A new class of supramolecular ligands stabilizes 14-3-3 protein–protein interactions by up to two orders of magnitude†
A.
Gigante
 a,
J.-N.
Grad
a,
J.-N.
Grad
 b,
J.
Briels
ac,
M.
Bartel
ac,
D.
Hoffmann
b,
J.
Briels
ac,
M.
Bartel
ac,
D.
Hoffmann
 *b,
C.
Ottmann
*b,
C.
Ottmann
 *ac and
C.
Schmuck
*ac and
C.
Schmuck
 *a
*a
aInstitute of Organic Chemistry, University of Duisburg Essen, Universitätstr. 7, 45141, Essen, Germany. E-mail: carsten.schmuck@uni-due.de; christian.ottmann@uni-due.de
bBioinformatics and Computational Biophysics, University of Duisburg Essen, Universitätstr. 7, 45141, Essen, Germany. E-mail: daniel.hoffmann@uni-due.de
cDepartment of Biomedical Engineering and Institute for Complex Molecular Systems, Technical University Eindhoven, P.O. Box 513, 5600 MB Eindhoven, The Netherlands. E-mail: c.ottmann@tue.nl
First published on 26th November 2018
Abstract
We report the first supramolecular stabilizers of the interaction between 14-3-3ζ and two of its effectors, Tau and C-Raf, which are involved in neurodegenerative diseases and proliferative signal transduction, respectively. These supramolecular ligands open up an opportunity to modulate functions of 14-3-3 with these effectors.
Due to the key role of protein–protein interactions (PPIs) in the homeostasis of biological systems, their dysfunction is involved in many pathologies and diseases like cancer, neurodegeneration or viral/parasitic infections.1,2 In the last years a number of successful examples of molecules that are able to modulate PPIs have been carried on into the clinic.3,4 Thus, protein surface recognition has become a promising tool to modulate protein function by disrupting or stabilizing PPIs.5
The human 14-3-3 protein family is one of the regulatory elements in intracellular signalling pathways which recognize Ser/Thr-phosphorylated proteins like those implicated in the mitogen-activated protein kinase (MAPK) pathway.6,7 To date approx. 300 potential physiological effectors of this family have been identified.8,9 Currently, only a few small molecules have been shown to stabilize PPIs of some 14-3-3/effector complexes.10–13 In all these studies the described effectors present the length that is involved in the interaction and not the full sequence. The first reported examples were the fungal toxin Fusicoccin12 and the plant growth regulator Cotylenin-A,13 which promote the interaction of 14-3-3 proteins with the human potassium channel Task314,15 and the proto-oncogene C-Raf,13 respectively. Specifically, Fusicoccin showed an EC50 value of 2 μM for the stabilization of 14-3-3/Task3 complex, while cotylenin-A is less effective with an EC50 of 65 μM for the 14-3-3/C-Raf interaction. More recently, new fusicoccin derivatives (fusicoccanes)12 and trisubstituted pyrrolidinones (as fusicoccin mimetics)16,17 have been described as result of different PPIs stabilizer optimization programs. However, their EC50 values for stabilization were similar to those of the natural compounds.
Currently more than 100 crystal structures of 14-3-3 proteins with different binding partners and also with their respective inhibitors or stabilizers are available in the PDB18 (see Table S1 in the ESI†). For example, by analysing the crystal structure of the complex formed by 14-3-3 and C-Raf with cotylenin-A, we observed that the natural product forms two hydrogen bonds with Asp213.12 In addition, around the central binding channel of 14-3-3 other surface-exposed negatively charged residues are located. Therefore, the design of positively charged compounds that could bind specifically to this anionic region in the vicinity of the central binding channel of 14-3-3 could lead to the modulation (inhibition or stabilization of PPI) of these 14-3-3 interactions with different effectors (Fig. 1).
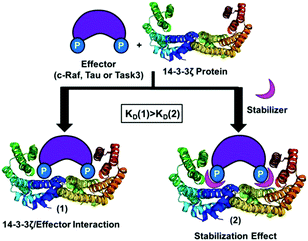 | ||
| Fig. 1 Stabilizing effect between14-3-3ζ protein (PDB id: 4JDD) and a binding partner (C-Raf, Tau or Task3). | ||
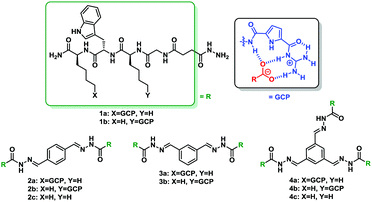
To identify molecules that could modulate these physiologically highly relevant 14-3-3 PPIs, we performed a screening of cationic supramolecular ligands originating from our home-made library. For the screening we selected ligands which contain the guanidiniocarbonyl pyrrole aka GCP moiety, an arginine mimetic developed by our group.21 It forms very stable ion pairs with oxyanions such as carboxylates and phosphates significantly more stable than simple guanidinium cations or ammonium ions (as in the natural amino acids arginine or lysine) (Table 1).19–23 Hence we usually incorporate this binding motif to improve the affinity of peptide-based ligands for proteins. In the search for stabilizers of 14-3-3 protein–protein interactions fluorescence polarization (FP) measurements were carried out, employing 14-3-3ζ/effector systems in which the complex had not formed yet (see Fig. S1, ESI†). The binding partners were fluorescein-labelled, monophosphorylated synthetic peptides comprising the phosphorylation sites C-RafpS259 and Task3pS373, as well as the diphosphorylated peptide TaupS214pS324. Binding of these peptides to 14-3-3ζ in the presence and absence of molecules from our library was tested. Several ligands containing the GCP group were identified as new stabilizers of 14-3-3ζ/effector complexes with the best results for C-Raf and Tau binding partners (Table 1).
| Cmpds | C-Raf-EC50a (μM) | Tau-EC50 (μM) | Task3-EC50 (μM) |
|---|---|---|---|
| a 50% effective concentration or concentration of these compounds which is required to stabilize 50% of the interaction between 14-3-3ζ and these effectors. | |||
| 1a | >1000 | >1000 | >1000 |
| 2a | 50.0 ± 4.1 | 89.0 ± 9.9 | 150.0 ± 7.3 |
| 3a | 47.0 ± 2.9 | >1000 | >300 |
| 4a | 8.4 ± 0.1 | 1.4 ± 0.2 | >200 |
| 1b | >1000 | >1000 | >1000 |
| 2b | >1000 | 22.9 ± 3.9 | >300 |
| 3b | 55.6 ± 0.5 | 14.5 ± 0.5 | 147.0 ± 9.0 |
| 4b | 30.9 ± 2.1 | 4.7 ± 2.3 | >200 |
| 2c | >1000 | >200 | >600 |
| 4c | >1000 | >200 | >1000 |
First, these data highlight the importance of the arginine mimetic in these ligands for the stabilization effect. Compounds which do not have a GCP group, such as 2c and 4c, do not show stabilization of the interaction between 14-3-3ζ protein with C-Raf, Tau and Task3; while their corresponding counterparts which have a GCP moiety, such as 2a, 2b, 4a and 4b, show EC50 values in the lower micromolar range. Second, the multivalency of the compounds seems to correlate with their PPI-stabilizing activity. For example, 4a which has three arms containing one GCP group each increases the stability of 14-3-3ζ/C-Raf complex to a larger extent compared to 3a or 2a which only have two arms. In addition, the latter substances show a higher stabilization effect on this interaction than 1a which possesses a single arm. The same behavior is noted for compounds belonging to series b and also for both series with the 14-3-3ζ partner peptide derived from Tau. Moreover, in terms of selectivity, compound 3a is noteworthy since it stabilizes specifically the 14-3-3ζ/C-Raf complex with an EC50 value of 47.0 ± 2.0 μM without showing an effect on 14-3-3 binding to the peptides derived from Tau and Task3. However, 4a and 4b are the most potent compounds of this family, being the first synthetic stabilizers for Tau peptide to 14-3-3ζ and being more potent in comparison with the described Cotylenin A for 14-3-3 ζ/C-Raf interaction, which stabilized the complex with an EC50 of around 65 μM.15 Also the same assay was carried out in absence of 14-3-3ζ protein for the latest compounds, as a control experiment, to ensure that these data are not a response of the binding of these ligands with C-Raf or Tau effectors (see Fig. S3, ESI†). As a result 4a and 4b did not bind to C-Raf or Tau.
Subsequently, we focused on the 14-3-3ζ interactions with the peptide derivates from C-Raf and Tau and determined the apparent KD values in the absence and presence of our compounds (Table 2). In the presence of 50 μM of these multi-armed compounds we observed an enhancement of the stability of these interactions as reflected in lower apparent KD values. In line with the EC50 data the apparent affinity confirmed the multivalency effect of this family in stabilizing the interactions. Accordingly, compounds with three arms showed a higher stabilization than compounds with two arms, while ligands with one arm were ineffective. 4a, the most potent compound, enhanced the interaction of 14-3-3ζ with C-Raf and Tau approximately 84-fold and 26-fold, respectively, from a KD of 16.00 ± 0.60 μM to 0.19 ± 0.01 μM for the binding partner C-Raf and from a KD of 7.00 ± 0.20 μM to 0.27 ± 0.01 μM for Tau. Similarly, 4b increased the stability of these interactions, 56-fold in the case of C-Raf and 20-fold for Tau, from a KD of 8.90 ± 0.50 μM to 0.16 ± 0.01 μM for the binding partner C-Raf and from a KD of 3.50 ± 0.10 μM to 0.17 ± 0.01 μM for Tau.
| Cmpds | C-Raf: KD app (μM) | IS | Tau: KD app (μM) | IS |
|---|---|---|---|---|
| Control | 16.00 ± 0.60 | — | 7.00 ± 0.20 | — |
| 8.90 ± 0.50a | 3.50 ± 0.10a | |||
| a Experiments carried out in a different time and with different batch of protein, effectors and ligands. | ||||
| 1a | 14.40 ± 0.60 | ×1 | 5.90 ± 0.10 | ×1 |
| 2a | 3.70 ± 0.40 | ×4 | 0.56 ± 0.02 | ×12 |
| 3a | 7.30 ± 0.20 | ×2 | 1.35 ± 0.03 | ×5 |
| 4a | 0.19 ± 0.01 | ×84 | 0.27 ± 0.01 | ×26 |
| 1b | 5.70 ± 0.30a | ×1.6 | 2.30 ± 0.10a | ×1.5 |
| 2b | 4.50 ± 0.20a | ×2 | 0.49 ± 0.01 | ×14 |
| 3b | 1.20 ± 0.03 | ×13 | 0.37 ± 0.01 | ×18 |
| 4b | 0.16 ± 0.01a | ×56 | 0.17 ± 0.01a | ×20 |
These observations regarding 4b were supported by isothermal titration calorimetry (ITC). The observed KD decreased from 17.04 to 0.36 μM for the complex 14-3-3ζ/C-Raf in the presence of 4b, enhancing the stabilization 47-fold (Fig. 2). In the case of Tau the KD decreases from 11.65 to 0.60 μM increasing the stabilization 20-fold (see Fig. S6, ESI†).
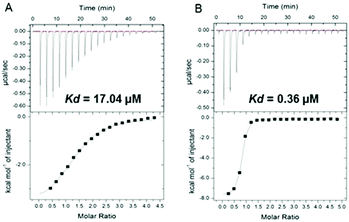 | ||
| Fig. 2 (A) Isothermal titration calorimetry of binding C-Raf to 14-3-3ζ and (B) the same interaction in the presence of 50 μM of 4b compound. | ||
Then, we determined by microscale thermophoresis (MST) the binding constants of some of these ligands with 14-3-3ζ (see Fig. S7 and S8, ESI†). The KD values of 4b, 4a and 3a are 5 μM, 22 μM and 41 μM, respectively. However, 1a did not show binding to 14-3-3ζ. These results explain why the 3-arm ligands stabilize these interactions better, as their affinity for this protein is higher than its 2-arm counterpart. Conversely, the single arm 1a does not bind and therefore cannot stabilize. Besides, it is interesting that although the affinity of 4b is higher than 4a, 4a stabilizes the C-Raf interaction more. This could be because its affinity for this complex is higher than in the case of 4b.
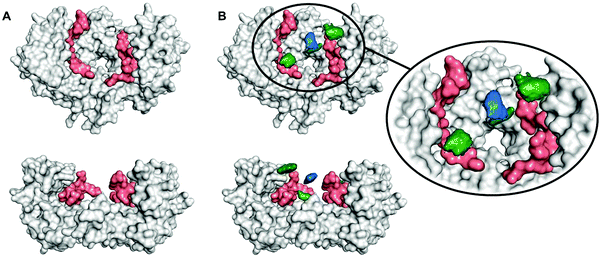
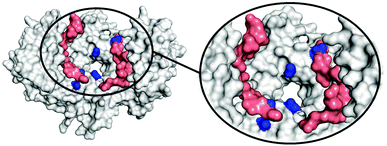
To determine the region of the protein where these compounds most likely bind, molecular dynamics (MD) simulations were carried out using a procedure detailed previously.27 We studied the complex of 14-3-3ζ/C-Raf (Fig. 3A) in the presence of 4b (Fig. 3B). However, the flexibility of this molecular system precluded the identification of a single binding pose. Instead, we monitored the volumes explored by the peptidic arms containing the GCP moiety and the central benzene ring in six MD simulations. An exemplary MD simulation is represented in Fig. 3B (see Fig. S9 for all MDs, ESI†). The 3-armed ligand 4b binds indeed to both the protein (grey surface) and one C-Raf peptide (salmon surface). The central benzene ring of 4b (blue volume) and one arm (green volume under the blue volume) are located close to the pore of the 14-3-3ζ dimer attaching the ligand to the protein. Another arm of the ligand binds to the C-Raf peptide's C-terminal carboxylic acid (salmon, bottom left corner in the zoomed region), while the third arm is not involved in a salt bridge. The six simulations consistently show the ligand anchoring itself to the protein electrostatically and preventing C-Raf release through steric effects. This is due to the GCP units in ligand 4b that interact more frequently than Lys units with the anionic Asp/Glu side-chains around the central pore and in regions where the C-Raf peptides also bind (see Fig. S10A, ESI†). The acidic residues most frequently engaged in salt bridges with GCP according to these MD simulations are marked in dark blue in Fig. 4. This binding pattern is fully consistent with the observed stabilization of the 14-3-3/C-Raf interaction by our ligand.
Even though the third arm in 4b does not form stable salt bridges compared to the other two arms (see Fig. S10B, ESI†), its presence statistically increases the chances of forming salt bridges during the initial binding, and during unbinding/rebinding events (positive cooperative binding). This could be the reason for the higher affinity of the 3-armed ligand compared to that of the 2-armed ligand.
In summary, we identified a new and potent family of stabilizers of the interaction between 14-3-3ζ and C-Raf or Tau with EC50 values in the low micromolar range. Ligands with a higher level of multivalency such as 4a and 4b (three arms) are more potent stabilizers for this PPI as shown by both FP measurements and ITC data. The interaction of the protein with the effector peptides is enhanced by nearly two orders of magnitude in the best case. In addition, the area where 4b most likely is located in the 14-3-3ζ/C-Raf complex was identified in silico by Epitopsy, followed by MD to sample the binding events with atomistic resolution, providing a consistent explanation of the observed stabilization. These results should now allow further modifications of this new class of stabilizers to obtain even more potent and selective compounds. A clear advantage of artificial ligands is the much higher synthetic accessibility23–26 over highly complex natural products which have predominantly been described as PPI stabilizers. To what extent the advantage of natural product PPI stabilizers – cell availability and selectivity – can be achieved with our supramolecular ligands will be the subject of future studies.
Conflicts of interest
There are no conflicts to declare.Notes and references
- D. C. Fry and L. T. Vassilev, J. Mol. Med., 2005, 83, 955–963 CrossRef CAS PubMed
.
- D. P. Ryan and J. M. Matthews, Curr. Opin. Struct. Biol., 2005, 15, 441–466 CrossRef CAS
.
- J.-C. Carry and C. Garcia-Echeverria, Bioorg. Med. Chem. Lett., 2013, 23, 2480–2485 CrossRef CAS
.
- G. Fischer, M. Rossmann and M. Hyvönen, Curr. Opin. Biotechnol., 2015, 35, 78–85 CrossRef CAS
.
- M. Aeluri, S. Chamakuri, B. Dasari, S. K. R. Guduru, R. Jimmidi, S. Jogula and P. Arya, Chem. Rev., 2014, 114, 4640–4694 CrossRef CAS
.
- D. Berg, C. Holzmann and O. Riess, Nat. Rev. Neurosci., 2003, 4, 752–762 CrossRef CAS
.
- X. Yang, W. H. Lee, F. Sobott, E. Papagrigoriou, C. V. Robinson, J. G. Grossmann, M. Sundström, D. A. Doyle and J. M. Elkins, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 17237–17242 CrossRef CAS
.
- G. P. H. van Heusden, IUBMB Life, 2005, 57, 623–629 CrossRef CAS PubMed
.
- C. Johnson, S. Crowther, M. J. Stafford, D. G. Campbell, R. Toth and C. MacKintosh, Biochem. J., 2010, 427, 69–78 CrossRef CAS PubMed
.
- A. Kaplan, B. Morquette, A. Kroner, S. Leong, C. Madwar, R. Sanz, S. L. Banerjee, J. Antel, N. Bisson, S. David and A. E. Fournier, Neuron, 2017, 93, 1082–1093 CrossRef CAS
.
- M. Mori, G. Vignaroli and M. Botta, Drug Discovery Today: Technol., 2013, 10, e541–e547 CrossRef
.
- R. G. Doveston, A. Kuusk, S. A. Andrei, S. Leysen, Q. Cao, M. P. Castaldi, A. Hendricks, L. Brunsveld, H. Chen, H. Boyd and C. Ottmann, FEBS Lett., 2017, 591, 2449–2457 CrossRef CAS
.
- L. M. Stevers, E. Sijbesma, M. Botta, C. MacKintosh, T. Obsil, I. Landrieu, Y. Cau, A. J. Wilson, A. Karawajczyk, J. Eickhoff, J. Davis, M. Hann, G. O’Mahony, R. G. Doveston, L. Brunsveld and C. Ottmann, J. Med. Chem., 2018, 61, 3755–3778 CrossRef CAS
.
- C. Anders, Y. Higuchi, K. Koschinsky, M. Bartel, B. Schumacher, P. Thiel, H. Nitta, R. Preisig-Müller, G. Schlichthörl, V. Renigunta, J. Ohkanda, J. Daut, N. Kato and C. Ottmann, Chem. Biol., 2013, 20, 583–593 CrossRef CAS
.
- M. Molzan, S. Kasper, L. Röglin, M. Skwarczynska, T. Sassa, T. Inoue, F. Breitenbuecher, J. Ohkanda, N. Kato, M. Schuler and C. Ottmann, ACS Chem. Biol., 2013, 8, 1869–1875 CrossRef CAS
.
- S. Bittner, T. Budde, H. Wiendl and S. G. Meuth, Brain Pathol., 2010, 20, 999–1009 CrossRef CAS
.
- M. Kilisch, O. Lytovchenko, B. Schwappach, V. Renigunta and J. Daut, Pflügers Arch. Eur. J. Physiol., 2015, 467, 1105–1120 CrossRef CAS
.
- R. Rose, S. Erdmann, S. Bovens, A. Wolf, M. Rose, S. Hennig, H. Waldmann and C. Ottmann, Angew. Chem., 2010, 49, 4129–4132 CrossRef CAS
.
- A. Richter, R. Rose, C. Hedberg, H. Waldmann and C. Ottmann, Chem, 2012, 18, 6520–6527 CrossRef CAS
.
- PDB: 5J31, 5EWZ, 5EXA, 5D2D, 5D3F, 4ZDR, 4WRQ, 4N7G, 4N7Y, 4N84, 4IEA, 4IHL… (see ESI†).
- M. Ehlers, J.-N. Grad, S. Mittal, D. Bier, M. Mertel, L. Ohl, M. Bartel, J. Briels, M. Heimann, C. Ottmann, E. Sanchez-Garcia, D. Hoffmann and C. Schmuck, ChemBioChem, 2017, 19, 591–595 CrossRef
.
- C. Schmuck, Coord. Chem. Rev., 2006, 250, 3053–3067 CrossRef CAS
.
- Q. Q. Jiang, L. Bartsch, W. Sicking, P. R. Wich, D. Heider, D. Hoffmann and C. Schmuck, Org. Biomol. Chem., 2013, 11, 1631–1639 RSC
.
- Q. Q. Jiang, W. Sicking, M. Ehlers and C. Schmuck, Chem. Sci., 2015, 6, 1792–1800 RSC
.
- M. Li, M. Ehlers, S. Schlesiger, E. Zellermann, S. K. Knauer and C. Schmuck, Angew. Chem., 2016, 55, 598–601 CrossRef CAS
.
- M. Li, S. Schlesiger, S. K. Knauer and C. Schmuck, Angew. Chem., 2015, 54, 2941–2944 CrossRef CAS
.
- J.-N. Grad, A. Gigante, C. Wilms, J. N. Dybowski, L. Ohl, C. Ottmann, C. Schmuck and D. Hoffmann, J. Chem. Inf. Model., 2018, 58, 315–327 CrossRef CAS
.
Footnote |
| † Electronic supplementary information (ESI) available: Details of the synthetic procedures and characterization data, as well as details of FP and ITC measurements, and description of the molecular dynamic simulations. See DOI: 10.1039/c8cc07946c |
| This journal is © The Royal Society of Chemistry 2019 |