Chemically-informed data-driven optimization (ChIDDO): leveraging physical models and Bayesian learning to accelerate chemical research†
Abstract
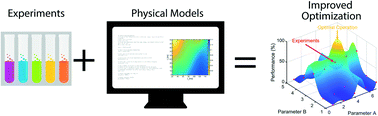
Current methods of finding optimal experimental conditions, Edisonian systematic searches, often inefficiently evaluate suboptimal design points and require fine resolution to identify near optimal conditions. For expensive experimental campaigns or those with large design spaces, the shortcomings of the status quo approaches are more significant. Here, we extend Bayesian optimization (BO) and introduce a chemically-informed data-driven optimization (ChIDDO) approach. This approach uses inexpensive and low-fidelity information obtained from physical models of chemical processes and subsequently combines it with expensive and high-fidelity experimental data to optimize a common objective function. Using common optimization benchmark objective functions, we describe scenarios in which the ChIDDO algorithm outperforms the traditional BO approach, and then implement the algorithm on a simulated electrochemical engineering optimization problem.
- This article is part of the themed collections: Emerging Investigator Series and Digitalization in Reaction Engineering


 Please wait while we load your content...
Please wait while we load your content...