Open Access Article
Open Access ArticleAlpha-hemolysin nanopore allows discrimination of the microcystins variants†
Janilson J. S. Júniorad,
Thereza A. Soares b,
Laércio Pol-Fachinbc,
Dijanah C. Machadoa,
Victor H. Rusub,
Juliana P. Aguiarad and
Cláudio G. Rodrigues
b,
Laércio Pol-Fachinbc,
Dijanah C. Machadoa,
Victor H. Rusub,
Juliana P. Aguiarad and
Cláudio G. Rodrigues *ad
*ad
aDepartamento de Biofísica e Radiobiologia, Universidade Federal de Pernambuco, Avenida Prof. Moraes Rego, s/n, Cidade Universitária, Recife, Pernambuco 50670-901, Brazil. E-mail: cgrufpe@gmail.com; Fax: +55 81 2126 8560; Tel: +55 81 2126 8535
bDepartamento de Química Fundamental, Universidade Federal de Pernambuco, Cidade Universitária 50740-560, Recife, PE, Brazil
cCentro de Pesquisas Aggeu Magalhães, Fundação Oswaldo Cruz, Cidade Universitária 50740-465, Recife, PE, Brazil
dPrograma de Pós-graduação em Inovação Terapêutica, Universidade Federal de Pernambuco, Brazil
First published on 10th May 2019
Abstract
Microcystins (MCs) are a class of cyclic heptapeptides with more than 100 variants produced by cyanobacteria present in surface waters. MCs are potent hepatotoxic agents responsible for fatal poisoning in animals and humans. Several techniques are employed in the detection of MCs, however, there is a shortage of methods capable of discriminating variants of MCs. In this work we demonstrate that the α-hemolysin (αHL) nanopore can detect and discriminate the variants (LR, YR and RR) of MCs in aqueous solution. The discrimination process is based on the analysis of the residence times of each variant of MCs within the unitary nanopore, as well as, on the amplitudes of the blockages in the ionic current flowing through it. Simulations of molecular dynamics and calculation of the electrostatic potential revealed that the variants of MCs present different charge distribution and correlated with the three patterns on the amplitudes of the blockages in the ionic current. Additionally, molecular docking analysis indicates different patterns of interaction of the variants of MCs with two specific regions of the nanopore. We conclude that αHL nanopore can discriminate variants of microcystins by a mechanism based mainly on electrostatic interaction. Finally, we propose the use of nanopore-based technology as a promising method for analyzing microcystins in aqueous solutions.
Introduction
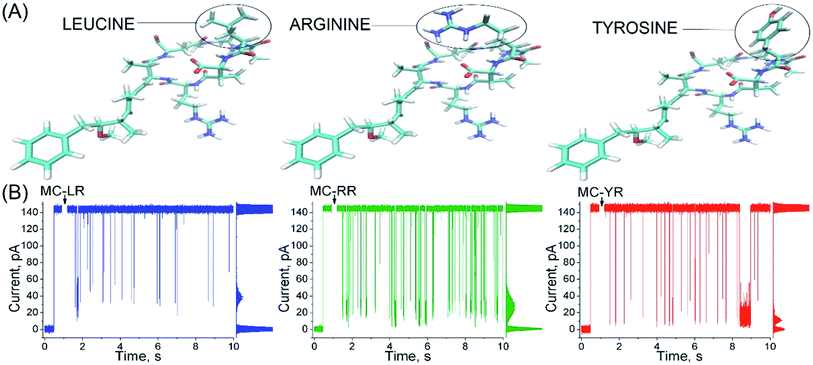
Microcystins (MCs) are constituted by cyclic heptapeptides containing several non-proteinogenic amino acids, and represent a class of exotoxins produced by cyanobacteria species commonly present in surface waters of tropical regions in the northern and southern hemispheres.1,2 MCs general sequence is cyclo-(D-Ala1-X2-D-MeAsp3-Z4-Adda5-D-Glu6-Mdha7) where the numbers indicate the position of a given amino acid in the sequence (Fig. 1). The modified residues MeAsp (Masp), Adda and Mdha corresponds to D-erythro-beta methylisoaspartic acid, (2S,3S,8S,9S)-3-amino-9-methoxy-2,6,8-trimethyl-10-phenyldeca-(4E,6E)-dienoic acid and methyldehydroalanine, respectively. Positions X2 and Z4 are occupied by variable L-amino acids, resulting in variants with different toxicity profiles.3 Currently over 100 variants of MCs have been described which differ from each other with respect to the identity of the two variable residues and chemical modifications in the other amino acids.4 The variant Microcystin-LR (MC-LR, 995 Da) containing the residues Leu2 and Arg4 is the most common and toxic of MCs.1,4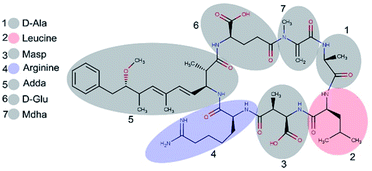 | ||
| Fig. 1 Structure of microcystin-LR. Variants differ with respect to the identity of L-amino acids in position 2 and 4. Numbers represent the position of the amino acid residue. | ||
MCs are potent hepatotoxic and nephrotoxic agents, which have been associated to liver, digestive and skin diseases, neurological impairment and death.5 It has been associated with most of the poisoning incidents worldwide.6 In February 1996, one of the largest toxic diseases caused by MCs in humans occurred in Caruaru in the Northeast of Brazil, resulting in the deaths of 60 patients.7 MCs exert their toxicity via irreversible inhibition of protein phosphatases 1 and 2A, affecting mainly liver cells, which can lead to death.8 Additionally, long-term exposure to low concentrations of MCs has been implicated in tumour growth promotion.9 The World Health Organization (WHO) recommends an upper limit of 1 μg L−1 of MCs in drinking water for human health protection.10 Eutrophication and climatic changes has led to the rapid proliferation and dominance of harmful blooms of cyanobacteria in freshwater and marine ecosystems.11,12 Temperatures ranging between 15–30 °C can selective promote cyanobacterial proliferation as the growth rates of these bacteria are optimal at relatively high temperatures.13 MCs are a growing threat to the use of freshwater ecosystems and reservoirs for drinking water, irrigation, and fishing. Therefore, fish and water consumption can be an important route of microcystin exposure for humans.14
The low concentration and large number of variants represent a considerable challenge for the monitoring of MCs in water bodies, to which several analytical methods have been employed.15 Among the most sensitive techniques are high performance liquid chromatography coupled with mass spectrometry (HPLC-MS), enzyme-linked immunosorbent assay (ELISA) and phosphatase assay. Some of these techniques offer excellent sensitivity and specificity but at high cost, time-consuming sample preparation procedures and lengthy analysis.16 Furthermore, the use of HPLC-MS for the detection and identification of unknown MCs in environmental samples is hampered mainly by the lack of analytical standards for many MC variants.3,16 In addition, HPLC-MS requires highly skilled personal and expensive equipment. Protein phosphatase inhibition assays are susceptible to high rates of false-positive results, and cannot distinguish between different MC variants.3,17 Likewise, immunoassay techniques may present cross reactivity to MCs variants.17 Therefore, there is a critical demand for the development of low-cost analytical devices for MC monitoring in water, which features high sensitivity and specificity, easiness to use and great portability. Nanopore based-systems have been successfully employed in the stochastic biosensing18,19 Stochastic biosensing is an approach that relies on the observation of individual binding events between analyte molecules and a single bioreceptor.20 Recently, solid-state nanopore with a diameter of approximately 10 nm was used to detect MC-LR via aptamer-conjugated with gold nanoparticles.21 In here, we used α-hemolysin (αHL) nanopore with diameter approximately 1 nm to discrimination microcystins variants.22 Nanopore formed by Staphylococcus aureus α-hemolysin (αHL) is the most commonly protein nanopore employed in the stochastic biosensing technology.23 Because these pores are so small, their interaction with an analyte can alter the ionic current that would otherwise flow freely. This nanopore can be used as analytical tools for a wide variety of analytes (e.g. mass spectrometry of polymers,24 DNA sequencing,25 detection of enantiomers,26 cocaine27 and poly(ethylene glycol)).28 In this work we study the interaction of MCs with the αHL-nanopore, aiming to develop a system for the detection and characterization of these cyanotoxins in aqueous systems. The choice of variants (LR, RR, YR) is due to higher occurrence in environmental water and also higher toxicity.
Results and discussion
αHL nanopores exhibited uniform single-channel conductance state of long duration at both negative and positive transmembrane potentials.20 In 40 mV an unobstructed αHL nanopore has a constant current of approximately 150 pA and conductance of ∼4 nS in KCl 4 M, pH 7.5, and the values of these parameters are in agreement with several previous reports.24,29,30 The addition of MC-LR, MC-RR or MC-YR to the trans side of nanopore induces reversible partial blockades of the ionic current, and similarly to studies with other analytes, this clearly indicates that this nanopore is capable of detecting MCs, so it works as a molecular counter31–33 (Fig. 2).The profile of the microcystin-induced ionic current blockages differ considerably from those reported in other studies using synthetic linear peptides with a number of amino acids similar to microcystins, suggesting that the nanopore is capable of discriminating the analytes by the molecular configuration.34,35 Although the MCs variants differ in only a single amino acid (Fig. 2A), each variants exhibits a typical characteristic blockages of the ionic current (Fig. 2B). Observe that the amplitude of the blockages of the ionic current caused by MC-YR is larger compared to the other variants (Table 1, right column). The MC-YR has blocked about ∼91.6% of the ionic current, followed by ∼81.6% of the MC-RR and ∼70.1% of MC-LR, respectively. Then, the magnitude of the ionic current blockages amplitude in the nanopore depends on the MC variant. As expected the percentage reduction in ionic current induced by microcystin is higher than that of acyclic peptides.34
| Nanopore state | Volumea (Å3) | Percentage of pore obstruction (theoretical) | Percentage of the ionic current blockages amplitude (experimental) |
|---|---|---|---|
| a A total of 84 residues were considered for theoretical analysis, embracing amino acids 110 to 115 and 145 to 150 from the seven αHL monomers. | |||
| Unoccupied | 1113 | — | — |
| Occupied with LR | 548 | 51% | 70.1% |
| Occupied with RR | 115 | 90% | 81.6% |
| Occupied with YR | 80 | 93% | 91.6% |
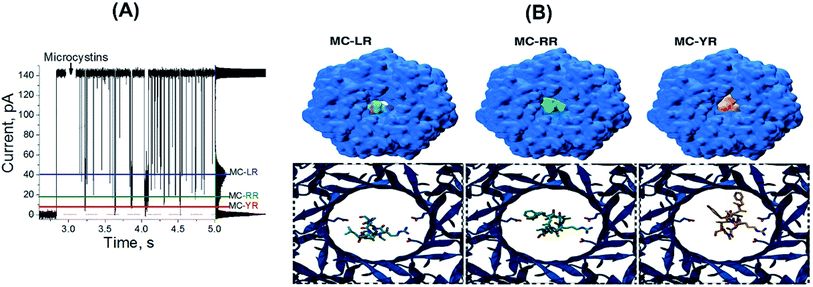
In order to rationalize the molecular basis of specific blockage types for each MCs by the αHL nanopore, molecular docking experiments were conducted. Such approach is the most common method employed to model the interaction between a small molecule and a protein at the atomic level, allowing the characterization of ligands behaviour in the binding site of target receptors, and generating novel information to help elucidating biological and biotechnological processes.36 Molecular docking is conducted through two steps: (1) sampling conformations of the ligand in the active site of the protein, which should be able to reproduce the experimental binding mode; and (2) ranking these conformations via a scoring function, which should also rank it highest among all generated conformations.37 From our calculations, the lowest energy docked conformations for each of the three MCs indeed fitted the region of constriction of the αHL nanopore (Fig. 3B), in which the Arg residues in MCs would interact with Glu111 residues. Such site of interaction was also experimentally observed and/or proposed for other compounds by previous studies.24–30
Aiming to correlate the molecular docking results and the experimental findings, a volume exclusion assumption based on “Solvent Extractor” tool output in “Voss Volume Volexator” website was performed.38 Such approach conceit that the volume not occupied by MCs should be available to allow water molecules and ionic current. A similar approach was employed for the biological nanopore MspA, by characterizing the amount of bulky-like water molecules in the protein constriction region, along with DNA molecules, during molecular dynamics simulations.38 For our molecular docking poses, a quantitative correlation was observed for MC-YR in terms of percentage of pore obstruction (93%), while both MC-RR (90%) and MC-LR (51%) presented the same trend (Table 1, third column). We can observe that the theoretical data are in agreement with the experimental results, as shown in more detail in the Table 1. Additionally, when three MCs variants are added simultaneously at equimolar concentrations, resulting in three blockages pattern (Fig. 3A). Amplitude of each of these three blocking patterns is like the results shown in Fig. 2B. So, αHL nanopore can distinguish different MCs variants simultaneously present in aqueous solutions.
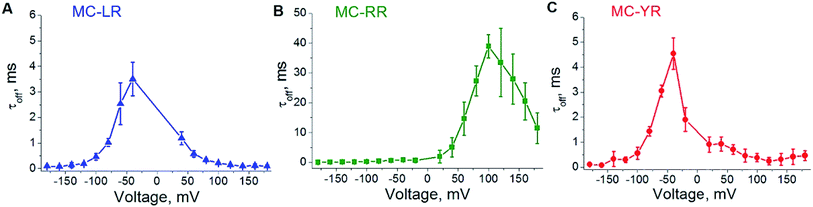
To analysing the dynamics of the interaction between microcystins and nanopore, we considering the process as on a simple bimolecular complexation reaction.29,39 A potential of −180 to 180 mV was applied at increments of 20 mV and measure the dwell time (τoff) of the MCs variants within the αHL nanopore (Fig. 4). For MC-RR, τoff is the highest among the MCs variants at the voltage of 100 mV (τoff = 38.96 ± 3.80 ms) (Fig. 4B). In relation to the variants LR and YR, on the potential of −40 mV the values of this parameters were (τoff = 3.50 ± 0.65 ms) (Fig. 4A) and (τoff = 4.54 ± 0.63 ms) (Fig. 4C), respectively.
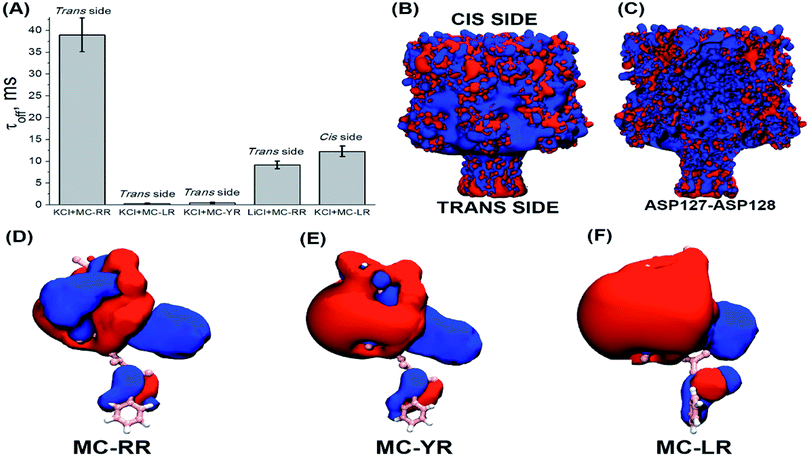
These results show a voltage-dependent behaviour of the residence time of MC variants within αHL nanopore. Analysis of the dwell times (τoff) for the three MC variants has been performed at the same transmembrane potential (Fig. 5A). It indicates that the MCs variants interact differently with the nanopore. It is known that the nanopore presents the negatively charged trans region due to the presence of the amino acid lysine and aspartic acid.40–42 So, we resolved to evaluate the contribution of electrostatic forces in the interaction of microcystin with the nanopore and for this we performed calculations of electrostatic potential of these molecules. Electrostatic potential and molecular docking calculations were performed for the αHL nanopore and MCs variants to elucidate the molecular basis of their differential interactions. The basal region on the trans side of the nanopore is rich in negatively charged residues, so, a potential of 100 mV applied on the same side as the microcystin was added, may influence their interaction with the nanopore (Fig. 5B and C). Since the net charge of the MC variants differs, these molecules will exhibit different interaction patterns with the nanopore, but also with the applied electrical field. In our experimental solution of pH 7.5, the net charge of MC-RR is 0 whereas while MC-YR and MC-LR have a net charge of −1.43 The electrostatic potentials calculated for the two latter variants are similar though quite dissimilar from the potential obtained for MC-RR (Fig. 5D–F).
The variant MC-RR shows a mostly positive potential due to the presence of two Arg residues while the electrostatic potential generated by MC-YR and MC-LR are negative. In fact, the molecular docking poses at Fig. 3B indicate that each of the two Arg residues in MC-RR interact with two Asp127 residues in opposite sides of the nanopore, thus possibly increasing the MCs-nanopore interaction and residence time with respect to MC-YR and MC-LR. This is entirely consistent with the measurements of τoff (Fig. 5A) and suggests an electrostatic interaction between MC-RR and αHL nanopore, in agreement with the experimental data. As the inner surface (lumen) of the nanopore is positively charged, then the MC-RR, is more likely to remain longer in the entrance than in the intermediate region of the nanopore (Fig. 5C). Moreover, the predominantly negative potential surface of MC-YR and MC-LR should be less attractive to the negative potential in the trans side of the nanopore, in which their Arg residue interacts with a single Asp127 residue. As depicted in Fig. 5B, the basal region of the nanopore (trans side) is rich in aspartic acid residues, which gives a net negative charge of the trans end of the nanopore.40–42
Additionally, recent studies have shown that lithium can shield charges in base of the stem region of the αHL nanopore.42 Based on this knowledge, we have changed the bath solution (KCl 4 M to LiCl 4 M), in order to shielding the charges of the stem region of nanopore and verified their influence on the residence time of the MC-RR. We can observe in Fig. 5A that there was a decrease in the residence time of the MC-RR, since without the negatively charged region, the electrostatic contribution of the interaction between the MC-RR and the nanopore is lower. Furthermore, we observed a significant increase in residence time of MC-LR, so negatively charged (Fig. 5A), by simply changing the experimental condition, by adding the MC-LR to the cis side of the nanopore. It is noteworthy that the MD-generated structural ensembles of the MC variants are quite similar with respect to the backbone conformation. The backbone conformation is maintained due to the presence of stable intra-ring hydrogen bonds between residue pairs (D-Masp![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Oδ-DGlu
Oδ-DGlu![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH, D-Masp
NH, D-Masp![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Oδ-ADDA
Oδ-ADDA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH, DGlu
NH, DGlu![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) O–X
O–X![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH where X is either Arg, Tyr or Leu) (Fig. 1). These interactions are persistent throughout the 10 ns simulation time. Therefore, the calculated electrostatic potentials for the MC variants are representative of the average conformation of these cyclic peptides in solution.
NH where X is either Arg, Tyr or Leu) (Fig. 1). These interactions are persistent throughout the 10 ns simulation time. Therefore, the calculated electrostatic potentials for the MC variants are representative of the average conformation of these cyclic peptides in solution.
Although the MC-LR and MC-YR present a similar electrostatic potential, their residence times were different (Fig. 5A), suggesting that non-electrostatic factors also influence the interaction of these toxins with the nanopore.
Aiming to elucidate the contribution of non-electrostatic factors in the interaction of microcystin with nanopore, we decided to analyse the role of hydration in the interaction of the analyte with the nanopore. In this case, we adopt the same approach of our previous studies,23,39 in which we show that the larger hydration of the analyte decreases the probability of formation and stabilization of analyte–nanopore complex. So, presence of the leucine residue gives greater hydrophobicity to MC-LR, leaving it less hydrated and increasing its residence time.
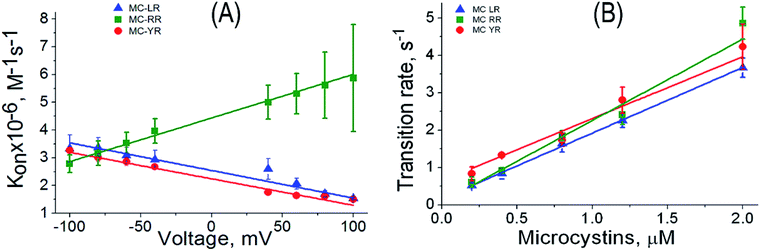
Another important parameter to analysing the kinetic process is the association rate constant (kon). This parameter is correlated with the frequency of blockages caused by analytes.39,44 The kon was calculated by assuming on a simple bimolecular complexation reaction between MCs and αHL nanopore at the single-molecule level, so that kon = 1/(τon × [MC]), where [MC] is the MC concentration and τon is the characteristic time between successive blockages.30,39 The frequency of blockages of the αHL nanopore decrease with the increasing potential for MC-LR and MC-YR, but not for MC-RR (Fig. 6). This behaviour can be expected based on the distinct net charge of the MC variants. It is known that under the solutions conditions used in our experiments (pH 7.5) the MCs LR, YR and RR exhibit net charges of −1, −1 and 0, respectively.43,45 However, our results indicate that MC-RR can acquire a positive charge (Fig. 6A). We suggest that this may occur because MC-RR forms complex with K+, such as occurs with some neutral molecules in the presence of 4 M KCl.30,39,44 In this context is possible to use the nanopore as molecular tool for the rapid probing of the capabilities of different MCs variants to form complexes with different ions. One of the parameters used to evaluate the sensing capacity of the αHL nanopore is to evaluate the behaviour of the transition rate as a function of analyte concentration.29,39 In our case, the slope of the transition rate-MC concentration increases linearly and depends on the MC variant (Fig. 6B), thus providing a basis for quantifying these cyanotoxins.
Recently method based in solid-state nanopore was used to detect MC-LR via aptamer-conjugated with gold nanoparticles and showed sensing capacity at nanomolar level.21 Our case, αHL nanopore method detected microcystin in the micromolar range above the levels of microcystins usually found in ambient waters. On the other hand, this method does not involve labelling, modifying or conjugating microcystins with nanoparticles or antibodies, simplifying the assay procedures and reducing the time of analysis. Furthermore, several strategies, for example change in the ionic solution composition,29 may be used in future studies for improvements in the present sensing capacity of the αHL nanopore.
Experimental
Materials and methods
| ∇ε(r)∇Φ(r) = −4πρint(r) + λ(r)k2Φ(r) | (1) |
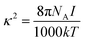 | (2) |
Conclusions
In summary, we demonstrate that the αHL nanopore can detect and discriminate three structural variants (LR, YR, RR) of microcystins simultaneously, by the difference in the blocking profiles in the ionic current induced by each of the variants. Using computational methods, it was found that electrostatic interactions contribute to the interaction between MCs and the base of the stem region nanopore. The positively charged profile acquired by MC-RR facilitates their interaction with the nanopore, which does not occur with the MCs LR and YR. In addition, being less hydrated, MC-LR has more interaction with the nanopore than the MC-YR. Finally, we concluded that αHL nanopore discriminates variants of microcystins by a mechanism based mainly on electrostatic interaction. Therefore, we propose the use of the nanopore-based technology as an auxiliary method of MCs analysis in aqueous solutions.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This work was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Grant 470008/2012-1 and, Fundação de Amparo à Ciência e Tecnologia do Estado de Pernambuco (FACEPE), Grant APQ-0707-2.09/15, Brazil.Notes and references
- J. Fawell, R. Mitchell, D. Everett and R. Hill, The toxicity of cyanobacterial toxins in the mouse: I Microcystin-LR, Hum. Exp. Toxicol., 1999, 18, 162–167 CrossRef CAS PubMed.
- A. Srivastava, S. Singh, C. Y. Ahn, H. M. Oh and R. K. Asthana, Monitoring approaches for a toxic cyanobacterial bloom, Environ. Sci. Technol., 2013, 47(16), 8999–9013 CrossRef CAS PubMed.
- J. McElhiney and L. A. Lawton, Detection of the cyanobacterial hepatotoxins microcystins, Toxicol. Appl. Pharmacol., 2005, 203(3), 219–230 CrossRef CAS PubMed.
- J. Puddick, M. R. Prinsep, S. A. Wood, S. A. F. Kaufononga, S. C. Cary and D. P. Hamilton, High levels of structural diversity observed in microcystins from microcystis CAWBG11 and characterization of six new microcystin congeners, Mar. Drugs, 2014, 12(11), 5372–5395 CrossRef PubMed.
- W. W. Carmichael, S. M. Azevedo, J. S. An, R. J. Molica, E. M. Jochimsen, S. Lau, K. L. Rinehart, G. R. Shaw and G. K. Eaglesham, Human Fatalities from Cyanobacteria: Chemical and Biological Evidence for Cyanotoxins, Environ. Health Perspect., 2001, 109(7), 663–668 CrossRef CAS PubMed.
- D. Drobac, N. Tokodi, J. Simeunović, V. Baltić, D. Stanić and Z. Svirčev, Human exposure to cyanotoxins and their effects on health, Arh. Hig. Rada Toksikol., 2013, 64(2), 305–316 CAS.
- S. Pouria, A. De Andrade, J. Barbosa, R. L. Cavalcanti, V. T. S. Barreto, C. J. Ward, W. Preiser, G. K. Poon, G. H. Neild and G. A. Codd, Fatal microcystin intoxication in haemodialysis unit in Caruaru, Brazil, Lancet, 1998, 352(9121), 21–26 CrossRef CAS.
- R. M. Dawson, The toxicology of microcystins, Toxicon, 1998, 36(7), 953–962 CrossRef CAS PubMed.
- M. M. Gehringer, Microcystin-LR and okadaic acid-induced cellular effects: A dualistic response, FEBS Lett., 2004, 557, 1–8 CrossRef CAS PubMed.
- H. G. Gorchev and G. Ozolins, Guidelines for Drinking-water Quality, WHO, 2011, vol. 1, p. 564 Search PubMed.
- H. W. Paerl and J. Huisman, Blooms like it hot, Science, 2008, 320(5872), 57–58 CrossRef CAS PubMed.
- H. W. Paerl, Mitigating Harmful Cyanobacterial Blooms in a Human- and Climatically-Impacted World, Life, 2014, 4(4), 988–1012 CrossRef PubMed.
- C. Butterwick, S. I. Heaney and J. F. Talling, Diversity in the influence of temperature on the growth rates of freshwater algae, and its ecological relevance, Freshwater Biol., 2005, 50(2), 291–300 CrossRef.
- A. E. Poste, R. E. Hecky and S. J. Guildford, Evaluating microcystin exposure risk through fish consumption, Environ. Sci. Technol., 2011, 45(13), 5806–5811 CrossRef CAS PubMed.
- S. Singh, A. Srivastava, H. M. Oh, C. Y. Ahn, G. G. Choi and R. K. Asthana, Recent trends in development of biosensors for detection of microcystin, Toxicon, 2012, 60(5), 878–894 CrossRef CAS PubMed.
- L. M. Alvarenga, J. Muzard, A. Ledreux, C. Bernard and P. Billiald, Colorimetric engineered immunoprobe for the detection and quantification of microcystins, J. Immunol. Methods, 2014, 406, 124–130 CrossRef CAS PubMed.
- J. McElhiney, L. A. Lawton and A. J. R. Porter, Detection and quantification of microcystins (cyanobacterial hepatotoxins) with recombinant antibody fragments isolated from a naive human phage display library, FEMS Microbiol. Lett., 2000, 193, 83–88 CAS.
- Nanopore-based Technology, ed. M. E. Gracheva, Humana Press, New York, 2012 Search PubMed.
- J. P. Aguiar, J. J. S. Júnior, D. C. Machado, M. C. A. Melo and C. G. Rodrigues, Biossensoriamento estocástico via nanoporo proteico individual no desenvolvimento de ferramentas analíticas, Quim. Nova, 2015, 38(6), 817–827 CAS.
- L. Q. Gu, O. Braha, S. Conlan, S. Cheley and H. Bayley, Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter, Nature, 1999, 398(6729), 686–690 CrossRef CAS PubMed.
- F. He, L. Liang, S. Zhou, W. Xie, S. He, Y. Wang, C. Tlili, S. Tong and D. Wang, Label-Free Sensitive Detection of Microcystin-LR via Aptamer-Conjugated Gold Nanoparticles Based on Solid-State Nanopores, Langmuir, 2018, 34(49), 14825–14833 CrossRef CAS PubMed.
- P. G. Merzlyak, L. N. Yuldasheva, C. G. Rodrigues, C. M. Carneiro, O. V. Krasilnikov and S. M. Bezrukov, Polymeric nonelectrolytes to probe pore geometry: application to the alpha-toxin transmembrane channel, Biophys. J., 1999, 77(6), 3023–3033 CrossRef CAS PubMed.
- P. A. Gurnev and E. M. Nestorovich, Channel-forming bacterial toxins in biosensing and macromolecule delivery, Toxins, 2014, 6(8), 2483–2540 CrossRef CAS PubMed.
- J. W. F. Robertson, C. G. Rodrigues, V. M. Stanford, K. A. Rubinson, O. V. Krasilnikov and J. J. Kasianowicz, Single-molecule mass spectrometry in solution using a solitary nanopore, Proc. Natl. Acad. Sci. U. S. A., 2007, 104(20), 8207–8211 CrossRef CAS PubMed.
- H. Bayley, Nanopore sequencing: From imagination to reality, Clin. Chem., 2015, 61(1), 25–31 CrossRef CAS PubMed.
- X. F. Kang, S. Cheley, X. Guan and H. Bayley, Stochastic detection of enantiomers, J. Am. Chem. Soc., 2006, 128(33), 10684–10685 CrossRef CAS PubMed.
- R. Kawano, T. Osaki, H. Sasaki, M. Takinoue, S. Yoshizawa and S. Takeuchi, Rapid detection of a cocaine-binding aptamer using biological nanopores on a chip, J. Am. Chem. Soc., 2011, 133(22), 8474–8477 CrossRef CAS PubMed.
- G. Baaken, I. Halimeh, L. Bacri, J. Pelta, A. Oukhaled and J. C. Behrends, High-Resolution Size-Discrimination of Single Nonionic Synthetic Polymers with a Highly Charged Biological Nanopore, ACS Nano, 2015, 9(6), 6443–6449 CrossRef CAS PubMed.
- C. G. Rodrigues, D. C. Machado, A. M. B. Silva, J. J. S. Júnior and O. V. Krasilnikov, Hofmeister effect in confined spaces: Halogen ions and single molecule detection, Biophys. J., 2011, 100(12), 2929–2935 CrossRef CAS PubMed.
- C. G. Rodrigues, D. C. Machado, S. F. Chevtchenko and O. V. Krasilnikov, Mechanism of KCl enhancement in detection of nonionic polymers by nanopore sensors, Biophys. J., 2008, 95(11), 5186–5192 CrossRef CAS PubMed.
- S. M. Bezrukov, Ion channels as molecular coulter counters to probe metabolite transport, J. Membr. Biol., 2000, 174(1), 1–13 CrossRef CAS PubMed.
- D. Branton, D. W. Deamer, A. Marziali, H. Bayley, S. A. Benner, T. Butler, M. Di Ventra, S. Garaj, A. Hibbs, X. Huang, S. B. Jovanovich, P. S. Krstic, S. Lindsay, X. S. Ling, C. H. Mastrangelo, A. Meller, J. S. Oliver, Y. V. Pershin, J. M. Ramsey, R. Riehn, G. V. Soni, V. Tabard-Cossa, M. Wanunu, M. Wiggin and J. Schloss, The potential and challenges of nanopore sequencing, Nat. Biotechnol., 2008, 26(10), 1146–1153 CrossRef CAS PubMed.
- L. Q. Gu, M. Wanunu, M. X. Wang, L. McReynolds and Y. Wang, Detection of miRNAs with a nanopore single-molecule counter, Expert Rev. Mol. Diagn., 2012, 12(6), 573–584 CrossRef CAS PubMed.
- L. Movileanu, J. P. Schmittschmitt, J. M. Scholtz and H. Bayley, Interactions of peptides with a protein pore, Biophys. J., 2005, 89(2), 1030–1045 CrossRef CAS PubMed.
- Q. Zhao, D. A. Jayawardhana, D. Wang and X. Guan, Study of peptide transport through engineered protein channels, J. Phys. Chem. B, 2009, 113(11), 3572–3578 CrossRef CAS PubMed.
- B. McConkey, V. Sobolev and M. Edelman, The Performance of Current Methods in Ligand–Protein Docking, Curr. Sci., 2002, 83(7), 845–856 CAS.
- X.-Y. Meng, H.-X. Zhang, M. Mezei and M. Cui, Molecular Docking: A powerful approach for structure-based drug discovery, Curr. Comput.-Aided Drug Des., 2011, 7(2), 146–157 CrossRef CAS PubMed.
- S. Bhattacharya, J. Yoo and A. Aksimentiev, Water Mediates Recognition of DNA Sequence via Ionic Current Blockade in a Biological Nanopore, ACS Nano, 2016, 10(4), 4644–4651 CrossRef CAS PubMed.
- D. C. Machado, J. J. S. Júnior, M. C. A. Melo, A. M. B. Silva, A. Fontes and C. G. Rodrigues, Effects of alkali and ammonium ions in the detection of poly(ethyleneglycol) by alpha-hemolysin nanopore sensor, RSC Adv., 2016, 6(61), 56647–56655 RSC.
- S. Y. Noskov, W. Im and B. Roux, Ion Permeation through the a-Hemolysin Channel: Theoretical Studies Based on Brownian Dynamics and Poisson-Nernst-Plank Electrodiffusion Theory, Biophys. J., 2004, 87, 2299–2309 CrossRef CAS PubMed.
- L. R. Teixeira, P. G. Merzlyak, A. Valeva and O. V. Krasilnikov, Interaction of heparins and dextran sulfates with a mesoscopic protein nanopore, Biophys. J., 2009, 97(11), 2894–2903 CrossRef CAS PubMed.
- S. Bhattacharya, J. Muzard, L. Payet, J. Math, U. Bockelmann, A. Aksimentiev and V. Viasnoff, Rectification of the Current in α-Hemolysin Pore Depends on the Cation Type: The Alkali Series Probed by Molecular Dynamics Simulations and Experiments, J. Phys. Chem., 2011, 115(10), 4255–4264 CAS.
- P. G. J. Maagd, A. J. Hendriks, W. Seinena and D. T. H. M. Sijma, pH-Dependent hydrophobicity of the cyanobacteria toxin microcystin-LR, Water Res., 1999, 33(3), 677–680 CrossRef.
- M. F. Breton, F. Discala, L. Bacri, D. Foster, J. Pelta and A. Oulchaled, Exploration of Neutral Versus Polyelectrolyte Behavior of Poly(ethylene glycol)s in Alkali Ion Solutions using Single-Nanopore Recording, J. Phys. Chem. Lett., 2013, 4(13), 2202–2208 CrossRef CAS.
- I. Liu, L. A. Lawton and P. K. Robertson, Mechanistic studies of the photocatalytic oxidation of microcystin-LR: an investigation of byproducts of the decomposition process, Environ. Sci. Technol., 2003, 37(14), 3214–3219 CrossRef CAS PubMed.
- G. B. Trogen, A. Annila, J. Eriksson, M. Kontteli, J. Meriluoto, I. Sethson, J. Zdunek and U. Edlund, Conformational studies of microcystin-LR using NMR spectroscopy and molecular dynamics calculations, Biochemistry, 1996, 35(10), 3197–3205 CrossRef CAS PubMed.
- C. Oostenbrink, T. A. Soares, N. F. A. Van Der Vegt and W. F. Van Gunsteren, Validation of the 53A6 GROMOS force field, Eur. Biophys. J., 2005, 34(4), 273–284 CrossRef CAS PubMed.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. C. Berendsen, GROMACS: Fast, flexible, and free, J. Comput. Chem., 2005, 26(16), 1701–1718 CrossRef CAS PubMed.
- B. Hess, C. Kutzner, D. Van Der Spoel and E. Lindahl, GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation, J. Chem. Theory Comput., 2008, 4(3), 435–447 CrossRef CAS PubMed.
- H. J. C. Berendsen, J. P. M. Postma, W. F. van Gunsteren and J. Hermans, Interaction models for water in relation to protein hydration, in Intermolecular Forces, ed. B. Pullman, Springer Netherlands, 1981, pp. 331–342 Search PubMed.
- H. J. C. Berendsen, J. P. M. Postma, W. F. van Gunsteren, A. DiNola and J. R. Haak, Molecular dynamics with coupling to an external bath, J. Chem. Phys., 1984, 81(8), 3684–3690 CrossRef CAS.
- W. Humphrey, A. Dalke and K. Schulten, VMD: Visual molecular dynamics, J. Mol. Graphics, 1996, 14(1), 33–38 CrossRef CAS PubMed.
- N. A. Baker, D. Sept, S. Joseph, M. J. Holst and J. A. McCammon, Electrostatics of nanosystems: application to microtubules and the ribosome, Proc. Natl. Acad. Sci. U. S. A., 2001, 98(18), 10037–10041 CrossRef CAS PubMed.
- F. Fogolari, A. Brigo and H. Molinari, The Poisson-Boltzmann equation for biomolecular electrostatics: A tool for structural biology, J. Mol. Recognit., 2002, 15(6), 377–392 CrossRef CAS PubMed.
- T. J. Dolinsky, J. E. Nielsen, J. A. McCammon and N. A. Baker, PDB2PQR: An automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations, Nucleic Acids Res., 2004, 32, 665–667 CrossRef PubMed.
- D. Sitkoff, K. A. Sharp and B. Honig, Accurate Calculation of Hydration Free Energies Using Macroscopic Solvent Models, J. Phys. Chem., 1994, 98(7), 1978–1988 CrossRef CAS.
- L. Song, M. R. Hobaugh, C. Shustak, S. Cheley, H. Bayley and J. E. Gouaux, Structure of staphylococcal alpha-hemolysin, a heptameric transmembrane pore, Science, 1996, 274(5294), 1859–1865 CrossRef CAS PubMed.
- C. S. Magalhães, D. M. Almeida, H. J. C. Barbosa and L. E. Dardenne, A dynamic niching genetic algorithm strategy for docking highly flexible ligands, Inf. Sci., 2014, 289, 206–224 CrossRef.
- N. R. Voss and M. Gerstein, 3V: Cavity, channel and cleft volume calculator and extractor, Nucleic Acids Res., 2010, 38, 555–562 CrossRef PubMed.
Footnote |
| † The authors dedicate this work to Dr Oleg Vladimirovich Krasilnikov (in memoriam). |
| This journal is © The Royal Society of Chemistry 2019 |