Highly sensitive optical biosensing of Staphylococcus aureus with an antibody/metal–organic framework bioconjugate
Abstract
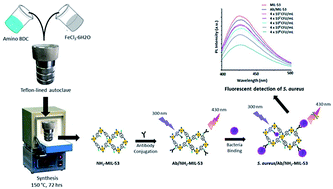
In this research, a new luminescent bioprobe was developed for the detection of S. aureus based on bio-conjugation of an amine functionalized metal–organic framework (NH2-MIL-53(Fe)) with an anti-S. aureus antibody (Ab). The formation of the desired bioprobe (Ab/NH2-MIL-53), in its liquid phase, has been verified with several spectroscopic and structural characterization techniques. The bioprobe was incubated with varying concentrations of S. aureus bacteria. The resulting antibody conjugated bioprobe (Ab/NH2-MIL-53) maintained a strong inverse correlation in which decreases in the fluorescence intensities were accompanied by an increase in the bacterial count. Thus, the potential of the herein developed probe has been successfully demonstrated for the detection of S. aureus with a low limit of detection (85 CFU mL−1) over a wide concentration range (4 × 102–4 × 108 CFU mL−1). It was further found to be reliable with regard to inter-/intra- precision assays and long-term stability. The feasibility of the method was further supported through the detection of S. aureus spiked in environmental samples (e.g., river water and cream pastry).


 Please wait while we load your content...
Please wait while we load your content...