Open Access Article
Open Access ArticleTeaching indicators to unravel the kinetic features of host–guest inclusion complexes†
Amrutha
Prabodh
 a,
Stephan
Sinn
a,
Stephan
Sinn
 *a,
Laura
Grimm
*a,
Laura
Grimm
 a,
Zsombor
Miskolczy
a,
Zsombor
Miskolczy
 b,
Mónika
Megyesi
b,
László
Biczók
b,
Mónika
Megyesi
b,
László
Biczók
 b,
Stefan
Bräse
b,
Stefan
Bräse
 cd and
Frank
Biedermann
cd and
Frank
Biedermann
 *a
*a
aKarlsruhe Institute of Technology (KIT), Institute of Nanotechnology (INT), Hermann-von-Helmholtz-Platz 1, 76344 Eggenstein-Leopoldshafen, Germany. E-mail: stephan.sinn@kit.edu; frank.biedermann@kit.edu
bInstitute of Materials and Environmental Chemistry Research Centre for Natural Sciences, Magyar tudósok körútja 2, 1117 Budapest, Hungary
cKarlsruhe Institute of Technology (KIT), Institute of Organic Chemistry (IOC), Fritz-Haber-Weg 6, 76131 Karlsruhe, Germany
dInstitute of Biological and Chemical Systems – Functional Molecular Systems (IBCS-FMS), Hermann-von-Helmholtz-Platz 1, 76344 Eggenstein-Leopoldshafen, Germany
First published on 15th July 2020
Abstract
Both thermodynamic and kinetic insights are needed for a proper analysis of association and dissociation processes of host–guest interactions. However, kinetic descriptions of supramolecular systems are scarce in the literature because suitable experimental protocols are lacking. We introduce here three time-resolved methods that allow for convenient determination of kinetic rate constants of spectroscopically silent or even insoluble guests with the macrocyclic cucurbit[n]uril family and human serum albumin (HSA) protein as representative hosts.
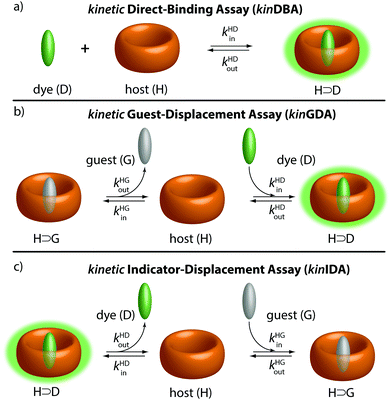
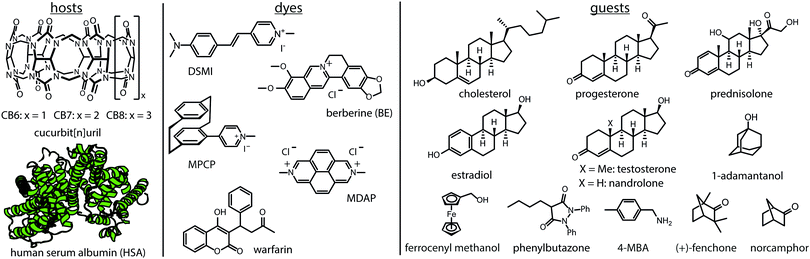
It has become clear that not only thermodynamic characteristics, e.g., binding affinities, but also the assessment of kinetic parameters (e.g., complexation and decomplexation rates) is required to obtain a full picture of supramolecular systems.1–4 For instance, kinetic rate constants of supramolecular complexes are key parameters for understanding catalysis5 and protein–ligand binding mechanisms,6–9 and stimuli-responsive materials.10,11 The design of out-of-equilibrium systems also requires knowledge of both Ka values and rate constants.12–15 However, except for CEST-active3 or slowly equilibrating systems that can be monitored by NMR (e.g., DOSY, EXSY, inversion recovery),1,16–20 kinetic rate constants of supramolecular systems are experimentally mostly only available for chromophoric or emissive systems.2,4,21–23 These experiments are typically conducted as time-resolved direct host–guest binding titration assays, herein abbreviated as kinDBA (Fig. 1a). In some cases, single molecule measurements with nanopores allowed for assessing the kinetic rate constants for complexation and decomplexation of entrapped host–guest complexes.15,24,25 Conversely, binding affinities (Ka) of host–guest complexes can be obtained for a wide range of hosts and guests by several different techniques, for instance, through NMR titrations and calorimetric measurements (ITC) as representative direct-binding assays26–28 or competitive-binding assays such as the indicator-displacement assay (IDA)28,29 and the recently by us introduced guest-displacement assay (GDA).30 Consequently, there is a strong mismatch between the number of reported binding affinities and kinetic parameters for any class of host–guest complexes. For instance, a survey for the cucurbit[n]uril (CBn)31,32 macrocyclic hosts (see Fig. 2 for their structure) on the supramolecular repository “SupraBank.org” revealed that only 3% of all entries for CBn–guest complexes included also kinetic rate constants, in agreement with the much larger number of Ka values versus kinetic parameters tabulated in reviews.
Herein, we show three novel competitive approaches through which kinetic rate constants of host–guest complexes, namely the complexation rate (kin) and decomplexation rate (kout) constants, can be accessed for spectroscopically silent guests. A competitive binding network consisting of a host (H), guest (G), and indicator dye (D) – see Fig. 1 – can be described both by thermodynamic30 and by kinetic equations (see ESI† for details). The binding affinities of the host-dye (H⊃D) and host–guest (H⊃G) complex are denoted as KHDa and KHGa, respectively. The complexation & decomplexation rate constants of the H⊃D and H⊃G complexes are symbolised by kHDin & kHDout and kHGin & kHGout, respectively. Note that an “SN1”-type, i.e., purely dissociative mechanism for the decomplexation step of the H⊃G and H⊃D complexes is implied by kinetic eqn (1)–(3).
| HG + D ⇄ HD + G | (1) |
 | (2) |
 | (3) |
| It = I0 + IHD·[HD]t + ID·[D]t | (4) |
Eqn (3) shows how the thermodynamic and kinetic parameters, i.e., affinity and rate constants, are coupled to each other. The mathematical expression for the background-corrected observable signal intensity It at time t is given by eqn (4), assuming that both the host and guest are spectroscopically silent. To kinetically characterize a supramolecular host–guest complex, it is, therefore, the task to obtain kHGin & kHGout by fitting an experimentally obtained signal-time curve of a non-equilibrated competitive binding network involving the host, guest, and dye.
The first, a conceptionally most intuitive method introduced here is the time-resolved guest-displacement assay, kinGDA. Fig. 3a shows the kinGDA traces that were obtained when the ultra-high-affinity dye MPCP33 was added to a solution of spectroscopically silent CB8⊃nandrolone complex. During the re-equilibration, nandrolone leaves the CB8 cavity, making room for the inclusion of indicator dye MPCP, which is the stronger binding guest. The detectable rate depends on (i) the concentrations of the host, guest, and dye, (ii) the rate constants kHDin and kHDout of the dye, which can be determined by a kinetic direct-binding assay (kinDBA) (see Table S3 and Fig. S2, S6–S8, S18, S24, S28, and S33, ESI†), and (iii) on the unknown rate constants kHGin and kHGout of the spectroscopically silent guest. The rate constants kHGin and kHGout can then be extracted from the time-resolved kinGDA curves through a mathematical fitting. Because the goodness of the fit improves when KHGa = kHGin/kHGout is used as an input parameter, prior KHGa determination, e.g., through competitive binding titrations such as GDA or IDA or direct binding assays (DBA) is recommended. Several host–guest pairs (Fig. S1–S34, ESI†) were analysed in this way, see Table 1. Note that the kinGDA method is extendable for determining the decomplexation rates of insoluble guests such as estradiol through precomplexation, e.g., see Table 1 for the rate constants kHGin and kHGout for the CB7⊃estradiol complex and Fig. S16 (ESI†) for the kinetic trace and fit. The applicability of kinGDA to insoluble guests is an asset it shares with the thermodynamic GDA method.30 The concept is transferable to protein–ligand interactions, as exemplified for human serum albumin (HSA) as a biological important carrier protein31,32 that is commercially available.35,36Fig. 3c demonstrates the determination of the kinetic rate constants for the binding of the anti-inflammatory drug phenylbutazone34,35 (PBZ) to HSA by kinGDA.
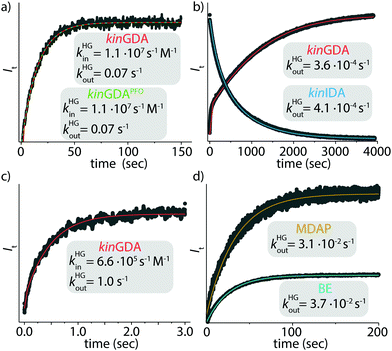 | ||
Fig. 3 Kinetic traces of (a) CB8⊃nandrolone (1 μM) and MPCP (50 μM) (●,  fitting for kinGDA and fitting for kinGDA and  fitting for kinGDAPFO), (b) CB7, nandrolone and BE (all 2 μM) either in kinGDA (●, fitting for kinGDAPFO), (b) CB7, nandrolone and BE (all 2 μM) either in kinGDA (●,  fitting) or kinIDA (●, fitting) or kinIDA (●,  fitting) mode, (c) HSA (20 μM), PBZ (40 μM) and warfarin (100 μM) in PBS in kinGDA mode (●, fitting) mode, (c) HSA (20 μM), PBZ (40 μM) and warfarin (100 μM) in PBS in kinGDA mode (●,  fitting), (d) CB7 (2 μM), nandrolone (2 μM) and MDAP (25 μM, ●, fitting), (d) CB7 (2 μM), nandrolone (2 μM) and MDAP (25 μM, ●,  fitting) or BE (50 μM, ●, fitting) or BE (50 μM, ●,  fitting), in sodium phosphate buffer (50 mM) in kinGDAPFO mode. T = 25 °C. See ESI† for details. fitting), in sodium phosphate buffer (50 mM) in kinGDAPFO mode. T = 25 °C. See ESI† for details. | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ka for host–guest and protein–ligand complexes determined by kinGDA, kinIDA and kinGDAPFO in aqueous media
Ka for host–guest and protein–ligand complexes determined by kinGDA, kinIDA and kinGDAPFO in aqueous media
| Guesta | Hosta | k HGin/s−1 M−1 | k HGout/s−1 | Methodb | log (Ka/M−1) |
|---|---|---|---|---|---|
| Errors (StDev) from triplicate experiments are ≤30% in kHGin and kHGout, see Table S3 (ESI). If not stated otherwise experiments were conducted in deionized water at 25 °C. Minor to no differences in guest binding kinetics have been found for non-desalted and desalted hosts.a See Fig. 2 for chemical structures.b See Table S3 (ESI) for indicator kinetics.c In deionized water with 8.23 μM HCl.d DSMI as dye.e See ESI for details.f H2O/ethanol (99.9/0.1; v/v) mixture.g BE as dye.h See ref. 30.i In water freshly distilled three times from dilute KMnO4 solution.j Desalted CB7/CB8.k MPCP as dye.l Determined by ITC.m CB7 (2 μM), nandrolone (log (KHGa/M−1) = 7.04;40 2 μM).n Dye (2 μM).o See ref. 40.p Dye (50 μM).q Dye (40 μM) likely associative mechanism also present, see text.r BE (50 μM) or MDAP (25 μM) in sodium phosphate buffer (50 mM).s Calculated using the formula presented in ref. 41.t MDAP as dye.u In phosphate buffered saline (PBS).v Warfarin as dye. | |||||
| 4-MBA | CB6c | 3.3 × 104 | 6.5 × 10−4 | kinIDAd | 7.7e |
| Cholesterol | CB7f | 7.0 × 104 | 8.7 × 10−2 | kinGDAg | 5.9h |
| 7.0 × 104 | 8.7 × 10−2 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g g |
|||
| Estradiol | CB7 | 4.2 × 104 | 2.0 × 10−2 | kinGDAg | 6.3h |
| 4.3 × 104 | 2.1 × 10−2 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g g |
|||
| (+)-Fenchone | CB7i | 9.2 × 104 | 3.2 × 10−3 | kinGDAg | 7.5e |
| Norcamphor | CB7i | 1.5 × 107 | 9.8 × 10−2 | kinGDAg | 8.2e |
| Adamantanol | CB7j | 1.7 × 105 | 6.6 × 10−6 | kinIDAg | 10.4e |
| CB8j | 1.2 × 107 | 1.97 | kinGDAk | 6.8l | |
| 1.2 × 107 | 1.92 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) k k |
|||
| Nandrolonem | CB7n | 4.1 × 103 | 3.6 × 10−4 | kinGDAg | 7.1o |
| 4.5 × 103 | 4.1 × 10−4 | kinIDAg | |||
| CB7p | 2.3 × 103 | 2.0 × 10−4 | kinGDAg | 7.1o | |
| 2.4 × 103 | 2.1 × 10−4 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g g |
|||
| CB7q | (9 × 103) | (8 × 10−4) | kinGDAt | 7.1o | |
| (9 × 103) | (8 × 10−4) |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) t t |
|||
| CB7r | 3.0 × 103 | 3.7 × 10−2 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g g |
5.2s | |
| 2.5 × 103 | 3.1 × 10−2 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) t t |
|||
| CB8 | 1.1 × 107 | 6.8 × 10−2 | kinGDAk | 8.2h | |
| 1.1 × 107 | 7.1 × 10−2 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) k k |
|||
| Prednisolone | CB8 | 1.6 × 106 | 1.1 | kinGDAk | 6.2o |
| 1.5 × 106 | 1.1 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) k k |
|||
| Testosterone | CB8 | 6.4 × 105 | 5.8 × 10−3 | kinGDAk | 8.0o |
| 6.4 × 105 | 5.8 × 10−3 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) k k |
|||
| Ferrocenyl methanol | CB8j | 2.1 × 107 | 5.8 | kinGDAk | 6.6l |
| 2.0 × 107 | 5.7 |
kinGDAPFO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) k k |
|||
| Phenylbutazone | HSAu | 6.6 × 105 | 1.0 | kinGDAv | 5.8h |
The second competitive kinetic method, the pseudo-first order kinGDA (kinGDAPFO), has a close analogy to some literature reports,18,36 and allows for measuring kHGout values without explicit knowledge of the kinetic rate constants of the indicator dye. Firstly, host and guest are equilibrated, followed by the spiked addition of excess of a high-affinity dye. Use of excess of the indicator allows for decoupling guest and dye rate constants for (de)complexation through a pseudo-first order approximation (see eqn (S15)–(S22), ESI†).
The kinetic trace is recorded and then fitted by a simple exponential decay function
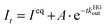 | (5) |
Finally, a third competitive method, the time-resolved indicator displacement assay (kinIDA), can be employed for obtaining kinetic rate constants. In kinIDA, a pre-equilibrated host-dye pair is mixed with the guest, to which the binding network responds with dye displacement (Fig. 1c). Indeed, comparable results were obtained for kinIDA and kinGDA for the system composed of nandrolone (G), CB7 (H) and berberine (D), see Fig. 3b.
The kinetic methods introduced herein provide meaningful rate constants if the host–guest and host-dye displacement mechanism follow a strict dissociative and not an additional, occasionally observed,36 associative mechanism. Several tests can be adopted to validate a dissociative mechanism. (i) kinGDA can be conducted at different dye concentrations and should yield similar kHGin and kHGout parameters. (ii) The kinGDA method can be compared to the analogous kinIDA setup, see above. In many cases, the competitive methods can circumvent the need for stopped-flow equipment because the equilibration times in the competitive assay format are much longer than in kinDBA. Thus, kinetic characterizations of CBn–guest complexes can now also be conducted in laboratories that do not have access to specialized stopped-flow setups. For instance, the kinetic rate constants for the CB7⊃steroid and CB8⊃steroid complexes can be determined in a cuvette equipped with a magnetic stirrer by a standard fluorescence spectrometer. Conversely, explorative kinGDA and kinIDA experiments for β-cyclodextrin complexes with high-affinity guests such as adamantanol resulted in equilibration times that were even too fast (<100 ms at 298 K) for our stopped-flow setup. The investigations of CBn complexes and the protein–ligand complex HSA⊃PBZ show that kinGDA, kinGDAPFO, and kinIDA yield reliable fits for guest egression rates kHGout ≤ 10 s−1. The kinetic rate constants that became available through the use of presented methods were converted alongside literature data to Gibb's activation energies by Eyring's equation, see also eqn (S24), (S25) and Table S4 in the ESI.† The data displayed in Fig. 4 shows a clear decoupling of thermodynamic and kinetic features for the CBn–guest and HSA–guest complexes compiled, motivating future in-depth analysis of these host–guest inclusion complexes. A first assessment demonstrates that increased thermodynamic stability is not always correlated to an increase in the kinetic inertness of the CBn–guest complexes. In conclusion, it was shown that kinIDA, kinGDA, and kinGDAPFO provide an experimental assessment of kinetic rate constants of spectroscopically silent host–guest and protein–ligand pairs. These methods will find use in the supramolecular and protein community due to their ease and scope.
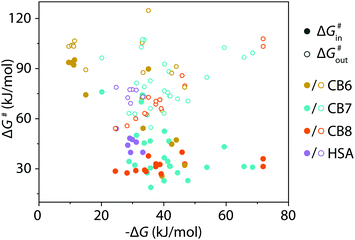 | ||
| Fig. 4 Correlation plot of Gibb's free energy (ΔG) of complex formation versus Gibb's energy of activation (ΔG#) for complexation (ΔG#in) and decomplexation (ΔG#out). Values were calculated from acquired and literature data (see ESI†) for CB6–8 and HSA. | ||
This work was financially supported through grants by the Emmy-Noether Programme of the DFG, the DAAD, NKFIH K123995, and the János Bolyai Research Scholarship Program.
Conflicts of interest
There are no conflicts of interest to declare.Notes and references
- C. Marquez, R. R. Hudgins and W. M. Nau, J. Am. Chem. Soc., 2004, 126, 5806–5816 Search PubMed.
- C. Bohne, Chem. Soc. Rev., 2014, 43, 4037–4050 Search PubMed.
- L. Avram, A. D. Wishard, B. C. Gibb and A. Bar-Shir, Angew. Chem. Int. Ed., 2017, 56, 15314–15318 Search PubMed.
- E. Masson, M. Raeisi and K. Kotturi, Isr. J. Chem., 2018, 58, 413–434 Search PubMed.
- K. I. Assaf and W. M. Nau, Chem. Soc. Rev., 2015, 44, 394–418 Search PubMed.
- H. J. Motulsky and L. C. Mahan, Mol. Pharmacol., 1984, 25, 1 Search PubMed.
- R. Casasnovas, V. Limongelli, P. Tiwary, P. Carloni and M. Parrinello, J. Am. Chem. Soc., 2017, 139, 4780–4788 Search PubMed.
- M. Bernetti, A. Cavalli and L. Mollica, MedChemComm, 2017, 8, 534–550 Search PubMed.
- P. J. Tonge, ACS Infect. Dis., 2019, 5, 796–808 Search PubMed.
- E. A. Appel, J. del Barrio, X. J. Loh and O. A. Scherman, Chem. Soc. Rev., 2012, 41, 6195–6214 Search PubMed.
- X. Yan, F. Wang, B. Zheng and F. Huang, Chem. Soc. Rev., 2012, 41, 6042–6065 Search PubMed.
- J. J. Armao IV and J.-M. Lehn, Angew. Chem. Int. Ed., 2016, 55, 13450–13454 Search PubMed.
- G. Ashkenasy, T. M. Hermans, S. Otto and A. F. Taylor, Chem. Soc. Rev., 2017, 46, 2543–2554 Search PubMed.
- J. H. van Esch, R. Klajn and S. Otto, Chem. Soc. Rev., 2017, 46, 5474–5475 Search PubMed.
- S. Borsley, J. A. Cooper, P. J. Lusby and S. L. Cockroft, Chem. – Eur. J., 2018, 24, 4542–4546 Search PubMed.
- D. J. Cram and G. M. Lein, J. Am. Chem. Soc., 1985, 107, 3657–3668 Search PubMed.
- A. Wu and L. Isaacs, J. Am. Chem. Soc., 2003, 125, 4831–4835 Search PubMed.
- J. S. Mugridge, R. G. Bergman and K. N. Raymond, Angew. Chem. Int. Ed., 2010, 49, 3635–3637 Search PubMed.
- T. J. Williams, A. D. Kershaw, V. Li and X. Wu, J. Chem. Educ., 2011, 88, 665–669 Search PubMed.
- T. Rama, E. M. Lopez-Vidal, M. D. Garcia, C. Peinador and J. M. Quintela, Chemistry, 2015, 21, 9482–9487 Search PubMed.
- Z. Miskolczy, L. Biczok and I. Jablonkai, Phys. Chem. Chem. Phys., 2017, 19, 766–773 Search PubMed.
- E. A. Appel, F. Biedermann, D. Hoogland, J. Del Barrio, M. D. Driscoll, S. Hay, D. J. Wales and O. A. Scherman, J. Am. Chem. Soc., 2017, 139, 12985–12993 Search PubMed.
- S. S. Thomas, H. Tang and C. Bohne, J. Am. Chem. Soc., 2019, 141, 9645–9654 Search PubMed.
- S. Borsley, M. M. Haugland, S. Oldknow, J. A. Cooper, M. J. Burke, A. Scott, W. Grantham, J. Vallejo, E. K. Brechin, P. J. Lusby and S. L. Cockroft, Chem, 2019, 5, 1275–1292 Search PubMed.
- Y. You, K. Zhou, B. Guo, Q. Liu, Z. Cao, L. Liu and H.-C. Wu, ACS Sens., 2019, 4, 774–779 Search PubMed.
- H.-J. Schneider and A. K. Yatsimirsky, Principles and methods in supramolecular chemistry, Wiley, Chichester, 2000 Search PubMed.
- P. Thordarson, Chem. Soc. Rev., 2011, 40, 1305–1323 Search PubMed.
- L. You, D. Zha and E. V. Anslyn, Chem. Rev., 2015, 115, 7840–7892 Search PubMed.
- S. Sinn and F. Biedermann, Isr. J. Chem., 2018, 58, 357–412 Search PubMed.
- S. Sinn, J. Kramer and F. Biedermann, Chem. Commun., 2020, 56, 6620–6623 Search PubMed.
- J. Kim, I. S. Jung, S. Y. Kim, E. Lee, J. K. Kang, S. Sakamoto, K. Yamaguchi and K. Kim, J. Am. Chem. Soc., 2000, 122, 540–541 Search PubMed.
- S. J. Barrow, S. Kasera, M. J. Rowland, J. del Barrio and O. A. Scherman, Chem. Rev., 2015, 115, 12320–12406 Search PubMed.
- S. Sinn, E. Spuling, S. Bräse and F. Biedermann, Chem. Sci., 2019, 10, 6584–6593 Search PubMed.
- V. Maes, Y. Engelborghs, J. Hoebeke, Y. Maras and A. Vercruysse, Mol. Pharmacol., 1982, 21, 100–107 Search PubMed.
- Å. Frostell-Karlsson, A. Remaeus, H. Roos, K. Andersson, P. Borg, M. Hämäläinen and R. Karlsson, J. Med. Chem., 2000, 43, 1986–1992 Search PubMed.
- Y.-C. Liu, W. M. Nau and A. Hennig, Chem. Commun., 2019, 55, 14123–14126 Search PubMed.
- M. V. Rekharsky, Y. H. Ko, N. Selvapalam, K. Kim and Y. Inoue, Supramol. Chem., 2007, 19, 39–46 Search PubMed.
- A. L. Koner, C. Márquez, M. H. Dickman and W. M. Nau, Angew. Chem. Int. Ed., 2011, 50, 545–548 Search PubMed.
- Z. Miskolczy, M. Megyesi, L. Biczok, A. Prabodh and F. Biedermann, Chemistry, 2020, 26, 7433–7441 Search PubMed.
- A. I. Lazar, F. Biedermann, K. R. Mustafina, K. I. Assaf, A. Hennig and W. M. Nau, J. Am. Chem. Soc., 2016, 138, 13022–13029 Search PubMed.
- S. Zhang, L. Grimm, Z. Miskolczy, L. Biczok, F. Biedermann and W. M. Nau, Chem. Commun., 2019, 55, 14131–14134 Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: Materials and methods, experimental details, as well as fitting equations. See DOI: 10.1039/d0cc03715j |
| This journal is © The Royal Society of Chemistry 2020 |