Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Ratiometric fluorescent probe for sensing Streptococcus mutans glucosyltransferase, a key factor in the formation of dental caries†
Lei
Feng‡
 ab,
Qingsong
Yan‡
ac,
Baojing
Zhang‡
a,
Xiangge
Tian
ac,
Chao
Wang
ab,
Qingsong
Yan‡
ac,
Baojing
Zhang‡
a,
Xiangge
Tian
ac,
Chao
Wang
 *ab,
Zhenlong
Yu
a,
Jingnan
Cui
*ab,
Zhenlong
Yu
a,
Jingnan
Cui
 b,
Dean
Guo
*c,
Xiaochi
Ma
b,
Dean
Guo
*c,
Xiaochi
Ma
 ac and
Tony D.
James
ac and
Tony D.
James
 *d
*d
aCollege of Pharmacy, Academy of Integrative Medicine, National & Local Joint Engineering Research Center for Drug Development of Neurodegenerative Disease, Dalian Medical University, Dalian 116044, China. E-mail: wach_edu@sina.com
bState Key Laboratory of Fine Chemicals, Dalian University of Technology, Dalian 116024, China
cChinese Acad Sci, Natl Engn Lab TCM Standardizat Technol, Shanghai Res Ctr Modernizat Tradit Chinese Med, Shanghai Inst Mat Med, Shanghai 201203, China. E-mail: daguo@simm.ac.cn
dDepartment of Chemistry, University of Bath, Bath BA2 7AY, UK. E-mail: t.d.james@bath.ac.uk
First published on 25th February 2019
Abstract
We report on a naphthalimide ratiometric fluorescent probe for the real-time sensing and imaging of pathogenic bacterial glucosyltransferases, which are associated with the development of dental caries. Using a high-throughput screening method, we identified that several natural polyphenols from green tea were GTFs inhibitors that could eventually lead to suitable oral treatments to prevent the development of dental caries.
Dental caries are the most common oral disease and are believed to be induced by pathogenic bacteria. Streptococcus mutans (S. mutans) exists in the oral cavity and is well known as the major bacteria of dental plaques resulting in the formation of dental caries.1 The formation process of dental caries has been investigated fully, and the glucosyltransferases (GTFs) have been determined to be the significant virulence factors, by the synthesis of extracellular glucan polymers.2 Several functions of glucan polymers have been implicated in the formation of dental caries, including promoting the adhesion of pathogenic bacteria to teeth and the formation of biofilms, which protects the pathogenic bacteria against antibacterial agents and facilitates the proliferation of bacteria in dental plaques.3 Therefore, glucan polymers are regarded as important in the formation of dental plaque and dental caries. While there are various pathogenic bacteria, including S. mutans widely occurring in the oral cavity, the cariogenicity of these bacteria is closely related to the expression levels of GTFs.4 Therefore, the real time detection and assay of GTFs could be an efficient approach for the prevention, diagnosis and treatment of dental caries.1 Moreover, the inhibiton of GTFs could be applied to effectively prevent dental plaque formation and to kill pathogenic bacteria that infect teeth.
Fluorescence systems have been extensively designed for the sensing of endogenous bioactive enzymes.5 In particular systems based on the ratio of fluorescence intensities at two wavelengths (ratiometric fluorescent probes) have been widely applied to assay biological enzymes activity, as well as the sensitive detection and imaging of endogenous enzymes due to the calibrating free determination facilitated by these ratiometric probes.6 Naphthalimide is an excellent ratiometric fluorescence skeleton that has previously been developed to image various biological enzymes,7 several of which were designed to detect glucosyltransferases in fungal, animal, and plant samples.8 However, there is no efficient detection method for bacterial glucosyltransferases relating to the treatment of oral caries and for the high-throughput screening of inhibitors.
In the current work, a naphthalimide derivate was designed for the sensitive and selective real-time detection and imaging of bacterial glucosyltransferases. Furthermore, a high-throughput screening method has been established suitable for evaluating new GTFs inhibitors, suitable for the effective prevention, and treatment of dental caries.
As shown in Fig. 1, with uridine-diphosphate glucose (UDPG) as the glucosyl donor, N-phenethyl-4-hydroxy-1,8-naphthalimide (PENA) could accept the glucosyl group with the formation of 4-O-β-D-glucopyranosyl-N-phenethyl-1,8-naphthalimide (PENA-G) mediated by GTFs. Both PENA and PENA-G display a strong fluorescence at 560 and 440 nm respectively (Fig. S1, ESI†). Under UV light (365 nm), the yellow PENA solution and blue PENA-G solution are observed respectively. The glucosylation of PENA mediated by GTFs was confirmed by HPLC-DAD (Fig. S2, ESI†), indicating the production of PENA-G with an excellent conversion rate. In silico molecular docking analysis has been performed with homeotic glucosyltransferase, which revealed PENA as a suitable substrate of GTFs, and amino acid residues ASN481, ASP477, and ARG475 of the GTFs as the key active sites are consistent with those reported previously (Fig. 1b).9
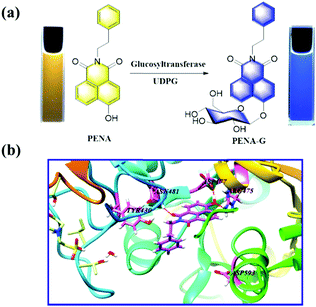 | ||
| Fig. 1 (a) Glucosylation of PENA by GTFs in the presence of UDPG (300 μM). (b) In silico docking analysis for the interaction between PENA and GTFs molecule. | ||
The fluorescence response of PENA toward GTFs with different GTFs concentrations (0–32 μg mL−1) and incubation time (0–80 min) were investigated, and a good linear relationship was observed for the fluorescence ratio (I440/I560) and GTFs concentration, incubation time respectively (Fig. S3 and S4, ESI†). Therefore, the positive correlation between fluorescence ratio and GTFs concentration could be used to determine the GTFs activity. In addition, the pH of the phosphate buffer and temperature of incubation were varied in order to optimize the yield of PENA-G (Fig. S5–S7, ESI†).
We then set out to use this fluorescence method to detect endogenous GTFs, and the suitability of various additives (amino acids, metal ions, and biological enzymes) as interferences for the glucosylation of PENA. It was obvious that common amino acids and metal ions could not influence the fluorescence response of PENA toward GTFs (Fig. S8, ESI†). Moreover, PENA was glucosylated selectively by GTFs in the presence of several biological enzymes including several common glycosidases (glucuronidases, UGT1A7, UGT1A10, UGT2B10, and UGT2B11), hydrolases (CES1b, CES1c, CES2, and DPP8), and bioactive proteins BSA and HSA (Fig. S9a, ESI†). The enzyme kinetics for the glucosylation of PENA were then investigated, and displayed Michaelis–Menten enzymatic behavior with the Vmax 10.01 μmol min−1 mg−1 and Km 8.7 μM (Fig. S9b, ESI†).
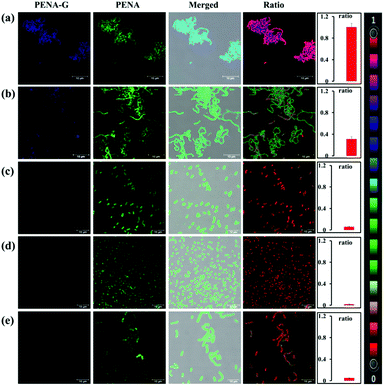
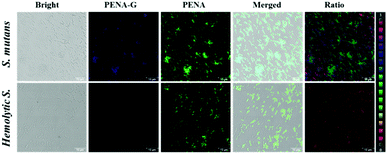
It has previously been reported that the levels of GTFs expressed in various bacteria are closely related to the cariogenicity of pathogenic bacteria. Thus, it is desirable to sense and image endogenous GTFs in various microorganisms, which could then be used as an efficient approach to analysis and detect the cariogenicity of bacteria. In this work, PENA was incubated with seven bacterial strains, including Streptococcus mutans, Hemolytic streptococcus, Lactobacillus amylovorus, Streptococcus pasteurianus strain 080205, Bacillus cereus 994000168 LBK, Escherichia coli DH5alpha BRL, Staphylococcus aureus ssp. aureus DSM 3463, to obtain fluorescence images (Fig. 2, Fig. S10 and S11, ESI†). From which it was obvious that Streptococcus mutans expressed the highest level of GTFs (ratio = 0.99), indicating the greatest potential cariogenicity and highlighting the key role of Streptococcus mutans as the pathogenic bacterium of dental caries. Moderate GTFs were observed for Hemolytic streptococcus with a ratio value of approximately 0.31. However, minimal GTFs were expressed in other bacteria. Therefore, PENA was successfully applied to sense and image endogenous GTFs in bacteria.
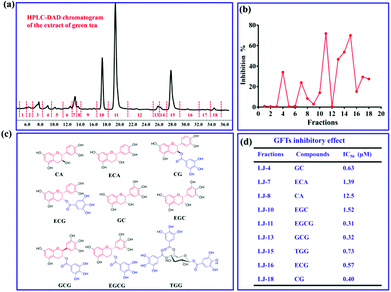
Bacterial GTFs as key virulence factor for the formation of dental caries, are suitable drug targets for the prevention, diagnosis, and therapy of dental caries. Using the current fluorescent probe PENA, a high-throughput screening method for GTFs inhibitors was established. It is well known that there are various toothpastes containing extracts of natural products, such as green tea, to prevent the formation of dental caries. Clearly indicating that green tea contains bioactive constituents for the effective prevention of dental caries. In order to discover new GTFs inhibitors, extracts from green tea were separated into 18 fractions using prep-HPLC (Fig. 3a). The inhibitory effects on GTFs for these fractions was then evaluated using a fluorescence high throughput screening method with PENA, and is shown in Fig. 3b. In particular, fraction 11 displayed 70% inhibition against GTFs at 500 μg mL−1. The major compounds within these bioactive fractions were then isolated using chromatographic techniques, and their structures were determined using 1H, 13C NMR and MS data (Fig. 3c). All of the compounds isolated were polyphenols, and most of them could be classified as catechin derivatives except for compound 1,4,6-tri-O-galloyl-β-D-glucose (TGG). The inhibitory effects of these isolated compounds on GTFs were evaluated, and indicate that gallocatechin gallate (GCG, IC50 0.32 μM) and epigallocatechin gallate (EGCG, IC50 0.31 μM) could be used as potential inhibitors of GTFs.
For the inhibitors EGCG and GCG, the inhibitory kinetics have been investigated to determine the inhibition types and Ki values (Fig. S21 and S22, ESI†). As shown in Fig. 4a and c, the intersection points were located in the second quadrant in the Dixon plot, which indicates competitive kinetics for both EGCG and GCG. In silico docking analysis was performed to investigate the preliminary inhibitory mechanism of polyphenols EGCG and GCG toward GTFs. The molecular docking experiments indicated that both EGCG and GCG could interact with the active pocket of GTFs, which overlapped with that of PENA, and several hydrogen bonds were formed between the phenolic hydroxyls and the amino acid residues of GTFs, such as ASP909, ASP593, ASP477, and TYR430. (Fig. 4b and d) So, the multiple phenolic hydroxyls, especial gallic acyls played key roles for the GTFs inhibition. The Ki values were determined for EGCG and GCG as 0.15 and 0.2 μM, respectively. Therefore, using the present fluorescent probe PENA, a high throughput screening system for the discovery of GTFs inhibitors has been established. In addition, two potential GTFs inhibitors extracted from green tea have been determined.
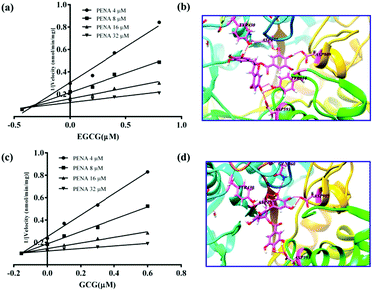 | ||
| Fig. 4 The inhibitory kinetics and in silico docking analysis of potential GTFs inhibitors EGCG (a and b) and GCG (c and d). | ||
As mentioned above, the catechin polyphenol EGCG displayed potential inhibitory effects against GTFs with competitive inhibition. The inhibitory effect on endogenous GTFs in Streptococcus genus bacteria was evaluated using PENA-G as the sensing product by confocal laser scanning microscopy (Fig. 5). For the cultures of Streptococcus mutans and Hemolytic streptococcus, EGCG (50 μM) and PENA (50 μM) were added with a co-incubation of 1 h. Then, the ratiometric fluorescence images were obtained respectively. The ratio values of Streptococcus mutans (ratio = 0.46) and Hemolytic streptococcus (ratio = 0.04) indicated a decreased yield of PENA-G and significant inhibitory effect of EGCG. Thus, EGCG isolated from green tea not only inhibited GTFs activity in vitro but also inhibited bacterial GTFs efficiently, offering great potential as new agent for the prevention of dental caries. The present inhibition experiment also confirmed the selective characteristic of the sensing of endogenous GTFs by the fluorescent probe PENA.
In summary, Streptococcus mutans together with over expressed GTFs play a key role for the formation of dental caries. An efficient method to sense the activity of GTFs as well as potential inhibitors is urgently needed. In the present work, a fluorescent naphthalimide derivate (PENA) was successfully designed as a ratiometric fluorescent probe for sensing and imaging endogenous GTFs in Streptococcus mutans. We demonstrated that PENA could be used as a high-throughput screening system for the investigation of GTFs inhibitors. Several bioactive compounds extracted from green tea were evaluated and EGCG and GCG were identified as potential inhibitors of GTFs. Their competitive inhibition kinetics, Ki values and inhibitory mechanisms were fully characterized. Our investigation provides a useful molecular tool for real-time sensing of endogenous GTFs in living pathogenic bacteria, and our high throughput screening approach has identified potential inhibitors for GTFs that could eventually lead to suitable oral treatments to prevent the development of dental caries.
This work was financially supported by National Natural Science Foundation of China (No. 81622047, 81872970 and 81503240), and State Key Laboratory of Fine Chemicals (KF1803, KF1705), and Program for High-level Talents of Dalian City (2017RQ119). TDJ wishes to thank the Royal Society for a Wolfson Research Merit Award.
Conflicts of interest
There are no conflicts to declare.Notes and references
- Z. Ren, T. Cui, J. M. Zeng, L. L. Chen, W. L. Zhang, X. Xu, L. Cheng, M. Y. Li, J. Y. Li, X. D. Zhou and Y. Q. Li, Antimicrob. Agents Chemother., 2016, 60, 126–135 CrossRef CAS PubMed.
- (a) K. Ito, S. Ito, T. Shimamura, S. Weyand, Y. Kawarasaki, T. Misaka, K. Abe, T. Kobayashi, A. D. Cameron and S. Iwata, J. Mol. Biol., 2011, 408, 177–186 CrossRef CAS PubMed; (b) A. Yanagida, M. Isozaki, Y. Shibusawa, H. Shindo and Y. Ito, J. Chromatogr. B: Anal. Technol. Biomed. Life Sci., 2004, 805, 155–160 CrossRef CAS PubMed.
- H. S. Kho, A. M. V. Smith, H. Koo, K. Scott-Anne and W. H. Bowen, Caries Res., 2005, 39, 411–416 CrossRef PubMed.
- T. Hoshino, Y. Kondo, K. Saito, Y. Terao, N. Okahashi, S. Kawabata and T. Fujiwara, Clin. Vaccine Immunol., 2011, 18, 1552–1561 CrossRef CAS PubMed.
- (a) T. Liu, Q. L. Yan, L. Feng, X. C. Ma, X. G. Tian, Z. L. Yu, J. Ning, X. K. Huo, C. P. Sun, C. Wang and J. N. Cui, Anal. Chem., 2018, 90, 9921–9928 CrossRef CAS PubMed; (b) L. Feng, Y. L. Yang, X. K. Huo, X. G. Tian, Y. J. Feng, H. W. Yuan, L. J. Zhao, C. Wang, P. Chu, F. D. Long, W. Wang and X. C. Ma, ACS Sens., 2018, 3, 1727–1734 CrossRef CAS PubMed; (c) T. Liu, J. Ning, B. Wang, B. Dong, S. Li, X. G. Tian, Z. L. Yu, Y. L. Peng, C. Wang, X. Y. Zhao, X. K. Huo, C. P. Sun, J. N. Cui, L. Feng and X. C. Ma, Anal. Chem., 2018, 90, 3965–3973 CrossRef CAS PubMed; (d) Y. Z. Jin, X. G. Tian, L. L. Jin, Y. L. Cui, T. Liu, Z. L. Yu, X. K. Huo, J. N. Cui, C. P. Sun, C. Wang, J. Ning, B. J. Zhang, L. Feng and X. C. Ma, Anal. Chem., 2018, 90, 3276–3283 CrossRef CAS PubMed; (e) J. J. Zhang, X. Z. Chai, X. P. He, H. J. Kim, J. Y. Yoon and H. Tian, Chem. Soc. Rev., 2019, 48, 683–722 RSC; (f) H. W. Liu, L. L. Chen, C. Y. Xu, Z. Li, H. Y. Zhang, X. B. Zhang and W. H. Tan, Chem. Soc. Rev., 2018, 47, 7140–7180 RSC; (g) J. Ning, T. Liu, P. Dong, W. Wang, G. Ge, B. Wang, Z. Yu, L. Shi, X. Tian, X. Huo, L. Feng, C. Wang, C. Sun, J. Cui, T. D. James and X. Ma, J. Am. Chem. Soc., 2019, 141, 1126–1134 CrossRef CAS PubMed.
- (a) S. J. Park, H. W. Lee, H. R. Kim, C. Kang and H. M. Kim, Chem. Sci., 2016, 7, 3703–3709 RSC; (b) S. M. Feng, Y. Fang, W. Y. Feng, Q. F. Xia and G. Q. Feng, Dyes Pigm., 2017, 146, 103–111 CrossRef CAS; (c) X. K. Huo, X. G. Tian, Y. N. Li, L. Feng, Y. L. Cui, C. Wang, J. N. Cui, C. P. Sun, K. X. Liu and X. C. Ma, Sens. Actuators, B, 2018, 262, 508–515 CrossRef CAS; (d) X. Han, X. Song, F. Yu and L. Chen, Chem. Sci., 2017, 8, 6991–7002 RSC; (e) H. D. Li, Y. Q. Li, Q. C. Yao, J. L. Fan, W. Sun, S. Long, K. Shao, J. J. Du, J. Y. Wang and X. J. Peng, Chem. Sci., 2019, 10, 1619–1625 RSC.
- (a) X. F. Wu, L. H. Li, W. Shi, Q. Y. Gong, X. H. Li and H. M. Ma, Anal. Chem., 2016, 88, 1440–1446 CrossRef CAS PubMed; (b) B. C. Zhu, C. C. Gao, Y. Z. Zhao, C. Y. Liu, Y. M. Li, Q. Wei, Z. M. Ma, B. Du and X. L. Zhang, Chem. Commun., 2011, 47, 8656–8658 RSC; (c) R. M. Duke, E. B. Veale, F. M. Pfeffer, P. E. Kruger and T. Gunnlaugsson, Chem. Soc. Rev., 2010, 39, 3936–3953 RSC; (d) X. F. Hou, Q. X. Yu, F. Zeng, C. M. Yu and S. Z. Wu, Chem. Commun., 2014, 50, 3417–3420 RSC; (e) Z. R. Dai, L. Feng, Q. Jin, H. L. Cheng, Y. Li, J. Ning, Y. Yu, G. B. Ge, J. N. Cui and L. Yang, Chem. Sci., 2017, 8, 2795–2803 RSC; (f) X. X. Zhang, H. Wu, P. Li, Z. J. Qu, M. Q. Tan and K. L. Han, Chem. Commun., 2016, 52, 9470 RSC.
- (a) X. Lv, L. Feng, C. Z. Ai, J. Hou, P. Wang, L. W. Zou, J. Cheng, G. B. Ge, J. N. Cui and L. Yang, J. Med. Chem., 2017, 60, 9664–9675 CrossRef CAS PubMed; (b) L. Feng, P. Li, J. Hou, Y. L. Cui, X. G. Tian, Z. L. Yu, J. N. Cui, C. Wang, X. K. Huo, J. Ning and X. C. Ma, Anal. Chem., 2018, 90, 13341–13347 CrossRef CAS PubMed; (c) L. Feng, X. Tian, Z. Yu, X. Zhao, C. Sun, J. Cui, J. Ning, C. Wang, B. Zhang and X. Ma, Sens. Actuators, B, 2019, 282, 112–121 CrossRef CAS.
- K. Ito, S. Ito, T. Shimamura, S. Weyand, Y. Kawarasaki, T. Misaka, K. Abe, T. Kobayashi, A. D. Cameron and S. Iwata, J. Mol. Biol., 2011, 408, 177–186 CrossRef CAS PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c9cc00440h |
| ‡ These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2019 |