A new approach to find biomarkers in chronic fatigue syndrome/myalgic encephalomyelitis (CFS/ME) by single-cell Raman micro-spectroscopy†
Abstract
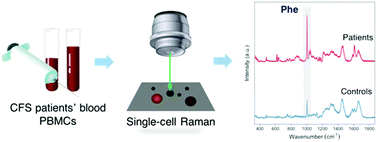
Chronic fatigue syndrome (CFS), also called myalgic encephalomyelitis (ME), is a debilitating disorder characterized by physical and mental exhaustion. Mitochondrial and energetic dysfunction has been investigated in CFS patients due to a hallmark relationship with fatigue; however, no consistent conclusion has yet been achieved. Single-cell Raman spectra (SCRS) are label-free biochemical profiles, indicating phenotypic fingerprints of single cells. In this study, we applied a new approach using single-cell Raman microspectroscopy (SCRM) to examine ρ0 cells that lack mitochondrial DNA (mtDNA), and peripheral blood mononuclear cells (PBMCs) from CFS patients and healthy controls. The experimental results show that Raman bands associated with phenylalanine in ρ0 cells and CFS patient PBMCs were significantly higher than those of the wild-type model and healthy controls. As similar changes were observed in the ρ0 cell model with a known deficiency in the mitochondrial respiratory chain as well as in CFS patients, our results suggest that the increase in cellular phenylalanine may be related to mitochondrial/energetic dysfunction in both systems. Interestingly, phenylalanine can be used as a potential biomarker for the diagnosis of CFS by SCRM. A machine learning classification model achieved an accuracy rate of 98% correctly assigning Raman spectra to either the CFS group or the control group. SCRM combined with a machine learning algorithm therefore has the potential to become a diagnostic tool for CFS.
- This article is part of the themed collection: Next wave advances in single cell analyses


 Please wait while we load your content...
Please wait while we load your content...