Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Protein encapsulation: a new approach for improving the capability of small-molecule fluorogenic probes†
Hai-Hao
Han‡
ac,
Adam C.
Sedgwick‡
 bd,
Ying
Shang‡
a,
Na
Li
e,
Tingting
Liu
c,
Bo-Han
Li
c,
Kunqian
Yu
c,
Yi
Zang
c,
James T.
Brewster
II
bd,
Ying
Shang‡
a,
Na
Li
e,
Tingting
Liu
c,
Bo-Han
Li
c,
Kunqian
Yu
c,
Yi
Zang
c,
James T.
Brewster
II
 d,
Maria L.
Odyniec
d,
Maria L.
Odyniec
 b,
Maria
Weber
b,
Maria
Weber
 b,
Steven D.
Bull
b,
Steven D.
Bull
 b,
Jia
Li
*c,
Jonathan L.
Sessler
b,
Jia
Li
*c,
Jonathan L.
Sessler
 *df,
Tony D.
James
*df,
Tony D.
James
 *b,
Xiao-Peng
He
*a and
He
Tian
a
*b,
Xiao-Peng
He
*a and
He
Tian
a
aKey Laboratory for Advanced Materials, Joint International Research Laboratory of Precision Chemistry and Molecular Engineering, Feringa Nobel Prize Scientist Joint Research Center, School of Chemistry and Molecular Engineering, East China University of Science and Technology, 130 Meilong Road, Shanghai 200237, P. R. China. E-mail: xphe@ecust.edu.cn
bDepartment of Chemistry, University of Bath, Bath, BA2 7AY, UK. E-mail: chstdj@bath.ac.uk
cNational Center for Drug Screening, State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, 189 Guo Shoujing Rd., Shanghai 201203, P. R. China. E-mail: jli@ecust.edu.cn
dDepartment of Chemistry, University of Texas at Austin, 105 E 24th Street A5300, Austin, TX 78712-1224, USA. E-mail: sessler@cm.utexas.edu
eNational Facility for Protein Science in Shanghai, Zhangjiang Laboratory, Shanghai 201210, P. R. China
fCenter for Supramolecular Chemistry and Catalysis, Department of Chemistry, Shanghai University, 99 Shang-Da Road, Shanghai 200444, P. R. China
First published on 27th November 2019
Abstract
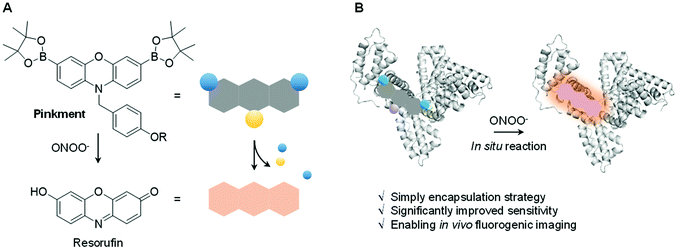
Herein, we report a protein-based hybridization strategy that exploits the host-guest chemistry of HSA (human serum albumin) to solubilize the otherwise cell impermeable ONOO− fluorescent probe Pinkment-OAc. Formation of a HSA/Pinkment-OAc supramolecular hybrid was confirmed by SAXS and solution-state analyses. This HSA/Pinkment-OAc hybrid provided an enhanced fluorescence response towards ONOO−versusPinkment-OAc alone, as determined by in vitro experiments. The HSA/Pinkment-OAc hybrid was also evaluated in RAW 264.7 macrophages and HeLa cancer cell lines, which displayed an enhanced cell permeability enabling the detection of SIN-1 and LPS generated ONOO− and the in vivo imaging of acute inflammation in LPS-treated mice. A remarkable 5.6 fold (RAW 264.7), 8.7-fold (HeLa) and 2.7-fold increased response was seen relative to Pinkment-OAc alone at the cellular level and in vivo, respectively. We anticipate that HSA/fluorescent probe hybrids will soon become ubiquitous and routinely applied to overcome solubility issues associated with hydrophobic fluorescent imaging agents designed to detect disease related biomarkers.
Introduction
Biochemical processes involve complex interactions between cellular components. Signalling molecules such as reactive oxygen species and reactive nitrogen species (ROS/RNS) play a crucial role in mediating these interactions and are required to maintain proper cell function. ROS/RNS consist of oxidative oxygen and nitrogen-based ions or free radicals.1,2 The overexpression of ROS/RNS has been implicated in an array of diseases, including cancer, inflammatory diseases, and CNS neurodegeneration.3,4 The widespread interest in biochemical processes where ROS/RNS play key roles, as well as deviations from homeostasis associated with various disease states, has provided an inspiration to develop probes that allow for the facile detection of these species. Fluorescence-based probes have received considerable attention in recent years in the context of ROS/RNS sensing. Ground-up synthetic methods have, for instance, allowed for the incorporation of reactive functionalities within the fluorophore scaffolds used to create such probes.5–9 These modifications have conferred utility for the detection and visualization of aberrant ROS/RNS levels considered causal in certain diseases.6–8 Despite significant progress, a number of issues remain to be addressed in the area of ROS/RNS probe design. In particular, standard fluorescent probes are often hydrophobic leading to poor cell permeability and aggregation induced quenching (ACQ).10 To overcome these limitations, a number of supramolecular-based strategies have been developed.11–17 Complementary to these efforts, researchers have utilized protein-based hybridization strategies (host–guest assembly) as a means to overcome the poor solubility and enhance the imaging capability of a contrast agent used in bioimaging applications.18,19,39 We thus envisaged that judicious design of an appropriate fluorescent probe–protein complex might permit enhanced sensitivity towards the detection of a disease-specific biomarker. Herein, we present the first demonstration of a fluorescent probe–protein complex strategy that results in the enhanced in vitro and in vivo detection of a key ROS/RNS signaling agent, in this case peroxynitrite (ONOO−) – Scheme 1.ONOO− is a RNS that is formed via the diffusion-controlled reaction between superoxide (O2˙−) and nitric oxide (NO˙). ONOO− is characterized by its deleterious properties, causing irreversible damage to a range of biological targets.40 Recent efforts have focused on the development of fluorescent probes for the detection of ONOO−.20–24 Towards this end, boronates have proven useful, allowing for a “turn-on” mechanism when incorporated within the sensing motif. The inherent reactivity of these substituents confers selectivity towards ONOO− over other ROS signaling molecules, such as hypochlorous acid (HOCl/ClO−) and hydrogen peroxide (H2O2).25 Recently, we developed a series ‘Pinkment’ fluorescent probes based on resorufin.26 We designed those ‘Pinkment’ probes to display ‘AND’-logic in the presence of two different biologically important species.26 In particular the Pinkment-OAc probe was designed to be activated by an esterase ‘AND’ H2O2 but the probe exhibited poor solubility and low cell permeability limiting its biological applications. Our aim with this research was to develop a general strategy to improve the sensitivity, solubility and uptake capabilities of a poorly performing fluorescent probe without changing its structure. Since, Pinkment-OAc could be selectively activated by ONOO− (via oxidation of the boronate and cleavage of the acetate ester – Scheme S1†), it represented a good example of an underperforming probe, suitable for development as an ONOO− selective fluorescent probe–protein hybrid with enhanced biological applicability.
Therefore, Pinkment-OAc was synthesized using the previously reported procedure.26 Incorporation of Pinkment-OAc within HSA to give HSA/Pinkment-OAc was then achieved via sonication of a simple mixture of the probe and HSA in water. In addition, Pinkment-OH was synthesized as previously reported26 and used to form a HSA/Pinkment-OH hybrid to illustrate the generality of our encapsulation strategy.
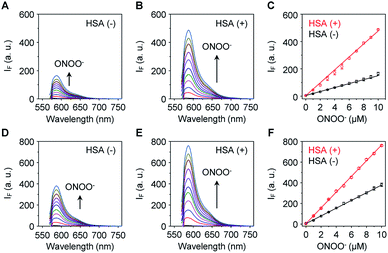
The sensitivity of these two probes towards ONOO− was then determined (Fig. 1). Unsurprisingly, Pinkment-OAc (Fig. 1A) and Pinkment-OH (Fig. 1D) gave rise to a dose dependent increase in fluorescence intensity when exposed to ONOO− in PBS (phosphate buffered saline, pH 7.4) solution. The effect, however, is modest. In contrast, a much stronger response was seen for both HSA/Pinkment-OAc (Fig. 1B) and HSA/Pinkment-OH (Fig. 1E). In the presence of ONOO− (10 μM), a 3-fold enhancement in fluorescence intensity was observed for HSA/Pinkment-OAc (Fig. 1C) whereas HSA/Pinkment-OH afforded a 2-fold increase in fluorescence intensity (Fig. 1F) when compared to each probe alone. This was taken as an initial indication of the strong affinity of HSA for each Pinkment probe. In support of this, the binding constant (Ka) between Pinkment-OAc and HSA was measured through isothermal titration calorimetry with HSA (Fig. S1†). As expected, strong binding between Pinkment-OAc and HSA was observed with a binding constant of Ka = 2.83 × 105 L mol−1. The quantum yield (ΦF; relative to quinine sulfate (probe w/o ONOO−) and rhodamine B (probe w/ ONOO−)) and extinction coefficient (ε) of HSA/Pinkment-OAc before and after addition of ONOO− were determined (Table S1†). Subsequently, the pH-dependence of Pinkment-OAc and HSA/Pinkment-OAc to detect ONOO− was evaluated and compared. Remarkably, a greater ONOO− sensitivity was observed for HSA/Pinkment-OAc when compared to Pinkment-OAc alone over a wide pH range of 3–9 (in particular from pH 5–8) (Fig. S2†).
In order to gain a greater understanding of the interactions between Pinkment-OAc and HSA, small-angle X-ray scattering (SAXS) measurements were carried out. SAXS is a powerful technique that enables conformational changes within biomacromolecules to be observed.27,28 The SAXS profile for homogeneous HSA and the HSA/Pinkment-OAc hybrid (w/w = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10) is shown in Fig. 2A and E, respectively. To explore the conformational changes of the hybrid, SREFLEX, a hybrid modeling program that systematically combines SAXS data and normal mode analysis (NMA) was used to obtain three-dimensional models of both HSA (Fig. 2B, middle) and the HSA/Pinkment-OAc hybrid (Fig. 2B, top).29 The three-dimensional SAXS model of HSA and HSA/Pinkment-OAc hybrid were then superimposed onto a resolved HSA atomic structure (PDB entry: 1n5u, Fig. 2B, bottom), respectively, using the program SUPCOMB. The modelling result obtained in this way revealed a significant conformational change of the HSA/Pinkment-OAc hybrid at the IIA region, and minimal change within the IIIA region with respect to the crystal structure of HSA (Fig. 2C). Conversely, the fitting of the SAXS model of HSA alone with its crystal structure led to good consistency for both the IIA and IIIA regions (Fig. 2G).
10) is shown in Fig. 2A and E, respectively. To explore the conformational changes of the hybrid, SREFLEX, a hybrid modeling program that systematically combines SAXS data and normal mode analysis (NMA) was used to obtain three-dimensional models of both HSA (Fig. 2B, middle) and the HSA/Pinkment-OAc hybrid (Fig. 2B, top).29 The three-dimensional SAXS model of HSA and HSA/Pinkment-OAc hybrid were then superimposed onto a resolved HSA atomic structure (PDB entry: 1n5u, Fig. 2B, bottom), respectively, using the program SUPCOMB. The modelling result obtained in this way revealed a significant conformational change of the HSA/Pinkment-OAc hybrid at the IIA region, and minimal change within the IIIA region with respect to the crystal structure of HSA (Fig. 2C). Conversely, the fitting of the SAXS model of HSA alone with its crystal structure led to good consistency for both the IIA and IIIA regions (Fig. 2G).
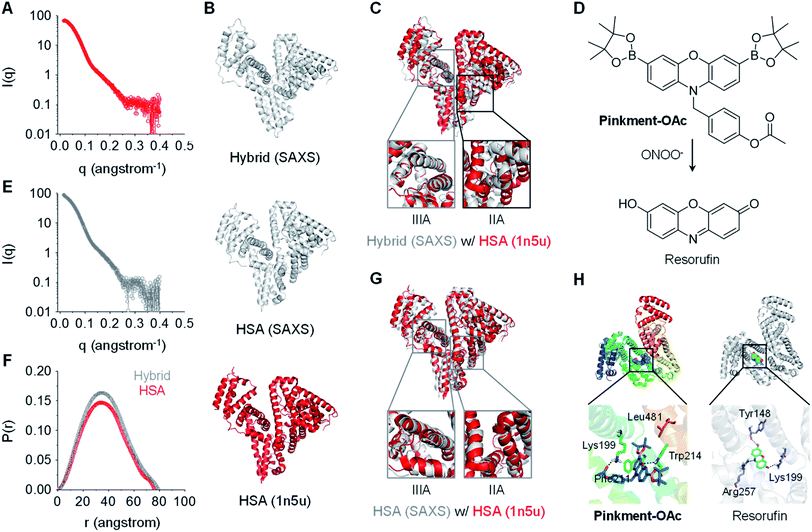 | ||
Fig. 2 X-Ray scattering patterns and molecular docking of Pinkment-OAc to HSA. (A) X-ray scattering pattern of HSA. (B) Fitting of atomic models of the hybrid (model refined with scattering data, top), HSA (model refined with scattering data, middle), and the crystal structure of HSA (PDB id 1n5u; bottom). (C) HSA crystal structure superimposed on an atomic model of HSA/Pinkment-OAc (w/w = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10) (NMA simulation; crystal structure used to fit the data: PDB id 1n5u). (D) Chemical structure of Pinkment-OAc and its reaction with ONOO−. (E) X-ray scattering pattern of HSA/Pinkment-OAc (w/w = 1 : 10). (F) Interatomic distance distribution function, P(r), of the X-ray scattering patterns of HSA and HSA/Pinkment-OAc (w/w = 1 10) (NMA simulation; crystal structure used to fit the data: PDB id 1n5u). (D) Chemical structure of Pinkment-OAc and its reaction with ONOO−. (E) X-ray scattering pattern of HSA/Pinkment-OAc (w/w = 1 : 10). (F) Interatomic distance distribution function, P(r), of the X-ray scattering patterns of HSA and HSA/Pinkment-OAc (w/w = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10). (G) HSA crystal structure superimposed with an atomic model of HSA (NMA simulation; crystal structure used to fit the data: PDB id 1n5u). (H) Proposed binding of Pinkment-OAc and resorufin to HSA showing interaction mode between Pinkment-OAc, resorufin, and selected amino acid residues of HSA. 10). (G) HSA crystal structure superimposed with an atomic model of HSA (NMA simulation; crystal structure used to fit the data: PDB id 1n5u). (H) Proposed binding of Pinkment-OAc and resorufin to HSA showing interaction mode between Pinkment-OAc, resorufin, and selected amino acid residues of HSA. | ||
We also compared the interatomic distance distribution function (P(r)) profile for both models (Fig. 2F). The results were consistent with a slightly increased maximum diameter (Dmax) for the HSA protein after inclusion of Pinkment-OAc to produce HSA/Pinkment-OAc (76.4 to 79.3 Å). The SAXS data thus provides support for the formation of a hybrid structure consisting of Pinkment-OAc and HSA. These data and the associated modeling also lead us to suggest that Pinkment-OAc is mainly bound to the IIA region of HSA. To complement the above studies, a competition assay was performed with a known IIA-region binder (phenylbutazone) and a known IIIA-region binder (ibuprofen). Satisfyingly, it was found that phenylbutazone was significantly better at competitively interrupting the association between Pinkment-OAc and HSA compared to ibuprofen (Fig. S3†). This was taken as further evidence that inclusion of Pinkment-OAc within HSA occurs predominantly at the IIA region of the protein.
Molecular docking studies were then carried out with the goal of determining how Pinkment-OAc is bound within the IIA region of HSA. On the basis of these studies, we propose that Pinkment-OAc enters the hydrophobic cavity of the HSA subdomain IIA and is stabilized by a hydrophobic interaction with the amino acid residue, Leu481 (Fig. 2H, left). Meanwhile, the carbonyl oxygen (OAc) and the oxygen on the phenoxazine unit are hydrogen bonded to Lys199 and Trp214, respectively. In addition, the benzene ring of Pinkment-OAc is involved in a π-cation interaction with the nitrogen atom of the polar amino acid residue Lys199 and a π–π interaction with Phe211 (Fig. 2H). Taken in concert, these inferred interactions provide a rationale for the inference that Pinkment-OAc is tightly bound to HSA at the IIA domain. The modeling studies also revealed that the polar groups of resorufin (the reaction product of Pinkment-OAc and ONOO−) interact with several polar residues in the IIA domain of HSA. These polar groups include the carbonyl oxygen, phenolic unit, and nitrogen atom of the aromatic core, which stabilize hydrogen bonding interactions with the residues Lys199, Tyr148, and Arg257, respectively (Fig. 2H, right). The resulting strong binding of resorufin within the hydrophobic pocket of HSA would account for the enhanced fluorescence seen when the HSA/Pinkment-OAc hybrid is exposed to ONOO− (Fig. 1).
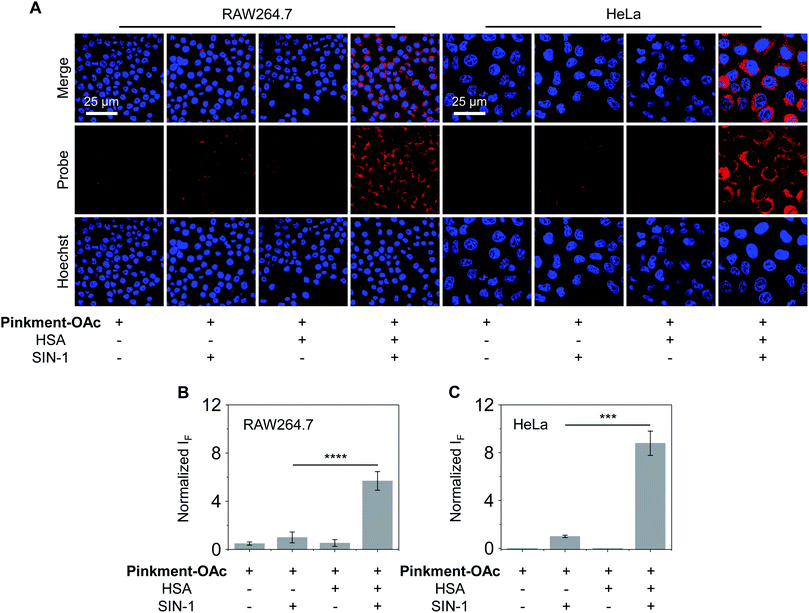
As noted above, hybridization between HSA and Pinkment-OAc leads to an enhanced fluorescence response in the presence of ONOO−. We therefore sought to test whether the HSA/Pinkment-OAc hybrid could be used to image ONOO−in vitro and in vivo. Towards this end, a macrophage cell line (RAW 264.7) and a human cervical cancer cell line (HeLa) were treated with Pinkment-OAc and the HSA/Pinkment-OAc hybrid. Each cell line was then evaluated with or without SIN-1 (an exogenous agent known to upregulate intracellular ONOO− concentrations).30 The cells were then visualized via confocal laser-scanning microscopy (CLSM) (Fig. 3). As expected, in the absence of SIN-1, no fluorescence was observed in either cell line; this was true for both free Pinkment-OAc and the HSA/Pinkment-OAc hybrid. Subsequent treatment with SIN-1 led to a remarkable fluorescence increase for HSA/Pinkment-OAc whereas a minimal fluorescence response was observed for Pinkment-OAc alone. Specifically, an overall 5.6-fold (RAW 264.7) and 8.7-fold (HeLa) difference in fluorescence intensity between HSA/Pinkment-OAc and Pinkment-OAc was observed for these two cell lines (Fig. 3B and C). In order to compare the sensitivity of HSA/Pinkment-OAc to that of Pinkment-OAc alone, we used high-content microscopy to quantify the fluorescence intensity of Pinkment-OAc and HSA/Pinkment-OAc in RAW 264.7 cells, both with and without treatment with SIN-1. As shown in Fig. S4,† a 5.4-fold increase in integrated fluorescence intensity for HSA/Pinkment-OAc was observed when treated with SIN-1 compared to Pinkment-OAc alone.
It has been reported that HSA can be actively endocytosed through clathrin-mediated pathway.31 To demonstrate that these HSA/Pinkment probes underwent a similar clathrin-mediated endocytosis, temperature-dependent fluorescence imaging experiments were carried out. In addition, fluorescence imaging experiments were carried out using a known clathrin inhibitor to inhibit clathrin-facilitated endocytosis. A decreased intracellular fluorescence response was observed, when the temperature was decreased to 4 °C (Fig. S5†), suggesting a kinetically controlled cell uptake process. Interestingly, the use of the known clathrin inhibitor chlorpromazine similarly led to a decreased fluorescence response (Fig. S6†), which suggests that HSA/Pinkment-OAc underwent a clathrin-mediated endocytosis to facilitate cellular uptake of Pinkment-OAc. Fluorescence co-localization experiments of HSA/Pinkment-OAc with a lysosome tracker (Fig. S7†) (Pearson's correlation (Rr) = 0.93), revealed that clathrin-mediated endocytosis resulted in accumulation of HSA/Pinkment-OAc in the lysosome of cells. This observation is in agreement with a previous report describing the lysosome-localization of HSA-based biomaterials.32
To demonstrate that both probe and hybrid displayed minimal toxicity towards cells, an MTS assay was performed using a RAW 264.7 cell line. As expected, little change in the cell viability was observed upon increasing concentrations of HSA/Pinkment-OAc from 5/5 μM to 80/80 μM (Fig. S8†). These findings further support our suggestion that the HSA hybridization strategy reported here may represent a general and robust approach that could be used to enhance the intracellular sensitivity of small molecule probes in vitro without triggering an adverse cellular toxicity response.
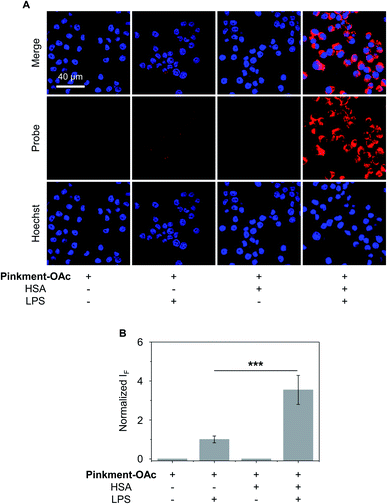
The over-production of intracellular ONOO− is believed to contribute to the development of a wide variety of inflammatory diseases.33–35 Therefore, we turned our attention to testing whether HSA/Pinkment-OAc and Pinkment-OAc could be used to detect endogenous ONOO−in vitro and in vivo. To achieve the imaging of endogenous ONOO−in vitro, RAW 264.7 cells were treated with lipopolysaccharide (LPS), which is known to induce an inflammatory response with acute upregulation of ONOO−.36 As shown in Fig. 4, a larger fluorescence increase was observed for HSA/Pinkment-OAc compared to Pinkment-OAc alone (Fig. 4). This again suggests the beneficial effect of the HSA encapsulation strategy for an increased cellular uptake of the probe and enhanced fluorescence imaging of endogenously produced ONOO−.
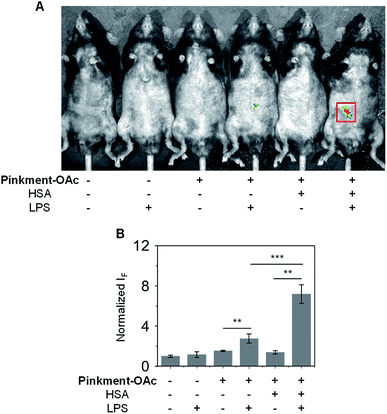
Subsequently, an in vivo experiment was carried out by treating C57BL/6J mice with LPS. In this way, a mouse model for acute inflammation was established according to the literature.37,38 The mice were divided into two groups; one was administered with pure saline intraperitoneally (i.p.) as a control, while the other was injected i.p. with LPS (200 μL, 2 mg mL−1) to induce acute inflammation. After 4 h, the mice were anaesthetized, and their abdominal fur was removed. Next, the two groups of mice were treated (via i.p. injection) with DMSO (control), Pinkment-OAc, or HSA/Pinkment-OAc. Images were acquired using the IVIS spectrum imaging system. In the case of the LPS-treated and saline-treated mice, no fluorescence was observed when injected with DMSO (Fig. 5). However, the LPS-treated mice that were injected with HSA/Pinkment-OAc produced an approximately 2.7-fold greater increase in fluorescence signal when compared to Pinkment-OAc alone (Fig. 5A and B). These in vivo experiments provide support for the contention that the present HSA/Pinkment-OAc hybrid approach allows ONOO− production to be monitored effectively in vivo and with greater sensitivity than provided by the constituent probe (Pinkment-OAc).
Conclusions
In this work, we have demonstrated the viability of a simple protein-based hybridization strategy, that involves incorporation of an insoluble ONOO− fluorescent probe (Pinkment-OAc) with HSA. The resulting HSA/Pinkment-OAc hybrid was characterized using SAXS analyses in conjunction with molecular docking studies. In solution, the HSA/Pinkment-OAc hybrid produced an enhanced fluorescence response towards ONOO− compared to the Pinkment-OAc probe alone. More importantly, the HSA/Pinkment-OAc hybrid permitted improved ONOO− detection in RAW 264.7 and HeLa cells relative to the free Pinkment-OAc probe. This enhancement is ascribed to improved cell permeability. In addition, our hybrid strategy was effective for the in vivo imaging of acute inflammation in LPS-treated mice, where again an improved performance was seen compared to the probe alone. On the basis of the present findings, we suggest that this approach represents a simple and general strategy to overcome the issues of low solubility and poor cell permeability that currently limit the use of hydrophobic imaging agents in vivo.Conflicts of interest
There are no conflicts to declare.Acknowledgements
All animal experiments were carried out under the Guidelines for Care and Use of Laboratory Animals of Shanghai Institute of Materia Medica (SIMM), Chinese Academy of Sciences and approved by the Institutional Animal Care and Use Committee (IACUC) of SIMM (Shanghai, China). RAW 264.7 and HeLa were obtained from ATCC (American Type Culture Collection). The authors thank the National Natural Science Foundation of China (Nos. 21788102, 91853201, 21722801, 81673489, 31871414 and U1832144), the Shanghai Municipal Science and Technology Major Project (No. 2018SHZDZX03), the International Cooperation Program of Shanghai Science and Technology Committee (No. 17520750100), the Youth Innovation Promotion Association of Chinese Academy of Sciences (No. 2017319), the Shanghai Science and Technology Committee (No. 19410712600) and the Fundamental Research Funds for the Central Universities (222201717003) for financial support. TDJ wishes to thank the Royal Society for a Wolfson Research Merit Award. The work in Austin was supported by the National Institutes of Health (RO1 GM103790) and the Robert A. Welch Foundation (F-0018). We would like to thank the EPSRC and the University of Bath for funding for ACS and MLO for studentships. MW would like to thank the EPSRC for funding (i) EP/L016354/1 and CDT in Sustainable Chemical Technologies.Notes and references
- M. Valko, D. Leibfritz, J. Moncol, M. T. D. Cronin, M. Mazur and J. Telser, Int. J. Biochem. Cell Biol., 2007, 39, 44–84 CrossRef CAS PubMed.
- B. Halliwell, Annu. Rev. Nutr., 1996, 16, 33–50 CrossRef CAS.
- M. Valko, C. J. Rhodes, J. Moncol, M. Izakovic and M. Mazur, Chem.-Biol. Interact., 2006, 160, 1–40 CrossRef CAS.
- J. E. Klaunig and L. M. Kamendulis, Annu. Rev. Pharmacol. Toxicol., 2004, 44, 239–267 CrossRef CAS.
- A. Romieu, Org. Biomol. Chem., 2015, 13, 1294–1306 RSC.
- A. C. Sedgwick, L. L. Wu, H. H. Han, S. D. Bull, X. P. He, T. D. James, J. L. Sessler, B. Z. Tang, H. Tian and J. Yoon, Chem. Soc. Rev., 2018, 47, 8842–8880 RSC.
- X. Q. Chen, X. Z. Tian, I. Shin and J. Yoon, Chem. Soc. Rev., 2011, 40, 4783–4804 RSC.
- D. Wu, A. C. Sedgwick, T. Gunnlaugsson, E. U. Akkaya, J. Yoon and T. D. James, Chem. Soc. Rev., 2017, 46, 7105–7123 RSC.
- J. L. Kolanowski, F. Liu and E. J. New, Chem. Soc. Rev., 2018, 47, 195–208 RSC.
- H. Nie, K. Hu, Y. J. Cai, Q. Peng, Z. J. Zhao, R. R. Hu, J. W. Chen, S. J. Su, A. J. Qin and B. Z. Tang, Mater. Chem. Front., 2017, 1, 1125–1129 RSC.
- H. J. Lv, X. T. Zhang, S. Wang and G. W. Xing, Analyst, 2017, 142, 603–607 RSC.
- M. H. Lee, E. J. Kim, H. Lee, H. M. Kim, M. J. Chang, S. Y. Park, K. S. Hong, J. S. Kim and J. L. Sessler, J. Am. Chem. Soc., 2016, 138, 16380–16387 CrossRef CAS.
- L. Skorpilova, S. Rimpelova, M. Jurasek, M. Budesinsky, J. Lokajova, R. Effenberg, P. Slepicka, T. Ruml, E. Kmonickova, P. B. Drasar and Z. Wimmer, Beilstein J. Org. Chem., 2017, 13, 1316–1324 CrossRef CAS.
- J. W. Zhou and H. Ritter, Polym. Chem., 2010, 1, 1552–1559 RSC.
- Z. C. Liu, K. Ren, Y. H. Chen, Y. X. Li, J. P. Xie, Y. J. Ding, L. L. Li, K. Li, W. P. Zhu, W. J. Yang and Z. H. Xu, RSC Adv., 2017, 7, 38160–38165 RSC.
- Z. C. Liu, W. P. Zhu, Y. H. Chen, Y. X. Li, Y. J. Ding, W. J. Yang and K. Li, Dalton Trans., 2015, 44, 16528–16533 RSC.
- D. R. Larson, H. Ow, H. D. Vishwasrao, A. A. Heikal, U. Wiesner and W. W. Webb, Chem. Mater., 2008, 20, 2677–2684 CrossRef CAS.
- Y. X. Fu, H. H. Han, J. J. Zhang, X. P. He, B. L. Feringa and H. Tian, J. Am. Chem. Soc., 2018, 140, 8671–8674 CrossRef CAS.
- F. F. An and X. H. Zhang, Theranostics, 2017, 7, 3667–3689 CrossRef CAS.
- A. C. Sedgwick, H. H. Han, J. E. Gardiner, S. D. Bull, X. P. He and T. D. James, Chem. Commun., 2017, 53, 12822–12825 RSC.
- L. L. Wu, H. H. Han, L. Y. Liu, J. E. Gardiner, A. C. Sedgwick, C. S. Huang, S. D. Bull, X. P. He and T. D. James, Chem. Commun., 2018, 54, 11336–11339 RSC.
- A. C. Sedgwick, W. T. Dou, J. B. Jiao, L. L. Wu, G. T. Williams, A. T. A. Jenkins, S. D. Bull, J. L. Sessler, X. P. He and T. D. James, J. Am. Chem. Soc., 2018, 140, 14267–14271 CrossRef CAS PubMed.
- A. C. Sedgwick, H. H. Han, J. E. Gardiner, S. D. Bull, X. P. He and T. D. James, Chem. Sci., 2018, 9, 3672–3676 RSC.
- A. C. Sedgwick, X. L. Sun, G. Kim, J. Yoon, S. D. Bull and T. D. James, Chem. Commun., 2016, 52, 12350–12352 RSC.
- A. Sikora, J. Zielonka, M. Lopez, J. Joseph and B. Kalyanaraman, Free Radical Biol. Med., 2009, 47, 1401–1407 CrossRef CAS PubMed.
- M. L. Odyniec, A. C. Sedgwick, A. H. Swan, M. Weber, T. M. S. Tang, J. E. Gardiner, M. Zhang, Y. B. Jiang, G. Kociok-Kohn, R. B. P. Elmes, S. D. Bull, X. P. He and T. D. James, Chem. Commun., 2018, 54, 8466–8469 RSC.
- L. Boldon, F. Laliberte and L. Liu, Nano Rev., 2015, 6 DOI:10.3402/nano.v6.25661.
- A. G. Kikhney and D. I. Svergun, FEBS Lett., 2015, 589, 2570–2577 CrossRef CAS.
- A. Panjkovich and D. I. Svergun, Phys. Chem. Chem. Phys., 2016, 18, 5707–5719 RSC.
- L. K. Cuddy, A. C. Gordon, S. A. Black, E. Jaworski, S. S. Ferguson and R. J. Rylett, J. Neurosci., 2012, 32, 5573–5584 CrossRef CAS PubMed.
- R. Yumoto, H. Nishikawa, M. Okamoto, H. Katayama, J. Nagai and M. Takano, Am. J. Physiol.: Lung Cell. Mol. Physiol., 2006, 290, L946–L955 CrossRef CAS PubMed.
- Q. Chen, C. Wang, Z. Zhan, W. He, Z. Cheng, Y. Li and Z. Liu, Biomaterials, 2014, 35, 8206–8214 CrossRef CAS PubMed.
- P. Pacher, J. S. Beckman and L. Liaudet, Physiol. Rev., 2007, 87, 315–424 CrossRef CAS PubMed.
- J. S. Beckman and W. H. Koppenol, Am. J. Physiol.: Cell Physiol., 1996, 271, C1424–C1437 CrossRef CAS PubMed.
- C. Szabo, H. Ischiropoulos and R. Radi, Nat. Rev. Drug Discovery, 2007, 6, 662–680 CrossRef CAS.
- A. Kumar, S. H. Chen, M. B. Kadiiska, J. S. Hong, J. Zielonka, B. Kalyanaraman and R. P. Mason, Free Radical Biol. Med., 2014, 73, 51–59 CrossRef CAS.
- Z. Li, S. H. Yan, C. Chen, Z. R. Geng, J. Y. Chang, C. X. Chen, B. H. Huang and Z. L. Wang, Biosens. Bioelectron., 2017, 90, 75–82 CrossRef CAS.
- D. Lee, S. Khaja, J. C. Velasquez-Castano, M. Dasari, C. Sun, J. Petros, W. R. Taylor and N. Murthy, Nat. Mater., 2007, 6, 765–769 CrossRef CAS PubMed.
- J. Miranda-Apodaca, N. Hananya, A. Velázquez-Campoy, D. Shabat and J. B. Arellano, Molecules, 2019, 24, 2422 CrossRef CAS PubMed.
- J. S. Beckman and W. H. Koppenol, Am. J. Physiol.: Cell Physiol., 1996, 271, C1424 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available: Synthetic procedures, 1H, 13C NMR, and mass spectra, UV-Vis and fluorescence spectroscopic analyses, and other materials. See DOI: 10.1039/c9sc03961a |
| ‡ These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2020 |