Raman spectroscopy as a tool for tracking cyclopropane fatty acids in genetically engineered Saccharomyces cerevisiae†
Abstract
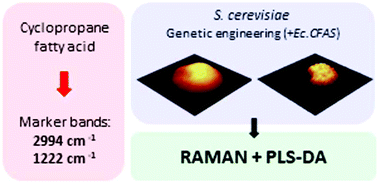
Cyclopropane fatty acids (CFAs) are a group of lipids with unique physical and chemical properties between those of saturated and monounsaturated fatty acids. The distinctive physicochemical characteristics of CFAs (e.g. oxidative stability, self-polymerization at high temperatures, etc.) results from the presence of a cyclopropane ring within their structure making them highly useful in industrial applications. CFAs are present in several species of plants and bacteria and are typically detected with standard lipid profiling techniques, such as gas or liquid chromatography. In this work we investigated several strains of S. cerevisiae, genetically modified to introduce the production of CFAs, in comparison to control strain using confocal Raman spectroscopy (CRS). The aim of our work was to demonstrate the potential of CRS not only to detect changes introduced due to the CFAs presence, but also to track CFAs within the cells. We present for the first time Raman and IR spectra of CFA standard (cis-9,10-methyleneoctadecanoic acid), completed with quantum chemical calculations and band assignment. We identified marker bands of CFA (e.g. 2992, 1222, 942 cm−1) attributed to the vibrations of the cyclopropyl ring. Furthermore, we analysed lipid bodies (LBs) from modified and control yeast using CRS imaging and identified multiple changes in size, number and composition of LBs from engineered strains. We observed a significant reduction in the degree of unsaturation of LBs using the ratio of bands located at 1660 cm−1 (ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C)) and 1448 cm−1 (δ(CH2)) in the modified cell lines. In addition, we were able to detect the presence of CFAs in LBs, using the established marker bands. CRS shows tremendous potential as technique to identify CFAs in lipid bodies providing a new way to track lipid production in genetically modified single yeast cells.
C)) and 1448 cm−1 (δ(CH2)) in the modified cell lines. In addition, we were able to detect the presence of CFAs in LBs, using the established marker bands. CRS shows tremendous potential as technique to identify CFAs in lipid bodies providing a new way to track lipid production in genetically modified single yeast cells.
- This article is part of the themed collection: Next wave advances in single cell analyses


 Please wait while we load your content...
Please wait while we load your content...