Open Access Article
Open Access ArticleFluorescence turn-on by photoligation – bright opportunities for soft matter materials
Vinh X.
Truong†
*ab,
Joshua O.
Holloway†
b and
Christopher
Barner-Kowollik
 *bc
*bc
aInstitute of Materials Research and Engineering (IMRE), Agency for Science, Technology and Research (A*STAR), 2 Fusionopolis Way, Singapore 138 634, Singapore. E-mail: vinh_truong@imre.a-star.edu.sg
bSchool of Chemistry and Physics, Centre for Materials Science, Queensland University of Technology (QUT), Brisbane, QLD 4000, Australia. E-mail: christopher.barnerkowollik@qut.edu.au
cInstitute of Nanotechnology, Karlsruhe Institute of Technology (KIT), Hermann-von-Helmholtz-Platz 1, 76344 Eggenstein-Leopoldshafen, Germany
First published on 26th October 2022
Abstract
Photochemical ligation has become an indispensable tool for applications that require spatially addressable functionalisation, both in biology and materials science. Interestingly, a number of photochemical ligations result in fluorescent products, enabling a self-reporting function that provides almost instantaneous visual feedback of the reaction's progress and efficiency. Perhaps no other chemical reaction system allows control in space and time to the same extent, while concomitantly providing inherent feedback with regard to reaction success and location. While photoactivable fluorescent properties have been widely used in biology for imaging purposes, the expansion of the array of photochemical reactions has further enabled its utility in soft matter materials. Herein, we concisely summarise the key developments of fluorogenic-forming photoligation systems and their emerging applications in both biology and materials science. We further summarise the current challenges and future opportunities of exploiting fluorescent self-reporting reactions in a wide array of chemical disciplines.
Introduction
Over the last few decades, the renaissance of photochemistry has laid the foundation for substantial innovations in research areas spanning from (cell) biology to soft matter materials.1,2 In the realm of biology, optical control of fluorescent probes is a valuable tool for detecting and imaging biomolecules with superior spatial and temporal resolution,3 which in turn enables the study of biological targets in cells, tissues and even entire intact organisms in a non-invasive manner.4 Thus, various photoactivable fluorescent probes, including organic small molecules,5,6 nucleic acids,7 and nanomaterials8,9 – whose fluorescent properties can be regulated by light irradiation – have been developed for bio-sensing and imaging applications. Most light responsive fluorescent probes are constructed by using either photocleavable groups to cage fluorogenic dyes, which can be subsequently uncaged by light,10 or photoswitches that express fluorogenic functionality in specific isomeric states that form under irradiation.11,12 The photon-excitable fluorescence characteristics of these probes are often not accompanied by molecular recognition or modification processes – they simply enable tracking of the targeted biomolecules.Starting from the pioneering work of light-induced neuronal stimulation by Richard Fork in 1971,13 there is an increasing interest in using light to not only report but also perturb biological activity.14 Thus, new photochemical systems that enable alteration of biological processes in their native environment – often referred to as photo-triggered bioorthogonal click chemistry, have become a critical research focus.15,16 Interestingly, despite the increasing number of light-triggered click reactions that have been introduced to date, only a limited number of photochemical ligations result in fluorescent products. Notably, the inherent fluorogenic function provides valuable visual feedback of the bio-modification process, without the need of re-engineering the biochemical pathways.14,17
Concurrent with their increasing use in new biological technology, advancements in fluorescence turn-on photoligation have also been implemented into the broader macromolecular and materials science fields. Indeed, our group and others have demonstrated the use of fluorescence turn-on by photochemical ligation in the preparation of inherently fluorescent hydrogels, single chain nanoparticles or microparticles.18–29 Herein, we discuss the recent developments in photochemical ligation that generate fluorogenic products alongside their applications in self-reporting (bio)materials designs.
Fluorescence turn-on via tetrazole-based photoligation
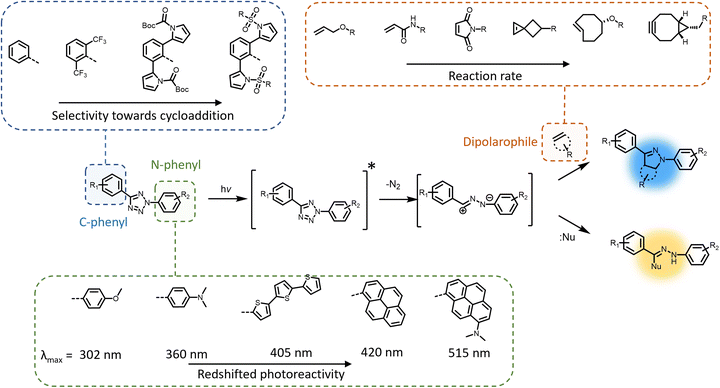
The photochemical ligation between 2,5-diaryl tetrazoles and alkenes is among the most prominent reactions that form fluorescent products, and substantial progress has been made in improving its photoreactivity, as well as exploiting its photoactivable fluorescent properties in different research areas.30 The photochemical reactivity of the tetrazole towards dipolarophiles was initially reported by Huisgen and coworkers, observing the UV light-induced 1,3-dipolar cycloaddition reaction between 2,5-diphenyltetrazole and methyl crotonate.31–33 An attractive feature of the tetrazoles is their accessibility – these compounds can be readily synthesised in a single reaction step via thermal coupling between inexpensive reagents, including benzensulphonyl hydrazones and arene diazonium salts.34,35 Under irradiation, the excited tetrazole moiety undergoes intersystem crossing (ISC) from the singlet state to the triplet state,36,37 followed by cycloreversion to generate nitrogen and a short-life 1,3-nitrile imine intermediate that can thermally react with either dipolarophiles or nucleophiles (Fig. 1). The nitrile imine intermediate has only been identified using transient spectroscopic techniques in the solid state38 and its electronic properties are still the object of investigation.39 Cycloadditions with dipolarophiles – most commonly electron deficient alkenes – result in the formation of fluorescent 2,5-diaryl pyrazolines, which can be used for the detection of photo-labelled (bio)macromolecules.40–43 The first biological application of tetrazole-based photoligation was reported in 2008 by Lin and co-workers, in which the authors demonstrated the photo-induced labelling of proteins by UV light (λmax = 302 nm) and the associated tracking of the ligation by monitoring the fluorescent adducts.44 In following reports, Lin's team further proved the utility of the tetrazole-based photoligation by labelling of living cells, including E. coli45 and HeLa cells.46 In this context, the authors referred to tetrazole-based photoligation as bioorthogonal “photoclick” chemistry. However, careful investigation of the photoinduced ligation between tetrazoles and simple olefins in physiological environment reveals significant side products from nucleophilic addition.47 Nevertheless, given its photo-induced reactivity towards biological nucleophiles including carboxyl and thiol groups to also form fluorescent adducts, tetrazoles may be used as general photo-labelling reagents for e.g., studying of proteins interactions48,49 or tracking of therapeutics in vivo.50–52The yield of cycloaddition adducts from nitrile imine reactions can be substantially improved by using electron deficient alkenes (Fig. 1, top right), such as acrylamides,53 methacrylates,54 and maleimides,55–57 or ring-strained alkenes/alkynes.58 For example, the cycloaddition of p-methylbenzoate tetrazole with allyl phenyl ether proceeds with a second order rate coefficient k2 = 0.002 M s−1, and this value increases significantly when reacting with acrylamide (k2 = 0.15 M s−1) under the employed reaction conditions.53 Further optimisation to increase the fidelity towards cycloaddition pathway includes shielding of the 1,3-nitrile imine intermediate with bulky substituents on the C-phenyl motif (Fig. 1, top left),59–62 or macrocyclic tetrazoles structure (Scheme 1).60 Coupled with electron-deficient or ring-strain alkenes/alkynes for extremely fast cycloadditions, the k2 value can be increased to 39![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 M s−1.59 In each of these optimisation processes, the applicability in protein labelling and live cell imaging with self-reporting function – via formation of the fluorescent adducts – was reported, validating the potential of this reaction type in biological research.
200 M s−1.59 In each of these optimisation processes, the applicability in protein labelling and live cell imaging with self-reporting function – via formation of the fluorescent adducts – was reported, validating the potential of this reaction type in biological research.
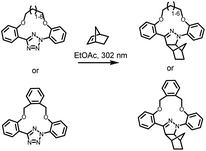 | ||
| Scheme 1 Tetrazole-containing macrocyclic structure for enhanced photoinduced 1,3-dipolar cycloaddition reactions with norbornene. | ||
To further enhance the utility of tetrazole-ene photoligation in bioengineering applications, substantial research has focused on redshifting the activation wavelength of tetrazole cycloreversion by modification of the N-phenyl structure (Fig. 1, bottom), including the addition of either electron donating substituents (methoxy or dimethylamine) at the para-position,63 or the design of extended π-structures such as styryl,64 conjugated thiophene derivatives,65,66 and pyrene.67 Highly conjugated naphthalene-based tetrazole compounds also feature strong two-photon absorption properties suitable for two-photon activated 1,3-dipolar cycloaddition using a 700 nm femtosecond pulsed laser.68 More recently, our team combined both strategies – conjugated structures via a pyrene and electron donating groups via a dimethylamine – introducing a dimethylamino pyrenyl aryl tetrazole moiety that can be activated by green light (λmax = 515 nm) for polymer end-group modification.69 Furthermore, the activation wavelength can be significantly redshifted when the photochemical system is assisted by up-conversion nanoparticles that convert near infrared (NIR) light (λmax = 974 nm) to UV light (λ = 330–370 nm), enabling an efficient and clean reaction between a pyrene aryl tetrazole and a maleimide.70 The NIR light-induced ligation was subsequently applied for the attachment of biotin to a polycaprolactone behind a tissue layer.
Modification of the N-phenyl moiety not only provides the opportunity to adjust the activation wavelength, yet improve the quantum efficiency and photostability of the resultant pyrazoline fluorophores, facilitating enhanced imaging of targeted cellular components.71,72 For example, the inclusion of a bis(trifluoromethyl)benzene substituent results in a 10-fold increase in the fluorescence quantum yield (φF = 0.42 vs. 0.04) of the pyrazole adduct compared to the non-substituted phenyl ring.71 It should be noted that even when a tetrazole exhibits absorption in the visible region, there is no guarantee for nitrile imine formation by irradiation within such a wavelength window, due to the potential vibrational (non-radiative) deactivation of the singlet excited state, as observed in dimethylamino-biphenyl-functionalised tetrazole.36 Furthermore, the preparation of tetrazole molecules with redshifted photoreactivity and bulky substituents can be demanding, often involving multiple synthesis steps with low to moderate yield (10–50%) in each step. In fact, the syntheses of tetrazoles with bulky substituents or conjugated structures have only been reported on the milligram scales.62,69 Consequently, scaling such photoreactive handles is a major obstacle in materials applications.
Towards the 2010s, the tetrazole-based photoligation has been increasingly applied in polymer chemistry for the preparation of “self-reporting” fluorescent materials.16,73 Early materials' engineering applications include fabrications of light-activated fluorescent hydrogel systems. Specifically, the team of Zhong designed a 4-arm poly(ethylene glycol) (PEG) functionalised with tetrazole for UV light (λmax = 365 nm) induced crosslinking with a 4-arm PEG methacrylate19 or a chitosan glycidyl methacrylated,18 forming noncytotoxic and fluorescent hydrogels (Fig. 2A). Zhong's team further incorporated the tetrazole-ene photoligation with microfluidic techniques to fabricate fluorescent hyaluronic acid microgels, which can be traced in situ when delivered to tumour sites of mice.74,75 Using a similar approach, Guaresti et al. photo-crosslinked complementary tetrazole and maleimide chitosan derivatives into fluorescent and self-healing hydrogels.76 In contrast, Zhang and co-workers utilised the intramolecular tetrazole-ene photoreaction77,78 to degrade a pre-formed hydrogel for 3D cell encapsulation and release.20,79 In their system, the photo-triggered tetrazole-to-pyrazoline transformation disrupt the hydrophobic interaction and π–π stacking of the self-assembled peptides, generating a highly fluorescent reporter (Fig. 2B). Critically, the self-reporting fluorescent properties present a powerful tool to obtain quantitative information on – or counting the number of crosslinking points within polymer networks. Estupiñán et al. established such a methodology by preparing polymer networks via tetrazole-maleimide photoligation that can be degraded by aminolysis, while leaving the pyrazoline intact (Fig. 2C).25 The amount of pyrazoline junctions can subsequently be quantified by fluorescence spectroscopy, providing insight into the efficiency of the photo-crosslinking via nitrile imine-mediated cycloaddition.
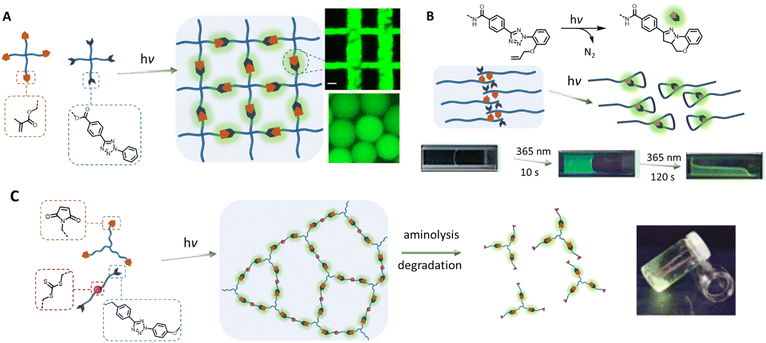 | ||
| Fig. 2 Application of tetrazole-ene photoligation in the preparation of photoresponsive hydrogels. (A) Photo-crosslinking of 4-arm PEG containing tetrazole and methacrylate end-groups to form fluorescent hydrogels, or microgels by integrating with microfluidic devices. (B) Photo-induced intramolecular tetrazole-ene leads to disruption of a self-assembled structure and consequently degradation of the physically crosslinked hydrogels. (C) Fluorescent polymer networks, formed by tetrazole-based photo-crosslinking, can be degraded by aminolysis of the trithiocarbonate functionalities within the structure, leading to soluble fragments. Analysis of the fragments by fluorescence spectroscopy can provide quantitative information on the number of crosslinking junctions. (Reproduced with permission from ref. 18, 19, 65 and 24. | ||
In addition to the fabrication of photoresponsive hydrogels, the tetrazole photoligation is a powerful tool for polymerisation and crosslinking to generate complex 3D soft matter structures. For example, our team has employed the tetrazole-ene photoreaction for the preparation of polymer-fullerene80 and polymer-cellulose27,81 hybrid networks. The critical advantages of the tetrazole-ene reaction, including mild reaction conditions, rapid reaction rate, and additive-free, further enable the synthesis of multifunctional microparticles24,82 with narrow dispersity and tuneable particle sizes (Fig. 3A). Along with their inherent fluorescent properties, these particles can be readily engineered for the attachment of bioactive molecules, or degradability in response to a specific chemical stimulus, such as hydrogen peroxide, with a chemiluminescent read-out function.29,83 In addition, our team has implemented the tetrazole-ene photoligation in folding of polymer chains into fluorescent nanoparticles,21,22,26,84 affording access to synthetic single chain nanoparticles (SNCPs) with potential applications in bioimaging and tracking. In a collaborative work with Stenzel's team, we developed glycopolymeric SCNPs and adsorbed these onto nanodiamond particles for monitoring of the nanoparticles within cellular environment.28 Landfester and co-workers further crosslinked proteins, via tetrazole-norbornene photoreaction, under miniemulsion polymerisation conditions to form fluorescent protein nanocarriers.23 These cargos have excellent drug uptake capacity and can be internalised by dendritic cells for controlled intracellular release.
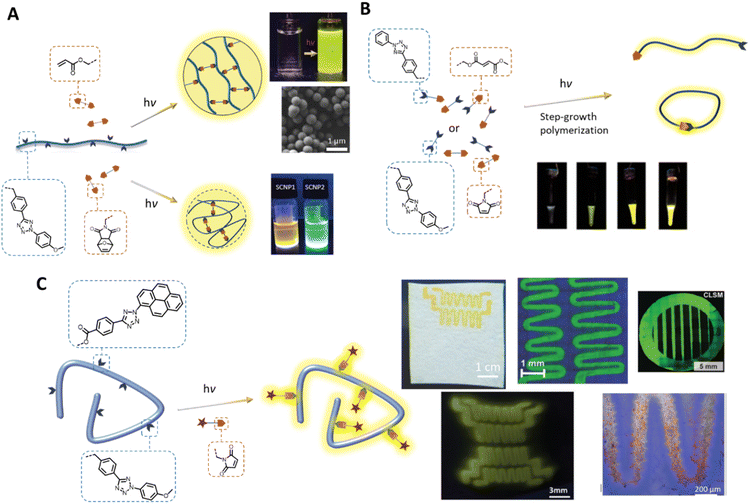 | ||
| Fig. 3 Application of tetrazole-ene photoligation in polymer synthesis and modification. (A) Photo-induced intermolecular or intramolecular crosslinking of polymer chains to form fluorescent microspheres (top) or single chain nanoparticles (bottom), respectively. (B) Step-growth photopolymerisation of AA and BB monomers, or an AB monomer containing a tetrazole and an electron-deficient alkene. The progress of photopolymerisation can be followed by the naked eye due to the fluorescence of the pyrazoline adducts. (C) Photo-patterning, via tetrazole-ene ligation, on cellulose-based substrates (top) or hydrogels (bottom left) to form fluorescent patterns, and attachment of bioactive component that enables cell attachment (bottom right). (Reproduced with permission from ref. 20, 26, 73, 77, 83 and 84. | ||
In addition to its utility in the fabrication of 3D polymer networks, the tetrazole-ene photoligation has been widely used in the preparation of fluorescent materials. Taking advantage of the rapid reaction kinetics of the reaction, our team introduced a facile strategy for the synthesis of self-reporting pyrazoline polymers by step-growth polymerisation of monomers (either AB monomer, or AA and BB monomers, Fig. 3B), containing tetrazole and complementary fumarate85 or maleimide86 moieties. The polymerisation progress can be monitored by following the fluorescence intensity, which was observed even by the naked eye, as the pyrazoline units formed under UV light irradiation. Mane et al. further incorporated a fumarate terminated PEG (B type) molecule as the chain stopper in the polymerisation of the AB monomer, forming an amphiphilic block copolymer that can be assembled into fluorescent nanoparticles.87 Subsequently, we demonstrated the utility of such nanoparticles as imaging probes for visualisation of the blood vessels in zebra fish. Other in vivo imaging of zebra fish using fluorescent pyrazoline-based probe was demonstrated by Zhang and co-workers, in which the authors performed DNA labelling and fluorescent turn-on directly on zebrafish embryo under UV (λmax = 350 nm) light irradiation with low cytotoxicity.88
The spatial and temporal control of the tetrazole-ene photoligation has been further employed in the self-reported patterning of hard (cellulose-based or modified silicon substrates)89–92 or soft (hydrogels) surfaces93,94 (Fig. 3C) with cell recognition function for cellular attachment.89,94 Critically, the various tetrazole components, based on different substituents on the N-phenyl group (refer to Fig. 1, bottom left), with different photoreactivities have enabled λ-orthogonal patterning on the silicon wafers, generating complex and interconnected patterns on the surface.90 More recently, emerging applications of tetrazole-ene photoligation include the light-induced synthesis of radiotracers for positron emission tomography (PET) imaging.95,96 Here, the rapid photoactivation of the tetrazole moiety enables direct photo- and radiochemical labelling of monoclonal antibodies. The resultant pyrazoline linker between the targeting ligand and the metal chelator can be used as the fluorescent core for optical imaging, thus minimising complex synthesis steps to incorporate the optical imaging function.
Fluorescent pyrazolines derived from diarylsydnones
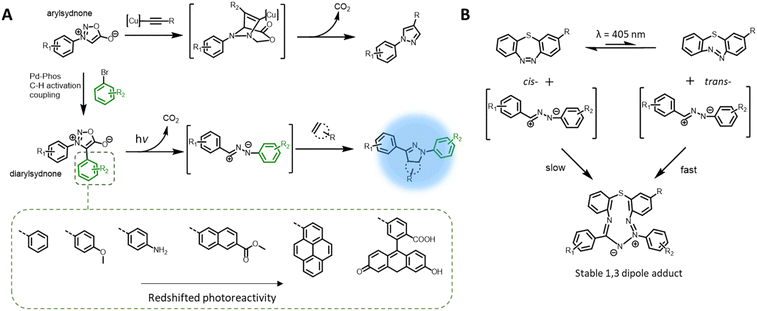
The nitrile imine intermediate can also be produced from various precursors via thermal and chemical activation.97 Of particular interest for light-induced ligation is the photo-generation of nitrile imines from sydnone derivatives, via cycloelimination of carbon dioxide (Fig. 4A), for cycloaddition with alkenes or nucleophiles. Although the photoreactivity of the sydnones was reported in the 1970s,98,99 these compounds have been mostly used in the copper-mediated cycloaddition with alkynes,100,101 or catalyst free cycloaddition with ring-strained alkynes102,103 as a substitute for azides. More recently, the team of Yu reinvestigated sydnone photochemical properties and revealed that diarylsdynone derivatives can efficiently react with various alkenes under light irradiation to form pyrazoline adducts.104 Compared to 2,4-diaryltetrazoles, the two ortho-substituted phenyl rings in the diarylsydnone scaffold are twisted around the sydnone core, facilitating tuning of both the reactivity and selectivity towards alkenes. The C-phenyl structure can be readily modified via a one-step procedure C–H activation coupling of monoarylsydnone (Fig. 4A, bottom), providing easy access to various diarylsydnones with enhanced photoreactivity for the photocycloaddition with various alkene substrates. The pyrazole adducts also feature high fluorescent quantum yields (φF = 0.16–0.56). Taking advantage of the straightforward synthetic procedure, Yu and co-workers prepared a library of some 15 sydnone compounds and screened their photoreactivity towards various alkenes. The authors identified several diarylsydnone molecules with excellent reactivity towards trans-cyclooctene, forming pyrazoline cycloadducts with an outstanding fluorescence turn-on effect. The utility of these compounds in bioimaging has been demonstrated via fluorogenic photolabeling of proteins105 and cell membranes.106 Yu's team further identified a ring-strained diazobenzene structure (dibenzo[b,f][1,4,5]thiadiazepine) that, in its trans-state, has high reactivity with nitrile imine (Fig. 4B). The cyclic diazobenzene is thermally stable in the cis-state, which shows negligible reactivity towards the nitrile imine, and undergoes isomerisation to the trans-state under irradiation of the same wavelength (λmax = 405 nm) used for activation of the diarylsydnone.107 The in situ photo-generation of the ring-strain structure thus enables visible light acceleration of bioorthogonal ligation in cellular environment.108,109 However, the resultant 1,3-dipole adducts do not possess fluorescent properties, and thus a fluorogenic dye was required for labelling of the targeted biomacromolecules.Photochemical pericyclic reactions with fluorescent adducts
While photochemical pericyclic reactions have been widely employed in soft matter materials for light responsive function,110 most cycloadducts do not possess fluorescent property. Nevertheless, recent advances in redshifting the activation wavelength of photocycloaddition have introduced new reaction types that form fluorogenic adducts, with significant potential in both biology and materials applications. In particular, Li et al. reported a photocycloaddition of 9,10-phenanthrenequinone (PQ) moiety with electron-rich vinyl ether (VE) under mild irradiation conditions (handheld LED light, white light) to form fluorogenic [4 + 2] cycloadducts (Fig. 5A, top).111 The photocycloaddition is highly selective, showing no cross reactivity with electron-deficient or ring-strained alkenes/alkynes. In addition, the PQ handle can be readily prepared on the gram scale with good yields (83%). These authors further demonstrated the bioorthogonality of the photoligation by protein labelling and fluorescent imaging of live epithelial cells. Furthermore, the PQ function can be readily attached to biomacromolecules, such as glycans, for spatiotemporal labelling of these molecules in live cells.112 However, it should be noted that the PQ can spontaneously and rapidly react with furan-2(3H)-one derivatives, via thermal cycloaddition, in water.113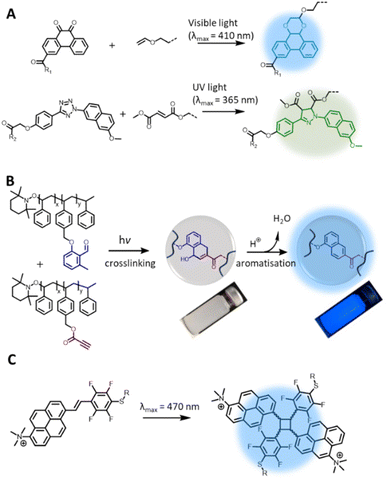 | ||
| Fig. 5 Recently reported photochemical pericyclic reactions that form (pro)fluorogenic products and their applications. (A) [4 + 2] Cycloaddition of 9,10-phenanthrenequinone and vinyl ether forming a fluorescent phenanthrodioxine moiety (top left); the PQ-VE photocycloaddition can take place orthogonally to the tetrazole-ene photoligation, enabling λ-orthogonal labelling of hydrogel networks (right). (B) Photochemical reaction of o-methylbenzaldehydes and propiolate ester pendent groups for polymer crosslinking, enabling microparticles formation; addition of acid triggers aromatisation of the cyclic adduct, forming the fluorescent moiety. (C) [2 + 2] photocycloaddition of 1-trimethylammonium-6-pentafluorstyryl-pyrene chromophore forming fluorescent cyclobutane structures (Reproduced with permission from ref. 110. | ||
Since its report in 2018, the PQ-VE photoligation has gained considerable traction in soft matter materials applications. Specifically, Adronov and co-workers prepared a conjugated polymer that contains the PQ moiety in the backbone, and effectively functionalised the polymer with several electron-rich alkenes under visible light irradiation, forming fluorescent pendent groups.114 The team of Feringa integrated the PQ-VE photoligation with flow chemistry for rapid synthesis of short-lived 18F-PET tracers.115 Using a flow micro-reactor equipped with automated module, the authors were able to achieve near complete conversion of the desired PET products within one minute, and the compounds have potential for multimodal imaging protocols due to their inherent fluorescence properties. In a recent report, Zheng et al. employed the PQ-VE photocycloaddition in the light-induced attachment of quantum dots to mosquito-transmitted Zika virus for viral labelling and real-time imaging of virus-host cell interactions.116 The high selectivity of PQ-VE photocycloaddition further allows for its use in conjunction with the tetrazole-ene photoligation in the λ-orthogonal ligation of bioactive molecules to a hydrogel substrate. Wu et al. demonstrated such a concept by constructing a double network hydrogel containing both the tetrazole and PQ moieties, and photo-patterned the hydrogel with complementary fumarate or VE compounds by UV light or visible light, respectively (Fig. 5A).117 Using this approach, the authors decorated a peptide-based hydrogel with various growth factors in a spatially resolved fashion, mimicking the architecture of native extracellular matrices, which is suitable for directing the growth and migration of encapsulated stem cells.
In our search for efficient photocycloadditions based on a photoenolisation process – whereby photo-excited o-methylbenzaldehydes (o-MBAs) reversibly transform to dienol species for [4 + 2] cycloaddition with dipolarophiles118 – we discovered a fluorogenic function that is activated under acidic conditions. Specifically, by using electron-deficient alkynes in the photoreaction with o-MBAs, we observed the formation of 1,4-dihydro-1-naphtholes that are rapidly converted to fluorescent naphthalenes by addition of catalytic amount of acids (Fig. 5B).119 The additional control over fluorogenic functions is highly interesting as it opens opportunities in e.g., sensor applications, where the fluorescence read-out can be triggered by a change in the environmental pH. As a proof-of-concept demonstration, our team has employed the photochemical ligation in the preparation of polystyrene microparticles with acid-activatable fluorescence properties.120
Within the broad spectrum of pericyclic photochemical reactions, [2 + 2] or [4 + 4] photocycloadditions generally result in cycloadducts with lower fluorescence intensity compared to the precursor chromophores due to the disruption of the π-conjugation.121–124 Interestingly, Ludwanowski et al. reported a [2 + 2] chromophore with a highly conjugated and electron-deficient push–pull system (Fig. 5C) that forms a strongly fluorescent product under visible light (λmax = 470 nm) irradiation.125 Therefore, the progress of the [2 + 2] photocycloaddition of such a chromophore was conveniently monitored by fluorescence intensity, which can be measured in a turbid solution and solid materials, compared to UV/Vis absorbance measurements, which require highly diluted and clear sample. The authors further employed the [2 + 2] photoligation in polymer crosslinking for the preparation of PEG hydrogels with inherent fluorescent properties.125 It should be noted that the [2 + 2] photocycloaddition reaction may lead to adducts with DNA or RNA, e.g. UV-induced crosslinking of thymine and uracil derivatives,126 and thus the biocompatibility of [2 + 2] chromophore should be investigated in the context of DNA damage for biological applications.
Outlook
The current Perspective has examined photochemical reactions that lead to products – ideally one defined product – that are fluorescent or at the least, pro-fluorescent. We have highlighted that a range of effective options are available in the current chemical toolbox to construct fluorescent entities via photochemical means. In the final section, we will explore what challenges lie ahead and what opportunities present themselves for the exploitation of fluorogenic reaction technology in the field of soft matter materials science.On the chemical level, the reaction systems need to improve further with regard to the quantum efficiency with which the fluorophores are generated, with the view to employing the longest possible wavelengths. Bright organic fluorophores, featuring a high fluorescence quantum yield and compatible with live-cell environments, are highly desirable for bioimaging applications, especially in single-molecule super resolution techniques.127 In many photochemical systems, perhaps most prevalent in the popular tetrazole-driven ligation, the highly reactive nitrile imine intermediate is able to react with a range of nucleophiles as well as double bonds, and some of the formed reaction products have limited or no fluorescence. Thus, limiting side reactions to non-fluorescent products is essential. While this can be achieved by modification of the phenyl structures adjacent to the tetrazole ring, the synthesis of such highly conjugated or bulky structures can be labour-intensive and time-consuming, thus diminishing the accessibility of these compounds in material applications. Alternatively, the recently reported diarylsydnones, which also generate reactive nitrile imine under light irradiation, are excellent candidates for fluorogenic photoligation due to their straightforward preparation procedures.
In our view, photochemically allowed pericyclic reactions such as [4 + 2] and [2 + 2] photocycloadditions are important avenues when it comes to generating one highly specific reaction product, however formation of fluorogenic cycloadducts is uncommon. We have outlined the most current and interesting examples of [4 + 2] and [2 + 2] reactions above and highlighted their potential in soft matter materials engineering, yet critically – this applies specifically to biological applications – the action wavelength for these reactions should be as long as possible. Nevertheless, redshifting the activation wavelength of photochemical ligations is an ongoing challenge in the realm of photochemical pericyclic reactions, where the current maximum ligation wavelength is close to 550 nm.128,129 We have recently introduced tetrazoles with activation wavelengths exceeding 500 nm69 and further efforts should concentrate on making such long wavelength activated systems available in aqueous environments through suitable hydrophilic handles. Similarly, the use of up-conversion nanoparticles can be an attractive option to push activation wavelengths in tetrazole systems. To avoid the use of metals, organic photosensitisers are an alternative and several suitable photosensitisers have been employed in photocatalytic activation of bioorthogonal ligation by red and far-red light (600–660 nm).130–133
While tracing reaction loci in biological systems is perhaps to date the most exploited application of fluorogenic product-generated photochemical reaction systems, opportunities in synthetic soft matter materials science abound, as exemplified by the extensive utilisation of tetrazole-ene photoligation within the last 15–20 years. In initial work in this realm, we have exploited the tetrazole ligation for counting chemical ligation points in polymer networks as a probe to quantitatively assess the formed bonds during network generation.25 This initial concept awaits to be expanded into an on-line monitoring tool for a wide range of network formations, including in the realm of single chain nanoparticles (SCNPs). SCNPs represent networks consisting of a single folded polymer chain and it is an outstanding challenge to quantitatively map the number of bonds formed within the SCNPs and correlate this number with their morphology. As SCNPs are currently discussed as potential reusable catalytic systems that fuse the advantages of hetero- and homogeneous catalysis, understanding the relationship between their inner chemical connectivity with their outer morphology is a matter of priority. In the network realm – whether on the macroscopic or microscopic scale – it is highly desirable that the generated fluorophores emit light at long wavelengths, as the longer the emission wavelength, the less any readouts will be affected by light penetration issues through a network or solution. To date, there exists ample opportunity to devise systems whose emission wavelength are well within the red part of the spectrum.
While we have above focused on photochemical reactions that generate a fluorophore as the result of chemical bond forming event, it is fascinating to explore the possibility that a photochemical bond cleavage event generates a fluorophore – a concept that has been utilised in live cell imaging,134 but is rarely seen in materials engineering. Such systems entail the enticing opportunity to detect photochemical damage to materials through irradiation, via a possibly quantitative readout of the fluorescence intensity. Here, the above-mentioned need for red-shifted fluorophore emission is a critical parameter. Such molecular damage self-reporting systems are in critical need, finding possible applications in all realms where soft-matter materials carry out critical functions in transport, including aerospace.
Author contributions
The perspective was written with contributions from all three authors.Conflicts of interest
There are no conflicts to declare.Acknowledgements
C. B.-K. acknowledges funding via an Australian Research Council (ARC) Laureate Fellowship underpinning his photochemical research program as well as continued support from QUT's Centre for Materials Science, the Deutsche Forschungsgemeinschaft (DFG) through the Excellence Cluster “3D Matter Made to Order” (EXC-2082/1-390761711) and the Karlsruhe Institute of Technology (KIT).References
- B. T. Tuten, S. Wiedbrauk and C. Barner-Kowollik, Prog. Polym. Sci., 2020, 100, 101183 CrossRef CAS.
- V. M. Lechner, M. Nappi, P. J. Deneny, S. Folliet, J. C. K. Chu and M. J. Gaunt, Chem. Rev., 2022, 122, 1752–1829 CrossRef CAS PubMed.
- F. M. Raymo, Phys. Chem. Chem. Phys., 2013, 15, 14840–14850 RSC.
- Z. Zou, Z. Luo, X. Xu, S. Yang, Z. Qing, J. Liu and R. Yang, TrAC, Trends Anal. Chem., 2020, 125, 115811 CrossRef CAS.
- F. M. Jradi and L. D. Lavis, ACS Chem. Biol., 2019, 14, 1077–1090 CrossRef CAS PubMed.
- E. M. S. Stennett, M. A. Ciuba and M. Levitus, Chem. Soc. Rev., 2014, 43, 1057–1075 RSC.
- H. Li and J. C. Vaughan, Chem. Rev., 2018, 118, 9412–9454 CrossRef CAS PubMed.
- T.-T. Chen, X. Tian, C.-L. Liu, J. Ge, X. Chu and Y. Li, J. Am. Chem. Soc., 2015, 137, 982–989 CrossRef CAS PubMed.
- Z. Tian and A. D. Q. Li, Acc. Chem. Res., 2013, 46, 269–279 CrossRef CAS PubMed.
- R. Weinstain, T. Slanina, D. Kand and P. Klán, Chem. Rev., 2020, 120, 13135–13272 CrossRef CAS PubMed.
- T. Fukaminato, J. Photochem. Photobiol., C, 2011, 12, 177–208 CrossRef CAS.
- J. Zhao, Z. Wang, Q. Ai, H. Li, P. Cai, J. Si, X. Yao, Q. He, X. Hu and Z. Liu, J. Lumin., 2022, 248, 118973 CrossRef CAS.
- R. L. Fork, Science, 1971, 171, 907–908 CrossRef CAS PubMed.
- A. Herner and Q. Lin, Top. Curr. Chem., 2015, 374, 1 CrossRef PubMed.
- B. D. Fairbanks, L. J. Macdougall, S. Mavila, J. Sinha, B. E. Kirkpatrick, K. S. Anseth and C. N. Bowman, Chem. Rev., 2021, 121, 6915–6990 CrossRef CAS PubMed.
- G. S. Kumar and Q. Lin, Chem. Rev., 2021, 121, 6991–7031 CrossRef CAS PubMed.
- C. P. Ramil and Q. Lin, Curr. Opin. Chem. Biol., 2014, 21, 89–95 CrossRef CAS PubMed.
- C. Deng, F. Meng, R. Cheng and Z. Zhong, J. Controlled Release, 2013, 172, e146–e147 Search PubMed.
- Y. Fan, C. Deng, R. Cheng, F. Meng and Z. Zhong, Biomacromolecules, 2013, 14, 2814–2821 CrossRef CAS PubMed.
- M. He, J. Li, S. Tan, R. Wang and Y. Zhang, J. Am. Chem. Soc., 2013, 135, 18718–18721 CrossRef CAS PubMed.
- C. Heiler, J. T. Offenloch, E. Blasco and C. Barner-Kowollik, ACS Macro Lett., 2017, 6, 56–61 CrossRef CAS PubMed.
- J. T. Offenloch, J. Willenbacher, P. Tzvetkova, C. Heiler, H. Mutlu and C. Barner-Kowollik, Chem. Commun., 2017, 53, 775–778 RSC.
- K. Piradashvili, J. Simon, D. Paßlick, J. R. Höhner, V. Mailänder, F. R. Wurm and K. Landfester, Nanoscale Horiz., 2017, 2, 297–302 RSC.
- C. Wang, M. M. Zieger, A. Schenzel, M. Wegener, J. Willenbacher, C. Barner-Kowollik and C. N. Bowman, Adv. Funct. Mater., 2017, 27, 1605317 CrossRef.
- D. Estupiñán, C. Barner-Kowollik and L. Barner, Angew. Chem., Int. Ed., 2018, 57, 5925–5929 CrossRef PubMed.
- C. Heiler, S. Bastian, P. Lederhose, J. P. Blinco, E. Blasco and C. Barner-Kowollik, Chem. Commun., 2018, 54, 3476–3479 RSC.
- D. Hoenders, J. Guo, A. S. Goldmann, C. Barner-Kowollik and A. Walther, Mater. Horiz., 2018, 5, 560–568 RSC.
- A. P. P. Kröger, M. I. Komil, N. M. Hamelmann, A. Juan, M. H. Stenzel and J. M. J. Paulusse, ACS Macro Lett., 2019, 8, 95–101 CrossRef PubMed.
- L. Delafresnaye, J. P. Hooker, C. W. Schmitt, L. Barner and C. Barner-Kowollik, Macromolecules, 2020, 53, 5826–5832 CrossRef CAS.
- V. Pirota, A. Benassi and F. Doria, Photochem. Photobiol. Sci., 2022, 21, 879–898 CrossRef CAS PubMed.
- R. Huisgen, M. Seidel, G. Wallbillich and H. Knupfer, Tetrahedron, 1962, 17, 3–29 CrossRef CAS.
- R. Huisgen, M. Seidel, J. Sauer, J. McFarland and G. Wallbillich, J. Org. Chem., 1959, 24, 892–893 CrossRef CAS.
- J. S. Clovis, A. Eckell, R. Huisgen and R. Sustmann, Chem. Ber., 1967, 100, 60–70 CrossRef CAS.
- S. Ito, Y. Tanaka, A. Kakehi and K.-i. Kondo, Bull. Chem. Soc. Jpn., 1976, 49, 1920–1923 CrossRef CAS.
- K. Koguro, T. Oga, S. Mitsui and R. Orita, Synthesis, 1998, 1998, 910–914 CrossRef.
- E. Blasco, Y. Sugawara, P. Lederhose, J. P. Blinco, A.-M. Kelterer and C. Barner-Kowollik, ChemPhotoChem, 2017, 1, 159–163 CrossRef CAS.
- A. Duque-Prata, C. Serpa and P. J. S. B. Caridade, Appl. Sci., 2021, 11 Search PubMed.
- S.-L. Zheng, Y. Wang, Z. Yu, Q. Lin and P. Coppens, J. Am. Chem. Soc., 2009, 131, 18036–18037 CrossRef CAS PubMed.
- R. C. Mawhinney, H. M. Muchall and G. H. Peslherbe, Chem. Commun., 2004, 1862–1863, 10.1039/B407302A.
- W. Siti, A. K. Khan, H.-P. M. de Hoog, B. Liedberg and M. Nallani, Org. Biomol. Chem., 2015, 13, 3202–3206 RSC.
- C. M. Geiselhart, H. Mutlu and C. Barner-Kowollik, ACS Macro Lett., 2021, 10, 1159–1166 CrossRef CAS PubMed.
- L. Stolzer, A. Vigovskaya, C. Barner-Kowollik and L. Fruk, Chem. - Eur. J., 2015, 21, 14309–14313 CrossRef CAS PubMed.
- M. Xiao, Y.-K. Zhang, R. Li, S. Li, D. Wang and P. An, Chem. - Asian J., 2022, 17, e202200634 CAS.
- W. Song, Y. Wang, J. Qu, M. M. Madden and Q. Lin, Angew. Chem., Int. Ed., 2008, 47, 2832–2835 CrossRef CAS PubMed.
- W. Song, Y. Wang, J. Qu and Q. Lin, J. Am. Chem. Soc., 2008, 130, 9654–9655 CrossRef CAS PubMed.
- W. Song, Y. Wang, Z. Yu, C. I. R. Vera, J. Qu and Q. Lin, ACS Chem. Biol., 2010, 5, 875–885 CrossRef CAS PubMed.
- Z. Li, L. Qian, L. Li, J. C. Bernhammer, H. V. Huynh, J.-S. Lee and S. Q. Yao, Angew. Chem., Int. Ed., 2016, 55, 2002–2006 CrossRef CAS PubMed.
- Y. Tian and Q. Lin, Chem. Commun., 2018, 54, 4449–4452 RSC.
- W. Gui, S. Shen and Z. Zhuang, J. Am. Chem. Soc., 2020, 142, 19493–19501 CrossRef CAS PubMed.
- A. Herner, J. Marjanovic, T. M. Lewandowski, V. Marin, M. Patterson, L. Miesbauer, D. Ready, J. Williams, A. Vasudevan and Q. Lin, J. Am. Chem. Soc., 2016, 138, 14609–14615 CrossRef CAS PubMed.
- Y. Y. Khine, R. Batchelor, R. Raveendran and M. H. Stenzel, Macromol. Rapid Commun., 2020, 41, 1900499 CrossRef CAS PubMed.
- K. Bach, B. L. H. Beerkens, P. R. A. Zanon and S. M. Hacker, ACS Cent. Sci., 2020, 6, 546–554 CrossRef CAS PubMed.
- X. S. Wang, Y.-J. Lee and W. R. Liu, Chem. Commun., 2014, 50, 3176–3179 RSC.
- M. M. Madden, C. I. Rivera Vera, W. Song and Q. Lin, Chem. Commun., 2009, 5588–5590, 10.1039/B912094G.
- F. Wendler, T. Rudolph, H. Görls, N. Jasinski, V. Trouillet, C. Barner-Kowollik and F. H. Schacher, Polym. Chem., 2016, 7, 2419–2426 RSC.
- S. Arndt and H.-A. Wagenknecht, Angew. Chem., Int. Ed., 2014, 53, 14580–14582 CrossRef CAS PubMed.
- B. Lehmann and H.-A. Wagenknecht, Org. Biomol. Chem., 2018, 16, 7579–7582 RSC.
- Z. Yu, Y. Pan, Z. Wang, J. Wang and Q. Lin, Angew. Chem., Int. Ed., 2012, 51, 10600–10604 CrossRef CAS PubMed.
- G. S. Kumar, S. Racioppi, E. Zurek and Q. Lin, J. Am. Chem. Soc., 2022, 144, 57–62 CrossRef CAS PubMed.
- Z. Yu, R. K. V. Lim and Q. Lin, Chem. – Eur. J., 2010, 16, 13325–13329 CrossRef CAS PubMed.
- P. An, T. M. Lewandowski, T. G. Erbay, P. Liu and Q. Lin, J. Am. Chem. Soc., 2018, 140, 4860–4868 CrossRef CAS PubMed.
- S. Jiang, X. Wu, H. Liu, J. Deng, X. Zhang, Z. Yao, Y. Zheng, B. Li and Z. Yu, ChemPhotoChem, 2020, 4, 327–331 CrossRef CAS.
- Y. Wang, W. J. Hu, W. Song, R. K. V. Lim and Q. Lin, Org. Lett., 2008, 10, 3725–3728 CrossRef CAS PubMed.
- Z. Yu, L. Y. Ho, Z. Wang and Q. Lin, Bioorg. Med. Chem. Lett., 2011, 21, 5033–5036 CrossRef CAS PubMed.
- P. An, Z. Yu and Q. Lin, Chem. Commun., 2013, 49, 9920–9922 RSC.
- P. An, Z. Yu and Q. Lin, Org. Lett., 2013, 15, 5496–5499 CrossRef CAS PubMed.
- P. Lederhose, K. N. R. Wüst, C. Barner-Kowollik and J. P. Blinco, Chem. Commun., 2016, 52, 5928–5931 RSC.
- Z. Yu, T. Y. Ohulchanskyy, P. An, P. N. Prasad and Q. Lin, J. Am. Chem. Soc., 2013, 135, 16766–16769 CrossRef CAS PubMed.
- P. W. Kamm, J. P. Blinco, A.-N. Unterreiner and C. Barner-Kowollik, Chem. Commun., 2021, 57, 3991–3994 RSC.
- P. Lederhose, Z. Chen, R. Müller, J. P. Blinco, S. Wu and C. Barner-Kowollik, Angew. Chem., Int. Ed., 2016, 55, 12195–12199 CrossRef CAS PubMed.
- Y.-K. Zhang, M. Li, L. Ruan and P. An, Chem. Commun., 2022, 58, 10404–10407 RSC.
- X. Shang, R. Lai, X. Song, H. Li, W. Niu and J. Guo, Bioconjugate Chem., 2017, 28, 2859–2864 CrossRef CAS PubMed.
- C. M. Geiselhart, H. Mutlu and C. Barner-Kowollik, Angew. Chem., Int. Ed., 2021, 60, 17290–17313 CrossRef CAS PubMed.
- J. Chen, K. Huang, Q. Chen, C. Deng, J. Zhang and Z. Zhong, ACS Appl. Mater. Interfaces, 2018, 10, 3929–3937 CrossRef CAS PubMed.
- J. Chen, Y. Zou, C. Deng, F. Meng, J. Zhang and Z. Zhong, Chem. Mater., 2016, 28, 8792–8799 CrossRef CAS.
- O. Guaresti, L. Crocker, T. Palomares, A. Alonso-Varona, A. Eceiza, L. Fruk and N. Gabilondo, J. Mater. Chem. B, 2020, 8, 9804–9811 RSC.
- Z. Yu, L. Y. Ho and Q. Lin, J. Am. Chem. Soc., 2011, 133, 11912–11915 CrossRef CAS PubMed.
- Z. He, Y. Chen, Y. Wang, J. Wang, J. Mo, B. Fu, Z. Wang, Y. Du and X. Zhou, Chem. Commun., 2016, 52, 8545–8548 RSC.
- Z. Zhou, Q. Yi, T. Xia, W. Yin, A. A. Kadi, J. Li and Y. Zhang, Org. Biomol. Chem., 2017, 15, 2191–2198 RSC.
- Y. Sugawara, K. Hiltebrandt, E. Blasco and C. Barner-Kowollik, Macromol. Rapid Commun., 2016, 37, 1466–1471 CrossRef CAS PubMed.
- A. Hufendiek, A. Carlmark, M. A. R. Meier and C. Barner-Kowollik, ACS Macro Lett., 2016, 5, 139–143 CrossRef CAS PubMed.
- J. P. Hooker, L. Delafresnaye, L. Barner and C. Barner-Kowollik, Mater. Horiz., 2019, 6, 356–363 RSC.
- L. Delafresnaye, C. W. Schmitt, L. Barner and C. Barner-Kowollik, Chem. – Eur. J., 2019, 25, 12538–12544 CrossRef CAS PubMed.
- J. Willenbacher, K. N. R. Wuest, J. O. Mueller, M. Kaupp, H.-A. Wagenknecht and C. Barner-Kowollik, ACS Macro Lett., 2014, 3, 574–579 CrossRef CAS PubMed.
- J. O. Mueller, D. Voll, F. G. Schmidt, G. Delaittre and C. Barner-Kowollik, Chem. Commun., 2014, 50, 15681–15684 RSC.
- D. Estupiñán, T. Gegenhuber, J. P. Blinco, C. Barner-Kowollik and L. Barner, ACS Macro Lett., 2017, 6, 229–234 CrossRef PubMed.
- S. R. Mane, I. L. Hsiao, M. Takamiya, D. Le, U. Straehle, C. Barner-Kowollik, C. Weiss and G. Delaittre, Small, 2018, 14, 1801571 CrossRef PubMed.
- Y. Wu, G. Guo, J. Zheng, D. Xing and T. Zhang, ACS Sens., 2019, 4, 44–51 CrossRef CAS PubMed.
- C. Rodriguez-Emmenegger, C. M. Preuss, B. Yameen, O. Pop-Georgievski, M. Bachmann, J. O. Mueller, M. Bruns, A. S. Goldmann, M. Bastmeyer and C. Barner-Kowollik, Adv. Mater., 2013, 25, 6123–6127 CrossRef CAS PubMed.
- P. Lederhose, D. Abt, A. Welle, R. Müller, C. Barner-Kowollik and J. P. Blinco, Chem. – Eur. J., 2018, 24, 576–580 CrossRef CAS PubMed.
- Y. Sugawara, N. Jasinski, M. Kaupp, A. Welle, N. Zydziak, E. Blasco and C. Barner-Kowollik, Chem. Commun., 2015, 51, 13000–13003 RSC.
- T. Tischer, C. Rodriguez-Emmenegger, V. Trouillet, A. Welle, V. Schueler, J. O. Mueller, A. S. Goldmann, E. Brynda and C. Barner-Kowollik, Adv. Mater., 2014, 26, 4087–4092 CrossRef CAS PubMed.
- J. O. Mueller, N. K. Guimard, K. K. Oehlenschlaeger, F. G. Schmidt and C. Barner-Kowollik, Polym. Chem., 2014, 5, 1447–1456 RSC.
- R. R. Batchelor, E. Blasco, K. N. R. Wuest, H. Lu, M. Wegener, C. Barner-Kowollik and M. H. Stenzel, Chem. Commun., 2018, 54, 2436–2439 RSC.
- L. Sun, J. Ding, W. Xing, Y. Gai, J. Sheng and D. Zeng, Bioconjugate Chem., 2016, 27, 1200–1204 CrossRef CAS PubMed.
- R. Fay and J. P. Holland, Chem. - Eur. J., 2021, 27, 4893–4897 CrossRef CAS PubMed.
- C. Jamieson and K. Livingstone, The Nitrile Imine 1,3-Dipole, Springer Cham, 1 edn, 2020 Search PubMed.
- Y. Huseya, A. Chinone and M. Ohta, Bull. Chem. Soc. Jpn., 1971, 44, 1667–1668 CrossRef CAS.
- M. Märky, H. J. Hansen and H. Schmid, Helv. Chim. Acta, 1971, 54, 1275–1278 CrossRef.
- S. Kolodych, E. Rasolofonjatovo, M. Chaumontet, M.-C. Nevers, C. Créminon and F. Taran, Angew. Chem., Int. Ed., 2013, 52, 12056–12060 CrossRef CAS PubMed.
- H. Liu, D. Audisio, L. Plougastel, E. Decuypere, D.-A. Buisson, O. Koniev, S. Kolodych, A. Wagner, M. Elhabiri, A. Krzyczmonik, S. Forsback, O. Solin, V. Gouverneur and F. Taran, Angew. Chem., Int. Ed., 2016, 55, 12073–12077 CrossRef CAS PubMed.
- S. Wallace and J. W. Chin, Chem. Sci., 2014, 5, 1742–1744 RSC.
- L. Plougastel, O. Koniev, S. Specklin, E. Decuypere, C. Créminon, D.-A. Buisson, A. Wagner, S. Kolodych and F. Taran, Chem. Commun., 2014, 50, 9376–9378 RSC.
- L. Zhang, X. Zhang, Z. Yao, S. Jiang, J. Deng, B. Li and Z. Yu, J. Am. Chem. Soc., 2018, 140, 7390–7394 CrossRef CAS PubMed.
- X. Zhang, X. Wu, S. Jiang, J. Gao, Z. Yao, J. Deng, L. Zhang and Z. Yu, Chem. Commun., 2019, 55, 7187–7190 RSC.
- Z. Yao, X. Wu, X. Zhang, Q. Xiong, S. Jiang and Z. Yu, Org. Biomol. Chem., 2019, 17, 6777–6781 RSC.
- J. Gao, Q. Xiong, X. Wu, J. Deng, X. Zhang, X. Zhao, P. Deng and Z. Yu, Commun. Chem., 2020, 3, 29 CrossRef CAS.
- J. Deng, X. Wu, G. Guo, X. Zhao and Z. Yu, Org. Biomol. Chem., 2020, 18, 5602–5607 RSC.
- Q. Xiong, T. Zheng, X. Shen, B. Li, J. Fu, X. Zhao, C. Wang and Z. Yu, Chem. Sci., 2022, 13, 3571–3581 RSC.
- V. X. Truong and C. Barner-Kowollik, Trends Chem., 2022, 4, 291–304 CrossRef CAS.
- J. Li, H. Kong, L. Huang, B. Cheng, K. Qin, M. Zheng, Z. Yan and Y. Zhang, J. Am. Chem. Soc., 2018, 140, 14542–14546 CrossRef CAS PubMed.
- F. Wang, H. Kong, X. Meng, X. Tian, C. Wang, L. Xu, X. Zhang, L. Wang and R. Xie, RSC Chem. Biol., 2022, 3, 539–545 RSC.
- Z. Xi, H. Kong, Y. Chen, J. Deng, W. Xu, Y. Liang and Y. Zhang, Angew. Chem., Int. Ed., 2022, 61, e202200239 CAS.
- D. Fong, A. Lang, K. Li and A. Adronov, Macromolecules, 2020, 53, 1760–1766 CrossRef CAS.
- Y. Fu, H. Helbert, N. A. Simeth, S. Crespi, G. B. Spoelstra, J. M. van Dijl, M. van Oosten, L. R. Nazario, D. van der Born, G. Luurtsema, W. Szymanski, P. H. Elsinga and B. L. Feringa, J. Am. Chem. Soc., 2021, 143, 10041–10047 CrossRef CAS PubMed.
- J. Zheng, R. Yue, R. Yang, Q. Wu, Y. Wu, M. Huang, X. Chen, W. Lin, J. Huang, X. Chen, Y. Jiang, B. Yang and Y. Liao, Front. in Bioengineering and Biotechnol., 2022, 10 Search PubMed.
- D. Wu, H. Lei, X. Xie, L. Zhou, P. Zheng, Y. Cao and Y. Zhang, Nano Res., 2022, 15, 4294–4301 CrossRef CAS.
- F. Feist, J. P. Menzel, T. Weil, J. P. Blinco and C. Barner-Kowollik, J. Am. Chem. Soc., 2018, 140, 11848–11854 CrossRef CAS PubMed.
- F. Feist, L. L. Rodrigues, S. L. Walden, T. W. Krappitz, T. R. Dargaville, T. Weil, A. S. Goldmann, J. P. Blinco and C. Barner-Kowollik, J. Am. Chem. Soc., 2020, 142, 7744–7748 CrossRef CAS PubMed.
- J. P. Hooker, F. Feist, L. Delafresnaye, F. Cavalli, L. Barner and C. Barner-Kowollik, Chem. Commun., 2020, 56, 4986–4989 RSC.
- V. X. Truong, F. Li and J. S. Forsythe, ACS Macro Lett., 2017, 6, 657–662 CrossRef CAS PubMed.
- V. X. Truong, F. Li, F. Ercole and J. S. Forsythe, ACS Macro Lett., 2018, 7, 464–469 CrossRef CAS PubMed.
- T. Doi, H. Kawai, K. Murayama, H. Kashida and H. Asanuma, Chem. – Eur. J., 2016, 22, 10533–10538 CrossRef CAS PubMed.
- H. Lu, X. Wang, X. Zhou, W. Zhang and X. Wang, Polym. Chem., 2020, 11, 5992–5997 RSC.
- S. Ludwanowski, D. Hoenders, K. Kalayci, H. Frisch, C. Barner-Kowollik and A. Walther, Chem. Commun., 2021, 57, 805–808 RSC.
- C. Moriou, C. Denhez, O. Plashkevych, S. Coantic-Castex, J. Chattopadhyaya, D. Guillaume and P. Clivio, J. Org. Chem., 2015, 80, 615–619 CrossRef CAS PubMed.
- R. Lincoln, M. L. Bossi, M. Remmel, E. D'Este, A. N. Butkevich and S. W. Hell, Nat. Chem., 2022, 14, 1013–1020 CrossRef CAS PubMed.
- V. X. Truong, J. Bachmann, A.-N. Unterreiner, J. P. Blinco and C. Barner-Kowollik, Angew. Chem., Int. Ed., 2022, 61, e202113076 CrossRef CAS PubMed.
- D. Kodura, L. L. Rodrigues, S. L. Walden, A. S. Goldmann, H. Frisch and C. Barner-Kowollik, J. Am. Chem. Soc., 2022, 144, 6343–6348 CrossRef CAS PubMed.
- V. X. Truong and C. Barner-Kowollik, ACS Macro Lett., 2021, 10, 78–83 CrossRef CAS PubMed.
- A. Jemas, Y. Xie, J. E. Pigga, J. L. Caplan, C. W. am Ende and J. M. Fox, J. Am. Chem. Soc., 2022, 144, 1647–1662 CrossRef CAS PubMed.
- V. X. Truong, K. M. Tsang, F. Ercole and J. S. Forsythe, Chem. Mater., 2017, 29, 3678–3685 CrossRef CAS.
- C. Wang, H. Zhang, T. Zhang, X. Zou, H. Wang, J. E. Rosenberger, R. Vannam, W. S. Trout, J. B. Grimm, L. D. Lavis, C. Thorpe, X. Jia, Z. Li and J. M. Fox, J. Am. Chem. Soc., 2021, 143, 10793–10803 CrossRef CAS PubMed.
- Y. Zhang and F. M. Raymo, Bioconjugate Chem., 2020, 31, 1052–1062 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2022 |