Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Highly efficient H2S scavengers via thiolysis of positively-charged NBD amines†
Ismail
Ismail‡
 a,
Zhuoyue
Chen‡
b,
Lu
Sun‡
a,
Zhuoyue
Chen‡
b,
Lu
Sun‡
 c,
Xiuru
Ji
c,
Haishun
Ye
b,
Xueying
Kang
b,
Haojie
Huang
b,
Haibin
Song
a,
Sarah G.
Bolton
d,
Zhen
Xi
a,
Michael D.
Pluth
c,
Xiuru
Ji
c,
Haishun
Ye
b,
Xueying
Kang
b,
Haojie
Huang
b,
Haibin
Song
a,
Sarah G.
Bolton
d,
Zhen
Xi
a,
Michael D.
Pluth
 d and
Long
Yi
d and
Long
Yi
 *b
*b
aState Key Laboratory of Elemento-Organic Chemistry and Department of Chemical Biology, College of Chemistry, National Pesticide Engineering Research Center, Collaborative Innovation Center of Chemical Science and Engineering, Nankai University, Tianjin 300071, China
bState Key Laboratory of Organic–Inorganic Composites, Beijing Key Lab of Bioprocess, Beijing University of Chemical Technology (BUCT), Beijing 100029, China. E-mail: yilong@mail.buct.edu.cn
cTianjin Key Laboratory on Technologies Enabling Development of Clinical Therapeutics and Diagnostics (Theranostics), School of Pharmacy, Tianjin Medical University, Tianjin 300070, China
dDepartment of Chemistry and Biochemistry, Materials Science Institute, Knight Campus for Accelerating Scientific Impact, Institute of Molecular Biology, University of Oregon, Eugene, OR 97403, USA
First published on 9th July 2020
Abstract
H2S is a well-known toxic gas and also a gaseous signaling molecule involved in many biological processes. Advanced chemical tools that can regulate H2S levels in vivo are useful for understanding H2S biology as well as its potential therapeutic effects. To this end, we have developed a series of 7-nitro-1,2,3-benzoxadiazole (NBD) amines as potential H2S scavengers. The kinetic studies of thiolysis reactions revealed that incorporation of positively-charged groups onto the NBD amines greatly increased the rate of the H2S-specific thiolysis reaction. We demonstrate that these reactions proceed effectively, with second order rate constants (k2) of >116 M−1 s−1 at 37 °C for NBD-S8. Additionally, we demonstrate that NBD-S8 can effectively scavenge enzymatically-produced and endogenous H2S in live cells. Furthering the biological significance, we demonstrate NBD-S8 mediates scavenging of H2S in mice.
Introduction
Hydrogen sulfide (H2S) has long been known as a toxic gas, but recent studies indicate that H2S has important physiological functions, leading to its inclusion as the third gasotransmitter along with nitric oxide (NO) and carbon monoxide (CO).1–3 Endogenous H2S is enzymatically generated by cystathionine γ-lyase (CSE), cystathionine-β-synthase (CBS) and 3-mercaptopyruvate sulfurtransferase (3-MST)/cysteine aminotransferase (CAT).4 H2S influences a wide range of physiological processes in mammals, ranging from vasorelaxation,5 cardioprotection,6 and neurotransmission7 to anti-inflammatory action8 and angiogenesis.9 Misregulation of H2S is associated with numerous diseases.10,11 Specially, low levels of endogenous H2S appears to exhibit pro-cancer effects, whereas higher concentrations of H2S can lead to cell apoptosis and have anti-cancer characteristics.11 Complementing the importance of H2S in mammals, H2S in bacteria and plants also plays many important functions.12,13 Due to its wide biodistribution and complex behaviors in many diseases, the physiological characters of H2S and the molecular mechanisms in which H2S is involved need further investigation. In addition, these factors support the development and refinement of advanced chemical tools that can visualize,14 scavenge15 or release16 H2S in vivo and in related complex environments.Significant attention has focused on developing chemical tools for H2S detection and delivery,14,16 but much less effort has focused on reducing H2S levels in complex environments.15,17 Such H2S elimination could be achieved by either inhibition of H2S biosynthesis or by the selective scavenging of H2S. Though CBS inhibitors have been reported to show anti-tumor activity, many of CBS inhibitors target the pyridoxal-5′-phosphate (PLP) cofactor and therefore have low specificity or unwanted side effects.11b Complementing these challenges, both CSE and 3-MST inhibitors are relatively underdeveloped.11a Moreover, H2S can be produced from non-enzymatic processes,14 which suggests that enzymatic inhibitors can only perturb certain pools of biological H2S genesis. Recently, sulfonyl azides were reported as highly efficient H2S scavengers in buffers, enzymatic systems, and living biological environments.15 It should be noted that the H2S-mediated reduction of aryl azides to amines results in generation of sulfane sulfur and polysulfides,15,18 both of which are important reactive sulfur species that might complicate such scavenging.19a Considering the in vivo biological complex,19 our goal was to develop efficient H2S scavengers utilizing different reaction mechanisms that not only scavenge H2S efficiently, but also do not generate other reactive sulfur species in the scavenging process with the long-term goal of applying these toward in vivo systems.
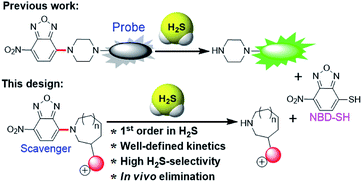
A major challenge in the development of H2S scavengers is developing fast chemical reactions that effectively differentiate the reactivity of biological nucleophiles (e.g. biothiols) from H2S. To this end, we as well as others have previously reported the H2S-specific thiolysis of 7-nitro-1,2,3-benzoxadiazole (NBD) amine in 2013 (Fig. 1),20 which has enabled, amongst other applications, the development of near-infrared H2S probes for imaging of H2S in vivo.20c To further advance the development of H2S scavengers, we report here that simple modifications of the NBD electrophiles can increase the reactivity of this platform to enable efficient H2S scavenging (Fig. 2) and apply these chemical tools to scavenge H2S in buffer, serum, cells, and mice. We believe that the present H2S scavengers could be useful tools to study H2S biology as well as potential therapeutic agents in the future.
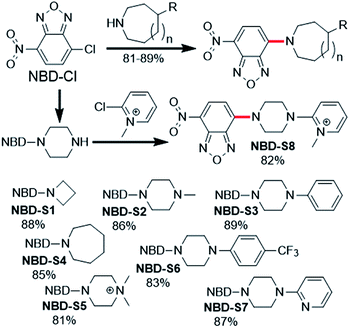 | ||
| Fig. 2 Synthesis of NBD amines NBD-S1 to NBD-S8. The C–N bond cleaved by H2S-mediated thiolysis is highlighted in red in each structure. | ||
Results and discussion
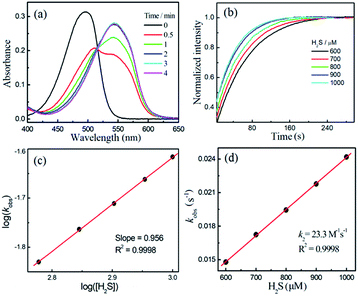
Compounds NBD-S1 to NBD-S8 were synthesized from commercially available reagents by simply coupling NBD-Cl to the desired amine or by reacting NBD-piperizine with 2-chloro-1-methylpyridinium salt (Fig. 2). Compound identity was confirmed by 1H and 13C NMR spectroscopy and HRMS. All scavengers were prepared in high yield in either one- or two-step syntheses, which readily enables access to large quantities of each scavenger.To measure the rate of the prepared compounds with H2S, we monitored the optical response of NBD-S1–NBD-S8 (5–10 μM) after addition of H2S (6–600 equiv., using Na2S) in PBS buffer (pH 7.4) at 25 °C. An example of the observed reactivity is shown for NBD-S2 (Fig. 3a), in which the starting absorbance peak at 490 nm from the NBD moiety shifts to 540 nm after reaction with H2S due to formation of NBD-SH.20b The reaction kinetics were monitored by measuring absorbance data at 540 nm or 490 nm (Fig. S1†) and showed a clear isosbestic point at 510 nm, which is consistent with a clean overall transformation. The pseudo-first-order rate constant, kobs was determined by fitting the intensity data to a single exponential function (Fig. 3b). Plotting log(kobs) versus log([H2S]) confirmed a first-order dependence in H2S (Fig. 3c). The linear fitting between kobs and H2S concentrations gives the second-order rate constant (k2) of 23.3 M−1 s−1 (Fig. 3d).
Using the above method, the H2S-mediated thiolysis rates for the prepared NBD amines were measured, and these data are summarized in Table 1. The thiolysis rates for the four, six, and seven-membered ring amines (NBD-S1 to NBD-S4) are significantly different, with the presence of the second amine group in the ring appearing to be particular important. We hypothesized that this increased rate could be due to protonation of the piperizine nitrogen at physiological pH (e. g.: the pKa of 1-methyl piperizine is ∼9).21 To verify this hypothesis, compounds NBD-S5 to NBD-S8 with the functionlized second amine were further tested. As expected, alkylation of the piperizine nitrogen (NBD-S5) or inclusion of a positively charged pyridinium (NBD-S8) significantly increased the thiolysis reaction rates. For example, k2 values for cationic NBD-S5 and NBD-S8 derivatives were 70.2 M−1 s−1 and 76.2 M−1 s−1 at 25 °C, respectively, which are significantly faster than the k2 values of the neutral NBD-S2 and NBD-S7 analogues. To further support these measurements, we also performed kinetic analysis using fluorescence methods (Fig. S9 and S10†) for NBD-S5 and NBD-S8, which afforded k2 values of 71.6 M−1 s−1 and 78.3 M−1 s−1, respectively, which are consistent with those from absorbance measurements. We also measured k2 for NBD-S8 at 37 °C, which yielded a significantly higher value of 116.1 M−1 s−1 (Fig. S11†), and suggests that this compound can scavenge H2S efficiently at physiological temperatures. The formation of the amine product during the thiolysis reaction was confirmed by HPLC and HRMS analysis (Fig. S12 and S13†). Taken together, these data suggest that NBD-S5 and NBD-S8 are highly reactive reagents toward H2S clearance.
| NBD-S1 | NBD-S2 | NBD-S3 | NBD-S4 | NBD-S5 | NBD-S6 | NBD-S7 | NBD-S8 | |
|---|---|---|---|---|---|---|---|---|
| a k obs: the value was obtained by fitting the time-dependent spectra data with single exponential function. b [H2S]: the H2S concentration used at the determination of the kobs value. c Slope: the slope value of the linear fitting of log(kobs) versus log([H2S]). d k 2: the slope value of the linear fitting of kobsversus [H2S]. e N.D.: no detection due to the very slow reaction. | ||||||||
| kobsa/s−1 | 4.2 × 10−3 | 2.4 × 10−2 | 2.7 × 10−2 | N.D.e | 9.1 × 10−3 | 2.2 × 10−2 | 3.6 × 10−2 | 9.3 × 10−3 |
| ([H2S]b) | 6.0 mM | 1.0 mM | 1.5 mM | 120 μM | 1.2 mM | 2.0 mM | 120 μM | |
| Slopec | 0.937 | 0.956 | 1.039 | N.D. | 0.898 | 1.066 | 1.048 | 0.979 |
| k 2 /M−1 s−1 | 0.67 | 23.3 | 18.0 | < 0.01 | 70.2 | 19.7 | 18.8 | 76.2 |
We also obtained crystals suitable for single-crystal X-ray diffraction studies of NBD-S7 and NBD-S8 (Fig. 4). In the solid state, the piperazinyl heterocycles in NBD-S7 and NBD-S8 are in twist-boat and chair conformations, respectively. Prior work with electrophilic piperazine-containing cyanine GSH probes has shown that the chair conformation is favorable for the thiolysis reaction,22a which may help explain the increased reaction rate of NBD-S8 when compared to NBD-S7. In addition, the positive charge of NBD-S8 may favor reaction with anionic HS−, and prior work with probes for F− has shown increased rates for cationic versions of probes when compared to neutral counterparts.22b
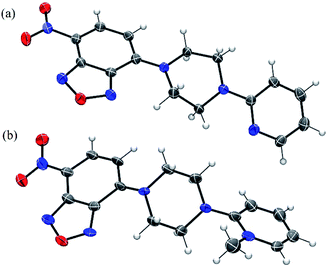 | ||
| Fig. 4 ORTEP diagram (50% ellipsoid) for the molecular structure of NBD-S7 (a) and NBD-S8 (b). The I− counterion in (b) is omitted for clarity. | ||
We also evaluated the stability, water-solubility, and selectivity of NBD-S5 and NBD-S8. The plots of absorbance intensity against the scavenger concentration (up to 100 μM) were linear (Fig. S14 and S15†), suggesting the good solubility of NBD-S5 and NBD-S8 in PBS buffer (50 mM, pH = 7.4, containing 2% DMSO). Both NBD-S5 and NBD-S8 were shown to be stable in PBS buffer, with no decomposition over 2 days (Fig. S16†). The high selectivity of the NBD amines toward H2S over biothiols were validated by HPLC analysis (Fig. S17†), suggesting the thiolysis of the NBD amines is highly H2S-specific.20c We propose that this high selectivity may originate from the intrinsic low thiolysis reactivity for C–N bond cleavage (e.g.NBD-S1), and that certain structural factors, such as the piperazine chair conformation and positively-charged group, may contribute to the faster reactivity with HS− (e.g.NBD-S5 and NBD-S8). Therefore, NBD-S5 and NBD-S8 were further employed as H2S scavengers in the following studies.
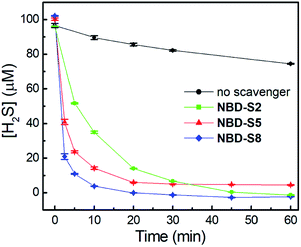
We first tested the efficiency of H2S clearance by NBD amines in PBS buffer (50 mM, pH 7.4), using the methylene blue assay (MBA)23 to quantify H2S concentrations (Fig. S18–S21†). The time-dependent H2S concentrations in the absence or presence of NBD-S2, NBD-S5, or NBD-S8 are shown in Fig. 5. The concentration of H2S slowly decreasd over time, which matches previous observations.24 Under these conditions, both NBD-S5 and NBD-S8 can scavenge more than 50% H2S within the first 2 min. Additionally, NBD-S8 can scavenge more than 90% H2S within 5 min, suggesting it is a highly efficient H2S scavenger in aqueous buffer (pH 7.4). Based on these parameters, we used NBD-S8 for subsequent applications. Considering H2S is a toxic gas,2b we tested the ability of the NBD-S8 solution to remove H2S in gaseous sample (Fig. S22†).25 The color change of NBD-S8 solution and MBA tests of the solution clearly suggested NBD-S8 could efficiently scavenge H2S when the H2S-containing gas was passed through the NBD-S8 solution.
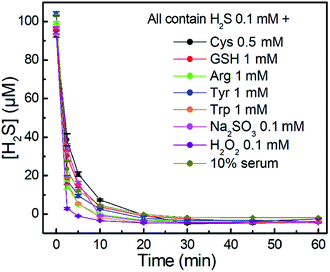
We further investigated the time-dependent H2S-scavenging ability of NBD-S8 in the presence of biologically relevant species. As shown in Fig. 6, the results suggest that the scavenging of H2S viaNBD-S8 is not significantly influenced by the presence of millimolar biothiols or other species. The scavenging of H2S in 10% fetal bovine serum (FBS) was also efficient. For all tested samples, NBD-S8 could scavenge more than 90% H2S within first 10 min. These data suggest that NBD-S8 should be an efficient H2S scavenger in complex biological-related systems.
Then we tested whether NBD-S8 could scavenge H2S in live cells. We first tested the cytotoxicity of NBD-S8 by the MTT assay using HeLa cells which contain nearly no endogenous H2S. The scavenger did not show cytotoxicity up to 50 μM concentration (Fig. S23a†). Furthermore, the cell viability in the presence of 100 μM and 200 μM NBD-S8 is still greater than 90% and 80%, respectively. These results suggest that NBD-S8 is biocompatible for scavenging H2S in living cells. To confirm that the thiolysis products of NBD-S8 (NBD-SH and S8-II) are not cytotoxic, we evaluated the cytotoxicity in HeLa cells for 24 hours. NBD-SH did not show any significant cytotoxicity below 25 μM (Fig. S23b†). The cell viability in the presence of 25 μM and 100 μM S8-II is still greater than 90% and 80%, respectively (Fig. S23c†). We note that under physiological conditions, the concentration of free H2S is significantly lower than 1 μM,19a meaning that we do not expect to generate high micromolar levels of NBD-SH under normal scavenging conditions.
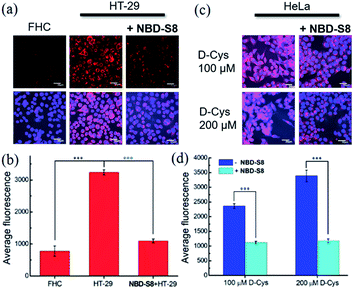
To measure H2S levels, we use the H2S-responsive fluorescence probe Cy7-NBD (Fig. S24†).20c We used HT-29 (human colonic carcinoma cells) cells for these experiments due to recent reports that colorectal cancer cell lines exhibit increased H2S production,11b which is likely due to the increased angiogenesis needed for cancer cell growth and proliferation.9 We used non-cancerous FHC (human fetal normal colonic mucosa) cells as a negative control, which we expected to lower basal H2S levels. To test whether NBD-S8 could function as an efficient H2S scavenger, we compared the fluorescence response from Cy7-NBD in FHC cells, HT-29 cells, and HT-29 cells treated with NBD-S8 (Fig. 7a and S25†). We observed a significantly higher response from Cy7-NBD in HT-29 cells when compared to FHC cells, as expected. Upon pre-treatment of HT-29 cells with 100 μM NBD-S8 for 10 min, the fluorescence signals in HT-29 cells significantly decreased (Fig. 7b), which supports that NBD-S8 can efficiently scavenge endogenous H2S in HT-29 cells.
To further support the ability of NBD-S8 to scavenge H2S in live cells, we also measured H2S levels in HeLa cells treated with D-Cys (100 μM or 200 μM), which can increase H2S biosynthesis from the 3-MST pathway.4b,26 As expected, we observed a strong red fluorescence signal from the D-Cys-treated cells in the absence of NBD-S8 (Fig. 7c and S26†).26 In contrast, co-incubation of D-Cys treated HeLa cells with 100 μM NBD-S8, the cells displayed significantly lower fluorescence than that of the control cells (Fig. 7d). Taken together with the data in HT-29 cells, these data suggest that NBD-S8 can be applied to efficiently scavenge enzymatically-produced and endogenous H2S in live cells.
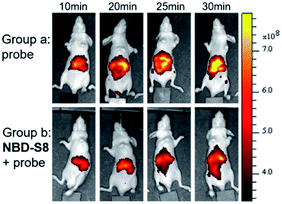
Encouraged by these results, we further examined the feasibility of NBD-S8 for H2S clearance in normal nude mice again using Cy7-NBD as an H2S reporter. Briefly, mice were intraperitoneally injected with probe only (negative control), Na2S followed by probe (positive control), or Na2S followed by NBD-S8 for 10 min and then probe. The mice were imaged using an IVIS spectrum imaging system (PerkinElmer, USA) with a 740 nm excitation filter. As expected, the mice injected with Na2S and NBD-S8 displayed weaker fluorescence signal than that of the mice injected with Na2S (Fig. S27†). These results implied that NBD-S8 still functions H2S clearance in vivo, and its feasibility to scavenge endogenous H2S in live mice was further examined. To investigate the ability to scavenge endogenous H2S, NBD-S8 was administered to 3 mice via tail vein injection. After 10 min, probe Cy7-NBD was injected to visualize the level of endogenous H2S. As a control group, another 3 mice were injected with Cy7-NBD only. As shown in Fig. 8, time-dependent fluorescence increase was detected primarily in the liver of mice (group a), which is consistent with previous results from tissue imaging.20c However, mice in (group b) exhibited comparatively lower fluorescence at each corresponding time point when compared to group a (Fig. S28†). These results preliminarily demonstrate that NBD-S8 can scavenge endogenous H2S at the organism level.
Conclusions
In summary, we have developed H2S scavengers based on the thiolysis of NBD amines. NBD-S8 shows clear reaction kinetics toward H2S with 2nd order rate constants of up to 116 M−1 s−1 at 37 °C and high selectivity toward H2S over other biologically relevant species. NBD-S8 can efficiently scavenge H2S in aqueous buffer, serum, gaseous sample, and live cells. Additionally, the high selectivity and reactivity of NBD-S8 toward H2S enables efficient H2S scavenging in live animals. Moreover, NBD-S8 showed better stability in 5 mM GSH containing PBS buffer (pH 7.4) than that of p-toluenesulfonyl azide (Fig. S29†), a recently reported H2S scavenger. Considering in vivo biological complexity,19 we propose that H2S scavengers via the H2S-specific thiolysis of NBD amines may represent an emerging class of efficient scavengers for investigating H2S in complex biological systems.Author contributions
L. Y., M. D. P., Z. X. supervised the work. I. I. synthesized the compounds; Z. C., H. Y., X. K. tested the kinetics and in vitro H2S scavenging; L. S., X. J., H. H. performed the bioimaging tests; I. I., H. S. solved crystal structures; S. G. B., X. J. tested the cytotoxicity; L. Y., M. D. P. wrote the manuscript; all authors involved in checking the manuscript and data.Ethical statement
All animal procedures were performed in accordance with the Guidelines for Care and Use of Laboratory Animals of Tianjin Medical University and approved by the Animal Ethics Committee of Tianjin Medical University.Conflicts of interest
The authors declare no competing financial interests.Acknowledgements
This work was supported by NSFC (21877008, 21837001), Beijing Natural Science Foundation (2192038) and the NIH (R01GM113030, T32GM007759).Notes and references
- (a) R. Wang, Physiol. Rev., 2012, 92, 791–896 CrossRef CAS PubMed; (b) O. Kabil and R. Banerjee, Antioxid. Redox Signal., 2014, 20, 770–782 CrossRef CAS PubMed; (c) H. Kimura, Antioxid. Redox Signal., 2010, 12, 1111–1123 CrossRef CAS PubMed.
- (a) L. Li, P. Rose and P. K. Moore, Annu. Rev. Pharmacol. Toxicol., 2011, 51, 169–187 CrossRef CAS PubMed; (b) C. Szabo, Biochem. Pharmacol., 2018, 149, 5–19 CrossRef CAS PubMed.
- (a) K. Módis, Y. J. Ju, A. Ahmad, A. A. Untereiner, Z. Altaany, L. Wu, C. Szabó and R. Wang, Pharmacol. Res., 2016, 113, 116–124 CrossRef PubMed; (b) G. K. Kolluru, X. G. Shen, S. C. Bir and C. G. Kevil, Nitric Oxide, 2013, 35, 5–20 CrossRef CAS PubMed.
- (a) H. Kimura, Amino Acids, 2011, 41, 113–121 CrossRef CAS PubMed; (b) N. Shibuya, S. Koike, M. Tanaka, M. Ishigami-Yuasa, Y. Kimura, Y. Ogasawara, K. Fukui, N. Nagahara and H. Kimura, Nat. Commun., 2013, 4, 1366–1372 CrossRef PubMed.
- (a) G. Yang, L. Wu, B. Jiang, W. Yang, J. Qi, K. Cao, Q. Meng, A. K. Mustafa, W. Mu, S. Zhang, S. H. Snyder and R. Wang, Science, 2008, 322, 587–590 CrossRef CAS PubMed; (b) D. Wu, Q. X. Hu, F. F. Ma and Y. Z. Zhu, Oxid. Med. Cell. Longevity, 2016, 2016, 7075682 Search PubMed; (c) L. Mys, N. A. Strutynska, Y. Goshovska and V. Sagach, Can. J. Physiol. Pharmacol., 2020, 98, 275–281 CrossRef CAS PubMed.
- (a) D. Wu, Q. X. Hu, Y. Xiong, D. Q. Zhu, Y. C. Mao and Y. Z. Zhu, Redox Biol., 2018, 15, 243–252 CrossRef CAS PubMed; (b) V. Citi, E. Piraqine, L. Testai, M. C. Breschi, V. Calderone and A. Martelli, Curr. Med. Chem., 2018, 25, 4380–4401 CrossRef CAS PubMed.
- (a) M. Wang, J. Zhu, Y. Pan, J. D. Dong, L. L. Zhang, X. R. Zhang and L. Zhang, J. Neurosci. Res., 2015, 93, 487–494 CrossRef CAS PubMed; (b) A. Salvi, P. Bankhele, J. M. Jamil, M. Kulkarni-Chitnis, Y. F. Njie-Mbye, S. E. Ohia and C. A. Opere, Neurochem. Res., 2016, 41, 1020–1028 CrossRef CAS PubMed.
- (a) E. Gugliandolo, R. Fusco, R. D'Amico, A. Militi, G. Oteri, J. L. Wallace, R. D. Paola and S. Cuzzocrea, Pharmacol. Res., 2018, 132, 220–231 CrossRef CAS PubMed; (b) C. Y. Zhang, Q. Z. Zhang, K. Zhang, L. Y. Li, M. D. Pluth, L. Yi and Z. Xi, Chem. Sci., 2019, 10, 1945–1952 RSC.
- A. Papapetropoulos, A. Pyriochou, Z. Altaany, G. Yang, A. Marazioti, Z. Zhou, M. G. Jeschke, L. K. Branski, D. N. Herndon, R. Wang and C. Szabó, Proc. Natl. Acad. Sci. U.S.A., 2009, 106, 21972–21977 CrossRef CAS PubMed.
- (a) C. Szabo, Nat. Rev. Drug Discov., 2016, 15, 185–203 CrossRef CAS PubMed; (b) L. M. Teigen, Z. Geng, M. J. Sadowsky, B. P. Vaughn, M. J. Hamilton and A. Khoruts, Nutrients, 2019, 11, 931 CrossRef CAS PubMed; (c) Y. Y. Han, Q. W. Shang, J. Yao and Y. Ji, Cell Death Dis., 2019, 10, 1–12 CrossRef PubMed; (d) A. Untereiner and L. Y. Wu, Antioxid. Redox Signal., 2018, 28, 1463–1482 CrossRef CAS PubMed.
- (a) X. Cao, L. Ding, Z. Z. Xie, Y. Yang, M. Whiteman, P. K. Moore and J. S. Bian, Antioxid. Redox Signal., 2019, 31, 1–38 CrossRef CAS PubMed; (b) M. R. Hellmich, C. Coletta, C. Chao and C. Szabo, Antioxid. Redox Signal., 2015, 22, 424–448 CrossRef CAS PubMed; (c) C. Szabo and A. Papapetropoulos, Pharmacol. Rev., 2017, 69, 497–564 CrossRef CAS.
- (a) K. Shatalin, E. Shatalina, A. Mironov and E. Nudler, Science, 2011, 334, 986–990 CrossRef CAS PubMed; (b) P. Shukla, V. S. Khodade, M. SharathChandra, P. Chauhan, S. Mishra, S. Siddaramappa, B. E. Pradeep, A. Singh and H. Chakrapani, Chem. Sci., 2017, 8, 4967–4972 RSC.
- (a) J. T. Hancock and M. Whiteman, Plant Physiol. Biochem., 2014, 78, 37–42 CrossRef CAS; (b) C. Alvarez, I. García, I. Moreno, M. E. Pérez-Pérez, J. L. Crespo, L. C. Romero and C. Gotor, Plant Cell, 2012, 24, 4621–4634 CrossRef CAS; (c) Z. G. Li, X. Min and Z. H. Zhou, Front. Plant Sci., 2016, 7, 1621 Search PubMed.
- (a) V. S. Lin, W. Chen, M. Xian and C. J. Chang, Chem. Soc. Rev., 2015, 44, 4596–4618 RSC; (b) F. B. Yu, X. Y. Han and L. X. Chen, Chem. Commun., 2014, 50, 12234–12249 RSC; (c) M. D. Hartle and M. D. Pluth, Chem. Soc. Rev., 2016, 45, 6108–6117 RSC; (d) P. Gao, W. Pan, N. Li and B. Tang, Chem. Sci., 2019, 10, 6035–6071 RSC; (e) R. Wang, K. Dong, G. Xu, B. Shi, T. Zhu, P. Shi, Z. Guo, W.-H. Zhu and C. Zhao, Chem. Sci., 2019, 10, 2785–2790 RSC; (f) Z. Chen, X. Mu, Z. Han, S. Yang, C. Zhang, Z. Guo, Y. Bai and W. He, J. Am. Chem. Soc., 2019, 141, 17973–17977 CrossRef CAS; (g) C. Wang, X. Cheng, J. Tan, Z. Ding, W. Wang, D. Yuan, G. Li, H. Zhang and X. Zhang, Chem. Sci., 2018, 9, 8369–8374 RSC; (h) W. Chen, T. Matsunaga, D. L. Neill, C. Yang, T. Akaike and M. Xian, Angew. Chem., Int. Ed., 2019, 58, 16067–16070 CrossRef CAS PubMed.
- C. T. Yang, Y. Y. Wang, E. Marutani, T. Ida, X. Ni, S. Xu, W. Chen, H. Zhang, T. Akaike, F. Ichinose and M. Xian, Angew. Chem., Int. Ed., 2019, 58, 10898–10902 CrossRef CAS PubMed.
- (a) Y. Zhao, T. D. Biggs and M. Xian, Chem. Commun., 2014, 50, 11788–11805 RSC; (b) C. M. Levinn, M. M. Cerda and M. D. Pluth, Acc. Chem. Res., 2019, 52, 2723–2731 CrossRef CAS PubMed; (c) C. R. Powell, K. M. Dillon and J. B. Matson, Biochem. Pharmacol., 2018, 149, 110–123 CrossRef CAS; (d) Y. Hu, X. Li, Y. Fang, W. Shi, X. Li, W. Chen, M. Xian and H. Ma, Chem. Sci., 2019, 10, 7690–7694 RSC; (e) Y. Zhao, M. M. Cerda and M. D. Pluth, Chem. Sci., 2019, 10, 1873–1878 RSC; (f) Y. Wang, K. Kaur, S. J. Scannelli, R. Bitton and J. B. Matson, J. Am. Chem. Soc., 2018, 140, 14945–14951 CrossRef CAS PubMed; (g) M. M. Cerda, T. D. Newton, Y. Zhao, B. K. Collins, C. H. Hendon and M. D. Pluth, Chem. Sci., 2019, 10, 1773–1779 RSC; (h) Y. Zheng, B. Yu, Z. Li, Z. Yuan, C. L. Organ, R. K. Trivedi, S. Wang, D. J. Lefer and B. Wang, Angew. Chem., Int. Ed., 2017, 56, 11749–11753 CrossRef CAS PubMed; (i) Y. Zhao, A. K. Steiger and M. D. Pluth, J. Am. Chem. Soc., 2019, 41, 13610–13618 CrossRef PubMed.
- S. Arndt, C. D. Baeza-Garza, A. Logan, T. Rosa, R. Wedmann, T. A. Prime, J. L. Martin, K. Saeb-Parsy, T. Krieg, M. R. Filipovic, R. C. Hartley and M. P. Murphy, J. Biol. Chem., 2017, 292, 7761–7773 CrossRef CAS PubMed.
- H. A. Henthorn and M. D. Pluth, J. Am. Chem. Soc., 2015, 137, 15330–15336 CrossRef CAS.
- (a) M. R. Filipovic, J. Zivanovic, B. Alvarez and R. Banerjee, Chem. Rev., 2018, 118, 1253–1337 CrossRef CAS PubMed; (b) O. F. Brandenberg, D. C. Miller, U. Markel, A. O. Chaib and F. H. Arnold, ACS Catal., 2019, 9, 8271–8275 CrossRef CAS PubMed; (c) V. Chalansonnet, J. Lowe, S. Orenga, J. D. Perry, S. N. Robinson, S. P. Stanforth, H. E. Sykes and T. V. Truong, Bioorg. Med. Chem. Lett., 2019, 29, 2354–2357 CrossRef CAS PubMed.
- (a) C. Wei, L. Wei, Z. Xi and L. Yi, Tetrahedron Lett., 2013, 54, 6937–6939 CrossRef CAS; (b) L. A. Montoya, T. F. Pearce, R. J. Hansen, L. N. Zakharov and M. D. Pluth, J. Org. Chem., 2013, 78, 6550–6557 CrossRef CAS; (c) K. Zhang, J. Zhang, Z. Xi, L. Y. Li, X. Gu, Q. Z. Zhang and L. Yi, Chem. Sci., 2017, 8, 2776–2781 RSC; (d) Y. L. Pak, J. Li, K. C. Ko, G. Kim, J. Y. Lee and J. Yoon, Anal. Chem., 2016, 88, 5476–5481 CrossRef CAS PubMed.
- F. Khalili, A. Henni and A. L. L. East, J. Chem. Eng. Data, 2009, 54, 2914–2917 CrossRef CAS.
- (a) J. Yin, Y. Kwon, D. Kim, D. Lee, G. Kim, Y. Hu, J. Ryu and J. Yoon, J. Am. Chem. Soc., 2014, 136, 5351–5358 CrossRef CAS PubMed; (b) Z. Wu and X. Tang, Anal. Chem., 2015, 87, 8613–8617 CrossRef CAS.
- L. M. Siegel, Anal. Biochem., 1965, 11, 126–132 CrossRef CAS.
- A. Drapala, D. Koszelewski, L. Tomasova, R. Ostaszewski, M. Grman, K. Ondrias and M. Ufnal, Acta Biochim. Pol., 2017, 64, 561–566 CrossRef CAS PubMed.
- T. D. Newton and M. D. Pluth, Chem. Sci., 2019, 10, 10723–10727 RSC.
- F. Augsburger and C. Szabo, Pharmacol. Res., 2020, 154, 104083 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available: Experimental details, photophysical data and fluorescence imaging figures. CCDC 1968528 and 1973453. For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/d0sc01518k |
| ‡ These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2020 |