Emerging on-chip surface acoustic wave technology for small biomaterials manipulation and characterization
Yu
Gao
 a,
Apresio K.
Fajrial
a,
Apresio K.
Fajrial
 a,
Tao
Yang
a,
Tao
Yang
 a and
Xiaoyun
Ding
a and
Xiaoyun
Ding
 *ab
*ab
aPaul M. Rady Department of Mechanical Engineering, University of Colorado Boulder, Boulder, CO 80309, USA. E-mail: xiaoyun.ding@colorado.edu
bBiomedical Engineering Program, University of Colorado Boulder, Boulder, CO 80309, USA
First published on 18th November 2020
Abstract
A surface acoustic wave (SAW) is a sound wave travelling on the surface of an elastic material. SAW offers a robust control of the acoustic energy leading to an unparalleled versatility. As an actuator, SAW can exert acoustic forces on particles and fluids thus enabling dexterous micro/nanoscale manipulations. As a sensor, SAW has a unique sensing capability upon changes in the environment. On-chip SAW technology, in which SAW is integrated with modern lab-on-a-chip (LOC), has drawn a lot of attention in recent years and found various exciting applications in micro/nanosystems. In particular, its well-known biocompatibility provides on-chip SAW technology as an exceptional platform for biomaterials research at the small-scale. In this minireview, we highlighted recent advances of on-chip SAW technology for biomaterials manipulation and characterization with a focus on cell-based (e.g. single-cell and multicellular) biomaterials. We also discussed and shared our perspective on future directions for this emerging research field.
1. Introduction
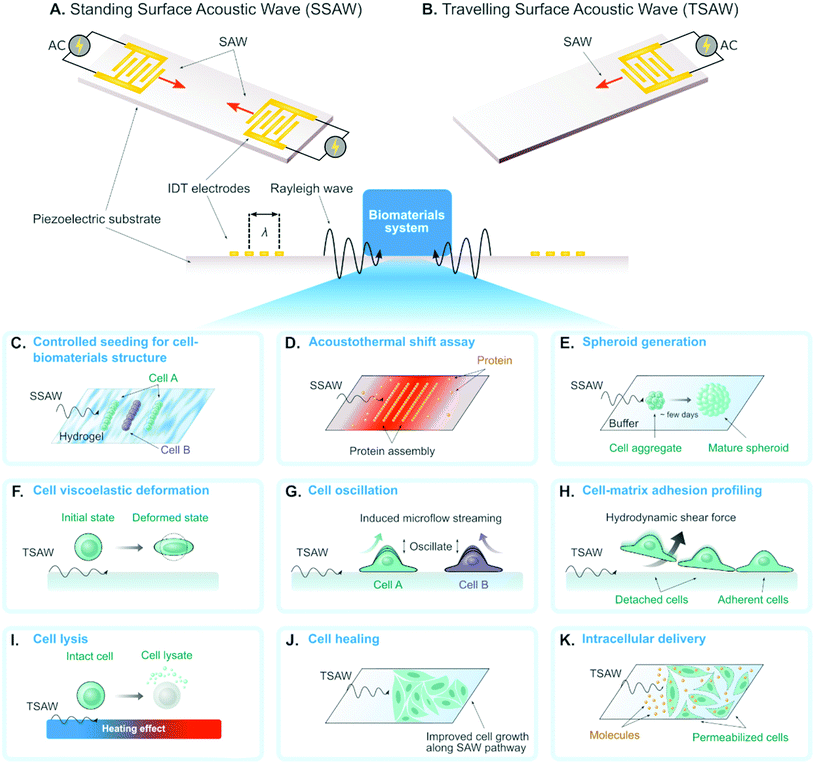
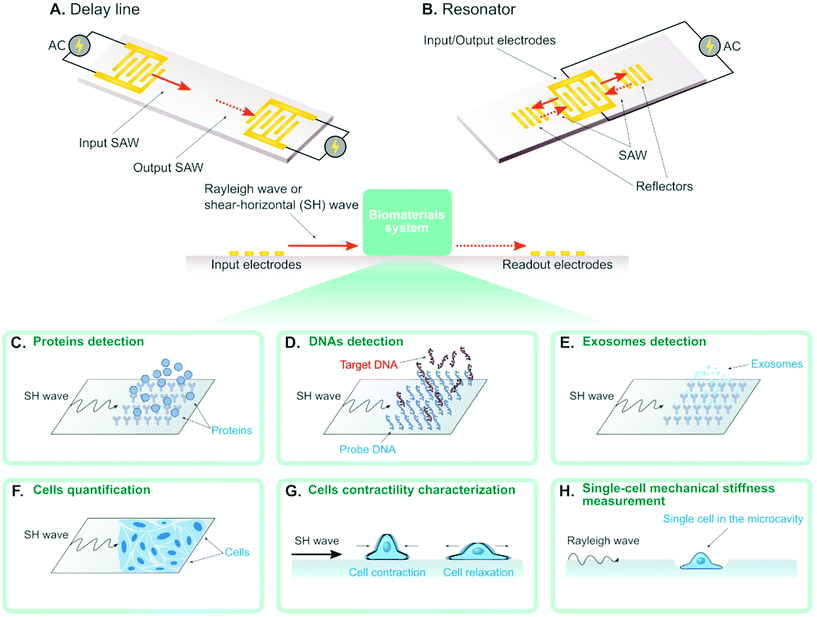
A surface acoustic wave (SAW) is an acoustic wave that propagates on the surface of an elastic material. By actuating interdigital transducers (IDTs) on the piezoelectric surface with alternating current (AC) within radio frequency range, SAW with a specified wavelength (λ) can be directly generated on piezoelectric crystals (see Fig. 1). Owing to its simple operation, SAW is often integrated with modern lab-on-a-chip (LOC) and microfluidic devices. Numerous advantages of SAW LOC and microfluidic devices have been summarized in the literature.1An important characteristic of SAW LOC is the dual role of SAW as both actuator and sensor. In a SAW LOC device, a target system is usually coupled on the piezoelectric surface where the SAW would propagate through. When the SAW passes the interface between elastic material and target system, two phenomena occur: (1) SAW exerts energy into the target system, enabling dexterous dynamics and object manipulation within the system; (2) The interaction between SAW and the target system would affect the properties of the propagating SAW, such as amplitude, velocity and phase, the change of which in turn can serve as indicators of the properties of the target system. The former is called a SAW actuator (Fig. 1) and the latter is called a SAW sensor (Fig. 2). There are two working modes in SAW actuators: standing surface acoustic wave (SSAW) by actuating pairs of IDTs or travelling surface acoustic wave (TSAW) by actuating a single IDT as shown in Fig. 1A and B respectively. In SAW sensors, there are also two configurations. The most common one is called delay line consisting of one pair of IDT as illustrated in Fig. 2A, where one IDT is used for SAW generation and the other is used as a receiver to transform the resulting SAW into electrical signal for analysis. The other one is called SAW resonator consisting of one IDT and some grating reflectors as illustrated in Fig. 2B, where the single IDT is used for both SAW generation and reception and the grating reflectors are used for SAW trapping to enhance the SAW effect. It is worth mentioning that there are multiple types of SAW generated in these SAW devices. Almost SAW actuators work with Rayleigh wave while a SAW sensor could work with either Rayleigh wave or shear-horizontal (SH) wave.1,2 Despite the similarity between the configurations of SAW actuators and SAW sensors, they function differently in real applications. More detailed information on the fundamentals of SAW devices, such as design parameters and working principles, have been described elsewhere.1,3–7
The unique versatility of SAW along with its noninvasiveness and biocompatibility8–10 establishes SAW-based miniaturized devices as emerging and powerful tools for research in biomedical fields such as bioanalytical chemistry.11 On-chip SAW technology also lends its merit to biomaterials science developments with respect to biomaterial manipulation and characterization in small scale. Albeit there emerge lots of exciting on-chip SAW designs leading this area to a hot topic, there has been limited attention to these discoveries possibly due to nonexistent literature review of the topic. Therefore, the objective of this minireview is to highlight recent on-chip SAW technology in biomaterials science, particularly manipulation and characterization of small biomaterials such as cell-based biomaterials. We will also discuss the challenges and future opportunities in this field. Ultimately, we hope this minireview can spur the adoption of on-chip SAW technology for small biomaterials manipulation and characterization.
2. SAW actuators for biomaterials
The development of SAW actuator began when people discovered that SAW could be coupled with microfluidics to provide effective control and manipulation of fluid as well as objects in the fluid around 1990s.1,7 This phenomenon is attributed to the trapping of most acoustic energy adjacent to the interface between the piezoelectric substrate and the fluid. The acoustic energy dissipates into the fluid to induce streaming effect and exert forces on any objects inside the fluid boundary without physical contact. Since then, SAW actuators have been widely used in microfluidic control and micromanipulation. Recently, various systems of small biomaterials have also been extensively studied with SAW actuators. In this section, different emerging SAW actuators for biomaterials will be introduced.2.1 Bioparticle patterning
The SAW actuators have a unique function of template-free and contactless bioparticle patterning in standing SAW mode (see Fig. 1A). By simply generating identical SAWs propagating towards each other, 1D (using one pair of IDTs) or 2D (using two pairs of IDTs) pattern of acoustic pressure nodes could be formed. The bioparticles with positive acoustic contrast factors,12 such as cells, will accumulate in these pressure nodes by acoustic forces to form the corresponding 1D/2D patterns with no pre-designed templates. The 1D pattern consists of parallel lines with equal spacing while the 2D pattern resembles checkerboard-like distribution of dots. The patterning performance highly depends on parameters in the on-chip SAW devices (e.g. such as the SAW wavelength and microfluidic channel dimensions8,13,14) and the operation modes (e.g. phase modulation of the SAW). Therefore, such patterning is highly tunable, which drives the following exciting biomaterial research:Researchers investigated the patterning as well as immobilization of cells in biodegradable, photocurable hydrogels, to create 3D biomimetic tissue structures (Fig. 1C). Typical successful cases include HeLa cells in poly(ethylene glycol) diacrylate (PEGDA) and methacrylated gelatin (GelMA) gels,15 cardiac cells in GelMA gels,16 co-aligned endothelial and stem cells in hyaluronic acid hydrogels,17 and leukemia cells in GelMA gel pillars.18 Li et al. proved the concept of co-culturing two cell types simultaneously by modulating the phase of SAW.24 This application of SAW for biofabrication facilitates vascularization in synthetic tissue that is important for long-term organ-on-chip study and artificial organ manufacturing.
Most recently, Ding et al.19 achieved to measure the protein stability by pattering the protein molecules in the pressure nodes and/or antinodes (Fig. 1D). The proteins unfolded and aggregated in the nodes due to the local heating effect and acoustic forces from SAW. In particular, the protein unfolding could be quantified by measuring the gray intensity of precipitated proteins in the nodes and the change of gray intensity as a function of SAW time was analyzed to reveal the protein thermal stability. Comparing to conventional protein thermal shift assay methods, they demonstrated that this novel acousto thermal shift assay (ATSA) enables ultrafast (with ∼1 minute) and highly sensitive (7–34 folds higher) monitoring of protein thermal stability change. Such superiority is promising for applications in fast diagnosis of blood disorder and hemoglobinopathies such as sickle cell disease.
By designing the SAW wavelength and channel height to the same order of cell diameter, single-cell 2D patterning can be achieved, where each node was occupied with a single cell.20 This unique patterning approach can be used for either assessing cellular response to effector molecules20 or growing neural networks.21
When the wavelength and the channel height are designed as tens of times of cell diameter, multicellular spheroid, which is an important 3D cell culture biomaterial for tissue engineering, could form atop the pressure nodes owing to a combination of acoustic force and SAW-induced microstreaming.14,22,23 Chen et al. recently demonstrated a high-throughput SAW actuator that can produce more than 6000 tumor spheroids in a single operation (Fig. 1E).23
2.2 Cell mechanical characterization
The mechanical properties of single-cell biomaterials provide a significant label-free marker for cell physiological conditions. Many on-chip platforms have been developed for single-cell mechanical characterization.25 Recently, SAW actuators have been emerging as a new approach to characterizing cell mechanics.With a novel design of SAW actuator, Link et al. quantitatively probed the viscoelasticity of single red blood cells (RBCs).26 A single RBC was trapped by the combined effects from SAW and fluid motion, and then was deformed by modulating the SAW power (see Fig. 1F). Typically, the degree of the deformation and the time required for the deformation process were recorded and analyzed to extract the elasticity and the viscosity of single RBCs respectively.
Salari et al. used SAW to oscillate an adherent MCF-7 cell in a microfluidic channel. This oscillation provided an indirect and qualitative way to probe cell stiffness as illustrated in Fig. 1G.27 The cell oscillation generated nontrivial acoustic flow in the channel, which was visualized by adding tracer (e.g. fluorescent particles) in the fluid. They systematically studied the correlation between the flow properties (e.g. pattern style, velocity magnitude) and the cell properties. One of the discoveries was that the oscillation of cells with different stiffness would generate flows with different maximum velocities.
SAW actuators have also been developed to characterize cell adhesive properties,28–31 where the working principle is to use SAW to induce hydrodynamic flow to detach adherent cells on the substrate (Fig. 1H). Generally, there are two approaches to evaluate cell adhesive properties. First, the SAW power was gradually increased until all the adherent cells were attached, and this critical power was recorded to characterize the cell adhesion. Second, SAW with constant power was applied for a certain period of time and the percentage of the detached cells was recorded to characterize the cell adhesion.
2.3 Cell lysis
SAW can be used to impose physical forces that dissociate cell membranes and produce chemical-free cell lysates (Fig. 1I). Applying SAW into a droplet containing cells induced shear force that is enough to lyse target cells. Usually, the use of SAW is combined with other physical structures or particles to enhance cell lysis efficiency. Reboud et al. demonstrated that patterning phononic lattices on the SAW propagation area intensified the generated vortex streaming.32,33 The phononic lattices consisted of circular wells in a square array configuration. The size of wells and the spacing between wells should be adjusted depending on the SAW frequency. They used circular wells with a radius of 82 μm and a pitch spacing of 203 μm for SAW frequency of 10 MHz. Another strategy was to incorporate diamond-shaped micropillars with sharp edges along the droplet streaming area intended to serve as disruption structures as reported by Luo and colleagues.34 The micropillars have 50 μm by 50 μm side length, 100 μm height, and 100 μm spacing in between. Instead of fabricating fixed microstructures, Wang et al. added microparticles to the droplet containing cells to increase the collision likelihood that would break the cells apart.35 The effective size of the microparticles was around 4–5 μm which resulted in a higher cell-microparticle collision frequency compared to particle sizes that are smaller (0.1 μm) or larger (15 μm). Overall, all these SAW-based techniques are capable to lyse cells up to 100% efficiency.2.4 Cell healing and growing
Controlling the amount of SAW energy applied to the cell can improve their growth as opposed to deteriorating the cell. Stamp et al. reported that exposing SAW directly to the cells increased their migration velocity leading to a 17% healing rate of an artificial wound in vitro as depicted in Fig. 1J.36 This positive effect of SAW in wound healing likely arose from mechanical and/or electrical stimulation as they claimed. The fluid streaming during SAW excitation can also dynamically recirculate the culture medium in a Petri dish thus increase the proliferation of suspension cells as demonstrated by Greco et al.37 The Petri dish was coupled to the SAW device using a thin layer (700 μm) of PDMS. The SAW-treated chamber has 36% more cell proliferation compared to the untreated control. The associated effects of SAW on living cells strongly depend on the type of cell being used. Devendran et al. reported the negative effects of SAW at a relatively high frequency (48.5 MHz), such as inhibited cell attachment and decrease in cell spreading, in mesenchymal stromal cells and mouse osteosarcoma.38 On the other hand, exposing SAW to human keratinocytes and mouse fibroblasts increased their metabolic activity. Therefore, further study to decipher the biological impacts of SAW in different cells is necessary.2.5 Intracellular delivery
Engineering cells as a living biomaterial, which is essential for cell-based tissue engineering, often requires intracellular delivery of biomolecules into target cells. Techniques for intracellular delivery include using virus vector, chemical carriers, or physical methods.39 Compared to other intracellular delivery methods, physical methods have shown various advantages for in vitro and ex vivo delivery such as high delivery performance and less dependence on cell or cargo types.40–42 In particular, intracellular delivery using acoustic waves does not require direct contact of physical probes to the target cell and is generally gentler thus preserving cell viability. Ramesan et al. demonstrated the use of SAW for intracellular delivery of lipid-siRNA complexes to adherent HEK cells using 10 MHz frequency for 5 minutes (see Fig. 1K).43 The internalization efficiency reached 39% with practically no decrease to cell viability. Unfortunately, they did not report the possibility to deliver naked siRNA using SAW which is more beneficial due to the elimination of the cost and preparation of the lipid vesicles. SAW also mediated intracellular transfer of cryopreservation agent into human umbilical cord matrix mesenchymal stem cells, a type of cells that is vital for cell-based therapies.44 The conventional method to load and unload cryopreservation agent required centrifugation and decreased the cell viability down to ∼25%. In contrast, the SAW-based method preserved cell viability up to ∼70%. When implemented at the tissue level, SAW can also assist drug delivery up to 1 mm tissue depth.45 By combining with responsive materials, such as temperature-sensitive liposome, SAW could also enable controlled drug release to target cells.462.6 Other applications
Besides the cases mentioned above, there are other designs of SAW actuators for some unique biomaterial applications. Ambattu et al. found that SAW could be used to promote exosome generation.47 Hu et al. patterned cells in versatile biomimietic arrays with six IDTs and phase modulation.48 Alhasan et al. proposed a new method to use SAW for spheroids generation, other than the patterning-based approach described previously, where the SAW induced flow in a tissue culture plate to assemble the cells.49 Also using SAW-induced flow generated by a C-shaped IDT design, Cai et al. was able to trap multiple spheroids.50 By proper alignment between the microfluidics and SAW propagation, researchers have demonstrated cell washing with high purity51 or cell coating with chemicals with high efficiency.523. SAW sensors for biomaterials
“SAW sensors” came out earlier than SAW actuators in the 1960s, when they were still regarded as SAW delay lines or SAW resonators instead of sensors and the main application was for signal processing and filtering in telecommunication industries.53,54 Later on, researchers realized the potential of these SAW devices as effective sensors and capitalized their sensing capabilities, which shed light on the field of SAW sensors. Compared to the bulk acoustic wave (BAW) sensor, such as quartz crystal microbalance, the SAW sensor is more advantageous due to its high sensitivity, low cost, minimal power consumption, precise control and ease of scaling down.55 The past thirty years have witnessed the rapid development of SAW sensors in numerous applications,2,56 where SAW sensors have been proven as promising tools for characterization of multiscale biomaterials, from molecular to cellular level.57–59 A common strategy adopted for the SAW sensor design is to immobilize biomaterial-specific molecules on the substrate to capture biomaterials from the suspension, which would lead to the phase shift of the SAW or the resonance frequency shift of the SAW sensor. On the one hand, only targeted biomaterial would be caught on the substrate so that the SAW sensor could be used for label-free detection of certain biomaterials. On the other hand, more biomaterials caught on the substrate will cause more change of the phase or the resonance frequency. In this way, SAW sensor could be used to quantify the biomaterials in suspension.In the past thirty years, detection and quantification have been the two major and conventional applications of SAW sensors for biomaterials characterization. Since then, many new SAW sensor designs have emerged. These sensors are characterizing either new biomaterials or new properties of biomaterials. In this section, different SAW sensors for biomaterials would be introduced based on the scale of the biomaterials. Particularly, this section would focus on these emerging new SAW sensors in the past several years. For more examples and details of the conventional SAW sensors for biomaterials, interested readers are referred to Länge et al.,57 Gronewold et al.,58 Rocha-Gaso et al.,59 and references therein.
3.1 Biomolecules
SAW sensors are able to detect different types of biomolecules (e.g. protein, antigen, DNA) by immobilizing the corresponding binding-molecules on the substrate. For example, the SAW sensor can detect proteins in solution when immobilizing the corresponding antigens on the substrate (Fig. 2C). Also, the SAW sensor can detect single-stranded DNAs (ssDNAs) in solution when immobilizing their complementary strands (i.e. probe DNA) on the substrates (Fig. 2D). Recently, several works show that the concentration of ssDNAs can actually be quantified, where the concentration is in proportional to the measured phase shift of the SAW.55,60–63 Some of these SAW sensors can detect the mismatch with the resolution of up to single base pair. Such high sensitivity can be achieved by applying GHz SAW as reported by Cai et al.,60 employing silver-nanoparticle amplification technique as reported by Zhang et al.62 and coating the piezoelectric surface with a sensitive layer as reported by Ji et al.61 Sensing the mismatch is important in modern structural DNA nanotechnology where ssDNAs are used to self-assemble complicated structures for different applications,64 as the mismatch is an indicator of the defect in the DNA-assembled biomaterials. From this perspective, SAW sensors hold great promise to become new characterizing tools for evaluating the quality of the DNA-based biomaterials.3.2 Exosomes
In the latest work, researchers reported the detection of exosomes for the first time using SAW sensor,65 where anti CD63 antibody was immobilized on the substrate to interact with the exosomes (Fig. 2E). The gold nanoparticle-amplification technique was integrated in the sensor to improve the sensitivity, which led to the final detection capability of 1.1 × 103 particles per mL exosomes. The exosome concentration in solution was also demonstrated to have a linear relationship with the phase shift of the SAW. This SAW sensor for exosome detection will promote new diagnostic applications such as early diagnosis of cancer in clinical practice.3.3 Biological cells
Recent advances show SAW can also be used to detect and quantify biological cells (see Fig. 2F).66–68 Chang et al. demonstrated the quantification of MCF-7 cells concentration from 1 × 102 cells per ml to 1 × 107 cells per ml, with a detection limit of 32 cells per ml.68 They used aptamers to increase the coupling between MCF-7 cells and the piezoelectric crystal substrate to improve the sensitivity. Wang et al. coupled cell culture well on SAW sensors and achieved real-time quantification of A549 cancer cell and RAW 264.7 macrophages in cell cultures.67 Future potential application of this technique is to incorporate SAW into 3D in vitro cancer models for continuous real-time measurement of cell growth in a 3D environment.The mechanical properties (e.g. viscoelasticity, stiffness) of biomaterials are important measures to understand their behavior in biological conditions. In recent years, the cell mechanical properties (e.g. stiffness, elasticity) have emerged as significant label-free markers for cell physiological conditions. Wei et al.69 used a SAW sensor to study the contractile properties of HL-1 cardiomyocytes as depicted in Fig. 2G. In the experiment, they added different chemicals that can change the contractile properties of cardiomyocytes in the solution and measured the amplitude shift and the phase shift of the SAW. They found that the cardiomyocytes with different contractile properties could be distinguished by the amplitude shift. Senveli et al. developed an approach for characterizing the single-cell mechanical stiffness by coupling the SAW sensor with a microcavity where a single cell can be trapped (Fig. 2H).70 They measured the phase shifts when the microcavity was filled with solution only and with a single cell in solution. By combining both experiment and simulation, they were able to estimate the single-cell elastic modulus of four different tumor cell types. Although the design of this SAW sensor is not conventional as the interaction between SAW and cell does not happen directly on the piezoelectric surface, this method poses a new paradigm to encourage more creative developments of SAW sensors to overcome challenges of the normal SAW sensor design. Conventionally, trapping single molecule, cell and organism on a bare piezoelectric surface is known to be challenging.
4. Conclusion
In the past decade, on-chip SAW technology indeed has permeated into the field of biomaterials science despite a limited attention from the scientific community. The unique flexibility of SAW, compared to other on-chip technologies, acting as both sensor and actuator brings this field various novel tools for biomaterials characterization and manipulation. Although SAW technology has shown promising applications in biomaterials science, there are many opportunities for continued development of the technology.First, trapping single molecule, cell and organism on the bare substrate directly is difficult when using SAW sensor to characterize their mechanical properties. The single-cell trapping proposed by Senveli et al. is inherently a gravity-assisted trapping,70 where the etched cell-trapping structure affects the SAW propagation, thus voiding the high sensitivity and versatility of SAW sensing. Ideally, the cell trapping is done without intruding the substrate so that first principles can be used to get an ideal estimate of cell mechanical properties.71 A possible solution as well as future direction is to integrate novel coupling approaches (e.g. electric, magnetic, optical, mechanical and etc.) for single molecule and single cell trapping on SAW sensors.
Moreoever, SAW can efficiently assemble simple multicellular structures such as spheroids. However, future SAW-assembly should aim for more sophisticated multicellular systems such as organoids with arbitrary shapes. Therefore, another potential research direction is to come up with new strategies for complex shape assembly, such as combining SAW with other force fields or programming the SAW in a specific way. Furthermore, another promising future direction is to integrate multiple SAW actuators or SAW sensors in a single chip platform which can handle multiple biomaterials tasks simultaneously. For example, one ideal chip platform may be able to use SAW actuation to manipulate different types of cells to different locations so that these cells can either be lysed or drug-treated by a second SAW actuation. Another ideal chip platform is expected to assemble biomaterials by SAW actuation and then characterize these biomaterials properties (e.g. mechanics, defects) using SAW sensing. Finally, we hope this minireview can spark interests from communities of both SAW microfluidics and biomaterials science to this emerging interdisciplinary topic which eventually fosters the adoption of SAW technology in biomaterials science to further advance its development.
Conflicts of interest
There are no conflicts to declare.References
- X. Ding, P. Li, S.-C. S. Lin, Z. S. Stratton, N. Nama, F. Guo, D. Slotcavage, X. Mao, J. Shi and F. Costanzo, Lab Chip, 2013, 13, 3626–3649 RSC.
- T. Nomura, A. Saitoh and Y. Horikoshi, Sens. Actuators, B, 2001, 76, 69–73 CrossRef CAS.
- M. J. Vellekoop, Ultrasonics, 1998, 36, 7–14 CrossRef.
- L. Bo, C. Xiao, C. Hualin, M. A. Mohammad, T. Xiangguang, T. Luqi, Y. Yi and R. Tianling, J. Semicond., 2016, 37, 21001 CrossRef.
- D. B. Go, M. Z. Atashbar, Z. Ramshani and H.-C. Chang, Anal. Methods, 2017, 9, 4112–4134 RSC.
- W. Connacher, N. Zhang, A. Huang, J. Mei, S. Zhang, T. Gopesh and J. Friend, Lab Chip, 2018, 18, 1952–1996 RSC.
- L. Y. Yeo and J. R. Friend, Annu. Rev. Fluid Mech., 2014, 46, 379–406 CrossRef.
- X. Ding, S.-C. S. Lin, B. Kiraly, H. Yue, S. Li, I.-K. Chiang, J. Shi, S. J. Benkovic and T. J. Huang, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 11105–11109 CrossRef CAS.
- H. Li, J. Friend, L. Yeo, A. Dasvarma and K. Traianedes, Biomicrofluidics, 2009, 3, 34102 CrossRef.
- M. Wiklund, Lab Chip, 2012, 12, 2018–2028 RSC.
- P. Li and T. J. Huang, Anal. Chem., 2018, 91, 757–767 CrossRef.
- D. Hartono, Y. Liu, P. L. Tan, X. Y. S. Then, L.-Y. L. Yung and K.-M. Lim, Lab Chip, 2011, 11, 4072–4080 RSC.
- X. Ding, J. Shi, S.-C. S. Lin, S. Yazdi, B. Kiraly and T. J. Huang, Lab Chip, 2012, 12, 2491–2497 RSC.
- K. Chen, M. Wu, F. Guo, P. Li, C. Y. Chan, Z. Mao, S. Li, L. Ren, R. Zhang and T. J. Huang, Lab Chip, 2016, 16, 2636–2643 RSC.
- J. P. Lata, F. Guo, J. Guo, P. Huang, J. Yang and T. J. Huang, Adv. Mater., 2016, 28, 8632–8638 CrossRef CAS.
- S. M. Naseer, A. Manbachi, M. Samandari, P. Walch, Y. Gao, Y. S. Zhang, F. Davoudi, W. Wang, K. Abrinia and J. M. Cooper, Biofabrication, 2017, 9, 15020 CrossRef.
- B. Kang, J. Shin, H.-J. Park, C. Rhyou, D. Kang, S.-J. Lee, Y. Yoon, S.-W. Cho and H. Lee, Nat. Commun., 2018, 9, 1–13 CrossRef.
- X. Hu, S. Zhao, Z. Luo, Y. F. Zuo, F. Wang, J. Zhu, L. Chen, D. Yang, Y. Zheng and Y. Zheng, Lab Chip, 2020, 20, 2228–2236 RSC.
- Y. Ding, K. A. Ball, K. J. Webb, Y. Gao, A. D'Alessandro, W. M. Old, M. H. B. Stowell and X. Ding, Small, 2020, 2003506 CrossRef CAS.
- D. J. Collins, B. Morahan, J. Garcia-Bustos, C. Doerig, M. Plebanski and A. Neild, Nat. Commun., 2015, 6, 1–11 Search PubMed.
- M. S. Brugger, S. Grundeen, A. Doyle, L. Theogarajan, A. Wixforth and C. Westerhausen, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2018, 98, 12411 CrossRef CAS.
- Y. Wu, Z. Ao, B. Chen, M. Muhsen, M. Bondesson, X. Lu and F. Guo, Nanotechnology, 2018, 29, 504006 CrossRef.
- B. Chen, Y. Wu, Z. Ao, H. Cai, A. Nunez, Y. Liu, J. Foley, K. Nephew, X. Lu and F. Guo, Lab Chip, 2019, 19, 1755–1763 RSC.
- S. Li, F. Guo, Y. Chen, X. Ding, P. Li, L. Wang, C. E. Cameron and T. J. Huang, Anal. Chem., 2014, 86, 9853–9859 CrossRef CAS.
- P. Pan, W. Wang, C. Ru, Y. Sun and X. Liu, J. Micromech. Microeng., 2017, 27, 123003 CrossRef.
- A. Link and T. Franke, Lab Chip, 2020, 20, 1991–1998 RSC.
- A. Salari, S. Appak-Baskoy, M. Ezzo, B. Hinz, M. C. Kolios and S. S. H. Tsai, Small, 2020, 16, 1903788 CrossRef CAS.
- N. Sivanantha, C. Ma, D. J. Collins, M. Sesen, J. Brenker, R. L. Coppel, A. Neild and T. Alan, Appl. Phys. Lett., 2014, 105, 103704 CrossRef.
- A. Bussonnière, Y. Miron, M. Baudoin, O. Bou Matar, M. Grandbois, P. Charette and A. Renaudin, Lab Chip, 2014, 14, 3556–3563 RSC.
- M. E. M. Stamp, A. M. Jötten, P. W. Kudella, D. Breyer, F. G. Strobl, T. M. Geislinger, A. Wixforth and C. Westerhausen, Diagnostics, 2016, 6, 38 CrossRef CAS.
- H. Cai, Z. Ao, Z. Wu, A. Nunez, L. Jiang, R. L. Carpenter, K. P. Nephew and F. Guo, Anal. Chem., 2019, 92, 2283–2290 CrossRef.
- J. Reboud, Y. Bourquin, R. Wilson, G. S. Pall, M. Jiwaji, A. R. Pitt, A. Graham, A. P. Waters and J. M. Cooper, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 15162–15167 CrossRef CAS.
- A. Salehi-Reyhani, F. Gesellchen, D. Mampallil, R. Wilson, J. Reboud, O. Ces, K. R. Willison, J. M. Cooper and D. R. Klug, Anal. Chem., 2015, 87, 2161–2169 CrossRef CAS.
- W. Wang, Y. Chen, U. Farooq, W. Xuan, H. Jin, S. Dong and J. Luo, Appl. Phys. Lett., 2017, 110, 143504 CrossRef.
- S. Wang, X. Lv, Y. Su, Z. Fan, W. Fang, J. Duan, S. Zhang, B. Ma, F. Liu and H. Chen, Small, 2019, 15, 1804593 CrossRef.
- M. E. M. Stamp, M. S. Brugger, A. Wixforth and C. Westerhausen, Biomater. Sci., 2016, 4, 1092–1099 RSC.
- G. Greco, M. Agostini, I. Tonazzini, D. Sallemi, S. Barone and M. Cecchini, Anal. Chem., 2018, 90, 7450–7457 CrossRef CAS.
- C. Devendran, J. Carthew, J. E. Frith and A. Neild, Adv. Sci., 2019, 6, 1902326 CrossRef CAS.
- M. P. Stewart, A. Sharei, X. Ding, G. Sahay, R. Langer and K. F. Jensen, Nature, 2016, 538, 183–192 CrossRef CAS.
- J. M. Meacham, K. Durvasula, F. L. Degertekin and A. G. Fedorov, J. Lab. Autom., 2014, 19, 1–18 CrossRef CAS.
- A. K. Fajrial, Q. Q. He, N. I. Wirusanti, J. E. Slansky and X. Ding, Theranostics, 2020, 10, 5532 CrossRef CAS.
- A. K. Fajrial and X. Ding, Nanotechnology, 2019, 30, 264002 CrossRef CAS.
- S. Ramesan, A. R. Rezk, C. Dekiwadia, C. Cortez-Jugo and L. Y. Yeo, Nanoscale, 2018, 10, 13165–13178 RSC.
- U. Farooq, Z. Haider, X. M. Liang, K. Memon, S. M. C. Hossain, Y. Zheng, H. Xu, A. Qadir, F. Panhwar and S. Dong, Small, 2019, 15, 1805361 CrossRef.
- S. Ramesan, A. R. Rezk and L. Y. Yeo, Lab Chip, 2018, 18, 3272–3284 RSC.
- L. Meng, Z. Deng, L. Niu, F. Li, F. Yan, J. Wu, F. Cai and H. Zheng, Theranostics, 2015, 5, 1203 CrossRef CAS.
- L. A. Ambattu, S. Ramesan, C. Dekiwadia, E. Hanssen, H. Li and L. Y. Yeo, Commun. Biol., 2020, 3, 1–9 CrossRef.
- X. Hu, J. Zhu, Y. Zuo, D. Yang, J. Zhang, Y. Cheng and Y. Yang, Lab Chip, 2020, 20, 3515–3523 RSC.
- L. Alhasan, A. Qi, A. Al-Abboodi, A. Rezk, P. P. Y. Chan, C. Iliescu and L. Y. Yeo, ACS Biomater. Sci. Eng., 2016, 2, 1013–1022 CrossRef CAS.
- H. Cai, Z. Wu, Z. Ao, A. Nunez, B. Chen, L. Jiang, M. Bondesson and F. Guo, Biofabrication, 2020, 12, 3 CrossRef.
- S. Li, X. Ding, Z. Mao, Y. Chen, N. Nama, F. Guo, P. Li, L. Wang, C. E. Cameron and T. J. Huang, Lab Chip, 2015, 15, 331–338 RSC.
- B. Ayan, A. Ozcelik, H. Bachman, S.-Y. Tang, Y. Xie, M. Wu, P. Li and T. J. Huang, Lab Chip, 2016, 16, 4366–4372 RSC.
- C. Campbell, Surface Acoustic Wave Devices for Mobile and Wireless Communications, Four-Volume Set, Academic press, 1998 Search PubMed.
- K. Hashimoto and K.-Y. Hashimoto, Surface acoustic wave devices in telecommunications, Springer, 2000 Search PubMed.
- J. Ji, C. Yang, F. Zhang, Z. Shang, Y. Xu, Y. Chen, M. Chen and X. Mu, Sens. Actuators, B, 2019, 281, 757–764 CrossRef CAS.
- D. S. Ballantine Jr. and H. Wohltjen, Anal. Chem., 1989, 61, 704A–715A Search PubMed.
- K. Länge, B. E. Rapp and M. Rapp, Anal. Bioanal. Chem., 2008, 391, 1509–1519 CrossRef.
- T. M. A. Gronewold, Anal. Chim. Acta, 2007, 603, 119–128 CrossRef CAS.
- M.-I. Rocha-Gaso, C. March-Iborra, Á. Montoya-Baides and A. Arnau-Vives, Sensors, 2009, 9, 5740–5769 CrossRef CAS.
- H.-L. Cai, Y. Yang, X. Chen, M. A. Mohammad, T.-X. Ye, C.-R. Guo, L.-T. Yi, C.-J. Zhou, J. Liu and T.-L. Ren, Biosens. Bioelectron., 2015, 71, 261–268 CrossRef CAS.
- J. Ji, Y. Pang, D. Li, X. Wang, Y. Xu and X. Mu, ACS Appl. Mater. Interfaces, 2020, 12, 12417–12425 CrossRef.
- Y. Zhang, F. Yang, Z. Sun, Y.-T. Li and G.-J. Zhang, Analyst, 2017, 142, 3468–3476 RSC.
- H.-L. Cai, Y. Yang, Y.-H. Zhang, C.-J. Zhou, C.-R. Guo, J. Liu and T.-L. Ren, Mod. Phys. Lett. B, 2014, 28, 1450056 CrossRef.
- A. V. Pinheiro, D. Han, W. M. Shih and H. Yan, Nat. Nanotechnol., 2011, 6, 763–772 CrossRef CAS.
- C. Wang, C. Wang, D. Jin, Y. Yu, F. Yang, Y. Zhang, Q. Yao and G.-J. Zhang, ACS Sens., 2020, 5, 362–369 CrossRef CAS.
- X. Zhang, J. Fang, L. Zou, Y. Zou, L. Lang, F. Gao, N. Hu and P. Wang, Biosens. Bioelectron., 2016, 77, 573–579 CrossRef CAS.
- T. Wang, R. Green, R. R. Nair, M. Howell, S. Mohapatra, R. Guldiken and S. S. Mohapatra, Sensors, 2015, 15, 32045–32055 CrossRef CAS.
- K. Chang, Y. Pi, W. Lu, F. Wang, F. Pan, F. Li, S. Jia, J. Shi, S. Deng and M. Chen, Biosens. Bioelectron., 2014, 60, 318–324 CrossRef CAS.
- X. Wei, J. Zhang, L. Zhuang, H. Wan and P. Wang, in 237th ECS Meeting with the 18th International Meeting on Chemical Sensors (IMCS 2020)(May 10-14, 2020), ECS, 2020.
- S. U. Senveli, Z. Ao, S. Rawal, R. H. Datar, R. J. Cote and O. Tigli, Lab Chip, 2016, 16, 163–171 RSC.
- H. Wu, H. Zu, J. H.-C. Wang and Q.-M. Wang, Eur. Biophys. J., 2019, 48, 249–260 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2021 |