Open Access Article
Open Access ArticleOxygen tolerant, photoinduced controlled radical polymerization approach for the synthesis of giant amphiphiles†
Alexis
Theodorou
 a,
Petros
Mandriotis
a,
Athina
Anastasaki
a,
Petros
Mandriotis
a,
Athina
Anastasaki
 b and
Kelly
Velonia
b and
Kelly
Velonia
 *a
*a
aLaboratory of Synthetic Biomaterials, Department of Materials Science and Technology, University of Crete, 70013 Heraklion, Crete, Greece. E-mail: velonia@materials.uoc.gr
bLaboratory of Polymeric Materials, Department of Materials, ETH Zurich, Vladimir-Prelog-Weg-5, 8093 Zurich, Switzerland. E-mail: athina.anastasaki@mat.ethz.ch
First published on 18th March 2021
Abstract
Amphiphilic protein–polymer conjugates form hybrid nanostructures with a broad range of potential applications. Herein, we expand the scope of amphiphilic protein–polymer bioconjugates via an oxygen tolerant, photoinduced RDRP method. A range of hydrophobic acrylic, methacrylic and acrylamide-based monomers are comprehensively screened to identify optimum reaction conditions that lead to quantitative biomacroinitiator consumption. By judiciously varying a number of components (e.g. monomer loading, catalyst concentration, pH, co-solvent, salt content), well-defined protein–polymer conjugates bearing different polymer moieties were obtained and their intrinsic properties were expressed through both their self-assembled architectures and esterase-like catalytic activity.
Introduction
Efficient construction of multifunctional, ordered nanoscale assemblies bearing intact biological building blocks is among the challenges of contemporary materials science.1 Amphiphilic protein–polymer conjugates, giant amphiphiles, have evolved to address this issue by merging the inherent physicochemical properties of both a biological and a synthetic component in a spontaneously forming nanoparticle.2 Such biopolymer assemblies are by design promising materials for bio- and nano-technological and medical applications such as delivery systems (e.g. protein or drug delivery), nanoreactors, biosensing and detection.3–10Ever since Abuchowski and Davis showed that polyethylene glycol (PEG) can be directly attached to Bovine Serum Albumin (BSA),11 a plethora of PEGylated,12–15 glycopolymer,16 responsive polymer17,18 or zwitterionic polymer19 bearing water-soluble biohybrids has been synthesized via direct bioconjugation processes. This merging of biological with synthetic matter often led to biohybrids with enhanced biochemical or material properties such as increased proteolytic resistance, increased thermal or pH stability or, increased activity.20 Despite employing the vast knowledge generated from the synthesis of hydrophilic bioconjugates, the synthesis of self-assembling, amphiphilic bioconjugate chimeras suffers from several shortcomings. Direct bioconjugation via orthogonal couplings (grafting to approach) mostly entails reaction between a pre-synthesized, orthogonally end-functionalized polymer, with one or more complementary reactive sites exposed on the protein surface.21,22 The hydrophobicity of the polymer mitigates its reactivity in aqueous solutions and a high percentage of an organic cosolvent is frequently employed to allow a more efficient reaction between unimers.23,24 As a result, low bioconjugation yields, tedious purification protocols and/or protein denaturation have often been reported.2,25
More recently, grafting from techniques have been used to synthesize hydrophilic bioconjugates. Reversible Deactivation Radical Polymerization (RDRP) approaches26–30 such as Atom Transfer Radical Polymerization (ATRP)31–38 or Reversible Addition–Fragmentation chain Transfer polymerization (RAFT)32,39–43 and olefin metathesis44 led to the synthesis of water-soluble protein–polymer conjugates in high yields under biologically relevant conditions. In 2008, we first reported on the synthesis of an amphiphilic protein–polymer conjugate in quantitative yields via ATRP grafting of styrene from a protein macroinitiator under conventional ATRP conditions.45 We demonstrated that this approach allowed isolation of the biohybrids through a simple dialysis step and, highlighting its efficiency, we also reported on the synthesis of chemically addressable giant amphiphiles46 and the in situ formation of nanoreactors and nanocarriers. This methodology was also found efficient for the synthesis of amphiphilic protein–polymer conjugates via ATRP grafting of the water-soluble monomer 2-hydroxypropyl methacrylate (HPMA).47–49
However, in practice the above-mentioned conventional polymerization techniques require specialized equipment that can be expensive, trained personnel and, most often, deoxygenation which, in the case of biohybrids could prove disruptive to proteins.50,51 Oxygen inhibits RDRPs by reacting with the propagating radicals to form less efficient peroxy-radicals and hydroperoxides while, in the case of ATRP, oxygen oxidizes the catalyst resulting in its rapid deactivation.52 The need to simplify polymerization protocols has led in the recent years to a surge of interest in developing new strategies for overcoming the deoxygenation procedures required for conventional RDRP. Recent literature reports elegantly demonstrate that the effects of oxygen triggered polymerization inhibition can be restricted in the presence of sacrificial organic compounds53 reducing agents54–56 or enzymes (e.g. glucose oxidase)57,58 that allow synthesis of low dispersity polymers. To implement such oxygen tolerant approaches to the synthesis of hydrophilic protein–polymer conjugates, reducing agents such as ascorbic acid58,59 or a biocatalytic system composed of glucose oxidase (GOx) and horseradish peroxidase (HRP) were previously employed.53,60 To eliminate the need of additives, scalable procedures which result in the controlled, copper mediated RDRP, (photo)polymerization of a wide range of monomers by simply adjusting the headspace of a reaction vessel were developed.61–68 Taking advantage of these recent reports we developed an oxygen tolerant, photoinduced, versatile RDRP grafting from approach to synthesize protein–polymer conjugates in quantitative yields.69 This simplified RDRP approach allowed grafting of styrene as well as of water soluble monomers from a variety of protein macroinitiators using ppm levels of copper(II) under UV, blue or sunlight irradiation within two hours. However, grafting of hydrophobic acrylates and methacrylates from proteins, which would give access to an unprecedented variety of giant amphiphiles via a single approach and pave the way for novel applications was not investigated.
Herein, we present a comparative new study that expands the applicability of the oxygen tolerant, photoinduced RDRP approach to a variety of hydrophobic and/or responsive acrylates, methacrylates and acrylamides (Scheme 1). We judiciously investigate the factors that allow tailoring the grafting of these monomer families to quantitative conversions expanding the scope of this method. It should be noted that by aiming at quantitative macroinitiator conversions we ensure at the same time a simple purification step which, together with the simplicity of this oxygen tolerant, photoinduced RDPR protocol, renders the polymerization accessible to all. We further demonstrate that the nature of the monomer/polymer has a direct effect on bioconjugate activity and assembly.
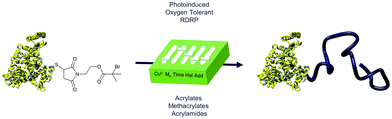 | ||
| Scheme 1 Schematic representation of the oxygen tolerant, photoinduced RDRP approach expanding the scope of amphiphilic protein–polymer conjugates. | ||
Results and discussion
Biomacroinitiator and synthetic approach
Bovine serum albumin (BSA), the most abundant translocator protein in blood, was used as model protein.70 BSA is a low-cost, stable globular protein, with remarkable ligand-binding properties and significant medical importance stemming from its high structural homology with human serum albumin (HSA). Its unique, accessible cysteine (Cys 34) allowed for the quantitative preparation of a biomacroinitiator (BSA-Br, Io) via a thiol Michael addition (ESI, Fig. S1†).45,69 To graft hydrophobic acrylates from BSA-Br (Io), the conditions reported in our recent study for the oxygen tolerant, photoinduced RDRP were used as starting point.69 More specifically, the grafting of styrene, its p-methoxy and m-nitro derivatives and the water soluble (oligo(ethylene glycol) methyl ether acrylate (OEOA480), 2-(dimethylamino)ethyl methacrylate, (DMAEMA) and acrylamide was reported to optimally proceed under UV light irradiation using 22 ppm Cu(II) and [monomer]/[BSA-Br]/[CuII]/[ligand] feed ratio equal to 2000/1/1.5/12. Tris[2-(dimethylamino)ethyl]amine (Me6TREN) was used as ligand in a 8-fold excess with respect to copper(II) bromide as it was previously proven essential for the in situ reduction of Cu(II) to Cu(I).71,72 All reactions were carried out under broad band UV irradiation (ca. 365 nm, 36 Watt), in 5 mL plastic syringes which were hermitically capped upon the addition of all reagents to ensure absence of headspace. Size Exclusion Chromatography (SEC) and native Polyacrylamide Gel Electrophoresis (PAGE) were used to follow the course of the reactions through the consumption of the biomacroinitiator, BSA-Br (Io).Acrylates
Methyl acrylate (MA), n-butyl acrylate (n-BuA) and t-butyl acrylate (t-BuA) were studied to explore the feasibility of grafting hydrophobic acrylates from a protein macroinitiator.In previous studies we postulated that formation of a stable monomer emulsion appeared to be essential for efficient grafting of the hydrophobic styrene and its derivatives from protein macroinitiators.45,69 To establish whether this pattern applies to other hydrophobic monomers, the grafting of MA from BSA-Br (Io) was studied at increasing monomer concentrations (Table 1, entries 1–5, Fig. S2†). While no product could be detected when a 100-fold excess of MA over the macroinitiator was employed using 22 ppm of Cu(II), increasingly higher yields were observed when increasing the monomer to initiator ratio (Table 1, entries 1–5, Fig. S2†). However, in contrast to what was previously demonstrated for styrene grafting,69 none of the above-mentioned reactions were quantitative for MA grafting while at the same time, the BSA-Br (Io) initiator displayed altered chromatographic behaviour upon exposure to MA in both electrophoresis (altered migration combined with smearing of the protein band) and SEC (lower retention times, Table 1, entries 1–4, Fig. S2†).
| Entry | Mn | Mn/Io/CuII/L | Cu (ppm) | Additive | BSA-Br (Io) Consumption |
|---|---|---|---|---|---|
| a Reaction time 300 min. b Reaction time 180 min. c 0.095% w/v SDS. d 5% v/v DMSO. e 5% v/v ethyl acetate (EtOAc). f 5 mM NaBr. g Lower concentration. | |||||
| 1a | MA | 100/1/1.5/12 | 22 | — | No reaction |
| 2a | MA | 500/1/1.5/12 | 22 | — | Partial |
| 3a | MA | 1000/1/1.5/12 | 22 | — | Partial |
| 4a | MA | 2000/1/1.5/12 | 22 | — | Near quantitative |
| 5a | MA | 5000/1/1.5/12 | 22 | — | Near quantitative |
| 6a | MA | 2000/1/1.5/12 | 22 | SDSc | Near quantitative |
| 7a | MA | 2000/1/1.5/12 | 22 | DMSOd | Near quantitative |
| 8b | MA | 2000/1/1.5/12 | 22 | EtOAce | Quantitative |
| 9a | MA | 2000/1/1.5/12 | 22 | NaBrf | Quantitative |
| 10b | MA | 1000/1/3/24 | 45 | — | Quantitative |
| 11a | MA | 2000/1/3/24 | 45 | — | Quantitative |
| 12a | MA | 5000/1/3/24 | 45 | — | Quantitative |
| 13a | MA | 2000/1/1.5/12 | 45 | pH 5.5 | Quantitative |
| 14a | t-BuA | 2000/1/1.5/12 | 22 | NaBrf | Quantitative |
| 15b | t-BuA | 1000/1/3/24 | 45 | — | Quantitative |
| 16b | t-BuA | 2000/1/3/24 | 45 | — | Quantitative |
| 17a | n-BuA | 2000/1/1.5/12 | 22 | NaBrf | Quantitative |
| 18b | n-BuA | 2000/1/3/24 | 45 | — | Near quantitative |
| 19b | n-BuA | 2000/1/4.5/42 | 45 | Quantitative | |
To address the issue of macroinitiator conversion, the concentration of Cu(II) was increased from 22 to 45 ppm resulting in quantitative BSA-Br grafting at higher rates (Table 1, entries 10–12, Fig. 1). To tune the reaction yield while at the same time retaining minimal levels of catalyst concentration, i.e. 22 ppm, we reasoned that the non-quantitative grafting expressed by partial biomacroinitiator consumption may be partly stemming from the formation of less stable emulsions. Addition of an emulsifier (sodium dodecyl sulphate, SDS, Table 1, entry 6, Fig. S3†) did not improve the reaction yield. In contrast, the addition of a low volume percentage of the non-miscible ethyl acetate (Table 1, entry 8, Fig. 1A), drove the reaction to full initiator consumption while keeping catalyst concentration minimal. In support of our initial assumption, the addition of a miscible organic cosolvent (dimethyl sulfoxide, DMSO, Table 1, entry 7, Fig. S3†) did not improve macroinitiator consumption. Interestingly, addition of 5 mM NaBr (Table 1, entry 9, Fig. 1) was also found to promote MA grafting, when using 22 ppm Cu(II), presumably due to the ability of the supplementary halide source to maintain polymerization control in aqueous media via suppressing the dissociation of the catalytic deactivator (Cu(II)/Me6TREN complex).72–76 FT-IR spectroscopy elucidated the structure of the produced bioconjugates via the characteristic intense stretching vibration of the ester carbonyl group of poly(MA) at 1725 cm−1 accompanied by the stretching of the corresponding ester oxygen (C–O) at 1159 cm−1 observed on the spectrum of the biohybrid (Fig. 1D). To gain qualitative information about the molecular weights of the polymer moieties, the protein was chemically digested and the resulting polymer was characterized with 1H-NMR and SEC (Fig. 2A and B). It should be noted that the polydispersity of the truncated polymers was not evaluated as the chemical digestion could have altered polymer end-group fidelity or, because of the colloidal nature of the amphiphiles, could be incomplete. 1H-NMR spectroscopy of the chemically digested BSA-poly(MA) bioconjugates verified (poly)MA formation.
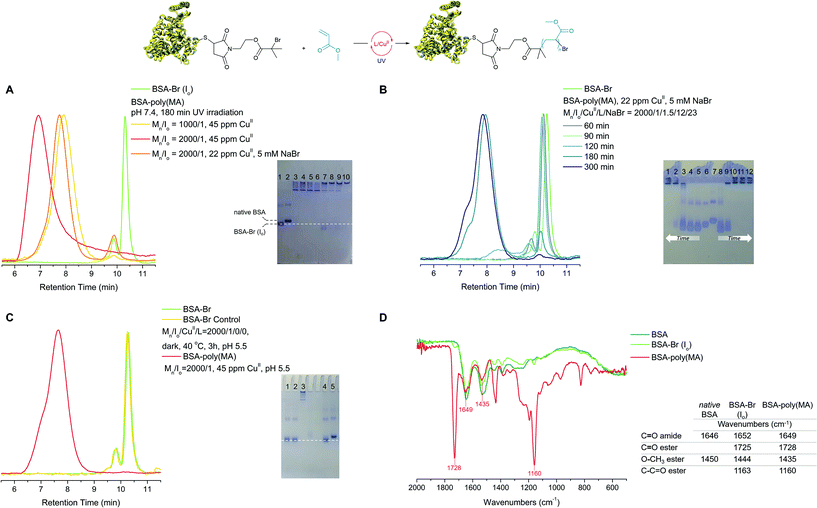 | ||
| Fig. 1 Optimization of the oxygen tolerant, photoinduced RDRP grafting of MA from BSA-Br (Io). (A) Left: SEC, green trace: BSA-Br (Io), yellow trace: Mn/Io = 1000/1, 45 ppm CuII, 180 min UV irradiation (Table 1, entry 10), red trace: Mn/Io = 2000/1, 45 ppm CuII (Table 1, entry 11), 180 min UV irradiation, orange trace: Mn/Io = 2000/1, 22 ppm CuII, 5 mM NaBr, 180 min UV irradiation (Table 1, entry 9); right: native PAGE: lane 1: BSA-Br (Io), lane 2: native BSA, lane 3: Mn/Io = 1000/1, 45 ppm CuII (Table 1, entry 10), lane 4: Mn/Io = 2000/1, 22 ppm CuII 5% v/v EtOAc (Table 1, entry 8). (B) BSA-Br (Io) consumption during the synthesis of BSA-poly(MA) via oxygen tolerant, photoinduced grafting of MA. Left: SEC, Mn/Io = 2000/1, 22 ppm CuII, 5 mM NaBr (Table 1, entry 11); right: Native PAGE, lanes 1–5: Mn/Io = 2000/1, 22 ppm CuII, 5 mM NaBr, 300, 180, 120, 90 and 60 min respectively (Table 1, entry 9), lane 6: BSA-Br (Io), lane 7: native BSA, lanes 8–12: Mn/Io = 2000/1, 45 ppm CuII, 60, 90, 120, 180 and 300 min respectively (Table 1, entry 11). (C) Blank experiment and polymerization reaction performed at pH 5.5. Left: SEC, green trace: BSA-Br (Io), yellow trace: MA/Io/CuII/L = 2000/1/3/24, 180 min at 40 °C (Table S2,† entry 7), pink trace: MA/Io/CuII/L = 2000/1/3/24, 180 min UV irradiation (Table S1,† entry 10); right: native PAGE, lane 1: MA/Io/CuII/L = 2000/1/0/0, 180 min UV irradiation (Table S2,† entry 8), lane 2: MA/Io/CuII/L = 2000/1/3/24, 180 min at 40 °C (Table S2,† entry 7), lane 3: MA/Io/CuII/L = 2000/1/3/24, 180 min UV irradiation (Table S1,† entry 10). (D) IR spectra, olive green trace: native BSA, light green trace: BSA-Br (Io), pink trace: BSA-poly(MA). | ||
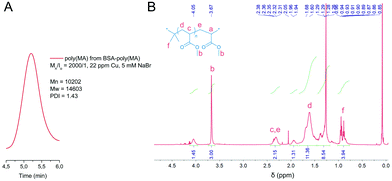 | ||
| Fig. 2 Characterization of the polymer isolated from BSA-poly(MA) by chemical digestion. (A) SEC chromatogram and, (B) 1H-NMR spectrum of poly(MA). | ||
Having optimized the conditions to drive the grafting of MA to full biomacroinitiator consumption, we next sought to address the question of protein stability upon exposure to MA. The observed chemical interaction between MA and BSA, which was expressed as smearing of the relevant protein band on PAGE, can probably be attributed to non-selective, multiple side-reactions of the monomer with the exposed lysines, histidines or the amino-terminus of BSA.77 By chromatographically following the course of the reaction (Table 1, entries 9 and 11, Fig. S4†) and performing a number of control experiments we observed the same effect upon incubation of BSA-Br or native BSA with MA under irradiation or heat (40 °C) in the presence or absence of the catalytic components (CuII or Me6TREN) under standard polymerization conditions (i.e. via headspace elimination, Table S2, entries 1–3, Fig. S5†). IR spectroscopy of the biomolecules obtained from these experiments supported the assumption of non-specific bioconjugation via the detection a new band that can be attributed to the stretching of ester carbonyls (1734 cm−1, Fig. S5†). Formation of BSA-poly(MA) was not observed in any of the blank experiments through either SEC or PAGE. Since we aimed to render this method universal, we addressed the protein charge rather than polymerization conditions to eliminate non-specific monomer binding to BSA. Taking into account that the most probable interactions between the protein and the acrylate monomer would rise from exposed amines, we studied the effect of pH and established a new set of conditions under which the side reactions between BSA and MA were eliminated and grafting of MA from BSA-Br was quantitative. Namely, when BSA-Br was incubated with MA at pH 5.5, blank experiments showed no effect on the protein and the grafting of MA under these conditions was quantitative (Fig. 1C, Table S2, entries 7 and 8, Fig. S6† and Table 1, entry 13).
To establish a universal set of conditions for the grafting of acrylates, n-BuA and t-BuA were also comprehensively studied (Table 1, entries 14–19, Fig. S7†). t-BuA was found to quantitatively yield BSA-poly(t-BuA) giant amphiphiles when using either increased catalyst loading (i.e. 45 ppm Cu(II), Mn/Io/CuII/L = 2000/1/3/24, Table 1, entries 15 and 16) or a supplementary halide source (5 mM NaBr, Table 1, entry 17). Traces of the biomacroinitiator were detected upon grafting n-BuA from the protein macroinitiator under the same conditions (Table 1, entry 18). Interestingly, the reaction was found to be quantitative when performed at lower concentrations (Table 1, entry 19). All bioconjugates were additionally characterized with FT-IR spectroscopy and the digested polymer moieties with 1H-NMR spectroscopy and SEC (Fig. S8†). It should be mentioned that by regulating the UV irradiation, excellent temporal control was observed in the photoactivated grafting of acrylates from BSA-Br (Io) (Fig. S7†).
Methacrylates
Optimization of the oxygen tolerant, photoinduced grafting of methacrylates proved to be even more demanding (ESI, Table S3,† entries 1–11). Methyl methacrylate (MMA) for example, did not afford any product under any of the optimal polymerization conditions established for styrenes and acrylates (Table 2, entry 1). This can most possibly be attributed to steric and electronic effects stemming from the interaction between the tertiary alkyl bromide initiator and the monomers.78 Addition of NaBr or ethyl acetate did not result in product formation. Increased Cu(II) concentrations (45 or 90 ppm, Table 2, entries 2 and 3) led to the formation of BSA-poly(MMA) with moderate biomacroinitiator consumption, while quantitative formation of BSA-poly(MMA) was only observed with 220 ppm Cu(II) upon prolonged irradiation (Table 2, entry 4). The IR spectrum of the produced BSA-poly(MMA) displayed the stretching vibrations for the ester group (1724 and 1143 cm−1) together with the characteristic bands of the methoxy side chain stretching at ca. 2948 cm−1 (Fig. 3C). The protein moiety of the biohybrid was chemically digested and the resulting poly(MMA) was characterized with 1H-NMR spectroscopy and SEC (Fig. S10†).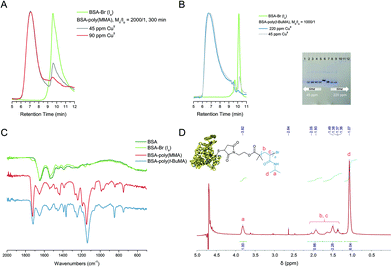 | ||
| Fig. 3 Characterization of polymer-biohybrids synthesized via the oxygen tolerant RDRP. (A) SEC of BSA-poly(MMA), green trace: BSA-Br (Io), grey trace: Mn/Io = 2000/1, 45 ppm CuII, (Table 2, entry 2), pink trace: Mn/Io = 2000/1, 90 ppm CuII, (Table 2, entry 3). (B) BSA-poly(t-BuMA). Left: SEC: green trace: BSA-Br (Io), blue trace: Mn/Io = 1000/1, 220 ppm CuII, (Table S3,† entry 14), grey trace: Mn/Io = 1000/1, 45 ppm CuII, (Table S3,† entry 13); right: study of the course of macroinitiator BSA-Br (Io) consumption with time, native PAGE, lanes 1–5: Mn/Io = 2000/1, 45 ppm CuII, 360, 235, 135, 45 and 20 min respectively (Table 2, entry 6), lane 6: native BSA, lane 7: BSA-Br (Io), lanes 8–12: Mn/Io = 2000/1, 220 ppm CuII, 20, 45, 135, 235 and 360 min respectively (Table 2, entry 7). (C) IR spectra, olive green trace: native BSA, light green: BSA-Br (Io), pink trace: BSA-poly(MMA), blue trace: BSA-poly(t-BuMA). (D) 1H-NMR spectrum of BSA-poly(NIPAM) in D2O. | ||
| Entry | Mn | Mn/Io/CuII/L | Cu (ppm) | Rxn time (min) | BSA-Br (Io) consumption |
|---|---|---|---|---|---|
| a 5 mM NaBr. | |||||
| 1a | MMA | 2000/1/1.5/12 | 22 | 300 | No reaction |
| 2 | MMA | 2000/1/3/24 | 45 | 300 | Moderate |
| 3 | MMA | 2000/1/6/48 | 90 | 300 | Moderate |
| 4 | MMA | 2000/1/15/120 | 220 | 360 | Quantitative |
| 5a | t-BuMA | 2000/1/1.5/12 | 22 | 300 | No reaction |
| 6 | t-BuMA | 2000/1/3/24 | 45 | 360 | Quantitative |
| 7 | t-BuMA | 2000/1/15/120 | 220 | 135 | Quantitative |
| 8 | NIPAM | 80/1/1.5/12 | 22 | 180 | No reaction |
| 9 | NIPAM | 80/1/3/24 | 45 | 180 | Quantitative |
| 10 | NIPAM | 400/1/3/24 | 45 | 180 | Quantitative |
The functional monomer t-butyl methacrylate (t-BuMA) was grafted in quantitative yields when using 45 ppm Cu(II) under the established photoinduced protocol conditions (Table 2, Fig. 3B, Fig. S9 and S10†), thus demonstrating the compatibility of our conditions to functional moieties. When 220 ppm Cu(II) were utilized for the grafting of t-BuMA, significantly faster macroinitiator consumption was observed. The IR spectrum of BSA-poly(t-BuMA) displayed the stretching vibrations for the ester group (1720 and 1139 cm−1) together with the characteristic bands of the tert-butyl side chains stretching at ca. 2949 cm−1 accompanied by the bending weak band at ca. 1392 cm−1 and a stronger band near 1364 cm−1 (Fig. 3C).
Acrylamides
We were further interested in evaluating the grafting of acrylamides and responsive monomers and for this reason we selected N-isopropylacrylamide (NIPAM). All reactions were performed at approximately 40 °C, i.e. under poly(NIPAM) aggregating conditions.79 No reaction was observed at standard styrene or MA polymerization conditions. Quantitative yields were obtained when 80 eq. or 400 eq. molar feed of NIPAM over the biomacroinitiator and 45 ppm Cu(II) were used. The product was characterized with PAGE and 1H-NMR (Fig. 3D, Fig. S11†) and its cloud point temperature as determined by UV was 30.4 °C.Assembly and activity
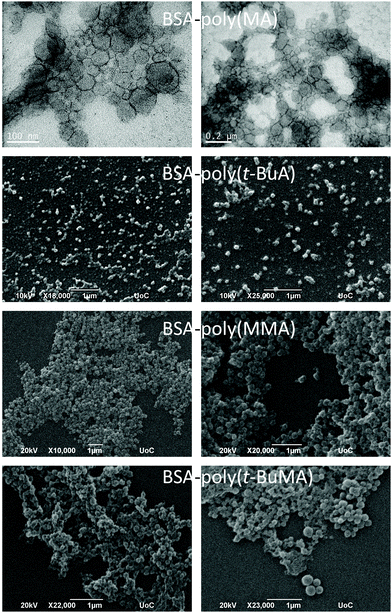
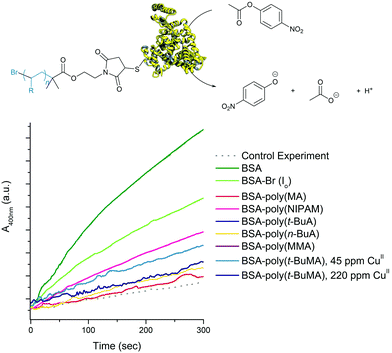
Expanding the scope of this oxygen tolerant methodology allowed for the first time to comparatively visualize the assembling architectures of amphiphilic polymer–protein biohybrids varying on the chemical identity of the polymer moiety. As shown in Fig. 4 (and Fig. S12–S15†), the biohybrids mostly formed spherical superstructures similar to those previously reported for BSA-PS giant amphiphiles.45,69 BSA-poly(MA) was challenging to image most probably due to the intrinsic soft nature of poly(MA). Transmission Electron Microscopy (TEM) of BSA-poly(MA) revealed micellar assemblies (diameter ca. 20 nm) and larger ill-defined superstructures with diameters varying from ca. 50 to 200 most probably stemming from secondary fusion of micelles. BSA-poly(t-BuA) was visualized as uniform spherical superstructures with diameters of ca. 50 nm, while BSA-poly(MMA) formed spherical superstructures with diameters of ca. 80 nm to 100 nm and BSA-poly(t-BuMA) more disperse superstructures with diameters between 80 nm and 200 nm.The effect of the polymer moiety on the biohybrid was more pronounced when studying the esterase-like activity of the produced biohybrids using p-nitrophenyl acetate (PNPA) as substrate. UV irradiation was not found to affect the esterase activity of BSA (Fig. S16†). Due to the amphiphilic nature of the biohybrids, any assay in solution can only bear qualitative results. Bearing this in mind, by performing triplicates of each catalytic assay we could safely conclude that biohybrid activity was affected by the nature of the polymer-protein amphiphiles (Fig. 5). More specifically, all biohybrids were found to retain part of the activity of the modified BSA-Br (Io) in the hydrolysis of PNPA. BSA-poly(MA) was found to be the less active bioconjugate. BSA-poly(NIPAM) on the other hand was found to retain the hydrolytic activity of its BSA head group a fact that can probably be attributed to its hydrophilic character under the activity assay conditions (20 °C i.e. below its cloud point temperature).
Conclusions
In conclusion, protein–polymer amphiphiles were synthesized via grafting of a variety of monomers from a protein macroinitiator through an oxygen tolerant, photoinduced, RDRP approach which requires ppm levels of copper(II) and UV irradiation. The universal character of this approach was demonstrated via grafting a variety of hydrophobic and/or responsive acrylates, methacrylates and acrylamides from a BSA macroinitiator. By careful optimization, reactions for all monomer families were tailored to quantitative conversions ensuring both a simple reaction and an easy purification protocol that render this polymerization accessible to all. More specifically, acrylates could be grafted using a minimal quantity of Cu(II) (22 ppm) when adding low concentrations of either ethyl acetate or NaBr. Additionally, quantitative conversions can be achieved for all monomers using 45 ppm Cu(II) loading, with the exception of MMA where 220 ppm of Cu(II) are required. We further demonstrated that the nature of the grafted monomer had a direct effect on bioconjugate activity and assembling properties.Conflicts of interest
There are no conflicts to declare.Acknowledgements
The authors acknowledge Eva Papadogiorgaki and Stefanos Papadakis at the Electron Microscopy Unit “Vassilis Galanopoulos” of the University of Crete, for assistance with TEM/SEM imaging.References
- T. A. Wright, R. C. Page and D. Konkolewicz, Polym. Chem., 2019, 10, 434–454 RSC.
- K. Velonia, A. E. Rowan and R. J. M. Nolte, J. Am. Chem. Soc., 2002, 124, 4224–4225 CrossRef PubMed.
- L. D. Blackman, S. Varlas, M. C. Arno, A. Fayter, M. I. Gibson and R. K. O'Reilly, ACS Macro Lett., 2017, 6, 1263–1267 CrossRef.
- C. K. Wong, A. J. Laos, A. H. Soeriyadi, J. Wiedenmann, P. M. G. Curmi, J. J. Gooding, C. P. Marquis, M. H. Stenzel and P. Thordarson, Angew. Chem., Int. Ed., 2015, 54, 5317–5322 CrossRef PubMed.
- S. Kulkarni, C. Schilli, B. Grin, A. H. E. Muller, A. S. Hoffman and P. S. Stayton, Biomacromolecules, 2006, 7, 2736–2741 CrossRef PubMed.
- S. J. Maassen, A. M. van der Ham and J. J. L. M. Cornelissen, ACS Macro Lett., 2016, 5, 987–994 CrossRef.
- A. C. Obermeyer and B. D. Olsen, ACS Macro Lett., 2015, 4, 101–110 CrossRef.
- Z. Liu, N. Chen, C. Dong, W. Li, W. Guo, H. Wang, S. Wang, J. Tan, Y. Tu and J. Chang, ACS Appl. Mater. Interfaces, 2015, 7, 18997–19005 CrossRef CAS.
- Z. Liu, C. Dong, X. Wang, H. Wang, W. Li, J. Tan and J. Chang, ACS Appl. Mater. Interfaces, 2014, 6, 2393–2400 CrossRef CAS.
- J. S. Kim, A. R. Sirois, A. J. Vazquez Cegla, E. Jumai'an, N. Murata, M. E. Buck and S. J. Moore, Bioconjugate Chem., 2019, 30, 1220–1231 CrossRef CAS.
- A. Abuchowski, T. van Es, N. C. Palczuk and F. F. Davis, J. Biol. Chem., 1977, 252, 3578–3581 CrossRef CAS.
- Z. Zarafshani, T. Obata and J.-F. Lutz, Biomacromolecules, 2010, 11, 2130–2135 CrossRef CAS.
- S. N. S. Alconcel, A. S. Baas and H. D. Maynard, Polym. Chem., 2011, 2, 1442–1448 RSC.
- T. H. Nguyen, S. Kim, C. G. Decker, D. Y. Wong, J. A. Loo and H. D. Maynard, Nat. Chem., 2013, 5, 221–227 CrossRef CAS.
- P. C. Nauka, J. Lee and H. D. Maynard, Polym. Chem., 2016, 7, 2352–2357 RSC.
- J. Geng, G. Mantovani, L. Tao, J. Nicolas, G. Chen, R. Wallis, D. A. Mitchell, B. R. G. Johnson, S. D. Evans and D. M. Haddleton, J. Am. Chem. Soc., 2007, 129, 15156–15163 CrossRef CAS.
- L. Tao, C. S. Kaddis, R. R. O. Loo, G. N. Grover, J. A. Loo and H. D. Maynard, Macromolecules, 2009, 42, 8028–8033 CrossRef CAS.
- M. Li, P. De, H. Li and B. S. Sumerlin, Polym. Chem., 2010, 1, 854–859 RSC.
- A. J. Keefe and S. Jiang, Nat. Chem., 2012, 4, 59–63 CrossRef CAS.
- P. Thordarson, B. Le Droumaguet and K. Velonia, Appl. Microbiol. Biotechnol., 2006, 73, 243–254 CrossRef CAS PubMed.
- M. D. Best, Biochemistry, 2009, 48, 6571–6584 CrossRef CAS.
- B. Le Droumaguet and K. Velonia, Macromol. Rapid Commun., 2008, 29, 1073–1089 CrossRef CAS.
- Y. Jiang, M. Liang, D. Svejkar, G. Hart-Smith, H. Lu, W. Scarano and M. H. Stenzel, Chem. Commun., 2014, 50, 6394–6397 RSC.
- L. Jose, A. Hwang, C. Lee, K. Shim, J. K. Song, S. S. A. An and H.-J. Paik, Polym. Chem., 2020, 11, 1580–1588 RSC.
- Q. Cao, N. He, Y. Wang and Z. Lu, Polym. Bull., 2018, 75, 2627–2639 CrossRef CAS.
- K. Parkatzidis, H. S. Wang, N. P. Truong and A. Anastasaki, Chem, 2020, 6, 1575–1588 CAS.
- N. Corrigan, K. Jung, G. Moad, C. J. Hawker, K. Matyjaszewski and C. Boyer, Prog. Polym. Sci., 2020, 111, 101311 CrossRef CAS.
- H. Li, M. Li, X. Yu, A. P. Bapat and B. S. Sumerlin, Polym. Chem., 2011, 2, 1531–1535 RSC.
- I. Cobo, M. Li, B. S. Sumerlin and S. Perrier, Nat. Mater., 2015, 14, 143–159 CrossRef CAS.
- R. M. Broyer, G. N. Grover and H. D. Maynard, Chem. Commun., 2011, 47, 2212–2226 RSC.
- S. Averick, A. Simakova, S. Park, D. Konkolewicz, A. J. D. Magenau, R. A. Mehl and K. Matyjaszewski, ACS Macro Lett., 2012, 1, 6–10 CrossRef CAS.
- B. S. Sumerlin, ACS Macro Lett., 2012, 11, 141–145 CrossRef.
- J. Nicolas, V. San Miguel, G. Mantovani and D. M. Haddleton, Chem. Commun., 2006, 4697–4699 RSC.
- M. L. Hovlid, J. L. Lau, K. Breitenkamp, C. J. Higginson, B. Laufer, M. Manchester and M. G. Finn, ACS Nano, 2014, 8, 8003–8014 CrossRef CAS.
- D. Cohen-Karni, M. Kovaliov, T. Ramelot, D. Konkolewiczc, S Graner and S Averick, Polym. Chem., 2017, 8, 3992–3998 RSC.
- Q. Zhang, M. Li, C. Zhu, G. Nurumbetov, Z. Li, P. Wilson, K. Kempe and D. M. Haddleton, J. Am. Chem. Soc., 2015, 137, 9344–9353 CrossRef CAS PubMed.
- S. Dadashi-Silab, I.-H. Lee, A. Anastasaki, F. Lorandi, B. Narupai, N. D. Dolinski, M. L. Allegrezza, M. Fantin, D. Konkolewicz, C. J. Hawker and K. Matyjaszewski, Macromolecules, 2020, 53, 5280–5288 CrossRef CAS.
- R. Whitfield, K. Parkatzidis, M. Rolland, N. P. Truong and A. Anastasaki, Angew. Chem., Int. Ed., 2019, 58, 13323–13328 CrossRef CAS.
- M. Li, H. Li, P. De and B. S. Sumerlin, Macromol. Rapid Commun., 2011, 32, 354–359 CrossRef CAS.
- C. Boyer, V. Bulmus, J. Liu, T. P. Davis, M. H. Stenzel and C. Barner-Kowollik, J. Am. Chem. Soc., 2007, 129, 7145–7154 CrossRef CAS.
- R. Whitfield, K. Parkatzidis, N. P. Truong, T. Junkers and A. Anastasaki, Chem, 2020, 6, 61340–61352 Search PubMed.
- J. Xu, K. Jung, N. A. Corrigan and C. Boyer, Chem. Sci., 2014, 5, 3568–3575 RSC.
- R. Falatach, C. McGlone, M. S. Al-Abdul-Wahid, S. Averick, R. C. Page, J. A. Berberich and D. Konkolewicz, Chem. Commun., 2015, 51, 5343–5346 RSC.
- S. A. Isarov and J. K. Pokorski, ACS Macro Lett., 2015, 4, 969–973 CrossRef CAS.
- B. Le Droumaguet and K. Velonia, Angew. Chem., Int. Ed., 2008, 47, 6263–6266 CrossRef CAS.
- E. Daskalaki, B. Le Droumaguet, D. Gerard and K. Velonia, Chem. Commun., 2012, 48, 1586–1588 RSC.
- C. Ma, X. Liu, G. Wu, P. Zhou, Y. Zhou, L. Wang and X. Huang, ACS Macro Lett., 2017, 6, 689–694 CrossRef CAS.
- X. Liu and W. Gao, ACS Appl. Mater. Interfaces, 2017, 9, 2023–2028 CrossRef CAS.
- C.-W. Chiang, X. Liu, J. Sun, J. Guo, L. Tao and W. Gao, Nano Lett., 2020, 20, 1383–1387 CrossRef CAS PubMed.
- T. Arakawa, S. J. Prestrelski, W. C. Kenney and J. F. Carpenter, Adv. Drug Delivery Rev., 2001, 46, 307–326 CrossRef CAS.
- Z. M. Tian, M. X. Wan, S. P. Wang and J. Q. Kang, Ultrason. Sonochem., 2004, 11, 399–404 CrossRef CAS PubMed.
- J. Yeow, R. Chapman, A. J. Gormley and C. Boyer, Chem. Soc. Rev., 2018, 47, 4357–4387 RSC.
- A. E. Enciso, L. Fu, A. J. Russell and K. Matyjaszewski, Angew. Chem., Int. Ed., 2018, 57, 933–936 CrossRef CAS PubMed.
- G. Ng, J. Yeow, J. Xu and C. Boyer, Polym. Chem., 2017, 8, 2841–2851 RSC.
- S. Fleischmann and V. Percec, Polym. Chem., 2010, 48, 2243–2250 CrossRef CAS.
- H. Qian and L. He, Anal. Chem., 2009, 81, 9824–9827 CrossRef CAS.
- R. Chapman, A. J. Gormley, K.-L. Herpoldt and M. M. Stevens, Macromolecules, 2014, 47, 8541–8547 CrossRef CAS.
- A. E. Enciso, L. Fu, S. Lathwal, M. Olszewski, Z. Wang, S. R. Das, A. J. Russell and K. Matyjaszewski, Angew. Chem., Int. Ed., 2018, 3, 16157–16161 CrossRef PubMed.
- K. M. Burridge, T. A. Wright, R. C. Page and D. Konkolewicz, Macromol. Rapid Commun., 2018, 39, 1800093 CrossRef.
- B. Zhu, D. Lu, J. Ge and Z. Liu, Acta Biomater., 2011, 7, 2131–2138 CrossRef CAS.
- E. Liarou, R. Whitfield, A. Anastasaki, N. G. Engelis, G. R. Jones, K. Velonia and D. M. Haddleton, Angew. Chem., Int. Ed., 2018, 57, 8998–9002 CrossRef CAS PubMed.
- E. Liarou, Y. Han, A. M. Sanchez, M. Walker and D. M. Haddleton, Chem. Sci., 2020, 11, 5257–5266 RSC.
- E. Liarou, A. Anastasaki, R. Whitfield, C. E. Iacono, G. Patias, N. G. Engelis, A. Marathianos, G. R. Jones and D. M. Haddleton, Polym. Chem., 2019, 10, 963–971 RSC.
- M. Rolland, R. Whitfield, D. Messmer, K. Parkatzidis, N. P. Truong and A. Anastasaki, ACS Macro Lett., 2019, 8, 1546–1551 CrossRef CAS.
- N. D. Dolinski, Z. A. Page, E. H. Discekici, D. Meis, I.-H. Lee, G. R. Jones, R. Whitfield, X. Pan, B. G. McCarthy, S. Shanmugam, V. Kottisch, B. P. Fors, C. Boyer, G. M. Miyake, K. Matyjaszewski, D. M. Haddleton, J. R. de Alaniz, A. Anastasaki and C. J. Hawker, J. Polym. Sci., Part A: Polym. Chem., 2019, 57, 268–273 CrossRef CAS.
- J. Xu, K. Jung, A. Atme, S. Shanmugam and C. Boyer, J. Am. Chem. Soc., 2014, 136, 5508–5519 CrossRef CAS.
- K. Matyjaszewski, S. Coca, S. G. Gaynor, M. Wei and B. E. Woodworth, Macromolecules, 1998, 31, 5967–5969 CrossRef CAS.
- K. Matyjaszewski, K. L. Beers, B. Woodworth and Z. Metzner, J. Chem. Educ., 2001, 78, 547–550 CrossRef CAS.
- A. Theodorou, E. Liarou, D. M. Haddleton, I. G. Stavrakaki, P. Skordalidis, R. Whitfield, A. Anastasaki and K. Velonia, Nat. Commun., 2020, 11, 1486 CrossRef CAS.
- J. Janatova, J. K. Fuller and M. J. Hunter, J. Biol. Chem., 1968, 243, 3612–3622 CrossRef CAS.
- X. Pan, N. Malhotra, A. Simakova, Z. Wang, D. Konkolewicz and K. J. Matyjaszewski, J. Am. Chem. Soc., 2015, 137, 15430–15433 CrossRef CAS.
- A. Anastasaki, V. Nikolaou, Q. Zhang, J. Burns, S. R. Samanta, C. Waldron, A. J. Haddleton, R. McHale, D. Fox, V. Percec, P. Wilson and D. M. Haddleton, J. Am. Chem. Soc., 2014, 136, 1141–1149 CrossRef CAS.
- A. Simula, A. Anastasaki and D. M. Haddleton, Macromol. Rapid Commun., 2016, 37, 356–361 CrossRef CAS.
- A. Simakova, S. E. Averick, D. Konkolewicz and K. Matyjaszewski, Macromolecules, 2012, 45, 6371–6379 CrossRef CAS.
- X. Pan, N. Malhotra, A. Simakova, Z. Wang, D. Konkolewicz and K. Matyjaszewski, J. Am. Chem. Soc., 2015, 137, 15430–15433 CrossRef CAS.
- G. R. Jones, R. Whitfield, A. Anastasaki and D. M. Haddleton, J. Am. Chem. Soc., 2016, 138, 7346–7352 CrossRef CAS.
- D. Liu, Y. Nashed-Samuel, P. V. Bondarenko, D. N. Brems and D. Ren, PDA J. Pharm. Sci. and Tech., 2012, 66, 12–19 CrossRef CAS.
- G. R. Jones, R. Whitfield, A. Anastasaki, N. Risangud, A. Simula, D. J. Keddie and D. M. Haddleton, Polym. Chem., 2018, 9, 2382–2388 RSC.
- H. G. Schild, Prog. Polym. Sci., 1992, 17, 163–249 CrossRef CAS.
Footnote |
| † Electronic supplementary information (ESI) available: Experimental procedures Tables S1–S3 and characterization Fig. S1–S16. See DOI: 10.1039/d0py01608j |
| This journal is © The Royal Society of Chemistry 2021 |