Y. H. Leung, A. M. Ng, X. Y. Xu, P. K. Lee, N. Degger and R. S. Wu, Toxicity of metal oxide nanoparticles: mechanisms, characterization, and avoiding experimental artefacts, Small, 2015, 11, 26–44 CrossRef CAS PubMed.
- Y. Li, W. Zhang, J. Niu and Y. Chen, Mechanism of Photogenerated Reactive Oxygen Species and Correlation with the Antibacterial Properties of Engineered Metal-Oxide Nanoparticles, ACS Nano, 2012, 6, 5164–5173 CrossRef CAS PubMed.
- M. Premanathan, K. Karthikeyan, K. Jeyasubramanian and G. Manivannan, Selective toxicity of ZnO nanoparticles toward Gram-positive bacteria and cancer cells by apoptosis through lipid peroxidation, Nanomedicine, 2011, 7, 184–192 CrossRef CAS PubMed.
- G. Applerot, A. Lipovsky, R. Dror, N. Perkas, Y. Nitzan, R. Lubart and A. Gedanken, Enhanced Antibacterial Activity of Nanocrystalline ZnO Due to Increased ROS-Mediated Cell Injury, Adv. Funct. Mater., 2009, 19, 842–852 CrossRef CAS.
- T. Xia, M. Kovochich, J. Brant, M. Hotze, J. Sempf, T. Oberley, C. Sioutas, J. I. Yeh, M. R. Wiesner and A. E. Nel, Comparison of the Abilities of Ambient and Manufactured Nanoparticles To Induce Cellular Toxicity According to an Oxidative Stress Paradigm, Nano Lett., 2006, 6, 1794–1807 CrossRef CAS PubMed.
- T. C. Long, N. Saleh, R. D. Tilton, G. V. Lowry and B. Veronesi, Titanium Dioxide (P25) Produces Reactive Oxygen Species in Immortalized Brain Microglia (BV2): Implications for Nanoparticle Neurotoxicity, Environ. Sci. Technol., 2006, 40, 4346–4352 CrossRef CAS PubMed.
- A. L. Neal, What can be inferred from bacterium–nanoparticle interactions about the potential consequences of environmental exposure to nanoparticles?, Ecotoxicology, 2008, 17, 362 CrossRef CAS PubMed.
- F. Thevenod, Catch me if you can! Novel aspects of cadmium transport in mammalian cells, BioMetals, 2010, 23, 857–875 CrossRef CAS PubMed.
- S. Sugiharto, T. M. Lewis, A. J. Moorhouse, P. R. Schofield and P. H. Barry, Anion-cation permeability correlates with hydrated counterion size in glycine receptor channels, Biophys. J., 2008, 95, 4698–4715 CrossRef CAS PubMed.
- R. Epsztein, E. Shaulsky, M. Qin and M. Elimelech, Activation behavior for ion permeation in ion-exchange membranes: Role of ion dehydration in selective transport, J. Membr. Sci., 2019, 580, 316–326 CrossRef CAS.
- S. Adapa and A. Malani, Role of hydration energy and co-ions association on monovalent and divalent cations adsorption at mica-aqueous interface, Sci. Rep., 2018, 8, 12198 CrossRef PubMed.
- Z. Qu and H. C. Hartzell, Anion permeation in Ca(2+)-activated Cl(-) channels, J. Gen. Physiol., 2000, 116, 825–844 CrossRef CAS PubMed.
- G. E. Brown, V. E. Henrich, W. H. Casey, D. L. Clark, C. Eggleston, A. Felmy, D. W. Goodman, M. Grätzel, G. Maciel, M. I. McCarthy, K. H. Nealson, D. A. Sverjensky, M. F. Toney and J. M. Zachara, Metal Oxide Surfaces and Their Interactions with Aqueous Solutions and Microbial Organisms, Chem. Rev., 1999, 99, 77–174 CrossRef CAS PubMed.
- Y. Mu, F. Wu, Q. Zhao, R. Ji, Y. Qie, Y. Zhou, Y. Hu, C. Pang, D. Hristozov, J. P. Giesy and B. Xing, Predicting toxic potencies of metal oxide nanoparticles by means of nano-QSARs, Nanotoxicology, 2016, 10, 1207–1214 CrossRef CAS PubMed.
- M. Balali-Mood, K. Naseri, Z. Tahergorabi, M. R. Khazdair and M. Sadeghi, Toxic Mechanisms of Five Heavy Metals: Mercury, Lead, Chromium, Cadmium, and Arsenic, Front. Pharmacol., 2021, 12, 643972 CrossRef CAS PubMed.
- M. Jaishankar, T. Tseten, N. Anbalagan, B. B. Mathew and K. N. Beeregowda, Toxicity, mechanism and health effects of some heavy metals, Interdiscip. Toxicol., 2014, 7, 60–72 CrossRef PubMed.
- S. J. Stohs and D. Bagchi, Oxidative mechanisms in the toxicity of metal ions, Free Radical Biol. Med., 1995, 18, 321–336 CrossRef CAS PubMed.
- A. Manuja, B. Kumar, R. Kumar, D. Chhabra, M. Ghosh, M. Manuja, B. Brar, Y. Pal, B. N. Tripathi and M. Prasad, Metal/metal oxide nanoparticles: Toxicity concerns associated with their physical state and remediation for biomedical applications, Toxicol. Rep., 2021, 8, 1970–1978 CrossRef CAS PubMed.
- A. Abdal Dayem, M. K. Hossain, S. B. Lee, K. Kim, S. K. Saha, G.-M. Yang, H. Y. Choi and S.-G. Cho, The Role of Reactive Oxygen Species (ROS) in the Biological Activities of Metallic Nanoparticles, Int. J. Mol. Sci., 2017, 18, 120 CrossRef PubMed.
- V. Forest, J.-F. Hochepied, L. Leclerc, A. Trouvé, K. Abdelkebir, G. Sarry, V. Augusto and J. Pourchez, Towards an alternative to nano-QSAR for nanoparticle toxicity ranking in case of small datasets, J. Nanopart. Res., 2019, 21, 95 CrossRef.
- R. R. Crichton, in Metal Chelation in Medicine, The Royal Society of Chemistry, 2017, pp. 1–23, 10.1039/9781782623892-00001.
- N. Mei, Y. Zhang, Y. Chen, X. Guo, W. Ding, S. F. Ali, A. S. Biris, P. Rice, M. M. Moore and T. Chen, Silver nanoparticle-induced mutations and oxidative stress in mouse lymphoma cells, Environ. Mol. Mutagen., 2012, 53, 409–419 CrossRef CAS PubMed.
- J. Roy and K. Roy, Assessment of toxicity of metal oxide and hydroxide nanoparticles using the QSAR modeling approach, Environ. Sci.: Nano, 2021, 8, 3395–3407 RSC.
- W.-S. Cho, R. Duffin, F. Thielbeer, M. Bradley, I. L. Megson, W. MacNee, C. A. Poland, C. L. Tran and K. Donaldson, Zeta Potential and Solubility to Toxic Ions as Mechanisms of Lung Inflammation Caused by Metal/Metal Oxide Nanoparticles, Toxicol. Sci., 2012, 126, 469–477 CrossRef CAS PubMed.
- I. Linkov, J. Steevens, G. Adlakha-Hutcheon, E. Bennett, M. Chappell, V. Colvin, J. M. Davis, T. Davis, A. Elder, S. Foss Hansen, P. B. Hakkinen, S. M. Hussain, D. Karkan, R. Korenstein, I. Lynch, C. Metcalfe, A. B. Ramadan and F. K. Satterstrom, Emerging methods and tools for environmental risk assessment, decision-making, and policy for nanomaterials: summary of NATO Advanced Research Workshop, J. Nanopart. Res., 2009, 11, 513–527 CrossRef CAS PubMed.
- M. Simkó, D. Nosske and W. G. Kreyling, Metrics, Dose, and Dose Concept: The Need for a Proper Dose Concept in the Risk Assessment of Nanoparticles, Int. J. Environ. Res. Public Health, 2014, 11, 4026–4048 CrossRef PubMed.
- Y. Feng, Y. Chang, K. Xu, R. Zheng, X. Wu, Y. Cheng and H. Zhang, Safety-by-Design of Metal Oxide Nanoparticles Based on the Regulation of their Energy Edges, Small, 2020, 16, 1907643 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d2en00897a |
| This journal is © The Royal Society of Chemistry 2023 |