Open Access Article
Open Access ArticleReproducibility and sensitivity of 36 methods to quantify the SARS-CoV-2 genetic signal in raw wastewater: findings from an interlaboratory methods evaluation in the U.S.†
Brian M.
Pecson‡
 *a,
Emily
Darby‡
*a,
Charles N.
Haas
*a,
Emily
Darby‡
*a,
Charles N.
Haas
 b,
Yamrot M.
Amha
c,
Mitchel
Bartolo
d,
Richard
Danielson
e,
Yeggie
Dearborn
e,
George
Di Giovanni
f,
Christobel
Ferguson
g,
Stephanie
Fevig
b,
Yamrot M.
Amha
c,
Mitchel
Bartolo
d,
Richard
Danielson
e,
Yeggie
Dearborn
e,
George
Di Giovanni
f,
Christobel
Ferguson
g,
Stephanie
Fevig
 g,
Erica
Gaddis
h,
Donald
Gray
i,
George
Lukasik
j,
Bonnie
Mull
j,
Liana
Olivas
c,
Adam
Olivieri
k,
Yan
Qu
c and
SARS-CoV-2 Interlaboratory Consortium§
g,
Erica
Gaddis
h,
Donald
Gray
i,
George
Lukasik
j,
Bonnie
Mull
j,
Liana
Olivas
c,
Adam
Olivieri
k,
Yan
Qu
c and
SARS-CoV-2 Interlaboratory Consortium§
aTrussell Technologies Inc., Oakland, California, USA. E-mail: brianp@trusselltech.com; emilyd@trusselltech.com
bDrexel University, Philadelphia, Pennsylvania, USA
cTrussell Technologies Inc., Pasadena, California, USA
dTrussell Technologies Inc., Solana Beach, California, USA
eCel Analytical Inc., San Francisco, California, USA
fMetropolitan Water District of Southern California, Los Angeles, California, USA
gThe Water Research Foundation, Alexandria, Virginia, USA
hUtah Department of Environmental Quality, Salt Lake City, Utah, USA
iEast Bay Municipal Utility District, Oakland, California, USA
jBCS Laboratories Inc., Gainesville, Florida, USA
kEOA Inc., Oakland, California, USA
First published on 20th January 2021
Abstract
In response to COVID-19, the international water community rapidly developed methods to quantify the SARS-CoV-2 genetic signal in untreated wastewater. Wastewater surveillance using such methods has the potential to complement clinical testing in assessing community health. This interlaboratory assessment evaluated the reproducibility and sensitivity of 36 standard operating procedures (SOPs), divided into eight method groups based on sample concentration approach and whether solids were removed. Two raw wastewater samples were collected in August 2020, amended with a matrix spike (betacoronavirus OC43), and distributed to 32 laboratories across the U.S. Replicate samples analyzed in accordance with the project's quality assurance plan showed high reproducibility across the 36 SOPs: 80% of the recovery-corrected results fell within a band of ±1.15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log10 genome copies per L with higher reproducibility observed within a single SOP (standard deviation of 0.13
log10 genome copies per L with higher reproducibility observed within a single SOP (standard deviation of 0.13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log10). The inclusion of a solids removal step and the selection of a concentration method did not show a clear, systematic impact on the recovery-corrected results. Other methodological variations (e.g., pasteurization, primer set selection, and use of RT-qPCR or RT-dPCR platforms) generally resulted in small differences compared to other sources of variability. These findings suggest that a variety of methods are capable of producing reproducible results, though the same SOP or laboratory should be selected to track SARS-CoV-2 trends at a given facility. The methods showed a 7
log10). The inclusion of a solids removal step and the selection of a concentration method did not show a clear, systematic impact on the recovery-corrected results. Other methodological variations (e.g., pasteurization, primer set selection, and use of RT-qPCR or RT-dPCR platforms) generally resulted in small differences compared to other sources of variability. These findings suggest that a variety of methods are capable of producing reproducible results, though the same SOP or laboratory should be selected to track SARS-CoV-2 trends at a given facility. The methods showed a 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log10 range of recovery efficiency and limit of detection highlighting the importance of recovery correction and the need to consider method sensitivity when selecting methods for wastewater surveillance.
log10 range of recovery efficiency and limit of detection highlighting the importance of recovery correction and the need to consider method sensitivity when selecting methods for wastewater surveillance.
Water impactWastewater surveillance can help assess community health during the COVID-19 pandemic, providing a critical diagnostic tool for resource-constrained settings where large-scale clinical testing is infeasible. This study showed multiple methods provide the reproducibility, sensitivity, and precision to quantify SARS-CoV-2 in raw wastewater. These methods have the potential to identify trends in community health and the effectiveness of public health interventions. |
1 Introduction
The international water community responded rapidly to the onset of the COVID-19 pandemic by developing methods to measure SARS-CoV-2 genome concentrations in wastewater.1–3 This effort was prompted by the identification of fecal shedding of SARS-CoV-2 in infected individuals.4–6 As a result, wastewater surveillance has the potential to complement clinical testing by providing a broad observational assessment of the community's health.3,7 Such knowledge could help guide public health agencies to identify and respond to outbreaks. Unlike clinical data—which may be biased toward the evaluation of symptomatic individuals—wastewater contains regular inputs from the entire population representing all stages of infection from symptomatic to pre-symptomatic to asymptomatic individuals. Furthermore, recent studies have shown that wastewater surveillance can provide an early warning of community infection, with wastewater concentrations spiking several days before identification through clinical testing.7–11In April, 2020, the Water Research Foundation (WRF) hosted an international summit to evaluate the use of wastewater surveillance as an indicator of the distribution of COVID-19 in communities.12 The participants identified two priority applications for the use of wastewater surveillance data: 1) tracking trends in occurrence and 2) assessing the degree of community prevalence. One of the prerequisites for these applications, however, is the identification of reliable, reproducible, and sensitive methods.10,12,13 To help address this issue, this study performed an interlaboratory evaluation of 36 different methods used to assess the genetic signal of SARS-CoV-2 in untreated wastewater. The nationwide study included 32 U.S. laboratories from 19 different states each processing split samples of two different raw wastewaters emanating from populations known to have high levels of infection. The project sought to identify if and how the SARS-CoV-2 findings were impacted by multiple methodological differences such as sample concentration method, pasteurization pre-treatment, primer/probe selection, and solids removal steps. The effort did not intend to standardize a single method, but evaluate whether the existing methods provide sufficient reliability and reproducibility to track trends in occurrence and assess the prevalence of community infection.
2 Methods
2.1 Participating labs
The 32 participating laboratories included 17 academic labs, 6 commercial labs, 4 non-municipal government labs, 3 municipalities, and 2 manufacturers of molecular tests (Table 1). Prior to the interlaboratory study, many of the labs were engaged in on-going monitoring efforts across the country. The participating labs agreed to follow the project's quality assurance project plan (QAPP) described below and process ten independent samples over a one-week period. The project QAPP is described in detail in this section in addition to an overview of the 36 individual standard operating procedures (SOPs) evaluated in the study.| Lab name | Lab type | State |
|---|---|---|
| Biological Consulting Services (BCS) Laboratories | Commercial | FL |
| Cel Analytical | Commercial | CA |
| City of Scottsdale | Government | AZ |
| City University of New York | Academic | NY |
| Columbia University | Academic | NY |
| Hampton Roads Sanitation District | Utility | VA |
| IDEXX Laboratories, Inc. | Manufacturer | ME |
| Los Angeles County Sanitation Districts | Utility | CA |
| Michigan State University | Academic | MI |
| Mycometrics | Commercial | NJ |
| New York City Department of Environmental Protection | Government | NY |
| Ohio State University | Academic | OH |
| Oregon State University | Academic | OR |
| Promega Corporation | Manufacturer | WI |
| Saginaw Valley State University | Academic | MI |
| SiREM | Commercial | TN |
| Source Molecular Corporation | Commercial | FL |
| Southern Nevada Water Authority | Utility | NV |
| Tulane University | Academic | LA |
| United States Environmental Protection Agency | Government | OH |
| University of California – Berkeley | Academic | CA |
| University of California – Irvine | Academic | CA |
| University of Colorado – Boulder | Academic | CO |
| University of Maryland | Academic | MD |
| University of Missouri | Academic | MO |
| University of Nebraska | Academic | NE |
| University of Nebraska – Medical Center | Academic | NE |
| University of Utah | Academic | UT |
| University of Wisconsin | Academic | WI |
| Utah State University | Academic | UT |
| Weck Labs | Commercial | CA |
| Wisconsin State Lab of Hygiene | Government | WI |
2.2 Microorganisms
Human betacoronavirus OC43 was used as a matrix spike to assess the recovery efficiency of each method. To prepare the OC43 matrix spike, a concentrated stock of OC43 (betacoronavirus 1 (ATCC® VR-1558™)) was grown in cell culture using HCT-8 cells (ATCC® CCL-244™), according to ATCC instructions. The concentration of OC43 genome copies (GC) in the stock was quantified by reverse transcription quantitative polymerase chain reaction (RT-qPCR) against a standard curve of quantitative genomic RNA from betacoronavirus 1 (ATCC® VR-1558DQ™) to determine the GC per ml of the stock. Eight labs concurrently evaluated additional matrix spike organisms, including bovine coronavirus (BCoV), heat-inactivated SARS-CoV-2, bacteriophage MS2, bacteriophage Phi6, in vitro transcribed RNA, and an engineered RNA virus.2.3 Sample collection, shipping, and handling
As detailed in the QAPP, raw wastewater samples were collected and distributed from two wastewater treatment plants (WWTPs) in Los Angeles County on two sampling days: (1) the Hyperion Water Reclamation Plant (operated by the City of Los Angeles Sanitation and Environment) on August 17, 2020 (Plant 1) and (2) the Joint Water Pollution Control Plant (operated by the Los Angeles County Sanitation Districts) on August 19, 2020 (Plant 2). These plants are two of the largest wastewater treatment plants on the west coast of the United States (Table 2). The sample collection location at both WWTPs was after grit removal prior to primary clarification. At both WWTPs, a single 40 gallon grab sample was collected at approximately 10:00 AM. The bulk sample was distributed into 1 gal containers (one for each lab) while mixing the bulk sample continuously to promote homogeneity. To confirm the homogeneity of the samples, 1 L aliquots were collected after the 1st, 17th, and 34th samples and the total suspended solids, temperature, and pH were measured as surrogates for sample homogeneity (Table 2). The 1 gallon samples were chilled on dry ice to a temperature of approximately 4°C and then blind-spiked with betacoronavirus OC43 to a final concentration of 2.8 × 108 GC/L. The samples were shipped to each laboratory with enough ice packs to maintain a temperature below 10°C. The participating labs were instructed to begin processing the sample between 8:00 AM and 12:00 PM Pacific time on the day after sample collection (i.e., 24 ± 2 h after sample collection).2.4 Sample analysis
The participating labs each processed a total of 10 sample replicates. Most of the labs achieved these 10 sample replicates by processing five sample replicates from Plant 1 and five from Plant 2. Eight laboratories evaluated the impact of heat pasteurization (60°C for 60 min) and so they achieved their 10 sample replicates by processing five sample replicates without heat pasteurization and five with heat pasteurization, all from Plant 1.Each of the participating labs followed their own SOP for sample pre-treatment, concentration, extraction, and molecular analysis. Four of the participating labs tested two different SOPs leading to a total of 36 SOPs evaluated across the 32 labs. The detailed SOPs can be found in the ESI.† The SOPs were organized into eight method groups based on the concentration step prior to RNA extraction and whether solids were removed prior to concentration. The key method steps and categorization of the 36 SOPs are shown in Table 3. Briefly, the starting sample volume ranged from 0.25 mL to 400 mL across the SOPs. The first step in sample processing was pre-treatment (e.g., heat pasteurization, solids removal, and/or chemical addition). Most labs did not pasteurize their samples before processing. SOPs involving heat pasteurization for all of the samples are marked with “H” and those involving heat pasteurization for half of the samples are marked with “(H)”. Approximately half of the SOPs involved the removal of solids (using either centrifugation, filtration, or both) prior to concentration. Method groups with SOPs involving solids removal are marked with an “S”. Many of the SOPs involved addition of chemicals to adjust the pH and/or the ionic composition of the matrix prior to concentration. After pre-treatment, the next major step in sample processing was concentration. The four main categories of concentration steps among these SOPs were 1) no concentration (i.e., direct extraction), 2) ultrafiltration, 3) filtration using an electronegative membrane (i.e., HA filter), and 4) PEG precipitation. The next step in sample processing was extraction. A variety of different extraction kits and in-house methods were used by the participating laboratories to extract the RNA from the sample. After extraction, the molecular analysis was conducted using either one-step or two-step RT-qPCR or reverse transcription digital PCR (RT-dPCR). All labs analyzed the native SARS-CoV-2 molecular signal using the N1 and N2 primer/probes sets and the OC43 matrix spike (Table 4). The concentration factors (CF) resulting from the different method steps of the SOPs, calculated using the equation below, ranged from 5 to 2100.
 | (1) |
| Method group | SOP | Sample volume (mL) | Pre-treatment | Concentration step | Extraction | Molecular analysise | Concentration factor | ||
|---|---|---|---|---|---|---|---|---|---|
| Pasteurization | Solids removal | Chemical addition | |||||||
| a SOP 1.3 centrifuged sample and analyzed solids. b SOP 2.3 separated solids and analyzed both solid and liquid fractions. c SOP 2S.5 used a concentrating pipette tip in the concentration step (similar principle to ultrafilter). d SOP 3S.1 filters the sample through an electropositive filter to remove solids and then elutes the viruses adsorbed to the filter with beef extract. The eluant is further concentrated with organic flocculation and ultrafiltration before extraction. e “Q” indicates reverse transcription quantitative PCR and “D” indicates reverse transcription digital PCR. | |||||||||
| 1 | 1.1 | 0.25 | No | Nonea | None or RNA shield | None | Zymo Quick-RNA Fecal/Soil Microbe Microprep kit | Q | 17 |
| 1.2(H) | 40 | Half the samples | PureYield Plasmid Midiprep system | Q | 500 | ||||
| 1.3 | 45 | No | Qiagen RNeasy PowerSoil Total RNA kit | Q | 450 | ||||
| 1S | 1S.1(H) | 40 | Half the samples | Yes (e.g., removal by centrifugation, filtration, or both) | None or salt addition (e.g., NaCl) prior to solids removal | PureYield Plasmid Midiprep system | Q | 500 | |
| 1S.2H | 40 | All samples | Zymo III-P silica column | Q | 200 | ||||
| 1S.3(H) | 2 | Half the samples | Qiagen QIAamp Viral RNA mini kit | D | 5 | ||||
| 2 | 2.1 | 30 | No | Noneb | Beef extract or phosphate buffered saline (PBS) | Zymo Quick-DNA/RNA Viral kit | Q | 60 | |
| 2.2 | 30 | No | Zymo Quick-DNA/RNA Viral kit | Q | 60 | ||||
| 2.3 | 225 | No | Qiagen AllPrep PowerViral kit and Qiagen RNeasy PowerWater kit | D | 1800 | ||||
| 2S | 2S.1 | 50 | No | Yes (e.g., removal by centrifugation, filtration, or both) | None | Ultrafiltrationc | Qiagen RNeasy mini kit | Q | 40–200 |
| 2S.2 | 105 | No | IDEXX Water DNA/RNA Magnetic Bead kit | Q | 380–980 | ||||
| 2S.3 | 150 | No | Invitrogen PureLink Viral RNA/DNA mini kit | Q | 220–630 | ||||
| 2S.4(H) | 50 | Half the samples | Agilent Absolutely RNA Miniprep kit | Q | 500 | ||||
| 2S.5 | 25 | No | TRIzol | Q | 63–280 | ||||
| 2S.6 | 30 | No | Zymo Quick-RNA Miniprep kit | Q | 16–18 | ||||
| 3 | 3.1 | 50 | No | None | Acid (HCl) to lower pH and (optionally) addition of salt (e.g., MgCl2) | HA filtrationd | NUCLISENS easyMAG | D | 250 |
| 3.2 | 100 | No | Qiagen QIAamp Viral RNA kit | Q | 880–2100 | ||||
| 3.3 | 50 | No | Qiagen AllPrep PowerViral DNA/RNA kit | Q | 280–470 | ||||
| 3.4 | 25 | No | Qiagen RNeasy PowerMicrobiome kit using PowerBead tubes | D | 420 | ||||
| 3.5 | 40 | No | Qiagen RNeasy PowerMicrobiome kit using BashingBead tubes | Q | 40–200 | ||||
| 3.6 | 30 | No | Applied Biosystems MagMAX Viral/Pathogen Nucleic Acid Isolation kit | D | 200–230 | ||||
| 3S | 3S.1 | 200 | No | Yes (e.g., removal by centrifugation, filtration, or both) | Acid (HCl) to lower pH after solids removal | Qiagen AllPrep PowerViral DNA/RNA kit | Q | 2000 | |
| 3S.2H | 100 | All samples | Phenol extraction | Q | 380–1300 | ||||
| 3S.3H | 50 | All samples | Phenol extraction | Q | 160–510 | ||||
| 4 | 4.1 | 100 | No | None | Salt (NaCl) and PEG | Qiagen QIAamp Viral RNA kit | D | 60–96 | |
| 4.2 | 100 | No | Qiagen QIAamp Viral RNA kit | D | 53 | ||||
| 4.3 | 100 | No | Qiagen QIAamp Viral RNA kit | D | 55–83 | ||||
| 4.4 | 282 | No | Qiagen QIAamp Viral RNA kit | Q | 220 | ||||
| 4S | 4S.1(H) | 40 | Half the samples | Yes (e.g., removal by centrifugation, filtration, or both) | Salt (NaCl) and PEG after solids removal | PEG precipitation | TRIzol | Q | 850–1300 |
| 4S.2(H) | 105 | Half the samples | IDEXX Water DNA/RNA Magnetic Bead kit | Q | 530 | ||||
| 4S.3 | 45 | No | Qiagen RNeasy PowerMicrobiome kit using PowerBead tubes | D | 130 | ||||
| 4S.4 | 36 | No | Qiagen QIAamp Viral RNA kit | Q | 590 | ||||
| 4S.5H | 40 | All samples | Qiagen AllPrep PowerViral DNA/RNA kit | Q | 670 | ||||
| 4S.6(H) | 200 | Half the samples | NucleoMag Pathogen RNA Isolation kit | Q | 170 | ||||
| 4S.7 | 40 | No | Invitrogen PureLink Viral RNA/DNA mini kit | Q | 34–170 | ||||
| 4S.8(H) | 400 | Half the samples | Qiagen QIAamp Viral RNA kit | D | 470 | ||||
| Target | Primer/probe sequences | Ref. |
|---|---|---|
| SARS-CoV-2 N1 | F: 5′-GAC CCC AAA ATC AGC GAA AT-3′ | 2019-nCoV CDC EUA kit, IDT Catalog No. 10006606 |
| R: 5′-TCT GGT TAC TGC CAG TTG AAT CTG-3′ | ||
| P: 5′-FAM-ACC CCG CAT TAC GTT TGG TGG ACC-BHQ1-3′ | ||
| Pdouble: FAM-ACC CCG CAT/ZEN/TAC GTT TGG TGG ACC-3IABkFQ | ||
| SARS-CoV-2 N2 | F: 5′-TTA CAA ACA TTG GCC GCA AA-3′ | 2019-nCoV CDC EUA kit, IDT Catalog No. 10006606 |
| R: 5′-GCG CGA CAT TCC GAA GAA-3′ | ||
| P: 5′-FAM-ACA ATT TGC CCC CAG CGC TTC AG-BHQ1-3′ | ||
| Pdouble: FAM-ACA ATT TGC/ZEN/CCC CAG CGC TTC AG-3IABkF | ||
| OC43 | F: 5′-CGATGAGGCTATTCCGACTAGGT-3′ | Dare, R.K. et al. J Infect. Diseases, 2007, 196: 1321–8 |
| R: 5′-CCTTCCTGAGCCTTCAATATAGTAACC-3′ | ||
| P: 5′-6FAM-TCCGCCTGGCACGGTACTCCCT-BHQ-3′ |
V sample before processing = original sample volume before processing (mL).
V after concentration = sample volume after concentration (mL).
V concentrate used for RNA extraction = volume of concentrate used for RNA extraction (mL).
V after RNA extraction = volume after RNA extraction (mL).
DF = dilution factor for RNA extract after extraction (i.e., [RNA extract volume + diluent volume]/RNA extract volume).
Some of the SOPs using direct extraction (i.e., without a concentration step prior to RNA extraction) had a comparable or greater CF than the SOPs with a concentration step. In these cases, high CFs were achieved by using a large sample volume for RNA extraction and concentrating the sample down to a small volume of RNA extract.
While each laboratory followed their own SOP, each lab was required to adhere to the project's QAPP that described the quality control requirements.14 The QAPP was constructed to ensure uniformity in sample collection, shipping and handling, quality control for the analytical methods, data management, and validation. Key elements of the QAPP included:
2.5 Data analysis
The following quality control exclusion criteria were used to determine which data were included in the method analysis.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–20
000–20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000); results below the limit of detection were considered NDs. Two thirds of the SOPs had at least one molecular replicate that was marked as non-detect due to these criteria.
000); results below the limit of detection were considered NDs. Two thirds of the SOPs had at least one molecular replicate that was marked as non-detect due to these criteria.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log impact on SARS-CoV-2 enumeration.15 Because the sample contained the OC43 matrix spike, it was decided that the recovery-corrected results would minimize the impact of this step and make it acceptable to include the findings in the analysis.
log impact on SARS-CoV-2 enumeration.15 Because the sample contained the OC43 matrix spike, it was decided that the recovery-corrected results would minimize the impact of this step and make it acceptable to include the findings in the analysis.
A summary of the results that were excluded from the analysis are presented in Table 5.
| SOPs excluded from method analysis | Quality control rationale |
|---|---|
| Two thirds of SOPs had at least one molecular replicate that were marked as non-detect due to the results falling outside of the range covered by the standard curve. NDs were not included in the analysis of SARS-CoV-2 results, but the SOPs were still included in method sensitivity analysis. | |
| 1S.1 (H) | Processed more than 24 h outside specified window |
| 3.2 (excluded N1 results only) | Positives in N1 NTC |
| 2S.1 (still included in method sensitivity analysis) | Low recovery (<0.01%) |
| 3S.1 (still included in method sensitivity analysis) | Low recovery (<0.01%) |
After applying the exclusion criteria, the results of the sample replicates from each WWTP were analyzed separately. In the eight cases where an SOP was tested with and without pasteurization, the results were analyzed independently. When analyzing data by method group, only the five replicates without pasteurization were included in the statistical analysis of the method groups so as to not give extra weight to those SOPs.
2.6 Statistical analysis
The statistical analysis was performed in R using the log10-transform of the SARS-CoV-2 concentration, recovery efficiency, and limit of detection.16 One-way ANOVA was used to compare the results of the eight method groups. A Tukey post hoc test was used to perform multiple pair-wise comparisons. Comparisons with a p-value less than 0.05 were considered significant. Two-way ANOVA, with an interaction term, was used to evaluate the impact of different method steps, specifically, heat pasteurization, solids removal, primer/probe target, PCR platform, and matrix spike selection. Two-way ANOVA allows for the evaluation of two independent variables. The difference between the two levels of the second independent variable are calculated at each level of the first independent variable and averaged to determine if the difference is significant. For each of the method steps evaluated, the first independent variable was either the SOP or the concentration step and the second independent variable was the method step of interest: heat pasteurization, solids removal, primer/probe target, PCR platform, and matrix spike surrogate. The dependent variable was either the SARS-CoV-2 concentration or the matrix spike recovery. When the design was unbalanced, a type III sum of squares approach was used for two-way ANOVA.3 Results
Over 2000 data points were produced from the interlaboratory analyses. This section addresses the reproducibility and sensitivity of the methods, both across all SOPs as well as within each of the eight major method groups. In addition, the impact of several other method steps—namely, pasteurization, primer/probe set, PCR platform, and matrix spike surrogate selection—was evaluated.3.1 Reproducibility
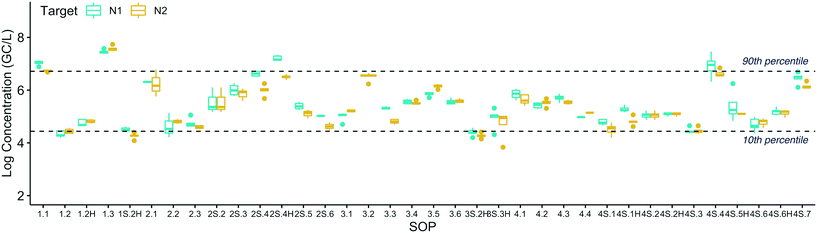
The reproducibility of the methods was evaluated at three different levels: 1) across all method groups, 2) within each method group, and 3) within each SOP.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log and 6.7
log and 6.7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log genome copies per liter (GC/L), respectively, for the combined N1 and N2 datasets (shown as dashed lines in Fig. 1). In other words, 80% of the values from 36 different SOPs fell within a ±1.15
log genome copies per liter (GC/L), respectively, for the combined N1 and N2 datasets (shown as dashed lines in Fig. 1). In other words, 80% of the values from 36 different SOPs fell within a ±1.15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log band (2.3
log band (2.3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log range). While a similar degree of reproducibility was observed at Plant 2, fewer SOPs were tested since those evaluating the impact of pasteurization only processed the Plant 1 sample and a greater percentage of the samples that were processed resulted in NDs (data not shown).
log range). While a similar degree of reproducibility was observed at Plant 2, fewer SOPs were tested since those evaluating the impact of pasteurization only processed the Plant 1 sample and a greater percentage of the samples that were processed resulted in NDs (data not shown).
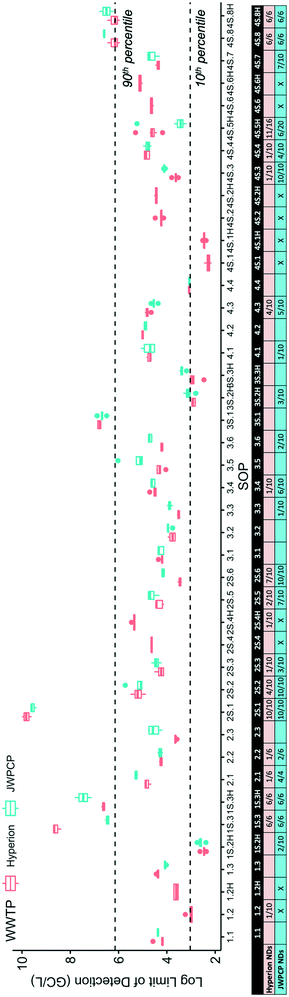
In contrast, the recovery efficiency of the SOPs spanned seven orders of magnitude (Fig. 2). Correcting for this source of methodological variability allowed the recovery-corrected concentrations to converge within a tighter minimum–maximum range than the uncorrected values (uncorrected data shown in Fig. S1†), highlighting the importance of correcting for recovery in obtaining reproducible results across SOPs.
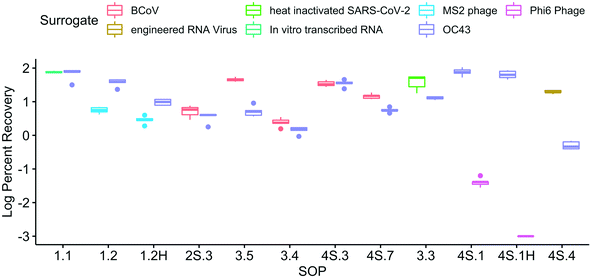
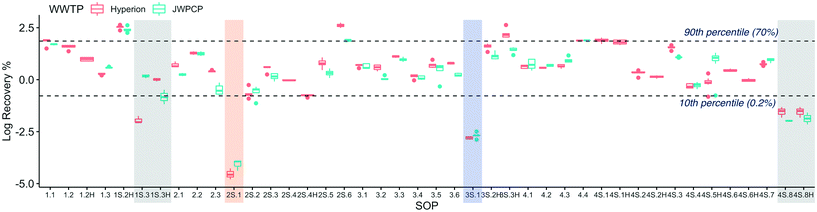 | ||
| Fig. 2 Log-transformed OC43 recovery efficiency at Plant 1 (Hyperion) and Plant 2 (JWPCP), measured by each SOP. The SARS-CoV-2 results from the SOPs highlighted are not represented in Fig. 1 due to the fact that the results were all non-detect (gray), the recovery was below the quality control cut-off of 0.01% (blue), or both (orange). | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log or less. Method group 1 had the lowest reproducibility with a 10th-to-90th percentile band of 3.2
log or less. Method group 1 had the lowest reproducibility with a 10th-to-90th percentile band of 3.2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) logs. The factors leading to higher reproducibility within some method groups was not clear from the analysis. Potential factors include features inherent in the methods that lend themselves towards higher reproducibility or greater similarity of the SOPs within that method group. For example, three laboratories in method group 4 used a very similar SOP and had been in communication with each other prior to this study. The high reproducibility observed within group 4 suggests that aligning the details of an SOP between participants and greater interlaboratory communication may help to further improve the reproducibility of methods.
logs. The factors leading to higher reproducibility within some method groups was not clear from the analysis. Potential factors include features inherent in the methods that lend themselves towards higher reproducibility or greater similarity of the SOPs within that method group. For example, three laboratories in method group 4 used a very similar SOP and had been in communication with each other prior to this study. The high reproducibility observed within group 4 suggests that aligning the details of an SOP between participants and greater interlaboratory communication may help to further improve the reproducibility of methods.
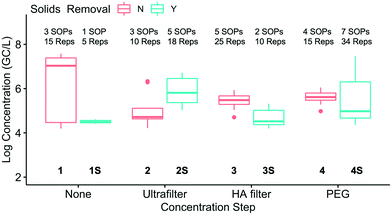
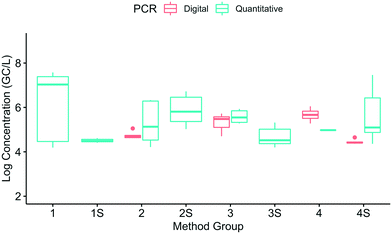
A box plot of the corrected SARS-CoV-2 N1 concentrations in eight method groups is shown in Fig. 3. Given the variability of the pooled samples within the method groups, the recovery-corrected results from the different method groups were not systematically impacted by solids removal or concentration. Of the 28 pairwise combinations, only six had significant differences: 1S and 1 (p = 0.00047), 2 and 1 (p = 0.0028), 3 and 1 (p = 0.031), 4S and 1 (p = 0.0074), 2S and 1S (p = 0.013), and 3S and 2S (p = 0.0027). In other words, multiple methods led to similar results if the results were corrected for recovery. Similar trends were observed at Plant 2 (data not shown). Because only one or two SOPs were present in method groups 1S and 3S, the variability within those groups was not as well characterized as the other groups. Further studies with additional SOPs per group could be used to confirm the impact of solids removal and concentration steps.
| Target | Uncorrected | Recovery-corrected |
|---|---|---|
| N1 | 0.15 [0.04–0.38] | 0.13 [0.032–0.60] |
| N2 | 0.14 [0.01–0.53] | 0.13 [0.033–0.51] |
3.2 Sensitivity
The sensitivity of each SOP was evaluated by quantifying the theoretical limit of detection (LOD), which was, in turn, a function of three variables: the recovery efficiency, the concentration factor (CF), and the instrument detection limit of the PCR platform. The recovery efficiency for each SOP was calculated as the percentage of the OC43 matrix spike that was detected by the method (Fig. 2). The concentration factor quantified the degree to which the SARS-CoV-2 concentrations increased as the raw wastewater was processed to produce the final RNA extract. Concentrations factors were SOP-dependent (Table 3). The instrument detection limit is the lowest concentration at which the PCR instrument can reliably distinguish a target signal from the background. Rigorous methods for quantifying instrument detection limits have been described previously,17 but were not evaluated during this study. In lieu of this, a theoretical instrument detection limit of one GC per 5 μl PCR assay was assumed.These three factors were used to calculate the theoretical LOD for each SOP:
 | (2) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log GC/L. To understand the sensitivity of the methods to detect lower concentrations than those present in the August 2020 wastewater samples, the log-difference between the measured SARS-CoV-2 concentrations and the theoretical LOD was determined for each SOP (shown in Table S1†). The median difference across all methods was 0.8
log GC/L. To understand the sensitivity of the methods to detect lower concentrations than those present in the August 2020 wastewater samples, the log-difference between the measured SARS-CoV-2 concentrations and the theoretical LOD was determined for each SOP (shown in Table S1†). The median difference across all methods was 0.8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) logs, though some methods could detect concentrations 2
logs, though some methods could detect concentrations 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log lower or more.
log lower or more.
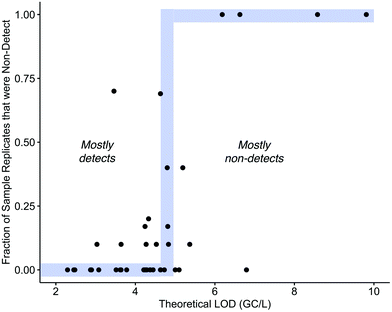
The variabilities in sensitivities can also be evaluated based on the frequency of sample replicates with NDs at each WWTP. As anticipated, SOPs with higher LODs (lower sensitivity) tended to have higher rates of NDs, and SOPs with lower LODs (higher sensitivity) tended to have fewer NDs (Fig. 5). Recall, the theoretical LOD is based on the observed OC43 recovery—the actual SARS-CoV-2 recovery was not directly measured. Therefore, the fact that a strong relationship is observed between the LOD and the frequency of NDs suggests that OC43 is generally providing an accurate reflection of the relative SARS-CoV-2 recovery across different methods. It should be noted, however, that other factors affecting OC43 recovery at each lab (e.g., sample-to-sample differences, shipping effects, sample handling) may also contribute to the differences in the calculated LODs.
To assess whether sensitivity was linked to methodological differences, the LODs for both WWTPs were compared by method group (Fig. 6). The LODs between method groups were generally indistinguishable, partially due to the high variability of LODs within the method groups with solids removal. In each of these solids removal groups, the large LOD range was driven by a single SOP in the group with a high LOD, specifically, 1S.3(H), 2S.1, 3S.1, and 4S.8(H). These SOPs all had NDs and/or recovery below 0.01%. Only three of the 28 pairwise combinations were significantly different and all were associated with method group 2S: 2S and 1 (p = 0.0011), 2S and 3 (p = 0.0062), and 2S and 4S (p = 0.011). The SOPs with highest sensitivity were not all associated with the same method group, meaning that multiple methods may be capable of achieving high sensitivities.
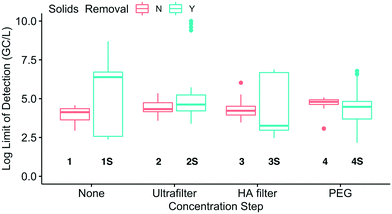 | ||
| Fig. 6 Comparison of the log-transformed theoretical limits of detection (combined for Plant 1 and Plant 2) for each of the eight method groups (grouped by concentration step and solids removal). | ||
3.3 Impact of other method steps
In addition to the main method steps differentiating the SOPs in this study (i.e., concentration step and solids removal), several other method steps were evaluated, namely heat pasteurization, primer set, PCR platform, and surrogate used as the matrix spike.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log for N1 and 0.31
log for N1 and 0.31![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log for N2) in the corrected SARS-CoV-2 concentrations after pasteurization (Fig. 7). Because there was no statistically significant difference in the uncorrected results with and without pasteurization (Fig. S2†), the slight increase in the corrected pasteurized values was due to the lower recovery efficiencies in the pasteurized samples compared to the unpasteurized samples (Fig. S2†).
log for N2) in the corrected SARS-CoV-2 concentrations after pasteurization (Fig. 7). Because there was no statistically significant difference in the uncorrected results with and without pasteurization (Fig. S2†), the slight increase in the corrected pasteurized values was due to the lower recovery efficiencies in the pasteurized samples compared to the unpasteurized samples (Fig. S2†).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log greater than N2 at Plant 1 and 0.12
log greater than N2 at Plant 1 and 0.12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log greater at Plant 2.
log greater at Plant 2.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log and 0.47
log and 0.47![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log higher than OC43, respectively). One systematic difference was that OC43 was added upon sample collection before shipment to the labs whereas the second matrix spike was added upon receipt by the individual labs. A lower recovery for OC43 could be the result of decay that occurred in the sample during shipment that was not accounted for by the second surrogate. In comparison to the other betacoronaviruses, other surrogates had larger differences in recovery than OC43. For example, enveloped bacteriophage Phi6 had a recovery that was 3.9
log higher than OC43, respectively). One systematic difference was that OC43 was added upon sample collection before shipment to the labs whereas the second matrix spike was added upon receipt by the individual labs. A lower recovery for OC43 could be the result of decay that occurred in the sample during shipment that was not accounted for by the second surrogate. In comparison to the other betacoronaviruses, other surrogates had larger differences in recovery than OC43. For example, enveloped bacteriophage Phi6 had a recovery that was 3.9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log lower than the OC43 recovery. It is important to note that differences in surrogate recovery may be SOP-dependent, meaning that a surrogate may behave similarly to another in one SOP but differently in another. These findings suggest that multiple surrogates may be acceptable, but highlight the differences between some of the commonly used selections.
log lower than the OC43 recovery. It is important to note that differences in surrogate recovery may be SOP-dependent, meaning that a surrogate may behave similarly to another in one SOP but differently in another. These findings suggest that multiple surrogates may be acceptable, but highlight the differences between some of the commonly used selections.
4 Discussion
This study demonstrated that a diverse set of 36 methods was able to quantify the SARS-CoV-2 genetic signal in raw wastewater with a high degree of reproducibility. 80% of the data from the eight different method groups fell within a band of approximately ±1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log GC/L when corrected for recovery. This finding bodes well for the nationwide interest in tracking SARS-CoV-2 in raw wastewater since a single standardized method may not be critical for obtaining comparable results between laboratories. Access to multiple, reliable methods may also increase the number of labs capable of participating in monitoring efforts and provide resilience against supply chain issues that have beset these efforts during the pandemic.
log GC/L when corrected for recovery. This finding bodes well for the nationwide interest in tracking SARS-CoV-2 in raw wastewater since a single standardized method may not be critical for obtaining comparable results between laboratories. Access to multiple, reliable methods may also increase the number of labs capable of participating in monitoring efforts and provide resilience against supply chain issues that have beset these efforts during the pandemic.
The findings also show, however, that methods-related hurdles remain before using the data for watershed-based epidemiology and modeling (e.g., estimating incidence and prevalence). This end use requires obtaining accurate information on the absolute concentration of SARS-CoV-2 genetic material in raw wastewater in addition to other information such as fecal shedding rates as noted below. Unfortunately, the accuracy of the methods—i.e., their ability to correctly quantify the true number of SARS-CoV-2 genome copies—could not be assessed because the actual concentrations in the raw wastewater samples were unknown. Despite the relatively tight band of results (80% within ±1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log), this 2
log), this 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log range may be too wide for estimating community infection since 2
log range may be too wide for estimating community infection since 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) logs represents the difference between 1% and 100% of the population being infected. Additional data gaps must also be addressed for accurately modeling community infections including information on a) viral shedding rates in feces during different stages of infection,6,20,21 b) how the genetic signal changes during travel through the wastewater collection system,22–24 and c) sewershed modeling to estimate travel time and dilution. Multiple efforts should be pursued to address these knowledge gaps.
logs represents the difference between 1% and 100% of the population being infected. Additional data gaps must also be addressed for accurately modeling community infections including information on a) viral shedding rates in feces during different stages of infection,6,20,21 b) how the genetic signal changes during travel through the wastewater collection system,22–24 and c) sewershed modeling to estimate travel time and dilution. Multiple efforts should be pursued to address these knowledge gaps.
The findings are encouraging, however, for tracking changes or trends in virus concentrations. For this purpose, the absolute numbers quantified are not as important as identifying when and to what degree those numbers are increasing or decreasing.25 The collection of SARS-CoV-2 wastewater concentrations could be used in conjunction with clinical data to provide complementary information on the extent of community infection and the effectiveness of public health interventions. The data could also be used to identify “hot spots” within a collection system where higher virus concentrations are measured.7–9 This knowledge could be used to trigger additional investigations of the populations within that sub-sewershed to identify and respond to communities experiencing higher infection rates. One benefit of this type of tracking is that the changes in wastewater concentrations may precede the clinical evidence of infection by multiple days, allowing for more responsive and focused public health interventions. A related use of this approach is confirmation of ongoing low community prevalence of SARS-CoV-2 in areas, such as small rural regions, for which testing rates are low. The use of wastewater surveillance as a sentinel for community infection has been described in Utah and at the University of Arizona.11
This study's findings would suggest that the same method or laboratory be used to assess the SARS-CoV-2 concentrations over time at a given set of locations. For example, use method A to assess trends within the sewersheds in region X over time rather than switching between methods A, B, and C over the monitoring period. Other regions (e.g., region Y) could select different methods, but should then use the same method over the entire testing period to facilitate the tracking of trends. One exception to this may be cases in which multiple laboratories use a similar SOP and have demonstrated a high degree of reproducibility across labs, such as SOPs 4.1, 4.2, and 4.3. Given the high degree of intra-method reproducibility observed (standard deviation <0.2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log GC/L), many methods have sufficient precision to sensitively detect when changes in virus concentrations are occurring. Collecting samples at multiple locations will also help identify where they are occurring.
log GC/L), many methods have sufficient precision to sensitively detect when changes in virus concentrations are occurring. Collecting samples at multiple locations will also help identify where they are occurring.
Factors promoting reproducibility
The high inter-method reproducibility was the result of three key factors: 1) the results were largely unaffected by methodological differences, 2) only data passing all QA/QC checks were included in the analysis, and 3) the QAPP normalized the findings to account for important sources of variability.Another positive finding was that the use of pasteurization prior to processing led to only modest impacts on virus enumeration when recovery correction was incorporated. This variability of approximately 0.3 to 0.4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) logs may be acceptable, particularly if pasteurization pre-treatment is a requirement for lab safety. Multiple participants in the interlaboratory comparison noted that their institutions mandated pre-pasteurization (per CDC guidelines) to minimize the lab staffs' exposure to the infectious agents in the raw wastewater (both SARS-CoV-2 and other pathogenic viruses and microorganisms). One concern was that pasteurization steps have been previously shown to impact both the infectivity and genetic signal of other viruses when heated at 72 °C.26 The QAPP prescribed lower temperature, longer duration conditions for pasteurization (60 °C for 60 minutes) since it was hypothesized that higher temperature, shorter duration conditions may have a greater impact on virus fate.27–30 While pasteurization led to a lower recovery efficiency of OC43, the uncorrected SARS-CoV-2 concentrations were not statistically different from the unpasteurized samples (Fig. S2†). This finding suggests that pasteurization does not have an important impact on the ability of the methods to detect SARS-CoV-2. Future studies could be used to confirm acceptable pasteurization conditions and quantify their impact on recovery and sensitivity.
logs may be acceptable, particularly if pasteurization pre-treatment is a requirement for lab safety. Multiple participants in the interlaboratory comparison noted that their institutions mandated pre-pasteurization (per CDC guidelines) to minimize the lab staffs' exposure to the infectious agents in the raw wastewater (both SARS-CoV-2 and other pathogenic viruses and microorganisms). One concern was that pasteurization steps have been previously shown to impact both the infectivity and genetic signal of other viruses when heated at 72 °C.26 The QAPP prescribed lower temperature, longer duration conditions for pasteurization (60 °C for 60 minutes) since it was hypothesized that higher temperature, shorter duration conditions may have a greater impact on virus fate.27–30 While pasteurization led to a lower recovery efficiency of OC43, the uncorrected SARS-CoV-2 concentrations were not statistically different from the unpasteurized samples (Fig. S2†). This finding suggests that pasteurization does not have an important impact on the ability of the methods to detect SARS-CoV-2. Future studies could be used to confirm acceptable pasteurization conditions and quantify their impact on recovery and sensitivity.
The two primer sets developed by the CDC for clinical diagnosis were used in this study. While the N1 primer set led to significantly higher concentrations than N2, these differences were considered to be minimal (approximately 0.1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log difference) compared to the other sources of variability. These findings suggest that future efforts may not need to evaluate both primer sets for tracking wastewater concentrations of SARS-CoV-2. Reducing the number of total PCR reactions per assay may be of particular interest for resource-constrained settings, though care should be taken to ensure that primer/probe sets account for mutational changes in the RNA sequence. The study also included methods using both qPCR and dPCR. Given the low number of dPCR methods evaluated, there was not sufficient statistical power to compare the results from the two platforms. Based on a preliminary analysis of the data, no clear pattern of differences emerged between the two quantification platforms suggesting both may be acceptable for future monitoring.
log difference) compared to the other sources of variability. These findings suggest that future efforts may not need to evaluate both primer sets for tracking wastewater concentrations of SARS-CoV-2. Reducing the number of total PCR reactions per assay may be of particular interest for resource-constrained settings, though care should be taken to ensure that primer/probe sets account for mutational changes in the RNA sequence. The study also included methods using both qPCR and dPCR. Given the low number of dPCR methods evaluated, there was not sufficient statistical power to compare the results from the two platforms. Based on a preliminary analysis of the data, no clear pattern of differences emerged between the two quantification platforms suggesting both may be acceptable for future monitoring.
Moving forward, additional elements could be specified in the QAPP that may further improve the reproducibility across methods. For example, specifying the type of standards to be used, the RNA extraction methods, and how the samples are shipped and stored prior to processing may further control variability. The high reproducibility between SOPs 4.1, 4.2, and 4.3 also suggests that greater consistency between SOPs and improved coordination between labs can further improve reproducibility.
One challenge with correcting for recovery is that it assumes that the matrix spike behaves similarly to the target virus. Additional studies are needed to assess how well OC43 mimics SARS-CoV-2 behavior in wastewater matrices, meaning that correcting based on OC43 (or any other viral surrogate) may also introduce some degree of variability in the results. For example, differences between SARS-CoV-2 and the matrix spike organism in terms of solids association, thermal sensitivity, extraction efficiency and surface properties may lead to variability when correcting for recovery after solids removal steps, pasteurization, and concentration methods, respectively. Nevertheless, the differences between SARS-CoV-2 and OC43 are likely to have a smaller net impact on the results than differences in recovery efficiency. The similarity in recovery efficiencies of the three betacoronaviruses tested in this study (OC43, BCoV, and heat-inactivated SARS-CoV-2) provides some assurance that OC43 may behave in a similar fashion to SARS-CoV-2. In a post-study poll of the laboratory participants, 87% supported the practice of reporting and correcting for recovery efficiency. Additional work to confirm the selection of matrix spike organisms is recommended.
Evolving the methods
Demonstrating the high degree of reproducibility between methods is an important step because it confirms that multiple methods can be used to obtain similar results in these complex matrices. This does not mean, however, that all of the methods are equally suited for all future efforts. One of the most promising end uses for these methods is to track SARS-CoV-2 concentrations in wastewater as a bellwether for community health. Ideally, methods employed for such uses would have both high precision to identify upward or downward trends in the data as well as high sensitivity to quantify concentrations in both epidemic (high community infection) and endemic (low community infection) settings. To understand how the sensitivity of these methods translates to potential application of this tool in endemic settings, the prevalence of COVID-19 in Los Angeles County at the time of sampling was estimated. Assuming infected individuals shed SARS-CoV-2 in in their feces for at least 27 days,6 then 61![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 people with confirmed infections were shedding SARS-CoV-2 in the wastewater samples collected during the study.32 In a population of ten million people, this corresponds to 1 in 160 people. At this level of community infection, nearly all of the methods were able to achieve quantifiable results of virus concentrations. The degree to which the concentration in the wastewater (and consequently the percent of the population infected) could decrease while still obtaining quantifiable numbers will vary across the methods.
000 people with confirmed infections were shedding SARS-CoV-2 in the wastewater samples collected during the study.32 In a population of ten million people, this corresponds to 1 in 160 people. At this level of community infection, nearly all of the methods were able to achieve quantifiable results of virus concentrations. The degree to which the concentration in the wastewater (and consequently the percent of the population infected) could decrease while still obtaining quantifiable numbers will vary across the methods.
The methods showed a sizable range of theoretical limits of detection with most falling in the 103 to 106 GC/L range (in comparison, the measured SARS-CoV-2 were generally in the range of 104 to 106 GC/L). Methods with theoretical LODs as low as 102 GC/L were also identified that would offer a 10- to 1000-fold improvement over those methods. Although methods with varying LODs reported similar corrected values in this study, it should be emphasized that the use of higher-sensitivity methods will reduce the probability of obtaining NDs. Consequently, the selection of more sensitive methods should be prioritized to track trends over a range of concentrations. To make this selection, one should target methods with low LODs (Fig. 4). Additional studies should identify the methods best suited for tracking trends, particularly those that offer high precision, reproducibility, and sensitivity. As the call for more expansive state- and nationwide monitoring programs increases, methods that offer higher throughput and lower processing time may also rise to the top.
The findings can also be used to identify methods that are best suited for areas with greater resource constraints, including those without the financial, technical, and material resources available in large U.S. cities. Through this lens, methods that have lower material costs, fewer and simpler steps, and require less specialized knowledge could offer important advantages. For example, the direct extraction methods forego the use of downstream concentration steps eliminating the need for filtration devices, centrifuges, and additional chemicals. Consequently, these methods may be cheaper, faster, and easier to run. Further research is needed to show if these methods can also provide sufficient precision, reproducibility, and sensitivity, to be the methods of choice for the diversity of locations across the country and globe.
5 Conclusions
• A nationwide interlaboratory comparison of methods for the quantification of SARS-CoV-2 genetic signal in wastewater showed a high degree of reproducibility. 80% of the results from eight method groups (36 different methods) fell within a band of approximately ±1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log GC/L when corrected for recovery. These findings suggest that a variety of methods are capable of producing reproducible results, though the same SOP or laboratory should be selected to track SARS-CoV-2 trends at a given facility.
log GC/L when corrected for recovery. These findings suggest that a variety of methods are capable of producing reproducible results, though the same SOP or laboratory should be selected to track SARS-CoV-2 trends at a given facility.
• Based on the seven order of magnitude range of recovery efficiencies reported in this study, it is recommended that future methods include matrix spikes to quantify and correct for recovery in order to obtain reproducible numbers between methods.
• Recovery-corrected results did not show a systematic impact from solids removal or concentration method used. Additional methods steps that were evaluated (e.g., pasteurization, primer set selection, and PCR platform) generally resulted in small differences compared to other sources of variability.
• Factors leading to greater interlaboratory reproducibility include a) the relative insensitivity of the findings to methodological differences, b) the implementation of strict QA/QC requirements, c) the use of a quality assurance project plan to normalize the findings and account for important sources of variability, and d) implementing a shared SOP among different laboratories.
• The findings support the use of wastewater surveillance for tracking trends in the concentrations of SARS-CoV-2 within communities. They also highlight methodological challenges related to modeling incidence and prevalence.
• Additional metrics should be used to select the best methods for future efforts including method sensitivity, cost, equipment requirements, and simplicity.
6 Disclaimer
This manuscript has been reviewed by the U.S. EPA and approved for publication. Approval does not signify that the contents reflect the views of the Agency, nor does mention of trade names or commercial products constitute endorsement or recommendation for use.7 Additional information
Group author details: SARS-CoV-2 interlaboratory consortium
Tiong Gim Aw: Environmental Health Sciences, School of Public Health and Tropical Medicine, Tulane University, New Orleans, LA; Nichole E. Brinkman: Office of Research and Development, U.S. Environmental Protection Agency, Cincinnati, OH; Kartik Chandran: Earth and Environmental Engineering, Columbia University, New York, NY; Francoise Chauvin: Bureau of Wastewater Treatment, New York City Department of Environmental Protection, Flushing, NY; John J. Dennehy: Biology, Queens College and The Graduate Center of The City University of New York, Queens, NY; Phil Dennis: SiREM Laboratory, Guelph, ON; Shuchen Feng: School of Freshwater Sciences, University of Wisconsin-Milwaukee, Milwaukee, WI; Matthew T. Flood: Fisheries and Wildlife, Michigan State University, East Lansing, MI; Raul Gonzalez: Hampton Roads Sanitation District, Virginia Beach, VA; Joe Hernandez: Microbiology, City of Scottsdale Water Campus, Scottsdale, AZ; Kayley H. Janssen: Wisconsin State Laboratory of Hygiene, University of Wisconsin-Madison, Madison, WI; Sunny Jiang: Civil and Environmental Engineering, University of California – Irvine, Irvine, CA; Marc C. Johnson: Molecular Microbiology and Immunology, University of Missouri, Columbia, MO; Devrim Kaya: Civil and Environmental Engineering, University of Maryland – College Park, College Park, MD; Huiling R. Lee: Mycometrics, LLC, Monmouth Jct, NJ; Jiyoung Lee: Division of Environmental Health Sciences, College of Public Health & Department of Food Science and Technology, Ohio State University, Columbus, OH; Xu Li: Civil and Environmental Engineering, University of Nebraska-Lincoln, Lincoln, NE; Cresten Mansfeldt: Civil. Environmental, and Architectural Engineering, University of Colorado – Boulder, Boulder, CO; Subhanjan Mondal: Promega Corporation, Fitchburg, WI; Kara L Nelson: Civil and Environmental Engineering, University of California – Berkeley, Berkeley, CA; Katerina Papp: Applied Research and Development Center, Southern Nevada Water Authority, Las Vegas, NV; Agustin E. Pierri: Weck Laboratories, Inc., Industry, CA; Catherine B. Pratt: College of Public Health, University of Nebraska Medical Center, Omaha, NE; Anda Quintero: Source Molecular Corporation, Miami Lakes, FL; Tyler Radniecki: School of Chemical, Biological and Environmental Engineering, Oregon State University, Corvallis, OR; Ryan A. Reinke: Microbiology, Los Angeles County Sanitation Districts, Whittier, CA; D. Keith Roper: Biological Engineering, Utah State University, Logan, UT; Tami L. Sivy: Chemistry, Saginaw Valley State University, University Center, MI; Brian M. Swalla: IDEXX Laboratories, Inc., Westbrook, ME; Jennifer Weidhaas: Civil and Environmental Engineering, University of Utah, Salt Lake City, UT.Statement of author contributions
Brian Pecson and Emily Darby developed the project plan, analyzed the data, and wrote the manuscript; Charles Haas participated in the statistical analysis, interpretation, and presentation of the findings; Yamrot Amha, Liana Olivas, and Yan Qu were responsible for the collection, preparation, and distribution of the wastewater samples; Mitchel Bartolo supported the study planning and data analysis; Richard Danielson and Yeggie Dearborn provided input on the project's Quality Assurance Project Plan and processed the wastewater samples alongside the SARS-CoV-2 interlaboratory consortium; George Lukasik and Bonnie Mull prepared the matrix spike and other shared reagents, provided input on the project's Quality Assurance Project Plan, and processed the wastewater samples alongside the SARS-CoV-2 interlaboratory consortium; George Di Giovanni, Erica Gaddis, Don Gray, and Adam Olivieri were members of the WRF Project Advisory Committee that advised on project planning and data interpretation; Christobel Ferguson and Stephanie Fevig provided coordination and organizational support between the research team, Project Advisory Committee, and laboratories; the SARS-CoV-2 interlaboratory consortium (group author list) processed the wastewater samples.Conflicts of interest
There are no conflicts of interest to declare.Acknowledgements
The authors would like to thank The Water Research Foundation (project No. 5089) and the Bill & Melinda Gates Foundation for funding this research. We thank Mia Mattioli (Centers for Disease Control and Prevention) for her guidance on the project advisory committee, and Hunter Johnson and Mark Keller (interns at Trussell Technologies) for support with the project planning and sample collection. We thank the City of Los Angeles Sanitation and Environment and the Los Angeles County Sanitation District for supporting the sample collection.References
- W. Ahmed, N. Angel, J. Edson, K. Bibby, A. Bivins, J. W. O'Brien, P. M. Choi, M. Kitajima, S. L. Simpson, J. Li, B. Tscharke, R. Verhagen, W. J. M. Smith, J. Zaugg, L. Dierens, P. Hugenholtz, K. V. Thomas and J. F. Mueller, First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community, Sci. Total Environ., 2020, 728, 138764 CrossRef CAS.
- G. Medema, L. Heijnen, G. Elsinga, R. Italiaander and A. Brouwer, Presence of SARS-Coronavirus-2 in sewage, medRxiv, 2020, DOI:10.1101/2020.03.29.20045880.
- F. Wu, A. Xiao, J. Zhang, X. Gu, W. L. Lee, K. Kauffman, W. Hanage, M. Matus, N. Ghaeli, N. Endo, C. Duvallet, K. Moniz, T. Erickson, P. Chai, J. Thompson and E. Alm, SARS-CoV-2 titers in wastewater are higher than expected from clinically confirmed cases, medRxiv, 2020, DOI:10.1101/2020.04.05.20051540.
- R. Wölfel, V. M. Corman, W. Guggemos, M. Seilmaier, S. Zange, M. A. Müller, D. Niemeyer, T. C. Jones, P. Vollmar, C. Rothe, M. Hoelscher, T. Bleicker, S. Brünink, J. Schneider, R. Ehmann, K. Zwirglmaier, C. Drosten and C. Wendtner, Virological assessment of hospitalized patients with COVID-2019, Nature, 2020, 581, 465–469, DOI:10.1038/s41586-020-2196-x.
- R. Zang, M. F. G. Castro, B. T. McCune, Q. Zeng, P. W. Rothlauf, N. M. Sonnek, Z. Liu, K. F. Brulois, X. Wang, H. B. Greenberg, M. S. Diamond, M. A. Ciorba, S. P. J. Whelan and S. Ding, TMPRSS2 and TMPRSS4 mediate SARS-CoV-2 infection of human small intestinal enterocytes, bioRxiv, 2020, DOI:10.1101/2020.04.21.054015.
- Y. Xu, X. Li, B. Zhu, H. Liang, C. Fang, Y. Gong, Q. Guo, X. Sun, D. Zhao, J. Shen, H. Zhang, H. Liu, H. Xia, J. Tang, K. Zhang and S. Gong, Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding, Nat. Med., 2020, 26, 502–505 CrossRef CAS.
- R. Gonzalez, K. Curtis, A. Bivins, K. Bibby, M. H. Weir, K. Yetka, H. Thompson, D. Keeling, J. Mitchell and D. Gonzalez, COVID-19 surveillance in Southeastern Virginia using wastewater-based epidemiology, Water Res., 2020, 186, 9 CrossRef.
- M. Hellmér, N. Paxéus, L. Magnius, L. Enache, B. Arnholm, A. Johansson, T. Bergström and H. Norder, Detection of pathogenic viruses in sewage provided early warnings of hepatitis A virus and norovirus outbreaks, Appl. Environ. Microbiol., 2014, 80, 6771–6781 CrossRef.
- A. F. Brouwer, J. N. S. Eisenberg, C. D. Pomeroy, L. M. Shulman, M. Hindiyeh, Y. Manor, I. Grotto, J. S. Koopman and M. C. Eisenberg, Epidemiology of the silent polio outbreak in Rahat, Israel, based on modeling of environmental surveillance data, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, E10625–E10633 CrossRef CAS.
- I. Michael-Kordatou, P. Karaolia and D. Fatta-Kassinos, Sewage analysis as a tool for the COVID-19 pandemic response and management: the urgent need for optimised protocols for SARS-CoV-2 detection and quantification, J. Environ. Chem. Eng., 2020, 8, 104306 CrossRef CAS.
- J. Weidhaas, Z. Aanderud, D. Roper, J. VanDerslice, E. Gaddis, J. Ostermiller, K. Hoffman, R. Jamal, P. Heck, Y. Zhang, K. Torgersen, J. Vander Laan and N. LaCross, Correlation of SARS-CoV-2 RNA in wastewater with COVID-19 disease burden in sewersheds, Sci. Total Environ., 2020 DOI:10.21203/rs.3.rs-40452/v1.
- Water Research Foundation, Wastewater surveillance of the COVID-19 genetic signal in sewersheds, Water Research Foundation, Alexandria, VA, 2020 Search PubMed.
- M. Kitajima, W. Ahmed, K. Bibby, A. Carducci, C. P. Gerba, K. A. Hamilton, E. Haramoto and J. B. Rose, SARS-CoV-2 in wastewater: State of the knowledge and research needs, Sci. Total Environ., 2020, 739, 19 CrossRef.
- Trussell Technologies, WRF 5089 Quality Assurance Project Plan, https://www.waterrf.org/research/projects/interlaboratory-and-methods-assessment-sars-cov-2-genetic-signal-wastewater, (accessed October 19, 2020).
- G. Medema, personal communication, 2020.
- R Core Team, R: a language and environment for statistical computing, Vienna, Austria, 2016 Search PubMed.
- A. Forootan, R. Sjöback, J. Björkman, B. Sjögreen, L. Linz and M. Kubista, Methods to determine limit of detection and limit of quantification in quantitative real-time PCR (qPCR), Biomol. Detect. Quantif., 2017, 12, 1–6 CrossRef CAS.
- M. A. Jahne, N. E. Brinkman, S. P. Keely, B. D. Zimmerman, E. A. Wheaton and J. L. Garland, Droplet digital PCR quantification of norovirus and adenovirus in decentralized wastewater and graywater collections: Implications for onsite reuse, Water Res., 2020, 169, 115213 CrossRef CAS.
- J. A. Steele, A. D. Blackwood, J. F. Griffith, R. T. Noble and K. C. Schiff, Quantification of pathogens and markers of fecal contamination during storm events along popular surfing beaches in San Diego, California, Water Res., 2018, 136, 137–149 CrossRef CAS.
- K. S. Cheung, I. F. N. Hung, P. P. Y. Chan, K. C. Lung, E. Tso, R. Liu, Y. Y. Ng, M. Y. Chu, T. W. H. Chung, A. R. Tam, C. C. Y. Yip, K.-H. Leung, A. Y.-F. Fung, R. R. Zhang, Y. Lin, H. M. Cheng, A. J. X. Zhang, K. K. W. To, K.-H. Chan, K.-Y. Yuen and W. K. Leung, Gastrointestinal Manifestations of SARS-CoV-2 Infection and Virus Load in Fecal Samples From a Hong Kong Cohort: Systematic Review and Meta-analysis, Gastroenterology, 2020, 159, 81–95 CrossRef CAS.
- F. Xiao, M. Tang, X. Zheng, Y. Liu, X. Li and H. Shan, Evidence for Gastrointestinal Infection of SARS-CoV-2, Gastroenterology, 2020, 158, 1831–1833.e1833 CrossRef CAS.
- L. M. Casanova and S. R. Weaver, Inactivation of an enveloped surrogate virus in human sewage, Environ. Sci. Technol. Lett., 2015, 2, 76–78 CrossRef CAS.
- Y. Ye, R. M. Ellenberg, K. E. Graham and K. R. Wigginton, Survivability, Partitioning, and Recovery of Enveloped Viruses in Untreated Municipal Wastewater, Environ. Sci. Technol., 2016, 50, 5077–5085 CrossRef CAS.
- W. Ahmed, P. M. Bertsch, K. Bibby, E. Haramoto, J. Hewitt, F. Huygens, P. Gyawali, A. Korajkic, S. Riddell, S. P. Sherchan, S. L. Simpson, K. Sirikanchana, E. M. Symonds, R. Verhagen, S. S. Vasan, M. Kitajima and A. Bivins, Decay of SARS-CoV-2 and surrogate murine hepatitis virus RNA in untreated wastewater to inform application in wastewater-based epidemiology, Environ. Res., 2020, 191, 110092 CrossRef CAS.
- P. M. Lago, H. E. Gary Jr., L. S. Pérez, V. Cáceres, J. B. Olivera, R. P. Puentes, M. B. Corredor, P. Jímenez, M. A. Pallansch and R. G. Cruz, Poliovirus detection in wastewater and stools following an immunization campaign in Havana, Cuba, Int. J. Epidemiol., 2003, 32, 772–777 CrossRef.
- B. M. Pecson, L. V. Martin and T. Kohn, Quantitative PCR for Determining the Infectivity of Bacteriophage MS2 upon Inactivation by Heat, UV-B Radiation, and Singlet Oxygen: Advantages and Limitations of an Enzymatic Treatment To Reduce False-Positive Results, Appl. Environ. Microbiol., 2009, 75, 5544–5554 CrossRef CAS.
- A. S. Jureka, J. A. Silvas and C. F. Basler, Propagation, Inactivation, and Safety Testing of SARS-CoV-2, Viruses, 2020, 12, 622 CrossRef CAS.
- H. F. Rabenau, J. Cinatl, B. Morgenstern, G. Bauer, W. Preiser and H. W. Doerr, Stability and inactivation of SARS coronavirus, Med. Microbiol. Immunol., 2005, 194, 1–6 CrossRef CAS.
- M. E. R. Darnell, K. Subbarao, S. M. Feinstone and D. R. Taylor, Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV, J. Virol. Methods, 2004, 121, 85–91 CrossRef CAS.
- A. Brié, I. Bertrand, M. Meo, N. Boudaud and C. Gantzer, The Effect of Heat on the Physicochemical Properties of Bacteriophage MS2, Food Environ. Virol., 2016, 8, 251–261 CrossRef.
- S. W. Hasan, Y. Ibrahim, M. Daou, H. Kannout, N. Jan, A. Lopes, H. Alsafar and A. F. Yousef, Detection and quantification of SARS-CoV-2 RNA in wastewater and treated effluents: Surveillance of COVID-19 epidemic in the United Arab Emirates, Sci. Total Environ., 2020, 142929, DOI:10.1016/j.scitotenv.2020.142929.
- CDPH, California open data portal — COVID-19 cases, https://data.ca.gov/dataset/covid-19-cases, (accessed October 9, 2020).
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/d0ew00946f |
| ‡ These authors contributed equally. |
| § Details of the participating individuals and institutions are provided in Table 1 and Section 7. |
| This journal is © The Royal Society of Chemistry 2021 |