Open Access Article
Open Access ArticleAssessing potential inhibitors of SARS-CoV-2 main protease from available drugs using free energy perturbation simulations†
Son Tung
Ngo
 *ab,
Hung Minh
Nguyen
cd,
Le Thi
Thuy Huong
ef,
Pham Minh
Quan
*ab,
Hung Minh
Nguyen
cd,
Le Thi
Thuy Huong
ef,
Pham Minh
Quan
 ef,
Vi Khanh
Truong
ef,
Vi Khanh
Truong
 g,
Nguyen Thanh
Tung
g,
Nguyen Thanh
Tung
 h and
Van V.
Vu
h and
Van V.
Vu
 *i
*i
aLaboratory of Theoretical and Computational Biophysics, Ton Duc Thang University, Ho Chi Minh City 700000, Vietnam. E-mail: ngosontung@tdtu.edu.vn
bFaculty of Applied Sciences, Ton Duc Thang University, Ho Chi Minh City 700000, Vietnam
cCenter for Molecular Biology, Institute of Research and Development, Duy Tan University, Da Nang, 550000, Vietnam
dFaculty of Pharmacy, Duy Tan University, Da Nang, 550000, Vietnam
eInstitute of Natural Products Chemistry, Vietnam Academy of Science and Technology, Hanoi 100000, Vietnam
fGraduate University of Science and Technology, Vietnam Academy of Science and Technology, Hanoi 100000, Vietnam
gSchool of Science, RMIT University, GPO Box 2476, Melbourne 3001, Australia
hInstitute of Materials Science, Vietnam Academy of Science and Technology, Hanoi 100000, Vietnam
iNTT Hi-Tech Institute, Nguyen Tat Thanh University, Ho Chi Minh City 700000, Vietnam. E-mail: vanvu@ntt.edu.vn
First published on 9th November 2020
Abstract
The main protease (Mpro) of the novel coronavirus SARS-CoV-2, which has caused the COVID-19 pandemic, is responsible for the maturation of its key proteins. Thus, inhibiting SARS-CoV-2 Mpro could prevent SARS-CoV-2 from multiplying. Because new inhibitors require thorough validation, repurposing current drugs could help reduce the validation process. Many recent studies used molecular docking to screen large databases for potential inhibitors of SARS-CoV-2 Mpro. However, molecular docking does not consider molecular dynamics and thus can be prone to error. In this work, we developed a protocol using free energy perturbation (FEP) to assess the potential inhibitors of SARS-CoV-2 Mpro. First, we validated both molecular docking and FEP on a set of 11 inhibitors of SARS-CoV-2 Mpro with experimentally determined inhibitory data. The experimentally deduced binding free energy exhibits significantly stronger correlation with that predicted by FEP (R = 0.94 ± 0.04) than with that predicted by molecular docking (R = 0.82 ± 0.08). This result clearly shows that FEP is the most accurate method available to predict the binding affinity of SARS-CoV-2 Mpro + ligand complexes. We subsequently used FEP to validate the top 33 compounds screened with molecular docking from the ZINC15 database. Thirteen of these compounds were predicted to bind strongly to SARS-CoV-2 Mpro, most of which are currently used as drugs for various diseases in humans. Notably, delamanid, an anti-tuberculosis drug, was predicted to inhibit SARS-CoV-2 Mpro in the nanomolar range. Because both COVID-19 and tuberculosis are lung diseases, delamanid has higher probability to be suitable for treating COVID-19 than other predicted compounds. Analysis of the complexes of SARS-CoV-2 Mpro and the top inhibitors revealed the key residues involved in the binding, including the catalytic dyad His14 and Cys145, which is consistent with the structural studies reported recently.
Introduction
The novel coronavirus SARS-CoV-2, which causes the disease COVID-19, has spread all over the world, resulting in one of the most devastating pandemics since the Spanish flu pandemic in 1918. COVID-19 has disrupted almost all sectors for more than three quarters of 2020. The virus causes severe acute respiratory syndrome, but many infected patients do not show symptoms, which renders the control of the virus extremely difficult. To date, there are more than 33 million people infected and more than 1 million deaths.1 Vaccines for SARS-CoV-2 are being developed and tested in clinical trials. However, there are not yet any efficient vaccine reported to date. Moreover, some studies show that the antibodies for SARS-CoV-2 developed in the patients disappear within a couple of months,2 which would result in a lot of complications in vaccine development and usage. Thus, finding drugs for SARS-CoV-2 is of great interests.The genomes of coronaviruses are 26–32 kb long, the largest genomes among RNA viruses,3,4 and encode more than 20 proteins. Polyproteins 1a and 1ab are synthesized from the first open reading frame, which are then processed by the main protease (Mpro) and a papain-like protease to form structural and non-structural proteins.4 Mpro, also called 3CLPPro, thus plays a vital role in the replication process of coronaviruses4,5 and has been proposed as a drug target.5,6 The X-ray diffraction structure of SARS-CoV-2 Mpro has been quickly determined,7,8 which serves as the foundation for the investigation of its inhibitors. Several inhibitors of SARS-CoV-2 Mpro were recently synthesized and characterized, some of which exhibits nano-molar affinity.7–10 However, these identified Mpro inhibitors have not been clinically evaluated as drugs for COVID-19. Side-effects of some of these inhibitors have not been reported.
Alternative to synthesizing new SARS-CoV-2 inhibitors, potential drugs can be screened from various databases. Because it is not feasible to experimentally test each of the numerous available compounds, these databases need to be screened by computational methods. Several SARS-CoV-2 Mpro inhibitors screening studies reported recently mainly relied on molecular docking methods.11–16 Although docking is rapid and often used for initial screening of a large database,17 it does not consider molecular dynamics and thus does not guarantee the accurate results. Therefore, the docking results require further validations by more accurate methods. Several other studies utilized different molecular dynamics methods, such fast pulling of ligand (FPL),18,19 to validate the docking results.
Among computational methods for studying the binding affinity protein–ligand complexes, free energy perturbation (FEP) has proved to be the most accurate.20,21 However, FEP requires tremendous computational resources and cannot be applied to a large database. We recently applied FEP on several SARS-CoV-2 Mpro inhibitors identified by FPL from a database of compounds found in Vietnamese herbs and obtained some promising results.18 In this work, we used FEP to assess a larger number of potential SARS-CoV-2 inhibitors, which were screened by molecular docking from 6363 compounds in the ZINC15 database.22
Results and discussion
Validation of docking and FEP methods with characterized inhibitors
To date, there are 11 inhibitors of SARS-CoV-2 Mpro that have been experimentally characterized.7–10 The half maximal inhibitory concentration (IC50) were reported, which can be used to deduce the binding free energy (Table 1). Docking of these compounds to SARS-CoV-2 was carried out using AutoDock Vina as described in our recent study.18 The binding free energy obtained from docking simulations (ΔGDock) (Table 1) exhibits a reasonably good correlation to that deduced from the experimental IC50 value (ΔGEXP), with a coefficient of R = 0.82 ± 0.08 (Fig. 1).| No. | Name | IC50 (μM) | ΔGEXPa | ΔGDock | ΔGFEP |
|---|---|---|---|---|---|
a ΔGEXP was calculated from the IC50 values obtained experimentally7–10 using the equation ΔGExp = RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(ki), where the inhibition constant ki is assumed to be equal to the corresponding IC50 value, R is the gas constant, and the temperature T is set at 298 K. The error is the standard error of the mean.
b Previously reported.18 The unit of free energy is kcal mol−1. ln(ki), where the inhibition constant ki is assumed to be equal to the corresponding IC50 value, R is the gas constant, and the temperature T is set at 298 K. The error is the standard error of the mean.
b Previously reported.18 The unit of free energy is kcal mol−1.
|
|||||
| 1 | 11r | 0.18 ± 0.02 | −9.23 | −7.1b | −13.3 ± 2.58b |
| 2 | 13a | 2.39 ± 0.63 | −7.70 | −6.7b | −8.18 ± 2.20b |
| 3 | 13b | 0.67 ± 0.18 | −8.45 | −6.9b | −9.18 ± 2.48b |
| 4 | 11a | 0.053 ± 0.005 | −9.96 | −7.2 | −14.1 ± 0.39 |
| 5 | 11b | 0.040 ± 0.002 | −10.1 | −7.2 | −14.8 ± 0.74 |
| 6 | Carmofur | 1.82 ± 0.06 | −7.86 | −5.6 | −5.16 ± 0.51 |
| 7 | Disulfiram | 9.35 ± 0.18 | −6.89 | −3.8 | −4.30 ± 0.34 |
| 8 | Ebselen | 0.67 ± 0.09 | −8.45 | −5.6 | −6.84 ± 0.28 |
| 9 | PX-12 | 21.39 ± 7.06 | −6.39 | −3.7 | −4.52 ± 0.80 |
| 10 | Shikonin | 15.75 ± 8.22 | −6.58 | −5.4 | −3.46 ± 0.55 |
| 11 | Tideglusib | 1.55 ± 0.30 | −7.95 | −6.6 | −5.58 ± 0.26 |
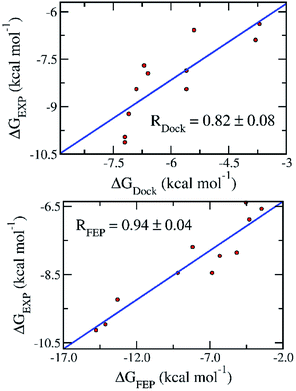 | ||
| Fig. 1 Correlation between the binding free energy deduced from experimental inhibitory data (ΔGEXP) and that obtained with docking (ΔGDock) (top) and FEP (ΔGFEP) (bottom) simulations. Experimental results were reported in recent publications.7,9,10 | ||
FEP was applied on the 11 experimentally characterized ligands. Two independent molecular dynamics (MD) trajectories were first carried out for each SARS-CoV-2 Mpro + inhibitor complex, which converged after 10 ns (Fig. S2†). The last snapshots of MD simulations were then used as the starting conformations for FEP calculation. Sixteen independent 2.0 ns MD simulations were carried out for the solvated SARS-CoV-2 Mpro + inhibitor complex systems and solvated inhibitor systems using various coupling parameter λ.23
The BAR method was used to calculate free energy every 100 ps.24 It almost converged after the first 1.0 ns interval. The data collected in the last 1.0 ns interval was used for determining the binding free energy. The difference between the de-solvation free energy of the ligand from the solvated complex and that of the solvated ligand system is the binding free energy ΔGFEP.23 The correlation coefficient between ΔGFEP and ΔGEXP is R = 0.94 ± 0.04, which is significantly higher than that between ΔGDock and ΔGEXP (Fig. 1). One thousand of bootstrapping rounds were used to calculate the error of correlation coefficient. The RMSE between ΔGFEP and ΔGEXP is 2.84 ± 0.38 kcal mol−1. The obtained results indicate the FEP calculation is the best method thus far to predict the ligand-binding free energy for SARS-CoV-2 Mpro.
Assessing potentials inhibitors of SARS-CoV-2 Mpro in the ZINC15 database
Given the high accuracy of our approach as tested with the experimentally characterized inhibitors, we continued to use it to assess the potential Mpro inhibitors in a database of available drugs, particularly the ZINC15 database that contains 6363 compounds. Docking studies show that most of the ZINC15 compounds has weak binding affinity to SARS-CoV-2 Mpro (Fig. S1†). We selected the top 33 compounds with binding affinity larger than 8.3 kcal mol−1 (Tables 2 and S1†) and subjected to FEP simulations (Fig. S2–S4†). The binding free energy derived from FEP simulations (ΔGFEP) ranges from ca. −1 kcal mol−1 to ca. −25 kcal mol−1, while ΔGDock falls in a narrow range of −8.3 to −9.0 kcal mol−1. This result clearly show the limits of molecular docking.| No. | ZINC ID | Name | ΔGDock | ΔGCou | ΔGvdW | ΔGFEP | Predicted ki range |
|---|---|---|---|---|---|---|---|
| a The error of computations is the standard error of the mean. The ki is predicted using the equation ki = e(ΔGFEP/RT), where R is the gas constant and the temperature T is set at 298 K. The unit is in kcal mol−1. The compounds in the second part of the table (entries 16–33) are not discussed here due to their weaker predicted binding affinity. | |||||||
| 1 | ZINC000169289767 | Trypan blue | −9.0 | −18.97 | −6.35 | −25.32 ± 4.6 | Sub attomolar |
| 2 | ZINC000100067477 | Alatrofloxacin | −8.5 | −15.34 | −5.82 | −21.16 ± 1.49 | Sub-femtomolar |
| 3 | ZINC000004215257 | Cefpiramide | −8.3 | −11.68 | −7.86 | −19.54 ± 0.57 | Femtomolar |
| 4 | ZINC000011616152 | Novobiocin, sodium salt | −8.3 | −8.80 | −9.18 | −17.97 ± 1.00 | High-femtomolar |
| 5 | ZINC000014880002 | Dihydroergotoxine | −8.5 | −10.05 | −6.82 | −16.87 ± 1.33 | Sub-picomolar |
| 6 | ZINC000004099104 | Sn38-glucuronide | −8.4 | −2.53 | −9.25 | −11.78 ± 2.15 | Nanomolar |
| 7 | ZINC000043100810 | Delamanid | −8.7 | −3.85 | −7.66 | −11.51 ± 0.16 | Nanomolar |
| 8 | ZINC000022058728 | Npc | −8.4 | −3.27 | −7.38 | −10.65 ± 1.37 | High-nanomolar |
| 9 | ZINC000019632618 | Imatinib | −8.3 | −4.15 | −5.71 | −9.86 ± 0.26 | High-nanomolar |
| 10 | ZINC000001612996 | Irinotecan | −8.7 | −3.83 | −5.75 | −9.58 ± 0.38 | Sub-micromolar |
| 11 | ZINC000008101127 | Indocyanine green | −8.3 | −2.78 | −6.77 | −9.55 ± 0.49 | Sub-micromolar |
| 12 | ZINC000052955754 | Ergotamine | −8.4 | −1.08 | −8.37 | −9.45 ± 1.04 | Sub-micromolar |
| 13 | ZINC000118915330 | Pregnanediol-3-glucuronide | −8.4 | 0.95 | −10.28 | −9.33 ± 0.85 | Sub-micromolar |
| 14 | ZINC000021981222 | N-Desmethyl imatinib | −8.5 | −0.60 | −8.55 | −9.15 ± 2.36 | Sub-micromolar |
| 15 | ZINC000003978005 | Dihydroergotamine | −8.7 | −4.67 | −4.13 | −8.81 ± 1.87 | Sub-micromolar |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||||
| 16 | ZINC000004099009 | Teniposide | −8.4 | 2.12 | −9.91 | −7.79 ± 1.31 | Micromolar |
| 17 | ZINC000084668739 | Lifitegrast | −8.3 | 0.53 | −8.30 | −7.77 ± 2.03 | Micromolar |
| 18 | ZINC000031425360 | 4-Hydroxyphenytoin glucuronide | −8.4 | 1.11 | −8.62 | −7.51 ± 0.81 | Micromolar |
| 19 | ZINC000001482077 | Gliquidone | −8.3 | −1.97 | −5.13 | −7.11 ± 1.00 | Micromolar |
| 20 | ZINC000198970879 | Olmutinib | −8.3 | −1.12 | −5.86 | −6.98 ± 1.31 | Micromolar |
| 21 | ZINC000000896717 | Zafirlukast | −8.3 | −1.54 | −5.04 | −6.59 ± 1.54 | High-micromolar |
| 22 | ZINC000006745272 | Regorafenib | −8.3 | 2.40 | −8.62 | −6.22 ± 0.48 | High-micromolar |
| 23 | ZINC000299818022 | N/A | −8.3 | 1.35 | −7.00 | −5.65 ± 0.18 | High-micromolar |
| 24 | ZINC000253975480 | Rifaximin | −8.4 | 8.38 | −13.67 | −5.29 ± 2.03 | Sub-millimolar |
| 25 | ZINC000064033452 | Lumacaftor | −8.3 | 3.68 | −8.68 | −5.00 ± 0.90 | Sub-millimolar |
| 26 | ZINC000118915340 | HMDB0010338 | −8.4 | 4.79 | −9.22 | −4.43 ± 1.66 | Sub-millimolar |
| 27 | ZINC000164528615 | Glecaprevir | −8.3 | 1.43 | −5.72 | −4.29 ± 8.70 | Sub-millimolar |
| 28 | ZINC000035051264 | N/A | −8.4 | 1.66 | −5.61 | −3.95 ± 0.82 | Sub-millimolar |
| 29 | ZINC000013515299 | 17-Beta-estradiol glucuronide | −8.5 | 5.18 | −7.02 | −1.84 ± 1.86 | High-millimolar |
| 30 | ZINC000003831231 | CHEMBL35025 | −8.3 | 4.82 | −6.56 | −1.73 ± 0.92 | High-millimolar |
| 31 | ZINC000006716957 | Nilotinib | −8.3 | 6.49 | −7.85 | −1.35 ± 0.97 | Sub-molar |
| 32 | ZINC000040165255 | Estriol-17-glucuronide | −8.3 | 4.74 | −6.06 | −1.32 ± 0.28 | Sub-molar |
| 33 | ZINC000096015174 | Glycyrrhizic acid | −8.3 | 5.73 | −6.88 | −1.15 ± 1.5 | Sub-molar |
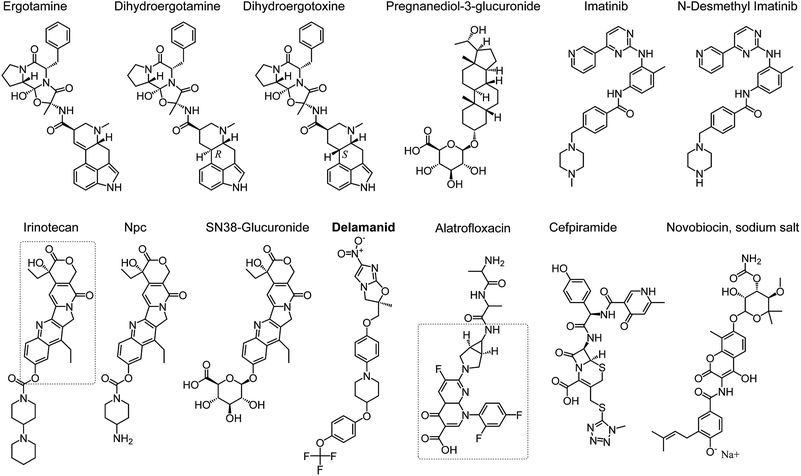
Fifteen compounds were identified by FEP to have binding affinity larger than −8.0 kcal mol−1, which would correspond to an inhibition constant ki in the sub-micromolar range or smaller (Table 2 and Fig. 2). Two of these compounds, no. 1 and 11 in Table 2, are common biological staining dyes that are known to bind to cell surface protein,25,26 which might not be suitable to use as drugs. The structures of the other 13 compounds are shown in Fig. 2.
Ergotamine27 and its derivatives dihydroergotamine28 and dihydroergotoxine29 are drugs for treating migraine. They narrow blood vessel and help modulate blood flow pattern. The binding affinity to SARS-CoV-2 Mpro of dihydroergotamine and ergotamine is in the sub-micromolar range, which increases substantially to the sub-picomolar range. Dihyroergotoxine only differs from dihydroergotamine in the 3-D configuration of one carbon chiral center (Fig. 2), which leads to substantial difference in their conformations in their complexes with SARS-CoV-2 Mpro. Dihydroergotoxine forms multiple H-bonds to SARS-CoV-2 Mpro, while dihydroergotamine only forms H-bonds with an atom of Mpro. This is an important insight of SARS-CoV-2 Mpro-ligand binding, which can serve as the basis for further design of new inhibitors. It is worth noting that dihydroergotoxine exhibits an extraordinary binding affinity for SARS-CoV-2 Mpro, which is more than 3 orders of magnitude higher than the best experimentally characterized inhibitors of this protease (Table 1).
Beside ergotamine and its two derivatives, seven other compounds screened herein exhibit binding affinity for SARS-CoV-2 Mpro ranging from sub-micromolar to nanomolar levels (Table 2). Pregnanediol-3-glucuronide is a metabolite of progesterone predominant in urine during pregnancy.30 Other compounds in this group have been used as drugs. Imatinib31 and its metabolite N-desmethyl-imatinib,32 which inhibit BCR/AbI complex and leucine-rich S/T kinase, are currently used as oral chemotherapy medications for cancer. Irinotecan is also a cancer drug, used to treat metastatic colon and rectal cancer.33 Npc and Sn38-glucuronide are compounds in the irinotecan metabolism pathway, which also contain the Sn38 as in irinotecan (indicated with a dashed rectangle in the structure of irinotecan in Fig. 2). It appears that the Sn38 moiety contribute the most to the high binding affinity of irinotecan, Npc, and Sn38-glucuronide to SARS-CoV-2 Mpro.
Notably, among the inhibitors with nanomolar affinity is delamanid, an anti-tuberculosis agent.34 Because both tuberculosis (TB) and SARS-CoV-2 infect the respiratory systems, a drug against TB that has strong affinity for Mpro could be more suitable to be used against SARS-CoV-2 than other drugs. The predicted nanomolar binding affinity for SARS-CoV-2 Mpro of delamanid is comparable to that of the best characterized inhibitors (Table 1). Thus, delamanid is one of the most promising new drug candidates for COVID-19.
Three antibiotics were predicted by FEP to have remarkably high binding affinity for SARS-CoV-2 Mpro in the femto-molar scale. Alatrofloxacin (no. 2 in Table 2) is a product of Pfizer, which is rapidly converted to trovafloxacin in plasma.35 Thus, although alatrofloxacin might have extraordinary binding affinity to SARS-CoV-2 Mpro, it might not have activity against SARS-CoV-2 in vivo. Instead, its metabolite trovafloxacin should be considered in further studies. Novobiocin is a coumarin-based antibiotic that can be used orally and can be used to treat methicillin-resistant Staphylococcus aureus MRSA.36 Cefpiramide is a potent wide-spectrum antibiotics.37 Both novobiocin and cefpiramide could be considered for further studies on SARS-CoV-2 inhibition.
Interactions of the inhibitors with SARS-CoV-2 Mpro
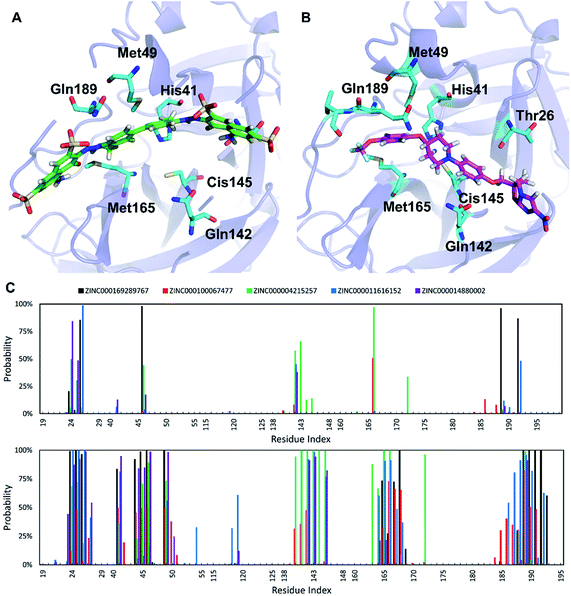
The ligands investigated herein bind to the same binding cleft on the surface of SARS-CoV-2 Mpro (Fig. 3), consistent with the structurally characterized complex reported recently.7 The conformation of the binding cleft changes sizably between the complexes, indicating the flexibility of this cleft and the ability to accommodate a wide range of ligands. The strongest binding ligand identified here, trypan blue, spans the entire cleft. Delamanid binds in a similar pose, but it is shorter than trypan blue and has significantly smaller binding affinity (Table 2). Analysis of the interactions between the top five ligands and SARS-CoV-2 Mpro reveals key residues contributing to ligand binding. The main interactions are side chain contact involving the residues around the positions T24–T25–T26, H41, C44–T45–S46, M49, N142, C145, H164–M165–E166–L167–P168, and V186–D187–R188–Q189–T190 (Fig. 3). Some of these residues also form hydrogen bonds to the inhibitors. The Mpro catalytic dyad His41 and Cys145 are among the residues forming the most interaction with the substrate, which is consistent with previous structural studies.8,10Experimental
Input structures
The X-ray diffraction structure of SARS-CoV-2 main protease was downloaded from Protein Data Bank (ID 6Y2F).7 The ligand structures were downloaded from ZINC15 and PubChem database.22,38Molecular docking simulations
The binding position and affinity of ligands to SARS-CoV-2 Mpro were predicted using the Autodock Vina package,39 and parameters described in our recent studies.17,18 The exhaustiveness was set at 8. The optimally docked conformation was chosen with the largest binding affinity. The grid center was set at the center of mass of the compound 13b.7 The grid size was set as 26 × 26 × 26 Å.Molecular dynamics simulations
The SARS-CoV-2 Mpro + ligand complexes were simulated using GROMACS 5.1.5.40 SARS-CoV-2 Mpro was topologized in the Amber99SB-ILDN force field.41 AmberTools18 and ACPYPE were used to paramterize the ligands in the general Amber force field (GAFF).42,43 The Restrained Electrostatic Potential (RESP) method44 and quantum chemical calculation (B3LYP functional at 6-31G(d,p) level of theory inimplicit water model with ε = 78.4) were carried out to assign the atomic charge of the ligands. The parameterized SARS-CoV-2 Mpro + inhibitor complex was placed in a 820 nm3 dodecahedron periodic boundary (PBC) condition box. The solvated complex consists of 1 SARS-CoV-2 Mpro molecule, 1 inhibitor molecule, 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 water molecules, and an appropriate number of Na+ ions to neutralize the system. In total, each solvated Mpro + inhibitor system contains more than 80
300 water molecules, and an appropriate number of Na+ ions to neutralize the system. In total, each solvated Mpro + inhibitor system contains more than 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 atoms. Likewise, the inhibitor was also put into a ca. 85 nm3 dodecahedron PBC box. The solvated inhibitor system consists of 1 inhibitor molecule, 2700 water molecules, and an appropriate number of Na+ or Cl− ions to neutralize the system. In total, each solvated ligand system contained ca. 8300 atoms.
000 atoms. Likewise, the inhibitor was also put into a ca. 85 nm3 dodecahedron PBC box. The solvated inhibitor system consists of 1 inhibitor molecule, 2700 water molecules, and an appropriate number of Na+ or Cl− ions to neutralize the system. In total, each solvated ligand system contained ca. 8300 atoms.
MD simulations were carried out using the parameters described in previous works.23 The MD time step was set at 2 fs. The nonbonded interaction cutoff was set at 0.9 nm. The fast particle-mesh Ewald electrostatics scheme was used to calculate the Coulomb interactions.45 The solvated system was then subjected to EM, 0.1 ns NVT, and 0.1 ns NPT simulations. A small harmonic force (1000 kJ mol−1 nm−2 per dimensions) was used to restrain the Cα atoms of SARS-CoV-2 Mpro. The relaxed SARS-CoV-2 Mpro + inhibitor system was then subjected to unrestrained MD simulations. The MD simulations were carried out for 20 ns for each solvated complex system and 5 ns for each solvated inhibitor system. During simulations, the systemic coordinates were logged every 10 ps.
Free energy perturbation (FEP) simulations
The last snapshots of MD simulations were subjected to FEP simulations.23,46 Details of FEP calculations were described in our previous study.23 Free energy calculations involve two inhibitor annihilation processes.47 The coupling parameter λ, ranging from 0 to 1, was utilized to annihilate the inhibitor from the solvated systems. The free energy change from the full-interaction state (λ = 0) to the non-interaction state (λ = 1) during λ-alteration simulations was then calculated by using the Bennett's acceptance ratio (BAR) method.24 The difference of free energy changes over two annihilation processes is the binding free energy ΔGFEP of an inhibitor to SARS-CoV-2 Mpro. Nine values of λ, including 0.00, 0.10, 0.25, 0.35, 0.50, 0.65, 0.75, 0.90, and 1.00, were used to change the electrostatic potential. The van der Waals (vdW) interaction was altered via eight values of λ, including 0.00, 0.10, 0.25, 0.35, 0.50, 0.65, 0.75, 0.90, and 1.00. Each λ-alteration simulation was performed for 2 ns. The FEP calculation for a complex was repeated two times in order to guarantee the simulation sampling.Structural analysis
Chemicalize webserver (ChemAxon) was used to predict the protonation states of the inhibitors. The protein surface charge was computed using Adaptive Poisson–Boltzmann Solver (APBS) package.48 The binding pose of the SARS-CoV-2 Mpro + ligand complexes was evaluated using PyMOL package.49 The hydrogen bonds (HB) were calculated an acceptor (A) and a donor (D) is within 0.35 nm from one another and the angle ∠A–H–D is greater than 135°. The side chain (SC) contact was recorded when the distance between a non-hydrogen atom of a side chain of SARS-CoV-2 Mpro and that of the ligand was smaller than 0.45 nm.Conclusions
We have carried out validation of FEP and docking on 11 experimentally characterized inhibitors of SARS-CoV-2 Mpro. The binding free energy of the SARS-CoV-2 Mpro + ligand complexes was deduced from experimental IC50 value (ΔGExp), molecular docking (ΔGDock), and FEP (ΔGFEP). ΔGExp strongly correlates to ΔGFEP (R = 0.94 ± 0.04), while it only moderately correlates to ΔGDock (R = 0.82 ± 0.08). This result indicates that FEP is the most appropriate method to predict the inhibitors of SARS-CoV-2 Mpro. We subsequently used FEP to assess the top 33 compounds identified by docking from 6363 compounds in the ZINC15 database. While ΔGDock of these compounds falls in a narrow range from −8.3 kcal mol−1 to −8.5 kcal mol−1, ΔGFEP varies substantially from ca. −1 kcal mol−1 to −25 kcal mol−1. This result further shows the limitations of molecular docking in predicting binding free energy. Thirteen of the top 33 compounds exhibits strong affinity for SARS-CoV-2 Mpro, which falls in the sub-micromolar range. It is worth emphasizing that delamanid is the most promising candidates because it has nanomolar binding affinity and is a drug for tuberculosis, a respiratory disease. The binding cleft of SARS-CoV-2 Mpro is flexible and able to bind a large variety of ligands. The key residues involved in ligand binding includes the catalytic dyad His41 and Cys145.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work was supported by Vietnam National Foundation for Science & Technology Development (NAFOSTED) grant #104.99-2019.57.Notes and references
- https://coronavirus.jhu.edu/data, accessed October 3, 2020.
- J. Seow, C. Graham, B. Merrick, S. Acors, K. J. A. Steel, O. Hemmings, A. Bryne, N. Kouphou, S. Pickering, R. Galao, G. Betancor, H. D. Wilson, A. W. Signell, H. Winstone, C. Kerridge, N. Temperton, L. Snell, K. Bisnauthsing, A. Moore, A. Green, L. Martinez, B. Stokes, J. Honey, A. Izquierdo-Barras, G. Arbane, A. Patel, L. Oconnell, G. O Hara, E. MacMahon, S. Douthwaite, G. Nebbia, R. Batra, R. Martinez-Nunez, J. D. Edgeworth, S. J. D. Neil, M. H. Malim and K. Doores, 2020, medRxiv, DOI:10.1101/2020.07.09.20148429.
- D. Schoeman and B. C. Fielding, Virology, 2019, 16, 69 CrossRef PubMed
.
- C. M. Fauquet and D. Fargette, Virology, 2005, 2, 64 CrossRef CAS PubMed
.
- A. Zhavoronkov, B. Zagribelnyy and A. Zhebrak, Potential Non-Covalent SARS-CoV-2 3C-like Protease Inhibitors Designed Using Generative Deep Learning Approaches and Reviewed by Human Medicinal Chemist in Virtual Reality, ChemRxiv, 2020 DOI:10.26434/chemrxiv.12301457.v1
.
- V. Nukoolkarn, V. S. Lee, M. Malaisree, O. Aruksakulwong and S. Hannongbua, J. Theor. Biol., 2008, 254, 861–867 CrossRef CAS PubMed
.
- L. Zhang, D. Lin, X. Sun, U. Curth, C. Drosten, L. Sauerhering, S. Becker, K. Rox and R. Hilgenfeld, Science, 2020, 368, 409–412 CrossRef CAS PubMed
.
- Z. Jin, X. Du, Y. Xu, Y. Deng, M. Liu, Y. Zhao, B. Zhang, X. Li, L. Zhang, C. Peng, Y. Duan, J. Yu, L. Wang, K. Yang, F. Liu, R. Jiang, X. Yang, T. You, X. Liu, X. Yang, F. Bai, H. Liu, X. Liu, L. W. Guddat, W. Xu, G. Xiao, C. Qin, Z. Shi, H. Jiang, Z. Rao and H. Yang, Nature, 2020, 582, 289–293 CrossRef CAS PubMed
.
- Z. Jin, Y. Zhao, Y. Sun, B. Zhang, H. Wang, Y. Wu, Y. Zhu, C. Zhu, T. Hu, X. Du, Y. Duan, J. Yu, X. Yang, X. Yang, K. Yang, X. Liu, L. W. Guddat, G. Xiao, L. Zhang, H. Yang and Z. Rao, Nat. Struct. Mol. Biol., 2020, 27, 529–532 CrossRef CAS PubMed
.
- W. Dai, B. Zhang, H. Su, J. Li, Y. Zhao, X. Xie, Z. Jin, F. Liu, C. Li, Y. Li, F. Bai, H. Wang, X. Cheng, X. Cen, S. Hu, X. Yang, J. Wang, X. Liu, G. Xiao, H. Jiang, Z. Rao, L.-K. Zhang, Y. Xu, H. Yang and H. Liu, Science, 2020, eabb4489, DOI:10.1126/science.abb4489
.
- R. Alexpandi, J. F. De Mesquita, S. K. Pandian and A. V. Ravi, Front. Microbiol., 2020, 1 DOI:10.3389/fmicb.2020.01796
.
- S. Shahinshavali, K. A. Hossain, A. V. D. N. Kumar, A. G. Reddy, D. Kolli, A. Nakhi, M. V. B. Rao and M. Pal, Tetrahedron Lett., 2020, 61, 152336 CrossRef CAS PubMed
.
- K. F. Azim, S. R. Ahmed, A. Banik, M. M. R. Khan, A. Deb and S. R. Somana, Informatics in Medicine Unlocked, 2020, 20, 100367 CrossRef PubMed
.
- W. R. Ferraz, R. A. Gomes, A. L. S. Novaes and G. H. G. Trossini, Future Med. Chem., 2020, 12, 1815–1828 CrossRef CAS PubMed
.
- O. O. Olubiyi, M. Olagunju, M. Keutmann, J. Loschwitz and B. Strodel, Molecules, 2020, 25, 3193 CrossRef CAS PubMed
.
- E. M. Marinho, J. Batista de Andrade Neto, J. Silva, C. Rocha da Silva, B. C. Cavalcanti, E. S. Marinho and H. V. Nobre Júnior, Microb. Pathog., 2020, 148, 104365 CrossRef CAS PubMed
.
- N. T. Nguyen, T. H. Nguyen, T. N. H. Pham, N. T. Huy, M. V. Bay, M. Q. Pham, P. C. Nam, V. V. Vu and S. T. Ngo, J. Chem. Inf. Model., 2020, 60, 204–211 CrossRef CAS PubMed
.
- S. T. Ngo, N. Quynh Anh Pham, L. Thi Le, D.-H. Pham and V. V. Vu, J. Chem. Inf. Model., 2020 DOI:10.1021/acs.jcim.0c00491
.
- M. Q. Pham, K. B. Vu, T. N. Han Pham, L. T. Thuy Huong, L. H. Tran, N. T. Tung, V. V. Vu, T. H. Nguyen and S. T. Ngo, RSC Adv., 2020, 10, 31991–31996 RSC
.
- R. W. Zwanzig, J. Chem. Phys., 1954, 22, 1420–1426 CrossRef CAS
.
- D. L. Beveridge and F. M. DiCapua, Annu. Rev. Biophys. Biophys. Chem., 1989, 18, 431–492 CrossRef CAS PubMed
.
- T. Sterling and J. J. Irwin, J. Chem. Inf. Model., 2015, 55, 2324–2337 CrossRef CAS PubMed
.
- S. T. Ngo, T. H. Nguyen, N. T. Tung, P. C. Nam, K. B. Vu and V. V. Vu, J. Comput. Chem., 2020, 41, 611–618 CrossRef CAS PubMed
.
- C. H. Bennett, J. Comput. Phys., 1976, 22, 245–268 CrossRef
.
- https://pubchem.ncbi.nlm.nih.gov/compound/Trypan-blue, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/indocyanine%20green, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Ergotamine, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Dihydroergotamine, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Dihydroergotoxine, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Pregnanediol-3-glucuronide, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Imatinib, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/N-desmethyl-imatinib, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Irinotecan, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Delamanid, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Alatrofloxacin, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Novobiocin, accessed July 25, 2020.
- https://pubchem.ncbi.nlm.nih.gov/compound/Cefpiramide, accessed July 25, 2020.
- S. Kim, P. A. Thiessen, E. E. Bolton, J. Chen, G. Fu, A. Gindulyte, L. Han, J. He, S. He, B. A. Shoemaker, J. Wang, B. Yu, J. Zhang and S. H. Bryant, Nucleic Acids Res., 2016, 44, D1202–D1213 CrossRef CAS PubMed
.
- O. Trott and A. J. Olson, J. Comput. Chem., 2010, 31, 455–461 CAS
.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef
.
- A. E. Aliev, M. Kulke, H. S. Khaneja, V. Chudasama, T. D. Sheppard and R. M. Lanigan, Proteins: Struct., Funct., Bioinf., 2014, 82, 195–215 CrossRef CAS PubMed
.
-
D. A. Case, I. Y. Ben-Shalom, S. R. Brozell, D. S. Cerutti, T. E. Cheatham, V. W. D. Cruzeiro III, T. A. Darden, R. E. Duke, D. Ghoreishi, M. K. Gilson, H. Gohlke, A. W. Goetz, D. Greene, R. Harris, N. Homeyer, Y. Huang, S. Izadi, A. Kovalenko, T. Kurtzman, T. S. Lee, S. LeGrand, P. Li, C. Lin, J. Liu, T. Luchko, R. Luo, D. J. Mermelstein, K. M. Merz, Y. Miao, G. Monard, C. Nguyen, H. Nguyen, I. Omelyan, A. Onufriev, F. Pan, R. Qi, D. R. Roe, A. Roitberg, C. Sagui, S. Schott-Verdugo, J. Shen, C. L. Simmerling, J. Smith, R. SalomonFerrer, J. Swails, R. C. Walker, J. Wang, H. Wei, R. M. Wolf, X. Wu, L. Xiao, D. M. York and P. A. Kollman, AMBER 18, University of California, San Francisco, 2018 Search PubMed
.
- A. W. Sousa da Silva and W. F. Vranken, BMC Res. Notes, 2012, 5, 1–8 CrossRef PubMed
.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef CAS PubMed
.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS
.
- R. W. Zwanzig, J. Chem. Phys., 1954, 22, 1420–1426 CrossRef CAS
.
- F. M. Ytreberg, J. Chem. Phys., 2009, 130, 164906 CrossRef PubMed
.
- N. A. Baker, D. Sept, S. Joseph, M. J. Holst and J. A. McCammon, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 10037–10041 CrossRef CAS PubMed
.
-
Schrödinger LLC, The PyMOL molecular graphics system, Versio1 1.3r1, 2010 Search PubMed
.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/d0ra07352k |
| This journal is © The Royal Society of Chemistry 2020 |