Enzymatically degradable, starch-based layer-by-layer films: application to cytocompatible single-cell nanoencapsulation†
Abstract
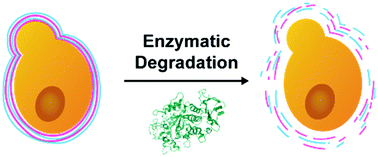
The build-up and degradation of cytocompatible nanofilms in a controlled fashion have great potential in biomedical and nanomedicinal fields, including single-cell nanoencapsulation (SCNE). Herein, we report the fabrication of biodegradable films of cationic starch (c-ST) and anionic alginate (ALG) by electrostatically driven layer-by-layer (LbL) assembly technology and its application to the SCNE. The [c-ST/ALG] multilayer nanofilms, assembled either on individual Saccharomyces cerevisiae or on the 2D flat gold surface, degrade on demand, in a cytocompatible fashion, via treatment with α-amylase. Their degradation profiles are investigated, while systematically changing the α-amylase concentration, by several surface characterization techniques, including quartz crystal microbalance with dissipation monitoring (QCM-D) and ellipsometry. DNA incorporation in the LbL nanofilms and its controlled release, upon exposure of the nanofilms to an aqueous α-amylase solution, are demonstrated. The highly cytocompatible nature of the film-forming and -degrading conditions is assessed in the c-ST/ALG-shell formation and degradation of S. cerevisiae. We envisage that the cytocompatible, enzymatic degradation of c-ST-based nanofilms paves the way for developing advanced biomedical devices with programmed dissolution in vivo.


 Please wait while we load your content...
Please wait while we load your content...