Open Access Article
Open Access ArticleApplication of scanning electrochemical microscopy for topography imaging of supported lipid bilayers†
Zahra
Nasri
*a,
Seyedali
Memari
ab,
Johanna
Striesow
a,
Klaus-Dieter
Weltmann
a,
Thomas
von Woedtke
 ac and
Kristian
Wende
ac and
Kristian
Wende
 *a
*a
aCenter for Innovation Competence (ZIK) plasmatis, Leibniz Institute for Plasma Science and Technology (INP), Greifswald, Germany. E-mail: zahra.nasri@inp-greifswald.de; kristian.wende@inp-greifswald.de
bDepartment of Chemistry, University of Hamburg, Hamburg, Germany
cInstitute for Hygiene and Environmental Medicine, University Medicine Greifswald, Greifswald, Germany
First published on 24th February 2022
Abstract
Oxidative stress in cellular environments may cause lipid oxidation and membrane degradation. Therefore, studying the degree of lipid membrane morphological changes by reactive oxygen and nitrogen species will be informative in oxidative stress-based therapies. This study introduces the possibility of using scanning electrochemical microscopy as a powerful imaging technique to follow the topographical changes of a solid-supported lipid bilayer model induced by reactive species produced from gas plasma. The introduced strategy is not limited to investigating the effect of reactive species on the lipid bilayer but could be extended to understand the morphological changes of the lipid bilayer due to the action of membrane proteins or antimicrobial peptides.
Introduction
Pro-oxidant therapies, such as radiotherapy, chemotherapy, and phototherapy, employ sudden and intense oxidative events by elevating reactive oxygen and nitrogen species (RONS) levels in cancer cells to induce their death.1 Cold physical plasma (CPP) is a partially ionized gas at body temperature producing similar levels of RONS and could present a promising alternative for the treatment of tumors.2 Besides cancer therapy, CPP's medical applications cover chronic wounds and other skin-related diseases, including precancerous lesions.3 It is adopted that CPP-generated RONS, such as hydrogen peroxide, ozone, hydroxyl radical, superoxide, singlet oxygen, atomic oxygen, peroxynitrite, and nitric oxide, are the critical elements in the mentioned CPP applications.4 The CPP-generated reactive species will be produced or transported across the plasma-target interface and subsequently propagate into the living tissue according to their lifetime.5 Thus, the cell membrane is among the first targets for the CPP-produced RONS. However, the related mechanisms for the delivery of RONS into the cells are not entirely known and are the subject of many studies.6 Moreover, RONS are involved in many processes such as redox signaling, aging, carcinogenesis, and neurodegeneration.7 As lipid membrane is the target of RONS attack,8 studying their interactions with lipid bilayer is of considerable interest. In persuading this, and due to the significant complexity of cell membranes and the huge diversity of lipids, simplified models usually comprising one phospholipid type have been developed. The most well-known biomimetic systems used for such purposes are lipid vesicles, lipid monolayers, and supported lipid bilayers (SLB).9 SLBs are lipid bilayer films immobilized on solid supports that can mimic the key features of the cell membranes.10 Moreover, they are compatible with various surface-sensitive techniques such as electrochemical methods and scanning electrochemical microscopy (SECM). SECM is a powerful and versatile electrochemical imaging platform with high temporal and spatial resolutions.11,12 In comparison with other scanning probe techniques, SECM enables topographical surface imaging and gathers chemical information with high spatial resolution.13 Although considerable progress has been made in the analysis of biological samples, following the local topography of the membrane is still a challenge. So far, SECM has been used to characterize the changes in the membrane permeability of the cells using permeable electrochemical mediators.14,15 Also, high resolution imaging of ion transport through porous nanocrystalline silicon membrane by SECM is reported.16 Herein, we deploy a robust and straightforward strategy using SECM to elucidate the topographic changes at the model supported lipid bilayer under oxidative stress. Moreover, the enzymatic damage of a membrane protein (phospholipase A2/PLA2) on the model lipid bilayer is studied, which could be extended to investigate the action of other membrane proteins or antimicrobial peptides on the supported lipid bilayer.Experimental
Reagents
1,2-Dioleoyl-sn-glycero-3-phosphoethanolamine (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (Δ9-cis) PE (DOPE)) was from Avanti Polar Lipids and ordered from Otto Nordwald (Otto Nordwald GmbH, Germany). HPLC grade chloroform and ethanol were purchased from Carl Roth (Carl Roth GmbH + Co. KG, Germany). Methanol (99.95%) was purchased from Th. Geyer (Th. Geyer GmbH & Co. KG, Germany). Phospholipase A2 from honey bee venom (Apis mellifera, UniProt P00630), cysteine and potassium hexacyanoferrate(II) trihydrate were purchased from Sigma-Aldrich (Sigma-Aldrich Chemie GmbH, Germany). Ferrocenemethanol was from Acros Organics and ordered from Fisher Scientific (Fisher Scientific GmbH, Germany). All other reagents were of analytical grade and used without further purification.
1 (Δ9-cis) PE (DOPE)) was from Avanti Polar Lipids and ordered from Otto Nordwald (Otto Nordwald GmbH, Germany). HPLC grade chloroform and ethanol were purchased from Carl Roth (Carl Roth GmbH + Co. KG, Germany). Methanol (99.95%) was purchased from Th. Geyer (Th. Geyer GmbH & Co. KG, Germany). Phospholipase A2 from honey bee venom (Apis mellifera, UniProt P00630), cysteine and potassium hexacyanoferrate(II) trihydrate were purchased from Sigma-Aldrich (Sigma-Aldrich Chemie GmbH, Germany). Ferrocenemethanol was from Acros Organics and ordered from Fisher Scientific (Fisher Scientific GmbH, Germany). All other reagents were of analytical grade and used without further purification.
Preparation of gold supported lipid bilayer
Langmuir–Blodgett (LB) and Langmuir–Schaefer (LS) deposition techniques were employed for the transfer of the first and second lipid monolayers onto the Au(111) (arrandee metal GmbH + Co. KG, Germany) substrate to prepare the gold supported lipid bilayer. Surface pressure-area isotherms were monitored during the transfer using a Langmuir trough (KSV NIMA, LOT-QuantumDesign GmbH, Germany). The Au (111) substrate was first flame-annealed, cleaned in the piranha solution, and then transferred into the trough.17 In the next step, a sufficient quantity of 1 mg mL−1 lipid solution in a mixture of 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 chloroform–methanol was drop wised onto the surface of the subphase. The solvent was allowed to evaporate, and then the barriers were closed to obtain the target pressure of 37 mN m−1. Next, the gold substrate was raised through the trough at the speed of 2 mm min−1, while the pressure was constant at the target pressure. Then, the substrate was dried in an argon atmosphere for 30 min. A transfer ratio of 1.0 ± 0.1 indicated a successful transfer of the lipid to the substrate. Next, LS deposition (horizontal dip) was performed at the speed of 0.5 mm min−1 to transfer the second monolayer. Finally, the obtained gold supported lipid bilayer was placed immediately into the electrochemical cell.
1 chloroform–methanol was drop wised onto the surface of the subphase. The solvent was allowed to evaporate, and then the barriers were closed to obtain the target pressure of 37 mN m−1. Next, the gold substrate was raised through the trough at the speed of 2 mm min−1, while the pressure was constant at the target pressure. Then, the substrate was dried in an argon atmosphere for 30 min. A transfer ratio of 1.0 ± 0.1 indicated a successful transfer of the lipid to the substrate. Next, LS deposition (horizontal dip) was performed at the speed of 0.5 mm min−1 to transfer the second monolayer. Finally, the obtained gold supported lipid bilayer was placed immediately into the electrochemical cell.
Electrochemical measurements
Electrochemical measurements were carried out using an AUTOLAB PGSTAT302N bipotentiostat (Deutsche METROHM GmbH & Co. KG, Germany). All experiments were performed in a three-electrode system consisting of a leakless Ag/AgCl miniature reference electrode, a platinum wire auxiliary electrode (eDAQ Europe, Poland), and the gold supported lipid bilayer as the working electrode. Differential pulse voltammograms (DPVs) were recorded in a solution of 10 mM K4[Fe(CN)6] in phosphate-buffered saline (PBS).SECM measurements
SECM experiments were performed using a Sensolytics SECM workstation (Sensolytics GmbH, Germany) coupled with the AUTOLAB PGSTAT302N bipotentiostat with an ECD module providing minimum current resolution of 0.3 fA, a piezoelectric positioning system, and a video for visual control of tip prepositioning. SECM measurements, as well as cyclic voltammograms (CVs) were carried out in 0.5 mM ferrocenemethanol solution in 100 mM KCl using 1 μm diameter platinum ultra-microelectrode tip (Sensolytics, Germany), and a Ag/AgCl (sat. KCl) (QCM, Deutsche METROHM GmbH & Co. KG, Germany) and a platinum wire as the reference and counter electrode, respectively.CPP treatment
The kINPen09 (neoplas tools GmbH, Germany) was applied as a well-characterized plasma source2,18 using pure argon (Air Liquide, 99.999%) as the feed gas running at a flux of 3.0 standard liters per minute (slm). The gold supported lipid bilayer electrode was placed at the bottom of the electrochemical cell filled with 2.0 mL of PBS solution. Plasma treatment was done at a distance of 9 mm between the jet nozzle and the buffer surface for 5 and 10 min plasma treatment. The electrochemical cell was connected to a peristaltic pump to compensate for the water evaporation during plasma treatment and keep the buffer volume constant.Lipid bilayer extraction
A volume of 2 mL methyl tert-butyl ether (MTBE) solution was added to the DOPE lipid bilayer. After shaking the solution for 15 min, aqueous and organic phases appeared. Then the MTBE phase was separated, and MTBE was removed under the nitrogen stream. The obtained lipid film was kept at −80 °C for further analysis.Liquid chromatography-tandem mass spectrometry (LC-MS) acquisition of lipids
Lipid films were resuspended in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 chloroform
4 chloroform![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) methanol
methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) isopropanol (v/v/v). DOPE and lipid oxidation products were separated on a Vanquish UHPLC with a reversed-phase C30 column (150 × 2.1 mm, 2.6 μm) equipped with a pre-column (10 × 2.1 mm, all obtained from Thermo Fisher, Germany). The column oven temperature was 50 °C. The 23 min gradient used was the following: 0–1 min 30% B, 1–2 min ramp to 50% B, 2–15 min ramp to 99% B, hold 99% B for 5 min, then decrease to 30% B until 23 min. The flow rate was set to 350 μL min−1, and the injection volume was 2 μL. Separated lipids were acquired on a QExactive Plus mass spectrometer equipped with a HESI II ionization source (both from Thermo Fisher, Germany). Lipids were measured in positive and negative ionization mode with a top 10 DDA method. Resolution in MS1 was 70.000 (at 200 m/z) with an AGC target of 1 × 106 and an ion time of 100 ms. Scan range was from 100 to 1000 m/z. Precursors with the highest abundance were subjected to fragmentation at a resolution of 17.500 (at 200 m/z) with an AGC target of 1 × 105 and an ion time of 50 ms. A normalized fragmentation energy was applied from 22 to 26%. The minimum AGC target was 8 × 103, and dynamic exclusion was 10 s.
isopropanol (v/v/v). DOPE and lipid oxidation products were separated on a Vanquish UHPLC with a reversed-phase C30 column (150 × 2.1 mm, 2.6 μm) equipped with a pre-column (10 × 2.1 mm, all obtained from Thermo Fisher, Germany). The column oven temperature was 50 °C. The 23 min gradient used was the following: 0–1 min 30% B, 1–2 min ramp to 50% B, 2–15 min ramp to 99% B, hold 99% B for 5 min, then decrease to 30% B until 23 min. The flow rate was set to 350 μL min−1, and the injection volume was 2 μL. Separated lipids were acquired on a QExactive Plus mass spectrometer equipped with a HESI II ionization source (both from Thermo Fisher, Germany). Lipids were measured in positive and negative ionization mode with a top 10 DDA method. Resolution in MS1 was 70.000 (at 200 m/z) with an AGC target of 1 × 106 and an ion time of 100 ms. Scan range was from 100 to 1000 m/z. Precursors with the highest abundance were subjected to fragmentation at a resolution of 17.500 (at 200 m/z) with an AGC target of 1 × 105 and an ion time of 50 ms. A normalized fragmentation energy was applied from 22 to 26%. The minimum AGC target was 8 × 103, and dynamic exclusion was 10 s.
Results and discussion
Effect of plasma treatment on the permeability of the model lipid bilayer
In order to determine the effect of CPP on the permeability of the lipid bilayer, model lipid bilayers from 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE) were transferred onto the gold electrode surface by Langmuir–Blodgett and Langmuir–Schaefer deposition techniques. Optimum deposition pressure was 37 mN m−1, where significant changes in the plot of surface pressure as a function of area per lipid molecule (π–A isotherm) were observed. The successful transfer of the lipid bilayer to the electrode surface (gold) was confirmed by recording the DPV of K4[Fe(CN)6] as the redox probe. As indicated in Fig. 1, the recorded DPV showed no significant peak current, pointing out that the transferred lipid bilayer was blocking the redox probe's access to the electrode surface, demonstrating that the gold surface was fully covered with the lipid bilayer. Then, the gold supported lipid bilayers were placed at the bottom of the electrochemical cells filled with 2.0 mL of PBS and treated for 5 min and 10 min with an argon plasma. After washing the lipid bilayers to remove the plasma-generated reactive species, the DPVs of plasma-treated gold supported lipid bilayers were recorded. The observed increase in the DPVs current indicated the enhanced permeability of the lipid bilayer due to the activity of plasma-produced RONS and subsequently easier access of the redox probe to the gold surface.19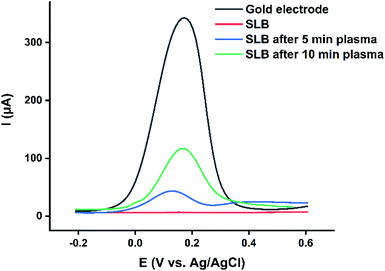 | ||
| Fig. 1 DPVs of the gold electrode, and gold supported lipid bilayer electrodes (SLB) before and after 5 and 10 min plasma treatment recorded in 10 mM K4[Fe(CN)6] at the scan rate of 0.05 V s−1. | ||
Topography imaging by SECM
The experiments were designed based on the positive feedback mode of the SECM, in which suitable potentials were applied to the tip and substrate. As the substrate was the gold supported lipid bilayer, and the lipids were on the surface of gold only by hydrophilic interactions, applying a continuous potential to the substrate led to the detachment of the lipids. Based on that, a new strategy was employed: first, the gold supported lipid bilayer was placed at the bottom of the electrochemical cell filled with 2.0 mL of PBS and treated with a distance of 9 mm between the plasma jet nozzle and the buffer surface. Then, the treated lipid bilayer was washed with PBS gently using a peristaltic pump to remove excess reactive species. After that, a solution of cysteine in PBS was injected on the surface of the bilayer, realizing a final concentration of 1 mM. The cysteine molecules have free thiol groups that can self-assemble and orient on the gold surface, wherever there are hole-like structures in the lipid bilayer due to the plasma treatment. Where the bilayer remained intact, access of the cysteine molecules to the gold surface was denied. After overnight incubation, the lipid bilayer was washed with isopropanol and ethanol several times to remove the lipids from the surface of the gold electrode. With that procedure, holes in the lipid layer due to plasma treatment were replaced by cysteine molecules (Scheme 1). The resulting substrate was transferred to the SECM setup for topography imaging.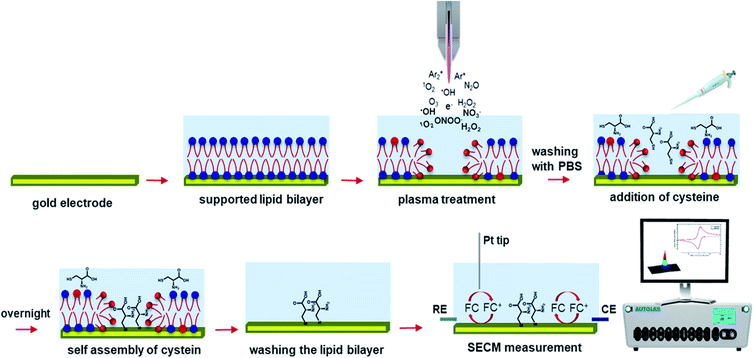 | ||
| Scheme 1 Schematic illustration of the strategy used to record the topography image of the model lipid bilayer after plasma treatment. | ||
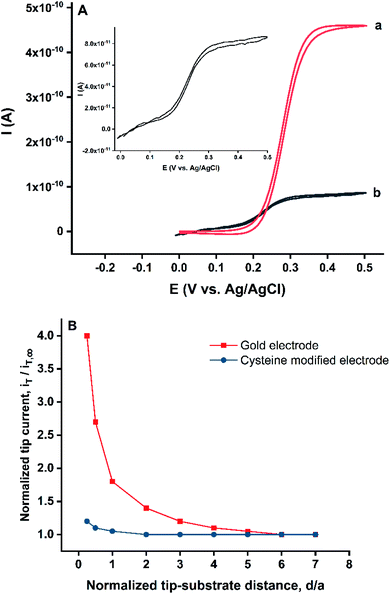
The gold electrode was placed as the substrate at the bottom of the SECM cell, filled with a solution of 0.5 mM ferrocenemethanol (FcMeOH) in 100 mM KCl. The 1 μm diameter platinum tip was used to approach the substrate. The tip was positioned at two distances, far from the substrate and at the surface of the substrate. The feedback mode of SECM was employed by applying the potential of −0.1 V to the substrate where the tip current was monitored. The CVs of the probe when the tip was located in close proximity to the substrate and when the tip was far from the gold surface were recorded and presented in Fig. 2A. The current of recorded CV at the surface of the gold electrode (position a) was enhanced significantly compared to the CV at the bulk (position b) because of the regeneration of FcMeOH that happened at the gold surface in this potential.
The approach curves were plotted by holding the tip and the substrate at potentials of 0.5 V and −0.1 V, respectively. The results indicated the notable increase of the tip signal at the surface of the electrode. The same approach curve was plotted for the cysteine modified gold electrode. Where the gold surface was covered with cysteine, the regeneration of FcMeOH did not take place to a significant extent (Fig. 2B). The negligible signal amplification in cysteine modified electrode is the consequence of the partial permeability of FcMeOH into the cysteine layer.20 Thus, by scanning the surface of the substrate by the tip and applying the appropriate potentials to the tip and substrate, it is possible to recognize the cysteine molecules on the gold substrate based on the drop in the probe current.
The constant-height mode of SECM was employed to obtain the array scans. Before any array scan experiment, the tilt control was performed by recording three approach curves at different substrate locations to examine the direction of the sample's tilt. The tilt control minimizes the influences of the tilted substrates on the current signal for the constant-height imaging mode of SECM. After that, the tip was positioned in the feedback interaction range. The constant height of 500 nm from the substrate (d/a = 1) was achieved using a piezoelectric positioning system with the max speed of 10 nm s−1 and the step width of 5 nm. This working distance led to the desired imaging contrast.
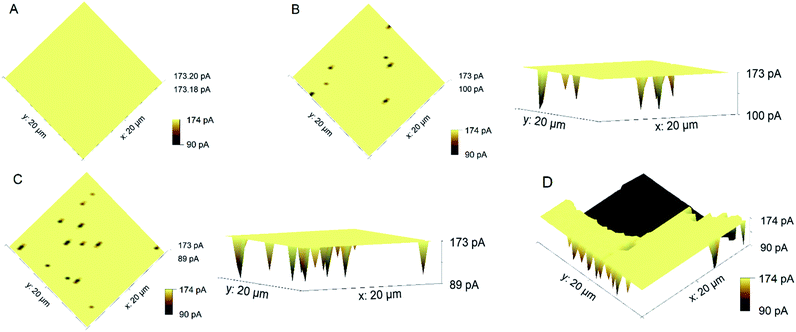
The obtained array scans of the untreated and plasma-treated gold supported lipid bilayers for 5 and 10 min were depicted in Fig. 3. The untreated lipid bilayer was blocking the access of cysteine molecules to the surface of gold. Thus, after the washing step, there was only the gold electrode, and the observed current is related to the regeneration of FcMeOH at the gold. After plasma treatment, several holes were induced, where the access of cysteine to the gold surface was possible. The observed drop in the current of recorded array scans (Fig. 3B and C) indicated the presence of holes in the lipid bilayer after plasma treatment. The results indicated that the plasma treatment led to pore formation which their number and size were dependent on the treatment time. As the resolution of the technique is limited to the size of the tip, pores in the order of micrometers were detectable. However, potential nanopores were not observed due to the resolution limit.21–23 Here also, the detection of nanopores is maybe possible by switching to the nanoelectrodes. Imaging the self-assembled monolayers on gold using SECM was reported earlier24–26 but was not employed for the topography imaging of supported lipid bilayers.
Another application of scanning probe techniques is to study the protein's behavior in media that mimic in vivo systems. Phospholipase A2 (PLA2) consists of membrane proteins that cleave the glycerophospholipids to fatty acids and lysophospholipids.27 Recently, we have studied the PLA2 activity to damage the model lipid bilayer.28 In this regard, the introduced method was employed to visualize the degradation of the lipid bilayer due to the action of PLA2. A solution of 10−4 mg mL−1 PLA2 in Tris buffer [0.010 M Tris (pH 8.9), 0.150 M NaCl, 0.005 M CaCl2] was in contact with the lipid bilayer for four hours. After that, the changes in the topography of the lipid bilayer were monitored using the introduced method (Fig. 3D). It is reported that hydrolysis of the lipid bilayer by PLA2 is initiated at regions where there are traces of structural defects or depressions in the bilayer. After that, the existing structural defects are enlarged as the top lipid layer is hydrolyzed and the bottom layer spontaneously desorbs.29 Here also, a large hole in the lipid bilayer was observed due to the action of the enzyme. The obtained topography image supports the applicability of the introduced method to monitor the morphological changes of the model lipid bilayer due to the activity of membrane proteins or antimicrobial peptides using SECM.
LC-MS analysis of lipids
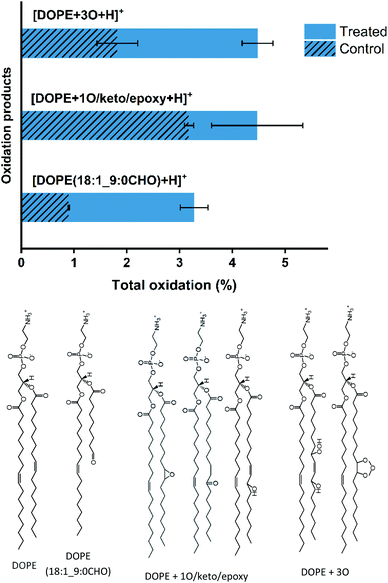
To demonstrate that the observed increase of permeability and topographic changes of the lipid bilayer is the result of lipid oxidation by RONS, high-resolution LC-MS profiling was performed. In Fig. 4, the type and total percentage of oxidations induced by plasma on DOPE lipid bilayer after 10 min plasma treatment are shown, and the structure of the oxidation products of [DOPE(18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1_9
1_9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0CHO)], [DOPE + 1O/keto/epoxy] and [DOPE + 3O] are detected. The corresponding extracted ion chromatograms and MS/MS fragment spectra are presented in Fig. S1.†
0CHO)], [DOPE + 1O/keto/epoxy] and [DOPE + 3O] are detected. The corresponding extracted ion chromatograms and MS/MS fragment spectra are presented in Fig. S1.†
The plasma-induced reactive species primarily attacked the unsaturated bond at the C-9 position of DOPE due to the presence of two allylic hydrogen atoms at C-8 and C-11. The abstraction of allylic hydrogen atoms led to the addition of plasma-produced reactive species, such as atomic oxygen and OH radicals. Subsequent rearrangements (Hock cleavage, homolytic β-scission, etc.) led to the formation of aldehyde products.30 The oxidation of lipids to the aldehyde products with shortened chains promotes direct defects on the membrane and can be considered a precursor for pore formation. When the concentration of truncated-chain lipids increases, micelles form. Their desorption paves the way to the observed pore formation in the lipid bilayer.22,23,31
Conclusions
The obtained results confirm that the provided method allows the topography imaging of the model supported lipid bilayer under oxidative stress or due to the action of membrane proteins. The utilization of SECM enabled the evaluation of the micrometer scale membrane pores in the plasma-treated lipid bilayer, which is limited to the diameter of the platinum tip. Additionally, high-resolution LC-MS confirmed that aldehyde lipoxidation products are related to the RONS-induced pore formation in lipid bilayers.Author contributions
Z. N., S. M., and K. W. conceived and designed the experiments. Z. N. and S. M. performed all electrochemical and SECM experiments. J. S. designed and performed LC-MS analysis of lipids. All authors discussed the results. Z. N. and K. W. conceived and wrote the main manuscript and prepared figures. All authors reviewed the manuscript.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work is funded by the German Federal Ministry of Education and Research (BMBF) (grant numbers 03Z22DN12).Notes and references
- G. Manda, G. Isvoranu, M. V. Comanescu, A. Manea, B. Debelec Butuner and K. S. Korkmaz, Redox Biol., 2015, 5, 347–357 CrossRef CAS PubMed.
- S. Reuter, T. von Woedtke and K. D. Weltmann, J. Phys. D: Appl. Phys., 2018, 51, 233001–233051 CrossRef.
- T. von Woedtke, A. Schmidt, S. Bekeschus, K. Wende and K. D. Weltmann, In Vivo, 2019, 33, 1011–1026 CrossRef CAS PubMed.
- K. D. Weltmann and T. von Woedtke, Plasma Phys. Controlled Fusion, 2017, 59, 014031–014041 CrossRef.
- A. M. Hirst, F. M. Frame, M. Arya, N. J. Maitland and D. O'Connell, Tumor Biology, 2016, 37, 7021–7031 CrossRef CAS PubMed.
- G. Bauer, D. Sersenova, D. B. Graves and Z. Machala, Sci. Rep., 2019, 9, 14210 CrossRef PubMed.
- R. M. Cordeiro, Biochim. Biophys. Acta, 2014, 1838, 438–444 CrossRef CAS PubMed.
- A. Bogaerts, M. Yusupov, J. Razzokov and J. Van der Paal, Front. Chem. Sci. Eng., 2019, 13, 253–263 CrossRef CAS.
- J. Sarkis and V. Vie, Front. Bioeng. Biotechnol., 2020, 8, 270 CrossRef PubMed.
- E. T. Castellana and P. S. Cremer, Surf. Sci. Rep., 2006, 61, 429–444 CrossRef CAS PubMed.
- T.-E. Lin, S. Rapino, H. H. Girault and A. Lesch, Chem. Sci., 2018, 9, 4546–4554 RSC.
- M. A. O'Connell and A. J. Wain, Anal. Methods, 2015, 7, 6983–6999 RSC.
- F. Conzuelo, A. Schulte and W. Schuhmann, Proc. Math. Phys. Eng. Sci., 2018, 474, 20180409 Search PubMed.
- F. P. Filice, J. D. Henderson, M. S. M. Li and Z. Ding, ACS Omega, 2019, 4, 2142–2151 CrossRef CAS PubMed.
- F. P. Filice, M. S. M. Li, J. M. Wong and Z. Ding, J. Inorg. Biochem., 2018, 182, 222–229 CrossRef CAS PubMed.
- M. Shen, R. Ishimatsu, J. Kim and S. Amemiya, J. Am. Chem. Soc., 2012, 134, 9856–9859 CrossRef CAS PubMed.
- E. Madrid and S. L. Horswell, J. Electroanal. Chem., 2018, 819, 338–346 CrossRef CAS.
- M. S. Mann, R. Tiede, K. Gavenis, G. Daeschlein, R. Bussiahn, K.-D. Weltmann, S. Emmert, T. v. Woedtke and R. Ahmed, Clinical Plasma Medicine, 2016, 4, 35–45 CrossRef.
- F. Scholz, G. Lopez de Lara Gonzalez, L. Machado de Carvalho, M. Hilgemann, K. Z. Brainina, H. Kahlert, R. S. Jack and D. T. Minh, Angew. Chem., Int. Ed. Engl., 2007, 46, 8079–8081 CrossRef CAS PubMed.
- C. Cannes, F. Kanoufi and A. J. Bard, J. Electroanal. Chem., 2003, 547, 83–91 CrossRef CAS.
- R. Tero, Y. Suda, R. Kato, H. Tanoue and H. Takikawa, Appl. Phys. Express, 2014, 7, 077001 CrossRef CAS.
- R. Tero, R. Yamashita, H. Hashizume, Y. Suda, H. Takikawa, M. Hori and M. Ito, Arch. Biochem. Biophys., 2016, 605, 26–33 CrossRef CAS PubMed.
- M. Ravandeh, G. Coliva, H. Kahlert, A. Azinfar, C. A. Helm, M. Fedorova and K. Wende, Biomolecules, 2021, 11, 276 CrossRef CAS PubMed.
- G. Wittstock, R. Hesse and W. Schuhmann, Electroanalysis, 1997, 9, 746–750 CrossRef CAS.
- A. Lesch, B. Vaske, F. Meiners, D. Momotenko, F. Cortés-Salazar, H. H. Girault and G. Wittstock, Angew. Chem., Int. Ed., 2012, 51, 10413–10416 CrossRef CAS PubMed.
- B. Vaske, M. Schaube, F. Meiners, J. H. Ross, J. Christoffers and G. Wittstock, ChemElectroChem, 2021, 8, 3192–3202 CrossRef CAS.
- E. A. Dennis, J. Cao, Y. H. Hsu, V. Magrioti and G. Kokotos, Chem. Rev., 2011, 111, 6130–6185 CrossRef CAS PubMed.
- Z. Nasri, S. Memari, S. Wenske, R. Clemen, U. Martens, M. Delcea, S. Bekeschus, K.-D. Weltmann, T. von Woedtke and K. Wende, Chem.–Eur. J., 2021, 27, 14702–14710 CrossRef CAS PubMed.
- K. Balashev, N. John DiNardo, T. H. Callisen, A. Svendsen and T. Bjornholm, Biochim. Biophys. Acta, 2007, 1768, 90–99 CrossRef CAS PubMed.
- M. Ravandeh, H. Kahlert, H. Jablonowski, J. W. Lackmann, J. Striesow, V. Agmo Hernandez and K. Wende, Sci. Rep., 2020, 10, 18683 CrossRef CAS PubMed.
- T. M. Tsubone, H. C. Junqueira, M. S. Baptista and R. Itri, Biochim. Biophys. Acta, Biomembr., 2019, 1861, 660–669 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/d2ay00154c |
| This journal is © The Royal Society of Chemistry 2022 |