 Open Access Article
Open Access ArticleProbing the competition between duplex and G-quadruplex/i-motif structures using a conformation-sensitive fluorescent nucleoside probe†
Pramod M.
Sabale‡
a,
Arun A.
Tanpure‡
ab and
Seergazhi G.
Srivatsan
 *a
*a
aDepartment of Chemistry, Indian Institute of Science Education and Research (IISER), Pune, Dr. Homi Bhabha Road, Pune 411008, India. E-mail: srivatsan@iiserpune.ac.in
bDepartment of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW, UK
First published on 21st May 2018
Abstract
Double-stranded segments of a genome that can potentially form G-quadruplex (GQ) and/or i-motif (iM) structures are considered to be important regulatory elements. Hence, the development of a common probe that can detect GQ and iM structures and also distinguish them from a duplex structure will be highly useful in understanding the propensity of such segments to adopt duplex or non-canonical four-stranded structures. Here, we describe the utility of a conformation-sensitive fluorescent nucleoside analog, which was originally developed as a GQ sensor, in detecting the iM structures of C-rich DNA oligonucleotides (ONs). The analog is based on a 5-(benzofuran-2-yl)uracil scaffold, which when incorporated into C-rich ONs (e.g., telomeric repeats) fluorescently distinguishes an iM from random coil and duplex structures. Steady-state and time-resolved fluorescence techniques enabled the determination of transition pH for the transformation of a random coil to an iM structure. Furthermore, a qualitative understanding on the relative population of duplex and GQ/iM forms under physiological conditions could be gained by correlating the fluorescence, CD and thermal melting data. Taken together, this sensor could provide a general platform to profile double-stranded promoter regions in terms of their ability to adopt four-stranded structures, and also could support approaches to discover functional GQ and iM binders.
Introduction
Guanine- and cytosine-rich sequences, which are abundantly found in telomeres, promoter DNA regions and untranslated regions of mRNA of several proto-oncogenes, are known to form non-canonical four-stranded structures called G-quadruplexes (GQs) and i-motifs (iMs), respectively.1,2 The conservation of their position in various eukaryotes and recent biochemical investigations indicate that these structural motifs could play important roles in the regulation of essential cellular processes including replication, transcription and translation.3 Despite their colocalization in the genome, most studies have focused on understanding the structural and functional relevance of GQs, while iMs have received relatively less attention. The reason is twofold: (i) stable GQ structures can form under physiological conditions and iM structures usually form under mildly acidic conditions by the intercalation of hemiprotonated C·CH+ base-paired strands,4 and (ii) GQs have been identified in a cellular environment,5 whereas the existence of iMs in a cellular setting is yet to be unambiguously established. However, recent studies indicate that molecular crowding, negative superhelicity, ligand binding and certain modifications can induce the formation of the iM structure at physiological pH.3b,6 Several proteins including transcription factors have been identified to bind C-rich DNA sequences.7 For example, a factor named hnRNP LL preferentially binds to the BCL2 promoter iM-forming sequence and transcriptionally activates BCL2 expression.8 Furthermore, iM's role as a molecular switch in modulating oncogene expression has been demonstrated by using structure-specific small molecule binders.3b,9 These findings suggest that the iM structure could complement the GQ structure as a potential therapeutic target. In addition to their proposed biological roles, programmability and pH-dependent structure, switching properties have made iMs useful supramolecular synthons for designing functional nanodevices.10In terms of structure, like GQs, iMs exhibit different folding topologies and varying stability in vitro, which depend on the number of cytosine residues, loop length and pH.4a,11 Hence, it is reasonable to say that the formation of GQ and iM structures by G-rich and C-rich sequences coexisting in the double-stranded region of telomeres and promoters will depend on the relative stability of the duplex and GQ/iM forms.12 Studies also suggest that the balance between duplex and GQ/iM forms could be altered in the presence of protein factors and small molecule ligands that induce or stabilize GQ/iM structures.
Several biophysical techniques including circular dichroism (CD), fluorescence, NMR and X-ray crystallography have provided valuable information on the structure, stability, folding dynamics and recognition properties of individual G-rich and C-rich strands.13 Among these, fluorescence-based tools, which show changes in fluorescence properties (e.g., quantum yield, emission maximum and lifetime) during a conformational change, are advantageous as they not only enable the real-time monitoring of the formation of GQ and iM structures but also provide platforms to screen small molecule binders.13e,14 In particular, fluorescent nucleoside analogs incorporated into ONs offer efficient systems to study iM and GQ structures. Fluorescent purine surrogates (e.g., 6-methylisoxanthopterin and 2-aminopurine) and vinyl-, styryl- and heteroaryl-conjugated nucleoside analogs incorporated into G-rich ONs have been utilized in the study of DNA GQs.15 An exciplex signaling system made of a pair of pyrene-modified deoxyadenosines (PyA) has been used to monitor the pH-dependent structural transition from a random coil to an iM structure.16 In a similar strategy, photoinduced electron transfer in the iM structure has been studied by using pyrene- and anthraquinone-modified dU as a donor–acceptor pair.17 More recently, a novel “push–pull” fluorescent nucleoside analog, derived by fusing dimethylaniline to deoxycytidine (DMAC), has enabled the real-time tracking of the exchange of iM to duplex DNA.18 While the utility of these probes is undeniable, their implementation in assays to evaluate the propensity of double-stranded regions of the human genome to adopt iM/GQ structures and the competition between duplex and iM/GQ structures is a challenge.19 Therefore, we envisioned that the development of a conformation-sensitive fluorescent nucleoside analog, which (i) is structurally minimally perturbing, (ii) serves as both GQ and iM sensors, and importantly (iii) shows that the distinct fluorescence properties of GQ, iM and duplex forms will be highly useful in not only profiling various double-stranded regions of the human genome in terms of their ability to adopt duplex or GQ and iM structures, but also could facilitate setting up screening assays to identify efficient binders.
In this context, we recently introduced microenvironment-sensitive fluorescent 2′-deoxyuridine and uridine nucleoside analogs made of a 5-(benzofuran-2-yl)uracil core.20,21 These emissive analogs serve as excellent GQ sensors, and enable the photophysical discrimination of different GQ structures adopted by H-Telo DNA and RNA repeats. Furthermore, we devised a simple fluorescence assay using these probes to quantitatively estimate the topology- and nucleic acid-specific binding of ligands to GQ structures.22 Other groups have also used the conformation sensitivity of our probes to design assays to investigate the formation, stability and ligand binding ability of different RNA topologies.23 Encouraged by these key observations, we sought to evaluate the proficiency of a benzofuran-modified fluorescent nucleoside analog in not only detecting the formation of iM structures but also in distinguishing iM, GQ and duplex structures. Here, we describe the development of a fluorescence-based platform to detect the iM structures of C-rich DNA ONs by using the 5-benzofuran-modified 2′-deoxyuridine (1) analog. An emissive nucleoside incorporated into C-rich DNA ONs is minimally perturbing and fluorescently distinguishes the iM structure from random coil and duplex structures (Fig. 1). The iM probe also enabled the determination of transition pH (tpH) for the transformation of a random coil to an iM structure by both steady-state and time-resolved fluorescence techniques. Furthermore, the conformation-specific fluorescence readout of the nucleoside probe complemented by CD and thermal melting experiments provided a qualitative understanding of the relative population of duplex, GQ and iM forms in the G-rich−C-rich double-stranded region of the human telomeric (H-Telo) DNA ON repeat.
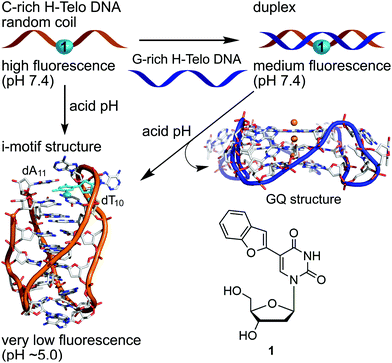 | ||
| Fig. 1 Schematic diagram showing the assay design to monitor the pH-induced transition from the random coil and duplex to iM structure by using a conformation-sensitive fluorescent nucleoside probe 1. The probe is placed in the loop region of C-rich DNA ONs. The iM structure of the C-rich H-Telo DNA ON repeat (PDB: 1EL2) is used as an example to illustrate the design (site of modification: second loop, T10 residue, cyan color).24 A random coil structure of C-rich H-Telo DNA ON shows high fluorescence compared to its perfect complementary duplex. Upon reducing the pH to ∼5.0, the random coil folds into an iM structure and exhibits very low fluorescence. The duplex can dissociate and form iM and GQ structures depending on the conditions (e.g., acidic pH) and sequence. | ||
Results and discussion
Synthesis of 5-benzofuran-2′-deoxyuridine-labeled C-rich DNA ONs
We chose to study two C-rich DNA ON sequences, (TCCCCC)4 and H-Telo (CCCTAA)4 repeats, which are known to adopt iM structures at near physiological pH or at slightly acidic pH (Fig. 2).11,13a Fluorescently labeled DNA ONs 2 and 3 were synthesized by using the phosphoramidite substrate of nucleoside analog 1. Purposely, we replaced one of the loop thymine residues with 1 as we envisioned that modification on cytosine residues could affect the efficiency of iM formation. The purity and integrity of the ONs synthesized by a solid-phase method were confirmed by HPLC and MALDI-TOF mass analyses (Fig. S1 and Table S1†).Fluorescence detection of the DNA iM structure
The ability of the nucleoside probe to report the transition from a random coil to an iM structure was first evaluated by monitoring the changes in the fluorescence of a model C-rich ON 2 as a function of pH. Solutions of ON 2 at different pH values (8.2 to 5.0) were prepared in phosphate or acetate buffer, which is commonly used to study iM formation at different pH values.13c,25 Upon exciting ON 2 at pH 8.2, a strong emission band centered at 442 nm was observed (Fig. 3A). As the pH was lowered from 8.2 to 5.0, ON 2 displayed a significant pH-dependent fluorescence quenching (∼23-fold at saturation) with no apparent change in emission maximum. The changes in fluorescence intensity as a function of pH followed a sigmoidal curve yielding a tpH of 7.13 ± 0.01, which is consistent with the cytosine content of the sequence (Fig. 3B, S2 and Table S2†).11 Furthermore, a discernible decrease in the excited-state lifetime of ON 2 was observed as the pH was lowered. A sigmoidal lifetime profile yielded a tpH of 6.92 ± 0.02, which is closer to the one obtained by steady-state analysis (Fig. 3B, S3, Tables S2 and S3†). To confirm whether the changes in fluorescence intensity and lifetime are indeed due to the formation of an iM structure, CD and UV-thermal melting profiles of emissive ON 2 and its control unmodified ON sequence 4 were recorded. At pH, 5.0 ONs 2 and 4 exhibited a characteristic CD profile for the iM structure with a strong positive peak at ∼285 nm and a negative peak at ∼264 nm (Fig. S4A†). Thermal melting experiments indicated that both modified and control unmodified DNA ONs formed stable iM structures with similar Tm values (Fig. S4C and Table S4†). The CD profiles and Tm values are consistent with literature reports.11,26 These results also indicate that benzofuran modification has a negligible impact on the iM structure and stability. Notably, at pH 7.0, a low fluorescence of ON 2 and the CD pattern largely resembling the iM form indicate that this sequence can adopt an iM structure at neutral pH (Fig. 3A and S4A†).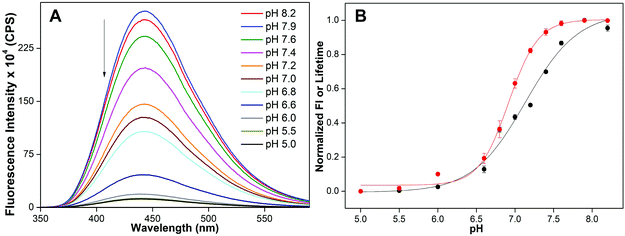 | ||
| Fig. 3 (A) Fluorescence spectra of 5-benzofuran-modified DNA ON 2 (1 μM) at different pH values. Samples were excited at 330 nm with excitation and emission slit widths of 3 nm and 4 nm, respectively. (B) tpH value was determined by fitting the curve obtained by plotting normalized fluorescence intensity at emission maximum (black) or lifetime (red) against pH. See Fig. S2† for individual curve fits. | ||
Next, we focused our attention on a biologically relevant C-rich H-Telo DNA ON repeat, which forms intramolecular iM structures in vitro. The H-Telo DNA ON repeat (C3TAA)4 under slightly acidic conditions folds into iM structures (5′E and 3′E topologies) in which the loop residues TAA show considerable differences in conformation.13c,24 Notably, the conformation of T10A11A12 residues, which form the second loop, is same in both the topologies (Fig. 1).24 While loop 2 is reasonably rigid with T10 stacked on the iM core and A11 stacked above them, loops 1 and 3 are fluxional. For these reasons, we synthesized labeled telomeric DNA ON 3 in which the loop residue T10 was replaced with benzofuran-modified nucleoside analog 1. The formation of the iM structure by C-rich H-Telo DNA ON was monitored by recording the changes in fluorescence intensity and lifetime at different pH values. As the pH was reduced from 8.2 to 5.0, DNA ON 3 displayed a significant reduction in fluorescence intensity and lifetime, which saturated at a pH nearly 5.5 (Fig. 4A). The tpH for the transition of a random coil to iM determined from steady-state (5.79 ± 0.01) and lifetime (5.80 ± 0.01) analyses is comparable to recent literature reports (tpH = 6.0–6.3, Fig. 4B. Fig. S5, Tables S2 and S3†).6c,18 CD and Tm analyses using modified (3) and control unmodified (5) ONs confirmed the formation of the iM structure at acidic pH (Fig. S4B, S4C and Table S4†). Furthermore, a significantly lower Tm exhibited by C-rich H-Telo ON 3 compared to C-rich ON 2 is consistent with the cytosine content of the sequences. Importantly, the fluorescence of free nucleoside 1 and benzofuran-labeled G-rich H-Telo DNA ON 6, which does not fold into the iM structure, was only marginally affected by changes in the pH of the medium (Fig. S6†). Collectively, these results confirm that 5-benzofuran-2′-deoxyuridine is structurally minimally invasive, and when incorporated into C-rich DNA ONs, faithfully reports the formation of iM structures with reliable tpH values.
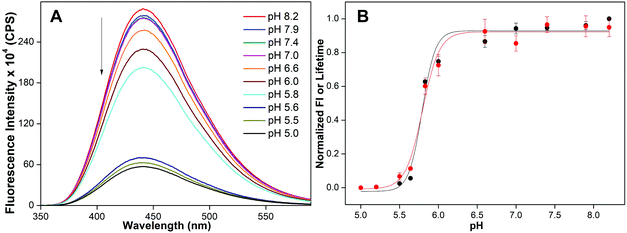 | ||
| Fig. 4 (A) Fluorescence spectra of 5-benzofuran-modified H-Telo DNA ON 3 (1 μM) at different pH values. Samples were excited at 330 nm with excitation and emission slit widths of 3 nm and 4 nm, respectively. (B) tpH value was determined by fitting the curve obtained by plotting normalized fluorescence intensity at emission maximum (black) or lifetime (red) against pH. See Fig. S5† for individual curve fits. | ||
Benzofuran-modified nucleoside distinguishes iM from random coil and duplex structures
The efficacy of emissive nucleoside 1 to distinguish the iM structure from random coil and duplex structures was evaluated by recording the fluorescence profile of single-stranded ONs (2 and 3) and corresponding duplexes at a pH above and below their respective tpH values. Random coils of ONs 2 and 3 displayed significantly higher fluorescence intensity compared to the corresponding duplexes (2·7 and 3·8) at basic pH (Fig. 5 and S7†). However, under acidic conditions (pH 5), the iM structures of DNA ONs 2 and 3 exhibited significant quenching in fluorescence intensity compared to random coil and duplex structures. The quantum yield and lifetime values were also found to be distinct for random coil, duplex and iM structures (Table 1).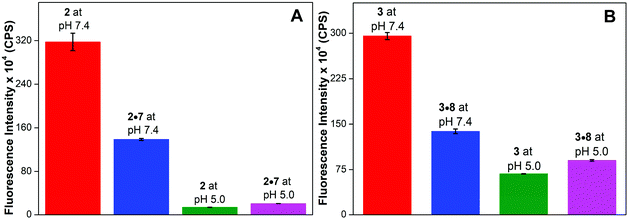 | ||
Fig. 5 Bar diagram showing the fluorescence intensity (1 μM) at λem of benzofuran-modified C-rich DNA ON 2 and H-Telo DNA ON 3 and the corresponding hybrids at basic and acidic pH. Samples of single stranded ON and the corresponding hybrids were prepared by heating the ON or a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of the respective ONs in buffers of different pH values at 90 °C for 3 min. The samples were cooled to RT slowly and incubated at RT for 1 h before analysis. All samples were excited at 330 nm. Excitation and emission slit widths were kept at 3 nm and 4 nm, respectively. See Fig. S7† for emission spectra. For λem see Table 1. 1 mixture of the respective ONs in buffers of different pH values at 90 °C for 3 min. The samples were cooled to RT slowly and incubated at RT for 1 h before analysis. All samples were excited at 330 nm. Excitation and emission slit widths were kept at 3 nm and 4 nm, respectively. See Fig. S7† for emission spectra. For λem see Table 1. | ||
| Sample | λ em (nm) | Φ | τ ave (ns) | Structural information based on fluorescence, CD and Tm |
|---|---|---|---|---|
| nd = not determined. Excited-state lifetime of the duplex could not be determined as it displayed very low fluorescence. | ||||
| 2 at pH 7.4 | 442 | 0.29 ± 0.01 | 5.32 ± 0.07 | Random coil |
| 2 at pH 5.0 | 442 | 0.015 ± 0.001 | 1.49 ± 0.03 | iM |
| 2·7 at pH 7.4 | 436 | 0.14 ± 0.002 | 2.88 ± 0.08 | Duplex (major) + iM/random coil and GQ (minor) |
| 2·7 at pH 5.0 | 436 | 0.023 ± 0.001 | 1.70 ± 0.02 | iM and GQ (major) + duplex (minor) |
| 3 at pH 7.4 | 442 | 0.28 ± 0.002 | 4.70 ± 0.04 | Random coil |
| 3 at pH 5.0 | 442 | 0.07 ± 0.01 | 3.55 ± 0.02 | iM |
| 3·8 at pH 7.4 | 434 | 0.16 ± 0.004 | 3.03 ± 0.04 | Duplex |
| 3·8 at pH 5.0 | 434 | 0.10 ± 0.001 | 2.54 ± 0.04 | Duplex (major) + iM and GQ (minor) |
| 6 at pH 7.4 | 438 | 0.13 ± 0.002 | 1.74 ± 0.03 | GQ |
| 6 at pH 5.0 | 438 | 0.14 ± 0.006 | 1.91 ± 0.04 | GQ |
| 6·5 at pH 7.4 | 436 | 0.003 ± 0.001 | nd | Duplex |
| 6·5 at pH 5.0 | 436 | 0.010 ± 0.001 | 1.57 ± 0.01 | Duplex (major) + iM and GQ (minor) |
Significantly lower fluorescence efficiency exhibited by the iM structures of C-rich ONs compared to the respective random coil and duplex structures could be due to the following reasons. While ONs 2 and 3 can potentially form different iM topologies, we preferred to provide a possible reason for the fluorescence outcome by using C-rich H-Telo DNA ON 3 as an example. The iM structure of native H-Telo DNA ON indicates that the T10 residue present in the second loop is nicely stacked between the iM core and adjacent A11 residue (Fig. S8A†).24 Hence, the benzofuran-modified nucleoside analog, which is structurally minimally perturbing, when placed at the T10 position could potentially experience a similar stacking interaction with neighbouring bases, thereby resulting in fluorescence quenching (Fig. S8B†). However, the stacking interaction between the emissive nucleoside and neighboring bases could be least in the random coil and moderate in the duplex structure. In the duplex structure, the base paired 5-benzofuran-modified nucleoside analog will be projected in the major groove and is likely to experience a partial stacking interaction, whereas in the random coil, the stacking interaction could be least as the analog is not restricted by base pairing.
It has been observed that dansyl-modified 2′-deoxycytidine and 5-(1-pyrenyl)-modified 2′-deoxyuridine exhibit dramatic quenching in the fluorescence intensity as the pH is lowered.27 The fluorescence quenching has been ascribed to an electron transfer process between the protonated pyrimidine moiety and the fluorophore. To test this possibility, a non-iM-forming control ON sequence was used, which contains the emissive nucleoside 1 flanked between the 2′-deoxycytidine residues (Fig. S9†). From basic pH to pH 6.0, there was no change in fluorescence intensity. However, as the pH of the buffer solution was lowered to 5.5 and 5.0, a noticeable decrease in fluorescence intensity was observed, which was not as dramatic as in the case of iM-forming ON sequences 2 and 3 at pH 5.5 or 5.0 (compared with Fig. 3A and 4A). At the nucleoside level, the fluorescence of 1 was not affected by changes in pH (pH 8.2–5.0, Fig. S6A†). Taken together, these results indicate that very low fluorescence exhibited by benzofuran-labeled iM-forming sequences under acidic conditions could be due to a combination of the stacking interaction with adjacent bases and the quenching effect of the C·CH+ base pair present in the neighboring environment.27,28
Duplex versus iM and GQ structures
Experimental setups based on NMR, laser tweezers, high-speed atomic force microscopy and SPR have been used to study the formation of tetraplexes in double-stranded DNA.29 These studies indicate that helical loosening, offset arrangements of G-rich and C-rich strands and inherent stability of the structures can influence the relative population of duplex and tetraplex forms. In another study, 2-aminopurine has been used to monitor the formation of GQ and iM structures in retinoblastoma susceptibility genes.30 While the fluorescence intensity of GQ and iM forms was significantly high compared to that of the duplex, the fluorescence of GQ and iM structures was similar. Since benzofuran-modified nucleoside analog 1 incorporated into G-rich and C-rich ONs shows distinct fluorescence properties for GQ, iM and duplex structures, we sought to understand the relative population of these structures at physiological and acidic pH.A solution of 2·7 at physiological pH displayed a strong emission band corresponding to a quantum yield of 0.14 (Fig. 5A, Table 1). At pH 5.0, it showed a significantly lower fluorescence intensity (Φ = 0.023), which was slightly higher than the iM form of ON 2 (Φ = 0.015). The CD profile of 2·7 at pH 7.4 displayed dominant peaks characteristic of a duplex structure (positive ∼260 nm and negative ∼237 nm) along with a shoulder near 285 nm (Fig. 6). The tpH of ON 2 is ∼7.1, and, hence, it is likely that a solution of 2·7 at pH 7.4 could potentially have a small population of the random coil/iM and GQ structures of 2 and 7, respectively, which is reflected in the form of a shoulder in the CD profile.29e,31 However, at pH 5.0, a solution of 2·7 revealed a CD profile mainly emanating from a combination of the iM and GQ forms of 2 and 7, respectively. It is important to mention here that the GQ structure of ON 7 is not affected by changes in pH (Fig. 6 and Table S5†). Hence, at acidic pH, a slightly higher fluorescence exhibited by a solution of 2·7 is due to the presence of a very large population of the weakly emissive iM form of ON 2 (along with the unmodified GQ of 7) and a small population of a more emissive duplex form. This notion is further supported by lifetime studies. A solution of 2·7 at pH 7.4, resembling mostly a duplex form, shows a lifetime of 2.88 ns (Table 1). At pH 5.0, it shows a lifetime closer to the iM form of 2, suggesting that a solution of 2·7 at acidic pH predominantly exists as iM and GQ structures. UV-thermal melting studies also corroborate the above results as the duplex structure of 2·7 at pH 7.4 and the iM structure of ON 2 at pH 5.0 display high and comparable Tm values (Table S4† and S5†). Based on the above information, we could make an approximate estimate of the competition between the duplex and tetraplex structures. A comparison of the quantum yield of 2·7 at pH 7.4 (predominantly duplex form), 2·7 at pH 5.0 (predominantly iM and GQ forms) and 2 at pH 5.0 (completely iM form) suggested that an acidic solution of 2·7 is composed of nearly 94% tetraplexes (iM and GQ forms) and 6% duplex (Table 1).
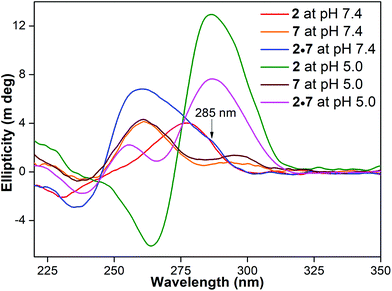 | ||
Fig. 6 CD spectra (5 μM) of C-rich ON 2, complementary G-rich ON 7 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 solution of 2 and 7 at pH 7.4 and 5.0. ON 2 is a random coil (red) at pH 7.4 and iM (green) at pH 5.0. CD profile of ON 7 at both the pH values shows a similar pattern (orange and brown) resembling a parallel GQ structure.15e A solution of 2·7 at pH 7.4 displays a typical duplex CD profile (blue) along with a shoulder near 285 nm, potentially arising from alternative structures, namely iM/random coil and GQ forms. At acidic pH, a CD profile (magenta) mainly resembling a combination of the iM and GQ forms of 2 and 7, respectively, is seen. 1 solution of 2 and 7 at pH 7.4 and 5.0. ON 2 is a random coil (red) at pH 7.4 and iM (green) at pH 5.0. CD profile of ON 7 at both the pH values shows a similar pattern (orange and brown) resembling a parallel GQ structure.15e A solution of 2·7 at pH 7.4 displays a typical duplex CD profile (blue) along with a shoulder near 285 nm, potentially arising from alternative structures, namely iM/random coil and GQ forms. At acidic pH, a CD profile (magenta) mainly resembling a combination of the iM and GQ forms of 2 and 7, respectively, is seen. | ||
A hybrid of benzofuran-labeled C-rich H-Telo and unmodified G-rich H-Telo DNA ONs 3 and 8, respectively, at pH 7.4 displayed a relatively higher fluorescence intensity (Φ = 0.16) compared to that under acidic conditions (Φ = 0.10, Fig. 5B, Table 1). Unlike 3·8, which exhibits multiple structures at pH 7.4 (vide supra, duplex form, major), the CD spectrum of a solution of 3·8 matched with the duplex structure (positive peak ∼265 nm and negative peak ∼240 nm, Fig. 7). This is because the tpH of ON 3 (5.8) is much lower than that of ON 2 (∼7.1) to facilitate the formation of an iM structure at physiological pH. Hence, the fluorescence intensity exhibited by 3·8 at pH 7.4 can be considered solely due to the duplex structure. However, the fluorescence of a solution of 3·8 at acidic pH was found to be discernibly higher (Φ = 0.10) compared to the iM form of ON 3 (Φ = 0.07, Fig. 5B, Table 1). The CD spectrum of 3·8 largely resembled a duplex structure along with a visible shoulder near 285 nm (Fig. 7). The shoulder band suggests the existence of alternative structures, namely iM and GQ forms.29e,31 This notion is supported by the fact that H-Telo DNA ON repeats 3 and 8 form stable iM and GQ structures, respectively, at acidic pH (Fig. 7, Tables S4 and S5†). However, relatively higher stability of duplex over iM and GQ structures suggests that the observed fluorescence of 3·8 under acidic conditions is due to a combination of duplex form (major) and iM-GQ structures (minor, Tables S4 and S5†). A time-resolved experiment using 3·8 at pH 5.0 gave a lifetime, which is closer to that of the duplex form, suggesting that this solution at acidic pH is mostly made of the duplex structure (Table 1).
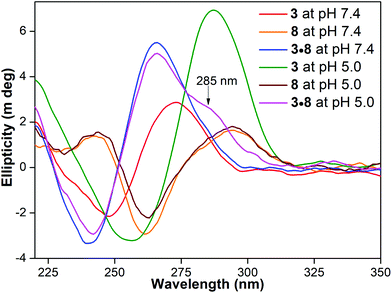 | ||
Fig. 7 CD spectra (5 μM) of H-Telo C-rich ON 3, complementary G-rich ON 8 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 solution of 3 and 8 at pH 7.4 and 5.0. ON 3 is a random coil (red) at pH 7.4 and iM (green) at pH 5.0. CD profile of ON 8 at both the pH values shows a similar pattern (orange and brown) resembling an antiparallel GQ structure.15c,22a A solution of 3·8 at pH 7.4 displays a profile corresponding to the duplex structure (blue). However, at acidic pH a duplex CD profile along with a shoulder near 285 nm is observed (magenta), potentially arising from alternative structures, namely iM and GQ forms. 1 solution of 3 and 8 at pH 7.4 and 5.0. ON 3 is a random coil (red) at pH 7.4 and iM (green) at pH 5.0. CD profile of ON 8 at both the pH values shows a similar pattern (orange and brown) resembling an antiparallel GQ structure.15c,22a A solution of 3·8 at pH 7.4 displays a profile corresponding to the duplex structure (blue). However, at acidic pH a duplex CD profile along with a shoulder near 285 nm is observed (magenta), potentially arising from alternative structures, namely iM and GQ forms. | ||
Next, a hybrid made of benzofuran-labeled G-rich H-Telo DNA ON 6 and unmodified C-rich H-Telo DNA ON 5 was subjected to fluorescence, CD and Tm measurements. In our earlier study, we have demonstrated that 5-benzofuran-modified nucleoside 1 incorporated into the G-rich strand of H-Telo DNA serves as a useful GQ sensor, wherein the GQ structure displays significantly higher fluorescence compared to its duplex at neutral pH.22 In the present study also, the GQ structure of 6 showed significantly higher fluorescence (Φ = 0.13) compared to that of duplex 6·5 at pH 7.4 (Φ = 0.003, Table 1, Fig. 8A). Interestingly, a solution of 6·5 at acidic pH exhibited a noticeable increase in fluorescence (Φ = 0.010) compared to that at pH 7.4. This increase in fluorescence is likely due to the dissociation of a small amount of the duplex, which leads to the formation of a highly emissive GQ structure of ON 6. The formation of the GQ structure, in a way, is also assisted by the formation of a stable iM structure by the complementary ON strand 5 at acidic pH (vide supra, Fig. S4†). These observations are supported by CD and Tm studies. CD and Tm measurements indicate that 6·5 forms only a duplex structure under physiological conditions (Fig. 8B and Table S5†). However, at pH 5.0, an additional shoulder band near 285 nm indicates the presence of small amounts of alternative forms, namely GQ and iM structures.29e,31
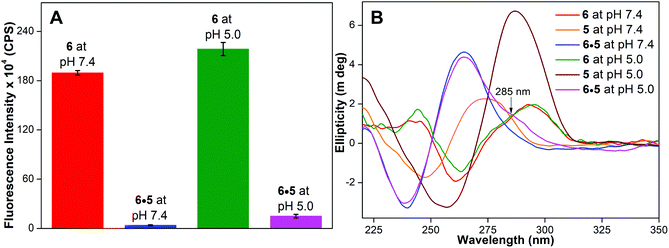 | ||
Fig. 8 (A) Bar diagram showing the fluorescence intensity (1 μM) at λem of benzofuran-modified H-Telo G-rich DNA ON 6 and corresponding hybrid with complementary ON 5 at pH 7.4 and 5.0. Samples were excited at 330 nm, and excitation and emission slit widths were kept at 4 nm and 5 nm, respectively. (B) CD spectra (5 μM) of H-Telo G-rich ON 6, complementary C-rich ON 5 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 solution of 6 and 5 at pH 7.4 and 5.0. CD profile of ON 6 at both the pH values shows a similar pattern (red and green) resembling an antiparallel GQ structure.22b ON 5 is a random coil (orange) at pH 7.4 and iM (brown) at pH 5.0. A solution of 6·5 at pH 7.4 displays a profile corresponding to a duplex structure (blue). However, at acidic pH a duplex CD profile along with a shoulder near 285 nm is observed (magenta), potentially arising from alternative structures, namely iM and GQ forms. 1 solution of 6 and 5 at pH 7.4 and 5.0. CD profile of ON 6 at both the pH values shows a similar pattern (red and green) resembling an antiparallel GQ structure.22b ON 5 is a random coil (orange) at pH 7.4 and iM (brown) at pH 5.0. A solution of 6·5 at pH 7.4 displays a profile corresponding to a duplex structure (blue). However, at acidic pH a duplex CD profile along with a shoulder near 285 nm is observed (magenta), potentially arising from alternative structures, namely iM and GQ forms. | ||
Since the duplex form of the telomeric repeats (6·5) is very weakly fluorescent, the observed fluorescence of a solution of 6·5 under acidic conditions is more or less due to the free GQ form of 6. A comparison of the quantum yield of 6 at pH 5.0 (GQ form), 6·5 at pH 5.0 (predominantly duplex form + iM-GQ forms) and 6·5 at pH 7.4 (completely duplex form) suggests that an acidic solution of 6·5 is composed of nearly 95% duplex form and 5% GQ and iM structures (Table 1). These conclusions corroborate with the results obtained using benzofuran-labeled C-rich H-Telo DNA ON 3, which further substantiates that at physiological pH the double-stranded telomeric region is duplex in nature, whereas under acidic conditions along with the duplex form there exists a small population of GQ and iM structures.
Based on the fluorescence, CD and thermal melting experiments, we could perform a qualitative analysis of the relative population of duplex, iM and GQ forms in G-rich and C-rich double-stranded systems at different pH values. The model system 2·7, containing more C and G residues, at physiological pH is largely made of the duplex form along with a small fraction of iM/random coil and GQ structures. However, under acidic conditions, iM and GQ structures dominate the overall population with the duplex form being the minor component. In the case of double-stranded telomeric repeats (3·8 or 6·5), only the duplex form exists at physiological pH. Albeit in small amounts, the telomeric repeats have the tendency to form stable iM and GQ structures at acidic pH.
Conclusions
We have devised a simple fluorescence-based platform to monitor the formation of the iM structures of C-rich DNA ONs by using the conformation sensitivity and minimally perturbing nature of the benzofuran-modified 2′-deoxyuridine analog. In addition to serving as a useful topology-specific GQ sensor, the nucleoside analog, incorporated into C-rich DNA ONs, photophysically shows the pH-induced transition of a single-stranded structure to an iM structure and provides reliable tpH values. Importantly, this probe exhibits distinct fluorescence properties for iM, GQ, random coil and duplex structures. This feature of the probe, when combined with CD and thermal melting studies, provided information on the propensity of a given G-rich−C-rich double-stranded system to adopt a duplex form or iM and GQ forms or a combination of these structures. Our results indicate that the formation of iM and GQ structures by G-rich−C-rich double-stranded systems under physiological conditions will depend on the relative stability of the duplex, iM and GQ forms. For example, the double-stranded region of the human telomere is significantly more stable such that the formation of iM and GQ structures seems difficult unless assisted by protein factors or small molecule binders, which induce or stabilize such non-canonical structures.3c,32 A similar scenario can also be envisioned for double-stranded promoter regions capable of supporting GQ/iM structures.33 Taken together, our GQ-iM sensor will be a very important addition to the resources and tools available for profiling non-canonical four-stranded structures in the genome. It is expected that this nucleoside probe could also be useful in setting up discovery approaches to identify efficient GQ and iM binders.Experimental section
Solid-phase synthesis of 5-benzofuran-modified DNA ONs
Benzofuran-modified DNA ONs 2, 3 and 6 were synthesized by following our earlier reports.22 All ONs were purified by polyacrylamide gel electrophoresis (PAGE) and analyzed by HPLC (Fig. S1†). The integrity of modified ONs was confirmed by MALDI-TOF mass analysis. See Table S1† for ε260 and MALDI-TOF mass data of the modified ONs.Photophysical analysis of fluorescently-modified C-rich (2, 3) and G-rich (6) DNA ONs
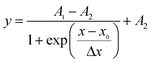 | (1) |
| Φ(x) = (As/Ax)(Fx/Fs)(nx/ns)2Φ(s) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of the respective ONs in different buffers at 90 °C for 3 min. The samples were cooled to RT and incubated at RT for 1 h before CD analysis. CD spectra were collected from 350 to 220 nm on a JASCO J-815 CD spectrometer using 1 nm bandwidth at 20 °C. Experiments were performed in duplicate wherein each spectrum was an average of five scans. The spectrum of buffer was subtracted from all sample spectra.
1 mixture of the respective ONs in different buffers at 90 °C for 3 min. The samples were cooled to RT and incubated at RT for 1 h before CD analysis. CD spectra were collected from 350 to 220 nm on a JASCO J-815 CD spectrometer using 1 nm bandwidth at 20 °C. Experiments were performed in duplicate wherein each spectrum was an average of five scans. The spectrum of buffer was subtracted from all sample spectra.
Conflicts of interest
There are no conflicts to declare.Acknowledgements
P. M. S. and A. A. T. thank the CSIR, India for a graduate research fellowship. S. G. S. thanks the Wellcome Trust-DBT India Alliance (IA/S/16/1/502360) and DST, India (EMR/2014/000419) for research grants.Notes and references
- T. A. Brooks, S. Kendrick and L. Hurley, FEBS J., 2010, 277, 3459–3469 CrossRef PubMed
.
- G. W. Collie and G. N. Parkinson, Chem. Soc. Rev., 2011, 40, 5867–5892 RSC
.
-
(a) V. K. Yadav, J. K. Abraham, P. Mani, R. Kulshrestha and S. Chowdhury, Nucleic Acids Res., 2008, 36, D381–D385 CrossRef PubMed
; (b) D. Sun and L. H. Hurley, J. Med. Chem., 2009, 52, 2863–2874 CrossRef PubMed
; (c) S. Balasubramanian, L. H. Hurley and S. Neidle, Nat. Rev. Drug Discovery, 2011, 10, 261–275 CrossRef PubMed
.
-
(a) M. Guéron and J.-L. Leroy, Curr. Opin. Struct. Biol., 2000, 10, 326–331 CrossRef
; (b) S. Dzatko, M. Krafcikova, R. Hänsel-Hertsch, T. Fessl, R. Fiala, T. Loja, D. Krafcik, J.-L. Mergny, S. Foldynova-Trantirkova and L. Trantirek, Angew. Chem., Int. Ed., 2018, 57, 2165–2169 CrossRef PubMed
.
-
(a) Y. Xu, Y. Suzuki, K. Ito and M. Komiyama, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 14579–14584 CrossRef PubMed
; (b) E. Y. N. Lam, D. Beraldi, D. Tannahill and S. Balasubramanian, Nat. Commun., 2013, 4, 1796 CrossRef PubMed
; (c) A. Henderson, Y. Wu, Y. C. Huang, E. A. Chavez, J. Platt, F. B. Johnson Jr., R. M. Brosh, D. Sen and P. M. Lansdrop, Nucleic Acids Res., 2014, 42, 860–869 CrossRef PubMed
; (d) A. Laguerre, K. Hukezalie, P. Winckler, F. Katranji, G. Chanteloup, M. Pirrotta, J.-M. Perrier-Cornet, J. M. Y. Wong and D. Monchaud, J. Am. Chem. Soc., 2015, 137, 8521–8525 CrossRef PubMed
; (e) H.-Y. Liu, Q. Zhao, T.-P. Zhang, Y. Wu, Y.-X. Xiong, S.-K. Wang, Y.-L. Ge, J.-H. He, P. Lv, T.-M. Ou, J.-H. Tan, D. Li, L.-Q. Gu, J. Ren, Y. Zhao and Z.-S. Huang, Cell Chem. Biol., 2016, 23, 1261–1270 CrossRef PubMed
; (f) H.-L. Bao, T. Ishizuka, T. Sakamoto, K. Fujimoto, T. Uechi, N. Kenmochi and Y. Xu, Nucleic Acids Res., 2017, 45, 5501–5511 CrossRef PubMed
.
-
(a) A. Rajendran, S. Nakano and N. Sugimoto, Chem. Commun., 2010, 46, 1299–1301 RSC
; (b) J. A. Brazier, A. Shah and G. D. Brown, Chem. Commun., 2012, 48, 10739–10741 RSC
; (c) L. Lannes, S. Halder, Y. Krishnan and H. Schwalbe, ChemBioChem, 2015, 16, 1647–1656 CrossRef PubMed
; (d) E. P. Wright, H. A. Day, A. M. Ibrahim, J. Kumar, L. J. E. Boswell, C. Huguin, C. E. M. Stevenson, K. Pors and Z. A. E. Waller, Sci. Rep., 2016, 6, 39456, DOI:10.1038/srep39456
.
-
(a) L. Lacroix, H. Liénard, E. Labourier, M. Djavaheri-Mergny, J. Lacoste, H. Leffers, J. Tazi, C. Hélène and J.-L. Mergny, Nucleic Acids Res., 2000, 28, 1564–1575 CrossRef PubMed
; (b) D. J. Uribe, K. Guo, Y.-J. Shin and D. Sun, Biochemistry, 2011, 50, 3796–3806 CrossRef PubMed
; (c) Y. M. K. Yoga, D. A. K. Traore, M. Sidiqi, C. Szeto, N. R. Pendini, A. Barker, P. J. Leedman, J. A. Wilce and M. C. J. Wilce, Nucleic Acids Res., 2012, 40, 5101–5114 CrossRef PubMed
.
- S. Kendrick, H.-J. Kang, M. P. Alam, M. M. Madathil, P. Agrawal, V. Gokhale, D. Yang, S. M. Hecht and L. H. Hurley, J. Am. Chem. Soc., 2014, 136, 4161–4171 CrossRef PubMed
.
-
(a) Y. Chen, K. Qu, C. Zhao, L. Wu, J. Ren, J. Wang and X. Qu, Nat. Commun., 2012, 3, 1074, DOI:10.1038/ncomms2091
; (b) H.-J. Kang, S. Kendrick, S. M. Hecht and L. H. Hurley, J. Am. Chem. Soc., 2014, 136, 4172–4185 CrossRef PubMed
; (c) H. A. Day, P. Pavlou and Z. A. E. Waller, Bioorg. Med. Chem., 2014, 22, 4407–4418 CrossRef PubMed
.
-
(a) Y. Krishnan and F. C. Simmel, Angew. Chem., Int. Ed., 2011, 50, 3124–3156 CrossRef PubMed
; (b) S. Modi, C. Nizak, S. Surana, S. Halder and Y. Krishnan, Nat. Nanotechnol., 2013, 8, 459–467 CrossRef PubMed
; (c) L. Tao and M. Famulok, J. Am. Chem. Soc., 2013, 135, 1593–1599 CrossRef PubMed
; (d) Y. Dong, Z. Yang and D. Liu, Acc. Chem. Res., 2014, 47, 1853–1860 CrossRef PubMed
.
- A. M. Fleming, Y. Ding, R. A. Rogers, J. Zhu, J. Zhu, A. D. Burton, C. B. Carlisle and C. J. Burrows, J. Am. Chem. Soc., 2017, 139, 4682–4689 CrossRef PubMed
.
- T. A. Brooks, S. Kendrick and L. Hurley, FEBS J., 2010, 277, 3459–3469 CrossRef PubMed
.
-
(a) K. Gehring, J.-L. Leroy and M. Guéron, Nature, 1993, 363, 561–565 CrossRef PubMed
; (b) M. Adrian, B. Heddi and A. T. Phan, Methods, 2012, 57, 11–14 CrossRef PubMed
; (c) A. L. Lieblein, J. Buck, K. Schlepckow, B. Furtig and H. Schwalbe, Angew. Chem., Int. Ed., 2012, 51, 250–253 CrossRef PubMed
; (d) M. Vorlíčková, I. Kejnovská, J. Sagi, D. Renčiuk, K. Bednářová, J. Motlová and J. Kypr, Methods, 2012, 57, 64–75 CrossRef PubMed
; (e) B. R. Vummidi, J. Alzeer and N. W. Luedtke, ChemBioChem, 2013, 14, 540–558 CrossRef PubMed
; (f) S. Neidle, J. Med. Chem., 2016, 59, 5987–6011 CrossRef PubMed
; (g) A. Dembska, P. Bielecka and B. Juskowiak, Anal. Methods, 2017, 9, 6092–6106 RSC
.
-
(a) E. Largy, A. Granzhan, F. Hamon, D. Verga and M.-P. Teulade-Fichou, Top. Curr. Chem., 2013, 330, 111–177 CrossRef PubMed
; (b) A. D. Rache and J.-L. Mergny, Biochimie, 2015, 115, 194–202 CrossRef PubMed
; (c) I. J. Lee, S. P. Patil, K. Fhayli, S. Alsaiari and N. M. Khashab, Chem. Commun., 2015, 51, 3747–3749 RSC
.
-
(a) T. Kimura, K. Kawai, M. Fujitsuka and T. Majima, Chem. Commun., 2006, 401–402 RSC
; (b) R. D. Gray, L. Petraccone, J. O. Trent and J. B. Chaires, Biochemistry, 2010, 49, 179–194 CrossRef PubMed
; (c) A. Nadler, J. Strohmeier and U. Diederichsen, Angew. Chem., Int. Ed., 2011, 50, 5392–5396 CrossRef PubMed
; (d) A. Dumas and N. W. Luedtke, Nucleic Acids Res., 2011, 39, 6825–6834 CrossRef PubMed
; (e) J. Johnson, R. Okyere, A. Joseph, K. Musier-Forsyth and B. Kankia, Nucleic Acids Res., 2013, 41, 220–228 CrossRef PubMed
; (f) M. Sproviero, K. L. Fadock, A. A. Witham and R. A. Manderville, ACS Chem. Biol., 2015, 10, 1311–1318 CrossRef PubMed
.
-
(a) J. Lee II and B. H. Kim, Chem. Commun., 2012, 48, 2074–2076 RSC
; (b) J. W. Park, Y. J. Seo and B. H. Kim, Chem. Commun., 2014, 50, 52–54 RSC
.
- J. Choi, A. Tanaka, D. W. Cho, M. Fujitsuka and T. Majima, Angew. Chem., Int. Ed., 2013, 52, 12937–12941 CrossRef PubMed
.
- G. Mata and N. W. Luedtke, J. Am. Chem. Soc., 2015, 137, 699–707 CrossRef PubMed
.
- J. Zhou, S. Amrane, D. N. Korkut, A. Bourdoncle, H.-Z. He, D.-L. Ma and J.-L. Mergny, Angew. Chem., Int. Ed., 2013, 52, 7742–7746 CrossRef PubMed
.
- A. A. Tanpure and S. G. Srivatsan, Chem. – Eur. J., 2011, 17, 12820–12827 CrossRef PubMed
.
- A. A. Tanpure and S. G. Srivatsan, ChemBioChem, 2012, 13, 2392–2399 CrossRef PubMed
.
-
(a) A. A. Tanpure and S. G. Srivatsan, Nucleic Acids Res., 2015, 43, e149 CrossRef PubMed
; (b) S. Manna, C. H. Panse, V. A. Sontakke, S. Sangamesh and S. G. Srivatsan, ChemBioChem, 2017, 18, 1604–1615 CrossRef PubMed
.
-
(a) T. Kanamori, H. Ohzeki, Y. Masaki, A. Ohkubo, M. Takahashi, K. Tsuda, T. Ito, M. Shirouzu, K. Kuwasako, Y. Muto, M. Sekine and K. Seio, ChemBioChem, 2015, 16, 167–176 CrossRef PubMed
; (b) C. S. Eubanks, J. E. Forte, G. J. Kapral and A. E. Hargrove, J. Am. Chem. Soc., 2017, 139, 409–416 CrossRef PubMed
; (c) C. S. Eubanks and A. E. Hargrove, Chem. Commun., 2017, 53, 13363–13366 RSC
.
- A. T. Phan, M. Guéron and J.-L. Leroy, J. Mol. Biol., 2000, 299, 123–144 CrossRef PubMed
.
- S. Chakraborty and Y. Krishnan, Biochimie, 2008, 90, 1088–1095 CrossRef PubMed
.
- S. Kendrick, Y. Akiyama, S. M. Hecht and L. H. Hurley, J. Am. Chem. Soc., 2009, 131, 17667–17676 CrossRef PubMed
.
-
(a) N. Amann, E. Pandurski, R. Fiebig and H.-A. Wagenknecht, Angew. Chem., Int. Ed., 2002, 41, 2978–2980 CrossRef
; (b) K. Kim, H. W. Kim, D. Moon, Y. M. Rhee and B. H. Kim, Org. Biomol. Chem., 2013, 11, 5605–5614 RSC
.
- J. Lee II, J. W. Yi and B. H. Kim, Chem. Commun., 2009, 5383–5385 RSC
.
-
(a) K. Halder and S. Chowdhury, Nucleic Acids Res., 2005, 33, 4466–4474 CrossRef PubMed
; (b) M. Endo, X. Xing, X. Zhou, T. Emura, K. Hidaka, B. Tuesuwan and H. Sugiyama, ACS Nano, 2015, 9, 9922–9929 CrossRef PubMed
; (c) Y. Cui, D. Kong, C. Ghimire, C. Xu and H. Mao, Biochemistry, 2016, 55, 2291–2299 CrossRef PubMed
; (d) S. Selvam, S. Mandal and H. Mao, Biochemistry, 2017, 56, 4616–4625 CrossRef PubMed
; (e) H. A. Assi, R. El-Khoury, C. Gonźalez and M. J. Damha, Nucleic Acids Res., 2017, 45, 11535–11546 CrossRef PubMed
.
- Y. Xu and H. Sugiyama, Nucleic Acids Res., 2006, 34, 949–954 CrossRef PubMed
.
- W. Li, P. Wu, T. Ohmichi and N. Sugimoto, FEBS Lett., 2002, 526, 77–81 CrossRef PubMed
.
-
(a) D. Broccoli, A. Smogorzewska, L. Chong and T. de Lange, Nat. Genet., 1997, 17, 231–235 CrossRef PubMed
; (b) I. M. Pedroso, W. Hayward and T. M. Fletcher, Nucleic Acids Res., 2009, 37, 1541–1554 CrossRef PubMed
; (c) G. Biffi, D. Tannahill and S. Balasubramanian, J. Am. Chem. Soc., 2012, 134, 11974–11976 CrossRef PubMed
.
- A. Arora, D. R. Nair and S. Maiti, FEBS J., 2009, 276, 3628–3640 CrossRef PubMed
.
- M. G. Pawar and S. G. Srivatsan, J. Phys. Chem. B, 2013, 117, 14273–14282 CrossRef PubMed
.
Footnotes |
| † Electronic supplementary information (ESI) available: Supplementary figures, tables and experimental procedures. See DOI: 10.1039/c8ob00646f |
| ‡ These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2018 |

