Single crystal structure and photocatalytic behavior of grafted uranyl on the Zr-node of a pyrene-based metal–organic framework†
Abstract
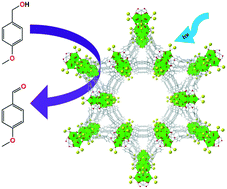
Accurately characterizing actinide oxides bound to metal–organic frameworks (MOFs) is important for designing MOFs as radioactive waste sorbents and catalytic supports. In this work, the zirconium MOF NU-1000 was post-synthetically modified through solvothermal deposition to include the uranyl (UO22+) ion and characterized via single-crystal X-ray diffraction. Bond lengths derived from the diffraction pattern and Raman spectroscopy indicate that UO22+ maintains its chemical properties upon deposition, while alcohol oxidation photocatalysis reveals photo-interactions between the pyrene linkers and the UO22+ ion.
- This article is part of the themed collection: Editor’s Collection: Zirconium based MOFs for catalysis


 Please wait while we load your content...
Please wait while we load your content...